Abstract
Background/Objectives: Gamma-aminobutyric acid (GABA) is a bioactive, non-proteinaceous amino acid with potential health benefits. Weissella cibaria UF-274 is an important lactic acid bacterium isolated from Balinese fermented sausage (urutan) with GABA-producing abilities. The aim of this study was to enhance GABA synthesis in skim milk as a basal substrate, as well as whole genome sequencing and analysis to evaluate the functionality and safety of the strain. Methods: A Box–Behnken response surface design was used to enhance GABA accumulation in skim milk. Results: The optimum conditions for GABA production were at concentrations of glucose of 23.91 g/L, monosodium glutamate concentrations of 2.32 g/L and pyridoxal-5′-phosphate at 46 μM. The genome assembly produced a high-quality draft with a 2.53 Mb circular chromosome and 2378 coding sequences. A whole genome analysis revealed that the strain possesses a glutamine amidotransferase (puuD-like) as an alternative route linked to the GABA pathway. AntiSMASH prediction results showed that the strain has two biosynthetic gene clusters including terpene and type III polyketide synthases. Several bioinformatic approaches predicted no antibiotic resistance genes, while van genes encoding vancomycin resistance were detected with low pathogen risk with one approach. Conclusions: Weissella cibaria UF-274 is a promising GABA producer with genomic evidence and a good candidate for functional food development.
1. Introduction
Gamma-aminobutyric acid (GABA) is recognized as a primary inhibitory neurotransmitter within the mammalian central nervous system. In plants and microorganisms, GABA functions as a defense mechanism, being excreted under stress conditions [1]. This non-proteinaceous amino acid exhibits well-documented physiological effects in humans, including the modulation of neuronal excitability and potential benefits for blood pressure regulation, obesity management, and stress reduction [2,3,4]. GABA biosynthesis primarily occurs through the decarboxylation of L-glutamate, a reaction catalyzed by L-glutamic acid decarboxylase (GAD; EC 4.1.1.15), an enzyme encoded by the gadA or gadB genes [5]. This enzymatic reaction necessitates a cofactor, pyridoxal-5′-phosphate, to initiate the decarboxylation. Following synthesis, the gadC-encoded glutamate/GABA antiporter is responsible for secreting GABA from the cell [6,7]. Nevertheless, certain microbial species, including Escherichia coli and Aspergillus oryzae, utilize an alternative glutamine-putrescine pathway involving a series of enzymes to convert arginine, ornithine, and agmatine into GABA [8,9]. The specific metabolic pathways employed for GABA synthesis vary across different organisms, with bacteria often utilizing distinct enzymatic cascades compared to fungi or plants.
Food-based delivery of GABA has attracted increasing interest, as fermentation by lactic acid bacteria (LAB) offers a pathway to naturally enriched functional foods without the need for chemical additives [10]. In contrast, chemical synthesis can lead to unwanted by-products. LAB are recognized as efficient cell factories for GABA production and generally hold the Generally Recognized As Safe (GRAS) status. Several GABA-producing LAB species and strains, including Lactiplantibacillus plantarum, Lactobacillus brevis, Lactobacillus paracasei, Lactobacillus pentosus, and Lactococcus lactis, have been identified [11,12,13,14,15]. Traditional fermented foods serve as a rich reservoir of unique LAB species and strains possessing adaptive traits. Several GABA-producing LAB have been isolated from various fermented foods like kimchi [16,17], fermented fish [18,19], fermented soybean [20,21], fermented sausages [22], and fermented milk [23]. Among these, LAB species/strains showed variability in GAD yield and characteristic in food matrix applications, suggesting a highly strain-dependent flexibility for food applications.
In this study, the strain Weissella cibaria UF-274 was isolated from Balinese fermented sausage (urutan) and shown to possess GABA-producing ability. A niche artisanal matrix often selects for strains with distinctive metabolic and stress-resistance characteristics relevant to food fermentation. However, the utilization of these strains for other food products necessitates applied optimization within a target matrix. Skim milk, for instance, offers a commercially relevant, and cheap substrate where protein and mineral composition can significantly influence microbial metabolism [24,25,26]. While W. cibaria has indeed emerged as a novel GABA-producing LAB, its promising potential as well as its full ability for GABA production has not yet been thoroughly investigated, particularly within different food systems. This gap in knowledge suggests a novel avenue for further investigation. The synthesis of this metabolite is primarily affected by fermentation parameters such as the supplementation of carbon sources, nitrogen sources and cofactors, as well as by the incubation temperature and incubation time. Furthermore, the specific supplementation of various nutrients in the medium of a food matrix can significantly influence GABA yield. Consequently, it is important to investigate the optimum nutrient conditions for GABA production in W. cibaria UF-274.
Whole genome sequencing (WGS) is crucial for empirically optimizing GABA production in W. cibaria UF-274. It enables the detection of GABA synthesis pathways, comprehensive genomic characterization, and the assessment of functional characteristics like secondary metabolite production [27,28]. Furthermore, WGS facilitates the prediction of antimicrobial resistance or virulence determinants, which are essential for compliance with food safety regulations [29,30]. In this preliminary study, we evaluate the potential of this strain as a starter or adjunct starter in skim milk as a basal substrate in addition to WGS-based analysis to maximize GABA yield under food-relevant conditions and establish genomic evidence for biosynthetic potential and strain safety.
2. Materials and Methods
2.1. Taxonomy Profiles in Fermented Urutan
Fermented urutan was obtained from a local household located in Baturity, Tabanan regency, Bali, Indonesia. The fermented sausages were vacuum packed and stored at 4 °C prior to metagenomic analysis. Analysis of the bacterial composition in fermented urutan was conducted using the genomic extraction kit DNA Miniprep (Zymo Research, D4300, Irvine, CA, USA) according to the manufacturer instructions and subsequent sequencing. The concentration of the extracted DNA was measured using a NanoDrop spectrophotometer (Thermo Fisher Scientific, Waltham, MA, USA), and it was stored at –20 °C until further analysis. For identification at the species level, long-read sequencing targeting 16S rRNA was performed. The gene was amplified using the forward primer 27F: 5′—AGAGTTTGATCMTGGCTCAG—3′ and the reverse primer 1492R: 5′—GGTTACCTTGTTACGACTT—3′. Subsequently, barcoding the PCR process was done using the PCR barcoding expansion kit 1-96 (Nanopore, Oxford, UK), followed by purification of the PCR products. An adapter-binding PCR was performed by ligation using a Ligation Sequencing Kit (Nanopore, Oxford, UK). The purified product was loaded into the Flow Cell MinIon (Nanopore) device operated by MinKNOW software version 22.05.7 (Nanopore) [31].
2.2. Isolation, Screening and Identification of GABA-Producing LAB
Ten grams of fermented urutan were suspended in 90 mL of sterile 0.85% NaCl solution and homogenized. This was followed by serial 10-fold dilutions and spreading on de Man Rogosa and Sharpe (MRS) agar supplemented with 0.5% (w/v) monosodium glutamate (MSG, Ajinomoto, Indonesia) and bromocresol purple (BCP). The plates were incubated at 37 °C for 24–48 h. LAB isolates with GABA-forming potential were selected based on the appearance of a yellow-clear zone around the colonies. The selected isolates were streaked on the same medium to obtain pure colonies. The purified colonies were stored at −80 °C in a 30% glycerol solution for further analysis.
To determine GABA production, each LAB isolate was inoculated in MRS broth medium containing 0.5% MSG and incubated at 37 °C for 24 h. The cultured MRS medium was then centrifuged at 8000× g for 5 min at 4 °C, and 1–2 μL of the supernatants was spotted onto silica plates 60 F254 (Merck, Darmstadt, Germany). The plates were placed in a chamber filled with the mobile phase containing a mixture of 1-butanol:acetic acid:aquadest (5:2:2). After separation, spots were visualized with 0.5% (w/v) ninhydrin and heated at 100 °C for 5 min. LAB with GABA-producing ability were further identified, and the amount of GABA was quantified using HPLC [32].
DNA barcoding of 16S rRNA was performed to identify GABA-producing LAB using the GeneJET DNA kit (Thermo Fisher, MA, USA). The isolated genomic was amplified using universal primers 27F: 5′—AGAGTTTGATCMTGGCTCAG—3′ and reverse primer 1492R: 5′—GGTTACCTTGTTACGACTT—3′. The partial sequences of 16S rRNA of each LAB were compared with the GenBank (NCBI) database, followed by the construction of a phylogenetic tree using MEGA software (v. 4).
2.3. Effect of Nutrients on GABA Production
Skim milk was sterilized and inoculated with 0.5% (v/v) of an active culture of W. cibaria UF-274. The skim milk was fermented at 37 °C for 24 h. Based on preliminary studies, glucose and monosodium glutamate (MSG) were selected as carbon and nitrogen sources. The effects of glucose concentration, MSG concentration and pyridoxal-5′-phosphate (PLP) concentration on GABA production in skim milk were determined. The experimental range of the three factors studied was as follows: glucose concentrations were set up at 0–40 g/L, MSG concentrations at 0–3 g/L, and PLP concentrations were at 0–60 μM. All experiments were performed in triplicate.
To obtain optimum conditions for GABA production, a Box–Behnken test design with three factors was set up using Stat Ease 360 v25. 0.1 (Stat Ease Inc., Minneapolis, MN, USA). The results were analyzed using ANOVA and response surface plots. Optimal conditions for GABA formation were determined by analyzing response surface plots from the response surface model (RSM).
2.4. GABA Quantification
GABA concentration in the supernatant was quantified using High Pressure Liquid Chromatography (HPLC, Thermo Dionex Ultimate 3000, Thermo Fisher, MA, USA). Fifty μL of supernatant was derivatized using 300 μL of an o-phthalaldehyde (OPA) solution (containing 1 mL methanol, 4 mL borate buffer pH 9.0 and 30 μL 2-mercaptoethanol). The derivatized samples (20 μL) were injected into the HPLC with a mobile phase consisting of gradient A (CH3OH; 50 mM natrium acetate: tetrahydrofuran pH 6.8) and gradient B (65% CH3OH). The system was operated at a flow rate of 1.5 mL/min with an excitation wavelength at 300 nm and emission wavelength at 500 nm. GABA standard solutions were prepared in various concentration (0–60 mg/L) and fitted by a standard curve (y = 9280x – 85,753, R= 0.9946) [33].
2.5. Whole Genome Analysis of W. cibaria UF-274
The whole genome sequencing of strain UF-274 was carried out using Oxford Nanopore Technology (ONT). The gDNA samples were used as the input for library preparation using the Library Preparation kit from Oxford Nanopore Technologies. gDNA was repaired using an end prep enzyme mix, generating DNA with 5′-phosphorylated, 3′-dA-tailed ends. The repaired DNA was ligated with an ONT-compatible adapter. The library was quantified with a Qubit Fluorometer before loading onto the flow cell. Sequencing was performed using PromethION (ONT) until the desired yield was achieved. The quality of sequencing was filtered with Nanoplot. Read correction and assembly were performed using Canu (v2.2) and Flye (v2.8), respectively. The assembled sequence was polished four times with Racon and three times with Medaka. Mapping was performed using minimap2. The quality of the assembled sequence was determined using Quast (v5.3.0) and Qualimap (v2.3). Annotation and visualization were conducted using Rast, Prokka (v1.14.5) and Genovi (v0.4.3), respectively. Additional Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) annotation were performed using EggNOG-mapper, while Biosynthetic Gene Cluster (BGC) analysis was done using AntiSmash (v8.0) [28,34,35].
Safety assessment of W. cibaria UF-274 was performed using the ResFinder v4.60 software and Resistance Gene Identifier (RGI) tool v6.0.3 software in the Comprehensive Antibiotic Resistance Database (CARD). Genes linked to resistance to different classes of antibiotics were analyzed. The RGI results were classified into three types: “Perfect hit” (100% identical with reference), “String hit” (bit score > 450 and not identical), and “Loose hit” (bit score < 450, matched in some regions). In addition, ResFinder can also be used to predict antimicrobial resistance genes (AMR) with a 90% threshold and a 60% minimum length. The pathogenicity of the strain was assessed using PathogenFinder (v0.6.0), with the addition of prophage region, Clustered Regularly Interspaced Palindromic Repeats (CRISPRs) and insertion sequences using Proksee server and PHASTER.
3. Results
3.1. Bacterial Composition in Fermented Urutan
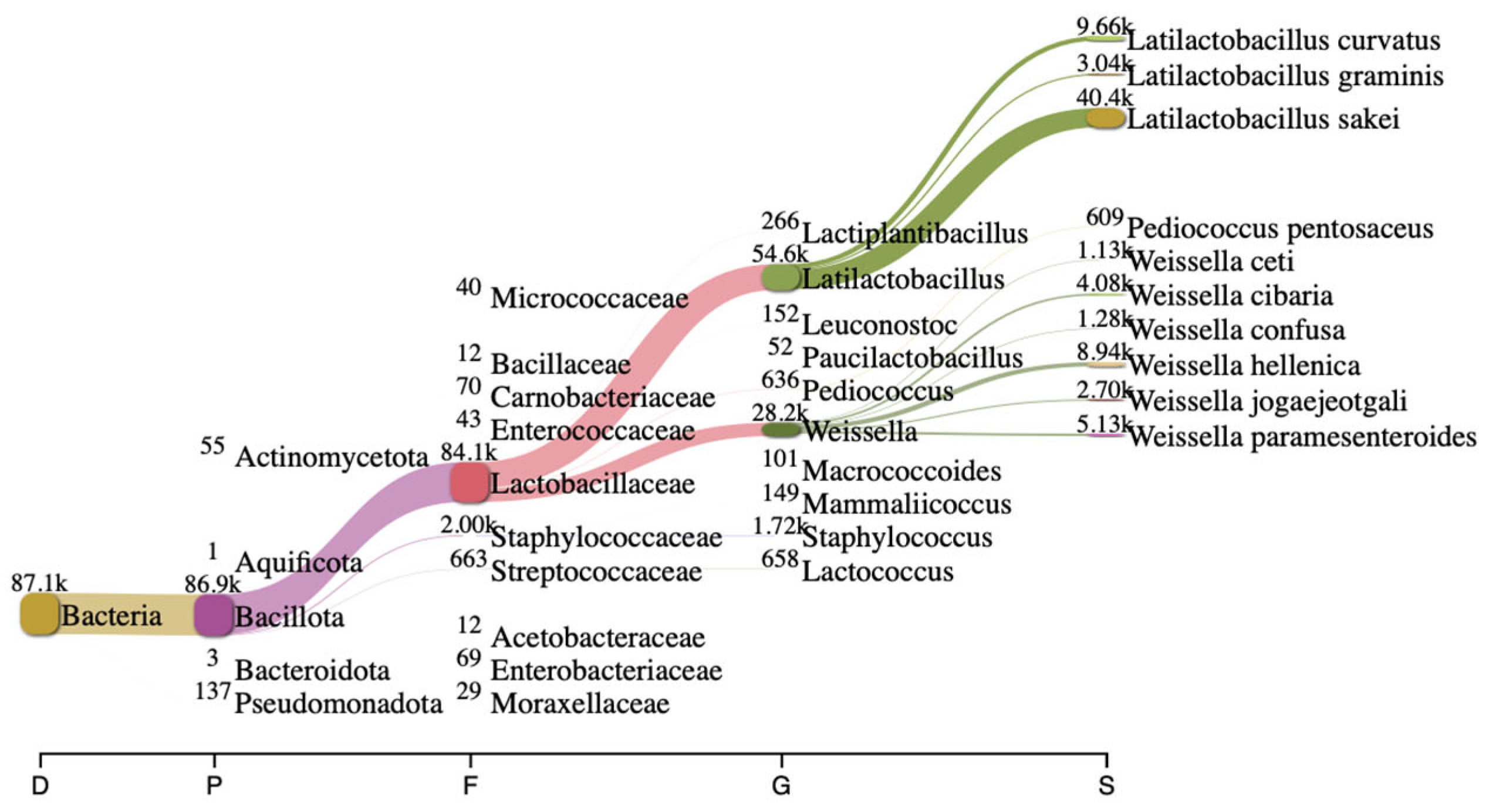
Long-read 16S rRNA sequencing using Next-Generation Sequencing (NGS) is a powerful tool to analyze bacterial composition within communities with a broader profile for species-level identification (Figure 1). In this study, the bacterial composition of artisanal urutan revealed a community structure dominated by Firmicutes, with lactic acid bacteria (LAB)—primarily Weissella, Lactococcus, Lactobacillaceae, Latilactobacillus and Leuconostoc—representing the dominant functional group. At the species level, W. hellenica (8940) was dominant, followed by W. paramesenteroides (5130) and W. cibaria (4080), respectively.
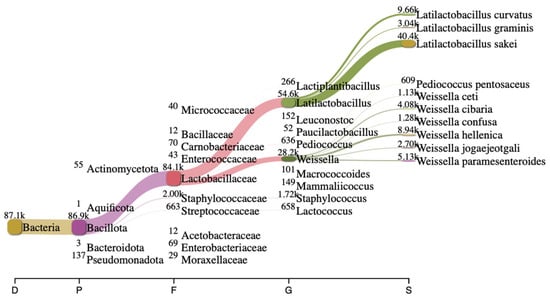
Figure 1.
Bacterial composition in fermented urutan. Sankey plot analysis showing different communities at different hierarchical taxonomies in fermented urutan.
3.2. Screening of GABA-Producing Lactic Acid Bacteria
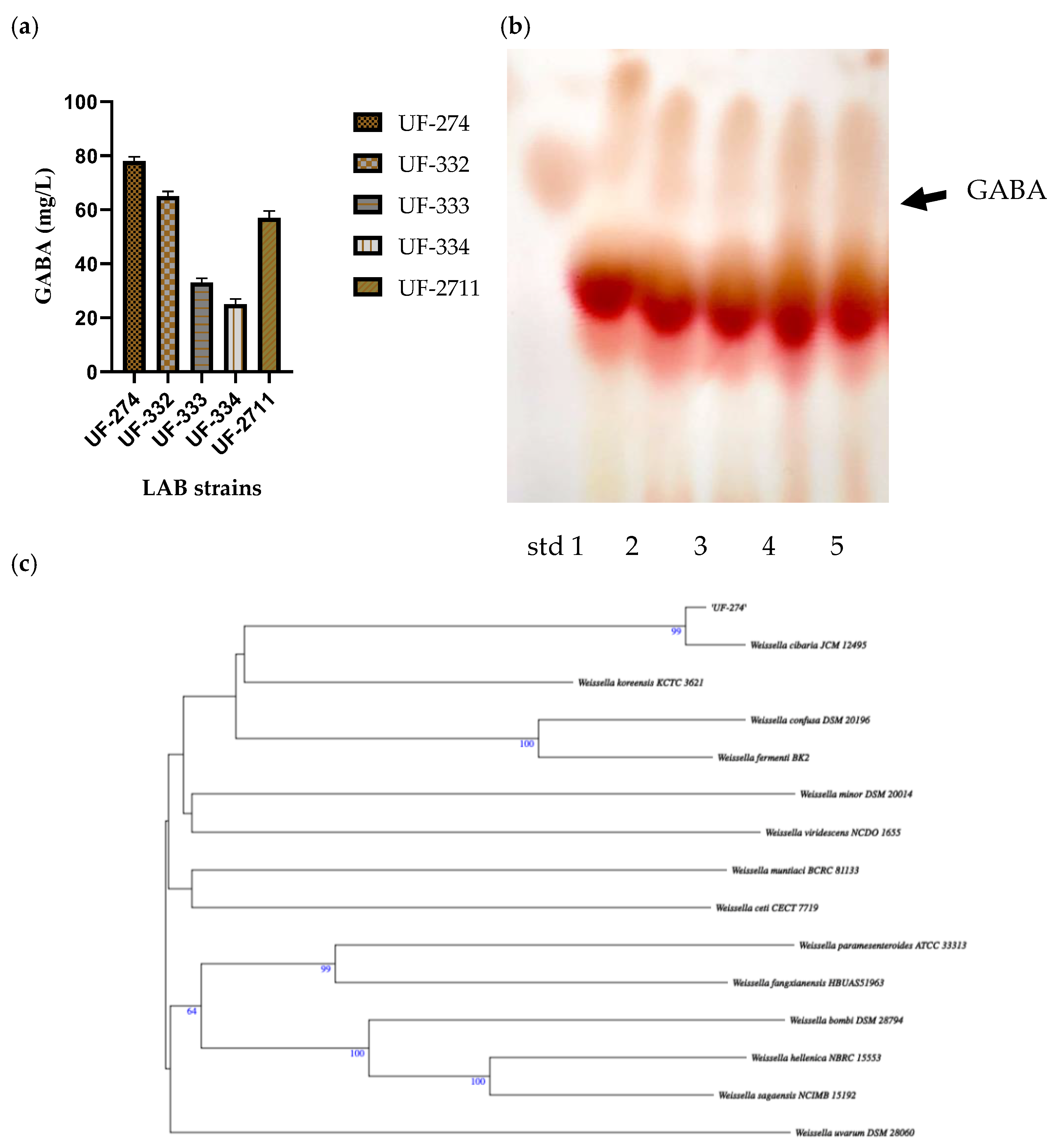
Thirty-two isolates were obtained and screened for their ability to perform GABA synthesis. Five strains were identified as positive GABA producers when using TLC (Figure 2). Quantitative analysis showed that the ability of the strains to produce GABA varied depending on the strains and species, and the best producing strain was UF-274. Strain UF-274 produced 78 mg/L of GABA during 24 h of cultivation in MRS broth medium supplemented with 1% MSG. Other good producers are strains UF-332 and UF-2711 with GABA production of 65.08 mg/L and 56.76 mg/L, respectively. All strains were identified using 16S rDNA barcoding. Four strains (UF-274, UF-332, UF-334, UF-2711) were Weissella cibaria species and one strain, UF-333, was identified as Lactobacillus sakei. Based on a phylogenetic tree, strain W. cibaria UF-274 showed 99% similarity to strain W. cibaria JCM 12495. In addition, this result aligned with a Sankey plot analysis (Figure 1), indicating that the diagram provides an integrated visual confirmation that Weissella is a dominant, culturable taxon in fermented urutan, and culturing successfully retrieved a functionally relevant strain (UF-274) responsible for the observed GABA phenotype.
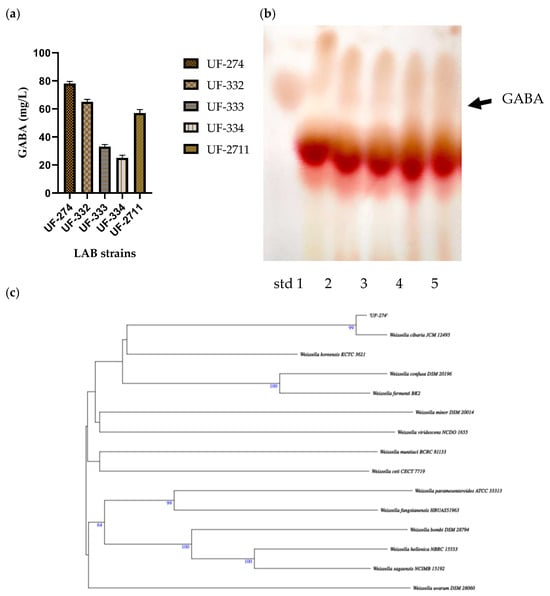
Figure 2.
GABA production in strains W. cibaria (UF—274, 332, 334, 2711) and L. sakei UF-333 (a), TLC screening of their GABA-producing ability, lane 1–5; UF-274, 332, 333, 334, 2711 (b), and (c) analysis of the phylogenetic tree of W. cibaria UF-274 using barcoding 16S rDNA, the tree was constructed using MEGA X software (v. 4).
3.3. Optimization of GABA Production in Skim Milk
To optimize GABA production in skim milk, the three factors glucose, monosodium glutamate and PLP addition to skim milk were selected as independent variables and studied for their effect on GABA enhancement. A Box–Behnken design was used to analyze these variables and to determine the optimum concentrations. A total of sixteen experiments with different concentrations of glucose, MSG and PLP was conducted in triplicate.
The proposed model obtained from the experimental results is shown in Table 1, with the adjustment coefficient value of 0.9251, indicating that the high correlation coefficient (R2) confirms the model reliability for identifying and forecasting the optimal medium composition. The F-value was 8.23, which again suggests that the model is statistically significant, and the p-value was 0.0092. p-values of less than 0.0500 indicate that model terms are significant. In this case B, A2, B2 are significant model terms. The equation in terms of coded factors can be used to make predictions about the response for the given levels of each factor. By default, the high levels of the factors are coded as +1 and the low levels are coded as −1. The coded Equation (1) is useful for identifying the relative impact of the factors by comparing the factor coefficients. The final equation in terms of coded factors is presented in Equation (1):
GABA = +6.28 − 0.1865A + 1.94B + 0.1554C − 0.4690AB − 0.7174AC +
0.6199BC − 1.73A2 − 1.11B2 − 0.8055C2.
0.6199BC − 1.73A2 − 1.11B2 − 0.8055C2.

Table 1.
Analysis of variance (ANOVA) for GABA production.
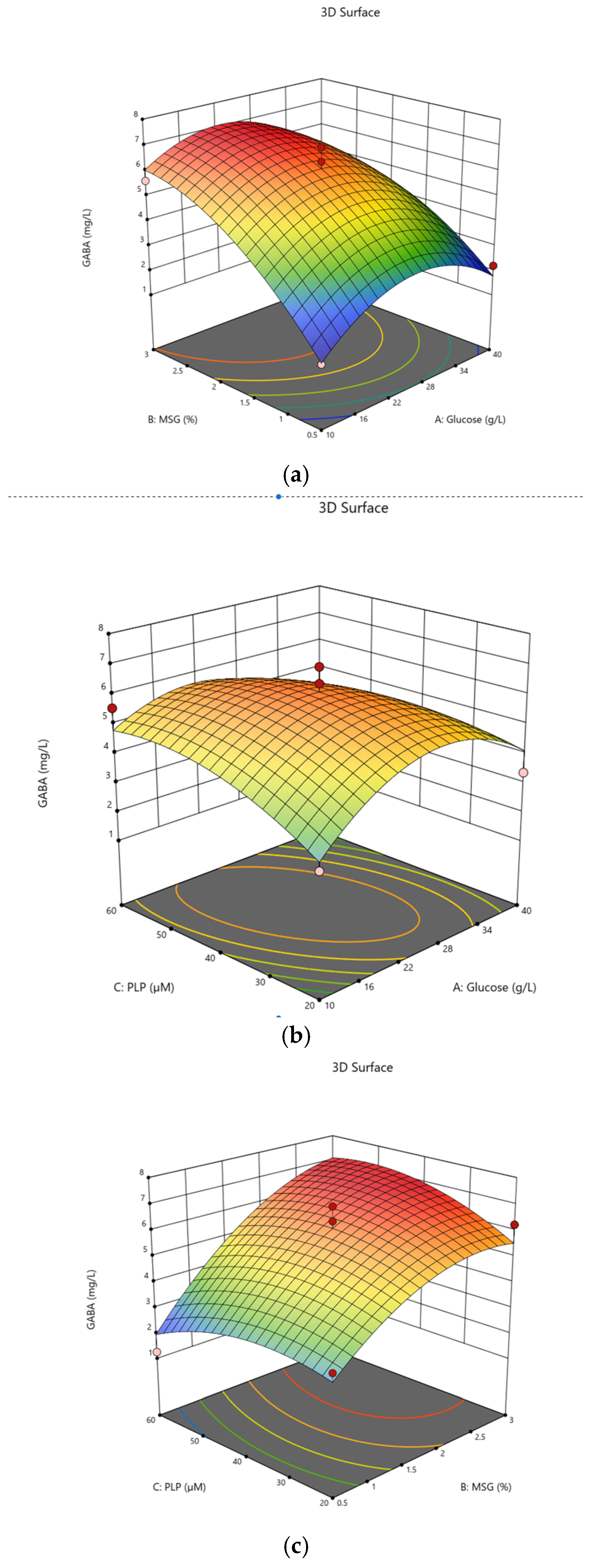
The equation can be used to predict GABA production, where A, B and C code for glucose, MSG and PLP concentrations, respectively. Response surface plots were used to display the interaction amongst glucose, MSG and PLP in skim milk. The addition of glucose slightly increases GABA content in skim milk (Figure 3a). The effects of PLP and glucose on GABA production are presented in Figure 3b. Higher levels of PLP did not significantly affect the GABA content. The effects of MSG and PLP showed a significant interaction (Figure 3c). The availability of PLP suggests that the cofactor modulates GAD activity to convert MSG to GABA.
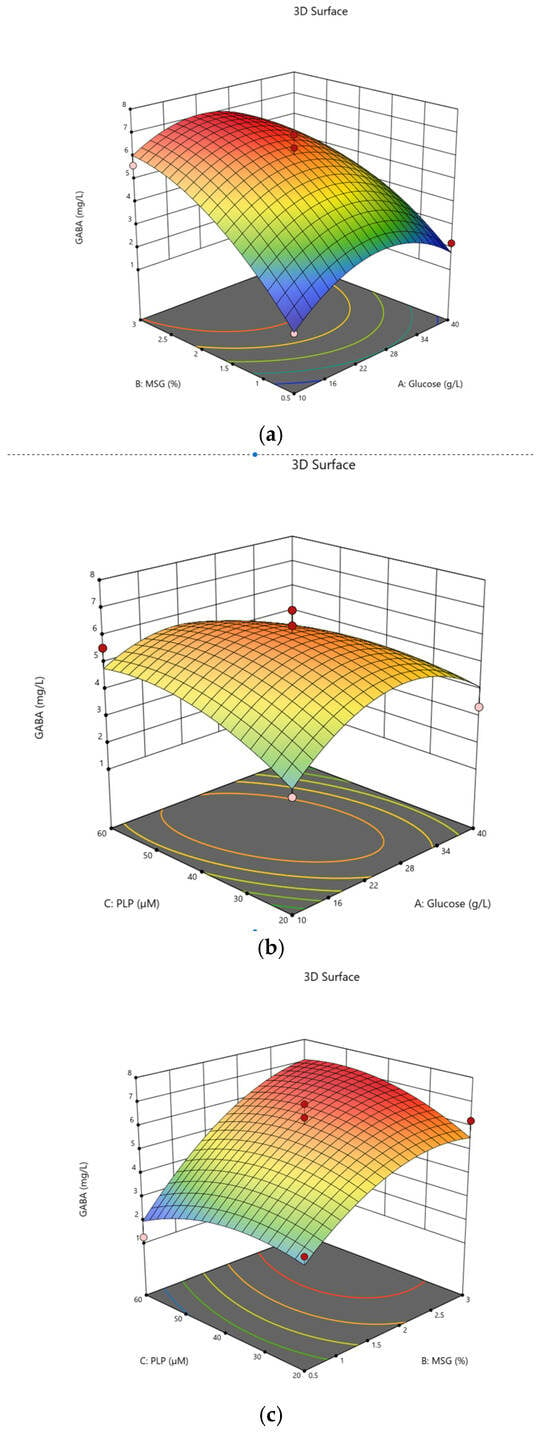
Figure 3.
Contour lines and the response surface for the GABA production in skim milk; (a) GABA production with combination of glucose and MSG; (b) PLP and glucose and (c) combination of MSG and PLP.
The optimum conditions for GABA synthesis according to RSM are a glucose concentration of 23.91 g/L, MSG at 2.32 g/L and PLP at 46 μM. The predicted GABA production under these optimal conditions is 7.040 mg/L. Verification of the actual GABA production was determined under optimal conditions with GABA content of 7.023 mg/L, suggesting that the predicted value is in excellent agreement with the actual experimental value.
3.4. Genomic Characteristic of W. cibaria UF-274
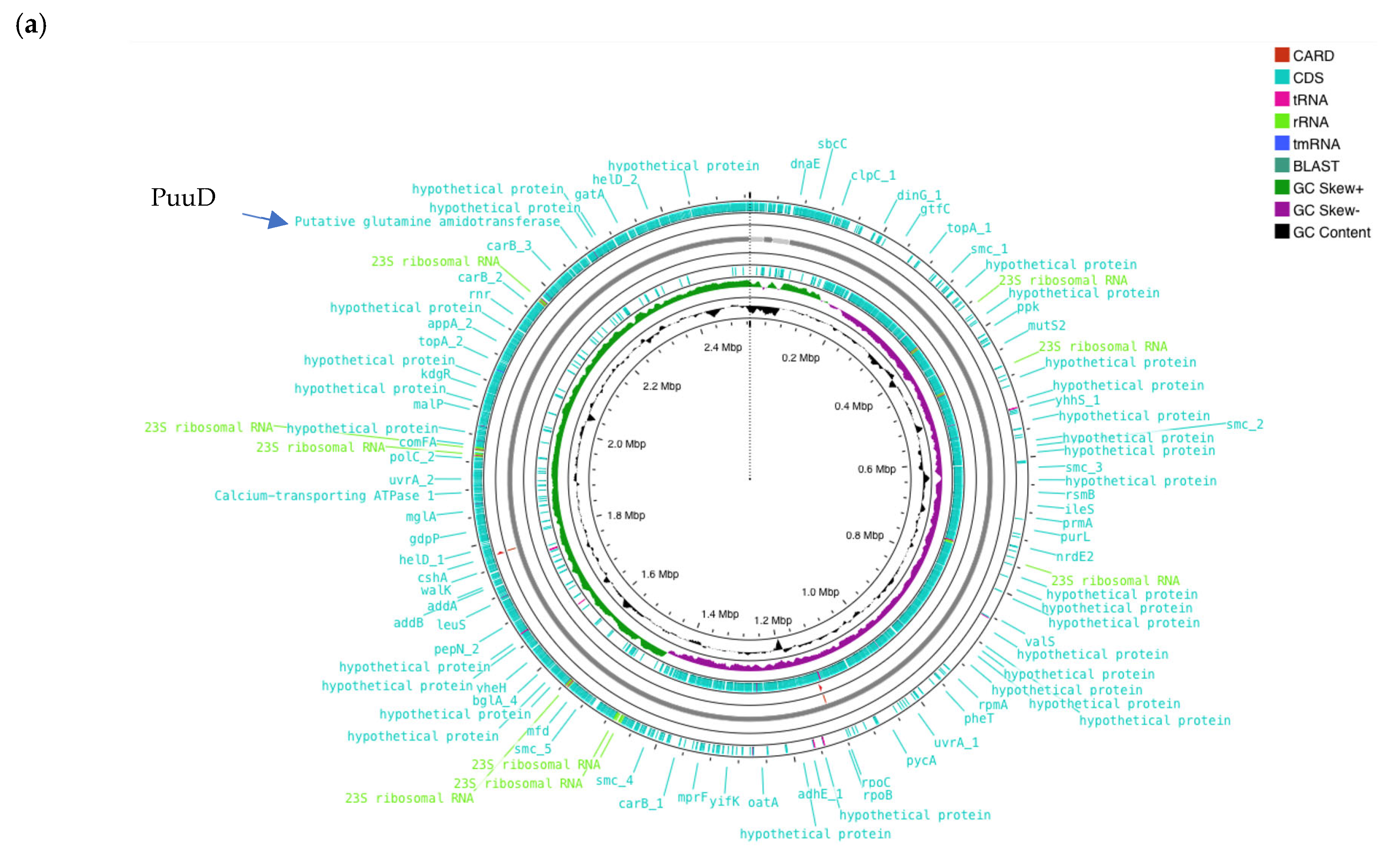
The whole genome sequencing of W. cibaria UF-274 generated a high-quality draft genome with a total length of 2.53 Mb and a GC content of 44.86% (Figure 4a). The genome consists of four contigs with the assembly quality as reflected by a contig N50 of 2.46 Mb and an L50 value of 1. Genome annotation identified 2378 coding sequences (CDS), of which 1577 genes (66.3%) were assigned putative functions, while 864 sequences were classified as hypothetical proteins (Table 2). The genome also encoded 87 tRNA genes and 28 rRNA genes, indicating a well-represented translational machinery typical for lactic acid bacteria. Benchmarking Universal Single-Copy Orthologs (BUSCO) v.5.8 and CheckM was performed to assess the assembly completeness with values of 96% and 98.3% completeness, respectively, with a contamination level of 0.72%.
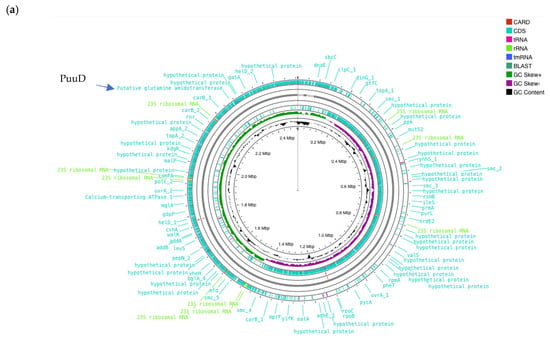
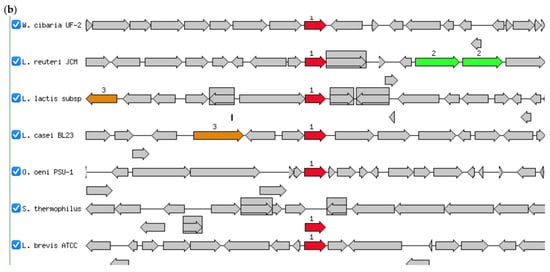
Figure 4.
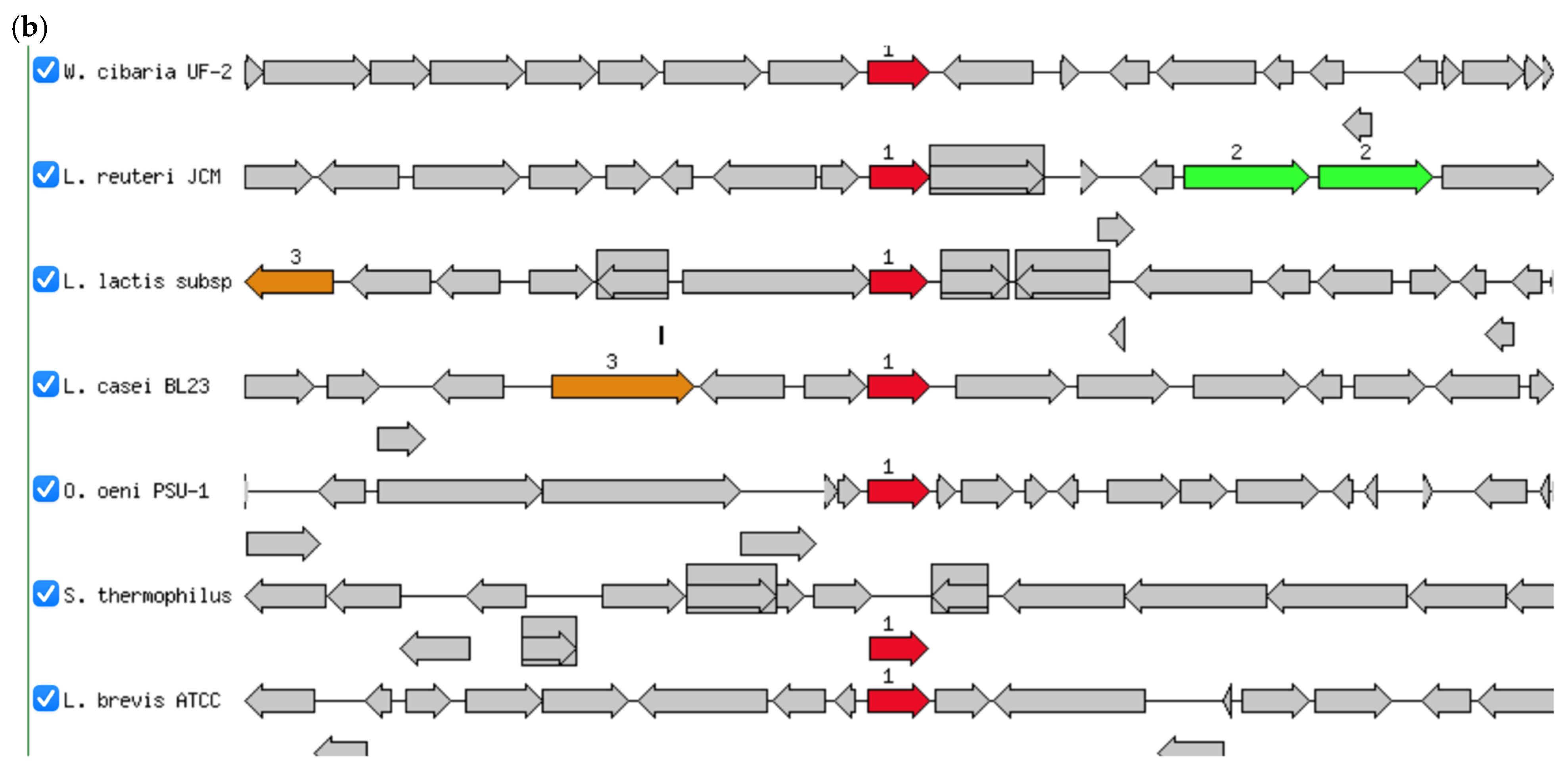
(a) Circular genome of W. cibaria UF-274. From outer to inner rings, open reading frame (ORF), GC content, GC skew – GC skew +. The red lines represent tRNA, green lines represent rRNA and blue lines represent tmRNA. (b) Comparative gene neighborhoods surrounding the glutamine amidotransferase (GATase; red arrows) in Weissella cibaria UF—274 and representative lactic acid bacteria. Genomes were annotated using RAST system. Sets of genes with similar sequence are grouped with the same number and color. Genes whose relative position is conserved in at least four other species are functionally coupled and share gray background boxes. Green arrows indicating alpha glucosyltransferase; brown arrows indicating pyruvate oxidase.

Table 2.
Genomic features of W. cibaria UF-274.
Of particular interest, annotation using RAST, Prokka and EggNOG revealed enzymes potentially involved in GABA metabolism. Genomic region centered on PuuD-like glutamine amidotransferase (GATase) family identified in UF–274 assembly (Figure 4b). The comparative neighborhoods map shows that homologous GATase–encoding genes are present in several LAB species. Although the neighboring genes differ between species, the GATase is consistently located in the same chromosomal region, suggesting it is part of a conserved metabolic module rather than a strain–specific insertion.
3.5. Functional Properties of Annotated Genes
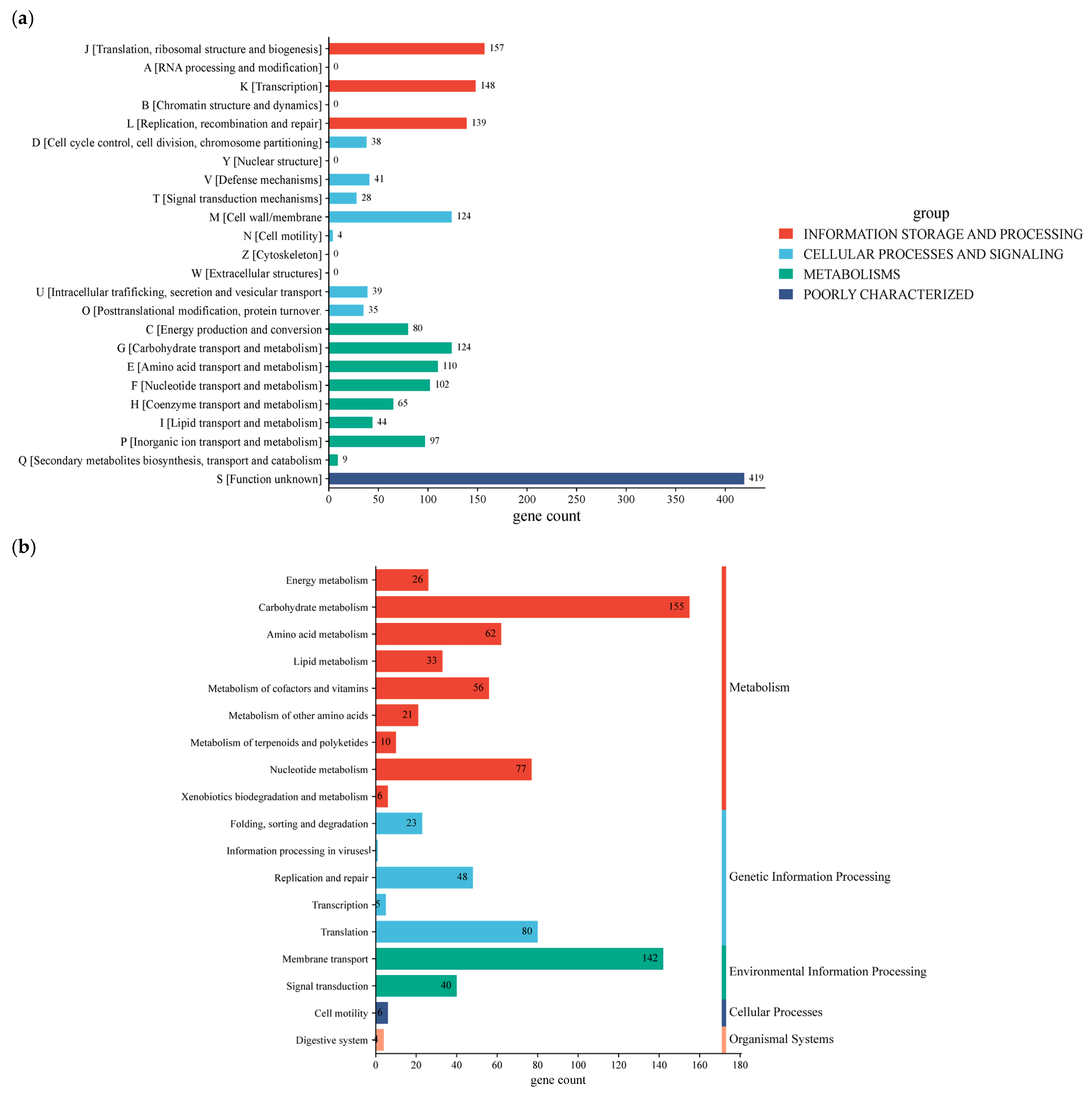
The functional properties of annotated genes were compared against the Clusters of Orthologous Group (COG) and the Kyoto Encyclopedia of Genes and Genomes (KEGG) databases (Figure 5a). Based on annotated gene classification, the chromosome of W. cibaria UF-274 classified into 18 categories, in which 446 genes in total are associated with metabolism, of which 155 genes are linked to carbohydrate metabolism, followed by amino acid metabolism (62 genes), nucleotide metabolism (77 genes), and cofactor as well as vitamin metabolism (56 genes), respectively. In addition, strain W. cibaria UF-274 revealed 156 genes and 148 genes responsible for genetic information processing and environmental information, respectively. Eighty genes are associated with the translation process and 142 genes are related to membrane transport of W. cibaria UF-274. Functional groups based on COG consist of “function unknown” [S] with 419 genes (17.6%), followed by genes that mostly associate with core information processing functions with 444 genes and cellular signaling with 309 genes. Genes assigned to Q (secondary metabolite biosynthesis) were few (nine genes), consistent with in silico antiSMASH detection of two relatively small BGCs (terpene and type-III PKS) in strain UF-274.
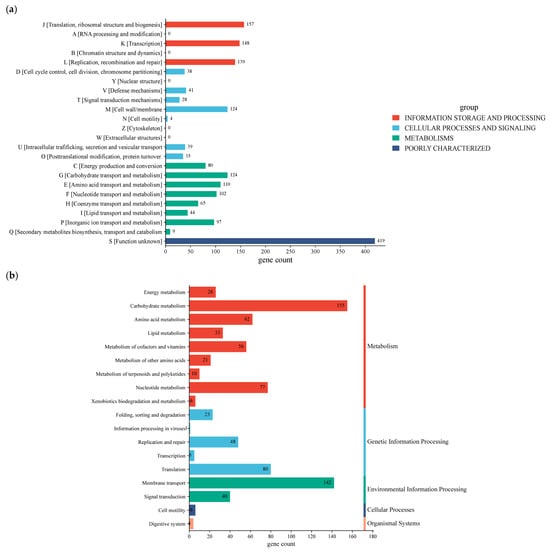
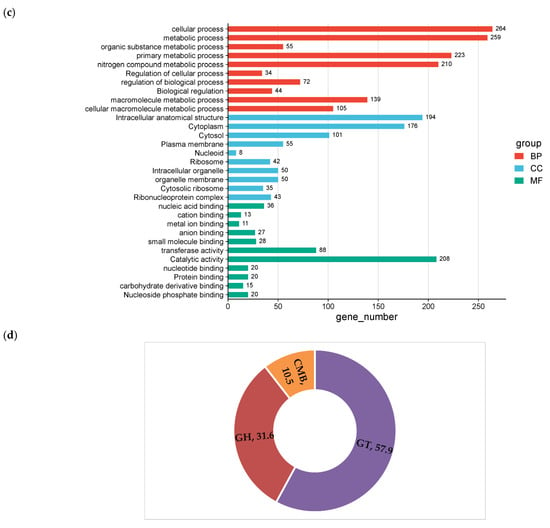
Figure 5.
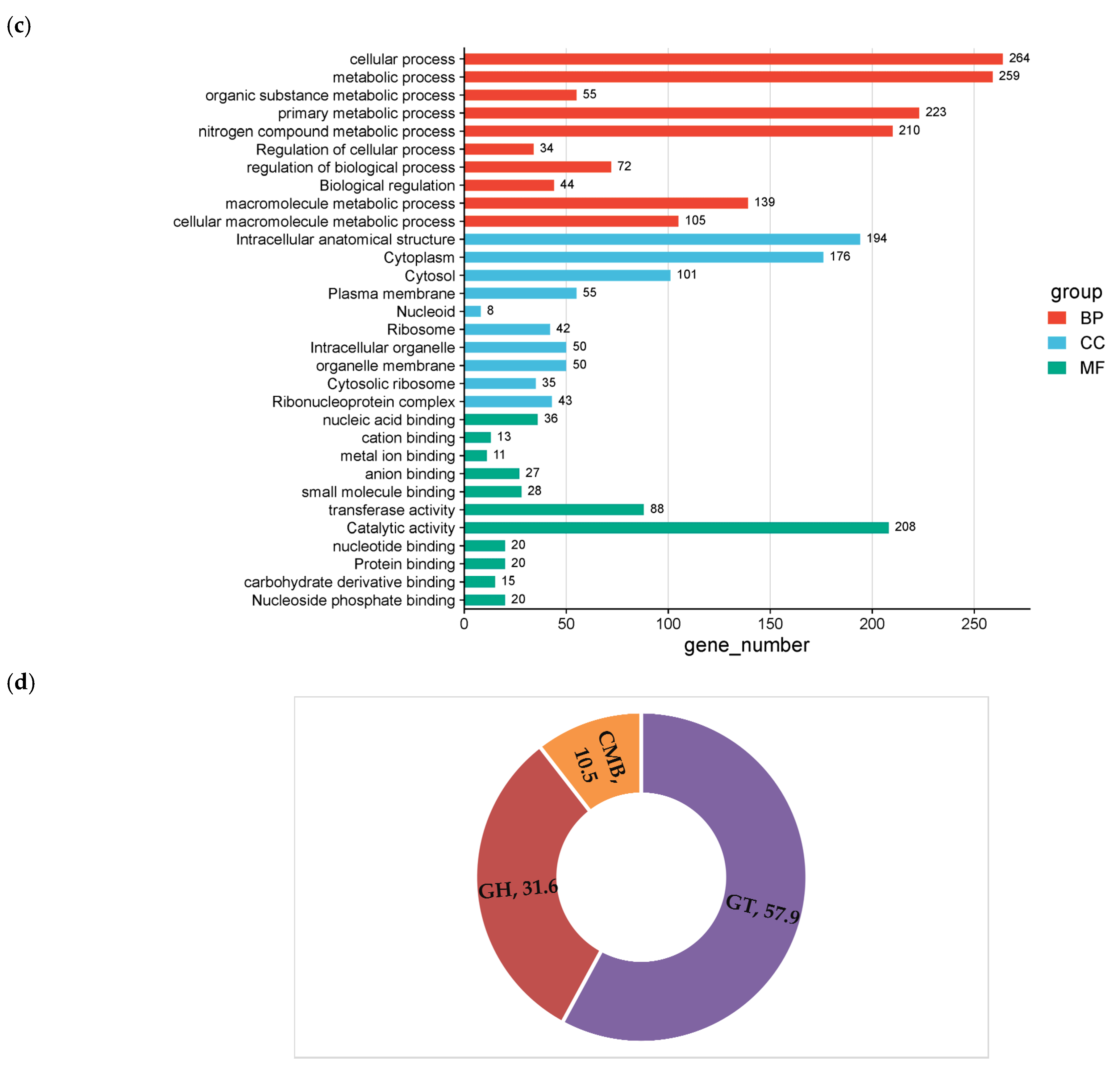
COG functional annotations (a); KEGG functional annotations (b); GO annotation of W. cibaria UF-274 (c); Carbohydrate-active enzymes annotation (d).
KEGG annotation results showed that the strain UF-274 has an active carbohydrate metabolism (155) with the highest results, followed by amino acid (62) and nucleotide metabolism (77), respectively (Figure 5b). In the GO analysis of the strain (Figure 5c), the majority of biological processes are in the category of cellular (264 genes) and metabolic processes (259 genes). The molecular function of the strain is dominated by catalytic activities (208 genes) and transferase activities (88 genes). In addition, the majority of the cellular components consist of intracellular anatomical structures (194) and the cytoplasm (176). A total of 52 proteins were identified as carbohydrate-active enzymes (CAZymes), resulting in a total of 57 CAZy family annotations (Figure 5d). Glycosyltransferases (GT) represent the most abundant class (57.9%), followed by glycoside hydrolases (GH) (31.6%). A clear predominance of GT over GH is commonly associated with cell wall biosynthesis, extracellular polysaccharide production and glycan restructuring rather than degradation of complex carbohydrates. A smaller proportion of carbohydrate-binding modules (CBM) was also detected (10.5%). The dominance of GT families, particularly GT1, GT4, and GT2, suggests that this organism possesses a strong capacity for glycan biosynthesis and carbohydrate modification rather than extensive polysaccharide degradation.
3.6. Biosynthetic Gene Clusters and Safety Analysis
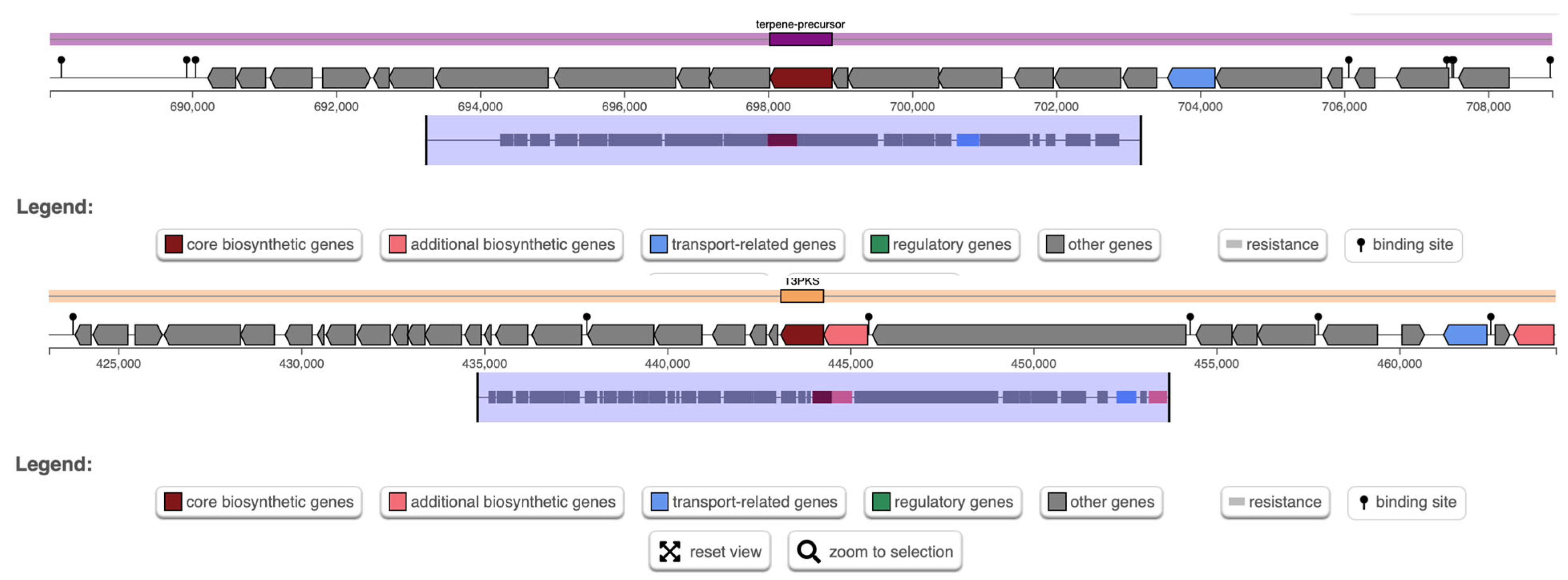
AntiSMASH analysis predicted two biosynthetic gene clusters (BGCs) in W. cibaria UF-274: a terpene cluster and a type-III polyketide synthase (T3PKS) cluster (Figure 6). The T3PKS region (region 4.1) encodes, in addition to the core polyketide synthase (PKS) enzyme, proteins with predicted hydrolase and ligase activities, including a methionine aminopeptidase (MetAP) and a hydroxymethylglutaryl-CoA (HMG-CoA) synthase. The terpene biosynthetic gene cluster (BGC) (region 4.2) contains core genes consistent with phytoene/carotenoid biosynthesis and predicts the potential to synthesize multiple classes of terpenoids, including sesquiterpene, diterpene, sesterterpene and triterpene precursors. Bacterial terpenes are assembled from universal prenyl diphosphate substrates (GPP, FPP, GGPP) by terpene synthases that catalyze ionization and rearrangement to generate diverse skeletons
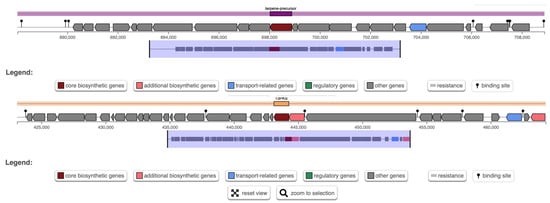
Figure 6.
Analysis of biosynthetic gene clusters in W. cibaria UF-274 genome. Prediction of BCG using AntiSMASH.
To assess the safety of the strain, the presence of antibiotic resistance genes was analyzed using ResFinder and CARD. ResFinder analysis did not detect any antibiotic resistance genes in the W. cibaria UF-274 genome, suggesting the safety of the strain as it cannot transfer antibiotic resistance gene. In contrast, CARD analysis revealed that the strain harbors vanT and vanY genes, encoding for vancomycin resistance. These genes are located on the complement strand in the genome with a length ranging from 1,064,635 to 1,065,420 bp. The product of the genes is D-alanyl-D-alanine carboxypeptidase, commonly found in the bacterial membrane or secreted into periplasmic space.
The genomic stability of the strain was assessed by the presence of prophages in the genome of UF-274. PHASTER analysis showed two prophage regions in the genome with intact prophage regions. These prophage regions have a GC content of 42.39% (region 1) and 42.31% (region 2), respectively. The prophage of each region is located in contig 4 and consists of 48 proteins in region 1 (region length 44.6 kb) and 44 proteins in region 2 (region length 37 kb).
4. Discussion
This study demonstrates the potential of Weissella cibaria UF-274 as a GABA-producing strain adapted to an artisanal fermented-meat niche and evaluates its performance in a skim-milk matrix together with whole-genome characterization. In this study, L. sakei was found abundance in Balinese fermented urutan, as this species is considered a major bacterium in fermented sausages [36,37,38]. Minor populations of non-LAB Firmicutes (including Staphylococcus spp.) were consistently detected across various samples. The presence of Staphylococcus and Lactobacillus is considered as dominant microflora in several types of traditional fermented sausages, where Staphylococcus spp. becomes one of the sub-major species in most fermented sausages [39,40]. Such dominance is consistent with a high protein content, mild salt concentrations, and an anaerobic microenvironment of fermented sausages, which favors heterofermentative and facultative heterofermentative LAB adapted to amino acid and peptide metabolism [39]. A number of GABA-producing LAB were successfully isolated from Nham (Thai fermented meat) such as Lactobacillus namurensis HN8 and Pediococcus pentosaceus NH2, which are considered as good GABA-producers and starters for fermented Nham [41]. To the best of our knowledge, this is the first report of GABA-producing species of W. cibaria and L. sakei isolated from fermented sausages.
The interaction between MSG and glucose indicates a synergistic effect, where MSG provides the substrate for L-glutamic acid decarboxylase (GAD) [11,42]. Furthermore, glucose controls cellular energy and central carbon fluxes that affect intracellular glutamate pools and cofactor regeneration. Gradual increase of PLP apparently did not affect GABA synthesis. Although PLP serves as cofactor for GABA synthesis, higher PLP concentrations did not enhance enzyme activity or exhibit negative physiological effects on cell stability [32]. This was also previously shown in strain L. plantarum FNCC 260 and L. plantarum FRT7, where excessive PLP did not enhance GABA production in MRS broth media [32,43]. The interaction between PLP and MSG suggests a dependence of PLP uptake on regeneration of the cellular metabolic status driven by glucose. In this study, we also observed a substantial reduction in GABA production from 78 mg/L in MRS broth to 7.023 mg/L in skim milk. This can be attributed to several limiting factors such as the presence of free glutamate or glutamate bound in peptides. MRS contains readily available free glutamate, whereas skim milk primarily contains glutamate bound within casein proteins, resulting in limited substrate availability [12,44]. Furthermore, the strain’s proteolytic activity may not be sufficient to release adequate amounts of free glutamate. The strong buffering capacity of milk may also attenuate acid stress induction, which is known to regulate the L-glutamic acid decarboxylase system. In addition, the absence of PLP supplementation and the complex nutrient competition in milk likely further constrained GABA biosynthesis. These factors collectively explain the markedly lower GABA yield observed in the dairy matrix. A similar study by Kanokwan et al. (2023) showed that the strain Lactobacillus futsaii CS3 did not produce GABA in skim milk medium without MSG, whereas the same strain yielded 11 g/L when 2% MSG was added [45]. Likewise, Kim et al. (2022) found that Lactobacillus plantarum Y7 produced 15 μg/mL of GABA in MRS medium supplemented with MSG, and only 6.85 μg/mL of GABA in skim milk medium [46]. These results suggest that MSG supplementation is indispensable and severely limits GABA yields.
Functional annotation of the genome indicates that the strain exhibits robust growth and metabolic versatility under fermentative conditions. A wide array of carbohydrate metabolism genes (G) and cell envelope biogenesis genes (M) may confer diverse carbohydrate substrates and adaptation to technological performance in food matrices such as adherence or tolerance to osmotic/acid stress [27,47]. The relatively high number of amino acid transport and metabolism genes (E) could contribute to GABA biosynthesis, either through the GAD pathway or via alternative gamma-glutamyl/putrescine-linked pathways (PuuD-like amidotransferase) [48,49]. Notably, in the CAZymes database, the strain showed various enzymes involved in the synthesis and degradation of various carbohydrate. The presence of GH families indicates the capability to hydrolyze α-glucans and other oligosaccharides, suggesting moderate carbohydrate utilization potential. Additionally, the detection of CBM families supports the functional role of CAZymes by facilitating substrate recognition and binding, thereby enhancing enzymatic efficiency [50,51].
The genomic analyses reported an absence of canonical gadA/gadB homologues alongside the identification of a puuD-like glutamine amidotransferase and related gamma-glutamyl metabolism genes. Although classical gadA/gadB L-glutamic acid decarboxylase homologs were not detected, the genome harbored genes encoding glutamine amidotransferase class I (peptidase family 26) as well as a gamma-glutamyl-GABA hydrolase-like enzyme (puuD-like) [48]. The enzymes belonging to this latter family are known to cleave gamma-glutamylated intermediates, suggesting that strain W. cibaria UF-274 may utilize an alternative, gamma-glutamyl-based pathway for GABA biosynthesis. The gene orientation revealed that W. cibaria UF-274 harbors the glutamine amidotransferase gene (red arrow) within a conserved syntenic locus shared among lactic acid bacteria [48,49]. The glutamine amidotransferase (GATase) gene is broadly conserved in the core genomes of diverse LAB (Figure 4b). Each species carries a highly similar GATase at a syntenic locus with different neighbors. For example, L. casei, L. brevis and L. reuteri show adjacent polyamine/arginine transport genes, suggesting the region is near arginine/ornithine metabolism genes in those species. We also observed that the gene order varied between lineages and linkage to the central one-carbon/amino acid metabolism in several genomes (for example, association with other aminotransferases). These findings suggest that GATase is present in a conserved chromosomal locus across these LAB species but may vary in flanking genes implying an ancestral gene with lineage-specific rearrangements.
The consistent gene orientation and neighboring decarboxylase-like and transporter genes suggest that this amidotransferase functions as part of an amino acid catabolic operon, potentially catalyzing gamma-glutamyl bond cleavage in gamma glutamyl-GABA intermediate [48]. The absence of gadA/gadB homologs and the presence of this puuD-like amidotransferase (blue arrow in Figure 4a) indicate that W. cibaria UF-274 may utilize an alternative GABA-forming pathway distinct from the conventional GAD system. Furthermore, the functionality of this enzyme is similar to that of GAD but with distinct evolutionary origins or structural characteristics [49]. This arrangement represents a potentially novel genomic organization for GABA biosynthesis within the Weissella genus and warrants further biochemical validation.
AntiSMASH analysis showed that methionine aminopeptidases in region 4.1 are ubiquitous metallopeptidases, required for protein maturation and are often considered therapeutic targets for several diseases such as cancer, obesity and microbial infections [52,53]. Their presence in the cluster may reflect accessory or tailoring functions. HMG-CoA synthase provides a key precursor for the mevalonate/isoprenoid pathway and is associated with biosynthesis of isoprenoid-derived cellular components such as menaquinones, ubiquinones and carotenoids, as well as cell-wall-associated isoprenoids. The region also contains genes showing similarity to bryostatin-like polyketide loci (GenBank DQ889942.1), indicating possible production of complex polyketide scaffolds, although the product identity must be confirmed experimentally [54,55,56]. The combined presence of a T3PKS and terpene biosynthetic machinery suggests that W. cibaria UF-274 harbors biosynthetic potential for structurally diverse secondary metabolites that could influence aroma, redox balance, membrane chemistry or inter-microbial interactions in fermented foods [54].
Safety analysis of the strain showed that the resistance determinants located on the chromosome of LAB are usually intrinsic and lack linkage to mobile genetic elements, meaning they are unlikely to be transmitted horizontally to other bacterial species [57,58]. Vancomycin resistance is commonly found in various Lactobacillus spp. and is considered to be intrinsic [29]. A number of studies have revealed that Weissella spp. are intrinsically resistant to vancomycin [59,60,61,62]. A recent study by Lee et al. (2026) showed that two W. cibaria strains (LB13201, LB3206) possess homologs of vanT and vanY with only 33% identity, which is below the threshold values. Furthermore, the two strains had minimum inhibitory concentration (MIC) ≥ 256 μg/mL suggesting that vancomycin resistance in these strains is intrinsic [63]. The presence of vanT and vanY genes in the strain genome should be interpreted with caution and therefore further transfer risk analysis is required. For this reason, the presence of chromosomal resistance genes is generally viewed as posing a lower biosafety risk than resistance carried on plasmids or transposons. In addition, the risk of the strain being a pathogen was determined with a risk score of 0.043, which is below the threshold and thus the risk of being a human pathogen is considered low. Moreover, VirulenceFinder (v3.2.0) analysis did not detect any virulence factors in the genome. These results suggest that the strain possesses safety characteristics that agree with a function as a starter or adjunct starter for functional foods application.
The presence of prophage may affect the genome stability in bacteria. Most of the prophages were dominated by hypothetical proteins and phage-related proteins from the genera Lactococcus and Enterococcus. Intact prophages exhibit various structural and functional properties and are commonly found in bacteria that inhabit diverse ecosystems [11,64]. Intact prophages found in the bacterial genome also serve as resistance mechanisms under harsh conditions and phage infections. CRISPR analysis showed that only one candidate of a CRISPR region (confirmed) could be found in the genome. This CRISPR region is located on the left flanking sequence with a length of 2,460,442 bp. Unfortunately, the system did not manage to detect the Cas locus in the genome. This could be due to a complete lack of Cas genes, or because the Cas genes are too few/divergent to meet the detection threshold. Genomic surveys of 160 Weissella spp. have shown no Cas gene cluster in their genome [65,66], while others possess a type II-A system (Cas1, Cas2, Cas9, Csn2). This finding indicates that the absence of Cas genes in the genome is a recurring pattern in Weissella and related LAB. The CRISPR-Cas system serves as an immune system in bacteria and archaea by degrading foreign DNA that enters the cell. The presence of a CRISPR-Cas system gives the strain some beneficial properties such as tolerance to phage infections and contributes to genomic stability under harsh conditions [47].
5. Conclusions
This study revealed that strain W. cibaria UF-274 is a promising GABA-producing LAB. Optimization using Box–Behnken in skim milk media can be formulated to enhance postbiotic GABA production. A whole genome analysis revealed that the strain employs an alternative gamma-glutamyl-linked route rather than the gadA/gadB system for GABA synthesis. Further genome mining analysis also revealed two small BGCs, terpene and T3PKS, which may affect ecological traits in fermentations. Safety analysis found no evidence of mobile antibiotic resistance genes; however, van genes and two intact prophages were detected with low pathogen risk scores. Thus, the safety of antibiotic susceptibility and transferable elements requires further validation. Overall, the strain W. cibaria UF-274 is a promising candidate for functional food applications such as a starter/adjunct to enrich GABA in dairy products.
Author Contributions
Conceptualization, I.B.A.Y., R.F., N.W.N. and I.G.A.W.K.; methodology, I.B.A.Y. and R.F.; software, I.B.A.Y. and R.F.; validation, I.B.A.Y., R.F., N.W.N. and I.G.A.W.K.; formal analysis, I.B.A.Y., R.F., N.W.N. and I.G.A.W.K.; investigation, I.B.A.Y., R.F., N.W.N. and I.G.A.W.K.; resources, I.B.A.Y., R.F, N.W.N. and I.G.A.W.K.; data curation, I.B.A.Y., R.F., N.W.N. and I.G.A.W.K.; writing—original draft preparation, I.B.A.Y., R.F., N.W.N., I.G.A.W.K. and D.H.; writing—review and editing, I.B.A.Y., R.F., N.W.N., I.G.A.W.K. and D.H.; visualization, I.B.A.Y., R.F., N.W.N., I.G.A.W.K. and D.H.; supervision, I.B.A.Y.; project administration, I.B.A.Y.; funding acquisition, I.B.A.Y. and I.G.A.W.K. All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by the Directorate General of Higher Education, Sciences and Technology of Indonesia through Fundamental Research scheme with grant number 129/C3/DT.05.00/PL/2025; 2166/LL8/AL.04/2025.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
The data presented in this study are available on request from the corresponding author.
Acknowledgments
The authors express their utmost gratitude to the Directorate General of Higher Education, Research, and Technology, Indonesia, for funding this work.
Conflicts of Interest
The authors declare no conflicts of interest.
Abbreviations
The following abbreviations are used in this manuscript:
| ANOVA | Analysis of Variance |
| BCP | Bromocresol Purple |
| BGC | Biosynthetic Gene Cluster |
| BUSCO | Benchmarking Universal Single-Copy Orthologs |
| CAZymes | Carbohydrate-Active enZymes |
| CDS | Coding Sequence |
| CRISPR | Clustered Regularly Interspaced Short Palindromic Repeats |
| GC | Guanine–Cytosine (content) |
| GABA | γ-Aminobutyric Acid |
| GAD | L-glutamic acid decarboxylase |
| gDNA | Genomic DNA |
| HPLC | High-Performance Liquid Chromatography |
| IS | Insertion Sequence |
| KEGG | Kyoto Encyclopedia of Genes and Genomes |
| MIC | Minimum Inhibitory Concentration |
| MSG | Monosodium Glutamate |
| N50 | Contig length at which 50% of the genome assembly is contained |
| ONT | Oxford Nanopore Technologies |
| OPA | o-Phthaldialdehyde |
| ORF | Open Reading Frame |
| PLP | Pyridoxal-5′-phosphate |
| PKS | Polyketide synthase |
| RSM | Response Surface Methodology |
| RNA | Ribonucleic Acid |
| rRNA | Ribosomal RNA |
| T3PKS | Type III Polyketide Synthase |
| TLC | Thin-Layer Chromatography |
| WGS | Whole Genome Sequencing |
References
- Hong, K.B.; Park, Y.; Suh, H.J. Sleep-promoting effects of the GABA/5-HTP mixture in vertebrate models. Behav. Brain Res. 2016, 310, 36–41. [Google Scholar] [CrossRef] [PubMed]
- Brambilla, P.; Perez, J.; Barale, F.; Schettini, G.; Soares, J.C. GABAergic dysfunction in mood disorders. Mol. Psychiatry 2003, 8, 721–737. [Google Scholar] [CrossRef] [PubMed]
- Tette, F.M.; Kwofie, S.K.; Wilson, M.D. Therapeutic Anti-Depressant Potential of Microbial GABA Produced by Lactobacillus rhamnosus Strains for GABAergic Signaling Restoration and Inhibition of Addiction-Induced HPA Axis Hyperactivity. Curr. Issues Mol. Biol. 2022, 44, 1434–1451. [Google Scholar] [CrossRef] [PubMed]
- Wu, C.; Huang, Y.; Lai, X.; Lai, R.; Zhao, W.; Zhang, M.; Zhao, W. Study on quality components and sleep-promoting effect of GABA Maoyecha tea. J. Funct. Foods 2014, 7, 180–190. [Google Scholar] [CrossRef]
- Yunes, R.A.; Poluektova, E.U.; Dyachkova, M.S.; Klimina, K.M.; Kovtun, A.S.; Averina, O.V.; Orlova, V.S.; Danilenko, V.N. GABA production and structure of gadB/gadC genes in Lactobacillus and Bifidobacterium strains from human microbiota. Anaerobe 2016, 42, 197–204. [Google Scholar] [CrossRef] [PubMed]
- Wu, Q.; Tun, H.M.; Law, Y.S.; Khafipour, E.; Shah, N.P. Common distribution of gad operon in Lactobacillus brevis and its GadA contributes to efficient GABA synthesis toward cytosolic near-neutral pH. Front. Microbiol. 2017, 8, 206. [Google Scholar] [CrossRef] [PubMed]
- Yogeswara, I.B.A.; Maneerat, S.; Haltrich, D. L-glutamic acid decarboxylase from lactic acid bacteria—A key enzyme in Gaba synthesis. Microorganisms 2020, 8, 1923. [Google Scholar] [CrossRef] [PubMed]
- Kurihara, S.; Kato, K.; Asada, K.; Kumagai, H.; Suzuki, H. A putrescine-inducible pathway comprising PuuE-YneI in which γ-aminobutyrate is degraded into succinate in Escherichia coli K-12. J. Bacteriol. 2010, 192, 4582–4591. [Google Scholar] [CrossRef] [PubMed]
- Kurihara, S.; Oda, S.; Kato, K.; Kim, H.G.; Koyanagi, T.; Kumagai, H.; Suzuki, H. A novel putrescine utilization pathway involves γ-glutamylated intermediates of Escherichia coli K-12. J. Biol. Chem. 2005, 280, 4602–4608. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Wang, Y.; Nan, B.; Cao, Y.; Piao, C.; Li, X.; Wang, Y. Optimization of fermentation for gamma-aminobutyric acid (GABA) production by Lactiplantibacillus plantarum Lp3 and the development of fermented soymilk. LWT 2024, 195, 115841. [Google Scholar] [CrossRef]
- Liu, H.; Liu, D.; Zhang, C.; Niu, H.; Xin, X.; Yi, H.; Liu, D.; Zhang, J. Whole-genome analysis, evaluation and regulation of in vitro and in vivo GABA production from Levilactobacillus brevis YSJ3. Int. J. Food Microbiol. 2024, 421, 110787. [Google Scholar] [CrossRef] [PubMed]
- Redruello, B.; Saidi, Y.; Sampedro, L.; Ladero, V.; Del Rio, B.; Alvarez, M.A. Gaba-producing lactococcus lactis strains isolated from camel’s milk as starters for the production of gaba-enriched cheese. Foods 2021, 10, 633. [Google Scholar] [CrossRef] [PubMed]
- Kanklai, J.; Somwong, T.C.; Rungsirivanich, P.; Thongwai, N. Screening of gaba-producing lactic acid bacteria from thai fermented foods and probiotic potential of levilactobacillus brevis f064a for gaba-fermented mulberry juice production. Microorganisms 2021, 9, 33. [Google Scholar] [CrossRef]
- Zhuang, K.; Jiang, Y.; Feng, X.; Li, L.; Dang, F.; Zhang, W.; Man, C. Transcriptomic response to GABA-producing Lactobacillus plantarum CGMCC 1.2437T induced by L-MSG. PLoS ONE 2018, 13, e0199021. [Google Scholar] [CrossRef] [PubMed]
- Yunes, R.A.; Poluektova, E.U.; Vasileva, E.V.; Odorskaya, M.V.; Marsova, M.V.; Kovalev, G.I.; Danilenko, V.N. A Multi-strain Potential Probiotic Formulation of GABA-Producing Lactobacillus plantarum 90sk and Bifidobacterium adolescentis 150 with Antidepressant Effects. Probiotics Antimicrob. Proteins 2020, 12, 973–979. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.J.; Jeon, H.S.; Yoo, J.Y.; Kim, J.H. Some important metabolites produced by lactic acid bacteria originated from kimchi. Foods 2021, 10, 2148. [Google Scholar] [CrossRef]
- Lim, H.S.; Cha, I.T.; Roh, S.W.; Shin, H.H.; Seo, M.J. Enhanced production of gamma-aminobutyric acid by optimizing culture conditions of Lactobacillus brevis HYE1 isolated from kimchi, a Korean fermented food. J. Microbiol. Biotechnol. 2017, 27, 450–459. [Google Scholar] [CrossRef]
- Tanamool, V.; Hongsachart, P.; Soemphol, W. Screening and characterisation of gamma-aminobutyric acid (GABA) producing lactic acid bacteria isolated from Thai fermented fish (Plaa-som) in Nong Khai and its application in Thai fermented vegetables (Som-pak). Food Sci. Technol. 2019, 40, 483–490. [Google Scholar] [CrossRef]
- Vo, T.T.T.; Park, J.H. Characteristics of potential gamma-aminobutyric acid-producing bacteria isolated from Korean and Vietnamese fermented fish products. J. Microbiol. Biotechnol. 2019, 29, 209–221. [Google Scholar] [CrossRef] [PubMed]
- Yogeswara, I.B.A.; Nursini, N.W.; Kusumaningsih, P.; Ayu Kusumawati, I.G. Isolation and Production of γ-Aminobutyric Acid (GABA) from Lactic Acid Bacteria from Seredele, a Balinese Fermented Soybean. Agritech 2025, 45, 341. [Google Scholar] [CrossRef]
- Kusumaningsih, P.; Jatmiko, Y.D.; Kusumawati, I.G.A.; Nursini, N.W.; Yogeswara, I.B.A. Metagenomic analysis of the bacterial community and gamma-aminobutyric acid (GABA) synthesis by lactic acid bacteria from seredele, a Balinese fermented soybean product. Biodiversitas 2024, 25, 4190–4198. [Google Scholar] [CrossRef]
- Ratanaburee, A.; Kantachote, D.; Charernjiratrakul, W.; Sukhoom, A. Selection of γ-aminobutyric acid-producing lactic acid bacteria and their potential as probiotics for use as starter cultures in Thai fermented sausages (Nham). Int. J. Food Sci. Technol. 2013, 48, 1371–1382. [Google Scholar] [CrossRef]
- Santos-Espinosa, A.; Beltrán-Barrientos, L.M.; Reyes-Díaz, R.; Mazorra-Manzano, M.Á.; Hernández-Mendoza, A.; González-Aguilar, G.A.; Sáyago-Ayerdi, S.G.; Vallejo-Cordoba, B.; González-Córdova, A.F. Gamma-aminobutyric acid (GABA) production in milk fermented by specific wild lactic acid bacteria strains isolated from artisanal Mexican cheeses. Ann. Microbiol. 2020, 70, 12. [Google Scholar] [CrossRef]
- Seo, M.J.; Nam, Y.D.; Park, S.L.; Lee, S.Y.; Yi, S.H.; Lim, S.I.L. γ-aminobutyric acid production in skim milk co-fermented with Lactobacillus brevis 877G and Lactobacillus sakei 795. Food Sci. Biotechnol. 2013, 22, 751–755. [Google Scholar] [CrossRef]
- Sun, T.; Zhao, S.; Wang, H.; Cai, C.; Chen, Y.; Zhang, H. ACE-inhibitory activity and gamma-aminobutyric acid content of fermented skim milk by Lactobacillus helveticus isolated from Xinjiang koumiss in China. Eur. Food Res. Technol. 2009, 228, 607–612. [Google Scholar] [CrossRef]
- De Souza Oliveira, R.P.; Perego, P.; De Oliveira, M.N.; Converti, A. Effect of inulin as a prebiotic to improve growth and counts of a probiotic cocktail in fermented skim milk. LWT 2011, 44, 520–523. [Google Scholar] [CrossRef]
- Lee, Y.D.; Park, J.H. Genomic analysis of WCP30 phage of Weissella cibaria for dairy fermented foods. Korean J. Food Sci. Anim. Resour. 2017, 37, 884–888. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Xu, M.; Tang, Y.; Shao, Y.; Wang, H.; Zhang, H. Genome Features and AntiSMASH Analysis of an Endophytic Strain Fusarium sp. R1. Metabolites 2022, 12, 521. [Google Scholar] [CrossRef] [PubMed]
- Stergiou, O.S.; Tegopoulos, K.; Kiousi, D.E.; Tsifintaris, M.; Papageorgiou, A.C.; Tassou, C.C.; Chorianopoulos, N.; Kolovos, P.; Galanis, A. Whole-Genome Sequencing, Phylogenetic and Genomic Analysis of Lactiplantibacillus pentosus L33, a Potential Probiotic Strain Isolated From Fermented Sausages. Front. Microbiol. 2021, 12, 746659. [Google Scholar] [CrossRef] [PubMed]
- EFSA Panel on Biological Hazards (EFSA BIOHAZ Panel); Koutsoumanis, K.; Allende, A.; Alvarez-Ordóñez, A.; Bolton, D.; Bover-Cid, S.; Chemaly, M.; Davies, R.; De Cesare, A.; Hilbert, F.; et al. Whole genome sequencing and metagenomics for outbreak investigation, source attribution and risk assessment of food-borne microorganisms. EFSA J. 2019, 17, e05898. [Google Scholar] [CrossRef]
- Yogeswara, I.B.A.; Nursini, N.W.; Kusumawati, I.G.A.W.; Kusumaningsih, P. Metagenomic Insights into the Bacterial Diversity of Balinese Fermented Sausage (Urutan) from the Household Industry. Fermentation 2024, 10, 629. [Google Scholar] [CrossRef]
- Yogeswara, I.B.A.; Kittibunchakul, S.; Rahayu, E.S.; Domig, K.J.; Haltrich, D.; Nguyen, T.H. Microbial production and enzymatic biosynthesis of γ-aminobutyric acid (GABA) using Lactobacillus plantarum FNCC 260 isolated from indonesian fermented foods. Processes 2021, 9, 22. [Google Scholar] [CrossRef]
- Sanchart, C.; Rattanaporn, O.; Haltrich, D.; Phukpattaranont, P.; Maneerat, S. Lactobacillus futsaii CS3, a New GABA-Producing Strain Isolated from Thai Fermented Shrimp (Kung-Som). Indian. J. Microbiol. 2017, 57, 211–217. [Google Scholar] [CrossRef] [PubMed]
- Yogeswara, I.B.A.; Mariyatun Pramesi, P.C.; Rahayu, E.S. Whole-Genome Sequence of Lactiplantibacillus plantarum Mut-3, Isolated from Indonesian Fermented Soybean (Tempeh). Microbiol. Resour. Announc. 2023, 12, e00513-22. [Google Scholar] [CrossRef]
- Blin, K.; Shaw, S.; Vader, L.; Szenei, J.; Reitz, Z.L.; Augustijn, H.E.; Cediel-Becerra, J.D.D.; de Crécy-Lagard, V.; Koetsier, R.A.; Williams, S.E.; et al. AntiSMASH 8.0: Extended gene cluster detection capabilities and analyses of chemistry, enzymology, and regulation. Nucleic Acids Res. 2025, 53, W32–W38. [Google Scholar] [CrossRef] [PubMed]
- Huang, Z.; Shen, Y.; Huang, X.; Qiao, M.; He, R.K.; Song, L. Microbial diversity of representative traditional fermented sausages in different regions of China. J. Appl. Microbiol. 2021, 130, 133–141. [Google Scholar] [CrossRef] [PubMed]
- Bassi, D.; Milani, G.; Daza, M.V.B.; Barbieri, F.; Montanari, C.; Lorenzini, S.; Šimat, V.; Gardini, F.; Tabanelli, G. Taxonomical Identification and Safety Characterization of Lactobacillaceae from Mediterranean Natural Fermented Sausages. Foods 2022, 11, 2776. [Google Scholar] [CrossRef]
- Coton, M.; Deniel, F.; Mounier, J.; Joubrel, R.; Robieu, E.; Pawtowski, A.; Jeuge, S.; Taminiau, B.; Daube, G.; Coton, E.; et al. Microbial Ecology of French Dry Fermented Sausages and Mycotoxin Risk Evaluation During Storage. Front. Microbiol. 2021, 12, 737140. [Google Scholar] [CrossRef] [PubMed]
- Xiao, Y.; Liu, Y.; Chen, C.; Xie, T.; Li, P. Effect of Lactobacillus plantarum and Staphylococcus xylosus on flavour development and bacterial communities in Chinese dry fermented sausages. Food Res. Int. 2020, 135, 109247. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.H.; Lee, E.S.; Kim, B.M.; Oh, M.H. Potential Correlation between Microbial Diversity and Volatile Flavor Compounds in Different Types of Korean Dry-Fermented Sausages. Foods 2022, 11, 3182. [Google Scholar] [CrossRef]
- Ratanaburee, A.; Kantachote, D.; Charernjiratrakul, W.; Sukhoom, A. Enhancement of γ-aminobutyric acid (GABA) in Nham (Thai fermented pork sausage) using starter cultures of Lactobacillus namurensis NH2 and Pediococcus pentosaceus HN8. Int. J. Food Microbiol. 2013, 167, 170–176. [Google Scholar] [CrossRef] [PubMed]
- Kittibunchakul, S.; Yuthaworawit, N.; Whanmek, K.; Suttisansanee, U.; Santivarangkna, C. Health beneficial properties of a novel plant-based probiotic drink produced by fermentation of brown rice milk with GABA-producing Lactobacillus pentosus isolated from Thai pickled weed. J. Funct. Foods 2021, 86, 104710. [Google Scholar] [CrossRef]
- Cai, H.; Li, X.; Li, D.; Liu, W.; Han, Y.; Xu, X.; Yang, P.; Meng, K. Optimization of Gamma-Aminobutyric Acid Production by Lactiplantibacillus plantarum FRT7 from Chinese Paocai. Foods 2023, 12, 3034. [Google Scholar] [CrossRef] [PubMed]
- Ushidee-Radzi, M.A.; Shin Yee, C.; Raja-Razali, R.B.; Zahia-Azizan, N.A.; Di Renzo, T.; Reale, A.; Nazzaro, S.; Marena, P.; Ilham, Z.; Abd Rahman, N.A.; et al. Advances in GABA-Enriched Yogurt and Frozen Yogurt: Microbial Biosynthesis, Functional Properties, and Health Perspectives—A Comprehensive Review. Foods 2025, 14, 4254. [Google Scholar] [CrossRef]
- Thongruck, K.; Maneerat, S. Enhanced Production of Gamma-Aminobutyric Acid (GABA) from Lactobacillus futsaii CS3 Using Agri-Food Industries By-Products Under Batch and Fed-Batch Fermentation. Indian J. Microbiol. 2023, 63, 467–482. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Yoon, Y.W.; Kim, M.S.; Lee, M.H.; Kim, G.A.; Bae, K.; Yoon, S.S. Gamma-aminobutyric acid fermentation in MRS-based medium by the fructophilic Lactiplantibacillus plantarum Y7. Food Sci. Biotechnol. 2022, 31, 333–341. [Google Scholar] [CrossRef] [PubMed]
- Surachat, K.; Kantachote, D.; Deachamag, P.; Wonglapsuwan, M. Genomic insight into Pediococcus acidilactici hn9, a potential probiotic strain isolated from the traditional thai-style fermented beef nhang. Microorganisms 2021, 9, 50. [Google Scholar] [CrossRef] [PubMed]
- Otaru, N.; Ye, K.; Mujezinovic, D.; Berchtold, L.; Constancias, F.; Cornejo, F.A.; Krzystek, A.; de Wouters, T.; Braegger, C.; Lacroix, C.; et al. GABA Production by Human Intestinal Bacteroides spp.: Prevalence, Regulation, and Role in Acid Stress Tolerance. Front. Microbiol. 2021, 12, 656895. [Google Scholar] [CrossRef] [PubMed]
- Diez-Gutiérrez, L.; San Vicente, L.; Luis, L.J.; Villarán Mdel, C.; Chávarri, M. Gamma-aminobutyric acid and probiotics: Multiple health benefits and their future in the global functional food and nutraceuticals market. J. Funct. Foods 2020, 64, 103669. [Google Scholar] [CrossRef]
- Lin, M.; Jama, S.M.; Cheng, Z.; Zong, Y.; Su, Y.; Liu, W.; Liu, L. Exploring CAZymes Differences in Pediococcus acidilactici Strain OM681363 and Lacticaseibacillus paracasei Strain ON606241 Based on Whole-Genome Sequencing. Fermentation 2025, 11, 64. [Google Scholar] [CrossRef]
- Liang, T.; Jiang, T.; Liang, Z.; Zhang, N.; Dong, B.; Wu, Q.; Gu, B. Carbohydrate-active enzyme profiles of Lactiplantibacillus plantarum strain 84-3 contribute to flavor formation in fermented dairy and vegetable products. Food Chem. X 2023, 20, 101036. [Google Scholar] [CrossRef] [PubMed]
- Sudek, S.; Lopanik, N.B.; Waggoner, L.E.; Hildebrand, M.; Anderson, C.; Liu, H.; Patel, A.; Sherman, D.H.; Haygood, M.G. Identification of the putative bryostatin polyketide synthase gene cluster from “Candidatus Endobugula sertula”, the uncultivated microbial symbiont of the marine bryozoan Bugula neritina. J. Nat. Prod. 2007, 70, 67–74. [Google Scholar] [CrossRef] [PubMed]
- Pillalamarri, V.; Reddy, C.G.; Bala, S.C.; Jangam, A.; Kutty, V.V.; Addlagatta, A. Methionine aminopeptidases with short sequence inserts within the catalytic domain are differentially inhibited: Structural and biochemical studies of three proteins from Vibrio spp. Eur. J. Med. Chem. 2021, 209, 112883. [Google Scholar] [CrossRef] [PubMed]
- Yamada, Y.; Kuzuyama, T.; Komatsu, M.; Shin-ya, K.; Omura, S.; Cane, D.E.; Ikeda, H. Terpene synthases are widely distributed in bacteria. Proc. Natl. Acad. Sci. USA 2015, 112, 857–862. [Google Scholar] [CrossRef] [PubMed]
- Helfrich, E.J.N.; Lin, G.M.; Voigt, C.A.; Clardy, J. Bacterial terpene biosynthesis: Challenges and opportunities for pathway engineering. Beilstein J. Org. Chem. 2019, 15, 2889–2906. [Google Scholar] [CrossRef] [PubMed]
- Martinelli, L.; Redou, V.; Cochereau, B.; Delage, L.; Hymery, N.; Poirier, E.; Le Meur, C.; Le Foch, G.; Cladiere, L.; Mehiri, M.; et al. Identification and Characterization of a New Type III Polyketide Synthase from a Marine Yeast, Naganishia uzbekistanensis. Mar. Drugs 2020, 18, 637. [Google Scholar] [CrossRef] [PubMed]
- Toropov, V.; Demyanova, E.; Shalaeva, O.; Sitkin, S.; Vakhitov, T. Whole-genome sequencing of lactobacillus helveticus D75 and D76 confirms safety and probiotic potential. Microorganisms 2020, 8, 329. [Google Scholar] [CrossRef] [PubMed]
- Chokesajjawatee, N.; Santiyanont, P.; Chantarasakha, K.; Kocharin, K.; Thammarongtham, C.; Lertampaiporn, S.; Vorapreeda, T.; Srisuk, T.; Wongsurawat, T.; Jenjaroenpun, P.; et al. Safety Assessment of a Nham Starter Culture Lactobacillus plantarum BCC9546 via Whole-genome Analysis. Sci. Rep. 2020, 10, 10241. [Google Scholar] [CrossRef] [PubMed]
- Kamboj, K.; Vasquez, A.; Balada-Llasat, J.M. Identification and significance of Weissella species infections. Front. Microbiol. 2015, 6, 1204. [Google Scholar] [CrossRef] [PubMed]
- Jang, Y.J.; Gwon, H.M.; Jeong, W.S.; Yeo, S.H.; Kim, S.Y. Safety evaluation of weissella cibaria jw15 by phenotypic and genotypic property analysis. Microorganisms 2021, 9, 450. [Google Scholar] [CrossRef]
- Abriouel, H.; Lerma, L.L.; Casado Muñoz, M.d.C.; Montoro, B.P.; Kabisch, J.; Pichner, R.; Cho, G.-S.; Neve, H.; Fusco, V.; Franz, C.M.A.P.; et al. The controversial nature of the Weissella genus: Technological and functional aspects versus whole genome analysis-based pathogenic potential for their application in food and health. Front. Microbiol. 2015, 6, 1197. [Google Scholar] [CrossRef] [PubMed]
- Tenea, G.N.; Hurtado, P. Next-Generation Sequencing for Whole-Genome Characterization of Weissella cibaria UTNGt21O Strain Originated From Wild Solanum quitoense Lam. Fruits: An Atlas of Metabolites with Biotechnological Significance. Front. Microbiol. 2021, 12, 675002. [Google Scholar] [CrossRef] [PubMed]
- Lee, M.G.; Yum, S.Y.; Gim, G.M.; Eom, K.H.; Jin, D.Y.; Kim, J.S.; Jang, G. Genomic and phenotypic analysis of Weissella cibaria LB13201 and LB13206 isolated from Hanwoo (native Korean cattle) with antimicrobial and anti-inflammatory capability. Front. Microbiol. 2026, 17, 1674601. [Google Scholar] [CrossRef]
- Hu, T.; Cui, Y.; Zhang, Y.; Qu, X.; Zhao, C. Genome Analysis and Physiological Characterization of Four Streptococcus thermophilus Strains Isolated From Chinese Traditional Fermented Milk. Front. Microbiol. 2020, 11, 184. [Google Scholar] [CrossRef] [PubMed]
- Ma, W.; Liu, X.; Jing, Y.; Zhang, M.; Zhang, X.; Wang, C.; Khan, M.Z.; Zhu, M. Weissella: From Beneficial Probiotics to Opportunistic Pathogens—A Review. Nutrients 2025, 17, 3162. [Google Scholar] [CrossRef] [PubMed]
- Vorapreeda, T.; Rampai, T.; Chamkhuy, W.; Nopgasorn, R.; Wannawilai, S.; Laoteng, K. Genomic Insights into the Probiotic Functionality and Safety of Lactiplantibacillus pentosus Strain TBRC 20328 for Future Food Innovation. Foods 2025, 14, 2973. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.