Abstract
Background: The identification of natural antibacterial agents from various sources that can act effectively against disease-causing foodborne bacteria is one of the major concerns throughout the world. In the present study, a unique phytobiotics mixture containing thymol, menthol, linalool, trans-anethole, methyl salicylate, 1,8-cineole, and p-cymene was evaluated for antibacterial activity against selected strains of Salmonella spp. Results: The phytobiotics mixture was effective against Salmonella enterica subsp. enterica serovars Enteritidis, Typhimurium, and Kentucky. The minimum inhibitory concentration (MIC) and minimum bactericidal concentration (MBC) values of this unique mixture for these three pathogens were 1:256. Among these three strains, one S. Kentucky presented the most extensive resistance profiles to 18 antibiotics belonging to 5 classes of antibiotics. One of S. Typhimurium presents extensive resistance profiles to 14 antibiotics belonging to 5 classes of antibiotics. Conclusions: The results suggest that the phytobiotics mixture used in the experiment can be used as a strong natural antibacterial agent against Gram-negative foodborne pathogens such as S. Typhimurium, S. Kentucky, and S. Enteritidis. This is a preliminary analysis of the effectiveness of a phytobiotic product in an in vitro model, which may be the starting point for further studies, including in vivo animal models.
1. Introduction
Salmonellosis is the second most common zoonotic disease after campylobacteriosis in the European Union (EU), mostly related to eggs and raw meat from poultry production. Each year in Europe and North America there are 7–10 cases of foodborne illness (per 100,000 of the population) related to non-typhoid Salmonella. It causes 16 million cases of inflammatory fever including salmonellosis, 1.3 billion cases of gastroenteritis, and 3 million deaths from Salmonella worldwide each year. An increase in the confirmed human salmonellosis cases noticed in the last decade in the EU necessitated the search for a possible effective Salmonella control in broiler chicken flocks using various methods, including substances obtained from plants.
For several years, national Salmonella control programs (also in Poland) in broiler flocks have shown that the problem is not disappearing and is even becoming one of the main microbiological threats in poultry production. This, together with the increasing antimicrobial resistance of Salmonella spp. (mostly in humans), forced the necessity to undertake work on alternative methods of controlling and eradicating Salmonella in poultry production using promising antibiotic alternatives including probiotics, prebiotics, symbiotics, organic acids, essential oils, cinnamaldehyde, chitosan, phages, nanoparticles, and vaccines.
Antimicrobial resistance (AMR) is a global growing health threat in both human and veterinary medicine. Poultry production is one of the fastest and most dynamically developing animal productions in the world. This, together with the various conditions of poultry farming such as ineffective biosecurity, a high stocking density, or the increasing threat of viral diseases causing immunosuppression, significantly contributes to the increase in the use of antibiotics in veterinary medicine, and indirectly to the overall increased antibiotic resistance. The latest analysis showed that more than 70% of global antimicrobials produced on Earth are used in food-animal production [1]. Data about AMR transmission pathways and finding about how its spread in different part of the world, can be found in Table 1.

Table 1.
Overview of antimicrobial resistance (AMR) of Salmonella transmission pathways originating from poultry production (based on [2]).
The antibiotics used in animals should be selected from those the World Health Organization (WHO) has listed as being “least important” to human health, and not from those classified and mentioned above as “highest priority critically important”. These antibiotics are often the last line, or one of limited treatments, available to treat serious bacterial infections in humans.
In 2019, WHO identified 32 antibiotics in clinical development that address the WHO list of priority pathogens. However, only six of them were classified as innovative. New antimicrobials agents, not only as classic antibiotics but also any alternatives (for, e.g., plant origin), are urgently needed in both human and veterinary medicine [3].
According to the Rapid Alert System for Food and Feed (RASFF) 2020 Annual report, the most incidents came from “accidental or environmental contamination” (30 incidents, 133 notifications), followed by “hazardous or unauthorized composition” (14 incidents, 42 notifications), and “foodborne outbreak” (9 incidents, 43 notifications). Among pathogenic microorganism groups related to the food sector analysed in the RASFF system, 788 notifications were related to—Salmonella, Listeria monocytogenes, Norovirus, and E. coli. The biggest number of Salmonella notifications (273) associated with poultry meat and poultry meat products was detected in Poland as a country of origin. Approximately half of these concerned Salmonella Enteritidis (149 notifications), for which a food safety criterion is set for fresh poultry. Sixteen operators were identified as recurrent. Number of notifications caused by Salmonella, associated to poultry meat and poultry meat products, divided to the by countries can be found in Table 2. These data clearly indicate that the problem of Salmonella in food of animal origin, in particular in poultry meat and its products, is still valid despite the introduction of many solutions both at the farm, slaughterhouse, and production plant level. [4].

Table 2.
Number of notifications counted for each combination of hazard/product category/notifying country.
The increasing resistance of microorganisms has generated the need to search for novel and, most importantly, effective solutions. As evidenced by current research, an effective alternative are substances produced by higher plants—phytoncides, phytoalexins, phytoanticipins, and other secondary metabolites. The term “phytoncide” was coined for the first time by the Soviet scientist B. P. Tokin [5]. This term describes substances secreted and excreted by telomeric plants that have antibacterial, antiprotozoal, and antifungal properties.
At the same time, scientists were also working on substances excreted by bacteria and fungi, trying to understand the mechanism of action and synthesize them. The milestone and first step to the “Golden Age” of antibiotics development was the discovery of penicillin by Alexander Fleming in 1928. On the other hand, the Second World War caused a huge demand for the substances with antimicrobial properties that can be easily used on the battlefield and prevent wounds infection. That reason forced the development of synthetic solutions over research on natural solutions. In the next 20 years, more than 25 antibiotics were discovered, and they dominated for the next decades [6,7]. Figure 1 shows the timeline and development of antibiotics in comparison to natural solutions.
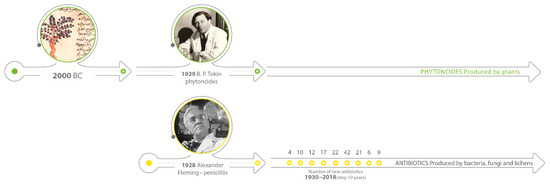
Figure 1.
Timeline of antibacterial natural and synthetic compounds development.
The most recent term to describe plant secondary metabolites possessing antimicrobial properties is phytoanticipins. It was introduced by van Etten et al. and describes low-molecular-weight compounds characterized by antimicrobial properties, which are present in the plant before the appearance of the elicitor, while at the time of pathogen attack they are released or constitute the main component of their synthesis [8]. Such compounds include glucosinolates commonly found in Brassicaceae plants [9,10,11,12] or saponins exemplified by avenacins and a-tomatine [9,12].
Research is currently being conducted on the use of the antimicrobial properties of, i.a. alkaloids [13,14,15], saponins [16,17,18], flavonoids [19,20], and essential oils [21,22,23]. In this article we will focus on essential oils, more precisely on their constituents. Essential oils are volatile, poorly water-soluble, lipophilic natural substances. They are used in many industries such as food, cosmetics, medicine, and pharmaceutics [24]. Essential oils are natural volatile substances obtained from almost every part of plants, including the leaves, buds, flowers, bark, seeds, roots, rhizomes, stems, and fruits by steam distillation, supercritical fluid extraction, and solvent extraction [25,26,27].
Chemically they are mainly mixtures of terpenes (mono-, di-, and sesqui-), terpenoids, phenylpropanoids, isoprenoids, phthalides, and phenolic compounds [25,26,27,28,29].
Figure 2 presents some of the proposed potential mechanisms of antimicrobial action have been observed against bacteria in general, but it is suspected that they may be consistent with those observed for Salmonella spp.
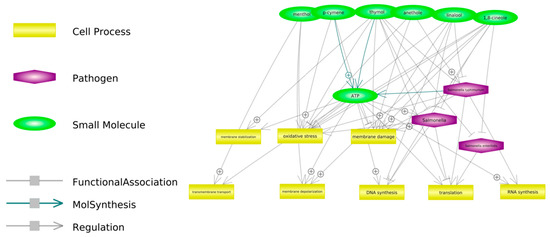
Figure 2.
Potential antibacterial activity of selected components of phytobiotics mixture used in the study. The links were created in the Pathway Studio Web (USA) software on the basis of available publications.
The aim of the present study was to determine the antibacterial activity of phytoncides mixture (thymol, menthol, linalool, anethole, methyl salicylate, 1,8 cineole, and p-cymene) in vitro against Salmonella spp. isolated from broiler farms.
2. Results
The chemical composition of menthol, 1,8-cineole, thymol, p-cymene, anethole, linalool, and methyl salicylate was effective against S. Enteritidis, S. Typhimurium, and S. Kentucky. The minimum inhibitory concentration (MIC) and minimum bactericidal concentration (MBC) values of this unique mixture for all serotypes of these three pathogens were 1:256. In seven strains, those dilutions were even lower, reaching 1:512. Moreover, MBC for the liver Salmonella Enteritidis was 1:1024. Table 3. shows the dilution which was effective against analysed Salmonella spp. Figures S1–S4 show the evaluation of the MIC of the phytoncides mixtures on the analysed Salmonella spp. and the negative control.

Table 3.
MIC of analyzed mixture.
2.1. Antimicrobial Resistance Profile
The isolates were subjected to antibiotic susceptibility tests against 33 antibiotics belonging to 10 different classes using the MIC method Merlin MICRONAUT (MERLIN Diagnostika GmbH, Niemcy, Germany) and the AST-GN 96 CARD and the VITEK2 system (Biomerieux, Marcy-l’Étoile, France). All Salmonella strains of the isolated species belong to 12 Salmonella enterica subsp. enterica and represented 3 serotypes (Typhimurium, Kentucky, Enteritidis). Antibiotic susceptibility testing conducted on the Salmonella strains showed that only two strains S. Typhimurium had resistance to two classes of antibiotics (CFX-CPH-GEN-NEO-STR) whereas other strains were resistant to three or more of the tested antibiotics. All isolated Salmonella were sensitive to imipenem (IMP) and colistin (COL)/polymyxin B (PB), cefequinome (CFQ), and trimethoprim/sulfamethoxazole (TR/SMX). Surprisingly, we detected that 100% of the Salmonella strains were phenotypically resistant to streptomycin and gentamycin. Amongst the resistant strains, only two resistance profiles were identical: CFX-CPH-GEN-NEO-STR (S. Typhimurium). S. Kentucky presented the most extensive resistance profiles to 18 antibiotics (AMP-AMX-AMX/CL-CFX-CFT-CPH-CFP-GEN-NEO-STR-ENR-UB-MRB-NOR-DOX-OXY-TET-LIN/SP) belonging to 5 classes of antibiotics. One of S. Typhimurium presents extensive resistance profiles to 14 antibiotics (AMP-AMX-CFX-CFT-CPH-CFT-CFP-GEN-STR-ENR-UB-MRB-FLR-LIN/SP) belonging to 5 classes of antibiotics. The classes to which it presented the highest resistance were β-lactams (AMP, AMX) and beta-lactam/beta-lactamase inhibitor combination (AMX/CL), I generation cefalexin (CFX-CFT-CPH), III generation cefalexin (CFT, CFP), aminoglycosides (GEN-NEO-STR), fluorochinolones (ENR-UB-MRB-NOR), and tetracyclines (DOX-OXY-TET). For individual serovars of Salmonella spp., Table 4 presents several multi antibiotic resistance patterns.

Table 4.
Multiple antibiotic resistance index (MAR index) and phenotype pattern of Salmonella enterica spp. Enterica, all identified serovars isolates from samples of poultry.
2.2. Prevalence of Multiple Drug Resistance
In our study, four of the isolates showed a multiple antibiotic resistance index (MAR index) greater than 0.3, whereas eight (all S. Enteritidis and two S. Typhimurium) showed a MAR index above 0.5. We observed a high prevalence of multiple antibiotic resistance amongst the isolates where four of the isolates were MDR, with resistance to five different classes of antibiotics.
2.3. Detection of Antibiotic Resistance Genes (ARGs)
Based on the antibiogram data, all isolates were studied for the presence of antibiotic resistance genes. The overall prevalence of ARGs amongst the investigated Salmonella isolates with their resistance phenotype is shown in Table 5. All the isolates were positive for at least one AMR gene. The gene blaCMY2, which confers resistance to ceftiofur (CFTI)/cefoperazone (CFP), was detected in four strains. However, the Salmonella Enteritidis strain did not exhibit phenotypic resistance to III generation cephalosporin. The genes aadA and strA/strB that confer resistance to streptomycin, were detected in all strains. The gene aadB was not detected in the strains. However, all Salmonella spp. strains were phenotypically resistant to gentamicin (GEN). All of the neomycin-resistant strains carried aphA1 and aphA2 genes. The tetA and tetB genes were detected in all strains resistant to doxycycline and oxytetracycline. The gene floR that confers resistance to florfenicol was detected in all strains resistant to florfenicol. The distribution of the various resistance genes and the prevalence of the corresponding serovars are shown in Table 5.

Table 5.
Distribution of resistance genes amongst Salmonella isolates.
3. Discussion
The results of the present study illustrated that the unique phytobiotic mixtures containing menthol, thymol, linalool, 1,8-cineole (eucalyptol), p-cymen, anethole, and methyl salicylate are effective against three Salmonella serotypes isolated from infected materials (birds, dust, and boot swabs) in an in vitro environment. To our best knowledge, there are no scientific reports showing bactericidal effects on Salmonella spp. of similar natural compounds composition.
The previous trend of studies related to searching for possible antibiotic alternatives (also used as antibiotic growth promoters) in animal production, so far has mostly focused on single components or mixtures of 2 to 3 active substances [30,31]. There are also reports with promising results, in the Salmonella spp. Control, with probiotics as an alternative for AGPs [32,33]. Analyzing the history of antibiotics from the discovery of penicillin in 1928 until today, when both human and veterinary medicine face the increasing drug resistance of many dangerous bacteria (including Salmonella spp.) and the lack of new, safe, and effective antibiotics, one should look at the dynamic development of phytobiotics. Phytobiotics, which include a wide range of plant-derived products such as essential oils, herbs, and other bioactive compounds, were first described in 1929 by Russian biochemist Boris Tokin who noticed that certain trees and plants release very active preservatives. This author conducted a number of studies in the 1950s, but only in the last 20 years have we observed a dynamic development of research on the potential effects of single phytobiotics or in simple combinations. Many recent studies show that the antimicrobial mechanism of action of phytobiotics is similar to the action of classic antibiotics [34,35], with the extremely important difference that in the case of phytobiotics, the phenomena of drug resistance and temporary accumulation of active substances in tissues (withdrawal period and antibiotic residues in products of origin) are not observed.
In our study, the sensitivity to 25 antibiotics were assessed. Penicillins (cloxacillin, penicillin G, and nafcillin), macrolides (erythromycin and tylvalosin), lincomycin, tiamulin, and tylvalosin were excluded from analysis, due to a natural lack of activity against Salmonella.
The results of the antibiotic resistance indicated that the Salmonella spp. strains isolated from birds could be categorized as resistant to MDR, that is, bacteria exhibiting resistance to one or more antibiotics from three or more classes of antibiotics. Phenotypic and genotypic profiles of resistance analysis showed that amongst these three strains, S. Kentucky presented the most extensive resistance profiles to 18 antibiotics belonging to 5 classes of antibiotics. One of S. Typhimurium presents extensive resistance profiles to 14 antibiotics belonging to 5 classes of antibiotics. These bacteria were resistant to β-lactams, aminoglycosides, cephalosporins, fluorochinolones, and tetracyclines. All strains were resistant to gentamicin, which is one of the major antibiotics used in the treatment of urinary infections in humans, and to streptomycin. Although streptomycin, an aminoglycoside, is not used for Salmonella treatment, streptomycin resistance has been widely used as an epidemiological marker [36]. For this reason, the results obtained in this study, indicating the antibacterial activity of the mixture of phytobiotics also against Salmonella Kentucky, seem to be extremely interesting and promising not only in terms of the control of salmonellosis in broiler breeding, but also due to the indirect reduction of resistance among this type of bacteria.
Finally, because these antibiotic phenotypes could be conferred by several ARGs, the detection of resistance genes was performed in order to confirm phenotypic antimicrobial resistance profiles.
In Salmonella, the main mechanism of resistance to β-lactams is encoded by the bla genes. The gene blaCMY2, encoding an extended-spectrum beta-lactamase that is responsible for resistance to ceftiofur (CFTI)/cefoperazone (CFP), was detected in four strains. However, the Salmonella Enteritidis strain did not exhibit phenotypic resistance to this antibiotic.
In our study, one S. Typhimurium and two S. Kentucky demonstrated the presence of the genes blaPSE-1 and blaTEM that encode beta-lactamases and confer resistance to ampicillin. The genes aadA and strA/strB that confer resistance to streptomycin were detected in all strains. The gene aadB was not detected in the strains. However, all Salmonella spp. strains were phenotypically resistant to gentamicin. This resistance may be mediated by other resistance genes, which were not assessed in this study.
The proposed composition shows unexpected and very high antibacterial properties against Salmonella spp. The composition was prepared based on experience of the authors with natural compounds and their uses in feed. All the components are allowed to be used as a feed additive according to Regulation (EC) no. 1831/2003 of the European Parliament and of The Council, Appendix I [37]. The composition based also on the scientific data provided by researchers about the used mentioned compounds against microorganisms. [38,39,40,41]. Similar trends have been reported by Kollanoor Johny et al. in their work on the plant-derived molecules trans-cinnamaldehyde, eugenol, carvacrol, and thymol [42]. However, their results test was conducted only on single compounds, not their mixtures and this might have been the reason for the lower efficacy of thymol. The synergistic effect of essential oils compositions was reported by Thanissery et al. The results show stronger antibacterial properties of a thyme–oregano blend against Salmonella, than both essential oils separately [43]. Mixtures with other natural compounds such as organic acids and surfactants present a very good effect against Salmonella spp. [44].
Our findings support the Aljumaah et al. hypothesis that supplementation of phytobiotic feed additives could be very effective in growth promotion, meat quality, and composition and what is most important, S. Typhimurium control [45].
The results obtained in this study, indicate that the antibacterial activity of the mixture of phytobiotics seems to be extremely interesting and promising not only in terms of the control of salmonellosis in broiler breeding, but also due to the indirect reduction of resistance among this type of bacteria.
4. Materials and Methods
4.1. Phytoncides Mixture
Seven common phytoncides were selected for the tests—thymol, menthol, linalool, trans-anethole, methyl salicylate, 1,8-cineole, and p-cymene. All compounds were purchased from Sigma-Aldrich (St. Louis, MO, USA) compliant to FCC and FG standards. Purity and percentage composition, according to supplier specification, was minimum ≥95%.
All phytoncides were mixed in equivalent amounts, heated, and left overnight. The prepared mixture was then mixed with an emulsifier (Polysorbat 80, Sigma-Aldrich) for easier dissolution in aqueous solutions and culture media.
4.2. Salmonella spp. Isolation and Identification
Salmonella spp. from environmental samples were isolated in accordance with PN-EN ISO 6579-1:2017-04. Microbiology of the food chain—horizontal method for the detection, enumeration, and serotyping of Salmonella—Part 1: Detection of Salmonella spp. Samples were taken from several infected broiler farms from different places (boot swabs and dust) and internal organs (intestines, liver, and spleen). Then, samples were suspended in 225 mL buffered peptone water (BPW GRASO, Starogard, Poland) in sterile stomacher bags for pre-enrichment (Whirl-Pak, NASco, Madison, WI, USA). The selective proliferation of Salmonella spp. was carried out using modified semisolid Rappaport–Vassiliadis (MSRV) agar (GRASO). Two selective enrichment media, xylose lysine deoxycholate agar (XLD, GRASO) and brilliant green agar (BGA, GRASO), were used as described [46]. Salmonella suspect colonies were transferred to nonselective nutrient agar (GRASO) to obtain the pure culture for testing flagellar antigens. Serotyping was performed by direct agglutination with commercial H poly antisera for verification of the genus Salmonella enterica (IBSS Biomed, Kraków, Poland), O group antisera to determine O group (IBSS Biomed, Kraków, Poland), and H phase and H factor antisera to determine H phase and H factor (IBSS Biomed, Kraków, Poland, Bio-Rad, Hercules, CA, USA), according to the White–Kauffmann–Le Minor scheme. Pure cultures were used for further biochemical and molecular tests.
4.3. Biochemical Strain Identification
Colonies showing morphology typical for Salmonella spp. on selective agars were subjected to biochemical identification using two commercially available tests: API 20E (BioMérieux, Craponne, France) and a VITEK2 COMPACT automated system for bacterial identification. VITEK® 2 GN cards (BioMérieux, Craponne, France) with reference strains for Salmonella Typhimurium ATCC 14028, Salmonella Enteritidis ATCC 13076, and Salmonella Kentucky ATTC 9263 served as a quality control. Both tests were used according to the manufacturer’s instructions.
4.4. Confirmation of Salmonella Identification with Molecular Biology Methods
A real-Time PCR method based on the detection of genes specific for Salmonella spp. was used to confirm biochemical identification. DNA for Real-time PCR was extracted from bacterial cells using an automated method (AutoPure96, Wuxi, China). For detection of Salmonella spp., a commercial Kylt® Salmonella spp. (Anicon, Emstek, Germany) kit was used. For simultaneous detection of Salmonella Enteritidis and Salmonella Typhimurium, a commercial Spp-Se-St PCR (BioChek, Reeuwijk, The Netherlands) kit was used. Both real-time PCR tests were performed according to the manufacturer’s instructions using an Applied Biosystems 7500 Fast Real-Time PCR System (Thermo, Waltham, MA, USA).
4.5. Phytoncides Mixture Test by Broth Microdilution Method
The antimicrobial activity of the phytoncides mixture was tested using the broth microdilution method described in ISO 20776-1:2006. In sterile vials, two-fold serial dilutions of the phytoncides mixture were prepared in Mueller Hinton II Broth (M-H Broth) with a final volume of 2 mL per vial. Next, fresh inoculum of each bacterial isolate was prepared by suspending colonies from an overnight culture on sheep blood agar in sterile saline (0.9% NaCl) and adjusting the turbidity 0.5 McFarland standard. Subsequently, the suspensions were diluted a hundredfold in M-H Broth by transferring 110 µL of the suspension into 11 mL M-H Broth. Then, 1 mL of this inoculum was transferred into each vial containing 1 mL of diluted product, resulting in the following test dilutions of the product, per row: 4, 8, 16, 32, 64, 128, 256, 512, 1024, 2048, 4096, and 8192. Vials containing 1 mL of M-H Broth only, without product, and 1 mL of inoculum were used as positive growth controls. Wells containing 1 mL of diluted product (a two-fold dilution series) and 1 mL of M-H Broth without any of the bacterial isolates were used as negative controls. Vials were incubated at 35 ± 1 °C for 21 ± 3 h. After incubation, the lowest concentration (the highest dilution) of the product that completely inhibits visible growth was recorded; the minimum inhibitory concentration (MIC). To check for purity after inoculation of the vials, bacterial suspensions made in saline were streaked onto Columbia agar with 5% sheep blood agar. Following overnight incubation at 37 °C, cultures were checked for morphologically characteristic colonies.
4.6. Antibiotic Resistance Test
The 8 classes of antimicrobials agents (β-lactams, aminoglycosides, polymyxins, fluoroquinolones, tetracyclines, macrolides, lincosamides, and sulfonamide) as well as florfenicol, tiamulin, and tylvalosin were used for the antimicrobial susceptibility test.
Antimicrobial susceptibility was assessed by determining the MIC values using a 96 well MICRONAUT Special Plates with antimicrobials: amoxicillin and clavulanic acid (AMX/CL), amoxicillin (AMX), cefquinome (CFQ), ceftiofur (CFTI), cephalexin (CFX), cloxacillin (CLO), colistin (COL), cefapiryna (CPH), docycycline (DOX), enrofloxacin (ENR), erythromycin (ERY), florfenicol (FLR), gentamicin (GEN), lincomycin (LIN), lincomycin/specinicin (LIN/SP), nafcillin (NAF), neomycin (NEO), norfloxacin (NOR), oxytetracycline (OXY), benzylpenicillin (PG), streptomycin (STR), trimethoprim-sulfamethoxazole (TR/SMX), tiamulin (TIA), tylosin (TYL), and tylvalosin (TYLV) (MERLIN Diagnostika GmbH, Bremen, Germany) was used. The MICs were interpreted according to the Clinical and Laboratory Standards Institute (CLSI) and FDA breakpoints [47].
Simultaneously, antimicrobial susceptibility was assessed by determining the MIC values using a VITEK® 2 System and AST-GN96 cards for Gram-negative bacteria (BioMérieux). The AST card is essentially a miniaturized and abbreviated version of the doubling dilution technique for MICs determined by the microdilution method [46]. With using AST-GN96 susceptibility for amoxicillin/clavulanic acid (AMX/CL); ampicillin (AM), cephalexin (CFX), cephalotin (CF), ceftriaxone (CFP), cefequinome (CFQ), ceftiofur (CFTI), enrofloxacin (ENR), florfenicol (FLR), flumequine (UB), gentamicin (GEN), imipenem (IPM), marbofloxacin (MRB), neomycin (NEO), polymyxin B (PB), tetracycline (TE), and trimethoprim/sulfamethoxazole (TR/SMX) and additional antibiotics, ampicillin (AM), cefalotin (CF), cefoperazone (CFP), imipenem (IPM), flumequine (UB), marbofloxacin (MRB), tetracycline (TE), polymyxin B (PB), and trimethoprim/sulfamethoxazole (TR/SMX) was assessed.
The isolates were subjected to antibiotic susceptibility tests against 33 antibiotics belonging to 13 different classes using MIC method Merlin MICRONAUT (MERLIN Diagnostika GmbH, Germany) and AST-GN 96 CARD and VITEK2 system (Biomerieux, France). The AST card is essentially a miniaturized and abbreviated version of the doubling dilution technique for MICs determined by the microdilution method [48]. The multiple antibiotics resistance (MAR) phenotypes were performed for isolates showing resistance to more than two antibiotics [49]. MAR index was using the formula:
4.7. Detection of Antimicrobial Resistance Genes by Multiplex PCR
Bacterial DNA isolation was performed using an automated method (AutoPure96, China). Fourteen resistance genes (aadA, strA/strB, aphA1, aphA2, aadB, tetA, tetB, sul1, sul2, floR, blaTEM, blaSHV, blaCMY-2, and blaPSE-1) were analyzed by conventional PCR using specific primer pairs in multiplex or single PCR reaction The primer sequences predicted PCR product sizes and references shown in Table 6.

Table 6.
Primers sequences for detection of antimicrobial resistance genes in the Salmonella spp. isolate and multiplex PCR annealing temperature (based on [50]).
5. Conclusions
Our results show the antibacterial effect (in vitro) of a unique mixture of phytobiotics on selected Salmonella strains isolated from material collected from flocks where Salmonella infection has been confirmed, as well as on classic reference strains used in the routine diagnosis of Salmonella in broiler chicken flocks. This is the first study where this unique phytobiotics mixture was used.
These promising observations are extremely important not only for poultry production, but also for the current situation in human and animal medicine related to increasing antibiotics resistance. In addition, due to the legal prohibition of treating Salmonella in broiler flocks in a classic way—with the use of antibiotics, the tested mixture of phytobiotics may potentially support the control and spread of this type of bacteria (as a part of non-antibiotic strategy), and ultimately reduce the incidence of infections in humans related to the consumption of poultry products. Therefore, the reduction of Salmonella from the farm seems to be crucial to contribute to food safety. What is also important is that the use of phytobiotics as an antibiotics alternative (also based on our results) seems to be essential in protecting the therapeutic effect of antibiotics and indirectly reducing (preventing) the growing antibiotics resistance.
In summary, our pilot observations on the in vitro model provide valuable information on new potentially effective solutions for the control of Salmonella in poultry production. Due to the fact that these are in vitro observations, the next step in assessing the effectiveness of this unique mixture of phytobiotics should include an in vivo model experiment, taking into account not only the potential antibacterial activity, but also (due to the rich composition of the mixture and the multidirectional action of phytobiotics) on production parameters and meat quality.
Supplementary Materials
The following supporting information can be downloaded at: https://www.mdpi.com/article/10.3390/antibiotics11070868/s1, Table S1: A summary of the results of biochemical reactions for the strain of Salmonella isolated from different samples. Figures S1–S4: MIC evaluation of the phytonicdes composition dilutions.
Author Contributions
Conceptualization, H.I., K.W., K.C. and H.R.; methodology, H.I., K.W., K.C. and H.R.; validation, H.I., K.W. and K.C.; formal analysis, H.I., K.W. and K.C.; investigation, H.I., K.W. and K.C.; data curation, H.I., K.W. and K.C.; writing—original draft preparation, H.I., K.W. and K.C.; writing—review and editing, H.I., K.W., K.C., T.N. and H.R.; visualization H.I., K.W. and K.C.; supervision, H.I., K.W., K.C. and H.R.; project administration, H.I., K.W. and K.C. All authors have read and agreed to the published version of the manuscript.
Funding
This research received no external funding.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
Not applicable.
Acknowledgments
The authors would like to thank Danuta Kukulka for her help in preparing the manuscript.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Van Boeckel, T.P.; Pires, J.; Silvester, R.; Zhao, C.; Song, J.; Criscuolo, N.G.; Gilbert, M.; Bonhoeffer, S.; Laxminarayan, R. Global trends in antimicrobial resistance in animals in low- and middle-income countries. Science 2019, 365. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hedman, H.D.; Vasco, K.A.; Zhang, L. A Review of Antimicrobial Resistance in Poultry Farming within Low-Resource Settings. Animals 2020, 10, 1264. [Google Scholar] [CrossRef] [PubMed]
- World Health Organization. 2019 Antibacterial Agents in Clinical Development: An Analysis of the Antibacterial Clinical Development Pipeline; World Health Organization: Geneva, Switzerland, 2019.
- European. Commission RASFF—The Rapid Alert System for Food and Feed—Annual Report 2020. 2021. Available online: https://ec.europa.eu/food/system/files/2021-08/rasff_pub_annual-report_2020.pdf (accessed on 6 May 2022).
- Duka, R.; Ardelean, D. Phytoncides and phytoalexins—Vegetal antibiotics. J. Med. Arad. 2010, XIII, 19–25. [Google Scholar]
- Katoch, R.; Rajagopalan, S. Warfare Injuries: History, Triage, Transport and Field Hospital Setup in the Armed Forces. Med. J. Armed Forces India 2010, 66, 304–308. [Google Scholar] [CrossRef] [Green Version]
- Hutchings, M.I.; Truman, A.W.; Wilkinson, B. Antibiotics: Past, present and future. Curr. Opin. Microbiol. 2019, 51, 72–80. [Google Scholar] [CrossRef]
- VanEtten, H.D.; Mansfield, J.W.; Bailey, J.A.; Farmer, E.E. Two Classes of Plant Antibiotics: Phytoalexins versus “Phytoanticipins”. Plant Cell 1994, 6, 1191–1192. [Google Scholar] [CrossRef]
- González-Lamothe, R.; Mitchell, G.; Gattuso, M.; Diarra, M.S.; Malouin, F.; Bouarab, K. Plant antimicrobial agents and their effects on plant and human pathogens. Int. J. Mol. Sci. 2009, 10, 3400–3419. [Google Scholar] [CrossRef]
- Chen, F.; Zhang, X.; Du, X.; Yang, L.; Zu, Y.; Yang, F. A new approach for obtaining trans-resveratrol from tree peony seed oil extracted residues using ionic liquid-based enzymatic hydrolysis in situ extraction. Sep. Purif. Technol. 2016, 170, 294–305. [Google Scholar] [CrossRef]
- Parama, D.; Girisa, S.; Khatoon, E.; Kumar, A.; Alqahtani, M.S.; Abbas, M.; Sethi, G.; Kunnumakkara, A.B. An overview of the pharmacological activities of scopoletin against different chronic diseases. Pharmacol. Res. 2022, 179, 106202. [Google Scholar] [CrossRef]
- Piasecka, A.; Jedrzejczak-Rey, N.; Bednarek, P. Secondary metabolites in plant innate immunity: Conserved function of divergent chemicals. New Phytol. 2015, 206, 948–964. [Google Scholar] [CrossRef]
- Dahmer, J.; Marangon, P.; Adolpho, L.O.; Reis, F.L.; Maldaner, G.; Burrow, R.A.; Mostardeiro, M.A.; Dalcol, I.I.; Morel, A.F. Alkaloids from the stem barks of Scutia buxifolia Reissek (Rhamnaceae): Structures and antimicrobial evaluation. Phytochemistry 2022, 196, 113071. [Google Scholar] [CrossRef] [PubMed]
- Long, L.N.; Kang, B.J.; Jiang, Q.; Chen, J.S. Effects of dietary Lycium barbarum polysaccharides on growth performance, digestive enzyme activities, antioxidant status, and immunity of broiler chickens. Poult. Sci. 2020, 99, 744–751. [Google Scholar] [CrossRef] [PubMed]
- Fan, K.; Ding, C.-F.; Deng, S.-Y.; Gao, W.; Tan, B.-Y.; Wu, H.; Guo, Y.; Song, J.-F.; Zhang, L.-C.; Zhang, R.-P.; et al. Monoterpene indole N-oxide alkaloids from Tabernaemontana corymbosa and their antimicrobial activity. Fitoterapia 2022, 158, 105178. [Google Scholar] [CrossRef] [PubMed]
- Sharma, P.; Tyagi, A.; Bhansali, P.; Pareek, S.; Singh, V.; Ilyas, A.; Mishra, R.; Poddar, N.K. Saponins: Extraction, bio-medicinal properties and way forward to anti-viral representatives. Food Chem. Toxicol. 2021, 150, 112075. [Google Scholar] [CrossRef]
- Zaiter, A.; Becker, L.; Baudelaire, E.; Dicko, A. Optimum polyphenol and triterpene contents of Hedera helix (L.) and Scrophularia nodosa (L.): The role of powder particle size. Microchem. J. 2018, 137, 168–173. [Google Scholar] [CrossRef]
- Saboora, A.; Sajjadi, S.-T.; Mohammadi, P.; Fallahi, Z. Antibacterial activity of different composition of aglycone and glycosidic saponins from tuber of Cyclamen coum Miller. Ind. Crops Prod. 2019, 140, 111662. [Google Scholar] [CrossRef]
- Bączek, K.B.; Kosakowska, O.; Przybył, J.L.; Pióro-Jabrucka, E.; Costa, R.; Mondello, L.; Gniewosz, M.; Synowiec, A.; Węglarz, Z. Antibacterial and antioxidant activity of essential oils and extracts from costmary (Tanacetum balsamita L.) and tansy (Tanacetum vulgare L.). Ind. Crops Prod. 2017, 102, 154–163. [Google Scholar] [CrossRef]
- Vieira, V.; Pereira, C.; Pires, T.C.S.P.; Calhelha, R.C.; Alves, M.J.; Ferreira, O.; Barros, L.; Ferreira, I.C.F.R. Phenolic profile, antioxidant and antibacterial properties of Juglans regia L. (walnut) leaves from the Northeast of Portugal. Ind. Crops Prod. 2019, 134, 347–355. [Google Scholar] [CrossRef]
- Kachur, K.; Suntres, Z. The antibacterial properties of phenolic isomers, carvacrol and thymol. Crit. Rev. Food Sci. Nutr. 2020, 60, 3042–3053. [Google Scholar] [CrossRef]
- Chidi, F.; Bouhoudan, A.; Khaddor, M. Antifungal effect of the tea tree essential oil (Melaleuca alternifolia) against Penicillium griseofulvum and Penicillium verrucosum. J. King Saud Univ. Sci. 2020, 32, 2041–2045. [Google Scholar] [CrossRef]
- Mangalagiri, N.P.; Panditi, S.K.; Jeevigunta, N.L.L. Antimicrobial activity of essential plant oils and their major components. Heliyon 2021, 7, e06835. [Google Scholar] [CrossRef] [PubMed]
- Ríos, J.-L. Chapter 1—Essential Oils: What They Are and How the Terms Are Used and Defined. In Essential Oils in Food Preservation, Flavor and Safety; Preedy, V.R., Ed.; Academic Press: San Diego, CA, USA, 2016; pp. 3–10. [Google Scholar]
- Tariq, S.; Wani, S.; Rasool, W.; Shafi, K.; Bhat, M.A.; Prabhakar, A.; Shalla, A.H.; Rather, M.A. A comprehensive review of the antibacterial, antifungal and antiviral potential of essential oils and their chemical constituents against drug-resistant microbial pathogens. Microb. Pathog. 2019, 134, 103580. [Google Scholar] [CrossRef] [PubMed]
- Carson, C.F.; Hammer, K.A. Chemistry and Bioactivity of Essential Oils. In Lipids and Essential Oils as Antimicrobial Agents; Thormar, H., Ed.; John Wiley & Sons, Ltd.: Hoboken, NJ, USA, 2011; pp. 203–238. [Google Scholar]
- Valdivieso-Ugarte, M.; Plaza-Diaz, J.; Gomez-Llorente, C.; Lucas Gómez, E.; Sabés-Alsina, M.; Gil, Á. In vitro examination of antibacterial and immunomodulatory activities of cinnamon, white thyme, and clove essential oils. J. Funct. Foods 2021, 81, 104436. [Google Scholar] [CrossRef]
- Srivastava, S.; Lal, R.K.; Yadav, K.; Pant, Y.; Bawitlung, L.; Kumar, P.; Mishra, A.; Gupta, P.; Pal, A.; Rout, P.K.; et al. Chemical composition of phenylpropanoid rich chemotypes of Ocimum basilicum L. and their antimicrobial activities. Ind. Crops Prod. 2022, 183, 114978. [Google Scholar] [CrossRef]
- Yan, J.; Wang, H.; Wang, Y.; Xu, S.; Wan, Y.; He, L.; Yu, L.; Zhu, W. Integrated metabolome and transcriptome analysis reveals candidate genes involved in metabolism of terpenoids and phthalides in celery seeds. Ind. Crops Prod. 2021, 172, 114011. [Google Scholar] [CrossRef]
- Aljumaah, M.R.; Alkhulaifi, M.M.; Abudabos, A.M.; Alabdullatifb, A.; El-Mubarak, A.H.; Al Suliman, A.R.; Stanley, D. Organic acid blend supplementation increases butyrate and acetate production in Salmonella enterica serovar Typhimurium challenged broilers. PLoS ONE 2020, 15, e0232831. [Google Scholar] [CrossRef] [PubMed]
- Ruvalcaba-Gómez, J.M.; Villagrán, Z.; Valdez-Alarcón, J.J.; Martínez-Núñez, M.; Gomez-Godínez, L.J.; Ruesga-Gutiérrez, E.; Anaya-Esparza, L.M.; Arteaga-Garibay, R.I.; Villarruel-López, A. Non-Antibiotics Strategies to Control Salmonella Infection in Poultry. Animals 2022, 12, 102. [Google Scholar] [CrossRef]
- Adhikari, B.; Hernandez-Patlan, D.; Solis-Cruz, B.; Kwon, Y.M.; Arreguin, M.A.; Latorre, J.D.; Hernandez-Velasco, X.; Hargis, B.M.; Tellez-Isaias, G. Evaluation of the Antimicrobial and Anti-inflammatory Properties of Bacillus-DFM (Norum™) in Broiler Chickens Infected With Salmonella KentuKen. Front. Vet. Sci. 2019, 6, 282. [Google Scholar] [CrossRef] [Green Version]
- Aljumaah, M.R.; Alkhulaifi, M.M.; Abudabos, A.M. In vitro Antibacterial Efficacy of Non-Antibiotic Growth Promoters in Poultry Industry. J. Poult. Sci. 2020, 57, 45–54. [Google Scholar] [CrossRef] [Green Version]
- Balta, I.; Linton, M.; Pinkerton, L.; Kelly, C.; Stef, L.; Pet, I.; Stef, D.; Criste, A.; Gundogdu, O.; Corcionivoschi, N. The effect of natural antimicrobials against Campylobacter spp. and its similarities to Salmonella spp, Listeria spp., Escherichia coli, Vibrio spp., Clostridium spp. and Staphylococcus spp. Food Control 2021, 121, 107745. [Google Scholar] [CrossRef]
- da Silva, A.P.S.A.; Nascimento da Silva, L.C.; Martins da Fonseca, C.S.; de Araújo, J.M.; Correia, M.T.D.S.; Cavalcanti, M.d.S.; Lima, V.L.d.M. Antimicrobial Activity and Phytochemical Analysis of Organic Extracts from Cleome spinosa Jaqc. Front. Microbiol. 2016, 7, 963. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Doran, G.; NiChulain, M.; DeLappe, N.; O’Hare, C.; Corbett-Feeney, G.; Cormican, M. Interpreting streptomycin susceptibility test results for Salmonella enterica serovar Typhimurium. Int. J. Antimicrob. Agents 2006, 27, 538–540. [Google Scholar] [CrossRef] [PubMed]
- European Union, Regulation (EC). No. 1831/2003 of the European Parliament and the council of 22 September 2003 on additives for use in animal nutrition. Off. J. Eur. Commun. 2003, 1, 29. [Google Scholar]
- El-Ashram, S.; Abdelhafez, G.A. Effects of phytogenic supplementation on productive performance of broiler chickens. J. Appl. Poult. Res. 2020, 29, 852–862. [Google Scholar] [CrossRef]
- Kim, D.K.; Lillehoj, H.S.; Lee, S.H.; Jang, S.I.; Park, M.S.; Min, W.; Lillehoj, E.P.; Bravo, D. Immune effects of dietary anethole on Eimeria acervulina infection. Poult. Sci. 2013, 92, 2625–2634. [Google Scholar] [CrossRef]
- Moharreri, M.; Vakili, R.; Oskoueian, E.; Rajabzadeh, G. Effects of microencapsulated essential oils on growth performance and biomarkers of inflammation in broiler chickens challenged with salmonella enteritidis. J. Saudi Soc. Agric. Sci. 2022, 21, 349–357. [Google Scholar] [CrossRef]
- Ed-Dra, A.; Filali, F.R.; Lo Presti, V.; Zekkori, B.; Nalbone, L.; Bouymajane, A.; Trabelsi, N.; Lamberta, F.; Bentayeb, A.; Giuffrida, A.; et al. Chemical composition, antioxidant capacity and antibacterial action of five Moroccan essential oils against Listeria monocytogenes and different serotypes of Salmonella enterica. Microb. Pathog. 2020, 149, 104510. [Google Scholar] [CrossRef]
- Kollanoor Johny, A.; Darre, M.J.; Donoghue, A.M.; Donoghue, D.J.; Venkitanarayanan, K. Antibacterial effect of trans-cinnamaldehyde, eugenol, carvacrol, and thymol on Salmonella Enteritidis and Campylobacter jejuni in chicken cecal contents in vitro1 1Mention of a trade name, proprietary product, or specific equipment does not constitute a guarantee or warranty by the USDA and does not imply its approval to the exclusion of other products that are suitable. J. Appl. Poult. Res. 2010, 19, 237–244. [Google Scholar] [CrossRef]
- Thanissery, R.; Kathariou, S.; Smith, D.P. Rosemary oil, clove oil, and a mix of thyme-orange essential oils inhibit Salmonella and Campylobacter in vitro. J. Appl. Poult. Res. 2014, 23, 221–227. [Google Scholar] [CrossRef]
- Lu, Y.; Wu, C. Reductions of Salmonella enterica on chicken breast by thymol, acetic acid, sodium dodecyl sulfate or hydrogen peroxide combinations as compared to chlorine wash. Int. J. Food Microbiol. 2012, 152, 31–34. [Google Scholar] [CrossRef]
- Aljumaah, M.R.; Suliman, G.M.; Abdullatif, A.A.; Abudabos, A.M. Effects of phytobiotic feed additives on growth traits, blood biochemistry, and meat characteristics of broiler chickens exposed to Salmonella typhimurium. Poult. Sci. 2020, 99, 5744–5751. [Google Scholar] [CrossRef] [PubMed]
- Pławińska-Czarnak, J.; Wódz, K.; Piechowicz, L.; Tokarska-Pietrzak, E.; Bełkot, Z.; Bogdan, J.; Wiśniewski, J.; Kwieciński, P.; Kwieciński, A.; Anusz, K. Wild Duck (Anas platyrhynchos) as a Source of Antibiotic-Resistant Salmonella enterica subsp. diarizonae O58—The First Report in Poland. Antibiotics 2022, 11, 530. [Google Scholar] [CrossRef] [PubMed]
- CLSI. Performance Standards for Antimicrobial Susceptibility Testing, 28th ed.; CLSI Supplement M100; Clinical and Laboratory Standards Institute: Wayne, PA, USA, 2018. [Google Scholar]
- Ramtahal, M.A.; Somboro, A.M.; Amoako, D.G.; Abia, A.L.K.; Perrett, K.; Bester, L.A.; Essack, S.Y. Molecular Epidemiology of Salmonella enterica in Poultry in South Africa Using the Farm-to-Fork Approach. Int. J. Microbiol. 2022, 2022, 5121273. [Google Scholar] [CrossRef] [PubMed]
- Akinola, S.A.; Mwanza, M.; Ateba, C.N. Occurrence, Genetic Diversities And Antibiotic Resistance Profiles Of Salmonella Serovars Isolated From Chickens. Infect. Drug Resist. 2019, 12, 3327–3342. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kozak, G.K.; Boerlin, P.; Janecko, N.; Reid-Smith, R.J.; Jardine, C. Antimicrobial resistance in Escherichia coli isolates from swine and wild small mammals in the proximity of swine farms and in natural environments in Ontario, Canada. Appl. Environ. Microbiol. 2009, 75, 559–566. [Google Scholar] [CrossRef] [Green Version]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).