Chlamydomonas Responses to Salinity Stress and Possible Biotechnological Exploitation
Abstract
1. Introduction
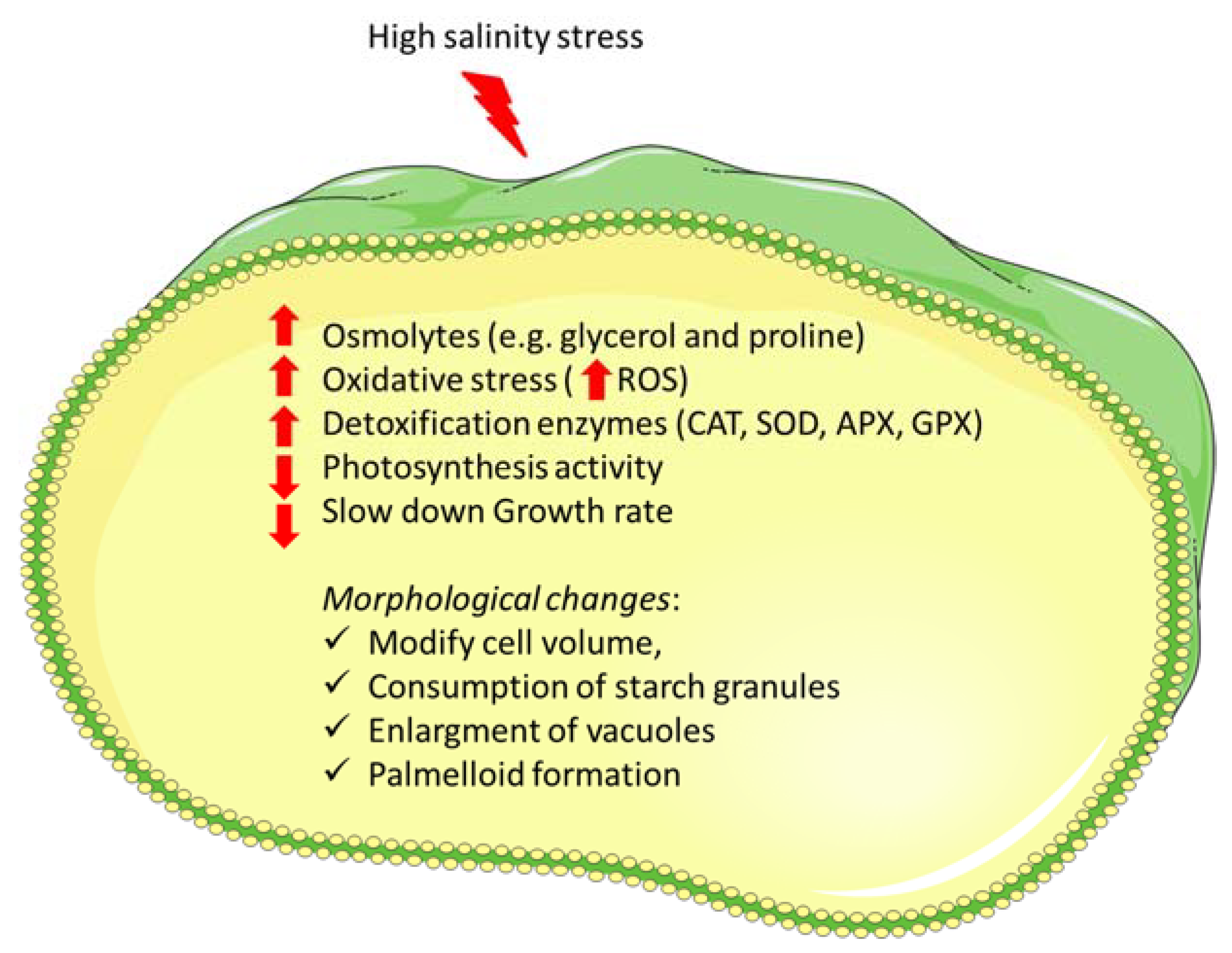
2. Main Physiological Responses to Salinity Stress Exposure
3. Molecular Studies on Salt Stress
3.1. Stress-Related and Antioxidant Genes
3.2. Genes Participating in Compatible Solutes and Lipid Accumulation
3.3. Salt Stress Signaling and Transcription Control
4. Omics Studies
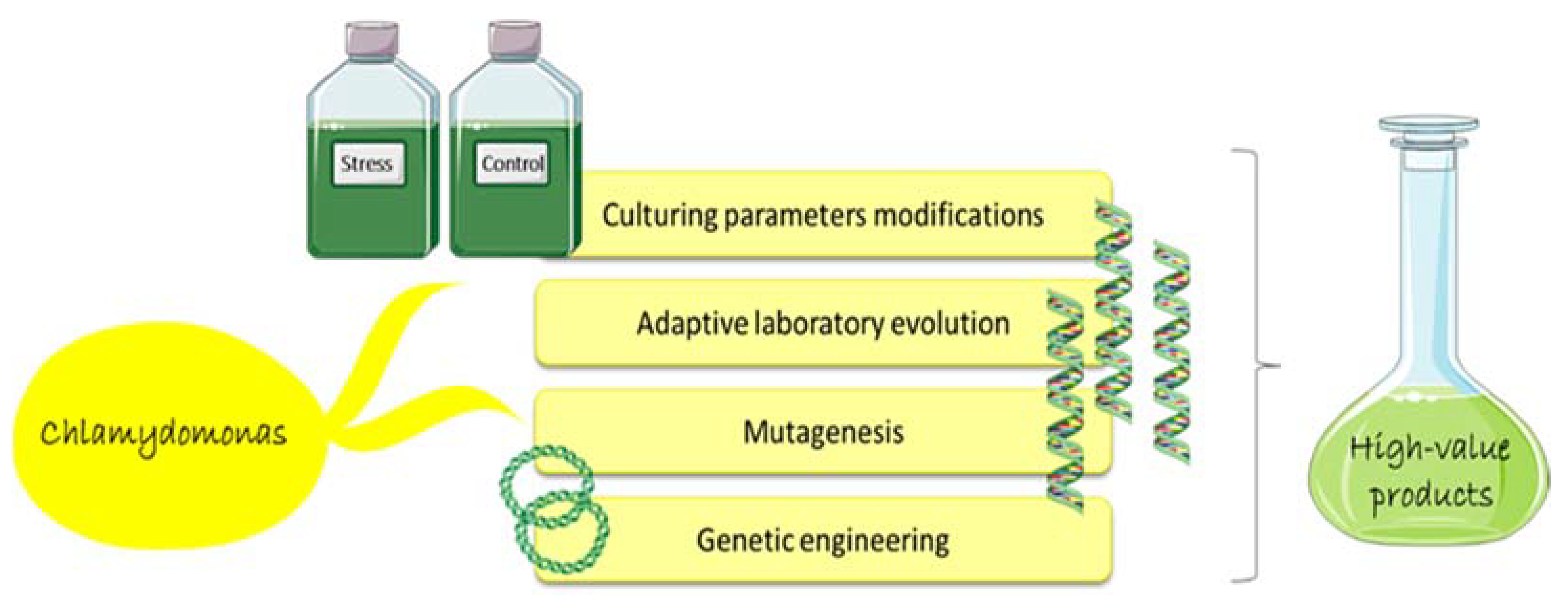
5. Chlamydomonas as Cell Factory
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Durack, P.J.; Wijffels, S.E.; Matear, R.J. Ocean Salinities Reveal Strong Global Water Cycle Intensification during 1950 to 2000. Science 2012, 336, 455–458. [Google Scholar] [CrossRef]
- Beardall, J.; Raven, J.A. The Potential Effects of Global Climate Change on Microalgal Photosynthesis, Growth and Ecology. Phycologia 2004, 43, 26–40. [Google Scholar] [CrossRef]
- Lauritano, C.; Ianora, A. Chemical Defense in Marine Organisms. Mar. Drugs 2020, 18, 518. [Google Scholar] [CrossRef]
- Lauritano, C.; Rizzo, C.; Lo Giudice, A.; Saggiomo, M. Physiological and Molecular Responses to Main Environmental Stressors of Microalgae and Bacteria in Polar Marine Environments. Microorganisms 2020, 8, 1957. [Google Scholar] [CrossRef]
- Harris, E.H. The Chlamydomonas Sourcebook, 2nd ed.; Harris, E.H., Stern, D.B., Witman, G.B., Eds.; Academic Press: London, UK, 2009; p. iii. ISBN 978-0-12-370873-1. [Google Scholar]
- Pröschold, T.; Marin, B.; Schlösser, U.G.; Melkonian, M. Molecular Phylogeny and Taxonomic Revision of Chlamydomonas (Chlorophyta). I. Emendation of Chlamydomonas Ehrenberg and Chloromonas gobi, and Description of Oogamochlamys Gen. Nov. and Lobochlamys Gen. Nov. Promoter of the Systematics of Thegenus. Protist 2001, 152, 265–300. [Google Scholar] [CrossRef]
- Merchant, S.S.; Prochnik, S.E.; Vallon, O.; Harris, E.H.; Karpowicz, S.J.; Witman, G.B.; Terry, A.; Salamov, A.; Fritz-Laylin, L.K.; Maréchal-Drouard, L.; et al. The Chlamydomonas Genome Reveals the Evolution of Key Animal and Plant Functions. Science 2007, 318, 245–250. [Google Scholar] [CrossRef]
- de Carpentier, F.; Lemaire, S.D.; Danon, A. When Unity Is Strength: The Strategies Used by Chlamydomonas to Survive Environmental Stresses. Cells 2019, 8, 1307. [Google Scholar] [CrossRef]
- Hellebust, J.A.; Le Gresley, S.M.L. Growth Characteristics of the Marine Rock Pool Flagellate Chlamydomonas pulsatilla Wollenweber (Chlorophyta). Phycologia 1985, 24, 225–229. [Google Scholar] [CrossRef]
- Ahmad, I.; Hellebust, J.A. The Role of Glycerol and Inorganic Ions in Osmoregulatory Responses of the Euryhaline Flagellate Chlamydomonas pulsatilla Wollenweber 1. Plant. Physiol. 1986, 82, 406–410. [Google Scholar] [CrossRef]
- Hellebust, J.A. Mechanisms of Response to Salinity in Halotolerant Microalgae. Plant. Soil 1985, 89, 69–81. [Google Scholar] [CrossRef]
- Miyasaka, H.; Ohnishi, Y.; Akano, T.; Fukatsu, K.; Mizoguchi, T.; Yagi, K.; Maeda, I.; Ikuta, Y.; Matsumoto, H.; Shioji, N.; et al. Excretion of Glycerol by the Marine Chlamydomonas sp. Strain W-80 in High CO2 Cultures. J. Ferment. Bioeng. 1998, 85, 122–124. [Google Scholar] [CrossRef]
- Yoshimura, K.; Miyao, K.; Gaber, A.; Takeda, T.; Kanaboshi, H.; Miyasaka, H.; Shigeoka, S. Enhancement of Stress Tolerance in Transgenic Tobacco Plants Overexpressing Chlamydomonas Glutathione Peroxidase in Chloroplasts or Cytosol. Plant. J. 2004, 37, 21–33. [Google Scholar] [CrossRef]
- Ho, S.-H.; Nakanishi, A.; Ye, X.; Chang, J.-S.; Hara, K.; Hasunuma, T.; Kondo, A. Optimizing Biodiesel Production in Marine Chlamydomonas sp. JSC4 through Metabolic Profiling and an Innovative Salinity-Gradient Strategy. Biotechnol. Biofuels 2014, 7, 97. [Google Scholar] [CrossRef]
- Xie, Y.; Lu, K.; Zhao, X.; Ma, R.; Chen, J.; Ho, S.-H. Manipulating Nutritional Conditions and Salinity-Gradient Stress for Enhanced Lutein Production in Marine Microalga Chlamydomonas sp. Biotechnol. J. 2019, 14, 1800380. [Google Scholar] [CrossRef]
- Cvetkovska, M.; Hüner, N.P.A.; Smith, D.R. Chilling out: The Evolution and Diversification of Psychrophilic Algae with a Focus on Chlamydomonadales. Polar Biol. 2017, 40, 1169–1184. [Google Scholar] [CrossRef]
- An, M.; Mou, S.; Zhang, X.; Zheng, Z.; Ye, N.; Wang, D.; Zhang, W.; Miao, J. Expression of Fatty Acid Desaturase Genes and Fatty Acid Accumulation in Chlamydomonas sp. ICE-L under Salt Stress. Bioresour Technol. 2013, 149, 77–83. [Google Scholar] [CrossRef]
- Hounslow, E.; Evans, C.A.; Pandhal, J.; Sydney, T.; Couto, N.; Pham, T.K.; Gilmour, D.J.; Wright, P.C. Quantitative Proteomic Comparison of Salt Stress in Chlamydomonas reinhardtii and the Snow Alga Chlamydomonas nivalis Reveals Mechanisms for Salt-Triggered Fatty Acid Accumulation via Reallocation of Carbon Resources. Biotechnol. Biofuels 2021, 14, 121. [Google Scholar] [CrossRef]
- Zhang, Z.; Qu, C.; Zhang, K.; He, Y.; Zhao, X.; Yang, L.; Zheng, Z.; Ma, X.; Wang, X.; Wang, W.; et al. Adaptation to Extreme Antarctic Environments Revealed by the Genome of a Sea Ice Green Alga. Curr. Biol. 2020, 30, 3330–3341.e7. [Google Scholar] [CrossRef]
- Khona, D.K.; Shirolikar, S.M.; Gawde, K.K.; Hom, E.; Deodhar, M.A.; D’Souza, J.S. Characterization of Salt Stress-Induced Palmelloids in the Green Alga, Chlamydomonas reinhardtii. Algal Res. 2016, 16, 434–448. [Google Scholar] [CrossRef]
- Neelam, S.; Subramanyam, R. Alteration of Photochemistry and Protein Degradation of Photosystem II from Chlamydomonas reinhardtii under High Salt Grown Cells. J. Photochem. Photobiol. B Biol. 2013, 124, 63–70. [Google Scholar] [CrossRef]
- Vega, J.M.; Garbayo, I.; Domínguez, M.J.; Vigara, J. Effect of Abiotic Stress on Photosynthesis and Respiration in Chlamydomonas reinhardtii: Induction of Oxidative Stress. Enzyme Microb. Technol. 2006, 40, 163–167. [Google Scholar] [CrossRef]
- Vavilala, S.L.; Gawde, K.K.; Sinha, M.; D’Souza, J.S. Programmed Cell Death Is Induced by Hydrogen Peroxide but Not by Excessive Ionic Stress of Sodium Chloride in the Unicellular Green Alga Chlamydomonas reinhardtii. Eur. J. Phycol. 2015, 50, 422–438. [Google Scholar] [CrossRef]
- Hema, R.; Senthil-Kumar, M.; Shivakumar, S.; Chandrasekhara Reddy, P.; Udayakumar, M. Chlamydomonas reinhardtii, a Model System for Functional Validation of Abiotic Stress Responsive Genes. Planta 2007, 226, 655–670. [Google Scholar] [CrossRef]
- Zuo, Z.-J.; Zhu, Y.-R.; Bai, Y.-L.; Wang, Y. Volatile Communication between Chlamydomonas reinhardtii Cells under Salt Stress. Biochem. Syst. Ecol. 2012, 40, 19–24. [Google Scholar] [CrossRef]
- Eddie, B.; Krembs, C.; Neuer, S. Characterization and Growth Response to Temperature and Salinity of Psychrophilic, Halotolerant Chlamydomonas sp. ARC Isolated from Chukchi Sea Ice. Mar. Ecol. Prog. Ser. 2008, 354, 107–117. [Google Scholar] [CrossRef]
- Kan, G.; Shi, C.; Wang, X.; Xie, Q.; Wang, M.; Wang, X.; Miao, J. Acclimatory Responses to High-Salt Stress in Chlamydomonas (Chlorophyta, Chlorophyceae) from Antarctica. Acta Oceanol. Sin. 2012, 31, 116–124. [Google Scholar] [CrossRef]
- Pittman, J.K.; Edmond, C.; Sunderland, P.A.; Bray, C.M. A Cation-Regulated and Proton Gradient-Dependent Cation Transporter from Chlamydomonas reinhardtii Has a Role in Calcium and Sodium Homeostasis. J. Biol. Chem. 2009, 284, 525–533. [Google Scholar] [CrossRef]
- Shetty, P.; Gitau, M.M.; Maróti, G. Salinity Stress Responses and Adaptation Mechanisms in Eukaryotic Green Microalgae. Cells 2019, 8, 1657. [Google Scholar] [CrossRef]
- León, R.; Galván, F. Halotolerance Studies on Chlamydomonas reinhardtii: Glycerol Excretion by Free and Immobilized Cells. J. Appl. Phycol. 1994, 6, 13–20. [Google Scholar] [CrossRef]
- Reynoso, G.T.; de Gamboa, B.A. Salt Tolerance in the Freshwater Algae Chlamydomonas reinhardii: Effect of Proline and Taurine. Comp. Biochem. Physiol. Part. A Physiol. 1982, 73, 95–99. [Google Scholar] [CrossRef]
- Wang, N.; Qian, Z.; Luo, M.; Fan, S.; Zhang, X.; Zhang, L. Identification of Salt Stress Responding Genes Using Transcriptome Analysis in Green Alga Chlamydomonas reinhardtii. Int. J. Mol. Sci. 2018, 19, 3359. [Google Scholar] [CrossRef]
- Zuo, Z.; Chen, Z.; Zhu, Y.; Bai, Y.; Wang, Y. Effects of NaCl and Na2CO3 Stresses on Photosynthetic Ability of Chlamydomonas reinhardtii. Biologia 2014, 69, 1314–1322. [Google Scholar] [CrossRef]
- Li, Z.; Keasling, J.D.; Niyogi, K.K. Overlapping Photoprotective Function of Vitamin E and Carotenoids in Chlamydomonas. Plant. Physiol. 2012, 158, 313–323. [Google Scholar] [CrossRef]
- Çakmak, Z.E.; Ölmez, T.T.; Çakmak, T.; Menemen, Y.; Tekinay, T. Antioxidant Response of Chlamydomonas reinhardtii Grown under Different Element Regimes. Phycol. Res. 2015, 63, 202–211. [Google Scholar] [CrossRef]
- Lauritano, C.; Procaccini, G.; Ianora, A. Gene Expression Patterns and Stress Response in Marine Copepods. Mar. Environ. Res. 2012, 76, 22–31. [Google Scholar] [CrossRef]
- Yoshida, K.; Igarashi, E.; Wakatsuki, E.; Miyamoto, K.; Hirata, K. Mitigation of Osmotic and Salt Stresses by Abscisic Acid through Reduction of Stress-Derived Oxidative Damage in Chlamydomonas reinhardtii. Plant. Sci. 2004, 167, 1335–1341. [Google Scholar] [CrossRef]
- Abu-Ghosh, S.; Iluz, D.; Dubinsky, Z.; Miller, G. Exogenous Abscisic Acid Confers Salinity Tolerance in Chlamydomonas reinhardtii during Its Life Cycle. J. Phycol. 2021, 57, 1323–1334. [Google Scholar] [CrossRef]
- Mou, S.; Zhang, X.; Ye, N.; Dong, M.; Liang, C.; Liang, Q.; Miao, J.; Xu, D.; Zheng, Z. Cloning and Expression Analysis of Two Different LhcSR Genes Involved in Stress Adaptation in an Antarctic Microalga, Chlamydomonas sp. ICE-L. Extremophiles 2012, 16, 193–203. [Google Scholar] [CrossRef]
- Cruz, J.A.; Salbilla, B.A.; Kanazawa, A.; Kramer, D.M. Inhibition of Plastocyanin to P700 +Electron Transfer in Chlamydomonas reinhardtii by Hyperosmotic Stress. Plant. Physiol. 2001, 127, 1167–1179. [Google Scholar] [CrossRef]
- Subramanyam, R.; Jolley, C.; Thangaraj, B.; Nellaepalli, S.; Webber, A.N.; Fromme, P. Structural and Functional Changes of PSI-LHCI Supercomplexes of Chlamydomonas reinhardtii Cells Grown under High Salt Conditions. Planta 2010, 231, 913–922. [Google Scholar] [CrossRef]
- Ji, C.; Mao, X.; Hao, J.; Wang, X.; Xue, J.; Cui, H.; Li, R. Analysis of BZIP Transcription Factor Family and Their Expressions under Salt Stress in Chlamydomonas reinhardtii. Int. J. Mol. Sci. 2018, 19, 2800. [Google Scholar] [CrossRef]
- Ivanov, A.G.; Hurry, V.; Sane, P.V.; Öquist, G.; Huner, N.P.A. Reaction Centre Quenching of Excess Light Energy and Photoprotection of Photosystem II. J. Plant. Biol. 2008, 51, 85. [Google Scholar] [CrossRef]
- Meijer, H.J.G.; Arisz, S.A.; Van Himbergen, J.A.J.; Musgrave, A.; Munnik, T. Hyperosmotic Stress Rapidly Generates Lyso-Phosphatidic Acid in Chlamydomonas. Plant. J. 2001, 25, 541–548. [Google Scholar] [CrossRef]
- Munnik, T.; Meijer, H.J.G.; Ter Riet, B.; Hirt, H.; Frank, W.; Bartels, D.; Musgrave, A. Hyperosmotic Stress Stimulates Phospholipase D Activity and Elevates the Levels of Phosphatidic Acid and Diacylglycerol Pyrophosphate. Plant. J. 2000, 22, 147–154. [Google Scholar] [CrossRef]
- Arisz, S.A.; Munnik, T. The Salt Stress-Induced LPA Response in Chlamydomonas Is Produced via PLA₂ Hydrolysis of DGK-Generated Phosphatidic Acid. J. Lipid Res. 2011, 52, 2012–2020. [Google Scholar] [CrossRef]
- Meijer, H.J.G.; van Himbergen, J.A.J.; Musgrave, A.; Munnik, T. Acclimation to Salt Modifies the Activation of Several Osmotic Stress-Activated Lipid Signalling Pathways in Chlamydomonas. Phytochemistry 2017, 135, 64–72. [Google Scholar] [CrossRef]
- Arisz, S.A.; van Himbergen, J.A.J.; Musgrave, A.; van den Ende, H.; Munnik, T. Polar Glycerolipids of Chlamydomonas moewusii. Phytochemistry 2000, 53, 265–270. [Google Scholar] [CrossRef]
- Xia, B.-B.; Wang, S.-H.; Duan, J.-B.; Bai, L.-H. The Relationship of Glycerol and Glycolysis Metabolism Patway under Hyperosmotic Stress in Dunaliella salina. Cent. Eur. J. Biol. 2014, 9, 901–908. [Google Scholar] [CrossRef]
- Foflonker, F.; Ananyev, G.; Qiu, H.; Morrison, A.; Palenik, B.; Dismukes, G.C.; Bhattacharya, D. The Unexpected Extremophile: Tolerance to Fluctuating Salinity in the Green Alga Picochlorum. Algal Res. 2016, 16, 465–472. [Google Scholar] [CrossRef]
- Pick, U.; Karni, L.; Avron, M. Determination of Ion Content and Ion Fluxes in the Halotolerant Alga Dunaliella salina. Plant. Physiol. 1986, 81, 92–96. [Google Scholar] [CrossRef]
- Gao, F.; Nan, F.; Feng, J.; Lü, J.; Liu, Q.; Liu, X.; Xie, S. Transcriptome Profile of Dunaliella salina in Yuncheng Salt Lake Reveals Salt-Stress-Related Genes under Different Salinity Stresses. J. Oceanol. Limnol. 2021, 21, 1–27. [Google Scholar] [CrossRef]
- Fu, W.; Paglia, G.; Magnúsdóttir, M.; Steinarsdóttir, E.A.; Gudmundsson, S.; Palsson, B.Ø.; Andrésson, Ó.S.; Brynjólfsson, S. Effects of Abiotic Stressors on Lutein Production in the Green Microalga Dunaliella salina. Microb. Cell Factories 2014, 13, 3. [Google Scholar] [CrossRef]
- Singh, P.; Khadim, R.; Singh, A.K.; Singh, U.; Maurya, P.; Tiwari, A.; Asthana, R.K. Biochemical and Physiological Characterization of a Halotolerant Dunaliella salina Isolated from Hypersaline Sambhar Lake, India. J. Phycol. 2019, 55, 60–73. [Google Scholar] [CrossRef]
- Ye, Z.-W.; Jiang, J.-G.; Wu, G.-H. Biosynthesis and Regulation of Carotenoids in Dunaliella: Progresses and Prospects. Biotechnol. Adv. 2008, 26, 352–360. [Google Scholar] [CrossRef]
- Hoffmann, X.-K.; Beck, C.F. Mating-Induced Shedding of Cell Walls, Removal of Walls from Vegetative Cells, and Osmotic Stress Induce Presumed Cell Wall Genes in Chlamydomonas. Plant. Physiol. 2005, 139, 999–1014. [Google Scholar] [CrossRef][Green Version]
- Miyasaka, H.; Kanaboshi, H.; Ikeda, K. Isolation of Several Anti-Stress Genes from the Halotolerant Green Alga Chlamydomonas by Simple Functional Expression Screening with Escherichia coli. World J. Microbiol. Biotechnol. 2000, 16, 23–29. [Google Scholar] [CrossRef]
- Takeda, T.; Yoshimura, K.; Yoshii, M.; Kanahoshi, H.; Miyasaka, H.; Shigeoka, S. Molecular Characterization and Physiological Role of Ascorbate Peroxidase from Halotolerant Chlamydomonas sp. W80 Strain. Arch. Biochem. Biophys. 2000, 376, 82–90. [Google Scholar] [CrossRef]
- Leisinger, U.; Rüfenacht, K.; Zehnder, A.J.B.; Eggen, R.I.L. Structure of a Glutathione Peroxidase Homologous Gene Involved in the Oxidative Stress Response in Chlamydomonas reinhardtii. Plant. Sci. 1999, 149, 139–149. [Google Scholar] [CrossRef]
- Takeda, T.; Miyao, K.; Tamoi, M.; Kanaboshi, H.; Miyasaka, H.; Shigeoka, S. Molecular Characterization of Glutathione Peroxidase-like Protein in Halotolerant Chlamydomonas sp. W80. Physiol. Plant. 2003, 117, 467–475. [Google Scholar] [CrossRef]
- Ding, Y.; Liu, Y.; Jian, J.-C.; Wu, Z.-H.; Miao, J.-L. Molecular Cloning and Expression Analysis of Glutathione Reductase Gene in Chlamydomonas sp. ICE-L from Antarctica. Mar. Genom. 2012, 5, 59–64. [Google Scholar] [CrossRef]
- Peng, Y.; Ding, Y.; Tang, X.; Jian, J.; Wang, J.; Wang, Q. Characteristic of Glutamate Cysteine Ligase Gene and Its Response to the Salinity and Temperature Stress in Chlamydomonas sp. ICE-L from Antarctica. Turk. J. Bot. 2018, 42, 371–381. [Google Scholar] [CrossRef]
- Huang, L.-F.; Lin, J.-Y.; Pan, K.-Y.; Huang, C.-K.; Chu, Y.-K. Overexpressing Ferredoxins in Chlamydomonas reinhardtii Increase Starch and Oil Yields and Enhance Electric Power Production in a Photo Microbial Fuel Cell. Int. J. Mol. Sci. 2015, 16, 19308–19325. [Google Scholar] [CrossRef]
- Tanaka, S.; Ikeda, K.; Miyasaka, H. Enhanced Tolerance Against Salt-Stress and Freezing-Stress of Escherichia coli Cells Expressing Algal Bbc1 Gene. Curr. Microbiol. 2001, 42, 173–177. [Google Scholar] [CrossRef]
- Tanaka, S.; Ikeda, K.; Miyasaka, H. Isolation of a New Member of Group 3 Late Embryogenesis Abundant Protein Gene from a Halotorelant Green Alga by a Functional Expression Screening with Cyanobacterial Cells. FEMS Microbiol. Lett. 2004, 236, 41–45. [Google Scholar] [CrossRef][Green Version]
- Liu, S.; Zhang, P.; Cong, B.; Liu, C.; Lin, X.; Shen, J.; Huang, X. Molecular Cloning and Expression Analysis of a Cytosolic Hsp70 Gene from Antarctic Ice Algae Chlamydomonas sp. ICE-L. Extremophiles 2010, 14, 329–337. [Google Scholar] [CrossRef]
- Herrera-Valencia, V.A.; Macario-González, L.A.; Casais-Molina, M.L.; Beltran-Aguilar, A.G.; Peraza-Echeverría, S. In Silico Cloning and Characterization of the Glycerol-3-Phosphate Dehydrogenase (GPDH) Gene Family in the Green Microalga Chlamydomonas reinhardtii. Curr. Microbiol. 2012, 64, 477–485. [Google Scholar] [CrossRef]
- Casais-Molina, M.L.; Peraza-Echeverria, S.; Echevarría-Machado, I.; Herrera-Valencia, V.A. Expression of Chlamydomonas reinhardtii CrGPDH2 and CrGPDH3 CDNAs in Yeast Reveals That They Encode Functional Glycerol-3-Phosphate Dehydrogenases Involved in Glycerol Production and Osmotic Stress Tolerance. J. Appl. Phycol. 2016, 28, 219–226. [Google Scholar] [CrossRef]
- Driver, T.; Trivedi, D.K.; McIntosh, O.A.; Dean, A.P.; Goodacre, R.; Pittman, J.K. Two Glycerol-3-Phosphate Dehydrogenases from Chlamydomonas Have Distinct Roles in Lipid Metabolism. Plant. Physiol. 2017, 174, 2083–2097. [Google Scholar] [CrossRef]
- Morales-Sánchez, D.; Kim, Y.; Terng, E.L.; Peterson, L.; Cerutti, H. A Multidomain Enzyme, with Glycerol-3-Phosphate Dehydrogenase and Phosphatase Activities, Is Involved in a Chloroplastic Pathway for Glycerol Synthesis in Chlamydomonas reinhardtii. Plant. J. 2017, 90, 1079–1092. [Google Scholar] [CrossRef]
- Raymond, J.A.; Morgan-Kiss, R.; Stahl-Rommel, S. Glycerol Is an Osmoprotectant in Two Antarctic Chlamydomonas Species From an Ice-Covered Saline Lake and Is Synthesized by an Unusual Bidomain Enzyme. Front. Plant. Sci. 2020, 11, 1259. [Google Scholar] [CrossRef]
- Miyasaka, H.; Ogata, T.; Tanaka, S.; Ohama, T.; Kano, S.; Kazuhiro, F.; Hayashi, S.; Yamamoto, S.; Takahashi, H.; Matsuura, H.; et al. Is Chloroplastic Class IIA Aldolase a Marine Enzyme? ISME J. 2016, 10, 2767–2772. [Google Scholar] [CrossRef]
- Ho, S.-H.; Nakanishi, A.; Kato, Y.; Yamasaki, H.; Chang, J.-S.; Misawa, N.; Hirose, Y.; Minagawa, J.; Hasunuma, T.; Kondo, A. Dynamic Metabolic Profiling Together with Transcription Analysis Reveals Salinity-Induced Starch-to-Lipid Biosynthesis in Alga Chlamydomonas sp. JSC4. Sci. Rep. 2017, 7, 45471. [Google Scholar] [CrossRef]
- Colina, F.; Amaral, J.; Carbó, M.; Pinto, G.; Soares, A.; Cañal, M.J.; Valledor, L. Genome-Wide Identification and Characterization of CKIN/SnRK Gene Family in Chlamydomonas reinhardtii. Sci. Rep. 2019, 9, 350. [Google Scholar] [CrossRef]
- He, Y.; Wang, Y.; Zheng, Z.; Liu, F.; An, M.; He, X.; Qu, C.; Li, L.; Miao, J. Cloning and Stress-Induced Expression Analysis of Calmodulin in the Antarctic Alga Chlamydomonas sp. ICE-L. Curr. Microbiol. 2017, 74, 921–929. [Google Scholar] [CrossRef]
- Li, Y.; Fei, X.; Wu, X.; Deng, X. Iron Deficiency Response Gene Femu2 Plays a Positive Role in Protecting Chlamydomonas reinhardtii against Salt Stress. Biochim. Biophys. Acta Gen. Subj. 2017, 1861, 3345–3354. [Google Scholar] [CrossRef]
- Tanaka, S.; Suda, Y.; Ikeda, K.; Ono, M.; Miyasaka, H.; Watanabe, M.; Sasaki, K.; Hirata, K. A Novel Gene with Antisalt and Anticadmium Stress Activities from a Halotolerant Marine Green Alga Chlamydomonas sp. W80. FEMS Microbiol. Lett. 2007, 271, 48–52. [Google Scholar] [CrossRef][Green Version]
- Suda, Y.; Yoshikawa, T.; Okuda, Y.; Tsunemoto, M.; Tanaka, S.; Ikeda, K.; Miyasaka, H.; Watanabe, M.; Sasaki, K.; Harada, K.; et al. Isolation and Characterization of a Novel Antistress Gene from Chlamydomonas sp. W80. J. Biosci. Bioeng. 2009, 107, 352–354. [Google Scholar] [CrossRef]
- Ishinishi, R.; Matsuura, H.; Tanaka, S.; Nozawa, S.; Tanada, K.; Kawashita, N.; Fujiyama, K.; Miyasaka, H.; Hirata, K. Isolation and Characterization of a Stress-Responsive Gene Encoding a CHRD Domain-Containing Protein from a Halotolerant Green Alga. Gene 2018, 640, 14–20. [Google Scholar] [CrossRef]
- Wang, Y.; Liu, X.; Gao, H.; Zhang, H.-M.; Guo, A.-Y.; Xu, J.; Xu, X. Early Stage Adaptation of a Mesophilic Green Alga to Antarctica: Systematic Increases in Abundance of Enzymes and LEA Proteins. Mol. Biol. Evol. 2020, 37, 849–863. [Google Scholar] [CrossRef]
- Chen, X.; Tian, D.; Kong, X.; Chen, Q.; Abd Allah, E.F.; Hu, X.; Jia, A. The Role of Nitric Oxide Signalling in Response to Salt Stress in Chlamydomonas reinhardtii. Planta 2016, 244, 651–669. [Google Scholar] [CrossRef]
- Shigeoka, S.; Takeda, T.; Hanaoka, T. Characterization and Immunological Properties of Selenium-Containing Glutathione Peroxidase Induced by Selenite in Chlamydomonas reinhardtii. Biochem. J. 1991, 275, 623–627. [Google Scholar] [CrossRef]
- Kalra, I.; Wang, X.; Cvetkovska, M.; Jeong, J.; McHargue, W.; Zhang, R.; Hüner, N.; Yuan, J.S.; Morgan-Kiss, R. Chlamydomonas sp. UWO 241 Exhibits High Cyclic Electron Flow and Rewired Metabolism under High Salinity. Plant. Physiol. 2020, 183, 588–601. [Google Scholar] [CrossRef]
- Sithtisarn, S.; Yokthongwattana, K.; Mahong, B.; Roytrakul, S.; Paemanee, A.; Phaonakrop, N.; Yokthongwattana, C. Comparative Proteomic Analysis of Chlamydomonas reinhardtii Control and a Salinity-Tolerant Strain Revealed a Differential Protein Expression Pattern. Planta 2017, 246, 843–856. [Google Scholar] [CrossRef]
- Perrineau, M.-M.; Zelzion, E.; Gross, J.; Price, D.C.; Boyd, J.; Bhattacharya, D. Evolution of Salt Tolerance in a Laboratory Reared Population of Chlamydomonas reinhardtii. Environ. Microbiol. 2014, 16, 1755–1766. [Google Scholar] [CrossRef]
- Zheng, Z.; Miao, J.; Kan, G.; Jin, Q.; Ding, Y.; Liu, F.; Wang, S.; Wang, Y. Effect of High Salinity on Cell Growth and Protein Production of Antarctic Ice Microalgae Chlamydomonas sp. ICE-L. Chin. J. Polar Sci. 2010, 21, 81–90. [Google Scholar]
- Mastrobuoni, G.; Irgang, S.; Pietzke, M.; Aßmus, H.E.; Wenzel, M.; Schulze, W.X.; Kempa, S. Proteome Dynamics and Early Salt Stress Response of the Photosynthetic Organism Chlamydomonas reinhardtii. BMC Genom. 2012, 13, 215. [Google Scholar] [CrossRef]
- Yokthongwattana, C.; Mahong, B.; Roytrakul, S.; Phaonaklop, N.; Narangajavana, J.; Yokthongwattana, K. Proteomic Analysis of Salinity-Stressed Chlamydomonas reinhardtii Revealed Differential Suppression and Induction of a Large Number of Important Housekeeping Proteins. Planta 2012, 235, 649–659. [Google Scholar] [CrossRef]
- Lu, N.; Wei, D.; Chen, F.; Yang, S.-T. Lipidomic Profiling and Discovery of Lipid Biomarkers in Snow Alga Chlamydomonas nivalis under Salt Stress. Eur. J. Lipid Sci. Technol. 2012, 114, 253–265. [Google Scholar] [CrossRef]
- Lu, N.; Wei, D.; Jiang, X.-L.; Chen, F.; Yang, S.-T. Regulation of Lipid Metabolism in the Snow Alga Chlamydomonas nivalis in Response to NaCl Stress: An Integrated Analysis by Cytomic and Lipidomic Approaches. Process. Biochem. 2012, 47, 1163–1170. [Google Scholar] [CrossRef]
- Lu, N.; Wei, D.; Jiang, X.-L.; Chen, F.; Yang, S.-T. Fatty Acids Profiling and Biomarker Identification in Snow Alga Chlamydomonas nivalis by NaCl Stress Using GC/MS and Multivariate Statistical Analysis. Anal. Lett. 2012, 45, 1172–1183. [Google Scholar] [CrossRef]
- Ves-urai, P.; Krobthong, S.; Thongsuk, K.; Roytrakul, S.; Yokthongwattana, C. Comparative Secretome Analysis between Salinity-Tolerant and Control Chlamydomonas reinhardtii Strains. Planta 2021, 253, 68. [Google Scholar] [CrossRef]
- Siaut, M.; Cuiné, S.; Cagnon, C.; Fessler, B.; Nguyen, M.; Carrier, P.; Beyly, A.; Beisson, F.; Triantaphylidès, C.; Li-Beisson, Y.; et al. Oil Accumulation in the Model Green Alga Chlamydomonas reinhardtii: Characterization, Variability between Common Laboratory Strains and Relationship with Starch Reserves. BMC Biotechnol. 2011, 11, 7. [Google Scholar] [CrossRef]
- Lauritano, C.; Ianora, A. Marine Organisms with Anti-Diabetes Properties. Mar. Drugs 2016, 14, 220. [Google Scholar] [CrossRef]
- Romano, G.; Costantini, M.; Sansone, C.; Lauritano, C.; Ruocco, N.; Ianora, A. Marine Microorganisms as a Promising and Sustainable Source of Bioactive Molecules. Mar. Environ. Res. 2017, 128, 58–69. [Google Scholar] [CrossRef]
- Lauritano, C.; Martín, J.; de la Cruz, M.; Reyes, F.; Romano, G.; Ianora, A. First Identification of Marine Diatoms with Anti-Tuberculosis Activity. Sci. Rep. 2018, 8, 2284. [Google Scholar] [CrossRef]
- Brillatz, T.; Lauritano, C.; Jacmin, M.; Khamma, S.; Marcourt, L.; Righi, D.; Romano, G.; Esposito, F.; Ianora, A.; Queiroz, E.F.; et al. Zebrafish-Based Identification of the Antiseizure Nucleoside Inosine from the Marine Diatom Skeletonema marinoi. PLoS ONE 2018, 13, e0196195. [Google Scholar] [CrossRef]
- Giordano, D.; Costantini, M.; Coppola, D.; Lauritano, C.; Núñez Pons, L.; Ruocco, N.; di Prisco, G.; Ianora, A.; Verde, C. Biotechnological Applications of Bioactive Peptides From Marine Sources. In Advances in Microbial Physiology; Elsevier: Amsterdam, The Netherlands, 2018; Volume 73, pp. 171–220. ISBN 978-0-12-815190-7. [Google Scholar]
- Martínez, K.A.; Lauritano, C.; Druka, D.; Romano, G.; Grohmann, T.; Jaspars, M.; Martín, J.; Díaz, C.; Cautain, B.; de la Cruz, M.; et al. Amphidinol 22, a New Cytotoxic and Antifungal Amphidinol from the Dinoflagellate Amphidinium carterae. Mar. Drugs 2019, 17, 385. [Google Scholar] [CrossRef]
- Vingiani, G.M.; De Luca, P.; Ianora, A.; Dobson, A.D.W.; Lauritano, C. Microalgal Enzymes with Biotechnological Applications. Mar. Drugs 2019, 17, 459. [Google Scholar] [CrossRef]
- Riccio, G.; Lauritano, C. Microalgae with Immunomodulatory Activities. Mar. Drugs 2020, 18, 2. [Google Scholar] [CrossRef]
- Lauritano, C.; Helland, K.; Riccio, G.; Andersen, J.H.; Ianora, A.; Hansen, E.H. Lysophosphatidylcholines and Chlorophyll-Derived Molecules from the Diatom Cylindrotheca closterium with Anti-Inflammatory Activity. Mar. Drugs 2020, 18, 166. [Google Scholar] [CrossRef]
- Saide, A.; Martínez, K.A.; Ianora, A.; Lauritano, C. Unlocking the Health Potential of Microalgae as Sustainable Sources of Bioactive Compounds. Int. J. Mol. Sci. 2021, 22, 4383. [Google Scholar] [CrossRef]
- Saide, A.; Lauritano, C.; Ianora, A. Pheophorbide a: State of the Art. Mar. Drugs 2020, 18, 257. [Google Scholar] [CrossRef]
- Riccio, G.; Ruocco, N.; Mutalipassi, M.; Costantini, M.; Zupo, V.; Coppola, D.; de Pascale, D.; Lauritano, C. Ten-Year Research Update Review: Antiviral Activities from Marine Organisms. Biomolecules 2020, 10, 1007. [Google Scholar] [CrossRef]
- Riccio, G.; De Luca, D.; Lauritano, C. Monogalactosyldiacylglycerol and Sulfolipid Synthesis in Microalgae. Mar. Drugs 2020, 18, 237. [Google Scholar] [CrossRef]
- Lauritano, C.; Andersen, J.H.; Hansen, E.; Albrigtsen, M.; Escalera, L.; Esposito, F.; Helland, K.; Hanssen, K.Ø.; Romano, G.; Ianora, A. Bioactivity Screening of Microalgae for Antioxidant, Anti-Inflammatory, Anticancer, Anti-Diabetes, and Antibacterial Activities. Front. Mar. Sci. 2016, 3, 68. [Google Scholar] [CrossRef]
- Ingebrigtsen, R.A.; Hansen, E.; Andersen, J.H.; Eilertsen, H.C. Light and Temperature Effects on Bioactivity in Diatoms. J. Appl. Phycol. 2016, 28, 939–950. [Google Scholar] [CrossRef]
- Fu, W.; Chaiboonchoe, A.; Khraiwesh, B.; Nelson, D.R.; Al-Khairy, D.; Mystikou, A.; Alzahmi, A.; Salehi-Ashtiani, K. Algal Cell Factories: Approaches, Applications, and Potentials. Mar. Drugs 2016, 14, 225. [Google Scholar] [CrossRef]
- Hounslow, E.; Kapoore, R.V.; Vaidyanathan, S.; Gilmour, D.J.; Wright, P.C. The Search for a Lipid Trigger: The Effect of Salt Stress on the Lipid Profile of the Model Microalgal Species Chlamydomonas reinhardtii for Biofuels Production. Curr. Biotechnol. 2016, 5, 305–313. [Google Scholar] [CrossRef]
- Kato, Y.; Ho, S.-H.; Vavricka, C.J.; Chang, J.-S.; Hasunuma, T.; Kondo, A. Evolutionary Engineering of Salt-Resistant Chlamydomonas Sp. Strains Reveals Salinity Stress-Activated Starch-to-Lipid Biosynthesis Switching. Bioresour. Technol. 2017, 245, 1484–1490. [Google Scholar] [CrossRef]
- Dragosits, M.; Mattanovich, D. Adaptive Laboratory Evolution–Principles and Applications for Biotechnology. Microb. Cell Factories 2013, 12, 64. [Google Scholar] [CrossRef]
- Sun, X.-M.; Ren, L.-J.; Zhao, Q.-Y.; Ji, X.-J.; Huang, H. Microalgae for the Production of Lipid and Carotenoids: A Review with Focus on Stress Regulation and Adaptation. Biotechnol. Biofuels 2018, 11, 272. [Google Scholar] [CrossRef]
- Velmurugan, N.; Sung, M.; Yim, S.S.; Park, M.S.; Yang, J.W.; Jeong, K.J. Systematically Programmed Adaptive Evolution Reveals Potential Role of Carbon and Nitrogen Pathways during Lipid Accumulation in Chlamydomonas reinhardtii. Biotechnol. Biofuels 2014, 7, 117. [Google Scholar] [CrossRef]
- Yu, S.; Zhao, Q.; Miao, X.; Shi, J. Enhancement of Lipid Production in Low-Starch Mutants Chlamydomonas reinhardtii by Adaptive Laboratory Evolution. Bioresour. Technol. 2013, 147, 499–507. [Google Scholar] [CrossRef]
- Satoh, K.; Oono, Y. Studies on Application of Ion Beam Breeding to Industrial Microorganisms at TIARA. Quantum Beam Sci. 2019, 3, 11. [Google Scholar] [CrossRef]
- Lee, B.; Choi, G.-G.; Choi, Y.-E.; Sung, M.; Park, M.S.; Yang, J.-W. Enhancement of Lipid Productivity by Ethyl Methane Sulfonate-Mediated Random Mutagenesis and Proteomic Analysis in Chlamydomonas reinhardtii. Korean J. Chem. Eng. 2014, 31, 1036–1042. [Google Scholar] [CrossRef]
- Abdrabu, R.; Sharma, S.; Khraiwesh, B.; Jijakli, K.; Nelson, D.; Alzahmi, A.; Koussa, J.; Sultana, M.; Khapli, S.; Jagannathan, R.; et al. Single-Cell Characterization of Microalgal Lipid Contents with Confocal Raman Microscopy; Springer: Berlin/Heidelberg, Germany, 2016; pp. 363–382. ISBN 978-3-662-49118-8. [Google Scholar]
- Sharma, S.K.; Nelson, D.R.; Abdrabu, R.; Khraiwesh, B.; Jijakli, K.; Arnoux, M.; O’Connor, M.J.; Bahmani, T.; Cai, H.; Khapli, S.; et al. An Integrative Raman Microscopy-Based Workflow for Rapid in Situ Analysis of Microalgal Lipid Bodies. Biotechnol. Biofuels 2015, 8, 164. [Google Scholar] [CrossRef]
- Darwish, R.; Gedi, M.A.; Akepach, P.; Assaye, H.; Zaky, A.S.; Gray, D.A. Chlamydomonas reinhardtii Is a Potential Food Supplement with the Capacity to Outperform Chlorella and Spirulina. Appl. Sci. 2020, 10, 6736. [Google Scholar] [CrossRef]
- Rasala, B.A.; Muto, M.; Lee, P.A.; Jager, M.; Cardoso, R.M.F.; Behnke, C.A.; Kirk, P.; Hokanson, C.A.; Crea, R.; Mendez, M.; et al. Production of Therapeutic Proteins in Algae, Analysis of Expression of Seven Human Proteins in the Chloroplast of Chlamydomonas reinhardtii. Plant. Biotechnol. J. 2010, 8, 719–733. [Google Scholar] [CrossRef]
- Baltz, A.; Dang, K.-V.; Beyly, A.; Auroy, P.; Richaud, P.; Cournac, L.; Peltier, G. Plastidial Expression of Type II NAD(P)H Dehydrogenase Increases the Reducing State of Plastoquinones and Hydrogen Photoproduction Rate by the Indirect Pathway in Chlamydomonas reinhardtii. Plant. Physiol. 2014, 165, 1344–1352. [Google Scholar] [CrossRef]
- Boynton, J.E.; Gillham, N.W.; Harris, E.H.; Hosler, J.P.; Johnson, A.M.; Jones, A.R.; Randolph-Anderson, B.L.; Robertson, D.; Klein, T.M.; Shark, K.B. Chloroplast Transformation in Chlamydomonas with High Velocity Microprojectiles. Science 1988, 240, 1534–1538. [Google Scholar] [CrossRef]
- Blowers, A.D.; Bogorad, L.; Shark, K.B.; Sanford, J.C. Studies on Chlamydomonas Chloroplast Transformation: Foreign DNA Can Be Stably Maintained in the Chromosome. Plant. Cell 1989, 1, 123–132. [Google Scholar] [CrossRef]
- Kindle, K.L.; Schnell, R.A.; Fernández, E.; Lefebvre, P.A. Stable Nuclear Transformation of Chlamydomonas Using the Chlamydomonas Gene for Nitrate Reductase. J. Cell Biol. 1989, 109, 2589–2601. [Google Scholar] [CrossRef]
- Randolph-Anderson, B.L.; Boynton, J.E.; Gillham, N.W.; Harris, E.H.; Johnson, A.M.; Dorthu, M.-P.; Matagne, R.F. Further Characterization of the Respiratory Deficient Dum-1 Mutation of Chlamydomonas reinhardtii and Its Use as a Recipient for Mitochondrial Transformation. Mol. Gen. Genet. MGG 1993, 236, 235–244. [Google Scholar] [CrossRef]
- Mayfield, S.P.; Franklin, S.E.; Lerner, R.A. Expression and Assembly of a Fully Active Antibody in Algae. Proc. Natl. Acad. Sci. USA 2003, 100, 438–442. [Google Scholar] [CrossRef]
- Sizova, I.; Greiner, A.; Awasthi, M.; Kateriya, S.; Hegemann, P. Nuclear Gene Targeting in Chlamydomonas Using Engineered Zinc-Finger Nucleases. Plant. J. 2013, 73, 873–882. [Google Scholar] [CrossRef]
- Gao, H.; Wright, D.A.; Li, T.; Wang, Y.; Horken, K.; Weeks, D.P.; Yang, B.; Spalding, M.H. TALE Activation of Endogenous Genes in Chlamydomonas reinhardtii. Algal Res. 2014, 5, 52–60. [Google Scholar] [CrossRef]
- Jiang, W.; Brueggeman, A.J.; Horken, K.M.; Plucinak, T.M.; Weeks, D.P. Successful Transient Expression of Cas9 and Single Guide RNA Genes in Chlamydomonas reinhardtii. Eukaryot. Cell 2014, 13, 1465–1469. [Google Scholar] [CrossRef]
- Ng, I.-S.; Tan, S.-I.; Kao, P.-H.; Chang, Y.-K.; Chang, J.-S. Recent Developments on Genetic Engineering of Microalgae for Biofuels and Bio-Based Chemicals. Biotechnol. J. 2017, 12, 1600644. [Google Scholar] [CrossRef]
- Kao, P.-H.; Ng, I.-S. CRISPRi Mediated Phosphoenolpyruvate Carboxylase Regulation to Enhance the Production of Lipid in Chlamydomonas reinhardtii. Bioresour. Technol. 2017, 245, 1527–1537. [Google Scholar] [CrossRef]
- Baek, K.; Kim, D.H.; Jeong, J.; Sim, S.J.; Melis, A.; Kim, J.-S.; Jin, E.; Bae, S. DNA-Free Two-Gene Knockout in Chlamydomonas reinhardtii via CRISPR-Cas9 Ribonucleoproteins. Sci. Rep. 2016, 6, 30620. [Google Scholar] [CrossRef]
- Shin, S.-E.; Lim, J.-M.; Koh, H.G.; Kim, E.K.; Kang, N.K.; Jeon, S.; Kwon, S.; Shin, W.-S.; Lee, B.; Hwangbo, K.; et al. CRISPR/Cas9-Induced Knockout and Knock-in Mutations in Chlamydomonas reinhardtii. Sci. Rep. 2016, 6, 27810. [Google Scholar] [CrossRef]
- Zhang, R.; Patena, W.; Armbruster, U.; Gang, S.S.; Blum, S.R.; Jonikas, M.C. High-Throughput Genotyping of Green Algal Mutants Reveals Random Distribution of Mutagenic Insertion Sites and Endonucleolytic Cleavage of Transforming DNA. Plant. Cell 2014, 26, 1398–1409. [Google Scholar] [CrossRef]
- Scaife, M.A.; Nguyen, G.T.D.T.; Rico, J.; Lambert, D.; Helliwell, K.E.; Smith, A.G. Establishing Chlamydomonas reinhardtii as an Industrial Biotechnology Host. Plant. J. 2015, 82, 532–546. [Google Scholar] [CrossRef]
- Lauritano, C.; Ferrante, M.I.; Rogato, A. Marine Natural Products from Microalgae: An -Omics Overview. Mar. Drugs 2019, 17, 269. [Google Scholar] [CrossRef]
- Lauritano, C.; De Luca, D.; Ferrarini, A.; Avanzato, C.; Minio, A.; Esposito, F.; Ianora, A. De Novo Transcriptome of the Cosmopolitan Dinoflagellate Amphidinium carterae to Identify Enzymes with Biotechnological Potential. Sci. Rep. 2017, 7, 11701. [Google Scholar] [CrossRef]
- Lauritano, C.; De Luca, D.; Amoroso, M.; Benfatto, S.; Maestri, S.; Racioppi, C.; Esposito, F.; Ianora, A. New Molecular Insights on the Response of the Green Alga Tetraselmis suecica to Nitrogen Starvation. Sci. Rep. 2019, 9, 3336. [Google Scholar] [CrossRef]
- Di Dato, V.; Di Costanzo, F.; Barbarinaldi, R.; Perna, A.; Ianora, A.; Romano, G. Unveiling the Presence of Biosynthetic Pathways for Bioactive Compounds in the Thalassiosira rotula Transcriptome. Sci. Rep. 2019, 9, 9893. [Google Scholar] [CrossRef]
- Elagoz, A.M.; Ambrosino, L.; Lauritano, C. De Novo Transcriptome of the Diatom Cylindrotheca closterium Identifies Genes Involved in the Metabolism of Anti-Inflammatory Compounds. Sci. Rep. 2020, 10, 1–9. [Google Scholar] [CrossRef]
- De Luca, D.; Lauritano, C. In Silico Identification of Type III PKS Chalcone and Stilbene Synthase Homologs in Marine Photosynthetic Organisms. Biology 2020, 9, 110. [Google Scholar] [CrossRef]
- Vingiani, G.M.; Štālberga, D.; De Luca, P.; Ianora, A.; De Luca, D.; Lauritano, C. De Novo Transcriptome of the Non-Saxitoxin Producing Alexandrium tamutum Reveals New Insights on Harmful Dinoflagellates. Mar. Drugs 2020, 18, 386. [Google Scholar] [CrossRef]
- Saggiomo, M.; Escalera, L.; Saggiomo, V.; Bolinesi, F.; Mangoni, O. Phytoplankton Blooms Below the Antarctic Landfast Ice During the Melt Season Between Late Spring and Early Summer. J. Phycol. 2021, 57, 541–550. [Google Scholar] [CrossRef]
- Corwin, D.L. Climate Change Impacts on Soil Salinity in Agricultural Areas. Eur. J. Soil Sci. 2021, 72, 842–862. [Google Scholar] [CrossRef]
- Falciatore, A.; Jaubert, M.; Bouly, J.-P.; Bailleul, B.; Mock, T. Diatom Molecular Research Comes of Age: Model Species for Studying Phytoplankton Biology and Diversity. Plant. Cell 2020, 32, 547–572. [Google Scholar] [CrossRef]
| Chlamydomonas sp. | Dunaliella salina | |
|---|---|---|
| Growth | Reduced | Reduced |
| Size | Variable | Variable/increased |
| Palmelloidy | Yes | No |
| Main osmolyte | Glycerol | Glycerol |
| Starch degradation | Increased | Increased |
| Antioxidant system | Enhanced | Enhanced |
| Photosynthesis | Decreased efficiency | Decreased efficiency at high stress levels |
| Pigments | Increased carotenoid content | Highly increased carotenoid content |
| Genes | Function | Species | Salinity Range | Exposure Time | Methods | Expression | Ref |
|---|---|---|---|---|---|---|---|
| Stress-related and detoxification genes | |||||||
| Anti-stress genes | Anti-stress | Chlamydomonas W80 | 500–1500 mM NaCl 5–6% in E. coli coltures | Growth time 3 days in E. coli | cDNA library functional expression screening | Conferred salt resistance to transformed E. coli colonies | [57] |
| Ascorbate peroxidase (APX) | Antioxidant | Chlamydomonas W80 | 1–7% in E. coli coltures | Growth time 3 days in E. coli | Expression of the recombinant APX in E. coli | Enhanced salt tolerance in transformed E. coli cells NaCl is needed for the expression of Chlamydomonas W80 APX in E. coli | [58] |
| Superoxide dismutase (SOD) Catalase (CAT) Ascorbate peroxidase (APX) | Antioxidant | C. reinhardtii IAM C-238 | 100 mM NaCl | 3–24 h | RT-PCR | Enhanced expression of CAT and SOD | [37] |
| Super oxide dismutase (Mn-SOD) catalase (CAT) ascorbate peroxidase (APX) | Antioxidant | C. reinhardtii | 200 mM NaCl | 1–24 h | RT-PCR | Enhanced expression of APX, CAT, and MnSOD | [23] |
| Glutathione peroxidase (gpxh) | Antioxidant | C. reinhardtii CC-325 | 200 mM NaCl | 0,5–3 h | Northern Blot | Weak enhancement of gpxh expression | [59] |
| Glutathione peroxidase-like protein (GPX-like) | Antioxidant | Chlamydomonas W80 | 1–5% in E. coli coltures | 10 h | Expression of the recombinant GPX-like protein in E. coli | Enhanced salt tolerance in transformed E. coli cells | [60] |
| 250 mM NaCl in N. tabacum | 24 h | Expression of the recombinant GPX-like protein in N. tabacum | Enhanced salt tolerance in transgenic tobacco plants | [13] | |||
| Glutathione peroxidase (ICE-LGPX) | Antioxidant | Chlamydomonas sp. ICE-L | 11‰–99‰ | 6–72 h | RT-qPCR | Overexpression under low and high salinity: ≈4-fold peak at 11‰ and 66‰ after 24 h ≈2-fold peak at 22‰ and 99‰ after 12 h | [32] |
| Glutathione reductase (ICE_LGR) | Antioxidant | Chlamydomonas sp. ICE-L | 11‰–99‰ | 6–96 h | RT-qPCR | Initial underexpression, subsequent overexpression under low and high salinity: >6-fold peak after 24 h at 11‰ ≈3-fold peak after 24 h at 22‰ ≈1.5-fold peak after 24 h at 66‰ ≈1.5-fold peak after 12 h at 99‰ | [61] |
| Glutamate cysteine ligase (ICE-LGCL) | Reduced glutathione (GSH) synthesis | Chlamydomonas sp. ICE-L | 11‰–99‰ | 6–72 h | RT-qPCR | Overexpression under low salinities: >2-fold peak after 48 h at 11‰ and 22‰ Underexpression under high salinities: <0.5-fold peak at 66‰ and 99‰ | [62] |
| Ferredoxins (PETF, FDX5) | Electron donors in the photosynthetic pathway and antioxidant system | C. reinhardtii CC125 | 120–240 mM NaCl | 12 h | Overexpression in transgenic Chlamydomonas | Overexpression of PETF and FDX5 enhances salt tolerance and starch and lipid production | [63] |
| Breast basic conserved (bbc1) | Protection against dehydration | Chlamydomonas W80 | 1–7% in E. coli coltures | 3 days | Expression of the recombinant BBC1 in E. coli | Enhanced salt tolerance in transformed E. coli cells | [64] |
| Group 3 late embryogenesis abundant (cw80lea3) | LEA-like protection against dehydration | Chlamydomonas W80 | 500–1500 mM NaCl 0.5–3% in S. PCC7942 | 6–24 h 7 days in in S. PCC7942 | cDNA library functional expression screening Northern blotting | Conferred salt resistance to transformed S. PCC7942 colonies Overexpression after 6 h Lowered expression after 24 h | [65] |
| Heat shock protein 70 (CiHsp70) | Molecular chaperon | Chlamydomonas sp. ICE-L | 31‰–93‰ | 2–36 h | RT-qPCR | Overexpression: 3-fold peak after 2 h at 66‰ ≈2.-fold peak after 2 h at 93‰ Expression levels gradually decreased over time | [66] |
| Stress-related members of the light-harvesting complex protein family (LhcSR1, LhcSR2) | Potential photoprotective role during stress | Chlamydomonas sp. ICE-L | 93‰ | 1–24 h | RT-qPCR | Overexpression peak of LhcSR1 and LhcSR2 at 15.68- and 12.72-fold, respectively, after 2 h. Gradual decrease after | [39] |
| Osmolytes or lipid synthesis-related genes | |||||||
| Glycerol-3-phosphate dehydrogenase (CrGPDH1, CrGPDH2, CrGPDH3) | G3P synthesis: glycerol and lipid precursor | C. reinhardtii CC-125 | 200 mM NaCl | 120 min | RT-PCR | Enhanced expression of CrGPDH2, CrGPDH3 | [67] |
| Glycerol-3-phosphate dehydrogenase (CrGPDH2, CrGPDH3) | G3P synthesis | C. reinhardtii CC-125 | 5–200 mM NaCl 200–800 mM NaCl in yeast | 5–120 min 4 h–4 days in yeast | RT-PCR Functional complementation of a gpdh-lacking yeast mutant | Enhanced expression of CrGPDH2, CrGPDH3 at all conditions Genetic complementation restored salt resistance and glycerol production | [68] |
| Glycerol-3-phosphate dehydrogenase (GPD1–5) | G3P synthesis | C. reinhardtii cw15 | 200 mM NaCl 700–1000 mM NaCl in yeast | 2 h 24 h in yeast | RT-qPCR Functional complementation of a gpdh1-lacking yeast mutant | GPD1 and GPD5 are constitutively expressed GPD2 is up-regulated under salt stress. GPD3 and GPD4 are down-regulated under salt stress. Functional complementation partly restored salt resistance and glycerol production | [69] |
| Genes encoding enzymes of glycerol metabolism | Glycerol, G3P and DHAP synthesis | C. reinhardtii CC-124 and CC-125 | 100 mM NaCl | 6 h | RT-PCR and RT-qPCR RNAi silencing of GPD2 and GPD3 | Up-regulation of GPD2 and glycerol kinase in the wild-type strain Reduced glycerol and TAGs accumulation under salinity in the RNAi strains | [70] |
| Glycerol-3-phosphate dehydrogenase (PSP-GPDH isoform 2) | G3P synthesis | Chlamydomonas sp. UWO241 | 10–1300 mM NaCl | Growth until mid-exponential phase | RT-qPCR | Overexpression: ≈ 4-fold peak. Dose-dependent increase. | [71] |
| Fructose-1, 6-bisphosphate aldolase (FBA) | Key enzyme in glucose metabolism and Calvin–Benson cycle | Chlamydomonas W80 | 50–500 mM NaCl | 72 h | RT-qPCR | Up-regulation of class I FBA at 50–75 mM NaCl Down-regulation of class IIA FBA at 50 mM NaCl | [72] |
| Fatty acid desaturases (Δ9ACPCiFAD, Δ6CiFAD, ω3CiFAD1, Δ12CiFAD, ω3CiFAD2) | Fatty acid desaturation | Chlamydomonas sp. ICE-L | 16‰–128‰ | 14 days | RT-qPCR | Overexpression of all genes with different patterns: Δ6CiFAD, Δ12CiFAD, ω3CiFAD2 expression increased with time Δ9ACPCiFAD overexpression is higher the first 2–4 days ω3CiFAD1 overexpression under high salinities | [17] |
| Lipid and starch metabolism—related genes | Starch synthesis, starch degradation, and lipid synthesis | Chlamydomonas sp. JSC4 | 2% Sea Salt | 1–7 days | RT-qPCR | Underexpression of starch-synthesis-related genes Overexpression of starch-degradation and lipid-synthesis-related genes | [73] |
| Signaling and transcription regulation | |||||||
| Sucrose nonfermenting-related kinase (CKINs/SnRK) | Energy sensing | C. reinhardtii | 250 mM NaCl | 48 h | RT-qPCR | Up-regulation of all CKIN, except for CKIN2.14 | [74] |
| Calmodulin (CaM) | Calcium binding protein | Chlamydomonas sp. ICE-L | 32‰–128‰ NaCl | 2–48 h | RT-qPCR | Overexpression at high salinities: ≈3-fold peak at 96‰ after 24 h ≈3-fold peak at 128‰ after 12 h | [75] |
| Basic leucine-region zipper (CrebZIPs) | Transcription factor | C. reinhardtii | 150 mM NaCl | 6–48 h | RT-qPCR | Overexpression of CrebZIP10, 11, and 16 Underexpression of CrebZIP4, 5, and 13 No obvious expression changes in 11 CrebZIP | [42] |
| Iron deficiency related gene (Femu2) | Transcription factor | C. reinhardtii CC124 | 50–2000 mM NaCl | 2–72 h | RT-qPCR Overexpression RNAi silencing | Overexpression enhanced by ABA addition (30-fold peak at 150 mM) Overexpression enhances salt tolerance Silencing reduces salt tolerance | [76] |
| Others | |||||||
| Hyp-rich glycoproteins (GAS28, GAS30, GAS31) | Cell-wall constituent | C. reinhardtii CC-620 (mt+) and CC-124 (mt-) | 10–50 mM NaCl | 2 h | Northern Blot | Increased transcript levels | [56] |
| Salt and cadmium stress related gene (scsr) | Unknown | Chlamydomonas W80 | 1–7% NaCl in E. coli coltures | 20 h | Expression of scsr in E. coli | Enhanced salt tolerance in transformed E. coli cells | [77] |
| WCFII | putative subunit of ATP synthase | Chlamydomonas W80 | 500–1500 mM NaCl 6% in E. coli coltures | 1–2 days | cDNA library functional expression screening | Conferred salt resistance to transformed E. coli colonies | [78] |
| cluster58 (CL58) | Unknown | Chlamydomonas W80 | 500–1500 mM NaCl | 8 h | RT-qPCR | Transcript level almost unchanged under salt stress | [79] |
| Species | Approach | Salinity | Exposure Time | Reference |
|---|---|---|---|---|
| C. reinhardtii CC-503 | Transcriptomic | 200 mM NaCl | 48 h: short-term 1255 generations (ca. 17 months): long-term | [85] |
| C. reinhardtii GY-D55 | Transcriptomic | 200 mM NaCl | 24 h | [32] |
| C. reinhardtii CC124, OE-62, RNAi-2 | Transcriptomic | 100 mM NaCl | 24 h | [76] |
| Chlamydomonas sp. ICE-L | Genomic Transcriptomic | 32,7‰, 64.0‰, 96.7‰ | [19] | |
| Chlamydomonas sp. ICE-L | Proteomic (preliminary) | 33‰, 66‰, 99‰, 132‰, and 165‰ | 2, 4, 6, 8, 10, 12, 14, 16, and 18 days | [86] |
| C. reinhardtii CC- 1618 | Proteomic Metabolomic | 0, 100, 150 mM NaCl | 1, 3, 8, 24 h | [87] |
| C. reinhardtii CC-503 | Proteomic | 300 Mm NaCl | 2 h | [88] |
| C. reinhardtii | Proteomic | 100 mM NaCl | 1, 3, 5 days | [81] |
| C. reinhardtii CC-503 | Proteomic | 300 Mm NaCl | Several generations | [84] |
| C. reinhardtii CC125 | Proteomic (on spent media) | 100 mM NaCl | 24 h 1 h de-stress | [20] |
| Chlamydomonas sp. UWO 241 | Proteomic Metabolomic | 700 mM NaCl | Until midlog phase | [83] |
| C. reinhardtii CC-4325 sta1-1 mt-[Ball I7] C. nivalis | Proteomic | 200 mM NaCl | 11, 18 h (C. reinhardtii) 80, 168 h (C. nivalis) | [18] |
| C. nivalis | Metabolomic (lipidomic) | NaCl at 0%, 0.25%, 0.5%, and 1.0% | 1, 7, and 15 h, respectively | [89] |
| C. nivalis | Metabolomic | NaCl at 0%, 0.25%, 0.50%, 0.75%, 1.00%, 1.25%, and 1.50% | 1, 3, 5, 7, 11, 15, 24, 48 h, respectively | [90] |
| C. nivalis | Metabolomic | NaCl at 0%, 0.25%, 0.50%, 0.75%, 1.00%, 1.25% | 1, 2, 3, 5, 7, 11, 15 h, respectively | [91] |
| Chlamydomonas sp. JSC4 | Metabolomic | 0%, 1%, 2% sea salt | 3, 5, 7 days | [73] |
| C. reinhardtii CC-503 | Secretomic | 300 Mm NaCl | Several generations | [92] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bazzani, E.; Lauritano, C.; Mangoni, O.; Bolinesi, F.; Saggiomo, M. Chlamydomonas Responses to Salinity Stress and Possible Biotechnological Exploitation. J. Mar. Sci. Eng. 2021, 9, 1242. https://doi.org/10.3390/jmse9111242
Bazzani E, Lauritano C, Mangoni O, Bolinesi F, Saggiomo M. Chlamydomonas Responses to Salinity Stress and Possible Biotechnological Exploitation. Journal of Marine Science and Engineering. 2021; 9(11):1242. https://doi.org/10.3390/jmse9111242
Chicago/Turabian StyleBazzani, Emma, Chiara Lauritano, Olga Mangoni, Francesco Bolinesi, and Maria Saggiomo. 2021. "Chlamydomonas Responses to Salinity Stress and Possible Biotechnological Exploitation" Journal of Marine Science and Engineering 9, no. 11: 1242. https://doi.org/10.3390/jmse9111242
APA StyleBazzani, E., Lauritano, C., Mangoni, O., Bolinesi, F., & Saggiomo, M. (2021). Chlamydomonas Responses to Salinity Stress and Possible Biotechnological Exploitation. Journal of Marine Science and Engineering, 9(11), 1242. https://doi.org/10.3390/jmse9111242








