Chilo suppressalis Population Dynamics Forecasting by Exponential Smoothing Decomposition and Multi-Stream Network
Abstract
1. Introduction
- We propose a novel deep learning-based framework for C. suppressalis population forecasting. The proposed framework employs a triple-stream network to effectively capture distinct multi-scale patterns, thereby improving prediction accuracy.
- We construct a novel real-world dataset, HNRP-6R. This dataset comprises over two decades (2000–2022) of rice pest monitoring data alongside 13 meteorological factors across six key rice-producing regions in Hunan Province, southern China.
- We conduct extensive experiments on the newly created HNRP-6R dataset, evaluating our framework against several baselines and state-of-the-art methods. The results demonstrate the significant superiority of our proposed framework, achieving state-of-the-art performance in C. suppressalis population forecasting.
2. Materials and Methods
2.1. Dataset and Preprocessing
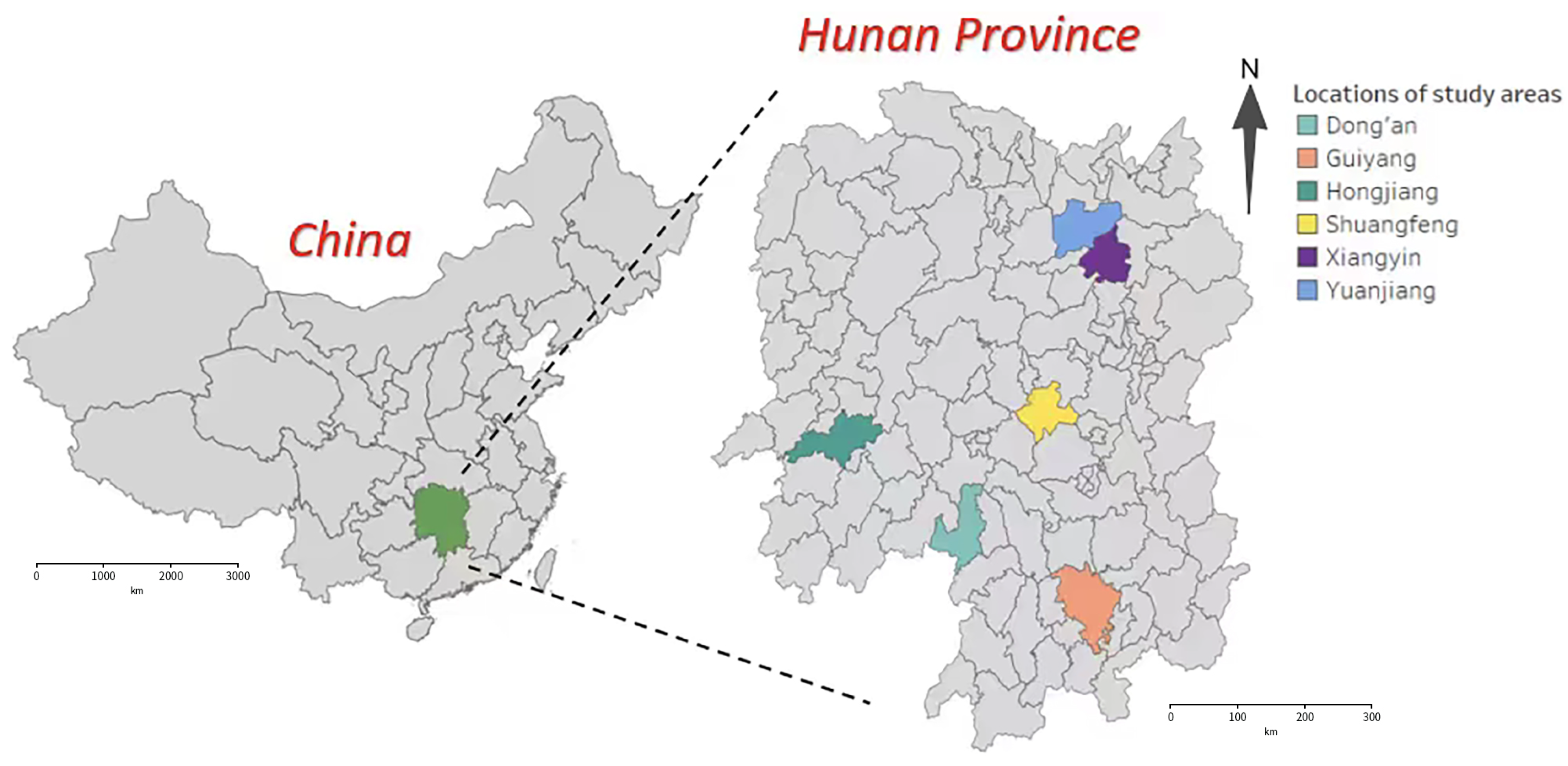
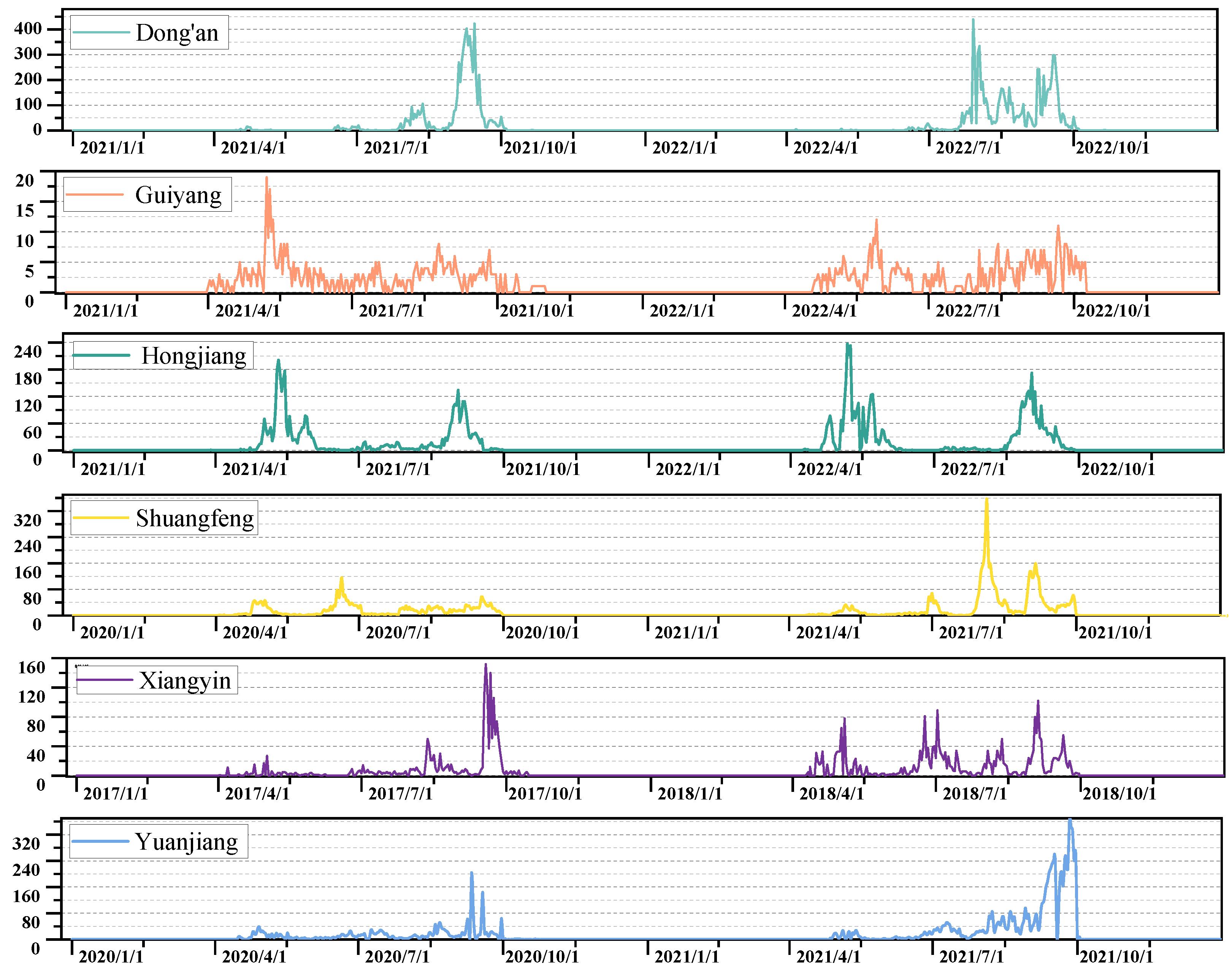
2.1.1. Study Area and Data Collection
2.1.2. Data Preprocessing
2.2. Methodology
2.2.1. Problem Formulation
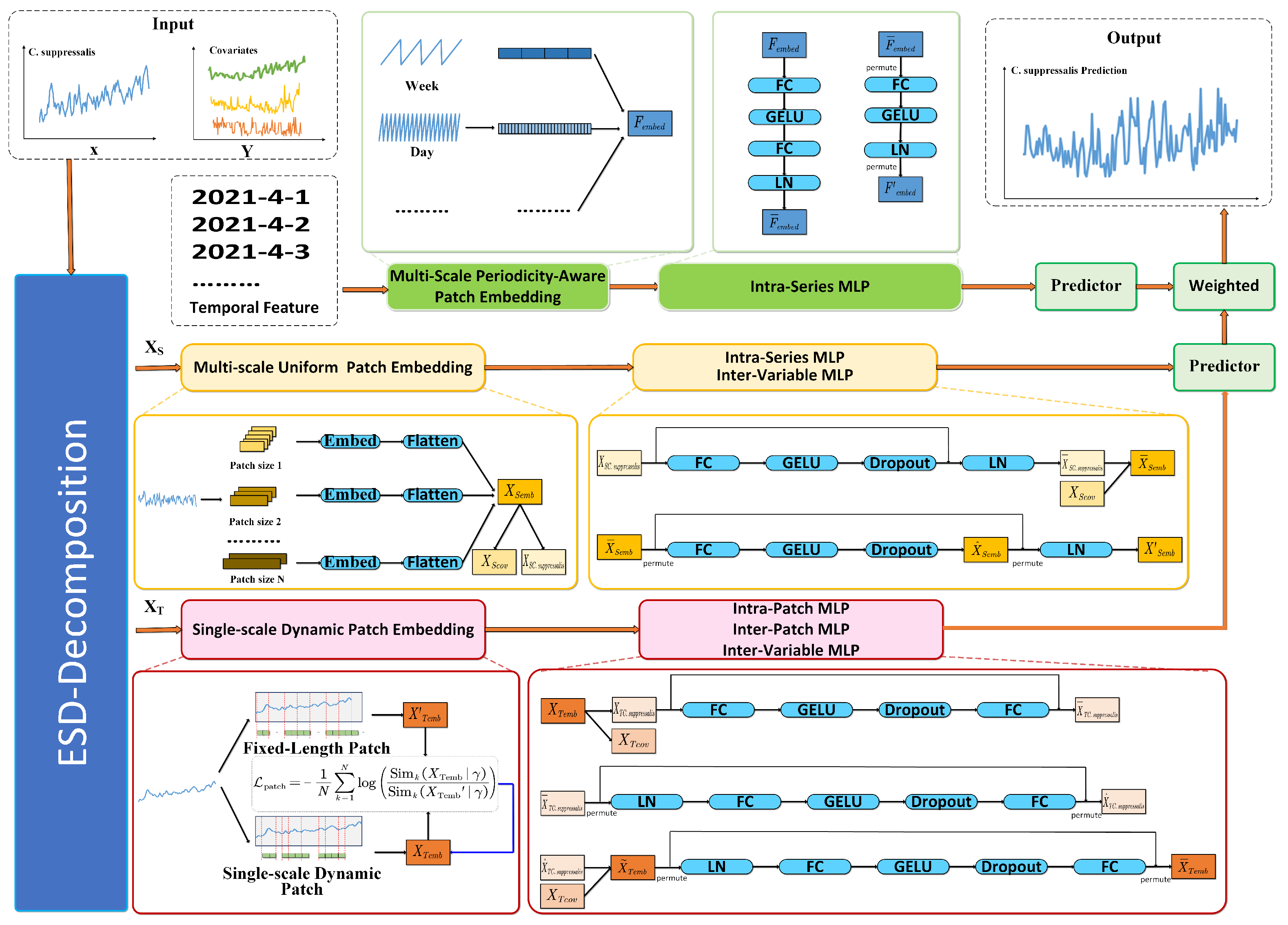
2.2.2. Overview of ESD-TripleStream
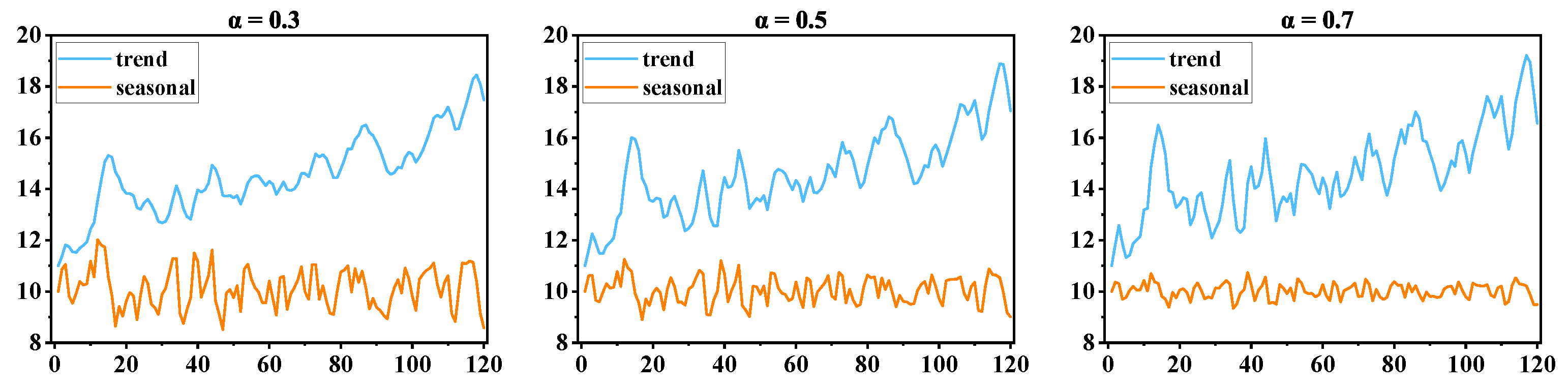
2.2.3. Trend-Seasonal Decomposition
2.2.4. Seasonal Stream
2.2.5. Trend Stream
2.2.6. Temporal Feature Stream
2.2.7. Predictor
3. Results and Discussion
3.1. Experimental Settings
- TimeXer (NeurIPS’24) innovatively uses a hybrid patch- and variable-level representation. It captures endogenous variables’ temporal dependencies via self-attention and exogenous variables’ global trends through cross-attention.
- iTransformer (ICLR’24) features series-wise representations. It treats each variable as a “variate token” and captures variate correlations via the attention mechanism.
- P-sLSTM (AAAI’25) enhances sLSTM with patching and channel independence, addressing traditional LSTM’s short memory limitations to improve medium-term time series forecasting performance.
- TimeLinear (arXiv’24) introduces the Time Stamp Forecaster (TimeSter) module to encode time-related features (e.g., seasons, hours, weekdays) and integrates them with a linear model for enhanced medium-term forecasting.
- Dlinear (AAAI’23) splits the time series into a trend series and a remainder series. Subsequently, it utilizes two single-layer linear networks to separately model these two sequences.
- PatchMLP (AAAI’25) innovatively employs Multi-Scale Patch Embedding to capture multiscale relationships in sequences, decomposes latent vectors into trend and seasonal components, processes them separately, and enhances variable interactions to improve forecasting accuracy.
- HDMixer (AAAI’24) devises a length-extensible patch module that module serves to enrich the boundary information of patches. Moreover, it makes use of the Hierarchical Dependency Explore module to effectively model hierarchical interactions.
- xPatch (AAAI’25) employs EMA for seasonal-trend decomposition, separates time-series components via an MLP–CNN dual-flow architecture with patching and channel independence, and introduces arctangent loss and sigmoid learning rate schemes to optimize training and boost performance.
3.2. Results and Analysis
3.2.1. Performance Comparison
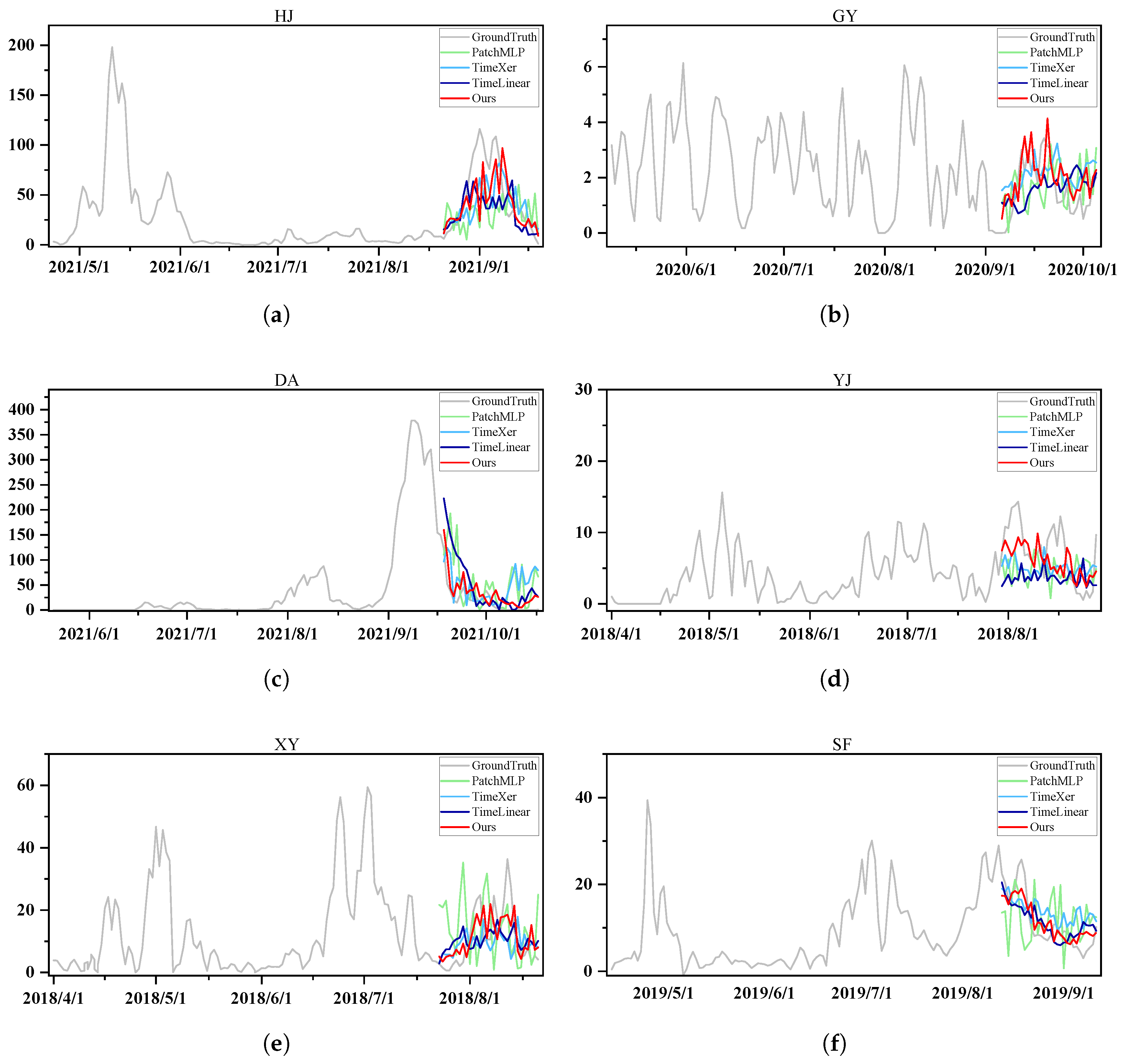
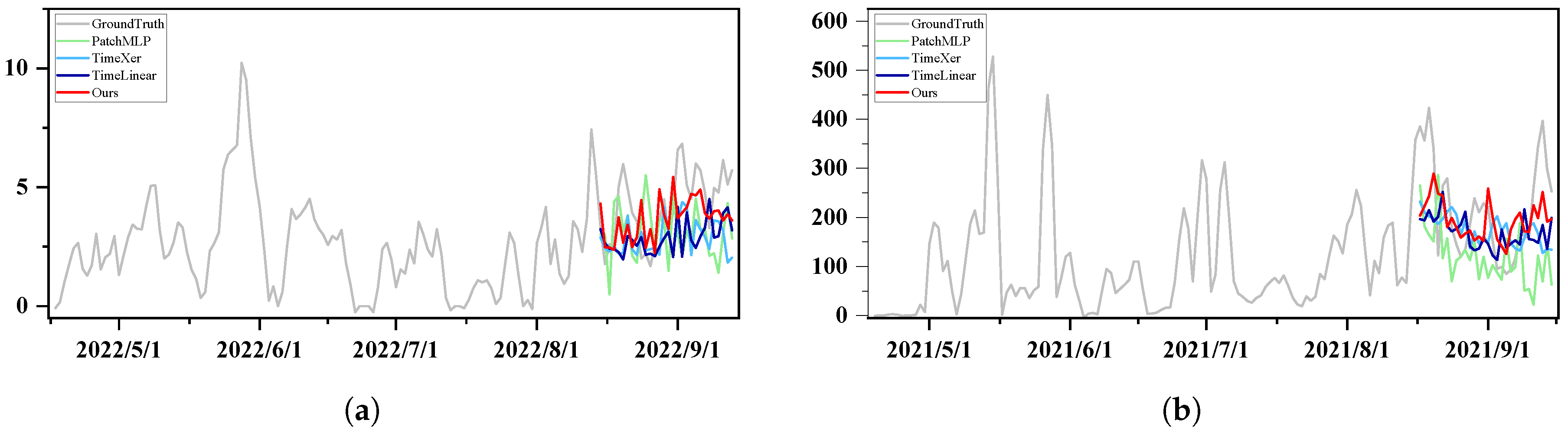
3.2.2. Qualitative Results
3.2.3. Ablation Analysis
- w/o TFS: Remove the Temporal Feature module means that the model does not consider time-related information at all during the prediction process.
- w/o SS: Remove the Seasonal Stream module and replace it with the Trend Stream module.
- w/o TS: Remove the Trend Stream module and replace it with the Seasonal Stream module.
- w/o ESD-S: Remove the ESD module, i.e., do not decompose the original sequence into trend and seasonal components, and use the Seasonal Stream module to process the original sequence.
- w/o ESD-T: Remove the ESD module, i.e., do not decompose the original sequence into trend and seasonal components, and use the Trend Stream module to process the original sequence.
- w/o TFS-Mod: Replace the modules in the temporal feature stream with two nonlinear hidden layers, a 1D convolutional layer, and a linear projection layer from the TimeLinear module.
3.2.4. Generalization Analysis
3.2.5. Model Limitations
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Conflicts of Interest
References
- Gnanamanickam, S.S. Rice and its importance to human life. In Biological Control of Rice Diseases; Springer: Dordrecht, The Netherlands, 2009; pp. 1–11. [Google Scholar]
- Liu, Q.; Wang, X.; Tzin, V.; Romeis, J.; Peng, Y.; Li, Y. Combined transcriptome and metabolome analyses to understand the dynamic responses of rice plants to attack by the rice stem borer Chilo suppressalis (Lepidoptera: Crambidae). BMC Plant Biol. 2016, 16, 259. [Google Scholar] [CrossRef]
- Douglas, A.E. Strategies for enhanced crop resistance to insect pests. Annu. Rev. Plant Biol. 2018, 69, 637–660. [Google Scholar] [CrossRef]
- Cohen, M.B.; Chen, M.; Bentur, J.S.; Heong, K.L.; Ye, G. Bt rice in Asia: Potential benefits, impact, and sustainability. In Integration of Insect-Resistant Genetically Modified Crops Within IPM Programs; Springer: Dordrecht, The Netherlands, 2008; pp. 223–248. [Google Scholar]
- Lin, K.; Hou, M.; Han, L.; Liu, Y. Research progress in host selection and underlying mechanisms, and factors affecting population dynamics of Chilo suppressalis. Plant Prot. 2008, 34, 22. [Google Scholar]
- Zhong, H.Y.; Zhang, J.F.; Li, F.; Yu, K.L.; Chen, J.M. The morphology and an improvement of breeding efficiency of larval Chilo suppressalis (Lepidoptera: Crambidae). Entomol. News 2025, 132, 253–265. [Google Scholar] [CrossRef]
- Xiang, X.; Liu, S.H.; Li, H.J.; Danso Ofori, A.; Yi, X.Q.; Zheng, A.P. Defense strategies of rice in response to the attack of the herbivorous insect, Chilo suppressalis. Int. J. Mol. Sci. 2023, 24, 14361. [Google Scholar] [CrossRef] [PubMed]
- Pandit, P.; Krishnamurthy, K.N.; Bakshi, B. Prediction of crop yield and pest-disease infestation. In AI, Edge and IoT-Based Smart Agriculture; Elsevier: Amsterdam, The Netherlands, 2022; pp. 375–393. [Google Scholar]
- Kopton, J.; de Bruin, S.; Schulz, D.; Luedeling, E. Combining spatio-temporal pest risk prediction and decision theory to improve pest management in smallholder agriculture. Comput. Electron. Agric. 2025, 236, 110426. [Google Scholar] [CrossRef]
- Zheng, Y.Q.; Zhang, X.Y.; Liu, X.; Qin, N.N.; Xu, K.F.; Zeng, R.S.; Liu, J.; Song, Y.Y. Nitrogen supply alters rice defense against the striped stem borer Chilo suppressalis. Front. Plant Sci. 2021, 12, 691292. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Liu, C.; Qiao, S.T.; Guo, F.R.; Xie, Y.; Sun, H.; Liu, Y.; Zhao, S.Q.; Zhou, L.Q.; He, L.F.; et al. The evolution and mechanisms of multiple-insecticide resistance in rice stem borer, Chilo suppressalis Walker (Lepidoptera: Crambidae). J. Agric. Food Chem. 2024, 72, 26475–26490. [Google Scholar] [CrossRef]
- Zhu, L.; Yuan, Z.C.; Yi, Z.Q.; Zhang, C.Y.; Wu, L.; Zhang, Y.; Boussaid, F.; Bennamoun, M.; Zhang, S.C. Information bottleneck-guided KNN contrastive hashing for unsupervised cross-modal retrieval. Knowl.-Based Syst. 2025, 329, 114347. [Google Scholar] [CrossRef]
- Jeong, W.J.; Kim, K.H. Determining the minimum data size for the development of artificial neural network-based prediction models for rice pests in Korea. Comput. Electron. Agric. 2024, 220, 108865. [Google Scholar] [CrossRef]
- Gao, J.; Gu, C.G.; Yang, H.J.; Weng, T.F. Prediction of spatial distribution of invasive alien pests in two-dimensional systems based on a discrete time model. Ecol. Eng. 2020, 143, 105673. [Google Scholar] [CrossRef]
- Ibrahim, E.A.; Salifu, D.; Mwalili, S.; Dubois, T.; Collins, R.; Tonnang, H.E.Z. An expert system for insect pest population dynamics prediction. Comput. Electron. Agric. 2022, 198, 107124. [Google Scholar] [CrossRef]
- Johnson, B.J.; Gomez, M.M.; Munch, S.B. Empirical dynamic modeling for prediction and control of pest populations. Ecol. Model. 2025, 504, 111081. [Google Scholar] [CrossRef]
- Khushboo, S.S.; Gupta, V.; Pandit, D. Forewarning of stripe rust (Puccinia striiformis) of wheat in Jammu plains. Indian Phytopathol. 2023, 76, 767–776. [Google Scholar] [CrossRef]
- Zhu, L.; Wu, R.B.; Liu, D.Y.; Zhang, C.Y.; Wu, L.; Zhang, Y.; Zhang, S.C. Textual semantics enhancement adversarial hashing for cross-modal retrieval. Knowl.-Based Syst. 2025, 317, 113303. [Google Scholar] [CrossRef]
- Wang, M.H.; Li, T. Pest and disease prediction and management for sugarcane using a hybrid autoregressive integrated moving average—a long short-term memory model. Agriculture 2025, 15, 500. [Google Scholar] [CrossRef]
- Narava, R.; DV, S.R.K.; Jaba, J.; P, A.K.; GV, R.R.; V, S.R.; Mishra, S.P.; Kukanur, V. Development of temporal model for forecasting of Helicoverpa armigera (Noctuidae: Lepidopetra) using Arima and Artificial Neural Networks. J. Insect Sci. 2022, 22, 2. [Google Scholar] [CrossRef] [PubMed]
- Quartey-Papafio, T.K.; Javed, S.A.; Liu, S.F. Forecasting cocoa production of six major producers through ARIMA and grey models. Grey Syst. Theory Appl. 2021, 11, 434–462. [Google Scholar] [CrossRef]
- Jain, S.; Ramesh, D. AI based hybrid CNN-LSTM model for crop disease prediction: An ML advent for rice crop. In Proceedings of the 2021 12th International Conference on Computing Communication and Networking Technologies (ICCCNT), Kharagpur, India, 6–8 July 2021; IEEE: Piscataway, NJ, USA, 2021; pp. 1–7. [Google Scholar]
- Guo, Q.C.; He, Z.F.; Wang, Z.S. Monthly climate prediction using deep convolutional neural network and long short-term memory. Sci. Rep. 2024, 14, 17748. [Google Scholar] [CrossRef]
- Guo, Q.C.; He, Z.F.; Wang, Z.S. Prediction of monthly average and extreme atmospheric temperatures in Zhengzhou based on artificial neural network and deep learning models. Front. For. Glob. Change 2023, 6, 1249300. [Google Scholar] [CrossRef]
- Kakarla, S.K.; Schall, E.; Dettweiler, A.; Stohl, J.; Glaser, E.; Adam, H.; Teubler, F.; Ingwersen, J.; Sauer, T.; Piepho, H.P.; et al. Dependence of the abundance of reed glass-winged cicadas (Pentastiridius leporinus (Linnaeus, 1761)) on weather and climate in the upper Rhine Valley, southwest Germany. Agriculture 2025, 15, 1323. [Google Scholar] [CrossRef]
- Pulighe, G.; Lupia, F.; Manente, V. Climate-driven invasion risks of Japanese beetle (Popillia japonica Newman) in Europe predicted through species distribution modelling. Agriculture 2025, 15, 684. [Google Scholar] [CrossRef]
- Neta, A.; Levi, Y.; Morin, E.; Morin, S. Seasonal forecasting of pest population dynamics based on downscaled SEAS5 forecasts. Ecol. Model. 2023, 480, 110326. [Google Scholar] [CrossRef]
- He, Z.P.; Wei, X.J.; Li, Y.P.; Deng, X.Q.; Zhuo, Z.H. Dynamics of Aromia bungii (Faldermann, 1835) (Coleoptera, Cerambycidae) distribution in China amidst climate change: Dual insights from MaxEnt and meta-analysis. Agriculture 2025, 15, 1224. [Google Scholar] [CrossRef]
- Tian, X.Y.; Gao, Y.; Ali, M.Y.; Li, X.H.; Hu, Y.L.; Li, W.B.; Wang, Z.J.; Shi, S.S.; Zhang, J.P. Impact of temperature on age–stage, two-sex life table analysis of a Chinese population of bean bug, Riptortus pedestris (Hemiptera: Alydidae). Agriculture 2022, 12, 1505. [Google Scholar] [CrossRef]
- Wang, J.L.; Zhang, D. Intelligent pest forecasting with meteorological data: An explainable deep learning approach. Expert Syst. Appl. 2024, 252, 124137. [Google Scholar] [CrossRef]
- Chen, P.; Xiao, Q.X.; Zhang, J.; Xie, C.J.; Wang, B. Occurrence prediction of cotton pests and diseases by bidirectional long short-term memory networks with climate and atmosphere circulation. Comput. Electron. Agric. 2020, 176, 105612. [Google Scholar] [CrossRef]
- Chen, Y.; Li, C.C.; Wang, C.; Xiao, Y.S.; Liu, T.B.; Li, J.Y.; Teng, K.; Cai, H.L.; Xiao, Z.P.; Zhou, H.; et al. The application of integrated deep learning models with the assistance of meteorological factors in forecasting major tobacco diseases. Comput. Electron. Agric. 2025, 236, 110429. [Google Scholar] [CrossRef]
- Wu, H.X.; Xu, J.H.; Wang, J.M.; Long, M.S. Autoformer: Decomposition transformers with auto-correlation for long-term series forecasting. Adv. Neural Inf. Process. Syst. 2021, 34, 22419–22430. [Google Scholar]
- Zhou, T.; Ma, Z.Q.; Wen, Q.S.; Wang, X.; Sun, L.; Jin, R. Fedformer: Frequency enhanced decomposed transformer for long-term series forecasting. In Proceedings of the International Conference on Machine Learning, PMLR, Baltimore, MD, USA, 17–23 July 2022; pp. 27268–27286. [Google Scholar]
- Zhang, X.Y.; Jin, X.Y.; Gopalswamy, K.; Gupta, G.; Park, Y.; Shi, X.J.; Wang, H.; Maddix, D.C.; Wang, Y.Y. First de-trend then attend: Rethinking attention for time-series forecasting. arXiv 2022, arXiv:2212.08151. [Google Scholar]
- Zeng, A.L.; Chen, M.X.; Zhang, L.; Xu, Q. Are transformers effective for time series forecasting? In Proceedings of the AAAI Conference on Artificial Intelligence, Washington, DC, USA, 7–14 February 2023; Volume 37, pp. 11121–11128. [Google Scholar]
- Zhu, L.; Wu, R.B.; Zhu, X.H.; Zhang, C.Y.; Wu, L.; Zhang, S.C.; Li, X.L. Bi-direction label-guided semantic enhancement for cross-modal hashing. IEEE Trans. Circuits Syst. Video Technol. 2024, 35, 3983–3999. [Google Scholar] [CrossRef]
- Stitsyuk, A.; Choi, J.S. xPatch: Dual-stream time series forecasting with exponential seasonal-trend decomposition. In Proceedings of the AAAI Conference on Artificial Intelligence, Philadelphia, PA, USA, 27 February–2 March 2025; Volume 39, pp. 20601–20609. [Google Scholar]
- Zeng, C.L.; Tian, Y.; Zheng, G.J.; Gao, Y.J. How much can time-related features enhance time series forecasting? arXiv 2024, arXiv:2412.01557. [Google Scholar] [CrossRef]
- Zhu, L.; Yu, W.R.; Zhu, X.H.; Zhang, C.Y.; Li, Y.D.; Zhang, S.C. MvHAAN: Multi-view hierarchical attention adversarial network for person re-identification. World Wide Web 2024, 27, 59. [Google Scholar] [CrossRef]
- Yu, W.J.; Wang, Q.N.; Zhu, J.; Cao, J.Z.; Liu, T.; Yi, S.X.; Sun, Q.F.; Cui, J.H.; Li, J.W.; Song, Y.L.; et al. Quantifying climate change impacts on rice chalkiness in southern China: Future trends and spatiotemporal patterns. Comput. Electron. Agric. 2025, 231, 109988. [Google Scholar] [CrossRef]
- Kim, T.S.; Kim, J.H.; Tae, Y.W.; Park, C.B.; Choi, J.H.; Choo, J.G. Reversible instance normalization for accurate time-series forecasting against distribution shift. In Proceedings of the International Conference on Learning Representations, Virtual, 3–7 May 2021. [Google Scholar]
- Mo, W.J.; Li, Q.; Lu, Z.X.; Ullah, F.; Guo, J.W.; Xu, H.X.; Lu, Y.H. Dynamic monitoring of Chilo suppressalis resistance to insecticides and the potential influencing factors. Plants 2025, 14, 724. [Google Scholar] [CrossRef]
- Tang, P.W.; Zhang, W.T. Unlocking the power of patch: Patch-based MLP for long-term time series forecasting. In Proceedings of the AAAI Conference on Artificial Intelligence, Philadelphia, PA, USA, 27 February–2 March 2025; Volume 39, pp. 12640–12648. [Google Scholar]
- Nie, Y.Q.; Nguyen, N.H.; Sinthong, P.; Kalagnanam, J. A time series is worth 64 words: Long-term forecasting with transformers. arXiv 2022, arXiv:2211.14730. [Google Scholar]
- Huang, Q.H.; Shen, L.; Zhang, R.X.; Cheng, J.H.; Ding, S.H.; Zhou, Z.Y.; Wang, Y. Hdmixer: Hierarchical dependency with extendable patch for multivariate time series forecasting. In Proceedings of the AAAI Conference on Artificial Intelligence, Vancouver, BC, Canada, 26–27 February 2024; Volume 38, pp. 12608–12616. [Google Scholar]
- Paszke, A.; Gross, S.; Massa, F.; Lerer, A.; Bradbury, J.; Chanan, G.; Killeen, T.; Lin, Z.M.; Gimelshein, N.; Antiga, L.; et al. Pytorch: An imperative style, high-performance deep learning library. Adv. Neural Inf. Process. Syst. 2019, 32, 8026–8037. [Google Scholar]
- Wang, Y.X.; Wu, H.X.; Dong, J.X.; Qin, G.; Zhang, H.R.; Liu, Y.; Qiu, Y.Z.; Wang, J.M.; Long, M.S. Timexer: Empowering transformers for time series forecasting with exogenous variables. Adv. Neural Inf. Process. Syst. 2024, 37, 469–498. [Google Scholar]
- Liu, Y.; Hu, T.G.; Zhang, H.R.; Wu, H.X.; Wang, S.Y.; Ma, L.T.; Long, M.S. itransformer: Inverted transformers are effective for time series forecasting. arXiv 2023, arXiv:2310.06625. [Google Scholar]
- Kong, Y.X.; Wang, Z.P.; Nie, Y.Q.; Zhou, T.; Zohren, S.; Liang, Y.X.; Sun, P.; Wen, Q.S. Unlocking the power of lstm for long term time series forecasting. In Proceedings of the AAAI Conference on Artificial Intelligence, Philadelphia, PA, USA, 27 February–2 March 2025; Volume 39, pp. 11968–11976. [Google Scholar]
| Name of Pest | Abbreviation | Number of Records |
|---|---|---|
| Chilo suppressalis | C. suppressalis | 3,443,619 |
| Nilaparvata lugens | N. lugens | 3,825,898 |
| Cnaphalocrocis medinalis | C. medinalis | 329,841 |
| Sesamia inferens | S. inferens | 90,015 |
| Tryporyza incertulas | T. incertulas | 22,747 |
| Naranga aenescens Moore | N. aenescens | 408,126 |
| Echinocnemus squameus Billberg | E. squameus | 15,242 |
| Lissorhoptrus oryzophilus | L. oryzophilus | 9415 |
| Orseoia oryzae | O. oryzophilus | 10,834 |
| Mythimna seperata | M. seperata | 26,630 |
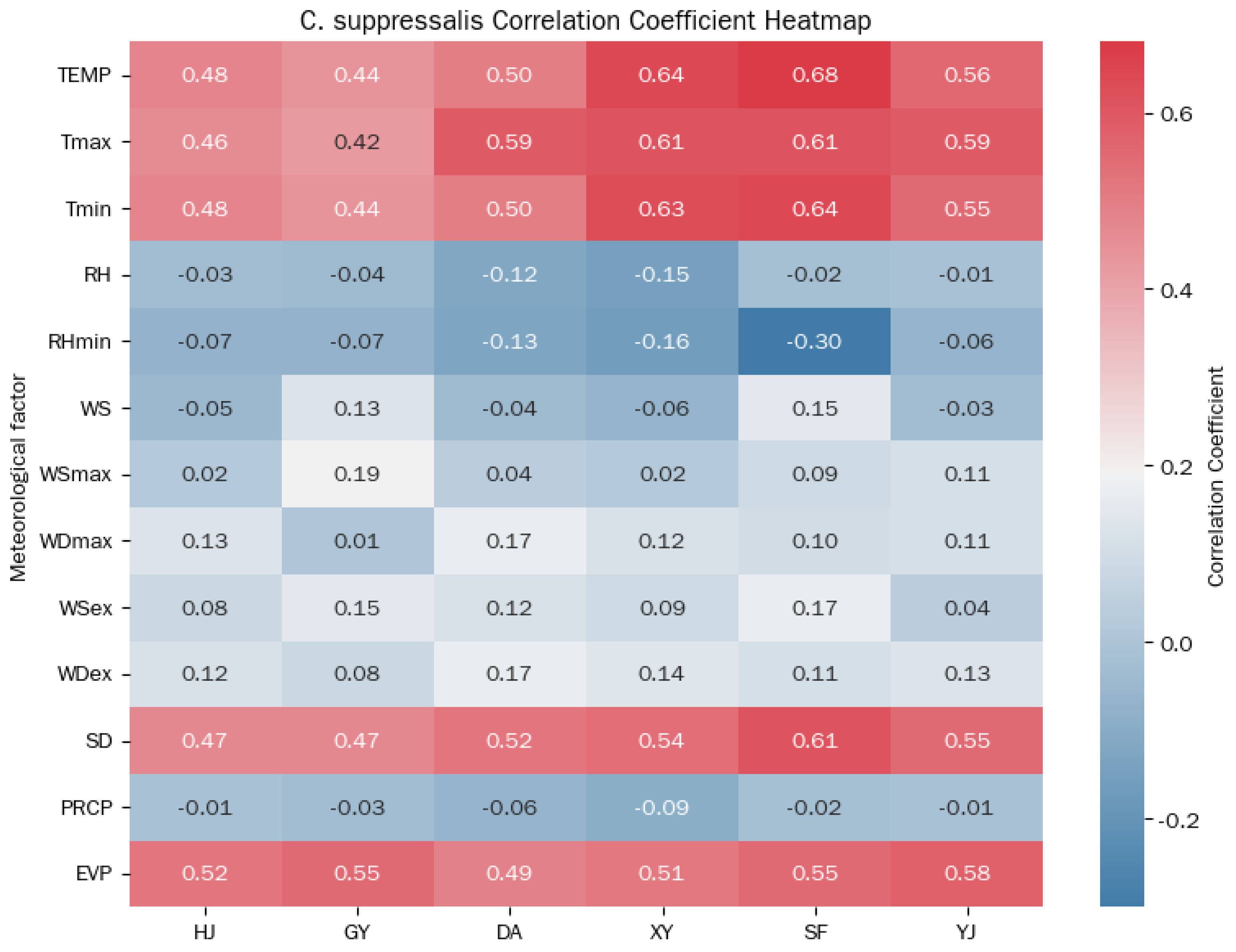
| Meteorological Factor | Abbreviation | Unit |
|---|---|---|
| Mean daily temperature | TEMP | °C |
| Daily maximum temperature | Tmax | °C |
| Daily minimum temperature | Tmin | °C |
| Mean daily relative humidity | RH | % |
| Daily minimum relative humidity | RHmin | % |
| Mean daily wind speed | WS | m/s |
| Daily maximum wind speed | WSmax | m/s |
| Daily extreme wind speed | WSex | m/s |
| Daily maximum wind direction | WDmax | Deg |
| Daily extreme wind direction | WDex | Deg |
| Daily sunshine duration | SD | h |
| Daily precipitation | PRCP | mm |
| Daily evaporation | EVP | mm |
| Location | Abbreviation | Time Span | Number of C. suppressalis |
|---|---|---|---|
| Hongjiang | HJ | 2007–2022 | 63,862 |
| Dong’an | DA | 2006–2022 | 56,013 |
| Yuanjiang | YJ | 2000–2022 | 47,341 |
| Xiangyin | XY | 2000–2019 | 45,082 |
| Shuangfeng | SF | 2010–2021 | 41,761 |
| Guiyang | GY | 2000–2022 | 14,720 |
| Model | Ours | TimeXer | iTransformer | PatchMLP | HDMixer | xPatch | P-sLSTM | TimeLinear | Dlinear | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Metric | MSE | MAE | MSE | MAE | MSE | MAE | MSE | MAE | MSE | MAE | MSE | MAE | MSE | MAE | MSE | MAE | MSE | MAE |
| HJ 120 | 0.212 | 0.228 | 0.221 | 0.248 | 0.216 | 0.244 | 0.258 | 0.231 | 0.218 | 0.249 | 0.367 | 0.322 | 0.324 | 0.406 | 0.216 | 0.232 | 0.288 | 0.410 |
| HJ 240 | 0.223 | 0.250 | 0.262 | 0.292 | 0.216 | 0.309 | 0.284 | 0.3331 | 0.255 | 0.296 | 0.393 | 0.355 | 0.511 | 0.52 | 0.265 | 0.296 | 0.283 | 0.386 |
| HJ 360 | 0.230 | 0.279 | 0.287 | 0.260 | 0.299 | 0.341 | 0.364 | 0.399 | 0.268 | 0.325 | 0.471 | 0.403 | 0.489 | 0.506 | 0.274 | 0.315 | 0.291 | 0.395 |
| HJ 480 | 0.230 | 0.278 | 0.287 | 0.327 | 0.321 | 0.368 | 0.352 | 0.401 | 0.283 | 0.344 | 0.492 | 0.443 | 0.559 | 0.552 | 0.288 | 0.337 | 0.315 | 0.421 |
| HJ 600 | 0.216 | 0.267 | 0.268 | 0.319 | 0.300 | 0.354 | 0.332 | 0.386 | 0.299 | 0.375 | 0.479 | 0.423 | 0.453 | 0.491 | 0.289 | 0.326 | 0.299 | 0.406 |
| GY 120 | 0.210 | 0.312 | 0.218 | 0.319 | 0.222 | 0.326 | 0.444 | 0.399 | 0.247 | 0.325 | 0.578 | 0.464 | 0.346 | 0.393 | 0.255 | 0.325 | 0.288 | 0.359 |
| GY 240 | 0.304 | 0.379 | 0.328 | 0.392 | 0.455 | 0.426 | 0.903 | 0.562 | 0.433 | 0.425 | 0.991 | 0.627 | 0.380 | 0.418 | 0.499 | 0.451 | 0.479 | 0.445 |
| GY 360 | 0.358 | 0.423 | 0.338 | 0.418 | 0.497 | 0.483 | 0.862 | 0.606 | 0.422 | 0.445 | 0.978 | 0.612 | 0.365 | 0.425 | 0.370 | 0.414 | 0.392 | 0.427 |
| GY 480 | 0.337 | 0.421 | 0.440 | 0.448 | 0.632 | 0.530 | 0.944 | 0.654 | 0.600 | 0.538 | 0.967 | 0.598 | 0.394 | 0.427 | 0.822 | 0.606 | 0.447 | 0.461 |
| GY 600 | 0.324 | 0.428 | 0.406 | 0.442 | 0.649 | 0.558 | 1.033 | 0.726 | 0.689 | 0.596 | 0.997 | 0.564 | 0.313 | 0.440 | 0.562 | 0.532 | 0.488 | 0.505 |
| DN 120 | 1.492 | 0.674 | 1.737 | 0.752 | 1.959 | 0.811 | 1.970 | 0.847 | 1.641 | 0.713 | 1.781 | 0.856 | 1.484 | 0.607 | 1.621 | 0.693 | 1.586 | 0.680 |
| DN 240 | 1.293 | 0.631 | 1.663 | 0.768 | 1.766 | 0.795 | 1.856 | 0.872 | 1.410 | 0.677 | 1.618 | 0.781 | 1.370 | 0.641 | 1.697 | 0.794 | 1.422 | 0.658 |
| DN 360 | 1.347 | 0.660 | 1.481 | 0.688 | 1.749 | 0.831 | 1.775 | 0.870 | 1.390 | 0.685 | 1.698 | 0.838 | 1.312 | 0.636 | 1.654 | 0.793 | 1.418 | 0.684 |
| DN 480 | 1.338 | 0.656 | 1.805 | 0.909 | 1.886 | 0.905 | 2.052 | 0.964 | 1.632 | 0.761 | 1.361 | 0.715 | 1.369 | 0.671 | 2.100 | 1.015 | 1.566 | 0.775 |
| DN 600 | 1.383 | 0.721 | 2.005 | 0.980 | 2.066 | 0.986 | 2.168 | 1.091 | 1.685 | 0.807 | 1.632 | 0.797 | 1.393 | 0.751 | 2.390 | 1.101 | 1.732 | 0.850 |
| YJ 120 | 0.918 | 0.441 | 1.014 | 0.480 | 1.234 | 0.582 | 1.293 | 0.605 | 0.932 | 0.449 | 1.162 | 0.575 | 1.140 | 0.544 | 0.929 | 0.446 | 0.938 | 0.457 |
| YJ 240 | 0.845 | 0.432 | 0.974 | 0.467 | 1.093 | 0.537 | 1.154 | 0.568 | 0.835 | 0.411 | 1.126 | 0.544 | 1.071 | 0.533 | 0.897 | 0.450 | 0.969 | 0.478 |
| YJ 360 | 0.917 | 0.437 | 0.965 | 0.467 | 1.081 | 0.515 | 1.056 | 0.524 | 0.932 | 0.444 | 1.196 | 0.586 | 1.047 | 0.542 | 0.921 | 0.434 | 0.951 | 0.482 |
| YJ 480 | 0.972 | 0.444 | 1.044 | 0.469 | 1.026 | 0.488 | 1.104 | 0.536 | 0.976 | 0.454 | 1.199 | 0.589 | 1.031 | 0.543 | 1.012 | 0.461 | 0.979 | 0.452 |
| YJ 600 | 0.916 | 0.446 | 0.991 | 0.475 | 1.212 | 0.711 | 1.039 | 0.512 | 0.963 | 0.472 | 0.990 | 0.476 | 1.011 | 0.554 | 1.023 | 0.479 | 1.091 | 0.534 |
| XY 120 | 0.592 | 0.416 | 0.595 | 0.423 | 0.609 | 0.430 | 0.765 | 0.533 | 0.603 | 0.422 | 0.754 | 0.564 | 0.647 | 0.512 | 0.599 | 0.428 | 0.604 | 0.479 |
| XY 240 | 0.581 | 0.437 | 0.613 | 0.461 | 0.678 | 0.498 | 0.728 | 0.542 | 0.619 | 0.448 | 0.776 | 0.582 | 0.796 | 0.611 | 0.612 | 0.463 | 0.593 | 0.485 |
| XY 360 | 0.578 | 0.468 | 0.667 | 0.487 | 0.756 | 0.545 | 0.842 | 0.627 | 0.741 | 0.526 | 0.895 | 0.654 | 0.828 | 0.623 | 0.713 | 0.542 | 0.714 | 0.563 |
| XY 480 | 0.593 | 0.480 | 0.699 | 0.507 | 0.765 | 0.552 | 0.862 | 0.645 | 0.807 | 0.558 | 0.884 | 0.659 | 0.799 | 0.600 | 0.766 | 0.574 | 0.802 | 0.610 |
| XY 600 | 0.635 | 0.530 | 0.999 | 0.629 | 1.211 | 0.709 | 1.149 | 0.778 | 1.443 | 0.725 | 1.196 | 0.831 | 0.843 | 0.616 | 1.229 | 0.697 | 1.154 | 0.733 |
| SF 120 | 0.506 | 0.341 | 0.514 | 0.347 | 0.445 | 0.319 | 0.523 | 0.369 | 0.435 | 0.304 | 0.617 | 0.419 | 0.549 | 0.463 | 0.501 | 0.333 | 0.460 | 0.354 |
| SF 240 | 0.499 | 0.352 | 0.502 | 0.354 | 0.504 | 0.375 | 0.567 | 0.418 | 0.507 | 0.386 | 0.654 | 0.463 | 0.584 | 0.506 | 0.506 | 0.374 | 0.511 | 0.374 |
| SF 360 | 0.509 | 0.347 | 0.519 | 0.367 | 0.557 | 0.410 | 0.628 | 0.468 | 0.533 | 0.398 | 0.663 | 0.469 | 0.640 | 0.540 | 0.532 | 0.400 | 0.527 | 0.401 |
| SF 480 | 0.509 | 0.356 | 0.515 | 0.372 | 0.571 | 0.437 | 0.681 | 0.507 | 0.541 | 0.403 | 0.745 | 0.558 | 0.682 | 0.550 | 0.541 | 0.407 | 0.522 | 0.420 |
| SF 600 | 0.496 | 0.351 | 0.506 | 0.418 | 0.556 | 0.461 | 0.695 | 0.539 | 0.524 | 0.376 | 0.762 | 0.565 | 0.760 | 0.588 | 0.547 | 0.424 | 0.620 | 0.453 |
| Model | Ours | TimeXer | iTransformer | PatchMLP | HDMixer | xPatch | P-sLSTM | TimeLinear | Dlinear | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Metric | MSE | MAE | MSE | MAE | MSE | MAE | MSE | MAE | MSE | MAE | MSE | MAE | MSE | MAE | MSE | MAE | MSE | MAE |
| HJ 45 | 0.225 | 0.257 | 0.264 | 0.296 | 0.272 | 0.302 | 0.307 | 0.355 | 0.273 | 0.311 | 0.321 | 0.374 | 0.569 | 0.562 | 0.251 | 0.285 | 0.283 | 0.394 |
| HJ 60 | 0.219 | 0.253 | 0.271 | 0.292 | 0.275 | 0.305 | 0.312 | 0.359 | 0.257 | 0.293 | 0.336 | 0.397 | 0.551 | 0.554 | 0.255 | 0.287 | 0.283 | 0.395 |
| HJ 75 | 0.225 | 0.253 | 0.256 | 0.283 | 0.278 | 0.309 | 0.314 | 0.360 | 0.263 | 0.299 | 0.331 | 0.386 | 0.518 | 0.523 | 0.265 | 0.296 | 0.287 | 0.399 |
| HJ 90 | 0.221 | 0.251 | 0.270 | 0.298 | 0.288 | 0.317 | 0.316 | 0.361 | 0.293 | 0.324 | 0.349 | 0.412 | 0.525 | 0.534 | 0.270 | 0.305 | 0.287 | 0.398 |
| GY 45 | 0.329 | 0.389 | 0.370 | 0.415 | 0.440 | 0.426 | 0.875 | 0.560 | 0.430 | 0.425 | 0.912 | 0.587 | 0.346 | 0.398 | 0.417 | 0.414 | 0.417 | 0.430 |
| GY 60 | 0.300 | 0.380 | 0.344 | 0.402 | 0.446 | 0.437 | 1.002 | 0.596 | 0.531 | 0.465 | 1.133 | 0.648 | 0.312 | 0.404 | 0.361 | 0.394 | 0.453 | 0.447 |
| GY 75 | 0.288 | 0.381 | 0.353 | 0.407 | 0.530 | 0.467 | 1.003 | 0.598 | 0.970 | 0.834 | 1.124 | 0.654 | 0.273 | 0.399 | 0.367 | 0.395 | 0.437 | 0.438 |
| GY 90 | 0.294 | 0.379 | 0.353 | 0.396 | 0.420 | 0.424 | 1.053 | 0.609 | 0.961 | 0.83 | 1.197 | 0.684 | 0.308 | 0.406 | 0.379 | 0.398 | 0.431 | 0.437 |
| DN 45 | 1.357 | 0.618 | 1.678 | 0.745 | 1.709 | 0.757 | 1.854 | 0.853 | 1.367 | 0.632 | 1.487 | 0.753 | 1.401 | 0.661 | 1.979 | 1.850 | 1.464 | 0.678 |
| DN 60 | 1.328 | 0.583 | 1.513 | 0.677 | 1.642 | 0.727 | 1.826 | 0.839 | 1.380 | 0.637 | 1.457 | 0.691 | 1.338 | 0.601 | 1.819 | 0.784 | 1.506 | 0.691 |
| DN 75 | 1.349 | 0.586 | 1.476 | 0.668 | 1.550 | 0.690 | 1.823 | 0.835 | 1.359 | 0.610 | 1.432 | 0.666 | 1.340 | 0.589 | 1.689 | 0.747 | 1.555 | 0.692 |
| DN 90 | 1.336 | 0.562 | 1.374 | 0.574 | 1.517 | 0.663 | 1.813 | 0.826 | 1.340 | 0.613 | 1.378 | 0.667 | 1.400 | 0.602 | 1.552 | 0.683 | 1.439 | 0.639 |
| YJ 45 | 0.916 | 0.449 | 1.004 | 0.470 | 1.077 | 0.527 | 1.169 | 0.567 | 0.876 | 0.434 | 1.072 | 0.898 | 1.027 | 0.532 | 0.871 | 0.428 | 1.006 | 0.491 |
| YJ 60 | 0.936 | 0.450 | 0.997 | 0.466 | 1.065 | 0.515 | 1.157 | 0.574 | 0.929 | 0.445 | 1.050 | 0.888 | 1.020 | 0.529 | 0.946 | 0.466 | 1.011 | 0.491 |
| YJ 75 | 0.952 | 0.450 | 0.992 | 0.481 | 1.074 | 0.513 | 1.159 | 0.563 | 1.028 | 0.478 | 1.057 | 0.887 | 1.025 | 0.526 | 0.978 | 0.467 | 1.015 | 0.489 |
| YJ 90 | 0.960 | 0.449 | 1.001 | 0.471 | 1.075 | 0.508 | 1.149 | 0.554 | 1.036 | 0.483 | 1.074 | 0.894 | 1.044 | 0.534 | 0.974 | 0.464 | 1.014 | 0.487 |
| XY 45 | 0.548 | 0.418 | 0.631 | 0.458 | 0.655 | 0.491 | 0.720 | 0.542 | 0.645 | 0.485 | 0.773 | 0.575 | 0.725 | 0.582 | 0.559 | 0.462 | 0.581 | 0.489 |
| XY 60 | 0.549 | 0.424 | 0.595 | 0.452 | 0.640 | 0.486 | 0.709 | 0.539 | 0.614 | 0.473 | 0.764 | 0.561 | 0.747 | 0.587 | 0.602 | 0.467 | 0.591 | 0.499 |
| XY 75 | 0.534 | 0.420 | 0.548 | 0.431 | 0.628 | 0.480 | 0.705 | 0.538 | 0.595 | 0.466 | 0.794 | 0.634 | 0.903 | 0.666 | 0.602 | 0.468 | 0.586 | 0.497 |
| XY 90 | 0.540 | 0.421 | 0.575 | 0.447 | 0.622 | 0.475 | 0.701 | 0.535 | 0.608 | 0.469 | 0.761 | 0.558 | 0.711 | 0.574 | 0.605 | 0.469 | 0.571 | 0.491 |
| SF 45 | 0.498 | 0.338 | 0.490 | 0.340 | 0.499 | 0.357 | 0.598 | 0.430 | 0.471 | 0.345 | 0.573 | 0.438 | 0.655 | 0.549 | 0.498 | 0.349 | 0.506 | 0.377 |
| SF 60 | 0.502 | 0.335 | 0.507 | 0.331 | 0.494 | 0.347 | 0.578 | 0.424 | 0.512 | 0.351 | 0.551 | 0.461 | 0.594 | 0.495 | 0.509 | 0.352 | 0.512 | 0.353 |
| SF 75 | 0.439 | 0.319 | 0.457 | 0.338 | 0.445 | 0.331 | 0.522 | 0.412 | 0.432 | 0.311 | 0.513 | 0.542 | 0.594 | 0.527 | 0.450 | 0.339 | 0.445 | 0.357 |
| SF 90 | 0.396 | 0.305 | 0.399 | 0.337 | 0.398 | 0.320 | 0.482 | 0.403 | 0.402 | 0.329 | 0.542 | 0.554 | 0.550 | 0.509 | 0.416 | 0.335 | 0.405 | 0.357 |
| Model | Full Version | w/o TSF | w/o SS | w/o TS | w/o ESD-S | w/o ESD-T | w/o TFS-Mod | |||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Metric | MSE | MAE | MSE | MAE | MSE | MAE | MSE | MAE | MSE | MAE | MSE | MAE | MSE | MAE |
| HJ 120 | 0.212 | 0.228 | 0.215 | 0.243 | 0.215 | 0.231 | 0.223 | 0.253 | 0.220 | 0.247 | 0.215 | 0.234 | 0.216 | 0.237 |
| HJ 240 | 0.223 | 0.250 | 0.254 | 0.294 | 0.236 | 0.268 | 0.243 | 0.274 | 0.261 | 0.292 | 0.227 | 0.257 | 0.229 | 0.255 |
| HJ 360 | 0.230 | 0.279 | 0.273 | 0.322 | 0.252 | 0.294 | 0.256 | 0.295 | 0.269 | 0.311 | 0.233 | 0.283 | 0.238 | 0.286 |
| HJ 480 | 0.230 | 0.278 | 0.259 | 0.315 | 0.244 | 0.293 | 0.257 | 0.302 | 0.277 | 0.320 | 0.231 | 0.278 | 0.229 | 0.278 |
| HJ 600 | 0.216 | 0.267 | 0.253 | 0.320 | 0.221 | 0.278 | 0.250 | 0.300 | 0.262 | 0.319 | 0.241 | 0.295 | 0.224 | 0.274 |
| GY 120 | 0.210 | 0.312 | 0.272 | 0.332 | 0.254 | 0.322 | 0.241 | 0.322 | 0.255 | 0.329 | 0.252 | 0.323 | 0.219 | 0.319 |
| GY 240 | 0.304 | 0.379 | 0.508 | 0.452 | 0.340 | 0.389 | 0.354 | 0.395 | 0.390 | 0.410 | 0.361 | 0.400 | 0.309 | 0.386 |
| GY 360 | 0.358 | 0.423 | 0.671 | 0.541 | 0.372 | 0.431 | 0.358 | 0.420 | 0.415 | 0.452 | 0.421 | 0.446 | 0.361 | 0.419 |
| GY 480 | 0.337 | 0.421 | 0.569 | 0.531 | 0.479 | 0.505 | 0.387 | 0.438 | 0.480 | 0.497 | 0.362 | 0.428 | 0.330 | 0.420 |
| GY 600 | 0.324 | 0.428 | 0.605 | 0.566 | 0.428 | 0.472 | 0.410 | 0.471 | 0.448 | 0.493 | 0.564 | 0.543 | 0.338 | 0.432 |
| DN 120 | 1.492 | 0.674 | 1.671 | 0.715 | 1.637 | 0.682 | 1.581 | 0.700 | 1.654 | 0.710 | 1.502 | 0.678 | 1.501 | 0.671 |
| DN 240 | 1.293 | 0.631 | 1.381 | 0.665 | 1.326 | 0.649 | 1.336 | 0.641 | 1.407 | 0.662 | 1.299 | 0.628 | 1.296 | 0.634 |
| DN 360 | 1.347 | 0.660 | 1.499 | 0.744 | 1.384 | 0.686 | 1.402 | 0.673 | 1.377 | 0.668 | 1.338 | 0.655 | 1.424 | 0.679 |
| DN 480 | 1.338 | 0.656 | 1.555 | 0.795 | 1.392 | 0.690 | 1.425 | 0.717 | 1.468 | 0.709 | 1.352 | 0.667 | 1.356 | 0.682 |
| DN 600 | 1.383 | 0.721 | 1.857 | 0.938 | 1.440 | 0.795 | 1.562 | 0.790 | 1.600 | 0.772 | 1.364 | 0.748 | 1.449 | 0.789 |
| YJ 120 | 0.918 | 0.441 | 0.942 | 0.458 | 0.939 | 0.435 | 0.942 | 0.446 | 0.948 | 0.455 | 0.964 | 0.448 | 0.924 | 0.448 |
| YJ 240 | 0.845 | 0.432 | 0.881 | 0.459 | 0.847 | 0.428 | 0.942 | 0.457 | 0.893 | 0.443 | 0.902 | 0.434 | 0.896 | 0.442 |
| YJ 360 | 0.917 | 0.437 | 0.945 | 0.464 | 0.927 | 0.439 | 0.924 | 0.438 | 0.929 | 0.446 | 0.929 | 0.439 | 0.941 | 0.438 |
| YJ 480 | 0.972 | 0.444 | 0.981 | 0.489 | 0.953 | 0.442 | 1.024 | 0.461 | 0.950 | 0.440 | 0.956 | 0.433 | 0.948 | 0.444 |
| YJ 600 | 0.916 | 0.446 | 0.936 | 0.459 | 0.921 | 0.449 | 1.050 | 0.466 | 0.952 | 0.452 | 0.967 | 0.462 | 0.959 | 0.458 |
| XY 120 | 0.592 | 0.416 | 0.618 | 0.437 | 0.564 | 0.420 | 0.606 | 0.420 | 0.610 | 0.425 | 0.594 | 0.414 | 0.597 | 0.412 |
| XY 240 | 0.581 | 0.437 | 0.640 | 0.483 | 0.591 | 0.457 | 0.596 | 0.450 | 0.619 | 0.468 | 0.577 | 0.445 | 0.580 | 0.439 |
| XY 360 | 0.578 | 0.468 | 0.688 | 0.544 | 0.641 | 0.491 | 0.620 | 0.489 | 0.636 | 0.500 | 0.636 | 0.495 | 0.582 | 0.473 |
| XY 480 | 0.593 | 0.480 | 0.685 | 0.543 | 0.592 | 0.477 | 0.634 | 0.506 | 0.660 | 0.525 | 0.596 | 0.483 | 0.597 | 0.483 |
| XY 600 | 0.635 | 0.530 | 0.750 | 0.585 | 0.694 | 0.545 | 0.730 | 0.573 | 0.770 | 0.584 | 0.669 | 0.542 | 0.648 | 0.543 |
| SF 120 | 0.506 | 0.341 | 0.497 | 0.342 | 0.489 | 0.329 | 0.520 | 0.338 | 0.509 | 0.343 | 0.499 | 0.335 | 0.521 | 0.375 |
| SF 240 | 0.499 | 0.352 | 0.507 | 0.369 | 0.488 | 0.357 | 0.515 | 0.363 | 0.537 | 0.377 | 0.492 | 0.354 | 0.503 | 0.356 |
| SF 360 | 0.509 | 0.347 | 0.522 | 0.378 | 0.504 | 0.364 | 0.549 | 0.375 | 0.551 | 0.391 | 0.506 | 0.359 | 0.497 | 0.456 |
| SF 480 | 0.509 | 0.356 | 0.524 | 0.376 | 0.511 | 0.380 | 0.541 | 0.368 | 0.552 | 0.392 | 0.517 | 0.364 | 0.515 | 0.359 |
| SF 600 | 0.635 | 0.530 | 0.750 | 0.585 | 0.694 | 0.545 | 0.730 | 0.573 | 0.770 | 0.584 | 0.669 | 0.542 | 0.648 | 0.543 |
| Model | Ours | TimeXer | iTransformer | PatchMLP | HDMixer | xPatch | P-sLSTM | TimeLinear | Dlinear | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Metric | MSE | MAE | MSE | MAE | MSE | MAE | MSE | MAE | MSE | MAE | MSE | MAE | MSE | MAE | MSE | MAE | MSE | MAE |
| HJ 120 | 0.369 | 0.148 | 0.392 | 0.159 | 0.375 | 0.171 | 0.467 | 0.230 | 0.371 | 0.152 | 0.552 | 0.321 | 0.373 | 0.227 | 0.367 | 0.149 | 0.479 | 0.322 |
| HJ 240 | 0.376 | 0.177 | 0.393 | 0.189 | 0.408 | 0.205 | 0.549 | 0.300 | 0.405 | 0.196 | 0.569 | 0.334 | 0.420 | 0.274 | 0.399 | 0.192 | 0.475 | 0.289 |
| HJ 360 | 0.379 | 0.184 | 0.397 | 0.221 | 0.426 | 0.234 | 0.560 | 0.334 | 0.456 | 0.259 | 0.597 | 0.387 | 0.445 | 0.313 | 0.449 | 0.253 | 0.469 | 0.297 |
| HJ 480 | 0.384 | 0.201 | 0.405 | 0.229 | 0.447 | 0.270 | 0.610 | 0.391 | 0.440 | 0.258 | 0.548 | 0.315 | 0.488 | 0.362 | 0.437 | 0.261 | 0.527 | 0.355 |
| HJ 600 | 0.391 | 0.242 | 0.434 | 0.274 | 0.473 | 0.306 | 0.617 | 0.423 | 0.426 | 0.258 | 0.579 | 0.353 | 0.508 | 0.380 | 0.456 | 0.286 | 0.520 | 0.357 |
| GY 120 | 0.014 | 0.058 | 0.015 | 0.059 | 0.015 | 0.057 | 0.019 | 0.075 | 0.016 | 0.059 | 0.033 | 0.144 | 0.033 | 0.148 | 0.016 | 0.067 | 0.049 | 0.194 |
| GY 240 | 0.014 | 0.058 | 0.015 | 0.059 | 0.016 | 0.060 | 0.018 | 0.072 | 0.016 | 0.060 | 0.031 | 0.143 | 0.043 | 0.153 | 0.016 | 0.060 | 0.029 | 0.141 |
| GY 360 | 0.014 | 0.057 | 0.015 | 0.058 | 0.016 | 0.060 | 0.019 | 0.078 | 0.016 | 0.059 | 0.037 | 0.168 | 0.063 | 0.191 | 0.016 | 0.059 | 0.036 | 0.164 |
| GY 480 | 0.013 | 0.055 | 0.014 | 0.058 | 0.015 | 0.059 | 0.018 | 0.077 | 0.016 | 0.064 | 0.038 | 0.171 | 0.060 | 0.188 | 0.015 | 0.057 | 0.039 | 0.174 |
| GY 600 | 0.013 | 0.055 | 0.014 | 0.056 | 0.015 | 0.058 | 0.020 | 0.082 | 0.015 | 0.060 | 0.047 | 0.192 | 0.059 | 0.194 | 0.014 | 0.057 | 0.045 | 0.193 |
| DN 120 | 0.034 | 0.078 | 0.033 | 0.082 | 0.034 | 0.081 | 0.043 | 0.109 | 0.032 | 0.083 | 0.321 | 0.523 | 0.101 | 0.274 | 0.034 | 0.079 | 0.251 | 0.477 |
| DN 240 | 0.030 | 0.068 | 0.032 | 0.075 | 0.032 | 0.083 | 0.041 | 0.110 | 0.031 | 0.080 | 0.291 | 0.496 | 0.069 | 0.202 | 0.035 | 0.081 | 0.223 | 0.448 |
| DN 360 | 0.030 | 0.069 | 0.032 | 0.071 | 0.032 | 0.083 | 0.039 | 0.106 | 0.031 | 0.076 | 0.278 | 0.475 | 0.067 | 0.181 | 0.037 | 0.085 | 0.290 | 0.520 |
| DN 480 | 0.031 | 0.071 | 0.030 | 0.078 | 0.032 | 0.091 | 0.038 | 0.108 | 0.032 | 0.088 | 0.289 | 0.483 | 0.079 | 0.199 | 0.039 | 0.092 | 0.261 | 0.487 |
| DN 600 | 0.031 | 0.074 | 0.034 | 0.078 | 0.032 | 0.091 | 0.036 | 0.077 | 0.036 | 0.097 | 0.253 | 0.432 | 0.073 | 0.191 | 0.038 | 0.086 | 0.244 | 0.471 |
| YJ 120 | 0.589 | 0.545 | 0.578 | 0.545 | 0.561 | 0.523 | 0.999 | 0.715 | 0.752 | 0.614 | 0.976 | 0.702 | 0.582 | 0.557 | 0.748 | 0.619 | 0.761 | 0.613 |
| YJ 240 | 0.585 | 0.547 | 0.574 | 0.537 | 0.658 | 0.571 | 1.030 | 0.753 | 0.668 | 0.570 | 1.112 | 0.814 | 0.599 | 0.577 | 0.748 | 0.618 | 0.851 | 0.664 |
| YJ 360 | 0.589 | 0.562 | 0.581 | 0.567 | 0.667 | 0.593 | 1.175 | 0.822 | 0.768 | 0.662 | 1.165 | 0.834 | 0.675 | 0.588 | 1.093 | 0.813 | 0.764 | 0.632 |
| YJ 480 | 0.572 | 0.551 | 0.585 | 0.569 | 0.672 | 0.590 | 1.157 | 0.828 | 0.716 | 0.642 | 1.177 | 0.867 | 0.766 | 0.625 | 0.889 | 0.718 | 0.833 | 0.667 |
| YJ 600 | 0.584 | 0.561 | 0.597 | 0.585 | 0.663 | 0.587 | 1.246 | 0.866 | 0.661 | 0.591 | 1.306 | 0.912 | 1.011 | 0.730 | 0.878 | 0.701 | 0.848 | 0.672 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
He, C.; Peng, Z.; Peng, L.; Liu, Y.; Zhang, C.; Zhu, L.; Tan, S.; Zou, L. Chilo suppressalis Population Dynamics Forecasting by Exponential Smoothing Decomposition and Multi-Stream Network. Agriculture 2025, 15, 2474. https://doi.org/10.3390/agriculture15232474
He C, Peng Z, Peng L, Liu Y, Zhang C, Zhu L, Tan S, Zou L. Chilo suppressalis Population Dynamics Forecasting by Exponential Smoothing Decomposition and Multi-Stream Network. Agriculture. 2025; 15(23):2474. https://doi.org/10.3390/agriculture15232474
Chicago/Turabian StyleHe, Chao, Ziang Peng, Longhuang Peng, Yi Liu, Chengyuan Zhang, Lei Zhu, Siqiao Tan, and Ling Zou. 2025. "Chilo suppressalis Population Dynamics Forecasting by Exponential Smoothing Decomposition and Multi-Stream Network" Agriculture 15, no. 23: 2474. https://doi.org/10.3390/agriculture15232474
APA StyleHe, C., Peng, Z., Peng, L., Liu, Y., Zhang, C., Zhu, L., Tan, S., & Zou, L. (2025). Chilo suppressalis Population Dynamics Forecasting by Exponential Smoothing Decomposition and Multi-Stream Network. Agriculture, 15(23), 2474. https://doi.org/10.3390/agriculture15232474






