Abstract
Viruses pose a significant threat to humans by causing numerous infectious and potentially fatal diseases. Understanding how the host’s innate immune system recognizes viruses is essential to understanding pathogenesis and ways to control viral infection. Innate immunity also plays a critical role in shaping adaptive immune responses induced by vaccines. Recently developed adjuvants often include nucleic acids that stimulate pattern recognition receptors which are essential components of innate immunity necessary for activating antigen-presentation cells and thereby bridging innate and adaptive immunity. Therefore, understanding viral nucleic acid sensing by cytosolic sensors is essential, as it provides the potential means for developing new vaccine strategies, including effective adjuvants.
1. Introduction
During virus infection, viral nucleic acids that are both released from virions during capsid disintegration and generated upon replication in host cells are recognized by specific receptors termed pattern recognition receptors (PRRs). PRRS play a critical role in detecting viral nucleic acids and, when activated, induce the production of type I interferons (IFNs) and cytokines, resulting in the expression of IFN-stimulated genes (ISGs) in sentinel cells such as monocytes and dendritic cells [1]. This early immune activation subsequently facilitates adaptive immune responses, including antigen trafficking to the lymphoid organs, antigen presentation, and the activation of T and B lymphocytes. The process is fundamental to the generation of virus-specific immunological memory, enhancing long-term protection against future infections.
Live attenuated and inactivated virus vaccines contain many of the same nucleic acid and protein molecules as replication-competent viruses and thus also have the potential to trigger innate immune responses and contribute to adaptive immunity. Moreover, the recently developed mRNA vaccines leverage PRR recognition mechanisms to induce effective immune responses [2]. These vaccines introduce RNA encoding viral proteins into host cells, where PRR recognition leads to IFN-I and cytokine production along with the biosynthesis of viral antigens [3,4,5,6]. This unique feature of mRNA vaccines—their ability to simultaneously activate both innate and adaptive immune responses—is a key factor in their rapid and effective induction of protective immunity.
Currently, attention is directed towards adjuvants that enhance PRR activation, facilitating robust immune responses upon vaccination [7,8]. This strategy offers the potential for more rapid and potent immune protection against viral pathogens. Thus, understanding the cellular recognition of viral nucleic acids and the subsequent signal transduction is essential for improving the design and efficacy of antiviral vaccines. Several excellent reviews that cover PRR recognition of pathogen molecules other than nucleic acids have recently been published [7,8]. In this review, we present a brief overview of the factors involved in recognizing viral nucleic acids and their signaling pathways and describe recent advances in the use of such antiviral responses in vaccine development.
2. The Role of Nucleic Acid Sensing in Pathogen Recognition and Immune Response
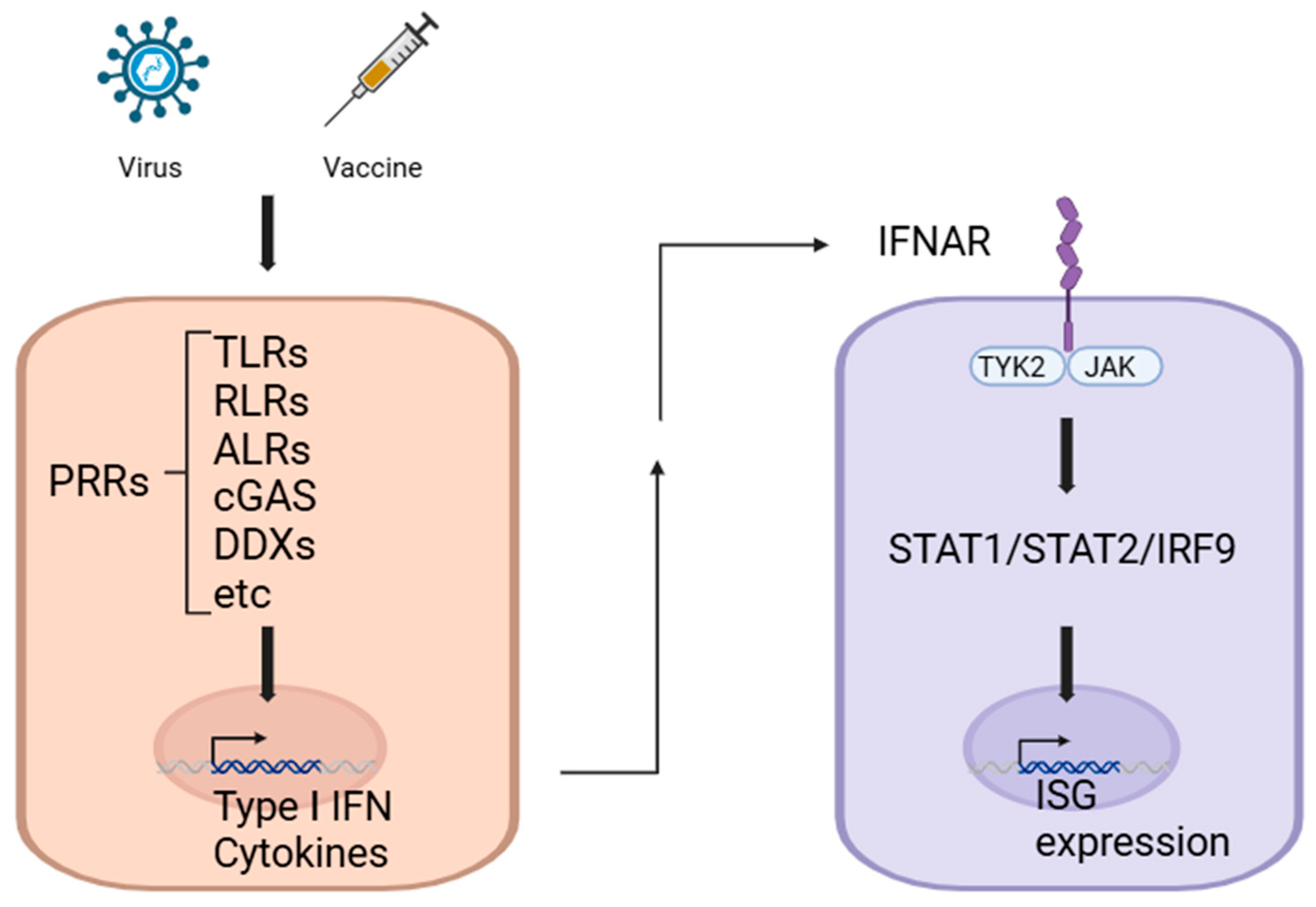
The innate immune system serves as a first line of defense against pathogenic infections by employing an array of PRRs and associated signaling pathways that detect the pathogen-associated molecular patterns (PAMPs) found in the various nucleic acids generated during virus infection, including single-stranded RNA (ssRNA), single-stranded DNA (ssDNA), RNA–DNA hybrids, double-stranded RNA (dsRNA), and double-stranded DNA (dsDNA). These PRRs, which include Toll-like receptors (TLRs), NOD-like receptors (NLRs), the retinoic acid-inducible gene I-like receptors (RLRs) retinoic acid-inducible gene-I (RIG-I) and melanoma differentiation-associated protein-5 (MDA5), absent in melanoma 2 (AIM2)-like receptors (ALRs), cyclic GMP–AMP synthase (cGAS), and receptors with DEAD box helicase (DDX) and zinc finger (ZNF) domains, are evolutionarily conserved and play an essential role in mounting a rapid innate immune response [1,9,10,11,12,13] (Figure 1).
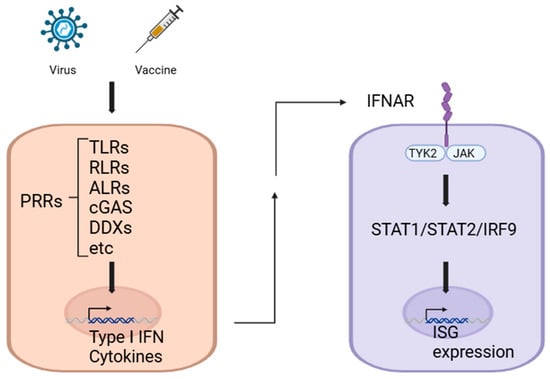
Figure 1.
Key roles of pattern recognition receptors in viral defense and vaccine-induced immunity. The innate immune system utilizes PRRs to detect PAMPs and initiate rapid immune responses. As an example, PRR activation triggers IFN-I production, which through IFNAR signaling induces antiviral gene expression, enhancing viral defense and vaccine efficacy. Created in BioRender. ENYA, T. (2025) Accessed on 9 December 2024. https://BioRender.com/r45j084.
Once PAMPs are recognized, PRR signaling initiates the production of IFNs, pro-inflammatory cytokines, and chemokines, which in turn activate transcription of additional antiviral genes. For example, type I IFNs (IFN-I) bind to the IFN α/β receptor (IFNAR), which activates Janus kinases (JAKs). JAKs then phosphorylate and activate the transcription factors signal transducer and activator of transcription 1 (STAT1) and STAT2, leading to the expression of IFN-stimulated genes (ISGs) [14,15,16] (Figure 1).
Pathogens infect different cell types and tissues, and many replicate in distinct cellular compartments. This, and the need for targeted responses to different pathogens, has likely led to the expansion of PRRs with different signaling pathways. These specific sensing and signaling pathways are also important targets for virus proteins that counteract their action and cause host immune evasion [1,17]. As a result, this greatly accelerates host–pathogen co-evolution [18,19,20,21].
3. TLRs in Innate Immunity: Detecting Pathogens and Triggering Immune Signaling Pathways
TLRs are a conserved family of type I transmembrane proteins that play crucial roles in innate immunity by recognizing a variety of PAMPs, thereby initiating immune responses against a broad spectrum of pathogens [9,13]. TLRs are strategically located either on the cell surface or within intracellular compartments such as endosomes and lysosomes. For example, TLR1, 2, 4, 5, 6, and 11 are primarily responsible for recognizing extracellular microbial components, while TLR3, 7, 8, and 9 recognize viral or bacterial nucleic acids within endosomal compartments. Upon ligand binding, TLRs undergo dimerization, forming either homodimers or heterodimers, which then trigger intracellular signaling cascades through their Toll/IL-1 receptor (TIR) domain. This interaction recruits specific adaptor proteins, activating downstream pathways such as the NF-κB signaling axis or the TBK1 pathway. These cascades culminate in the activation of transcription factors, including NF-κB, IRF3 and 7, driving the production of pro-inflammatory cytokines and IFN-I essential for mounting effective immune responses.
TLRs act as PRRs for a wide variety of PAMPs. TLR2 forms heterodimers with either TLR1 or TLR6, recognizing structurally diverse bacterial lipoproteins. TLR4, while primarily recognizing LPS from Gram-negative bacteria, also detects components from certain parasites and fungi, as well as proteins derived from viruses and host cells (see below) [9,13,22]. TLR5 binds to flagellin, a protein found in bacterial flagella, and TLR11 is known to recognize profilin-like proteins derived from protozoa [9,13,23]. TLR2 also interacts with viral proteins as a TLR2/6 heterodimer, mediating cytokine production during infection with some strains of lymphocytic choriomeningitis virus (LCMV), New World arenaviruses, measles virus, respiratory syncytial virus, and herpes simplex virus (HSV)-1 [24,25,26,27,28].
TLRs 3, 7, and 9 are important sensors of viral nucleic acids. TLR3 recognizes dsRNA, triggering immune responses in response to West Nile virus (WNV) and influenza A virus (IAV), as well as HSV [29,30,31,32,33]. TLR7 detects viral ssRNA, inducing the production of type I interferons and inflammatory cytokines during infections with IAV, vesicular stomatitis virus, and WNV, as well as in response to human immunodeficiency virus (HIV)-1 ssRNA transfection [34,35,36,37]. TLR9, the first identified DNA sensor, recognizes unmethylated CpG DNA as its ligand, participating in immune responses to bacterial and viral DNA, and plays a crucial role in inducing interferon and cytokine production during infections with HSV, adenovirus, and poxvirus [38,39,40].
4. NOD-like Receptors
The NLR family is classified into four subfamilies (NLRA, NLRB, NLRC, and NLRP) based on the structure of their N-terminal effector domains. NLRs are expressed in the cytoplasm of immune cells such as macrophages and dendritic cells, as well as in non-immune cells, including epithelial cells, and play a crucial role in detecting molecules associated with intracellular infections [41,42]. NLRs recognize PAMPs such as those found in bacterial cell walls and danger-associated molecular patterns (DAMPs) released from injured cells [43]. The activation of these NLRs forms an inflammasome complex, which stimulates pro-interleukin-1β (IL-1β) and IL-18 production, triggering an immune response and contributing to host defense against pathogens [44].
NOD2 and NLRP9b in particular are thought to serve as viral nucleic acid sensors. NOD2 plays a key role in recognizing both bacterial-derived muramyl dipeptide motifs and virus-derived ssRNA, and subsequently activates downstream signaling pathways, including NF-κB and MAPK, leading to the induction of IFN-I [41,45,46,47]. NLRP9b recognizes viral dsRNA through the RNA helicase DHX9 and forms an inflammasome that inhibits rotavirus replication in intestinal cells [48].
6. The STING Pathway
The STING pathway is downstream of several nucleic acid sensors, including cGAS, ALRs, and DDXs. Recent studies have identified the degradation pathway for STING after it is activated, wherein ubiquitinated STING is targeted by hepatocyte growth factor-regulated tyrosine kinase substrate (HRS), a crucial component of the ESCRT-0 complex, resulting in its lysosomal degradation [80,81]. STING signaling is also terminated through the ESCRT-dependent microautophagy of vesicles originating from recycling endosomes [82].
There have been many reports regarding the recognition and regulation of DNA virus infection by the STING pathway [83]. STING pathway activation suppresses hepatitis B virus replication in human liver cell lines and in vivo mouse models [84], cytomegalovirus (CMV) replication in primary human endothelial cells [85], and HSV-1 replication in murine microglial cells [86]. DNA viruses have also evolved various strategies to antagonize the STING pathway and many evasion mechanisms and immunomodulators targeting the pathway have been identified [83]. For instance, the HSV-1 protein ICP27 interacts with the STING–TBK1 complex to inhibit type I IFN induction [87]. The UL82 protein of human CMV impairs the translocation of STING from the ER to perinuclear microsomes and inhibits the recruitment of TBK1 and IRF3 to STING [88]. The EBV large tegument protein BPLF1 is a virulence factor that mitigates cGAS-induced IFN-β production by antagonizing IFN-β gene transcription through its deubiquitinase activity [89]; and the pseudorabies virus tegument protein UL13 inhibits antiviral responses by recruiting the E3 ligase RING-finger protein 5 to induce K27/K29-linked ubiquitination and degradation of STING, thereby suppressing STING-dependent signaling [90].
We recently discovered that in mice, one member of the ALR family, IFI207 (PYHINA), increases STING signaling by stabilizing it [20]. The Alr locus in mice has experienced rapid evolution over the past few million years, leading to the emergence of two novel members, MndaL and Ifi207. Notably, Ifi207 in particular exhibits substantial genetic variation even among closely related inbred mouse strains. We found that IFI207 is unique among ALRs due to a large repeat region separating the N-terminal PYD and C-terminal HIN domains that plays a crucial role in stabilizing STING. While IFI207/STING interaction did not affect the cytokine response to bacterial infections (S. aureus, P. aeruginosa, and K. pneumoniae), it contributed to the control of in vivo MLV infection. IFI207 enhanced the STING signaling pathway, leading to an increased antiviral response by stabilizing STING protein [20]. Whether humans or other species also encode ALRs or other factors that stabilize STING remains to be determined, although IFI16 has been reported to have the opposite effect of destabilizing STING [91].
7. The Role of Intracellular Nucleic Acid Sensors in Vaccine Development
The common mechanism underlying all effective vaccines is that the activation of innate immune responses serves as a crucial initiating event that alters the outcome of the adaptive immune response [92,93,94]. Vaccines are thought to utilize two primary types of immune triggers. The first involves PAMPs derived from the target pathogen, while the second pertains to vaccine components, such as specific adjuvants, that induce the release of endogenous DAMPs. Both PAMPs and DAMPs stimulate the innate immune system by activating PRRs [94]. Signals derived from PRRs are integrated at the level of antigen-presenting cells (APCs), thereby effectively modulating the adaptive immune response to the vaccine [94,95].
Recently, adjuvants have been developed that include additional substances that stimulate PRRs, such as the TLR3 agonist polyinosinic:polycytidylic acid (poly I:C) [7]. Moreover, defective interfering particles (DIPs), which are non-infectious virions that typically encapsidate subgenomic viral RNA molecules and are naturally found in measles virus, poliovirus, and influenza vaccine preparations, activate both TLR3 and TLR7 and are also being tested as adjuvants [7]. CpG oligodeoxynucleotide (CpG ODN 1018), a 22-mer sequence with a modified phosphorothioate backbone, is the only TLR9 agonist utilized in an approved vaccine for use in humans, specifically in the licensed hepatitis B vaccine [96,97]. Inactivated whole-virus influenza vaccines do not contain any added adjuvants, but the viral genomic RNA present in the vaccine formulation is believed to exhibit strong adjuvant activity, whereas this effect is weaker in inactivated or split vaccines [98]. The combination of the synthetic dsRNA, poly I:C, and monophosphoryl lipid A (a TLR4 agonist) is under development as an adjuvant for influenza vaccines [99]. Furthermore, cyclic dinucleotides (CDNs), including those activating cGAS, are expected to be developed as vaccine adjuvants due to their ability to induce safe, potent, and long-lasting humoral and cellular memory responses in both systemic and mucosal compartments [100].
DNA vaccines, which include the adjuvant effect of DNA plasmids to enhance immune responses, generate immunogens in vivo which are presented to the immune system, while also activating PRRs that respond to DNA molecules [101,102,103]. Specifically, upon DNA vaccination, Aim2 knockout mice exhibit significantly reduced levels of IFN-α/β, along with diminished humoral and cellular antigen-specific adaptive responses, indicating that the inflammatory responses induced by Aim2 play a crucial role in DNA vaccine efficacy [103].
Some studies have reported that the STING pathway plays a crucial role as an intracellular sensor for DNA vaccines. Notably, cytosolic double-stranded DNA strongly induces IFN-I in both immune and non-immune cells through the STING pathway, likely via interaction with cGAS. The pathway operates independently of traditional TLRs and CpG motifs, suggesting that STING is a key sensor for the innate immune response to DNA vaccines [103,104,105]. However, although it has been proposed that STING agonists might make good vaccine adjuvants, none are in clinical trials as of yet [106].
The advent of mRNA vaccines, particularly highlighted during the COVID-19 pandemic, has revolutionized vaccine technology by introducing viral genetic material into host cells to activate the innate immune system. The two key components of the currently approved mRNA vaccines for COVID-19 are nucleoside-modified mRNAs that encode the antigenic protein and lipid nanoparticles (iLNPs) containing ionizable lipids that facilitate the efficient delivery of intact mRNA to the cytoplasm of cells [3,4].
8. mRNA Vaccines and the Recognition Mechanisms of Non-Self RNA
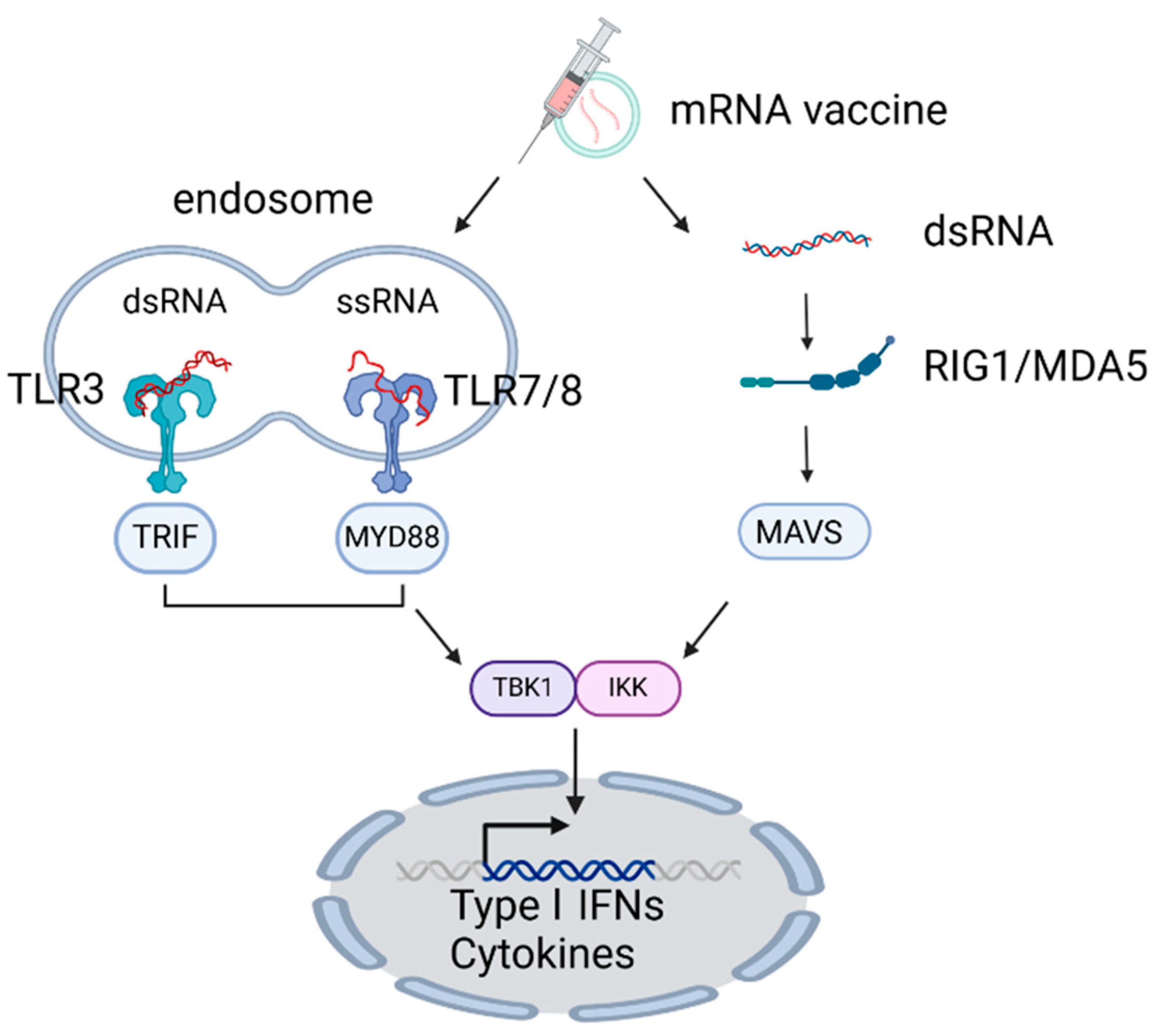
mRNA vaccines can also stimulate innate immunity through TLRs (TLR3, 7, and 8) and the RLRs (RIG-I and MDA5) [107,108,109]. RIG-I recognizes ssRNA and dsRNA bearing a 5-triphosphates, stimulating IFN-I production [109,110]. In contrast, MDA5 detects long dsRNA generated during RNA virus replication, as well as synthetic RNAs, including poly I:C; the recognition of dsRNA by MDA5 activates IRF3 and NFκB, also leading to increased production of IFN-I [109,110] (Figure 2).
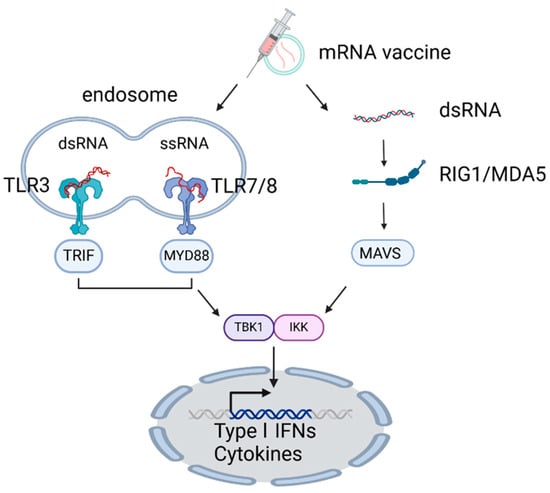
Figure 2.
Mechanisms of innate immunity triggered by mRNA vaccine. mRNA vaccine activates innate immunity via TLRs (TLR3, 7, 8) and RLRs (RIG-I, MDA5), inducing IFN-I production through IRF3 and NFκB signaling. Created in BioRender. ENYA, T. (2025). Accessed on 9 December 2024. https://BioRender.com/p79c529.
However, excessive activation of immune responses may compromise the safety of mRNA vaccines, highlighting the importance of understanding the distinctions between self and non-self mRNA structures. Notably, insights into this distinction have paved the way for RNA modification techniques [111]. Specifically, the recognition of non-self-nucleic acids is based on their structure, availability, and localization. Self-nucleic acids are typically degraded by nucleases before they can be sensed by nucleic acid receptors, preventing their recognition. In contrast, nucleic acid receptors recognize structural features characteristic of non-self RNAs, such as long dsRNA and 5′-triphosphate or 5′-diphosphate dsRNA, which trigger antiviral immune responses through these structural motifs [112]. In mRNA vaccines, modifications to structural elements such as the 5′ cap, 5′-and 3′-untranslated regions, coding region, and poly(A) tail prevent its recognition and degradation as non-self RNA by the immune system [113].
Furthermore, the dsRNA produced during in vitro is recognized in cells by PRRs such as TLR3 and RLRs, leading to the induction of IFN-I [106,107,108,109], as well as by protein kinase R leading to activation of the PKR–eIF2α pathway, which inhibits both cellular and viral translation, leading to apoptosis and thereby triggering the stress response [114]. Incorporation of pseudouridine and 2-thiouridine prevents the recognition of in vitro transcribed mRNA by TLR, RIG-I, and PKR, thereby improving vaccine effectiveness and minimizing the risk of inflammatory responses [115,116,117,118,119].
The mRNA cap structure is a major site of dynamic mRNA methylation and is classified into either Cap1 or Cap2 structures depending on whether the first transcribed nucleotide, or both the first and second transcribed nucleotides, are 2′-O-methylated [120]. The Cap1 structure on mRNA serves as an important marker for distinguishing self RNA from non-self RNAs such as viral RNAs which lack a cap, and by evading recognition by RLRs, viruses can effectively escape immune detection [121]. The dual methylation in Cap2 inhibits the binding of Cap1 to RIG-I, thereby suppressing the ability of endogenous RNA to activate the innate immune response [120].
In summary, these modifications when engineered into RNA vaccines enhance RNA stability, improve translation efficiency, and reduce the overall immunogenic response associated with RNA vaccines. While RNA modification techniques are important for enhancing RNA stability and modulating immunogenicity, the cytosolic sensors are also critical for ensuring the safety and efficacy of mRNA vaccines.
9. Conclusions
Cytosolic sensors regulate nucleic acid recognition and signaling pathways to appropriately control immune responses. Through these mechanisms, cells have evolved a multi-layered system to detect virus-derived components that serve as PAMPs, suppressing viral infection at an early stage via the innate immune system and facilitating the transition to adaptive immune responses. Thus, the mammalian innate immune system utilizes multiple PRRs to effectively control and eliminate viral infections, demonstrating sophisticated evolutionary adaptation.
Advancements in the understanding of cytosolic sensors and signaling mechanisms are essential for the development of effective and safe vaccines. Intracellular nucleic acid sensors are integral to the immunogenicity of live attenuated, killed, and nucleic acid-based vaccines. Their roles in mediating immune responses form the foundation for developing innovative strategies to enhance vaccine efficacy, facilitating the design of next-generation vaccines against infectious diseases and cancer. At the same time, the use of these molecules as adjuvants could lead to the excessive, uncontrolled activation of innate immune responses that may compromise vaccine safety. Future research is anticipated to further refine vaccine design based on these pathways, leading to more robust and sustained immune responses while minimizing untoward side effects.
Author Contributions
T.E.: Conceptualization; writing—review and editing; writing—original draft. S.R.R.: Conceptualization; writing—review and editing; funding acquisition; supervision. All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by NIH/NIAID 1R01AI174538.
Acknowledgments
We apologize for being unable to cite all the excellent papers on this subject. We thank Masaaki Miyazawa for their helpful comments on the manuscript.
Conflicts of Interest
The authors declare no conflicts of interest.
References
- Guy, C.; Bowie, A.G. Recent insights into innate immune nucleic acid sensing during viral infection. Curr. Opin. Immunol. 2022, 78, 102250. [Google Scholar] [CrossRef] [PubMed]
- Pardi, N.; Hogan, M.J.; Porter, F.W.; Weissman, D. mRNA vaccines—A new era in vaccinology. Nat. Rev. Drug Discov. 2018, 17, 261–279. [Google Scholar] [CrossRef] [PubMed]
- Cullis, P.R.; Hope, M.J. Lipid Nanoparticle Systems for Enabling Gene Therapies. Mol. Ther. 2017, 25, 1467–1475. [Google Scholar] [CrossRef] [PubMed]
- Verbeke, R.; Hogan, M.J.; Loré, K.; Pardi, N. Innate immune mechanisms of mRNA vaccines. Immunity 2022, 55, 1993–2005. [Google Scholar] [CrossRef] [PubMed]
- Lint, S.V.; Renmans, D.; Broos, K.; Dewitte, H.; Lentacker, I.; Heirman, C.; Breckpot, K.; Thielemans, K. The ReNAissanCe of mRNA-based cancer therapy. Expert Rev. Vaccines 2015, 14, 235–251. [Google Scholar] [CrossRef]
- Edwards, D.K.; Jasny, E.; Yoon, H.; Horscroft, N.; Schanen, B.; Geter, T.; Fotin Mleczek, M.; Petsch, B.; Wittman, V. Adjuvant effects of a sequence-engineered mRNA vaccine: Translational profiling demonstrates similar human and murine innate response. J. Transl. Med. 2017, 15, 1–18. [Google Scholar] [CrossRef] [PubMed]
- Vasou, A.; Sultanoglu, N.; Goodbourn, S.; Randall, R.E.; Kostrikis, L.G. Targeting Pattern Recognition Receptors (PRR) for Vaccine Adjuvantation: From Synthetic PRR Agonists to the Potential of Defective Interfering Particles of Viruses. Viruses 2017, 9, 186. [Google Scholar] [CrossRef] [PubMed]
- Hennessy, C.; McKernan, D.P. Anti-Viral Pattern Recognition Receptors as Therapeutic Targets. Cells 2021, 10, 2258. [Google Scholar] [CrossRef] [PubMed]
- Akira, S.; Uematsu, S.; Takeuchi, O. Pathogen Recognition and Innate Immunity. Cell 2006, 124, 783–801. [Google Scholar] [CrossRef] [PubMed]
- Loo, Y.M.; Gale, M., Jr. Immune signaling by RIG-I-like receptors. Immunity 2011, 34, 680–692. [Google Scholar] [CrossRef] [PubMed]
- Bhat, N.; Fitzgerald, K.A. Recognition of Cytosolic DNA by cGAS and other STING dependent sensors. Eur. J. Immunol. 2014, 44, 634–640. [Google Scholar] [CrossRef] [PubMed]
- Schattgen, S.A.; Fitzgerald, K.A. The PYHIN protein family as mediators of host defenses. Immunol. Rev. 2011, 243, 109–118. [Google Scholar] [CrossRef] [PubMed]
- Kawai, T.; Akira, S. The role of pattern-recognition receptors in innate immunity: Update on Toll-like receptors. Nat. Immunol. 2010, 11, 373–384. [Google Scholar] [CrossRef] [PubMed]
- Decker, T.; Stockinger, S.; Karaghiosoff, M.; Muller, M.; Kovarik, P. IFNs and STATs in innate immunity to microorganisms. J. Clin. Investig. 2002, 109, 1271–1277. [Google Scholar] [CrossRef] [PubMed]
- Bose, S.; Banerjee, A.K. Innate immune response against nonsegmented negative strand RNA viruses. J. Interferon Cytokine Res. 2003, 23, 401–412. [Google Scholar] [CrossRef] [PubMed]
- Rehwinkel, J.; Gack, M.U. RIG-I-like receptors: Their regulation and roles in RNA sensing. Nat. Rev. Immunol. 2020, 20, 537–551. [Google Scholar] [CrossRef]
- Zhu, J.; Chiang, C.; Gack, M.U. Viral evasion of the interferon response at a glance. J. Cell Sci. 2023, 136, jcs260682. [Google Scholar] [CrossRef] [PubMed]
- Cridland, J.A.; Curley, E.Z.; Wykes, M.N.; Schroder, K.; Sweet, M.J.; Roberts, T.L.; Ragan, M.A.; Kassahn, K.S.; Stacey, K.J. The mammalian PYHIN gene family: Phylogeny, evolution and expression. BMC Evol. Biol. 2012, 12, 140. [Google Scholar] [CrossRef]
- Phillips, K.P.; Cable, J.; Mohammed, R.S.; Herdegen-Radwan, M.; Raubic, J.; Przesmycka, K.J.; van Oosterhout, C.; Radwan, J. Immunogenetic novelty confers a selective advantage in host-pathogen coevolution. Proc. Natl. Acad. Sci. USA 2018, 115, 1552–1557. [Google Scholar] [CrossRef] [PubMed]
- Moran, E.A.; Salas-Briceno, K.; Zhao, W.; Enya, T.; Aguilera, A.N.; Acosta, I.; Alonzo, F.; Kiani, D.; Behnsen, J.; Alvarez, C.; et al. IFI207, a young and fast-evolving protein, controls retroviral replication via the STING pathway. mBio 2024, 15, e01209-24. [Google Scholar] [CrossRef]
- Melepat, B.; Li, T.; Vinkler, M. Natural selection directing molecular evolution in vertebrate viral sensors. Dev. Comp. Immunol. 2024, 154, 105147. [Google Scholar] [CrossRef] [PubMed]
- Brubaker, S.W.; Bonham, K.S.; Zanoni, I.; Kagan, J.C. Innate immune pattern recognition: A cell biological perspective. Annu. Rev. Immunol. 2015, 33, 257–290. [Google Scholar] [CrossRef]
- Yarovinsky, F.; Zhang, D.; Andersen, J.F.; Bannenberg, G.L.; Serhan, C.N.; Hayden, M.S.; Hieny, S.; Sutterwala, F.S.; Flavell, R.A.; Ghosh, S.; et al. TLR11 Activation of Dendritic Cells by a Protozoan Profilin-like Protein. Science 2005, 308, 1626–1629. [Google Scholar] [CrossRef] [PubMed]
- Zhou, S.; Halle, A.; Kurt-Jones, E.A.; Cerny, A.M.; Porpiglia, E.; Rogers, M.; Golenbock, D.T.; Finberg, R.W. Lymphocytic choriomeningitis virus (LCMV) infection of CNS glial cells results in TLR2-MyD88/Mal-dependent inflammatory responses. J. Neuroimmunol. 2008, 194, 70–82. [Google Scholar] [CrossRef] [PubMed]
- Cuevas, C.D.; Ross, S.R. Toll-like receptor 2-mediated innate immune responses against Junín virus in mice lead to antiviral adaptive immune responses during systemic infection and do not affect viral replication in the brain. J. Virol. 2014, 88, 7703–7714. [Google Scholar] [CrossRef]
- Bieback, K.; Lien, E.; Klagge, I.M.; Avota, E.; Schneider-Schaulies, J.; Duprex, W.P.; Wagner, H.; Kirschning, C.J.; Ter Meulen, V.; Schneider-Schaulies, S. Hemagglutinin protein of wild-type measles virus activates toll-like receptor 2 signaling. J. Virol. 2002, 76, 8729–8736. [Google Scholar] [CrossRef] [PubMed]
- Murawski, M.R.; Bowen, G.N.; Cerny, A.M.; Anderson, L.J.; Haynes, L.M.; Tripp, R.A.; Kurt-Jones, E.A.; Finberg, R.W. Respiratory syncytial virus activates innate immunity through Toll-like receptor 2. J. Virol. 2009, 83, 1492–1500. [Google Scholar] [CrossRef] [PubMed]
- Aravalli, R.N.; Hu, S.; Rowen, T.N.; Palmquist, J.M.; Lokensgard, J.R. Cutting Edge: TLR2-Mediated Proinflammatory Cytokine and Chemokine Production by Microglial Cells in Response to Herpes Simplex Virus. J. Immunol. 2005, 175, 4189–4193. [Google Scholar] [CrossRef]
- Alexopoulou, L.; Holt, A.C.; Medzhitov, R.; Flavell, R.A. Recognition of double-stranded RNA and activation of NF-κB by Toll-like receptor 3. Nature 2001, 413, 732–738. [Google Scholar] [CrossRef] [PubMed]
- Tabeta, K.; Georgel, P.; Janssen, E.; Du, X.; Hoebe, K.; Crozat, K.; Mudd, S.; Shamel, L.; Sovath, S.; Goode, J.; et al. Toll-like receptors 9 and 3 as essential components of innate immune defense against mouse cytomegalovirus infection. Proc. Natl. Acad. Sci. USA 2004, 101, 3516–3521. [Google Scholar] [CrossRef] [PubMed]
- Wang, T.; Town, T.; Alexopoulou, L.; Anderson, J.F.; Fikrig, E.; Flavell, R.A. Toll-like receptor 3 mediates West Nile virus entry into the brain causing lethal encephalitis. Nat. Med. 2004, 10, 1366–1373. [Google Scholar] [CrossRef]
- Goffic, R.L.; Balloy, V.; Lagranderie, M.; Alexopoulou, L.; Escriou, N.; Flavell, R.; Chignard, M.; Si-Tahar, M. Detrimental contribution of the Toll-like receptor (TLR)3 to influenza A virus-induced acute pneumonia. PLoS Pathog. 2006, 2, e53. [Google Scholar] [CrossRef] [PubMed]
- Reinert, L.S.; Harder, L.; Holm, C.K.; Iversen, M.B.; Horan, K.A.; Dagnæs-Hansen, F.; Ulhøi, B.P.; Holm, T.H.; Mogensen, T.H.; Owens, T.; et al. TLR3 deficiency renders astrocytes permissive to herpes simplex virus infection and facilitates establishment of CNS infection in mice. J. Clin. Investig. 2012, 122, 1368–1376. [Google Scholar] [CrossRef] [PubMed]
- Diebold, S.S.; Kaisho, T.; Hemmi, H.; Akira, S.; Sousa, C.R.E. Innate Antiviral Responses by Means of TLR7-Mediated Recognition of Single-Stranded RNA. Science 2004, 303, 1529–1531. [Google Scholar] [CrossRef]
- Lund, J.M.; Alexopoulou, L.; Sato, A.; Karow, M.; Adams, N.C.; Gale, N.W.; Iwasaki, A.; Flavell, R.A. Recognition of single-stranded RNA viruses by Toll-like receptor 7. Proc. Natl. Acad. Sci. USA 2004, 101, 5598–5603. [Google Scholar] [CrossRef] [PubMed]
- Town, T.; Bai, F.; Wang, T.; Kaplan, A.T.; Qian, F.; Montgomery, R.R.; Anderson, J.F.; Flavell, R.A.; Fikrig, E. Toll-like receptor 7 mitigates lethal West Nile encephalitis via interleukin 23-dependent immune cell infiltration and homing. Immunity 2009, 30, 242–253. [Google Scholar] [CrossRef]
- Heil, F.; Hemmi, H.; Hochrein, H.; Ampenberger, F.; Kirschning, C.; Akira, S.; Lipford, G.; Wagner, H.; Bauer, S. Species-Specific Recognition of Single-Stranded RNA via Tolllike Receptor 7 and 8. Science 2004, 303, 1526–1529. [Google Scholar] [CrossRef] [PubMed]
- Krug, A.; Luker, G.D.; Barchet, W.; Leib, D.A.; Akira, S.; Colonna, M. Herpes simplex virus type 1 activates murine natural interferon-producing cells through toll-like receptor 9. Blood 2004, 103, 1433–1437. [Google Scholar] [CrossRef] [PubMed]
- Samuelsson, C.; Hausmann, J.; Lauterbach, H.; Schmidt, M.; Akira, S.; Wagner, H.; Chaplin, P.; Suter, M.; O’Keeffe, M.; Hochrein, H. Survival of lethal poxvirus infection in mice depends on TLR9, and therapeutic vaccination provides protection. J. Clin. Invest. 2008, 118, 1776–1784. [Google Scholar] [CrossRef] [PubMed]
- Zhu, J.; Huang, X.; Yang, Y. Innate immune response to adenoviral vectors is mediated by both Toll-like receptor-dependent and -independent pathways. J. Virol. 2007, 81, 3170–3180. [Google Scholar] [CrossRef]
- Chen, G.; Shaw, M.H.; Kim, Y.G.; Nuñez, G. NOD-Like Receptors: Role in Innate Immunity and Inflammatory Disease. Annu. Rev. Pathol. 2009, 4, 365–398. [Google Scholar] [CrossRef] [PubMed]
- Ting, J.P.Y.; Lovering, R.C.; Alnemri, E.S.; Bertin, J.; Boss, J.M.; Davis, B.K.; Flavell, R.A.; Girardin, S.E.; Godzik, A.; Harton, J.A.; et al. The NLR gene family: A standard nomenclature. Immunity 2008, 28, 285–287. [Google Scholar] [CrossRef] [PubMed]
- Velloso, F.J.; Trombetta-Lima, M.; Anschau, V.; Sogayar, M.C.; Correa, R.G. NOD-like receptors: Major players (and targets) in the interface between innate immunity and cancer. Biosci. Rep. 2019, 39, BSR20181709. [Google Scholar] [CrossRef] [PubMed]
- Venuprasad, K.; Theiss, A.L. NLRP6 in host defense and intestinal inflammation. Cell Rep. 2021, 35, 109043. [Google Scholar] [CrossRef] [PubMed]
- Keestra-Gounder, A.M.; Tsolis, R.M. NOD1 and NOD2: Beyond Peptidoglycan Sensing. Trends Immunol. 2017, 38, 758–767. [Google Scholar] [CrossRef] [PubMed]
- Sabbah, A.; Chang, T.H.; Harnack, R.; Frohlich, V.; Tominaga, K.; Dube, P.H.; Xiang, Y.; Bose, S. Activation of innate immune antiviral responses by Nod2. Nat. Immunol. 2009, 10, 1073–1080. [Google Scholar] [CrossRef] [PubMed]
- Mukherjee, T.; Hovingh, E.S.; Foerster, E.G.; Abdel-Nour, M.; Philpott, D.J.; Girardin, S.E. NOD1 and NOD2 in inflammation, immunity and disease. Arch. Biochem. Biophys. 2019, 670, 69–81. [Google Scholar] [CrossRef] [PubMed]
- Zhu, S.; Ding, S.; Wang, P.; Wei, Z.; Pan, W.; Palm, N.W.; Yang, Y.; Yu, H.; Li, H.B.; Wang, G.; et al. Nlrp9b inflammasome restricts rotavirus infection in intestinal epithelial cells. Nature 2017, 546, 667–670. [Google Scholar] [CrossRef] [PubMed]
- Bruns, A.M.; Horvath, C.M. Antiviral RNA recognition and assembly by RLR family innate immune sensors. Cytokine Growth Factor Rev. 2014, 25, 507–512. [Google Scholar] [CrossRef] [PubMed]
- Solstad, A.; Hogaboam, O.; Forero, A.; Hemann, E.A. RIG-I-like Receptor Regulation of Immune Cell Function and Therapeutic Implications. J. Immunol. 2022, 209, 845–854. [Google Scholar] [CrossRef] [PubMed]
- Moran, E.A.; Ross, S.R. Insights into Sensing of Murine Retroviruses. Viruses 2020, 12, 836. [Google Scholar] [CrossRef] [PubMed]
- Itell, H.L.; Humes, D.; Overbaugh, J. Several cell-intrinsic effectors drive type I interferon-mediated restriction of HIV-1 in primary CD4+ T cells. Cell Rep. 2023, 42, 112556. [Google Scholar] [CrossRef] [PubMed]
- Hotter, D.; Bosso, M.; Jønsson, K.L.; Krapp, C.; Stürzel, C.M.; Das, A.; Littwitz-Salomon, E.; Berkhout, B.; Russ, A.; Wittmann, S.; et al. IFI16 targets the transcription factor Sp1 to suppress HIV-1 transcription and latency reactivation. Cell Host Microbe 2019, 25, 858–872.e13. [Google Scholar] [CrossRef] [PubMed]
- Coffin, J.M.; Hughes, S.H.; Varmus, H.E. Retroviruses; Cold Spring Harbor Laboratory Press: Long Island, NY, USA, 1997; ISBN-10: 0-87969-571-4. [Google Scholar]
- Rassa, J.C.; Meyers, J.L.; Zhang, Y.; Kudaravalli, R.; Ross, S.R. Murine retroviruses activate B cells via interaction with toll-like receptor 4. Proc. Natl. Acad. Sci. USA 2002, 99, 2281–2286. [Google Scholar] [CrossRef] [PubMed]
- Persaud, A.T.; Khela, J.; Fernandes, C.; Chaphekar, D.; Burnie, J.; Tang, V.A.; Colpitts, C.C.; Guzzo, C. Virion-incorporated CD14 enables HIV-1 to bind LPS and initiate TLR4 signaling in immune cells. J. Virol. 2024, 98, e0036324. [Google Scholar] [CrossRef] [PubMed]
- Wilks, J.; Lien, E.; Jacobson, A.N.; Fischbach, M.A.; Qureshi, N.; Chervonsky, A.V.; Golovkina, T.V. Mammalian lipopolysaccharide receptors incorporated into the retroviral envelope augment virus transmission. Cell Host Microbe 2015, 18, 456–462. [Google Scholar] [CrossRef] [PubMed]
- Sun, L.; Wu, J.; Du, F.; Chen, X.; Chen, Z.J. Cyclic GMP-AMP Synthase Is a Cytosolic DNA Sensor That Activates the Type I Interferon Pathway. Science 2013, 339, 786–791. [Google Scholar] [CrossRef] [PubMed]
- Tan, X.; Sun, L.; Chen, J.; Chen, Z.J. Detection of Microbial Infections Through Innate Immune Sensing of Nucleic Acids. Annu. Rev. Microbiol. 2018, 72, 447–478. [Google Scholar] [CrossRef] [PubMed]
- Xiao, T.S.; Fitzgerald, K.A. The cGAS-STING pathway for DNA sensing. Mol. Cell 2013, 51, 135–139. [Google Scholar] [CrossRef]
- Cai, X.; Chiu, Y.H.; Chen, Z.J. The cGAS-cGAMP-STING pathway of cytosolic DNA sensing and signaling. Mol. Cell 2014, 54, 289–296. [Google Scholar] [CrossRef] [PubMed]
- Moriyama, M.; Koshiba, T.; Ichinohe, T. Influenza A virus M2 protein triggers mitochondrial DNA-mediated antiviral immune responses. Nat. Commun. 2019, 10, 4624. [Google Scholar] [CrossRef]
- Zhu, T.; Fernandez-Sesma, A. Innate Immune DNA Sensing of Flaviviruses. Viruses 2020, 12, 979. [Google Scholar] [CrossRef]
- Barber, G.N. STING: Infection, inflammation and cancer. Nat. Rev. Immunol. 2015, 15, 760–770. [Google Scholar] [CrossRef] [PubMed]
- Brunette, R.L.; Young, J.M.; Whitley, D.G.; Brodsky, I.E.; Malik, H.S.; Stetson, D.B. Extensive evolutionary and functional diversity among mammalian AIM2-like receptors. J. Exp. Med. 2012, 209, 1969–1983. [Google Scholar] [CrossRef] [PubMed]
- Nakaya, Y.; Lilue, J.; Stavrou, S.; Moran, E.; Ross, S.R. AIM2-like receptors positively and negatively regulate the interferon response induced by cytosolic DNA. MBio 2017, 8, e00944-17. [Google Scholar] [CrossRef] [PubMed]
- Cagliani, R.; Forni, D.; Biasin, M.; Comabella, M.; Guerini, F.R.; Riva, S.; Pozzoli, U.; Agliardi, C.; Caputo, D.; Malhotra, S.; et al. Ancient and recent selective pressures shaped genetic diversity at AIM2-like nucleic acid sensors. Genome Biol. Evol. 2014, 6, 830–845. [Google Scholar] [CrossRef] [PubMed]
- Stavrou, S.; Blouch, K.; Kotla, S.; Bass, A.; Ross, S.R. Nucleic acid recognition orchestrates the anti-viral response to retroviruses. Cell Host Microbe 2015, 17, 478–488. [Google Scholar] [CrossRef] [PubMed]
- Unterholzner, L.; Keating, S.E.; Baran, M.; Horan, K.A.; Jensen, S.B.; Sharma, S.; Sirois, C.M.; Jin, T.; Latz, E.; Xiao, T.S.; et al. IFI16 is an innate immune sensor for intracellular DNA. Nat. Immunol. 2010, 11, 997–1004. [Google Scholar] [CrossRef]
- Hornung, V.; Ablasser, A.; Charrel-Dennis, M.; Bauernfeind, F.; Horvath, G.; Caffrey, D.R.; Latz, E.; Fitzgerald, K.A. AIM2 recognizes cytosolic dsDNA and forms a caspase1-activating inflammasome with ASC. Nature 2009, 458, 514–518. [Google Scholar] [CrossRef] [PubMed]
- Jin, T.; Perry, A.; Jiang, J.; Smith, P.; Curry, J.A.; Unterholzner, L.; Jiang, Z.; Horvath, G.; Rathinam, V.A.; Johnstone, R.W.; et al. Structures of the HIN domain: DNA complexes reveal ligand binding and activation mechanisms of the AIM2 inflammasome and IFI16 receptor. Immunity 2012, 36, 561–571. [Google Scholar] [CrossRef]
- Erdemci-Evin, S.; Bosso, M.; Krchlikova, V.; Bayer, W.; Regensburger, K.; Mayer, M.; Dittmer, U.; Sauter, D.; Kmiec, D.; Kirchhoff, F. A Variety of Mouse PYHIN Proteins Restrict Murine and Human Retroviruses. Viruses 2024, 16, 493. [Google Scholar] [CrossRef]
- Jakobsen, M.R.; Bak, R.O.; Andersen, A.; Berg, R.K.; Jensen, S.B.; Tengchuan, J.; Jin, T.; Laustsen, A.; Hansen, K.; Ostergaard, L.; et al. IFI16 senses DNA forms of the lentiviral replication cycle and controls HIV-1 replication. Proc. Natl. Acad. Sci. USA 2013, 110, E4571-80. [Google Scholar] [CrossRef] [PubMed]
- Gao, D.; Wu, J.; Wu, Y.T.; Du, F.; Aroh, C.; Yan, N.; Sun, L.; Chen, Z.J. Cyclic GMP-AMP synthase is an innate immune sensor of HIV and other retroviruses. Science 2013, 341, 903–906. [Google Scholar] [CrossRef]
- Knipe, D.M. Nuclear sensing of viral DNA, epigenetic regulation of herpes simplex virus infection, and innate immunity. Virology 2015, 479–480, 153–159. [Google Scholar] [CrossRef] [PubMed]
- Amurri, L.; Dumont, C.; Pelissier, R.; Reynard, O.; Mathieu, C.; Spanier, J.; Pályi, B.; Déri, D.; Karkowski, L.; Gonzalez, C.; et al. Multifaceted activation of STING axis upon Nipah and measles virus-induced syncytia formation. PLoS Pathog. 2024, 20, e1012569. [Google Scholar] [CrossRef]
- Stunnenberg, M.; Geijtenbeek, T.B.H.; Gringhuis, S.I. DDX3 in HIV-1 infection and sensing: A paradox. Cytokine Growth Factor Rev. 2018, 40, 32–39. [Google Scholar] [CrossRef] [PubMed]
- Stavrou, S.; Aguilera, A.N.; Blouch, K.; Ross, S.R. DDX41 Recognizes RNA/DNA Retroviral Reverse Transcripts and Is Critical for In Vivo Control of Murine Leukemia Virus Infection. MBio 2018, 9, e00923-18. [Google Scholar] [CrossRef] [PubMed]
- Ma, J.; Ross, S.R. Multifunctional role of DEAD-box helicase 41 in innate immunity, hematopoiesis and disease. Front. Immunol. 2024, 15, 1451705. [Google Scholar] [CrossRef]
- Balka, K.R.; Venkatraman, R.; Saunders, T.L.; Shoppee, A.; Pang, E.S.; Magill, Z.; Homman-Ludiye, J.; Huang, C.; Lane, R.M.; York, H.M.; et al. Termination of STING responses is mediated via ESCRT-dependent degradation. EMBO J. 2023, 42, e112712. [Google Scholar] [CrossRef] [PubMed]
- Gentili, M.; Liu, B.; Papanastasiou, M.; Dele-Oni, D.; Schwartz, M.A.; Carlson, R.J.; Al’Khafaji, A.M.; Krug, K.; Brown, A.; Doench, J.G.; et al. ESCRT-dependent STING degradation inhibits steady-state and cGAMP-induced signaling. Nat. Commun. 2023, 14, 611. [Google Scholar] [CrossRef] [PubMed]
- Kuchitsu, Y.; Mukai, K.; Uematsu, R.; Takaada, Y.; Shinojima, A.; Shindo, R.; Shoji, T.; Hamano, S.; Ogawa, E.; Sato, R.; et al. STING signalling is terminated through ESCRT-dependent microautophagy of vesicles originating from recycling endosomes. Nat. Cell Biol. 2023, 25, 453–466. [Google Scholar] [CrossRef] [PubMed]
- Zhang, K.; Huang, Q.; Li, X.; Zhao, Z.; Hong, C.; Sun, Z.; Deng, B.; Li, C.; Zhang, J.; Wangm, S. The cGAS-STING pathway in viral infections: A promising link between inflammation, oxidative stress and autophagy. Front. Immunol. 2024, 15, 1352479. [Google Scholar] [CrossRef] [PubMed]
- Zhao, L.; Yuan, H.; Wang, Y.; Geng, Y.; Yun, H.; Zheng, W.; Yuan, Y.; Lv, P.; Hou, C.; Zhang, H.; et al. HBV confers innate immune evasion through triggering HAT1/acetylation of H4K5/H4K12/miR-181a-5p or KPNA2/cGAS-STING/IFN-I signaling. J. Med. Virol. 2023, 95, e28966. [Google Scholar] [CrossRef] [PubMed]
- Lio, C.W.; McDonald, B.; Takahashi, M.; Dhanwani, R.; Sharma, N.; Huang, J.; Pham, E.; Benedict, C.A.; Sharmaet, S. cGAS-STING signaling regulates initial innate control of cytomegalovirus infection. J. Virol. 2016, 90, 7789–7797. [Google Scholar] [CrossRef]
- Reinert, L.S.; Lopušná, K.; Winther, H.; Sun, C.; Thomsen, M.K.; Nandakumar, R.; Mogensen, T.H.; Meyer, M.; Vægter, C.; Nyengaard, J.R.; et al. Sensing of HSV-1 by the cGAS–STING pathway in microglia orchestrates antiviral defence in the CNS. Nat. Commun. 2016, 7, 13348. [Google Scholar] [CrossRef] [PubMed]
- Christensen, M.H.; Jensen, S.B.; Miettinen, J.J.; Luecke, S.; Prabakaran, T.; Reinert, L.S.; Mettenleiter, T.; Chen, Z.J.; Knipe, D.M.; Sandri-Goldin, R.M.; et al. HSV-1 ICP27 targets the TBK1-activated STING signalsome to inhibit virus-induced type I IFN expression. EMBO J. 2016, 35, 1385–1399. [Google Scholar] [CrossRef]
- Fu, Y.Z.; Su, S.; Gao, Y.Q.; Wang, P.P.; Huang, Z.F.; Hu, M.M.; Luo, W.W.; Li, S.; Luo, M.H.; Wang, Y.Y.; et al. Human Cytomegalovirus Tegument Protein UL82 Inhibits STING-Mediated Signaling to Evade Antiviral Immunity. Cell Host Microbe 2017, 21, 231–243. [Google Scholar] [CrossRef] [PubMed]
- Lui, W.Y.; Bharti, A.; Wong, N.H.M.; Jangra, S.; Botelho, M.G.; Yuen, K.S.; Jin, D.Y. Suppression of cGAS- and RIG-I-mediated innate immune signaling by Epstein-Barr virus deubiquitinase BPLF1. PLoS Pathog. 2023, 19, e1011186. [Google Scholar] [CrossRef] [PubMed]
- Kong, Z.; Yin, H.; Wang, F.; Liu, Z.; Luan, X.; Sun, L.; Liu, W.; Shang, Y. Pseudorabies virus tegument protein UL13 recruits RNF5 to inhibit STING-mediated antiviral immunity. PLoS Pathog. 2022, 18, e1010544. [Google Scholar] [CrossRef]
- Panchanathan, R.; Liu, H.; Xin, D.; Choubey, D. Identification of a negative feedback loop between cyclic di-GMP-induced levels of IFI16 and p202 cytosolic DNA sensors and STING. Innate Immun. 2014, 20, 751–759. [Google Scholar] [CrossRef]
- Coffman, R.L.; Sher, A.; Seder, R.A. Vaccine adjuvants: Putting innate immunity to work. Immunity 2010, 33, 492–503. [Google Scholar] [CrossRef]
- Pulendran, B.; Ahmed, R. Immunological mechanisms of vaccination. Nature Immunol. 2011, 12, 509–517. [Google Scholar] [CrossRef] [PubMed]
- Desmet, C.J.; Ishii, K.J. Nucleic acid sensing at the interface between innate and adaptive immunity in vaccination. Nat. Rev. Immunol. 2012, 12, 479–491. [Google Scholar] [CrossRef]
- Iwasaki, A.; Medzhitov, R. Regulation of adaptive immunity by the innate immune system. Science 2010, 327, 291–295. [Google Scholar] [CrossRef]
- Kuo, T.Y.; Lin, M.Y.; Coffman, R.L.; Campbell, J.D.; Traquina, P.; Lin, Y.J.; Liu, L.T.C.; Cheng, J.; Wu, Y.C.; Wu, C.C. Development of CpG-adjuvanted stable prefusion SARS-CoV-2 spike antigen as a subunit vaccine against COVID-19. Sci. Rep. 2020, 10, 20085. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Liu, S.; Li, B.; Sun, X.; Pan, Q.; Zheng, Y.; Liu, J.; Zhao, Y.; Wang, J.; Liu, L.; et al. A novel CpG ODN compound adjuvant enhances immune response to spike subunit vaccines of porcine epidemic diarrhea virus. Front. Immunol. 2024, 15, 1336239. [Google Scholar] [CrossRef] [PubMed]
- Koyama, S.; Aoshi, T.; Tanimoto, T.; Kumagai, Y.; Kobiyama, K.; Tougan, T.; Sakurai, K.; Coban, C.; Horii, T.; Akira, S.; et al. Plasmacytoid Dendritic Cells Delineate Immunogenicity of Influenza Vaccine Subtypes. Sci. Transl. Med. 2010, 2, 25ra24. [Google Scholar] [CrossRef]
- Le, C.T.T.; Ahn, S.Y.; Ho, T.L.; Lee, J.; Lee, D.H.; Hwang, H.S.; Kang, S.M.; Ko, E.J. Adjuvant efects of combination monophosphoryl lipid A and poly I:C on antigen specifc immune responses and protective efcacy of infuenza vaccines. Sci. Rep. 2023, 13, 12231. [Google Scholar] [CrossRef] [PubMed]
- Gogoi, H.; Mansouri, S.; Jin, L. The Age of Cyclic Dinucleotide Vaccine Adjuvants. Vaccines 2020, 8, 453. [Google Scholar] [CrossRef]
- Sato, Y.; Roman, M.; Tighe, H.; Lee, D.; Corr, M.; Nguyen, M.D.; Silverman, G.J.; Lotz, M.; Carson, D.A.; Raz, E. Immunostimulatory DNA Sequences Necessary for Effective Intradermal Gene Immunization. Science 1996, 273, 352–354. [Google Scholar] [CrossRef] [PubMed]
- Takeuchi, O.; Akira, S. Pattern recognition receptors and inflammation. Cell 2010, 140, 805–820. [Google Scholar] [CrossRef] [PubMed]
- Suschak, J.J.; Wang, S.; Fitzgerald, K.A.; Lu, S. Identification of Aim2 as a Sensor for DNA Vaccines. J. Immunol. 2015, 194, 630–636. [Google Scholar] [CrossRef] [PubMed]
- Ishii, K.J.; Kawagoe, T.; Koyama, S.; Matsui, K.; Kumar, H.; Kawai, T.; Uematsu, S.; Takeuchi, O.; Takeshita, F.; Coban, C.; et al. TANK-binding kinase-1 delineates innate and adaptive immune responses to DNA vaccines. Nature 2008, 451, 725–729. [Google Scholar] [CrossRef] [PubMed]
- Li, X.D.; Wu, J.; Gao, D.; Wang, H.; Sun, L.; Chen, Z.J. Pivotal roles of cGAS-cGAMP signaling in antiviral defense and immune adjuvant effects. Science 2013, 341, 1390–1394. [Google Scholar] [CrossRef]
- McWhirter, S.M.; Jefferies, C.A. Nucleic acid sensors as therapeutic targes for human disease. Immunity 2020, 52, 78–97. [Google Scholar] [CrossRef]
- Karikó, K.; Ni, H.; Capodici, J.; Lamphier, M.; Weissman, D. mRNA Is an Endogenous Ligand for Toll-like Receptor 3. J. Biol. Chem. 2004, 279, 12542–12550. [Google Scholar] [CrossRef] [PubMed]
- Tatematsu, M.; Funami, K.; Seya, T.; Matsumoto, M. Extracellular RNA Sensing by Pattern Recognition Receptors. J. Innate Immun. 2018, 10, 398–406. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.; Maruggi, G.; Shan, H.; Li, J. Advances in mRNA Vaccines for Infectious Diseases. Front. Immunol. 2019, 10, 594. [Google Scholar] [CrossRef]
- Goubau, D.; Schlee, M.; Deddouche, S.; Pruijssers, A.J.; Zillinger, T.; Goldeck, M.; Schuberth, C.; Van der Veen, A.G.; Fujimura, T.; Rehwinkel, J.; et al. Antiviral immunity via RIG-I-mediated recognition of RNA bearing 5′-diphosphates. Nature 2014, 514, 372–375. [Google Scholar] [CrossRef] [PubMed]
- Luan, X.; Wang, L.; Song, G.; Zhou, W. Innate immune responses to RNA: Sensing and signaling. Front. Immunol. 2024, 15, 1287940. [Google Scholar] [CrossRef]
- Martin, S.; Hartmann, G. Discriminating self from non-self in nucleic acid sensing. Nat. Rev. Immunol. 2016, 16, 566–580. [Google Scholar] [CrossRef]
- Kim, S.C.; Sekhon, S.S.; Shin, W.R.; Ahn, G.; Cho, B.K.; Ahn, J.Y.; Kim, Y.H. Modifications of mRNA vaccine structural elements for improving mRNA stability and translation efficiency. Mol. Cell. Toxicol. 2022, 18, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Costa-Mattioli, M.; Walter, P. The integrated stress response: From mechanism to disease. Science 2020, 368, eaat5314. [Google Scholar] [CrossRef] [PubMed]
- Karikó, K.; Buckstein, M.; Ni, H.; Weissman, D. Suppression of RNA Recognition by Toll-like Receptors: The Impact of Nucleoside Modification and the Evolutionary Origin of RNA. Immunity 2005, 23, 165–175. [Google Scholar] [CrossRef] [PubMed]
- Liu, A.; Wang, X. The Pivotal Role of Chemical Modifications in mRNA Therapeutics. Front. Cell Dev. Biol. 2022, 10, 901510. [Google Scholar] [CrossRef] [PubMed]
- Hornung, V.; Ellegast, J.; Kim, S.; Brzózka, K.; Jung, A.; Kato, H.; Poeck, H.; Akira, S.; Conzelmann, K.K.; Schlee, M.; et al. 5’-Triphosphate RNA Is the Ligand for RIG-I. Science 2006, 314, 994–997. [Google Scholar] [CrossRef]
- Anderson, B.R.; Muramatsu, H.; Nallagatla, S.R.; Bevilacqua, P.C.; Sansing, L.H.; Weissman, D.; Karikó, K. Incorporation of pseudouridine into mRNA enhances translation by diminishing PKR activation. Nucleic Acids Res. 2010, 38, 5884–5892. [Google Scholar] [CrossRef]
- Lu, T.; Chen, A.; Wang, S.; Zhang, Y.; Yang, B.; Wang, J.; Fang, H.; Gong, Q.; Li, A.; Liu, X.; et al. N1-methylpseudouridine mRNA modification enhances efficiency and specificity of gene 1 overexpression by preventing Prkra-mediated global translation repression. BioRxiv 2024. [Google Scholar] [CrossRef]
- Despic, V.; Jaffrey, S.R. mRNA ageing shapes the Cap2 methylome in mammalian mRNA. Nature 2023, 614, 358–366. [Google Scholar] [CrossRef]
- Ramanathan, A.; Robb, G.B.; Chan, S.H. mRNA capping: Biological functions and applications. Nucleic Acids Res. 2016, 44, 7511–7526. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).