Prediction of Oleic Acid Content of Rapeseed Using Hyperspectral Technique
Abstract
:1. Introduction
2. Materials and Methods
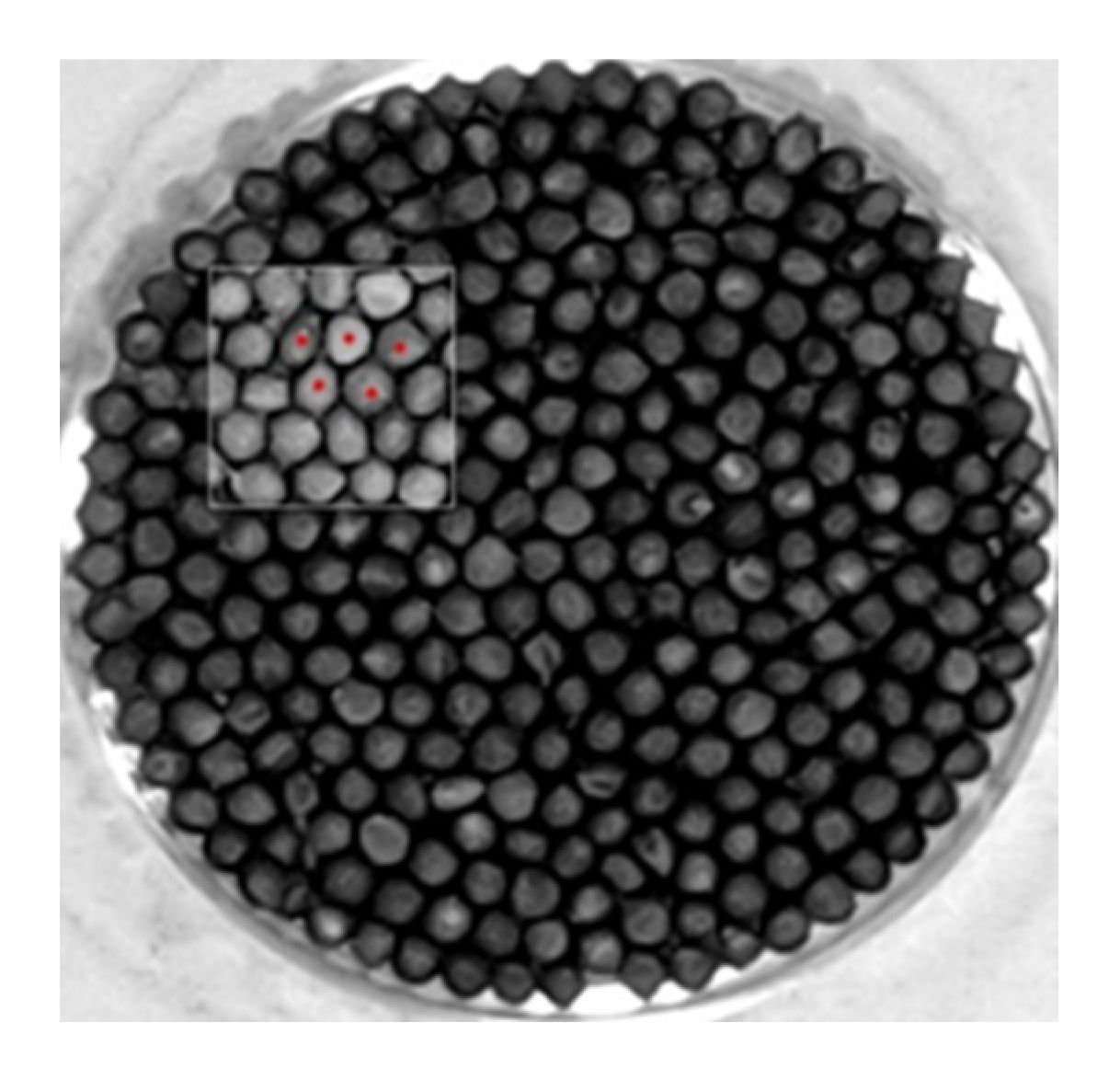
2.1. Materials
The Selected Materials
2.2. Data Collection
- Chromatography column: DB-5 Chromatography column (30 m × 0.25 mm × 0.25 μm, 5% Phenyl-95% Polydimethylsiloxane);
- Temperature programmed condition: The initial temperature is 50°. The temperature is then raised to 200 °C at 10 °C/min and held there for 10 min. The temperature is then raised to 300 °C at 10 °C/min and held there for 10 min again;
- Vaporization chamber temperature: 300 °C;
- Detector temperature: 300 °C;
- Carrier gas: high-purity nitrogen, gas flow ratio 100:1;
- Column flow rate: 1 mL/min;
- Cracking furnace temperature: 350 °C;
- Interface temperature: 300 °C;
- Fatty acid methyl esterification: 400 mL potassium hydroxide methanol solution and 800 mL ether petroleum ether solution.
2.3. Method
2.3.1. Multifractal Detrended Fluctuation Analysis (MF-DFA)
2.3.2. BP Neural Networks
2.4. Evaluate Indicator
3. Result and Discussion
3.1. Feature Selection
3.2. Correlation Analysis of Spectral Parameters and Oleic Acid Content
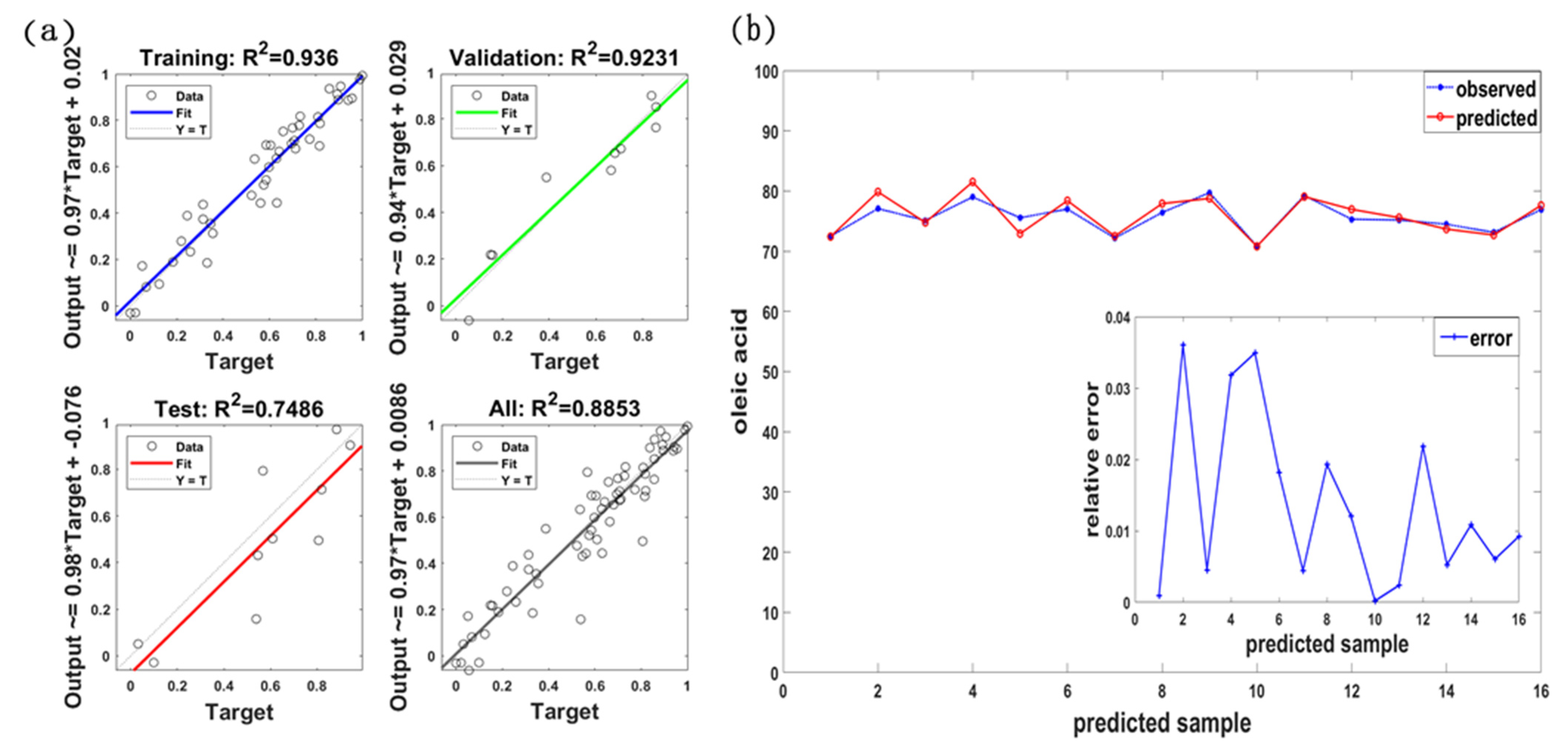
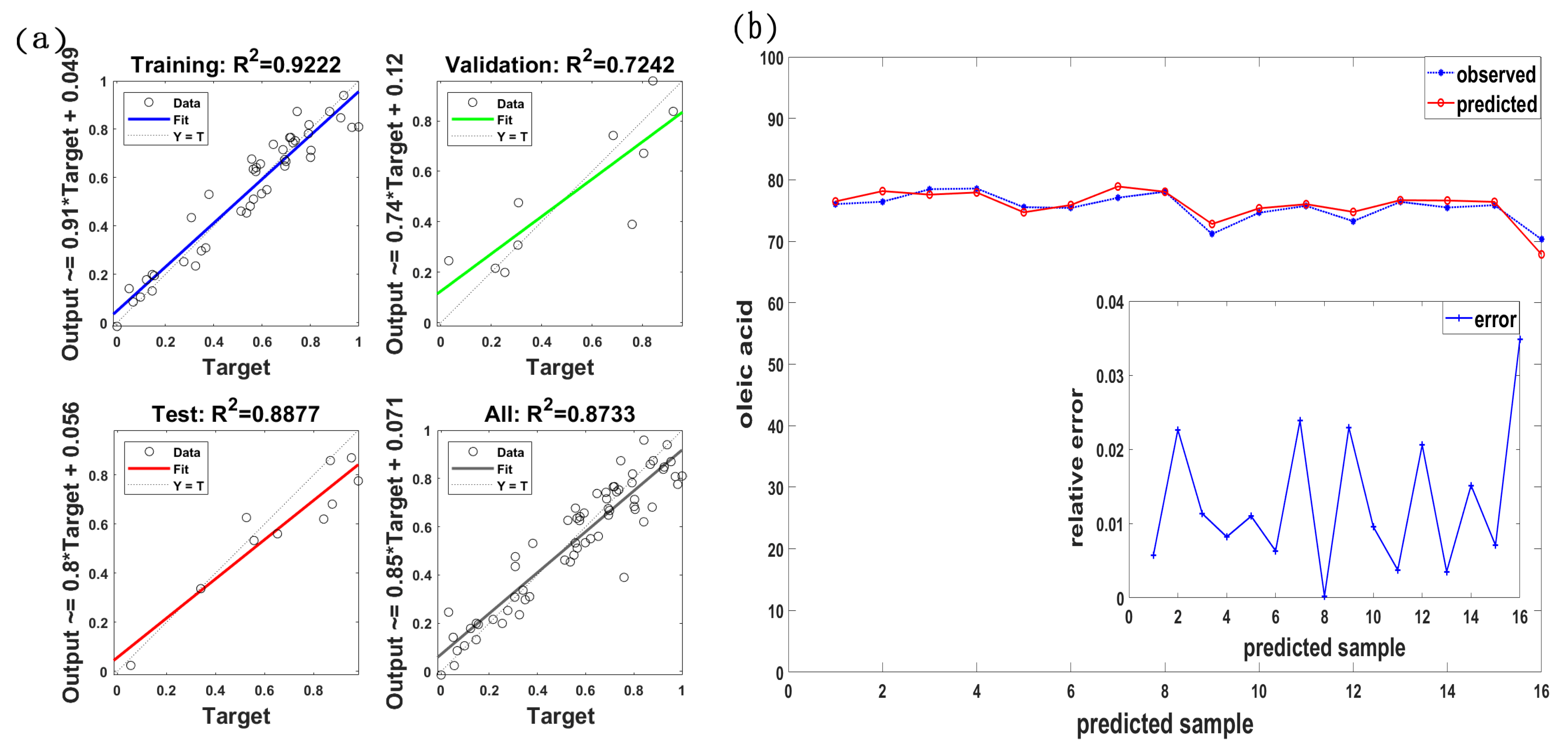
3.3. Estimation Model of Oleic Acid Content
4. Conclusions
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Wang, L.J.; Huang, L.J.; Chen, X.L.; Zhang, Z.Q. Effects of Different Cultivation Methods on Quality of High Oleic Acid Rape. Crop Res. 2019, 33, 274–277, (In Chinese with English abstract). [Google Scholar]
- Li, S.M.; Wang, N.H.; Zhang, M.; Liu, C.; Wang, R. Establishment of near infrared analytic model for major fatty acids content of Brassica napus L. Guangdong Agric. Sci. 2014, 41, 16–21, (In Chinese with English abstract). [Google Scholar]
- Liao, G.J.; Jiang, P.; Luo, Y.H.; Xu, N. Correlation between camellia seed aliphatic acid composition and hyperspectral reflectance. J. Hunan Agric. Univ. (Nat. Sci.) 2013, 39, 445–448, (In Chinese with English abstract). [Google Scholar] [CrossRef]
- Jin, H.L.; Ma, Y.S.; Li, L.L.; Cheng, J.H. Rapid and non-destructive determination of oil content of peanut (Arachis hypogaea L.) using hyperspectral imaging analysis. Food Anal. Methods 2016, 9, 2060–2067. [Google Scholar] [CrossRef]
- Sundaram, J.; Kandala, C.V.; Holser, R.A.; Butts, C.L.; Windham, W.R. Determination of In-Shell Peanut Oil and Fatty Acid Composition Using Near-Infrared Reflectance Spectroscopy. J. Am. Oil Chem. Soc. 2010, 87, 1103–1114. [Google Scholar] [CrossRef]
- Cheng, Q.; Xiao, G.; Zhang, Z.Q.; Chen, H.; Wang, Y.; Guan, C.Y. Study on the Fatty Acid Composition of Plant Seeds of Winter Rapeseed Based on Hyperspectral. Crop Res. 2017, 31, 265–271, (In Chinese with English abstract). [Google Scholar]
- Zhao, H.J.; Cai, H.; Li, N. Feature extraction method based on multifractal parameters for hyperspectral imagery. J. Beijing Univ. Aeronaut. Astronaut. Sci. 2012, 38, 1317–1320, (In Chinese with English abstract). [Google Scholar]
- Wang, X.Q.; Wang, F.; Liao, G.P.; Guan, C.Y. Multifractal Analysis of Rapeseed Spectrum for Chlorophyll Diagnosis Modeling. Spectrosc. Spectr. Anal. 2016, 36, 3657–3663. [Google Scholar]
- Jiang, S.; Wang, F.; Shen, L.M.; Liao, G.P. Local detrended fluctuation analysis for spectral red-edge parameters extraction. Nonlinear Dyn. 2018, 93, 995–1008. [Google Scholar] [CrossRef]
- Jiang, S.; Wang, F.; Shen, L.M.; Liao, G.P.; Wang, L. Extracting sensitive spectrum bands of rapeseed using multiscale multifractal detrended fluctuation analysis. J. Appl. Phys. 2017, 121, 104702. [Google Scholar] [CrossRef]
- Wang, F.; Fan, Q.; Stanley, H. Multiscale multifractal detrended-fluctuation analysis of two-dimensional surfaces. Phys. Rev. E 2016, 93, 042213. [Google Scholar] [CrossRef] [Green Version]
- Wang, F.; Wang, L.; Chen, Y. A DFA-based bivariate regression model for estimating the dependence of PM2.5 among neighbouring cities. Sci. Rep. 2018, 8, 7475. [Google Scholar] [CrossRef]
- Li, J.; Wang, F.; Li, J.W.; Zou, R.B.; Liao, G.P. Multifractal Methods for Rapeseed Nitrogen Nutrition Qualitative Diagnosis Modeling. Int. J. Biomath. 2016, 9, 1650064. [Google Scholar] [CrossRef]
- Wang, F.; Liao, G.; Li, J.H.; Li, X.C.; Zhou, T.J. Multifractal detrended fluctuation analysis for clustering structures of electricity price periods. Physical A 2013, 392, 5723–5734. [Google Scholar] [CrossRef]
- Sun, J.Q.; Zhang, Z.W.; Ai, W.W. Application of BP Neural Network in the Prediction of Oilseed Rape Florescence. Meteorol. Environ. Sci. (Chin.) 2019, 42, 22–26, (In Chinese with English abstract). [Google Scholar]
- Wang, D.C.; Wang, J.H.; Jin, N.; Wang, Q.; Li, C.J.; Huang, J.F.; Wang, Y.; Huang, F. ANN-based wheat biomass estimation using canopy hyperspectral vegetation indices. Trans. Chin. Soc. Agric. Eng. 2008, 24, 196–201, (In Chinese with English abstract). [Google Scholar]
- Yao, F.Q.; Zhang, Z.H.; Yang, R.Y.; Sun, J.W.; Cui, S.F. Hyperspectral models for estimating vegetation chlorophyll content based on red edge parameter. Trans. Chin. Soc. Agric. Eng. 2009, 25, 123–129, (In Chinese with English abstract). [Google Scholar]
- Chen, L.; Huang, J.F.; Wang, F.M.; Tang, Y.L. Comparison between back propagation neural network and regression models for the estimation of pigment content in rice leaves and panicles using hyperspectral data. Int. J. Remote Sens. 2007, 28, 3457–3478. [Google Scholar] [CrossRef]
- Rumelhart, D.; Hinton, G.; Williams, R. Parallel distributed processing: Explorations in the microstructure of cognition. Volume 1. Language 1986, 63, 45–76. [Google Scholar]
- Qi, B.; Zhang, N.; Zhao, T.J.; Xing, G.N.; Zhao, J.M.; Gai, J.Y. Prediction of Leaf Area Index Using Hyperspectral Remote Sensing in Breeding Programs of Soybean. Acta Agron. Sin 2015, 41, 1073–1085, (In Chinese with English abstract). [Google Scholar] [CrossRef]
- Li, Y.Y.; Chang, Q.R.; Liu, X.Y.; Yan, L.; Luo, D.; Wang, S. Estimation of maize leaf SPAD value based on hyperspectrum and BP neural network. Trans. Chin. Soc. Agric. Eng. 2016, 32, 135–142, (In Chinese with English abstract). [Google Scholar]
- Hu, Z.Z.; Pan, C.D.; Xiao, B.; Pan, X. Spectral characteristic parameter-based models for foliar nitrogen concentration estimation of Juglans regia. Trans. Chin. Soc. Agric. Eng. 2015, 31, 180–186, (In Chinese with English abstract). [Google Scholar]
- Guo, P.T.; Su, Y.; Cha, Z.Z.; Lin, Q.H.; Luo, W.; Lin, Z.M. Prediction of leaf phosphorus contents for rubber seedlings based on hyperspectral sensitive bands and back propagation artificial neural networks. Trans. Chin. Soc. Agric. Eng. 2016, 32, 177–183, (In Chinese with English abstract). [Google Scholar]
- Filella, I.; Penuelas, J. The red edge position and shape as indicators of plant chlorophyll content, biomass and hydric status. Int. J. Remote Sens. 1994, 15, 1459–1470. [Google Scholar] [CrossRef]
- Cheng, D.; Liu, Y.M.; Li, J.Z.; Mo, C.H. Extraction of spectral difference characteristics of Stellera chamaejasme in Qilian County of Qinghai Province, Northwest China. J. Appl. Ecol. 2015, 26, 2307–2313, (In Chinese with English abstract). [Google Scholar]
- An, H.B.; Li, F.; Zhao, M.L.; Liu, Y.J. Optimized Spectral Indices Based Estimation of Forage Grass Biomass. Spectrosc. Spectr. Anal. 2015, 35, 3155–3160. [Google Scholar]
- Richardson, A.J.; Wiegand, C.L. Distinguishing vegetation from soil background information. Photogramm. Eng. Remote Sens. 1977, 43, 1541–1552. [Google Scholar]
- Gutiérrez-Rodríguez, M.; Escalante-Estrada, J.A.; Rodriguez, M.T.; And, G.; Reynolds, M.P. Canopy reflectance índices and its relationship with yield in common bean plants (Phaseolus vulgaris L.) with phosphorous supply. J. Agric. Biol. 2006, 2, 203–207. [Google Scholar]



| Number | Min/% | Max/% | Mean/% | Standard Derivation/% | Coefficient of Variation |
|---|---|---|---|---|---|
| 80 | 65.827 | 80.085 | 75.176 | 3.753 | 0.0499 |
| Parameters | Symbol | Description | Ref |
|---|---|---|---|
| Multifractal feature | MF- | Generalized Hurst exponent | [8,9,10] |
| MF- | Generalized Hurst exponent | [8,9,10] | |
| MF- | Generalized Hurst exponent | [8,9,10] | |
| MF- | Generalized Hurst exponent | [8,9,10] | |
| MF- | Span of h(q) | [8,9,10] | |
| MF- | Maximum of index | [8,9,10] | |
| MF- | Minimum of index | [8,9,10] | |
| MF- | Span of | [8,9,10] | |
| Sensitive band | SB-R | Sensitive band of reflectance | [23] |
| SB-DR | Sensitive band of the first derivative reflectance | [23] | |
| Trilateral Parameter | TriP-rep | Red edge position | [24] |
| TriP-Dr | Red edge amplitude | [24] | |
| TriP-SDr | Red edge area | [24] | |
| TriP-yep | Yellow edge position | [25] | |
| TriP-Dy | Yellow edge amplitude | [25] | |
| TriP-SDy | Yellow edge area | [22] | |
| TriP-bep | Blue edge position | [25] | |
| TriP-Db | Blue edge amplitude | [25] | |
| TriP-SDb | Blue edge area | [22] | |
| Spectral index | SPI-RSI | Ratio spectral index | [26] |
| SPI-DSI | Difference spectral index | [27] | |
| SPI-NDSI | Normalized difference spectral index | [28] |
| Parameter | Correlation Coefficient |
|---|---|
| MF- | 0.7898 ** |
| MF- | 0.7442 ** |
| TriP-SDy | 0.7751 ** |
| TriP-SDr | 0.7442 ** |
| SPI-DSI (R572, R574) | 0.7862 ** |
| SPI-NDSI (R575, R576) | 0.7950 ** |
| SB-R818 | 0.7022 ** |
| SB-DR574 | 0.782 ** |
| Min | Max | Mean | STDEV | |
|---|---|---|---|---|
| MF-h(0) | 1.1976 | 1.8628 | 1.5511 | 0.1269 |
| MF- | 1.2306 | 2.0727 | 1.5900 | 0.1993 |
| TriP-SDy | 0.0176 | 0.5693 | 0.2328 | 0.1490 |
| TriP-SDr | 0.3019 | 1.0011 | 0.6179 | 0.1812 |
| SPI-DSI (R572, R574) | −0.0068 | 0.0118 | 0.0037 | 0.0037 |
| SPI-NDSI (R575, R576) | −0.0012 | 0.0047 | 0.0021 | 0.0017 |
| SB-R818 | 0.2867 | 0.8076 | 0.4937 | 0.1308 |
| SB-DR574 | −0.0002 | 0.0015 | 0.0005 | 0.0004 |
| TRAIN (N = 64) | TEST (N = 16) | |||||
|---|---|---|---|---|---|---|
| R2 | RMSE | RE (%) | R2 | RMSE | RE (%) | |
| {SPI-NDSI(R575, R576)} | 0.7777 | 1.3647 | 1.377 | 0.6799 | 1.55 | 1.728 |
| {MF-h(0), SB-DR574} | 0.8635 | 1.0603 | 1.128 | 0.7971 | 1.3388 | 1.678 |
| {MF-h(0), SB-DR574, SPI-NDSI(R575, R576)} | 0.8753 | 1.0301 | 1.047 | 0.8169 | 1.2922 | 1.497 |
| {MF-h(0), SB-DR574, TriP-SDr, SPI-NDSI(R575, R576)} | 0.8652 | 1.0486 | 1.085 | 0.8012 | 1.2706 | 1.526 |
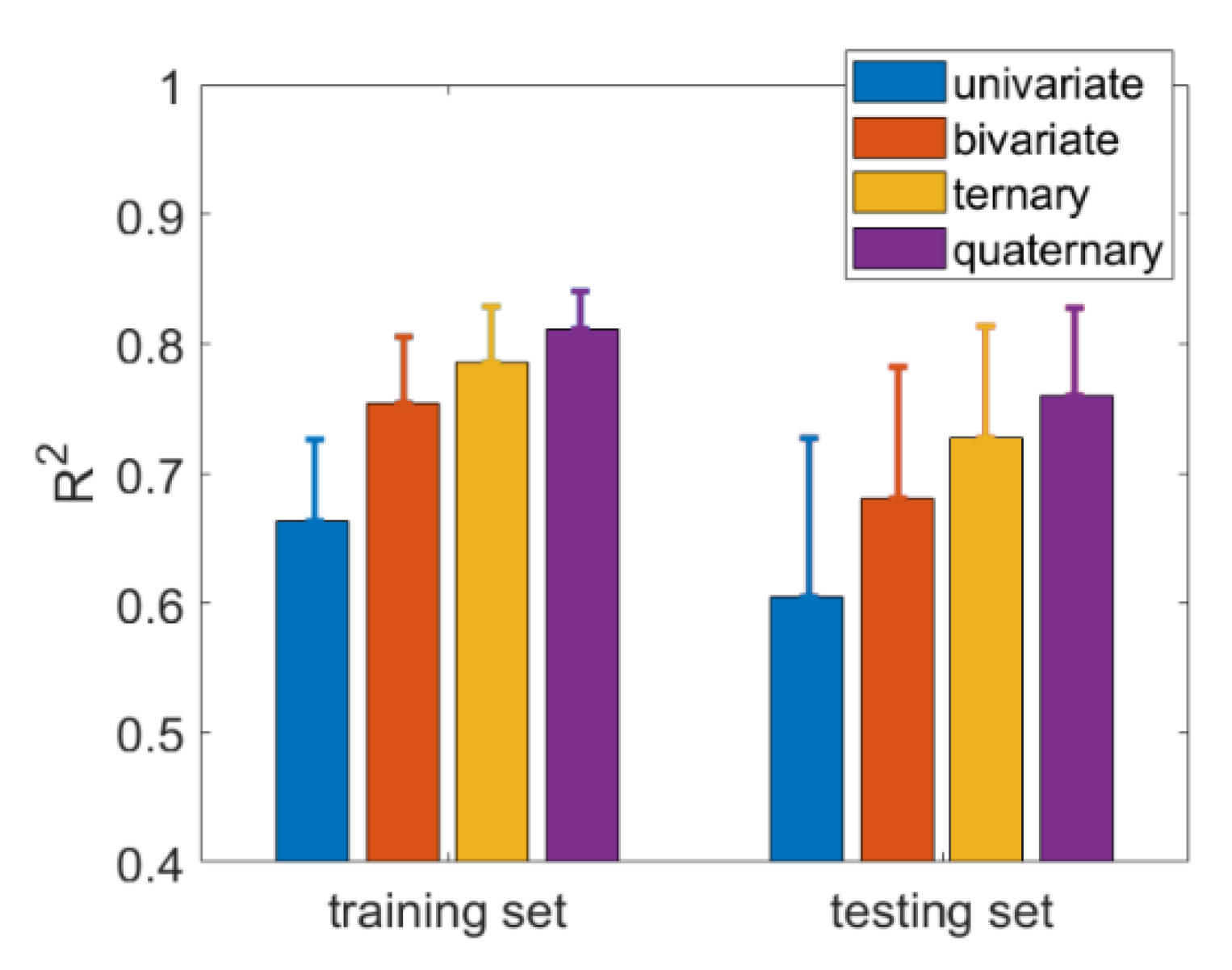
| MF-Features | Training Set (N = 64) | Testing Set (N = 16) | |||||
|---|---|---|---|---|---|---|---|
| R2 | RMSE | RE (%) | R2 | RMSE | RE (%) | ||
| Univariate | with | 0.6232 | 1.748 | 1.913 | 0.6043 | 1.8096 | 2.154 |
| without | 0.6789 | 1.6399 | 1.752 | 0.6201 | 1.8153 | 2.086 | |
| Bivariate | with | 0.7835 | 1.3451 | 1.432 | 0.7152 | 1.5610 | 1.816 |
| without | 0.7122 | 1.5326 | 1.634 | 0.6421 | 1.7567 | 2.274 | |
| Ternary | with | 0.8086 | 1.2498 | 1.353 | 0.7573 | 1.4679 | 1.685 |
| without | 0.7394 | 1.4754 | 1.601 | 0.6685 | 1.6988 | 1.983 | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, F.; Wang, F.; Liao, G.; Lu, X.; Yang, J. Prediction of Oleic Acid Content of Rapeseed Using Hyperspectral Technique. Appl. Sci. 2021, 11, 5726. https://doi.org/10.3390/app11125726
Liu F, Wang F, Liao G, Lu X, Yang J. Prediction of Oleic Acid Content of Rapeseed Using Hyperspectral Technique. Applied Sciences. 2021; 11(12):5726. https://doi.org/10.3390/app11125726
Chicago/Turabian StyleLiu, Fan, Fang Wang, Guiping Liao, Xin Lu, and Jiayi Yang. 2021. "Prediction of Oleic Acid Content of Rapeseed Using Hyperspectral Technique" Applied Sciences 11, no. 12: 5726. https://doi.org/10.3390/app11125726
APA StyleLiu, F., Wang, F., Liao, G., Lu, X., & Yang, J. (2021). Prediction of Oleic Acid Content of Rapeseed Using Hyperspectral Technique. Applied Sciences, 11(12), 5726. https://doi.org/10.3390/app11125726





