Abstract
In order to reduce the environmental burden associated with Cordyceps militaris residue, we conducted a study to explore the feasibility of Cordyceps militaris residue as a silage additive. The experimental treatments included distilled sterile water control (CK), Lactiplantibacillus plantarum (LP, 1 × 105 cfu/g), Cordyceps militaris residue (CM, 4% of fresh matter), and their combination (LP + CM). A total of 48 samples (4 treatments × 4 storage periods × 3 replicates) were prepared for the analysis of fermentation quality, chemical composition, microbial population, and bacterial community composition at days 2, 7, 28, and 45 of ensiling. Results showed that compared with the control, the CM and LP + CM treatments increased the lactic acid bacteria number and lactic acid content (p < 0.05), and a decrease in pH value and NH3-N content was observed (p < 0.05). The bacterial diversity in the CM group was higher and lower than that in the CK group in the early and late stages of fermentation, respectively, and maintained a higher relative abundance of Lactococcus on day 7. Lactobacillus remained the predominant bacterial community at the end of fermentation. Random forest analysis indicated that Lactobacillus was a key determinant of the pH, lactic acid, and acetic acid levels. Consequently, the addition of Cordyceps militaris residue alone, or combined with Lactiplantibacillus plantarum, improved the quality of alfalfa silage by increasing lactic acid and lowering NH3-N, providing a new approach for its development and utilization.
1. Introduction
Cordyceps militaris is a common medicinal fungus belonging to the phylum Ascomycota [1]. C. militaris contains several active ingredients, which are used to produce medicines and health products. Nowadays, C. militaris has gained growing recognition as a potential alternative to Cordyceps sinensis due to its comparable chemical capabilities and medicinal properties [2]. According to the market analysis report, global C. sinensis and C. militaris market was estimated to be valued at USD 1072.5 million in 2022 and was predicted to grow by 10.9% CAGR (compound annual growth rate) from 2022 to 2030 [3]. However, the expansion of artificial C. militaris cultivation has led to generate a large amount of solid medium residues and mycelial waste [4]. These residues are classified as waste and managed through conventional disposal methods such as incineration and landfilling, both of which lead to significant environmental concerns through either direct pollution or long-term ecological impacts [5,6,7,8]. Research has shown that many active substances remain in the residues after the processing of traditional Chinese medicines, and these can be repurposed as silage additives to enhance the fermentation quality and increase the feed value [9]. The application of herbal residues, astragalus residues, Rosa roxburghii pomace, and sea buckthorn pomace improved the fermentation quality of alfalfa (Medicago sativa L.) silage by means of favoring lactic acid bacteria activity and suppressing undesirable microorganisms [10,11,12,13]. Research indicates that C. militaris boasts various bioactive ingredients, including cordycepin, cordyceps polysaccharides, cordycepin acid, and sterols [14,15,16], which makes its residue suitable for development into additives. The development and utilization of C. militaris residue as a silage additive demonstrate significant environmental and economic benefits.
Alfalfa is widely used to feed livestock as a forage with high protein and abundant vitamins and minerals that animals need. However, its low dry matter and water-soluble carbohydrate contents, combined with a strong buffering capacity, make it notoriously challenging to ensile. During the fermentation of alfalfa silage, lactic acid bacteria are often unable to competitively dominate the microbial community and inhibit the reproduction of undesirable microorganisms, which can result in the failure of alfalfa ensiling [17]. Lactic acid bacteria has been applied in the production of silage, in particular, Lactiplantibacillus plantarum (formerly Lactobacillus plantarum) [18]. Guo et al. [19] found that inoculating with L. plantarum could alter the microbial composition and succession of alfalfa silage. Yang et al. [20] demonstrated that inoculating alfalfa silage with L. plantarum could enhance lactic acid bacteria proliferation, enabling them to rapidly dominate the bacterial community. L. plantarum inoculation in alfalfa silage significantly enhanced lactic acid bacteria dominance while effectively suppressing the growth and reproduction of Enterobacteria [21]. Taken together, the inoculation of L. plantarum inhibits the activity of putrefying bacteria and ensures the proliferation and competitive advantage of lactic acid bacteria, thus playing an integral role in successful alfalfa silage. Meanwhile, the bioactive compound cordycepin, present in C. militaris residue, can inhibit the growth of undesirable microorganisms such as Bacillus subtilis, Escherichia coli, and Staphylococcus aureus [22,23,24]. Drawing from these findings, we hypothesized that including C. militaris residue in alfalfa at ensiling could yield favorable impacts on the proliferation of lactic acid bacteria and help thwart the reproduction of harmful microorganisms. To our knowledge, no research on using C. militaris residue as a silage additive has previously been reported. Therefore, to access the impact of C. militaris residue, L. plantarum, and their combinations as silage additives on the fermentation property and bacterial community in alfalfa silage.
2. Materials and Methods
2.1. Silage Preparation
Alfalfa (Juneng No. 2) planted in an approximately 300 m2 experimental field in the Baicaoyuan Research Station (41°50′ N, 123°34′ E) of Shenyang Agricultural University (Liaoning, China) was managed with fertilization and irrigation and harvested through manually cutting at a location approximately 5 cm above ground at 10:00 a.m. Using a commercial grass chopper (Donghong No. 1, Donghong Mechanical Equipment Co., Ltd., Zhengzhou, China), the harvested alfalfa was processed into 1–2 cm segments. The experimental setup included four treatment groups: (i) sterile water control (CK); (ii) Lactiplantibacillus plantarum (LP, 1 × 105 cfu/g of fresh matter (FM)); (iii) C. militaris residues (CM, 4% of FM, Shenyang Ivy Biotechnology Co., Ltd.,. Shenyang, China); and (iv) Lactiplantibacillus plantarum and C. militaris residue (LP + CM). The sterile water containing additives was evenly sprayed onto the materials. Then, the mixed forage was manually packed into polyethylene bags (20 cm × 30 cm) and vacuum-sealed. There were 12 bags for each treatment, among which triplicate bags were opened and sampled after 2, 7, 28, and 45 days of ensiling, respectively. A total of 48 samples (4 treatments × 4 storage periods × 3 replicates) were prepared for analysis of fermentation quality, chemical composition, microbial population, and bacterial community composition.
2.2. Fermentation Quality and Nutrient Composition Analyses
To determine the dry matter (DM) content, fresh and ensiled samples were measured following a 48-h drying period at 65 °C. The determination of neutral detergent fiber (NDF) content and acid detergent fiber (ADF) content adhered to the method of Van Soest et al. [25], utilizing an ANKOM A200i fiber analyzer (ANKOM Technology Corp., Fairport, NY, USA). For the total nitrogen (TN) content, the Kjeldahl method [26] was followed, utilizing an autoanalyzer (Kjeltec 8400; FOSS Co., Ltd., Hillerød, Denmark). The total nitrogen (TN) values were subsequently multiplied by 6.25 to determine the crude protein (CP) content. Water-soluble carbohydrate (WSC) content was determined using the colorimetric method after fully reacting with anthrone reagents [27].
A 20-g sample of silage was combined with 180 mL of sterile distilled water and then refrigerated at 4 °C for 12 h. Following this, the pH of the resulting extract was measured with a PB-10 pH meter (Sartorius Group, Goettingen, Germany) [28]. Using high-performance liquid chromatography, the organic acid concentrations were analyzed, in accordance with the method of Bai et al. [29]. The ammonia nitrogen (NH3-N) levels were measured via colorimetry using the phenol–sodium hypochlorite method [30]. A 10-g sample of silage was blended with 90 mL of distilled water, agitated at 180 rpm for 30 min, and then serially diluted. Yeasts and lactic acid bacteria were counted by placing them on rose Bengal medium and MRS agar (Beijing Aoboxing Bio-tech Co., Ltd., Beijing, China) [31].
2.3. Bacterial Community Analysis
Subsamples (10 g) of silage samples were shaken well with 90 mL of sterile distilled water using a cryogenic oscillator (THZ-98C, Shanghai Yiheng Scientific Instrument Co., Ltd., Shanghai, China), maintained at 4 °C and 180 rpm, for a duration of 30 min. The mixtures were then strained through quadruple-layered sterile gauze. The resulting filtrate underwent centrifugation at 8000 rpm for 15 min in a refrigerated centrifuge (ST 16R, Thermo Fisher Scientific, Inc., Waltham, MA, USA) maintained at 4 °C to concentrate the sediment. These sediments were then dispatched for high-throughput sequencing analysis [32].
Following the guidelines provided by the manufacturer, the E.Z.N.A.® soil DNA Kit (Omega Bio-tek, Norcross, GA, USA) was utilized to isolate DNA from both the alfalfa and silage specimens. A polymerase chain reaction (PCR) was then conducted to amplify the V3–V4 sections of the bacterial 16S rRNA gene using tailored forward primer 341F (5′-CCTAYGGGRBGCASCAG-3′) and reverse primer 806R (5′-GGACTACNNGGGTATCTAAT-3′). The PCR product was retrieved from a 2% agarose gel, subjected to purification with the PCR Clean-Up Kit (YuHua, Shanghai, China), and subsequently quantified using a Qubit 4.0 fluorometer (Thermo Fisher Scientific, USA). The purified PCR products underwent sequencing on the Illumina Nextseq2000 platform (Illumina, San Diego, CA, USA). Quality checking of the paired-end raw sequencing data was conducted using fastp, followed by sequence assembly with FLASH (FLASH 1.2.11). Subsequently, UPARSE (UPARSE 7.1) was employed to cluster sequences at 97% similarity into operational taxonomic units (OTUs). Alpha diversity and beta diversity metrics were employed to assess the microbial diversity and community structure variations across samples. The resemblance between microbial populations in various samples was evaluated using principal coordinate analysis (PCoA). To identify significant variations in species composition across multiple sample groups, the Kruskal–Wallis H test was performed. R studio was used to analyze the OTU values of each sample genus level, and then AI software (version 26.0.1) was used to build the random forest model. The sequencing data have been deposited into the NCBI Sequence Read Archive database (accession: PRJNA750590).
2.4. Statistical Analysis
The experimental data were tested for normality and homogeneity of variance using SPSS (SPSS 22.0 program, SPSS Inc., Chicago, IL, USA), meeting the prerequisite requirements of normal distribution and homogeneity of variance for the two-way analysis of variance (ANOVA).The effects of additives, ensiling duration, and their interaction on fermentation quality, microbial counts, and bacterial community indices of silage were analyzed using two-way ANOVA in SPSS. The statistical model was Yıjh = µ + αι + βj + αβıj + εıjh, where Yıjh is an observation, µ is the overall mean, αι is the effect of the additives (ı = CK, LP, CM, LP + CM), βj is the number of ensiling days (j = 2, 7, 28, 45), αβıj is the additives × the number of ensiling days interaction, and εıjh is the error. Significant differences were determined using Duncan multiple range tests, with p < 0.05 considered statistically significant.
3. Results
3.1. Chemical and Microbial Characteristics of Alfalfa and Cordyceps militaris Residue Before Ensiling
The DM and CP contents were notably higher in the C. militaris residue compared with fresh alfalfa (p < 0.05) (Table 1). Conversely, the NDF and ADF content in C. militaris residue was significantly lower than that in fresh alfalfa (p < 0.05). C. militaris residue exhibited lower yeast counts compared with fresh alfalfa (p < 0.05).

Table 1.
Chemical and microbial characteristics of fresh alfalfa and Cordyceps militaris residue.
3.2. Fermentation Quality of Alfalfa Silage
During the second day of ensiling, the pH value was lower in the CM and LP + CM groups than that in the CK group (p < 0.05) (Table 2). The alfalfa silage treated with LP + CM showed a marked increase in lactic acid concentration over the CK group during the ensiling process (p < 0.05). After 7d of ensiling, the CM and LP + CM groups exhibited a significantly lower acetic acid content compared with the CK group (p < 0.05). After 28 days of ensiling, the NH3-N concentration in both the CM and LP + CM groups was substantially lower than in the CK group (p < 0.05). At 45 days of ensiling, the LP, CM, and LP + CM silages all demonstrated substantially greater lactic acid bacterial counts than the CK group (p < 0.05).

Table 2.
The effects of adding Cordyceps militaris residue on the fermentation quality of alfalfa silage during 45 days of ensiling.
3.3. Chemical Composition of Alfalfa Silage
The DM levels were notably greater in the CM and LP + CM treatments compared with the CK and LP groups (p < 0.05) (Table 3). The CP content in the CM and LP + CM treatments substantially exceeded that in the CK group (p < 0.05). The WSC content showed a marked increase in the CM and LP + CM groups relative to both CK and LP (p < 0.05).

Table 3.
The effects of adding Cordyceps militaris residue on the chemical composition of alfalfa silage after 45 days of ensiling.
3.4. The Microbial Community of Alfalfa Silage During Ensiling
The Chao index was noticeably higher in the CM group compared with the CK group at day 2 of ensiling (p < 0.05) (Table 4). By day 7 of ensiling, the Shannon index was significantly greater in the CM and LP + CM groups compared with CK (p < 0.05). With the extension of ensiling time, the alpha diversity of all treatment groups showed a decreasing trend.

Table 4.
Effect of additives and ensiling days on the bacterial alpha diversity of alfalfa silage during 45 days of ensiling.
The principal coordinate analysis of the alfalfa silage microbial communities at different fermentation stages showed that PC1 contributed 65.72% of the total variance, and PC2 explained 12.68% of the total variance (Figure 1). The CM and LP + CM microbial communities showed clear separation from CK within the initial 7 days. As the silage duration increased, microbial populations within each treatment largely stabilized.
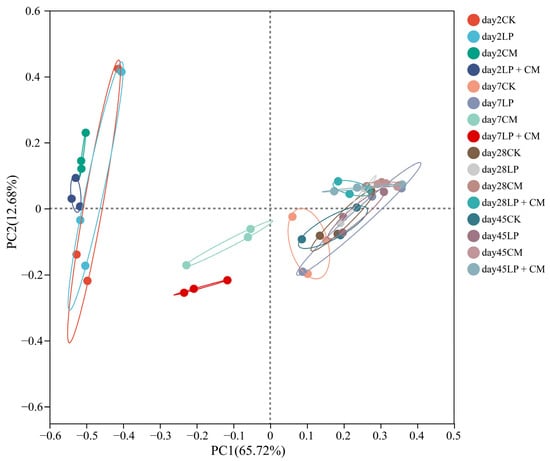
Figure 1.
Principal coordinate analysis of the bacterial community in alfalfa silage after ensiling for 2, 7, 28 and 45 days (CK, control; LP, L. plantarum; CM, Cordyceps militaris residue; LP + CM, L. plantarum and Cordyceps militaris residue).
By the second day of ensiling, the dominant phyla across all treatment groups were Firmicutes and Proteobacteria. After a week of fermentation, Firmicutes emerged as the predominant phylum within the microbial population (Figure 2A). By day 2 of the ensiling process, the predominant bacterial genera were primarily consisted of Weissella and Lactococcus. After seven days of fermentation, Lactobacillus and Weissella dominated all treatment groups. Following 28 days of ensiling, Lactobacillus emerged as the predominant bacterial community across all treatment groups (Figure 2B). On day 7 of ensiling, the LP + CM group showed a marked decline in Lactobacillus abundance compared with the CK group (p < 0.05) (Figure 2C). The CM group had a substantially greater abundance of Lactococcus than the other groups (p < 0.05). At 45 days of ensiling, the Weissella abundance in both the CM and LP + CM treatments was significantly reduced relative to the CK group (p < 0.05).
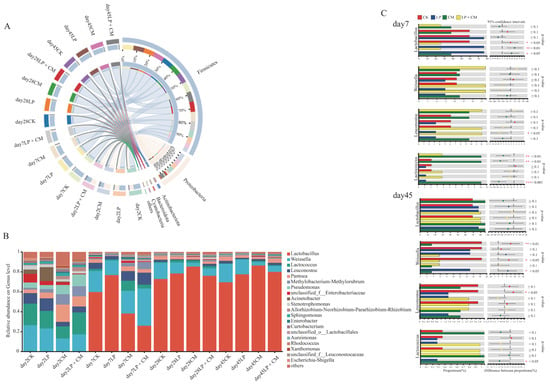
Figure 2.
Changes in the relative abundances of bacteria at the phylum level (A), genus level (B), and the significance of intergroup differences (C) in alfalfa silage after ensiling for 2, 7, 28, and 45 days (CK, control; LP, L. plantarum; CM, Cordyceps militaris residue; LP + CM, L. plantarum and Cordyceps militaris residue).
The pH levels exhibited an inverse relationship with Lactobacillus abundance, whereas they were positively associated with the abundances of Weissella, Lactococcus, and Leuconostoc (p < 0.05) (Figure 3). However, lactic acid, acetic acid, propylene glycol, ethanol, and NH3-N contents demonstrated opposite trends.
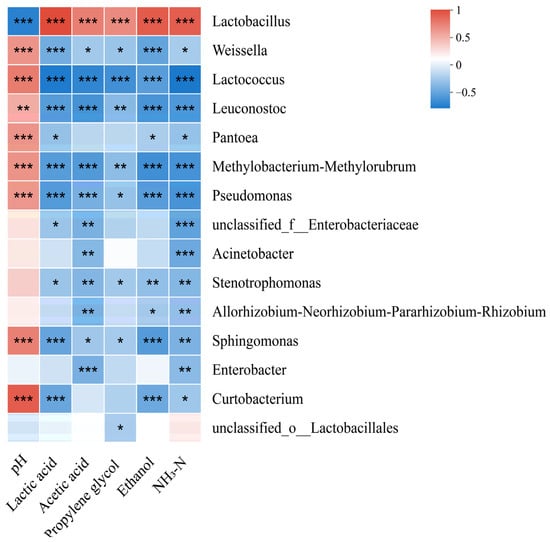
Figure 3.
Correlation of the bacterial community and silage fermentation (NH3-N, ammonia nitrogen; p-values are shown as follows: * p < 0.05; ** p < 0.01; *** p < 0.001).
Lactobacillus, Weissella, Xanthomonas, Acinetobacter, etc. were the primary microorganisms influencing pH (Figure 4A). The main microorganisms affecting lactic acid content were Lactobacillus, Leuconostoc, Lactococcus, Xanthomonas, and others (Figure 4B). Lactobacillus, Leuconostoc, Xanthomonas, and Lactococcus had the greatest impact on the acetic acid content (Figure 4C). Among these, Lactobacillus had a significant impact on the pH level as well as on the concentrations of lactic acid and acetic acid.
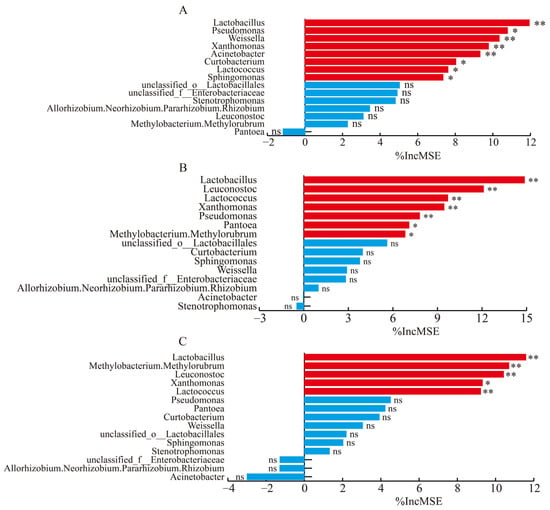
Figure 4.
The importance of microorganisms in the random forest model for pH (A), lactic acid (B), and acetic acid (C). Red bars indicate statistically significant contributions (p < 0.05), while blue bars indicate no statistically significant contributions (p > 0.05). Levels of significance are shown as follows: ns, no significance; * p < 0.05; ** p < 0.01.
4. Discussion
4.1. The Characteristics of Raw Material
The DM content of raw material prior to ensiling plays a pivotal role in determining the silage quality [33]. The DM content in alfalfa fell below the ideal range (30–40%) for silage preparation. The WSC content of alfalfa and C. militaris residue was 8.76 and 6.90% DM, falling within the recommended range for ensuring silage quality [34]. The lactic acid bacteria population in fresh alfalfa exceeded 105 cfu/g, ensuring the successful ensiling [31,32,33,34,35]. The amount of yeast in the C. militaris residue was lower than that in alfalfa, which is related to a high DM content. The higher DM content can effectively suppress the proliferation of undesirable microorganisms [36].
4.2. Fermentation Quality and Chemical Composition of Alfalfa Silage
The CM and LP + CM groups exhibited a lower pH level compared with the CK group, demonstrating that C. militaris residue supplementation can effectively enhance alfalfa silage fermentation. This outcome may be due to the increased number of lactic acid bacteria after adding C. militaris residue, thereby producing more lactic acid, ultimately lowering the pH [10].Moreover, the LP treatment group exhibited pH levels comparable to the control group, and this pH consistency was maintained throughout the ensiling process. This finding aligns with Ni et al. [13], suggesting that overly high moisture in alfalfa could hinder the pH-modulating effect of exogenous additives.
The pH decline in alfalfa silage results from the accumulation of fermentation acids, primarily lactic acid. Lactic acid is mainly produced by lactic acid bacteria consuming WSC [37]. In this study, the addition of C. militaris residue increased the lactic acid content compared with the CK group. That result was similar with the previous report, in which the bioactive elements in sea buckthorn pomace could boost the metabolic prowess of L. plantarum, which in turn results in greater lactic acid production in the silage [38]. Some bioactive components in C. militaris residue might also elicit comparable effects. Acetic acid is a product of hetero-fermentation during the ensiling process [39]. Research indicates that a high WSC content in silage could can slow the transition from homofermentative to heterofermentative processes [40]. In this study, the CM and LP + CM treatments demonstrated a notably greater WSC content than the CK group, which is the reason for the decrease in acetic acid levels observed in the CM and LP + CM groups compared with the CK group. NH3-N mainly results from the breakdown of CP caused by undesirable microorganisms [41]. After a 28-day period of silage, the NH3-N levels in both the CM and LP + CM treatments were significantly lower compared with the CK group, suggesting that adding C. militaris residue could effectively reduce protein degradation. This effect results from the lower pH levels suppressing harmful microbe proliferation, which in turn decreases its degradation of CP [42]. After 2 days of ensiling, the yeast counts in each treatment group rapidly decreased, which was due to acetic acid inhibiting the growth of yeast [43].
During the ensiling process, the main reason for DM loss during silage is the growth of harmful microorganisms including yeasts, molds, and Brucella [44]. Higher DM retention in alfalfa silage for the CM and LP + CM groups, as opposed to CK and LP, suggest that CM and LP + CM helped minimize DM loss during fermentation. Li et al. [45] also observed comparable results, noting that the incorporation of herbal residues inhibited harmful microbe proliferation, leading to decreased DM losses. In addition, the high dry matter content in C. militaris residue could also contribute to this result. Lee’s study [46] demonstrated that native proteolytic enzymes undergo denaturation under acidic conditions (pH < 4.5), thereby effectively suppressing protein degradation and improving the silage fermentation quality. This research provides a plausible explanation for the higher CP content in CM and LP + CM groups compared with the CK group. In this study, the water-soluble carbohydrate content in the alfalfa silage samples from the CM and LP + CM groups was higher when compared with those from the CK and LP groups. This striking difference could be explained by the swift drop in pH levels, which effectively curbed the proliferation of harmful microorganisms. This suppression, in turn, limited their consumption of WSC, leading to the conservation of a greater amount of WSC. Chen et al. [10] also observed comparable results, noting that sea buckthorn pomace supplementation effectively preserved more WSC by restricting WSC utilization by undesirable microbial populations.
4.3. The Microbial Community of Alfalfa Silage
Each group showed near-complete coverage (above 0.99), demonstrating that our results accurately depicted the sampled biological community [47]. In the initial phase of ensiling, the Shannon and Chao diversity indices were elevated in the CM and LP + CM groups compared with the CK group. This result was similar to Wu et al. [48], who reported that adding N. cadamba leaf meal to stylo silage could increase the bacterial diversity in the initial phase of ensiling. As the duration of ensiling increased, the alpha diversity of all treatment groups showed a decreasing trend, which may be due to the gradual dominance of lactic acid bacteria replacing the originally diverse microbial population in the silage [17]. Early on in the ensiling process, the CM and LP + CM groups exhibited a separation from the CK group, indicating a shift in the composition of microbial communities. As the silage period progressed, the microbial communities in each group started to converge. One possible reason is that as the pH declined across all treatment groups, Lactobacillus gradually dominated the bacterial community, resulting in convergent microbial compositions among treatments [49].
The main bacteria found in alfalfa silage are Firmicutes and Proteobacteria [50]. In this study, Firmicutes and Proteobacteria initially dominated the microbial population during the early fermentation phase. As the fermentation process continued, Firmicutes emerged as the predominant phylum within the microbial community. This is related to the anaerobic and acidic conditions developed during silage, which can promote the growth of Firmicutes [51,52]. During the initial phase of ensiling, Weissella and Lactococcus participates in the fermentation process, but as the pH level drops in silage, their proliferation slows, leading to their eventual replacement by more acid-tolerant lactic acid bacteria [53]. Our study yielded comparable findings, demonstrating that the dominant bacterial genera primarily consisted of Weissella and Lactococcus at day 2 of ensiling, whereas Lactobacillus had taken over as the dominant genus after one week of ensiling. The CM treatment group showed a markedly higher abundance of Lactococcus compared with the other treatments at 7 days of ensiling. This revealed that C. militaris residue supplementation in alfalfa silage effectively maintained a higher relative abundance of Lactococcus. Peng et al. [42] reported similar findings, who observed that the addition of sea buckthorn pomace could preserve a higher population of Lactococcus during the initial stages of the silage process. Cai et al. [35] carried out a study showing that coccoid lactic acid bacteria (primarily Lactococcus, Leuconostoc, and Weissella) exhibit vigorous growth during the early fermentation stage. At day 7 of ensiling, the dominant bacteria in the LP + CM group was Weissella, indicating that it was in the early stage of fermentation. This is the reason why the abundance of Lactobacillus in the LP + CM group was lower than that in the CK group (p < 0.05). This result indicates that incorporating the C. militaris residue did not facilitate Lactobacillus in quickly becoming the dominant species within the bacterial community, aligning with the earlier observations made by Chen et al. [10]. Based on this analysis, the addition of C. militaris residue boosted bacterial diversity early in the silage process. However, it could not cause Lactobacillus to rapidly become the dominant strain, thereby prolonging the transition of the lactic acid bacteria from acid-intolerant to acid-tolerant types. Following 28 days of ensiling, the pH reduction inhibited the growth of Weissella, leading to its eventual replacement by acid-resistant Lactobacillus late in silage [54]. At the 45-day mark of the silage process, the Weissella abundance in the CM and LP + CM groups showed a significant decrease compared with the CK group. There were no significant differences in the abundance of Weissella between the LP and CK groups. This suppression may stem from reduced pH in the CM and LP + CM groups, which further restricts the growth of Weissella [43].
The pH value was inversely related to Lactobacillus abundance but positively correlated the abundances of Weissella, Lactococcus, and Leuconostoc. Lactic acid and acetic acid demonstrated opposite trends. Weissella and Lactococcus have poor tolerance to acidic environments and cannot survive when the pH decreases [53]. Lactobacillus plays a pivotal role in the production of lactic acid within alfalfa silage, ultimately driving pH reduction [55,56]. Thus, Lactobacillus is pivotal in lowering pH levels in the late phase of silage fermentation.
During the ensiling process, Lactobacillus gradually dominate the microbial community, promoting lactic acid accumulation and thus reducing the pH [57]. Due to the diversity of sugars in the fermentation substrate, acetic acid serves as another critical metabolic product of Lactobacillus fermentation besides lactic acid [17]. This is the reason why Lactobacillus served as pivotal determinants in influencing the pH level as well as the concentrations of lactic acid and acetic acid in the random forest analysis.
5. Conclusions
In comparison to the CK group, both the CM and LP + CM treatment groups exhibited enhanced lactic acid accumulation and reduced pH levels, indicating enhanced fermentation quality. The incorporation of C. militaris residue decreased the alpha diversity at the end phase of fermentation. C. militaris residue could preserve more Lactococcus by day 7 of the ensiling process and inhibit the growth of Weissella at 45 days of ensiling. Lactobacillus still dominated the bacterial community in the late stage of fermentation. Thus, C. militaris residue can be applied as a silage additive to enhance the fermentation quality of alfalfa silage, providing a new approach for its development and utilization. Nevertheless, to develop and ensure the use of C. militaris residue as a silage additive, it is necessary to further evaluate the effect of combining C. militaris residue with other microbial strains as well as the aerobic stability of the treated silage and its impact on animal performance.
Author Contributions
Conceptualization, J.S. (Juanjuan Sun), Z.Y. and C.B.; Methodology, X.L., J.S. (Juanjuan Sun), Z.Y. and C.B.; Formal analysis, L.W.; Investigation, L.W., M.Z., D.G. and X.L.; Data curation, L.W., M.Z., J.S. (Jia Song) and D.G.; Writing—original draft, L.W. and C.B.; Writing—review and editing, J.S. (Juanjuan Sun) and C.B.; Funding acquisition, Z.Y. and C.B. All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by the Natural Science Foundation of Liaoning Province (2024-MSLH-215) and the National Natural Science Foundation of China (31872422).
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
The original contributions presented in this study are included in the article. Further inquiries can be directed to the corresponding author.
Conflicts of Interest
The authors declare no conflicts of interest.
References
- Liu, L.; Chen, Y.; Luo, Q.; Xu, N.; Zhou, M.Z.; Gao, B.; Wang, C.; Shi, Y. Fermenting liquid vinegar with higher taste, flavor and healthy value by using discarded Cordyceps militaris solid culture medium. Lwt-Food Sci. Technol. 2018, 98, 654–660. [Google Scholar] [CrossRef]
- Krishna, K.V.; Balasubramanian, B.; Park, S.; Bhattacharya, S.; Sebastian, J.K.; Liu, W.C.; Pappuswamy, M.; Meyyazhagan, A.; Kamyab, H.; Chelliapan, S.; et al. Conservation of Endangered Cordyceps sinensis Through Artificial Cultivation Strategies of C. militaris, an Alternate. Mol. Biotechnol. 2025, 67, 1382–1397. [Google Scholar] [CrossRef] [PubMed]
- Insights, C.M. Cordyceps Sinensis and Militaris Extract Market Analysis & Forecast 2032. Available online: https://www.coherentmarketinsights.com/market-insight/cordyceps-sinensis-and-militaris-extract-market-2578 (accessed on 31 July 2025).
- Zhu, Z.Y.; Liu, X.C.; Dong, F.Y.; Guo, M.Z.; Wang, X.T.; Wang, Z.; Zhang, Y.M. Influence of fermentation conditions on polysaccharide production and the activities of enzymes involved in the polysaccharide synthesis of Cordyceps militaris. Appl. Microbiol. Biotechnol. 2016, 100, 3909–3921. [Google Scholar] [CrossRef]
- Jin, Z.Q.; Li, Y.L.; Ren, J.H.; Qin, N. Yield, Nutritional Contant, and Antioxidant Activity of Pleurotus ostreatus on Corncobs Supplemented with Herb Residues. Mycobiology 2018, 46, 24–32. [Google Scholar] [CrossRef]
- Lin, Q.Y.; Long, L.K.; Wu, L.L.; Zhang, F.L.; Wu, S.L.; Zhang, W.M.; Sun, X.M. Evaluation of different agricultural wastes for the production of fruiting bodies and bioactive compounds by medicinal mushroom Cordyceps militaris. J. Sci. Food Agric. 2017, 97, 3476–3480. [Google Scholar] [CrossRef]
- Meng, F.J.; Yang, S.G.; Wang, X.; Chen, T.T.; Wang, X.L.; Tang, X.Y.; Zhang, R.J.; Shen, L. Reclamation of Chinese herb residues using probiotics and evaluation of their beneficial effect on pathogen infection. J. Infect. Public Health 2017, 10, 749–754. [Google Scholar] [CrossRef]
- Wu, F.C.; Chen, Y.L.; Chang, S.M.; Shih, I.L. Cultivation of Medicinal Caterpillar Fungus, Cordyceps militaris (Ascomycetes), and Production of Cordycepin Using the Spent Medium from Levan Fermentation. Int. J. Med. Mushrooms 2013, 15, 393–405. [Google Scholar] [CrossRef]
- Tao, G.; Ya, W.; Yi-Lin, Z.; Zhen, Z. The Reutilization Of Herbal Residues. In Proceedings of the 2nd International Conference on Energy and Environmental Protection (ICEEP), Guilin, China, 19–21 April 2013; pp. 2993–2996. [Google Scholar]
- Chen, L.Y.; Qu, H.; Bai, S.Q.; Yan, L.J.; You, M.H.; Gou, W.L.; Li, P.; Gao, F.Q. Effect of wet sea buckthorn pomace utilized as an additive on silage fermentation profile and bacterial community composition of alfalfa. Bioresour. Technol. 2020, 314, 123773. [Google Scholar] [CrossRef]
- Li, M.Y.; Wang, J.C.H.; Cheng, Q.M.; Long, Z.F.; Chen, C.; Xie, Y.X.; Lei, Y.; Chen, Y.L.; Zhao, Y.Y.; He, X.J.; et al. The potential of biogas production and effects of alfalfa silage under the synergistic influence of Lactobacillus acidophilus and Rosa roxburghii pomace waste on the fermentation quality and bacterial community. mSphere 2025, 10, 17. [Google Scholar] [CrossRef]
- Li, X.M.; Chen, F.; Xu, J.J.; Guo, L.N.; Xiong, Y.; Lin, Y.L.; Ni, K.K.; Yang, F.Y. Exploring the Addition of Herbal Residues on Fermentation Quality, Bacterial Communities, and Ruminal Greenhouse Gas Emissions of Paper Mulberry Silage. Front. Microbiol. 2022, 12, 820011. [Google Scholar] [CrossRef] [PubMed]
- Ni, K.K.; Wang, X.K.; Lu, Y.; Guo, L.N.; Li, X.M.; Yang, F.Y. Exploring the silage quality of alfalfa ensiled with the residues of astragalus and hawthorn. Bioresour. Technol. 2020, 297, 122249. [Google Scholar] [CrossRef] [PubMed]
- Barido, F.H.; Lee, S.K. Tenderness-related index and proteolytic enzyme response to the marination of spent hen breast by a protease extracted from Cordyceps militaris mushroom. Anim. Biosci. 2021, 34, 1859–1869. [Google Scholar] [CrossRef] [PubMed]
- Sun, J.B.; An, L.P.; Zhang, Z.X.; Zhao, N.X.; Yuan, G.X.; Du, P.G. Extraction methods and sedative-hypnotic effects of polysaccharide and total flavonoids of Cordyceps militaris. Biotechnol. Biotechnol. Equip. 2018, 32, 498–505. [Google Scholar] [CrossRef]
- Won, S.-Y.; Park, E.-H. Anti-inflammatory and related pharmacological activities of cultured mycelia and fruiting bodies of Cordyceps militaris. J. Ethnopharmacol. 2005, 96, 555–561. [Google Scholar] [CrossRef]
- Mcdonald, P.; Henderson, A.R.; Heron, S.J.E. The Biochemistry of Silage; Chalcombe Publications: Marlow, UK, 1991. [Google Scholar]
- Kung, L.M.; Shaver, R.D.; Grant, R.J.; Schmidt, R.J. Silage review: Interpretation of chemical, microbial, and organoleptic components of silages. J. Dairy Sci. 2018, 101, 4020–4033. [Google Scholar] [CrossRef]
- Guo, X.S.; Ke, W.C.; Ding, W.R.; Ding, L.M.; Xu, D.M.; Wang, W.W.; Zhang, P.; Yang, F.Y. Profiling of metabolome and bacterial community dynamics in ensiled Medicago sativa inoculated without or with Lactobacillus plantarum or Lactobacillus buchneri. Sci. Rep. 2018, 8, 357. [Google Scholar] [CrossRef] [PubMed]
- Yang, F.Y.; Zhao, S.S.; Wang, Y.; Fan, X.M.; Wang, Y.P.; Feng, C.S. Assessment of Bacterial Community Composition and Dynamics in Alfalfa Silages With and Without Lactobacillus plantarum Inoculation Using Absolute Quantification 16S rRNA Sequencing. Front. Microbiol. 2021, 11, 629894. [Google Scholar] [CrossRef]
- Agarussi, M.C.N.; Pereira, O.G.; Silva, V.P.D.; Leandro, E.S.; Santos, S.A. Fermentative profile and lactic acid bacterial dynamics in non-wilted and wilted alfalfa silage in tropical conditions. Mol. Biol. Rep. 2019, 46, 451–460. [Google Scholar] [CrossRef]
- Ahn, Y.J.; Park, S.J.; Lee, S.G.; Shin, S.C.; Choi, D.H. Cordycepin: Selective growth inhibitor derived from liquid culture of Cordyceps militaris against Clostridium spp. J. Agric. Food Chem. 2000, 48, 2744–2748. [Google Scholar] [CrossRef]
- Jiang, Q.; Lou, Z.X.; Wang, H.X.; Chen, C. Antimicrobial effect and proposed action mechanism of cordycepin against Escherichia coli and Bacillus subtilis. J. Microbiol. 2019, 57, 288–297. [Google Scholar] [CrossRef]
- Zhao, S.X.; Zhou, T.S. Biosorption of methylene blue from wastewater by an extraction residue of Salvia miltiorrhiza Bge. Bioresour. Technol. 2016, 219, 330–337. [Google Scholar] [CrossRef]
- Van Soest, P.J.; Robertson, J.B.; Lewis, B.A. Methods for dietary fiber, neutral detergent fiber, and nonstarch polysaccharides in relation to animal nutrition. J. Dairy Sci. 1991, 74, 3583–3597. [Google Scholar] [CrossRef] [PubMed]
- Sun, L.; Na, N.; Li, X.M.; Li, Z.Q.; Wang, C.; Wu, X.G.; Xiao, Y.Z.; Yin, G.M.; Liu, S.B.; Liu, Z.P.; et al. Impact of Packing Density on the Bacterial Community, Fermentation, and In Vitro Digestibility of Whole-Crop Barley Silage. Agriculture 2021, 11, 672. [Google Scholar] [CrossRef]
- Arthur Thomas, T. An automated procedure for the determination of soluble carbohydrates in herbage. J. Sci. Food Agric. 1977, 28, 639–642. [Google Scholar] [CrossRef]
- Zhang, Y.X.; Ke, W.C.; Bai, J.; Li, F.H.; Xu, D.M.; Ding, Z.T.; Guo, X.S. The effect of Pediococcus acidilactici J17 with high-antioxidant activity on antioxidant, α-tocopherol, β-carotene, fatty acids, and fermentation profiles of alfalfa silage ensiled at two different dry matter contents. Anim. Feed. Sci. Technol. 2020, 268, 114614. [Google Scholar] [CrossRef]
- Bai, C.S.; Pan, G.; Leng, R.X.; Ni, W.H.; Yang, J.Y.; Sun, J.J.; Yu, Z.; Liu, Z.G.; Xue, Y.L. Effect of Ensiling Density and Storage Temperature on Fermentation Quality, Bacterial Community, and Nitrate Concentration of Sorghum-Sudangrass Silage. Front. Microbiol. 2022, 13, 828320. [Google Scholar] [CrossRef] [PubMed]
- Feng, Q.X.; Zhang, J.; Ling, W.Q.; Degen, A.A.; Zhou, Y.; Ge, C.Y.; Yang, F.L.; Zhou, J. Ensiling hybrid Pennisetum with lactic acid bacteria or organic acids improved the fermentation quality and bacterial community. Front. Microbiol. 2023, 14, 1216722. [Google Scholar] [CrossRef]
- Cai, Y.; Benno, Y.; Ogawa, M.; Kumai, S. Effect of applying lactic acid bacteria isolated from forage crops on fermentation characteristics and aerobic deterioration of silage. J. Dairy Sci. 1999, 82, 520–526. [Google Scholar] [CrossRef]
- Wang, Q.; Garrity, G.M.; Tiedje, J.M.; Cole, J.R. Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl. Environ. Microbiol. 2007, 73, 5261–5267. [Google Scholar] [CrossRef]
- Yan, Y.H.; Li, X.M.; Guan, H.; Huang, L.K.; Ma, X.; Peng, Y.; Li, Z.; Nie, G.; Zhou, J.Q.; Yang, W.Y.; et al. Microbial community and fermentation characteristic of Italian ryegrass silage prepared with corn stover and lactic acid bacteria. Bioresour. Technol. 2019, 279, 166–173. [Google Scholar] [CrossRef]
- Wang, Y.; Wang, C.; Zhou, W.; Yang, F.Y.; Chen, X.Y.; Zhang, Q. Effects of Wilting and Lactobacillus plantarum Addition on the Fermentation Quality and Microbial Community of Moringa oleifera Leaf Silage. Front. Microbiol. 2018, 9, 1817. [Google Scholar] [CrossRef] [PubMed]
- Cai, Y.; Benno, Y.; Ogawa, M.; Ohmomo, S.; Kumai, S.; Nakase, T. Influence of Lactobacillus spp. from An inoculant and of weissella and leuconostoc spp. from forage crops on silage fermentation. Appl. Environ. Microbiol. 1998, 64, 2982–2987. [Google Scholar] [CrossRef]
- Zhang, X.Y.; Zhao, S.S.; Wang, Y.P.; Yang, F.Y.; Wang, Y.; Fan, X.M.; Feng, C.S. The Effect of Lactiplantibacillus plantarum ZZU203, Cellulase-Producing Bacillus methylotrophicus, and Their Combinations on Alfalfa Silage Quality and Bacterial Community. Fermentation 2023, 9, 287. [Google Scholar] [CrossRef]
- Besharati, M.; Palangi, V.; Salem, A.Z.M.; De Palo, P.; Lorenzo, J.M.; Maggiolino, A. Substitution of raw lucerne with raw citrus lemon by-product in silage: In vitro apparent digestibility and gas production. Front. Vet. Sci. 2022, 9, 1006581. [Google Scholar] [CrossRef]
- Markkinen, N.; Laaksonen, O.; Nahku, R.; Kuldjärv, R.; Yang, B. Impact of lactic acid fermentation on acids, sugars, and phenolic compounds in black chokeberry and sea buckthorn juices. Food Chem. 2019, 286, 204–215. [Google Scholar] [CrossRef]
- Da Silva, E.B.; Smith, M.L.; Savage, R.M.; Polukis, S.A.; Drouin, P.; Kung, L. Effects of Lactobacillus hilgardii 4785 and Lactobacillus buchneri 40788 on the bacterial community, fermentation and aerobic stability of high-moisture corn silage. J. Appl. Microbiol. 2021, 130, 1481–1493. [Google Scholar] [CrossRef]
- Mu, L.; Xie, Z.; Hu, L.; Chen, G.; Zhang, Z. Lactobacillus plantarum and molasses alter dynamic chemical composition, microbial community, and aerobic stability of mixed (amaranth and rice straw) silage. J. Sci. Food Agric. 2021, 101, 5225–5235. [Google Scholar] [CrossRef] [PubMed]
- Kung, L., Jr.; Myers, C.L.; Neylon, J.M.; Taylor, C.C.; Lazartic, J.; Mills, J.A.; Whiter, A.G. The effects of buffered propionic acid-based additives alone or combined with microbial inoculation on the fermentation of high moisture corn and whole-crop barley. J. Dairy Sci. 2004, 87, 1310–1316. [Google Scholar] [CrossRef]
- Peng, S.; Xie, L.L.; Cheng, Y.Y.; Wang, Q.Q.; Feng, L.; Li, Y.; Lei, Y.H.; Sun, Y.F. Effect of Lactiplantibacillus and sea buckthorn pomace on the fermentation quality and microbial community of paper mulberry silage. Front. Plant Sci. 2024, 15, 1412759. [Google Scholar] [CrossRef]
- Muck, R.E.; Nadeau, E.M.G.; McAllister, T.A.; Contreras-Govea, F.E.; Santos, M.C.; Kung, L. Silage review: Recent advances and future uses of silage additives. J. Dairy Sci. 2018, 101, 3980–4000. [Google Scholar] [CrossRef]
- Wang, J.; Chen, L.; Yuan, X.J.; Guo, G.; Li, J.F.; Bai, Y.F.; Shao, T. Effects of molasses on the fermentation characteristics of mixed silage prepared with rice straw, local vegetable by-products and alfalfa in Southeast China. J. Integr. Agric. 2017, 16, 664–670. [Google Scholar] [CrossRef]
- Li, X.; Chen, F.; Wang, X.; Xiong, Y.; Liu, Z.; Lin, Y.; Ni, K.; Yang, F. Innovative utilization of herbal residues: Exploring the diversity of mechanisms beneficial to regulate anaerobic fermentation of alfalfa. Bioresour. Technol. 2022, 360, 127249. [Google Scholar] [CrossRef]
- Lee, M.R.F.; Scott, M.B.; Tweed, J.K.S.; Minchin, F.R.; Davies, D.R. Effects of polyphenol oxidase on lipolysis and proteolysis of red clover silage with and without a silage inoculant (Lactobacillus plantarum L54). Anim. Feed. Sci. Technol. 2008, 144, 125–136. [Google Scholar] [CrossRef]
- He, L.W.; Lv, H.J.; Xing, Y.Q.; Chen, X.Y.; Zhang, Q. Intrinsic tannins affect ensiling characteristics and proteolysis of Neolamarckia cadamba leaf silage by largely altering bacterial community. Bioresour. Technol. 2020, 311, 123496. [Google Scholar] [CrossRef] [PubMed]
- Wu, S.; Gao, L.; Chen, D.D.; Xue, Y.L.; Kholif, A.E.; Zhou, W.; Chen, X.Y.; Zhang, Q. Effects of Forestry Waste Neolamarckia cadamba Leaf Meal as an Additive on Fermentation Quality, Antioxidant Activity, and Bacterial Community of High-Moisture Stylo Silage. Front. Environ. Sci. 2022, 10, 925400. [Google Scholar] [CrossRef]
- Si, Q.; Wang, Z.J.; Liu, W.; Liu, M.J.; Ge, G.T.; Jia, Y.S.; Du, S. Influence of Cellulase or Lactiplantibacillus plantarum on the Ensiling Performance and Bacterial Community in Mixed Silage of Alfalfa and Leymus chinensis. Microorganisms 2023, 11, 426. [Google Scholar] [CrossRef]
- McGarvey, J.A.; Franco, R.B.; Palumbo, J.D.; Hnasko, R.; Stanker, L.; Mitloehner, F.M. Bacterial population dynamics during the ensiling of Medicago sativa (alfalfa) and subsequent exposure to air. J. Appl. Microbiol. 2013, 114, 1661–1670. [Google Scholar] [CrossRef]
- Ali, N.; Wang, S.R.; Zhao, J.; Dong, Z.H.; Li, J.F.; Nazar, M.; Shao, T. Microbial diversity and fermentation profile of red clover silage inoculated with reconstituted indigenous and exogenous epiphytic microbiota. Bioresour. Technol. 2020, 314, 123606. [Google Scholar] [CrossRef]
- Keshri, J.; Chen, Y.R.; Pinto, R.; Kroupitski, Y.; Weinberg, Z.G.; Sela, S. Microbiome dynamics during ensiling of corn with and without Lactobacillus plantarum inoculant. Appl. Microbiol. Biotechnol. 2018, 102, 4025–4037. [Google Scholar] [CrossRef]
- Zhao, M.R.; Zhang, H.Y.; Pan, G.; Yin, H.; Sun, J.J.; Yu, Z.; Bai, C.S.; Xue, Y.L. Effect of exogenous microorganisms on the fermentation quality, nitrate degradation and bacterial community of sorghum-sudangrass silage. Front. Microbiol. 2022, 13, 1052837. [Google Scholar] [CrossRef] [PubMed]
- Ogunade, I.M.; Kim, D.H.; Jiang, Y.; Weinberg, Z.G.; Jeong, K.C.; Adesogan, A.T. Control of Escherichia coli O157:H7 in contaminated alfalfa silage: Effects of silage additives. J. Dairy Sci. 2016, 99, 4427–4436. [Google Scholar] [CrossRef] [PubMed]
- Cai, Y.M.; Du, Z.M.; Yamasaki, S.; Nguluve, D.; Tinga, B.; Macome, F.; Oya, T. Community of natural lactic acid bacteria and silage fermentation of corn stover and sugarcane tops in Africa. Asian-Australas. J. Anim. Sci. 2020, 33, 1252–1264. [Google Scholar] [CrossRef] [PubMed]
- Li, P.; Zhang, Y.; Gou, W.L.; Cheng, Q.M.; Bai, S.Q.; Cai, Y.M. Silage fermentation and bacterial community of bur clover, annual ryegrass and their mixtures prepared with microbial inoculant and chemical additive. Anim. Feed. Sci. Technol. 2019, 247, 285–293. [Google Scholar] [CrossRef]
- Li, D.T.; Xie, H.D.; Zeng, F.Q.; Luo, X.Q.; Peng, L.J.; Sun, X.W.; Wang, X.F.; Yang, C.J. An Assessment on the Fermentation Quality and Bacterial Community of Corn Straw Silage with Pineapple Residue. Fermentation 2024, 10, 242. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).