Abstract
Substances of organic origin are seeing increasing use in agriculture as rich sources of nutrients for plants. The aim of this study was to determine the microbiological contamination of sewage sludge and digestate to assess their safe use as fertilizers in Poland. The assessment of microbial soil, sewage sludge and digestate contamination was based on the total number of mesophilic bacteria and Gram-negative bacteria from the Enterobacteriaceae family. The presence of Escherichia coli and Salmonella spp. was identified via culture and the presence of Enterobacteriaceae species was determined via biochemical and molecular methods. In laboratory conditions, the survival of E. coli in soil fertilized with sewage sludge or digestate inoculated with a reference strain was determined. The average concentration of Enterobacteriaceae in soil, sewage sludge and digestate samples was 1.1 × 104 CFU/g, 9.4 × 105 CFU/g and 5.6 × 106 CFU/g, respectively. Escherichia coli was detected in all sample types. From the soil samples, Serratia, Enterobacter, Pantoea, Citrobacter and Pseudomonas genera were identified the most frequently, while in sewage sludge and digestate, E. coli was predominant. Based on the results of our laboratory experiment, it can be concluded that after three weeks, fertilization with organic waste in acceptable doses does not significantly increase soil contamination with Enterobacteriaceae.
Keywords:
Enterobacteriaceae; soil; sewage sludge; digestate; Escherichia coli; Salmonella; fertilizers 1. Introduction
Soil is a basic reservoir of microorganisms (e.g., bacteria, fungi, viruses, parasites), the diversity of which is the source of mechanisms regulating the impact of pathogens on other organisms [1]. The presence of indicator microorganisms in soil not only reflects the degree of contamination of the soil environment, but also provides information about the potential risk of contamination of agricultural produce and potential threats to human and animal health. In addition to naturally occurring microflora, the soil environment may also contain microorganisms introduced by improper sewage management or the use of contaminated manure, slurry or sewage sludge in agriculture.
Organic wastes contain plant nutrient contents and can be used to fertilize and improve soil properties in both raw and processed forms. Organic wastes are also introduced into the market as fertilizers and soil improvers, provided they meet quality requirements and contamination does not exceed permissible levels. In some cases, agricultural tests are necessary to confirm the suitability of the fertilizer for application to plants or for soil remediation. In addition to several quality parameters regarding minimum nutrient content and maximum heavy metal content, organic wastes and fertilizers cannot exceed the permissible values of biological contamination. Often, despite meeting all other quality requirements, organic wastes and fertilizers are disqualified from use because of the content of bacteria and parasite eggs [2].
In Poland, in accordance with the Regulation of the Minister of the Environment (2015) [3], sewage sludge may be used in agriculture and for land reclamation for agricultural purposes if Salmonella bacteria have not been isolated in a representative 100 g sludge sample and the total number of live eggs of intestinal parasites (e.g., Toxocara sp., Trichuris sp. Ascaris sp.) in 1 kg of dry matter is 0. There are no regulations requiring testing the concentration of Enterobacteriaceae bacteria or Escherichia coli as a representative species. In Europe, the Council Directive 86/278/EEC of June 1986 on the application of agricultural sewage sludge in agriculture [4] is still in force; while this directive establishes limit values for heavy metal concentrations, it does not provide indicators of biological origin. The presence of E. coli is a mandatory indicator of biological contamination of sewage sludge in only a few countries, such as Finland, Portugal and Lithuania, in amounts not exceeding 1000 CFU/g (colony-forming units per gram) or in no more than 100 CFU/g in Austria [5].
According to research by Estrada et al. [6], 80 days after the introduction of sewage sludge into the soil, the concentrations of most Enterobacteriaceae, E. coli and fecal coliform bacteria were below the detection limits in various conditions. Research conducted in Poland by Stańczyk-Mazanek and Stępniak [7] confirmed that the use of sewage sludge in doses not exceeding 40 t/ha should not cause soil contamination, but the use of higher doses may pose such a risk, especially from drug-resistant bacterial species. In turn, Michelon et al. [8] pointed out the need to limit and control enteric pathogens in organic substances introduced into the soil. The use of natural fertilizers should also consider the regional context, so that the introduced sewage does not present too much of a burden on the environment and result in, e.g., contamination of water bodies.
In terms of testing the number of Enterobacteriaceae (with a limit below 1000 CFU/g), Poland was subject, until 2024, to the provisions of the Regulation of the Minister of Agriculture and Rural Development (2008) [9] in the field of organic and organic–mineral fertilizers based on animal by-products. Pursuant to the 2007 Act on fertilizers and fertilization [10], digestate belongs to the group of manufactured fertilizers or fertilizers containing animal-derived products or by-products. Pursuant to the new ministerial regulation of August 2024 [11], fertilizers, plant cultivation aid products and post-fermentation products cannot contain live eggs of intestinal parasites and Salmonella, while the Enterobacteriaceae indicator has been removed.
In the current work, we focused exclusively on the contamination of soil, sewage sludge and digestate with bacteria from the Enterobacteriaceae family, and in particular, its representatives E. coli and Salmonella. The aim of the study was to determine the biodiversity of bacteria from the Enterobacteriaceae family in the tested samples and to determine safe limits of microbiological contamination of sludge and digestate based on an analysis of the risk of transfer of these pathogens to the soil in laboratory conditions. The results of this stage of the project will be used to verify current standards concerning regulations regarding the content of pathogenic bacteria in substances of organic origin intended for use as fertilizers in a way that does not pose a threat to human and animal health.
2. Materials and Methods
2.1. Determination of Bacteria Concentration Using Culture-Based Methods
2.1.1. Sample Collection
Samples of arable soil (82), sewage sludge (9) and digestate (9) were collected for microbiological examination in 2021. Soil samples from agricultural fields in northeastern Poland were gathered from the top layer (up to 20 cm depth) by a soil stick sampler. In accordance with the principles of soil sampling, at least 10 punctures were made to obtain an average sample. The sewage sludge and digestate samples were obtained from biological wastewater treatment plants and agriculture biogas plants, respectively. The samples were intended for testing immediately after their delivery to the laboratory. The soil samples were sieved through a sieve with a hole diameter of 2 mm.
2.1.2. Microbiological Culture
The assessment of microbial soil, sludge and digestate contamination was based on the following tests: total number of mesophilic bacteria, total number of Gram-negative bacteria from the Enterobacteriaceae family and presence of E. coli and Salmonella spp. Due to the lack of applicable procedures concerning microbiological testing of sewage sludge and digestates, the Polish standards pertaining to soil, food and feed research were used. Detection of Salmonella was performed according to the standard PN-Z-19000-1/2001 [12], Escherichia coli according to PN-EN ISO 16649-2:2004 [13], the total number of bacteria according to PN-EN ISO 4833-2:2013-12/AC [14] and Enterobacteriaceae according to PN-EN ISO 21528-2:2017-08 [15].
Two 10 g subsamples were taken from each sample for testing. One of the subsamples was suspended in 90 mL of Ringer’s solution, homogenized with a BagMixer 400 SW (Interscience, France) and intended for culture. The number of aerobic mesophilic bacteria was determined on nutrient agar plates (BTL, Łódź, Poland) incubated at 30 °C for 24 h. The presence of Gram-negative Enterobacteriaceae was determined on Violet, Red, Bile and Glucose (VRBG) agar plates (BioMaxima, Lublin, Poland), after incubation at 37 °C for 24 h. For E. coli, Tryptone Bile X-glucuronide (TBX) agar (BioMaxima, Lublin, Poland) was used, and the inoculated media were incubated at 44 °C for 24 h. The number of bacteria was expressed as the number of colony-forming units (CFU) in 1 g of sample.
The second subsample was suspended in 90 mL Selenite-F (SF) broth (BTL, Łódź, Poland) and incubated at 43 °C for 24 h. An inoculation loop full of each SF suspension with sample was streaked onto Salmonella Shigella (SS) agar (BTL, Łódź, Poland) and incubated at 37 °C for 24 h. Bacteria, isolated on VRBG, TBX and SS media, were subjected to genera/species identification.
2.2. Determination of Bacteria Species via Biochemical and Molecular Methods
2.2.1. Biochemical Tests
Preliminary identification of the isolated strains was carried out using the following sets of commercial kits: ENTEROtest 24N—for Salmonella, E. coli and other oxidase-negative bacteria from Enterobacteriaceae; NEFERMtest 24N—for oxidase-positive non-fermenting bacteria; OXItest—a supplementary test for detecting bacterial cytochrome oxidase; and INDOLtest—for detection of E. coli and screening differentiation of indole-positive and indole-negative bacterial genera (Erba-Lachema, Brno, Czech Republic). All tests were performed in accordance with the manufacturers’ recommendations using the ErbaScan absorbance microplate reader with a measurement range from 0.000 to 4.000 OD (Erba-Lachema, Brno, Czech Republic). The interpretation of the ENTEROtest 24N and NEFERMtest 24N results was performed using ErbaExpert microbiological software version 1.2.013 (Erba-Lachema, Brno, Czech Republic). In addition, an analysis of isolated strains was also performed using the Gen III BIOLOG system (BIOLOG Inc., Hayward, CA, USA). The results were read by MicroLog M 5.2 software (BIOLOG Inc., Hayward, CA, USA).
2.2.2. Molecular Tests
Isolation of DNA from bacterial cultures was performed using the Qiamp® DNA Mini Kit (Qiagen, Hilden Germany), according to the protocol for Gram-negative bacteria extraction. One culture loop from a 24-h bacterial culture was taken for isolation. Bacterial DNA was detected by amplification of the 16S rRNA gene fragment using the universal oligonucleotide primers p27f and p1525r according to the method by Chun and Goodfellow [16]. Each reaction has a volume of 50 µL and consisted of 1.5 U Taq DNA Polymerase, 1 × PCR buffer containing 15 mM MgCl2 (Qiagen, Hilden, Germany) and 0.2 mM dNTPs (Thermo Scientific, Waltham, MA, USA), 0.4 µM of each primer (Institute of Biochemistry and Biophysics, Warsaw, Poland) and 5 µL each of DNA template and nuclease-free water (Qiagen, Hilden, Germany). The reaction was conducted on a C1000 Thermal Cycler (BioRad, Hercules, CA, USA). Products of 1500 bp amplification were visualized in 1.5% agarose gel (Prona, Basica LE, Prona, Spain) after electrophoresis in standard conditions and staining with ethidium bromide solution (2 μg/mL).
The PCR sequencing reaction was performed using a BigDye™ Terminator v3.1 Cycle Sequencing Kit (Applied Biosystems, Waltham, MA, USA), and the reaction products were purified using a BigDye XTerminator™ Purification Kit (Applied Biosystems, Waltham, MA, USA). Sequencing was performed on the ABI PRISM 310 Genetic Analyzer (Applied Biosystems, Waltham, MA, USA). The nucleotide sequences were compared with sequences in GenBank using the Basic Local Alignment Search Tool (BLAST).
2.3. The Survival of E. coli Present in Organic Fertilizers on a Laboratory Scale
2.3.1. Samples
In the initial phase, the total number of Gram-negative Enterobacteriaceae and E. coli was determined in the soil (universal soil—used for, e.g., gardening—and clay), sewage sludge and digestate samples used in the experiment (Table 1). The sewage sludge and digestate samples were subjected to preliminary heat treatment at 121 °C for 15 min to remove natural microflora. Sterile samples were intended for inoculation with E. coli suspension. No Salmonella spp. was detected in any samples.

Table 1.
Initial bacterial contamination of soil, sewage sludge and digestate samples (before heat treatment).
2.3.2. Inoculum Preparation
The reference strain of E. coli ATCC 25922 was used to prepare the inoculum. From the 24-h culture, a suspension was prepared with an optical density of 0.5 McFarland (optical density at 550 nm: 0.125), measured with a Densi-La-Meter II densitometer (Erba-Lachema, Brno, Czech Republic). The initial suspension density (2.05 × 108 CFU/g) was determined based on the average concentration of mesophilic bacteria in the tested non-sterile sewage sludge and digestate samples (Table 2). When the E. coli suspension was added to the sterile sewage sludge and digestate samples, the final concentration was 1.8 × 106 CFU/g.

Table 2.
Concentration of bacteria in soil, sewage sludge and digestate samples.
2.3.3. Main Experiment
Four containers were prepared, filled with non-sterile soil (universal or clay) in a volume of 8.3 dm3, ensuring the mapping of the top 20 cm of the cultivated surface layer. Two containers were filled with soil, with the addition of 14.2 g and 88.5 g of sterile sewage sludge inoculated with 1.8 mL and 11.1 mL of the stock E. coli suspension of the same 2.05 × 108 CFU/g concentration, respectively. The same proportions were used to add digestate samples. The amount of the added sewage sludge or digestate sample was determined based on permissible doses of fertilizers (min. 3 t/ha; max. 20 t/ha) included in the Regulation of the Minister of the Environment [3]. The samples were mixed thoroughly and stored at a temperature of 20–25 °C during the day and 15–20 °C at night. Microbiological contamination was carried out after the 1st, 2nd and 3rd weeks of storage.
2.3.4. Control Group
The control group consisted of universal and clay soil samples with additives of non-sterile sewage sludge and digestate in amounts of 14.2 g and 88.5 g, with the concentration determined in the initial test (Table 1).
3. Results
3.1. Bacterial Concentration in Soil, Sewage Sludge and Digestate Samples
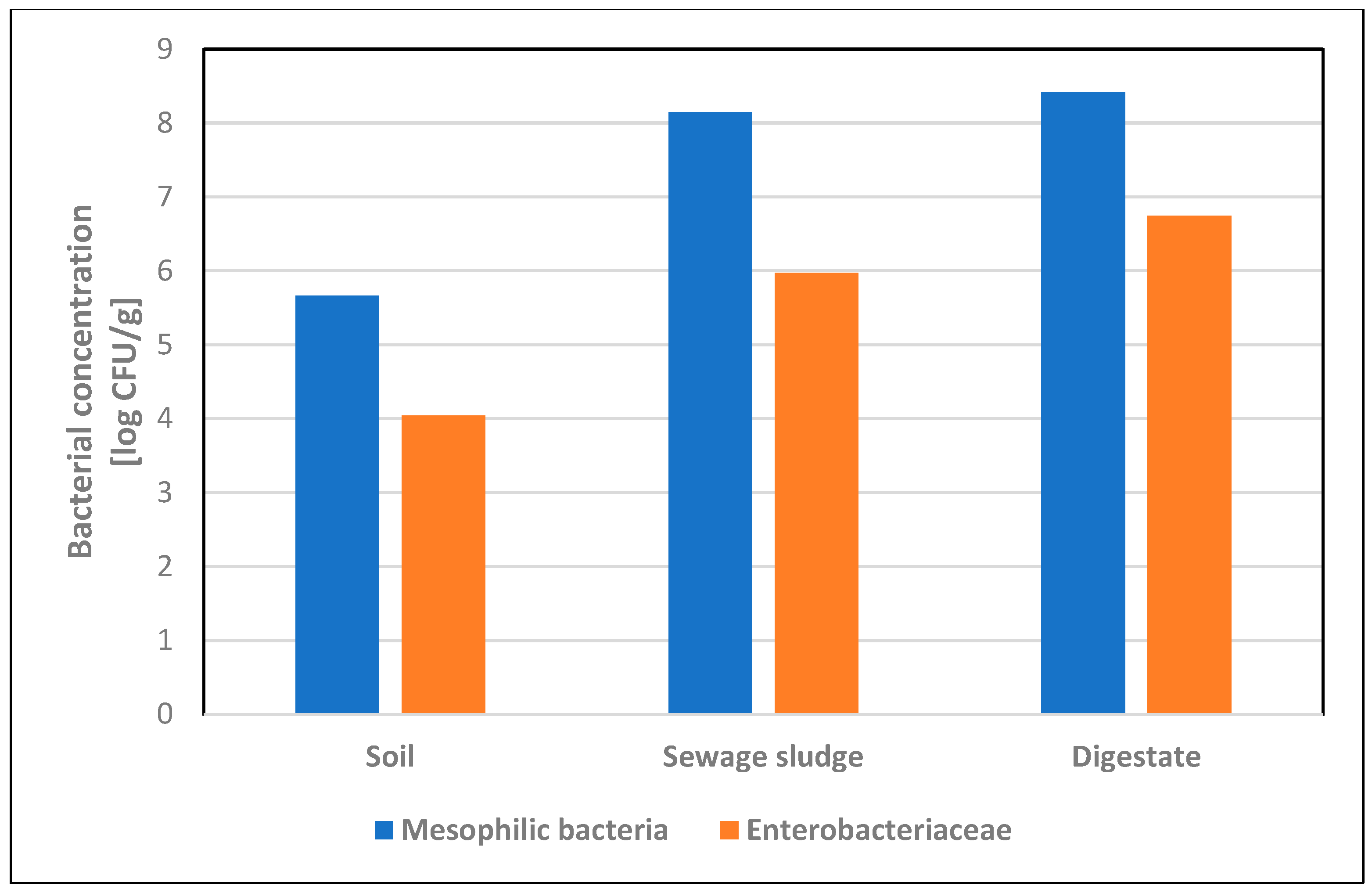
The average concentration of mesophilic bacteria in soil samples was 4.6 × 105 CFU/g, and the average concentration of Enterobacteriaceae was 1.1 × 104 CFU/g. Escherichia coli was detected in two soil samples, with an average concentration of 25.3 CFU/g. Microbiological analysis of sewage sludge showed the presence of mesophilic bacteria in eight (88.9%) samples, with an average concentration of 1.4 × 108 CFU/g, and Enterobacteriaceae in six samples (66.7%), with an average concentration of 9.4 × 105 CFU/g. Escherichia coli was detected in four (44.4%) samples, with an average concentration of 1.7 × 104 CFU/g. Microbiological analysis of digestate showed the presence of mesophilic bacteria and Enterobacteriaceae in all tested samples, obtaining average concentrations of 2.6 × 108 CFU/g and 5.6 × 106 CFU/g, respectively, while E. coli was detected in six (66.7%) samples (Table 2). Among all tested samples, Enterobacteriaceae isolated on VRBG medium constituted over 70% of the total number of mesophilic bacteria isolated on nutrient agar (Figure 1). In one digestate sample, the presence of Salmonella was confirmed [17].
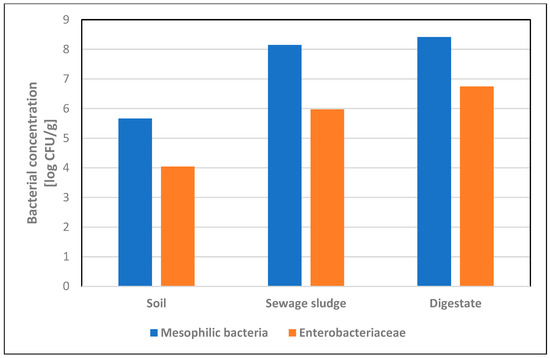
Figure 1.
Concentrations [log CFU/g] of mesophilic bacteria and Enterobacteriaceae in soil, sewage sludge and digestate samples.
3.2. Species Diversity of Enterobacteriaceae Isolated from Soil, Sewage Sludge and Digestate
From the soil samples, Gram-negative bacteria belonging to the Serratia (n = 44), Enterobacter (n = 37), Pantoea (n = 32), Citrobacter (n = 27) and Pseudomonas (n = 25) genera were identified the most frequently. Individual cases were confirmed for the genera Ewingella, Gibbsiella, Hafnia, Kluyvera and Yersinia. In 14 of the 82 samples tested, the presence of Escherichia coli, considered one of the main bacterial indicators of soil microbiological purity, was confirmed (Table 3), but in no sample did the concentration exceed the permissible value of 1000 CFU/g. Some species determined by biochemical methods could not be confirmed by sequencing (i.e., Burkholderia cepacia complex, Lelliottia amnigena, Chryseobacterium indologenes, Methylobacterium mesophilicum). The bacterial composition in each tested soil sample is provided in the Supplementary Materials (Table S1).
In both sewage sludge and digestate samples, the most frequently identified bacterium was E. coli. Some species were detected only in sewage sludge samples (Alcaligenes faecalis, Comamonas jiangduensis, Enterobacter cloacae, Hafnia alvei, Morganella morganii subsp. Morganii), and others only in digestates (Citrobacter freundii, C. gillenii, Ignatzschineria indica, Proteus mirabilis). The genera Klebsiella and Yersinia were isolated from both sample types but identified as different species. Salmonella enterica subsp. enterica (serotype Johannesburg) was identified in only one digestate sample (Table 4). Of all species identified via biochemical methods, Raoultella terrigena, Brevundimonas diminuta and Oligella urethralis were not confirmed by sequencing (Table S2) [18].

Table 3.
Enterobacteriaceae isolated from the soil samples.
Table 3.
Enterobacteriaceae isolated from the soil samples.
| Enterobacteriaceae Isolated from Soil Samples (n = 82) | Sequence Result (Similarity with 16S rRNA) | Impact on Human Health | ||||
|---|---|---|---|---|---|---|
| Aeromonas spp. (n = 13) | A. hydrophila | 100% | MT572504.1 | opportunistic pathogen | emerging foodborne pathogen capable of causing human gastroenteritis | [19] |
| A. sobria | 99.79% | OL778934.1 | food-borne illness, severe extraintestinal diseases: sepsis, peritonitis, meningitis | [20] | ||
| A. media | 99.82% | CP038448.1 | food-borne illness, diarrhea | [21] | ||
| A. veronii | 99.41% | KF853564.1 | gastroenteritis, bacteremia, septicemia, wound infections | [22] | ||
| A. salmonicida | 99.54% | MT576565.1 | strain isolated from the blood of the patient with endocarditis | [23] | ||
| A. encheleia | 100% | MT436428.1 | - | - | ||
| Achromobacter spp. (n = 2) | A. xylosoxidans subsp. xylosoxidans | 99.77% | CP054571.1 | opportunistic pathogen | bacteremia, meningitis, urinary tract infection, endocarditis, pneumonia, especially in immunocompromised patients | [24] |
| Buttiauxella spp. (n = 4) | B. agrestis | 99.57% | AP023184.1 | rare opportunistic pathogen | B. agrestis infection occurring at a post-cesarean surgical site | [25] |
| B. gaviniae | 100% | MK905440.1 | - | clinical strain isolated from a urine sample from a spinal cord patient with urinary bladder pathology | [26] | |
| Citrobacter spp. (n = 27) | C. braakii | 99.57% | MT534007.1 | rare pathogen, nosocomial infections | bacteremia in immunocompromised patients | [27,28] |
| C. freundii | 100% | MH045703.1 | opportunistic pathogen of a wide spectrum of nosocomial infections | food poisoning, diarrhea, urinary tract infections | [29,30] | |
| Enterobacter spp. (n = 37) | E. cloacae subsp. cloacae | 99.83% | MN181145.1 | opportunistic pathogen, nosocomial infections | lower respiratory tract infections, bacteremia | [31] |
| E. ludwigii | 99.71% | MN826154.1 | unusual human pathogen | nosocomial bloodstream infection | [32] | |
| E. asburiae | 99.67% | MN709316.1 | opportunistic pathogen | nosocomial infections | [33] | |
| E. amnigenus biovar 1 | 100% | MN658356.1 | unusual human pathogen | nosocomial infections | [34] | |
| Escherichia spp. (n = 14) | E. coli | 100% | MT192520.1 | opportunistic pathogen | enteritis, urinary tract infection, septicemia, neonatal meningitis, diarrhea | [35] |
| E. vulneris | 99.57% | KX357823.1 | possible opportunistic pathogen | wound infections; complicated diarrhea and sepsis in an infant | [36,37] | |
| Ewingella spp. (n = 1) | E. americana | 100% | MT998223.1 | rare opportunistic pathogen | bacteremia, pneumonia, conjunctivitis, Waterhouse–Friderichsen syndrome, peritonitis | [38] |
| Gibbsiella spp. (n = 1) | G. quercinecans | 99.78% | MN822736.1 | - | - | - |
| Hafnia spp. (n = 2) | H. alvei | 99.78% | CP050150.1 | potentially opportunistic pathogen | rare pneumonia cases | [39,40] |
| Klebsiella spp. (12) | K. pneumoniae subsp. pneumoniae | 99.79% | OM017200.1 | opportunistic pathogen | urinary and respiratory tract infections, liver abscess, endophthalmitis, meningitis | [41] |
| K. oxytoca | 100% | MT568561.1 | opportunistic pathogen | colitis, infective endocarditis, urinary and respiratory tract infections associated with nosocomial infections | [42] | |
| Kluyvera spp. (n = 2) | K. intermedia | 99.78% | LT899978.1 | potential opportunistic pathogen | soft tissue infections, urinary tract infections, intra-abdominal abscesses, catheter-associated bloodstream infections, septic shock in immunocompromised patients (nosocomial infections) | [43] |
| Pantoea spp. (n = 32) | P. agglomerans | 99.76% | MT635441.1 | opportunistic pathogen | septic arthritis, synovitis endophthalmitis, periostitis, endocarditis and osteomyelitis in the event of wound infection with plant material or as a hospital-acquired infection, mostly in immunocompromised individuals | [44] |
| Pseudomonas spp. (n = 25) | P. tolaasii | 99.56% | MT561438.1 | - | - | - |
| P. abietaniphila | 100% | MH379754.1 | - | - | - | |
| P. fluroescens | 99.73% | OM827287.1 | scarce clinical significance | nosocomial infections | [45] | |
| P. koreensis | 100% | MT501807.1 | - | possible nosocomial infections; a case of contact lens-related mixed infectious keratitis caused by A. fumigatus and P. koreensis | [46,47] | |
| P. brassicacearum subsp. neoaurantiaca | 99.80 | MT634587.1 | - | - | - | |
| P. chlororaphis | 100% | KJ530973.1 | - | - | - | |
| P. kilonensis | 99.57% | MT102732.1 | - | - | - | |
| Rahnella spp. (n = 18) | R. aquatilis | 99.77% | MN826573.1 | possible opportunistic pathogen | possible role as the pathogen responsible for ventilator-associated pneumonia associated with nosocomial infections in immunocompromised patients | [48] |
| R. victoriana | 99.78% | OK658118.1 | - | - | - | |
| Raoultella spp. (n = 5) | R. terrigena | 100% | MT545123.1 | opportunistic pathogen | cases of bloodstream, urinary tract, respiratory tract or bile tract infections mostly associated with nosocomial infections; subungual abscess caused by R. terrigena | [49,50] |
| R. ornithinolytica | 100% | MT568560.1 | ||||
| Serratia spp. (n = 44) | S. plymuthica | 99.58% | CP053398.1 | rare (unusual) human pathogen | associated with chronic osteomyelitis and cases of sepsis secondary to central venous catheter infection | [51] |
| S. liquefaciens | 100% | MT279350.1 | opportunistic pathogen | cause of transfusion-related sepsis, meningitis thrombophlebitis, corneal ulcers | [52] | |
| S. fonticola | 100% | MN227497.1 | rare (unusual) human pathogen | skin and soft tissue infections | [53,54] | |
| S. quinivorans | 99.71% | MT256279.1 | - | - | - | |
| S. proteamaculans | 100% | MK530287.1 | opportunistic pathogen | able to penetrate eukaryotic cells | [55] | |
| S. marcescens | 99.85% | MT598027.1 | opportunistic pathogen | urinary tract infections, pneumonia, intravenous catheter-associated infections, osteomyelitis, endocarditis | [51,56] | |
| S. entomophila | 99.83% | MK216954.1 | - | - | - | |
| Yersinia spp. (n = 2) | Y. enterocolitica | 99.14% | MN905014.1 | zoonotic pathogen | enteric infections, mesenteric lymphadenitis, reactive arthritis, erythema nodosum | [57] |

Table 4.
Enterobacteriaceae isolated from the sewage sludge and digestate samples.
Table 4.
Enterobacteriaceae isolated from the sewage sludge and digestate samples.
| Enterobacteriaceae Isolated from the Sewage Sludge and Digestate Samples (n = 18) | Sequence Result (Similarity with 16S rRNA) | Impact on Human Health | ||||
|---|---|---|---|---|---|---|
| Sewage Sludge Samples | Digestate Samples | |||||
| Aeromonas salmonicida | 100% | MT576565.1 | 100% | KF551980.1 | As in Table 2 | |
| Alcaligenes faecalis | 99.76% | MT277037.1 | - | - | sporadic cases of endocarditis, meningitis, chronic otitis, pyelonephritis, bacteremia, peritonitis, endophthalmitis, abscesses, often associated with nosocomial infections | [58] |
| Comamonas jiangduensis | 99.80% | NR_109655.1 | - | - | - | - |
| Citrobacter freundii | - | - | 100% | OM666544.1 | As in Table 2 | |
| Citrobacter gillenii | - | - | 100% | MT436425.1 | the strains were isolated from human stool, urine and blood | [59] |
| Enterobacter cloacae | 99.61% | MN006380.1 | - | - | As in Table 2 | |
| Escherichia coli | 100% | OM982954.1 | 100% | CP091756.1 | As in Table 2 | |
| Hafnia alvei | 99.39% | CP050150.1 | - | - | As in Table 2 | |
| Ignatzschineria indica | 99.75% | LC010924.1 | rare cases of bacteremia mostly associated with wound myiasis | [60] | ||
| Klebsiella oxytoca | 99.48% | MT509911.1 | - | - | As in Table 2 | |
| Klebsiella pneumoniae | - | - | 99.79% | OM978275.1 | As in Table 2 | |
| Morganella morganii subsp. morganii | 99.80% | CP043955.1 | - | - | opportunistic infections: urinary tract infection, wound infection, arthritis, prostatitis | [61] |
| Proteus mirabilis | - | - | 99.63% | OM882519.1 | catheter-associated urinary tract infections | [62] |
| Salmonella enterica subsp. enterica serovar Johannesburg | - | - | 100% | CP049308.1 | gastrointestinal infection—salmonellosis | [63] |
| Yersinia enterocolitica | - | - | 99.71% | MK910030.1 | As in Table 2 | |
| Yersinia frederiksenii | 100% | KC776774.1 | - | - | possible diarrheal diseases | [64] |
| Yersinia intermedia | - | - | 99.82% | MN416246.1 | strains were isolated from human stool and urine samples and from wound infections but rarely associated with human diseases | [65] |
3.3. The Survival of E. coli in Soil Samples Fertilized with Sewage Sludge and Digestate in Laboratory Conditions
3.3.1. Main Experiment
After the first week of fertilization with sewage sludge, an increase in the total number of Enterobacteriaceae was found compared to the initial result found in unfertilized soil (3.1 × 102 CFU/g): up to 9.7 × 103 CFU/g and 2.9 × 105 CFU/g for universal and clay soils, respectively. In the case of applying the minimum dose of sewage sludge, after the third week, the values did not exceed 100 CFU/g, while at the maximum dose, the final results were similar to the results of the unfertilized soil (1.3 × 102 and 6.0 × 102 CFU/g). The highest E. coli concentration values were obtained after the first week: up to 5.7 × 103 CFU/g and 1.8 × 105 CFU/g for universal and clay soils, respectively. After three weeks, the results decreased below 1 CFU/g, except for the application of the maximum dose of fertilizer in clay soil (2.8 × 102 CFU/g).
When digestate was used in all four variants, E. coli concentrations dropped to <1 or 16 CFU/g after three weeks. In the case of Enterobacteriaceae, after using the minimum dose of digestate, the bacterial concentration did not exceed 70 CFU/g. At the maximum dose, the final values were similar to those for unfertilized soil and amounted to an average of 2 × 102 CFU/g for both types of soil (Table 5).

Table 5.
Concentration of Enterobacteriaceae in soil samples fertilized with sterile sewage sludge/digestate inoculated with an E. coli suspension.
3.3.2. Control Experiment
In both types of soil with the addition of non-sterile sewage sludge or digestate, no growth of E. coli bacteria was observed. In universal soil with sewage sludge and clay soil with digestate at the maximum dose, the final concentration of Enterobacteriaceae obtained after three weeks was lower (<1.8 × 102 CFU/g) than the initial value for unfertilized soil (3.1 × 102 CFU/g). In the remaining variants, the final concentration of Enterobacteriaceae did not exceed 100 CFU/g (Table 6).

Table 6.
Concentration of Enterobacteriaceae in the soil samples fertilized with non-sterile sewage sludge and digestate.
4. Discussion
The use of sewage sludge and digestate as fertilizer on arable land promotes the functioning of the soil ecosystem, increasing crop productivity. In this study, high levels of contamination were found in sewage sludges and digestate: 1.4 × 108 CFU/g and 2.6 × 108 CFU/g, respectively. Slightly lower degrees of organic fertilizer contamination with mesophilic bacteria have been recorded in Spain (2.4 × 107 CFU/g for sludge) [66] and Germany (0.5 × 106 CFU/g for digestate) [67]. Direct sewage or digestate introduction into soil could increase the risk of environmental exposure to microbiological contamination, thus posing a threat to human and animal health. As indicated in Table 3 and Table 4, bacteria posing no direct health risk and those with pathogenic properties were isolated from the tested organic substances.
Currently, it is not possible for every organic sample introduced as fertilizer to be tested for bacterial species, including the quantitative assessment of individual species. Most legislation regulating the biological safety of fertilizers in terms of bacteria is based on tests for the detection of Salmonella. Of the samples tested, only one digestate sample confirmed the presence of Salmonella (S. enterica subsp. enterica serovar Johannesburg), which excludes the possibility of its introduction into the soil for agricultural purposes. In other studies, the presence of Salmonella spp. in digestates varies depending on the origin of the material and ranges from 8% (1/12) [68] to 100% (5/5) [69]. The presence of Salmonella spp. in sewage sludges is recorded at the level of 26.7% (4/15) [70] or 38.9% (21/54) [71].
The second frequently used indicator of microbiological contamination of soils and fertilizers is E. coli, which is a representative species of the Enterobacteriaceae family. The current study showed a higher average number of E. coli in the digestate samples, amounting to 1.7 × 106 CFU/g, than in the sewage sludge samples, which were determined to be 1.7 × 104 CFU/g. Studies on raw digestates collected from biogas plants in France showed mean E. coli counts ranging from 9.4 × 101 to 1.3 × 104 CFU/g [69]. The presence of E. coli strains in sewage sludge may suggest that treatment methods are not effective and that bacteria may be introduced into the soil environment, including pathogenic strains. The average counts of these bacterial strains may vary depending on the origin of the sewage sludge. Korzeniewska et al. [72] showed that the mean number of E. coli bacteria in untreated hospital sewage ranged from 6 × 102 to 1 × 105 CFU/mL, whereas in municipal sewage, it was higher and ranged from 1.1 × 103 to 1.3 × 105 CFU/mL. In a screening study in Sweden, the mean number of E. coli also varied depending on the sampling period in the effluent and ranged from 5.0 × 101 to 9.15 × 102 CFU/mL [73].
The results from our laboratory experiments revealed the presence of Enterobacteriaceae and E. coli at the levels below 1000 CFU/g recorded within 2 weeks of fertilization. After the first week, an increase in the number of E. coli specimens was noted in the universal soil sample. Similar results were obtained in the study conducted by Qiao et al. [74], who also showed an increase in the concentration of E. coli bacteria after 8 days. However, in the experiment conducted on clayey soil, the opposite results were observed, characterized by a decrease in the concentration of these microorganisms after a week. Moreover, higher concentrations of the tested microorganisms persisted longer in clay soil compared to sandy soil. The obtained results are consistent with those obtained by Alegbeleye and Sant’Ana [75], who also confirmed the higher survival of E. coli strains in clay soil compared to sandy soil. The persistent microbiological contamination of fertilized soil also depends on the dose of fertilizer applied. The use of smaller doses of fertilizers significantly accelerates the reduction in potentially pathogenic bacteria in the fertilized soil, and thus increases the safety of people in contact with it. However, further research in this area under natural conditions using experimental plots is necessary.
Taking into account the obtained diversity of the microbiome, the assessment of the microbiological purity of sediments and fermentates, referring mainly to E. coli and Salmonella spp., does not fully demonstrate the potential risk resulting from human exposure to pathogens. High levels of contamination based on the total number of microorganisms may indicate a need to improve sanitation methods used in biogas plants and sewage treatment plants. Despite these difficulties, sewage sludge and digestate are being increasingly used in agriculture as rich sources of plant nutrients and because the beneficial chemical elements they contain, including nitrogen, decompose slowly, providing nutrients over an extended period of time. Additionally, their use as fertilizers is an alternative to conventional waste disposal.
However, it is also important to appropriately adapt existing regulations regarding limiting potential human contact with pathogens [76,77]. Experimental field studies conducted in Poland on the effect of regular use of sewage sludge to fertilize agricultural soils showed significant quantitative and qualitative changes in the composition of the soil microbiota, disturbing its balance and influencing the processes occurring within it [75].
5. Conclusions
Based on the results of the experiments conducted in laboratory conditions, it can be concluded that adding the minimum dose (corresponding to a value of 3 t/ha) of sewage sludge or digestate with Enterobacteriaceae contamination below 2.5 × 106 CFU/g to soil, after an initial increase in the concentration of bacteria, does not result in bacterial concentrations exceeding the permissible value of 1000 CFU/g after weeks. Similarly, if the maximum dose (corresponding to a value of 20 t/ha) is used, the final results are at the same level found in unfertilized soil (2 × 102 CFU/g). Considering the species composition of soil, sewage sludge and digestate, including both pathogenic and non-pathogenic microorganisms, it is justified to change existing regulations by abolishing the obligation to quantitatively test samples for the presence of bacteria from the Enterobacteriaceae family. However, further research under natural conditions is necessary to confirm the biological safe use of sewage sludge and digestate as fertilizers.
Supplementary Materials
The following supporting information can be downloaded at: https://www.mdpi.com/article/10.3390/pathogens13121056/s1, Table S1: Results of quantitative and qualitative analysis of bacteriological contamination of soil samples; Table S2: Results of quantitative and qualitative analysis of bacteriological contamination of sewage sludge and digestate samples.
Author Contributions
Conceptualization, T.C. and A.W.-F.; methodology, A.W.-F., E.F., and K.K.; investigation, E.F., K.K., P.S., G.S., J.S., and J.M.Z.; resources, T.C., T.J., and J.M.Z.; writing—original draft preparation, A.W.-F., E.F., and K.K.; writing—review and editing, T.C. and J.S.; supervision, A.W.-F. and T.C.; funding acquisition, T.C., P.S., G.S., and A.W.-F. All authors have read and agreed to the published version of the manuscript.
Funding
This study was supported by the National Centre for Research and Development (Narodowe Centrum Badań i Rozwoju—NCBiR), Grant No.: GOSPOSTRATEG-III/0061/2020–00 OrgSafety: “Introduction of an innovative, cheap and environmentally-friendly method of hygienizing organic waste, enabling its use in fertilization”.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
All data generated or analyzed during this study are included in this published article and Supplementary Materials, and they are available from the Department of Health Biohazards and Parasitology (Institute of Rural Health, Lublin, Poland).
Conflicts of Interest
The authors declare no conflicts of interest.
References
- Samaddar, S.; Karp, D.S.; Schmidt, R.; Devarajan, N.; McGarvey, J.A.; Pires, A.F.A.; Scow, K. Role of soil in the regulation of human and plant pathogens: Soils’ contributions to people. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2021, 376, 20200179. [Google Scholar] [CrossRef] [PubMed]
- Chojnacka, K.; Skrzypczak, D.; Szopa, D.; Izydorczyk, G.; Moustakas, K.; Witek-Krowiak, A. Management of biological sewage sludge: Fertilizer nitrogen recovery as the solution to fertilizer crisis. J. Environ. Manag. 2023, 15, 116602. [Google Scholar] [CrossRef] [PubMed]
- Regulation of the Minister of the Environment of 6 February 2015 Regarding Municipal Sewage Sludge (Journal of Laws 2015, Item 257). Available online: https://isap.sejm.gov.pl/isap.nsf/DocDetails.xsp?id=wdu20150000257 (accessed on 10 September 2024). (In Polish)
- Council Directive of 12 June 1986 on the protection of the environment, and in particular of the soil, when sewage sludge is used in agriculture (86/278/EEC). Off. J. Eur. Communities 1986, 181, 6–12.
- Hudcová, H.; Vymazal, J.; Rozkošný, M. Present restrictions of sewage sludge application in agriculture within the European Union. Soil Water Res. 2019, 14, 104–120. [Google Scholar] [CrossRef]
- Estrada, I.B.; Aller, A.; Aller, F.; Gómez, X.; Morán, A. The survival of Escherichia coli, faecal coliforms and enterobacteriaceae in general in soil treated with sludge from wastewater treatment plants. Bioresour. Technol. 2004, 93, 191–198. [Google Scholar] [CrossRef] [PubMed]
- Stańczyk-Mazanek, E.; Stępniak, L. Analysis of migration of pathogenic drug-resistant bacteria to soils and groundwater after fertilization with sewage sludge. PLoS ONE 2021, 16, e0256936. [Google Scholar] [CrossRef]
- Michelon, W.; Peter, N.R.W.; Schneider, T.M.; Segalla, D.C.; Viancelli, A. Enterobacteria survival, percolation, and leaching on soil fertilized with swine manure. Int. J. Environ. Res. Public Health 2023, 20, 5283. [Google Scholar] [CrossRef]
- Regulation of the Minister of Agriculture and Rural Development of 18 June 2008 on the Implementation of Certain Provisions of the Act on Fertilizers and Fertilization. (Journal of Laws 2008 No. 119, Item 765). Available online: https://isap.sejm.gov.pl/isap.nsf/DocDetails.xsp?id=wdu20081190765 (accessed on 10 September 2024). (In Polish)
- Act of 10 July 2007 on Fertilizers and Fertilization (Journal of Laws of 2024, item 105, as Amended). Available online: https://isap.sejm.gov.pl/isap.nsf/DocDetails.xsp?id=WDU20240000105 (accessed on 10 September 2024). (In Polish)
- Regulation of the Minister of Agriculture and Rural Development of 9 August 2024 on the Implementation of Certain Provisions of the Act on Fertilizers and Fertilization (Journal of Laws 2024, Item 1261). Available online: https://isap.sejm.gov.pl/isap.nsf/DocDetails.xsp?id=WDU20240001261 (accessed on 10 September 2024). (In Polish)
- PN-Z-19000–1/2001; Jakość gleby. Ocena Stanu Sanitarnego Gleby: Wykrywanie Bakterii z Rodzaju Salmonella [Soil Quality—Assessment of the Soil Sanitary Conditions—Detection of the Salmonella Genus Bacteria]. Polish Standardization Committee: Warsaw, Poland, 2001. (In Polish)
- PN-ISO 16649-2:2004; Microbiology of Food and Animal Feeding Stuffs—Horizontal Method for the Enumeration of β-glucuronidase-Positive Escherichia coli—Part 2: Colony-Count Technique at 44 °C Using 5-bromo-4-chloro-3-indolyl β-D-glucuronide. ISO: Geneva, Switzerland, 2004.
- PN-EN ISO 4833-2:2013-12/AC; Microbiology of the Food Chain. Horizontal Method for the Enumeration of Microorganisms. Part 2: Colony Count at 30 °C by the Surface Plating Technique. ISO: Geneva, Switzerland, 2013.
- PN-EN ISO 21528-2:2017-08; Microbiology of the Food Chain—Horizontal Method for the Detection and Enumeration of Enterobacteriaceae. ISO: Geneva, Switzerland, 2017.
- Chun, J.; Goodfellow, M. A phylogenetic analysis of the genus Nocardia with 16S rRNA gene sequences. Int. J. Syst. Bacteriol. 1995, 45, 240–245. [Google Scholar] [CrossRef]
- Farian, E.; Wójcik-Fatla, A.; Kłapeć, T.; Zdybel, J.; Kowalczyk, K.; Sroka, J.; Cencek, T. Soil, sewage sludge and digestate samples contamination with Gram-negative bacteria from Enterobacteriaceae. In Proceedings of the 34th Annual Conference of the International Society for Environmental Epidemiology (ISEE 2022), Athens, Greece, 18–21 September 2022. [Google Scholar] [CrossRef]
- Wójcik-Fatla, A.; Kowalczyk, K.; Kłapeć, T.; Zdybel, J.; Farian, E.; Sroka, J.; Cencek, T. Species diversity of bacteria isolated from soil, sludge sewage and digestate. In Proceedings of the 34th Annual Conference of the International Society for Environmental Epidemiology (ISEE 2022), Athens, Greece, 18–21 September 2022. [Google Scholar] [CrossRef]
- Park, S.M.; Kim, H.W.; Choi, C.; Rhee, M.S. Pathogenicity and seasonal variation of Aeromonas hydrophila isolated from seafood and ready-to-eat sushi in South Korea. Food Res. Int. 2021, 147, 110484. [Google Scholar] [CrossRef]
- Kobayashi, H.; Seike, S.; Yamaguchi, M.; Ueda, M.; Takahashi, E.; Okamoto, K.; Yamanaka, H. Aeromonas sobria serine protease decreases epithelial barrier function in T84 cells and accelerates bacterial translocation across the T84 monolayer in vitro. PLoS ONE 2019, 14, e0221344. [Google Scholar] [CrossRef]
- Talagrand-Reboul, E.; Roger, F.; Kimper, J.L.; Colston, S.M.; Graf, J.; Latif-Eugenín, F.; Figueras, M.J.; Petit, F.; Marchandin, H.; Jumas-Bilak, E.; et al. Delineation of taxonomic species within complex of species: Aeromonas media and related species as a test case. Front. Microbiol. 2017, 8, 621. [Google Scholar] [CrossRef] [PubMed]
- Fernández-Bravo, A.; Figueras, M.J. Immune response of the monocytic cell line THP-1 against six Aeromonas spp. Front. Immunol. 2022, 13, 875689. [Google Scholar] [CrossRef]
- Salehi, M.R.; Shadvar, S.; Sadeghian, M.; Doomanlou, M.; Abdollahi, A.; Manshadi, S.A.D.; Sardari, A.; Rahdar, H.A.; Feizabadi, M.M. Endocarditis with Aeromonas salmonicida. IDCases 2019, 18, e00625. [Google Scholar] [CrossRef]
- Pickrum, A.M.; DeLeon, O.; Dirck, A.; Tessmer, M.H.; Riegert, M.O.; Biller, J.A.; Ledeboer, N.A.; Kirby, J.R.; Frank, D.W. Achromobacter xylosoxidans cellular pathology is correlated with activation of a type III secretion. Syst. Infect. Immun. 2020, 88, e00136-20. [Google Scholar] [CrossRef] [PubMed]
- Antonello, V.S.; Dallé, J.; Domingues, G.C.; Ferreira, J.A.; Fontoura Mdo, C.; Knapp, F.B. Post-cesarean surgical site infection due to Buttiauxella agrestis. Int. J. Infect. Dis. 2014, 22, 65–66. [Google Scholar] [CrossRef]
- De Baere, T.; Wauters, G.; Kämpfer, P.; Labit, C.; Claeys, G.; Verschraegen, G.; Vaneechoutte, M. Isolation of Buttiauxella gaviniae from a spinal cord patient with urinary bladder pathology. J. Clin. Microbiol. 2002, 40, 3867–3870. [Google Scholar] [CrossRef]
- Hirai, J.; Uechi, K.; Hagihara, M.; Sakanashi, D.; Kinjo, T.; Haranaga, S.; Fujita, J. Bacteremia due to Citrobacter braakii: A case report and literature review. J. Infect. Chemother. 2016, 22, 819–821. [Google Scholar] [CrossRef]
- Oyeka, M.; Antony, S. Citrobacter braakii bacteremia: Case report and review of the literature. Infect. Disord. Drug Targets 2017, 17, 59–63. [Google Scholar] [CrossRef]
- Liu, L.; Song, L.; Deng, R.; Lan, R.; Jin, W.; Tran Van Nhieu, G.; Cao, H.; Liu, Q.; Xiao, Y.; Li, X.; et al. Citrobacter freundii activation of NLRP3 inflammasome via the type VI secretion system. J. Infect. Dis. 2021, 223, 2174–2185. [Google Scholar] [CrossRef]
- Yang, L.; Li, P.; Liang, B.; Hu, X.; Li, J.; Xie, J.; Yang, C.; Hao, R.; Wang, L.; Jia, L.; et al. Multidrug-resistant Citrobacter freundii ST139 co-producing NDM-1 and CMY-152 from China. Sci. Rep. 2018, 8, 10653. [Google Scholar] [CrossRef]
- Mustafa, A.; Ibrahim, M.; Rasheed, M.A.; Kanwal, S.; Hussain, A.; Sami, A.; Ahmed, R.; Bo, Z. Genome-wide analysis of four Enterobacter cloacae complex type strains: Insights into virulence and niche adaptation. Sci. Rep. 2020, 10, 8150. [Google Scholar] [CrossRef]
- Flores-Carrero, A.; Labrador, I.; Paniz-Mondolfi, A.; Peaper, D.R.; Towle, D.; Araque, M. Nosocomial outbreak of extended-spectrum β-lactamase-producing Enterobacter ludwigii co-harbouring CTX-M-8, SHV-12 and TEM-15 in a neonatal intensive care unit in Venezuela. J. Glob. Antimicrob. Resist. 2016, 7, 114–118. [Google Scholar] [CrossRef] [PubMed]
- Zhu, B.; Wang, S.; Li, O.; Hussain, A.; Hussain, A.; Shen, J.; Ibrahim, M. High-quality genome sequence of human pathogen Enterobacter asburiae type strain 1497-78T. J. Glob. Antimicrob. Resist. 2017, 8, 104–105. [Google Scholar] [CrossRef]
- Capdevila, J.A.; Bisbe, V.; Gasser, I.; Zuazu, J.; Olivé, T.; Fernández, F.; Pahissa Berga, A. Enterobacter amnigenus. An unusual human pathogen. Enferm. Infecc. Microbiol. Clin. 1998, 16, 364–366. [Google Scholar]
- Allocati, N.; Masulli, M.; Alexeyev, M.F.; Di Ilio, C. Escherichia coli in Europe: An overview. Int. J. Environ. Res. Public Health 2013, 10, 6235–6254. [Google Scholar] [CrossRef]
- Pérez, C.; Melo, P.; Besomi, J.; Porte, L.; Wilhelm, J.P. Escherichia vulneris etiologic agent of septic arthritis in a child. Rev. Chilena Infectol. 2018, 35, 80–82. [Google Scholar] [CrossRef]
- Jain, S.; Nagarjuna, D.; Gaind, R.; Chopra, S.; Debata, P.K.; Dawar, R.; Sardana, R.; Yadav, M. Escherichia vulneris: An unusual cause of complicated diarrhoea and sepsis in an infant. A case report and review of literature. New Microbes New Infect. 2016, 13, 83–86. [Google Scholar] [CrossRef]
- Abrantes, C.; Freitas, J.; Silva, T.; Marques da Silva, L.; Carvalho, M.J.; Rodrigues, A.; Cabrita, A. A case of peritoneal dialysis-related peritonitis caused by Ewingella americana. Case Rep. Infect. Dis. 2022, 2022, 5607080. [Google Scholar] [CrossRef]
- Ramos-Vivas, J. Microbiology of Hafnia alvei. Enferm. Infecc. Microbiol. Clin. 2020, 38, 1–6. [Google Scholar] [CrossRef]
- Severiche-Bueno, D.F.; Vargas-Cuervo, M.T.; Medina-Lee, L.; Oliver-Hernandez, G.; Buitrago-Toro, K.; Insignares, D.A.; Conde-Camacho, R. Hafnia alvei pneumonia: From bees to human beings. Germs 2021, 11, 306–309. [Google Scholar] [CrossRef] [PubMed]
- Chew, K.L.; Octavia, S.; Lai, D.; Lin, R.T.P.; Teo, J.W.P. Genomic Characterization of Klebsiella quasipneumoniae from clinical specimens in Singapore. Antimicrob. Agents Chemother. 2021, 65, e0041221. [Google Scholar] [CrossRef]
- Neog, N.; Phukan, U.; Puzari, M.; Sharma, M.; Chetia, P. Klebsiella oxytoca and emerging nosocomial infections. Curr. Microbiol. 2021, 78, 1115–1123. [Google Scholar] [CrossRef]
- Thele, R.; Gumpert, H.; Christensen, L.B.; Worning, P.; Schønning, K.; Westh, H.; Hansen, T.A. Draft genome sequence of a Kluyvera intermedia isolate from a patient with a pancreatic abscess. J. Glob. Antimicrob. Resist. 2017, 10, 1–2. [Google Scholar] [CrossRef]
- Dutkiewicz, J.; Mackiewicz, B.; Lemieszek, M.; Golec, M.; Milanowski, J. Pantoea agglomerans: A mysterious bacterium of evil and good. Part III. Deleterious effects: Infections of humans, animals and plants. Ann. Agric. Environ. Med. 2016, 23, 197–205. [Google Scholar] [CrossRef]
- Donnarumma, G.; Buommino, E.; Fusco, A.; Paoletti, I.; Auricchio, L.; Tufano, M.A. Effect of temperature on the shift of Pseudomonas fluorescens from an environmental microorganism to a potential human pathogen. Int. J. Immunopathol. Pharmacol. 2010, 23, 227–234. [Google Scholar] [CrossRef] [PubMed]
- Fluit, A.C.; Rogers, M.R.C.; Díez-Aguilar, M.; Cantón, R.; Benaissa-Trouw, B.J.; Bayjanov, J.R.; Ekkelenkamp, M.B. Draft genome sequence of the strain 16-537536, isolated from a patient with bronchiectasis and its relationship to the Pseudomonas koreensis group of the Pseudomonas fluorescens complex. BMC Res. Notes 2020, 13, 10. [Google Scholar] [CrossRef]
- Khoo, L.W.; Srinivasan, S.S.; Henriquez, F.L.; Bal, A.M. A rare case of mixed infectious keratitis caused by Pseudomonas koreensis and Aspergillus fumigatus. Case Rep. Ophthalmol. 2020, 11, 600–605. [Google Scholar] [CrossRef]
- Martins, W.; Carvalhaes, C.G.; Cayô, R.; Gales, A.C.; Pignatari, A.C. Co-transmission of Rahnella aquatilis between hospitalized patients. Braz. J. Infect. Dis. 2015, 19, 648–650. [Google Scholar] [CrossRef]
- Appel, T.M.; Quijano-Martínez, N.; De La Cadena, E.; Mojica, M.F.; Villegas, M.V. Microbiological and clinical aspects of Raoultella spp. Front. Public Health 2021, 9, 686789. [Google Scholar] [CrossRef]
- Wang, Y.; Jiang, X.; Xu, Z.; Ying, C.; Yu, W.; Xiao, Y. Identification of Raoultella terrigena as a rare causative agent of subungual abscess based on 16S rRNA and housekeeping gene sequencing. Can. J. Infect. Dis. Med. Microbiol. 2016, 2016, 3879635. [Google Scholar] [CrossRef]
- Carrero, P.; Garrote, J.A.; Pacheco, S.; García, A.I.; Gil, R.; Carbajosa, S.G. Report of six cases of human infection by Serratia plymuthica. J. Clin. Microbiol. 1995, 33, 275–276. [Google Scholar] [CrossRef] [PubMed]
- Remuzgo-Martínez, S.; Aranzamendi-Zaldunbide, M.; Pilares-Ortega, L.; Icardo, J.M.; Acosta, F.; Martínez-Martínez, L.; Ramos-Vivas, J. Interaction of macrophages with a cytotoxic Serratia liquefaciens human isolate. Microbes Infect. 2013, 15, 480–490. [Google Scholar] [CrossRef] [PubMed]
- Aljorayid, A.; Viau, R.; Castellino, L.; Jump, R.L. Serratia fonticola, pathogen or bystander? A case series and review of the literature. IDCases 2016, 5, 6–8. [Google Scholar] [CrossRef]
- Hai, P.D.; Hoa, L.T.V.; Tot, N.H.; Phuong, L.L.; Quang, V.V.; Thuyet, B.T.; Son, P.N. First report of biliary tract infection caused by multidrug-resistant Serratia fonticola. New Microbes New Infect. 2020, 36, 100692. [Google Scholar] [CrossRef] [PubMed]
- Tsaplina, O.; Bozhokina, E. Bacterial outer membrane protein OmpX regulates β1 integrin and epidermal growth factor receptor (EGFR) involved in invasion of M-HeLa cells by Serratia proteamaculans. Int. J. Mol. Sci. 2021, 22, 13246. [Google Scholar] [CrossRef] [PubMed]
- Weakland, D.R.; Smith, S.N.; Bell, B.; Tripathi, A.; Mobley, H.L.T. The Serratia marcescens siderophore serratiochelin is necessary for full virulence during bloodstream infection. Infect. Immun. 2020, 88, e00117–e00120. [Google Scholar] [CrossRef]
- Leon-Velarde, C.G.; Jun, J.W.; Skurnik, M. Yersinia phages and food safety. Viruses 2019, 11, 1105. [Google Scholar] [CrossRef]
- Tena, D.; Fernández, C.; Lago, M.R. Alcaligenes faecalis: An unusual cause of skin and soft tissue infection. Jpn. J. Infect. Dis. 2015, 68, 128–130. [Google Scholar] [CrossRef]
- Brenner, D.J.; O’Hara, C.M.; Grimont, P.A.; Janda, J.M.; Falsen, E.; Aldova, E.; Ageron, E.; Schindler, J.; Abbott, S.L.; Steigerwalt, A.G. Biochemical identification of Citrobacter species defined by DNA hybridization and description of Citrobacter gillenii sp. nov. (formerly Citrobacter genomospecies 10) and Citrobacter murliniae sp. nov. (formerly Citrobacter genomospecies 11). J. Clin. Microbiol. 1999, 37, 2619–2624. [Google Scholar] [CrossRef]
- Deslandes, V.; Haney, C.; Bernard, K.; Desjardins, M. Ignatzschineria indica bacteremia in a patient with a maggot-infested heel ulcer: A case report and literature review. Access Microbiol. 2019, 2, acmi000078. [Google Scholar] [CrossRef]
- Li, X.; Chen, J. Septic shock induced by bacterial prostatitis with Morganella morganii subsp. morganii in a posttransplantation patient. Case Rep. Transplant. 2015, 2015, 850532. [Google Scholar] [CrossRef]
- Armbruster, C.E.; Mobley, H.L.T.; Pearson, M.M. Pathogenesis of Proteus mirabilis infection. EcoSal Plus 2018, 8, 10–1128. [Google Scholar] [CrossRef] [PubMed]
- Lamas, A.; Miranda, J.M.; Regal, P.; Vázquez, B.; Franco, C.M.; Cepeda, A. A comprehensive review of non-enterica subspecies of Salmonella enterica. Microbiol. Res. 2018, 206, 60–73. [Google Scholar] [CrossRef] [PubMed]
- Yeung, E.Y.H. A case series of diarrheal diseases associated with Yersinia frederiksenii. Infect. Dis. Rep. 2021, 13, 552–557. [Google Scholar] [CrossRef] [PubMed]
- Babujee, L.; Balakrishnan, V.; Kiley, P.J.; Glasner, J.D.; Perna, N.T. Transcriptome changes associated with anaerobic growth in Yersinia intermedia (ATCC29909). PLoS ONE 2013, 8, e76567. [Google Scholar] [CrossRef] [PubMed]
- Miguel, N.; Sarasa, J.; López, A.; Gómez, J.; Mosteo, R.; Ormad, M.P. Study of evolution of microbiological properties in sewage sludge-amended soils: A pilot experience. Int. J. Environ. Res. Public Health 2020, 17, 6696. [Google Scholar] [CrossRef] [PubMed]
- Pleissner, D.; Händel, N. Reduction of the microbial load of digestate by the cultivation of Galdieria sulphuraria under acidic conditions. Waste Biomass Valor. 2023, 14, 2621–2627. [Google Scholar] [CrossRef]
- Bonetta, S.; Ferretti, E.; Bonetta, S.; Fezia, G.; Carraro, E. Microbiological contamination of digested products from anaerobic co-digestion of bovine manure and agricultural by-products. Lett. Appl. Microbiol. 2011, 53, 552–557. [Google Scholar] [CrossRef]
- Le Maréchal, C.; Druilhe, C.; Repérant, E.; Boscher, E.; Rouxel, S.; Le Roux, S.; Poëzévara, T.; Ziebal, C.; Houdayer, C.; Nagard, B.; et al. Evaluation of the occurrence of sporulating and nonsporulating pathogenic bacteria in manure and in digestate of five agricultural biogas plants. Microbiologyopen 2019, 8, e872. [Google Scholar] [CrossRef]
- Santos, F.A.; Santos, P.C.; Matos, M.A.; Cardoso, O.; Quina, J.M. Effect of thermal drying and chemical treatments with wastes on microbiological contamination indicators in sewage sludge. Microorganisms 2020, 8, 376. [Google Scholar] [CrossRef]
- Krzyzanowski, F., Jr.; Zappelini, L.; Martone-Rocha, S.; Dropa, M.; Matté, M.H.; Nacache, F.; Razzolini, M.T. Quantification and characterization of Salmonella spp. isolates in sewage sludge with potential usage in agriculture. BMC Microbiol. 2014, 14, 263. [Google Scholar] [CrossRef] [PubMed]
- Korzeniewska, E.; Korzeniewska, A.; Harnisz, M. Antibiotic resistant Escherichia coli in hospital and municipal sewage and their emission to the environment. Ecotoxicol. Environ. Saf. 2013, 91, 96–102. [Google Scholar] [CrossRef] [PubMed]
- Flach, C.F.; Genheden, M.; Fick, J.; Joakim Larsson, D.G. A comprehensive screening of Escherichia coli isolates from Scandinavia’s largest sewage treatment plant indicates no selection for antibiotic resistance. Environ. Sci. Technol. 2018, 52, 11419–11428. [Google Scholar] [CrossRef] [PubMed]
- Qiao, C.; Azeem, M.; Zhang, H.; Yang, L.; Yang, S. Soil environment modulation by varying physicochemical attributes change the population dynamics of fecal Escherichia coli. Pol. J. Environ. Stud. 2013, 32, 225–232. [Google Scholar] [CrossRef]
- Alegbeleye, O.; Sant’Ana, A.S. Survival behavior of six enterotoxigenic Escherichia coli strains in soil and biochar-amended soils. Environ. Res. 2023, 223, 115443. [Google Scholar] [CrossRef]
- Bina, B.; Movahedian, H.I.; Kord, I. The effect of lime stabilization on the microbiological quality of sewage sludge. Iranian J. Environ. Health. Sci. Eng. 2004, 1, 34–38. [Google Scholar]
- Garrec, N.; Picard-Bonnaud, F.; Pourcher, A.M. Occurrence of Listeria sp and L monocytogenes in sewage sludge used for land application: Effect of dewatering, liming and storage in tank on survival of Listeria species. FEMS Immunol. Med. Microbiol. 2003, 35, 275–283. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).