Simple Summary
The two-spot cotton leafhopper, Amrasca biguttula (Ishida) (Hemiptera: Cicadellidae), is a regulated invasive pest recently detected in Florida and other southeastern states, USA. This insect attacks staple crops, such as okra, cotton, eggplant, and tropical hibiscus. While collecting infested okra plants in Homestead, Florida, one female Anagrus (Hymenoptera: Mymaridae) parasitoid was collected using a brush, whereas five parasitoids emerged from Am. biguttula eggs. The parasitoids were identified morphologically and molecularly as Anagrus vulneratus and Anagrus sp. near vulneratus. These parasitoids are native to North America and likely switched from unknown local hosts in southern Florida to parasitize Am. biguttula eggs. Future research is warranted to evaluate the efficiency of both parasitoids as natural enemies of Am. biguttula.
Abstract
The two-spot cotton leafhopper, Amrasca biguttula (Ishida) (Hemiptera: Cicadellidae), was recently detected in Florida and other southeastern states, USA. This is a quarantine pest of regulatory significance, since it can infest staple crops, such as okra, cotton, eggplant, and tropical hibiscus. While collecting infested okra plants in Homestead, Florida, five female Anagrus (Hymenoptera: Mymaridae) parasitoids emerged from eggs of Am. biguttula. The specimens were identified morphologically and molecularly by sequencing the cytochrome c oxidase subunit I (COI) gene and internal transcribed spacer 2 (ITS2) region of nuclear ribosomal RNA. Two Anagrus species were identified: Anagrus vulneratus and Anagrus sp. near vulneratus. These parasitoids are not known to occur in the Old World, the origin of Am. biguttula. Rather, they are native to North America. The available evidence suggests that the collected specimens switched from unknown local hosts in southern Florida to parasitize eggs of the invasive Am. biguttula. Future research to ascertain the identity of A. sp. near vulneratus and evaluate the efficiency of both parasitoids as natural enemies of Am. biguttula is warranted.
1. Introduction
The two-spot cotton leafhopper Amrasca biguttula (Ishida) (Hemiptera: Cicadellidae), also known under the common names two-spotted cotton leafhopper, cotton jassid, and okra leafhopper, is a serious pest of okra (Abelmoschus esculentus (L.) Moench), cotton (Gossypium hirsutum L.), eggplant (Solanum melongena L.), sunflower (Helianthus annuus L.), roselle (Hibiscus sabdariffa Rottler), and tropical hibiscus (Hibiscus rosa-sinensis L.) [1,2]. Both nymphs and adults feed on the underside of leaves. Using the piercing-sucking mouthparts, the leafhopper causes significant damage to the plants. The symptoms include leaf yellowing and curling, “hopperburn” (necrosis and drying of leaf margins), stunted growth, and crookedness of okra pods. In cotton, yield losses range from 19 to 49% [3,4], while in okra and eggplant, losses may reach up to 50% and 37%, respectively [5]. This pest is currently not known to vector any plant pathogens [1].
Amrasca biguttula is native to Asia and Oceania and has been reported from Afghanistan, Australia, Bangladesh, China, Christmas Island, French Polynesia, Guam, Iran, India, Indonesia, Japan, Laos, Malaysia, Myanmar, Nepal, Northern Mariana Islands, Pakistan, Philippines, Sri Lanka, Taiwan, Thailand, and Vietnam [6,7]. However, it is adventive in Iraq, West Africa, and the Caribbean [8,9], including Puerto Rico, the US Virgin Islands, Barbados, Antigua, Anguilla, and St. Kitts and Nevis [10], as well as in parts of Central America, where it has been confirmed at least in Honduras. Its recent detection in Florida in December of 2024 [10], in addition to its rapid spread in the Caribbean and southeastern USA [11], has raised concerns about its potential impact on staple American crops such as cotton, okra, and eggplant. In addition, a recent risk analysis highlights that Southwest Asia, sub-Saharan Africa, and South America are at high threat of invasion by Am. biguttula [12]. Moreover, rising temperatures are predicted to facilitate the pest’s establishment in other geographies.
Currently, Am. biguttula is considered a quarantinable pest of regulatory significance in the USA. In Florida, this pest is regulated by the Florida Department of Agriculture and Consumer Services-Division of Plant Industry (FDACS-DPI). Tropical hibiscus is an ornamental host plant intensively produced in Florida and shipped throughout the country; therefore, nurseries found with the leafhopper are being placed under “Stop Sale and Hold Order”. It is imperative to implement effective integrated pest management (IPM) programs to mitigate the economic damage caused by Am. biguttula. Notably, IPM programs developed should not rely on chemical control because there are reports of Am. biguttula resistance to pesticides across multiple insecticide classes [13]. Studies document moderate-to-very high resistance ratios to neonicotinoids in several cotton-growing regions, and recent work has demonstrated dimethoate (organophosphate, 1B) resistance with cross-resistance to some pyrethroids (3A) and novel modes of action [14,15,16]. Chemical control of Am. biguttula has relied heavily on systemic neonicotinoids (4A) (imidacloprid, thiamethoxam, acetamiprid) and, more recently, on newer chemistries such as dinotefuran, flonicamid and flupyradifurone, which generally show high field efficacy and rapid knockdown when applied as foliar sprays or seed treatments [17,18,19,20].
Natural enemies can play a pivotal role in regulating Am. biguttula populations. Fairyfly (Hymenoptera: Mymaridae) egg parasitoids have been repeatedly identified as the most important natural enemies attacking concealed Am. biguttula eggs, with higher parasitism rates observed on unsprayed or pesticide-free farms and clear reductions in leafhopper populations where parasitoids are abundant [21,22,23]. Several species of the genus Anagrus Haliday have been found to parasitize Am. biguttula eggs in Japan. Examples include Anagrus atomus (L.), Anagrus japonicus Sahad and Anagrus turpanicus Triapitsyn and Hu [22,23,24]. Adult Am. biguttula females insert their eggs into the veins of tender plant tissue [25]. The female Anagrus spp. parasitoids use the opening made by Am. biguttula to oviposit in the leafhopper eggs [26]. Within the family Mymaridae, Arescon enocki (Subba Rao and Kaur) and Stethynium empoascae Subba Rao have also been identified as egg parasitoids of Am. biguttula [23,24,27].
This study reports two Anagrus spp. for the first time in Florida and the entire New World associated with Am. biguttula. Here we present information on their morphological and molecular identification and discuss potential implications for the biological control of the invasive Am. biguttula.
2. Materials and Methods
2.1. Specimen Collection
On September 23, 2025, okra leaves infested with Am. biguttula nymphs and adults were collected from a 0.2 ha field at the Tropical Research and Education Center (TREC) of the University of Florida in Homestead (25°30′36″ N 80°30′22″ W, 2 m). No other leafhoppers were present on the okra plants. Leaves were inspected under a stereomicroscope (MZ6, Leica, Wetzlar, Germany) and one adult female parasitoid was collected using a fine brush (Winsor & Newton Cotman Series 111 Round #0000, Sheffield, UK). The specimen (Figure 1a) was slide-mounted using Hoyer’s medium [28] and kept at the Ornamental Entomology & Acarology laboratory at TREC.
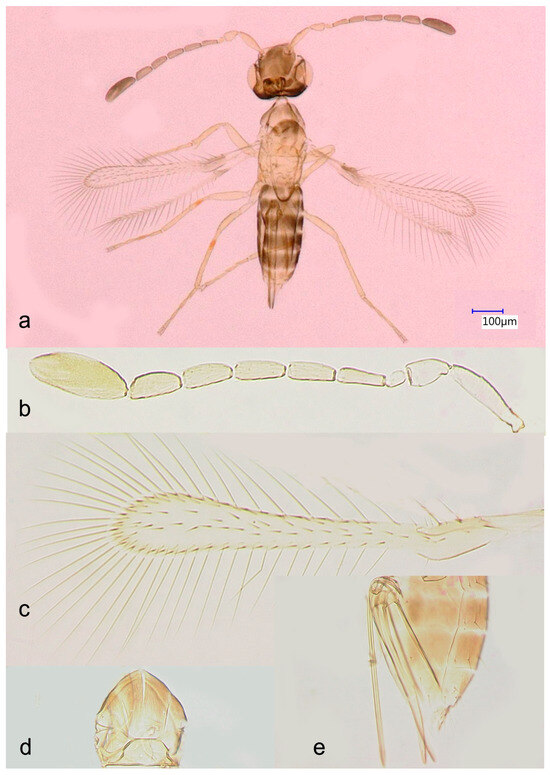
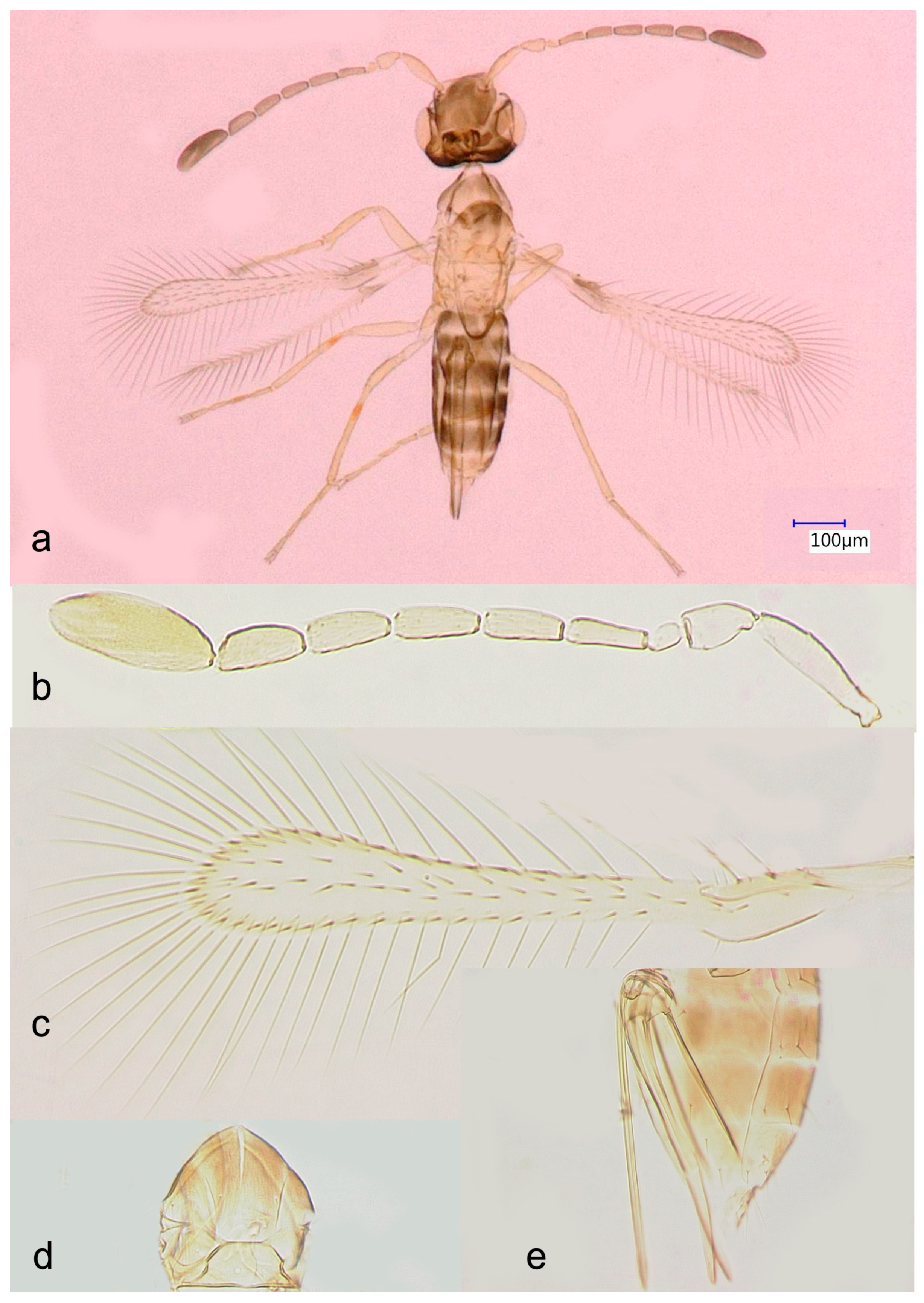
Figure 1.
Anagrus vulneratus from Homestead, FL, USA (female): (a), habitus; (b), antenna (length: 458 μm); (c), fore wing (length: 470 μm); (d), mesoscutum and scutellum (length: 111 μm); (e), ovipositor (length: 246 μm).
Subsequent okra leaf collections were held on September 29, October 2, 14, 29, and November 4, 2025. Leafhoppers were removed from the okra leaves, which were then placed with the abaxial side facing up in a Petri dish (52 × 37 mm) (Dynalon, New York, NY, USA) with 1% agar (Fisher Chemical A360500 Agar CAS 9002-18-0 Powder, Fisherbrand, Pittsburgh, PA, USA). To ensure ventilation, the Petri dish lids had two holes (21 mm diameter) covered with a fine mesh (100 μm diameter). To simulate the natural leaf position on the plant, the Petri dishes were placed upside-down inside an incubator (Percival I-36LL, Percival Geneva Scientific, Williams Bay, WI, USA) at 27 ± 1 °C, 12:12 h L:D, and 70% RH. The leaves were inspected for parasitoid emergence every 24 h for a total period of seven days. Emerging parasitoids from okra leaves were kept in the Petri dish for 24 h for observation, after which they were transferred to 4 mL glass vials (Wheaton, DWK Life Sciences, LLC., Millville, NJ, USA) containing 95% ethanol. In total, ten okra leaves were incubated per sampling date, and five adult female parasitoids were collected during October 2, 14, 29, 2025. Since leafhoppers insert their eggs in the leaves, it was not possible to count the number of laid and parasitized eggs.
2.2. Taxonomic Identification
The five ethanol-preserved female specimens of Anagrus sp. (at that point we presumed that they were all conspecific) were shipped to the Entomology Research Museum, Department of Entomology, University of California at Riverside (UCRC) for further identification to species level. DNA was extracted by P. F. Rugman-Jones from all five individuals using a nondestructive method described below. Consequently, these primary molecular vouchers (under P. F. Rugman-Jones’ numbers PR25-794, PR25-795, PR25-796, PR25-797, and PR25-798), each of which was also assigned a unique identifier for its repository (UCRC ENT number), were slide-mounted in Canada balsam to be used for morphological identification.
A tentative taxonomic identification was made based on the obtained molecular data and using the morphological descriptions and keys in Triapitsyn [29,30,31] and Triapitsyn et al. [32]. The morphological terms of Triapitsyn [31] were used, with the following abbreviations: F—funicle segment of the female antenna or flagellomere of the male antenna; mps—multiporous plate sensillum or sensilla on the antennal flagellar segments (=longitudinal sensillum or sensilla, or sensory ridge(s)).
Slide-mounted specimens of Anagrus spp. were examined under a Zeiss Axioskop 2 plus compound microscope (Carl Zeiss Microscopy, LLC., Thornwood, NY, USA) and photographed using the Auto-Montage® system (Syncroscopy, Princeton, NJ, USA). Photographs were retouched where necessary using Adobe Photoshop® (Adobe Systems, Inc., San Jose, CA, USA).
2.3. DNA Extraction, Amplification, and Sequencing
DNA was extracted from five individual specimens using the HotSHOT method of Truett et al. [33] in a total volume of 80 µL. Using the polymerase chain reaction (PCR), a section of the “barcoding” region of the mitochondrial cytochrome c oxidase subunit I (COI) gene was amplified using the primers C1-J-1718 and C1-N-2191, or LCO1490 and HCO2198, as described in Triapitsyn et al. [34]. In a separate PCR, the internal transcribed spacer 2 (ITS2) region of nuclear ribosomal RNA (rRNA) was also amplified as described in Triapitsyn et al. [35]. Amplification was confirmed by gel electrophoresis, after which the amplicons were purified using the PCR Product Pre-Sequencing Kit (Applied Biosystems, Foster City, CA, USA) and directly Sanger-sequenced in both directions at the Institute for Integrative Genome Biology, University of California at Riverside. The parity of forward and reverse reads was checked, and we initially used BLASTn searches (online version BLAST+ 2.17.0) to compare our sequences with records in two complementary public repositories to obtain molecular support/guidance for their identification. Sequences of the ITS2 were compared with those in the comprehensive GenBank library, while COI sequences were compared with those in the more focused Barcode of Life Data System (BOLD). Based on the outcome of these searches, specimens of two further species for which COI sequences were not already available were sequenced. These included specimens of A. vulneratus Triapitsyn originally collected in Grand Junction, CO, USA, and an undetermined species that we previously referred to as A. sp. near vulneratus originally collected in Sonora, Mexico [32,36] (misidentified as A. epos Girault). Respective DNA vouchers from Triapitsyn et al. [32] were retrieved from storage (A. vulneratus, PR08-004 and PR08-192; A. sp. near vulneratus, PR09-038) and several fresh extractions were made of specimens from each of the original collections. These included two male A. sp. near vulneratus (PR25-799 and PR25-800), and one male (PR25-801) and one female (PR25-802) A. vulneratus, from the same collections in Colorado and Sonora, respectively, as reported in Triapitsyn et al. [32]. We subsequently amplified and sequenced COI for all these specimens. The ITS2 was also sequenced for each of the fresh extractions to confirm conspecificity with the earlier material. All sequences generated herein were deposited in GenBank (accession numbers PX978890-900, COI; PX983467-475, ITS2).
2.4. Genetic Analysis
The COI sequences of the five Florida specimens were combined with those from additional authoritatively identified (SVT) Anagrus spp., produced herein or garnered from earlier studies [35,37,38]. The combined sequences were aligned in MAFFT version 7.050 [39] and trimmed to a uniform length, delimited at the 5′ end by the site of the C1-J-1718 primer, and at the 3′ end by the removal of a ~50 bp nucleotide-deficient overhanging region. The result was a final sequence data matrix containing 17 terminal taxa, 396 nucleotide positions, and no gaps. Although not the full length (~658 bp) barcoding region, shortened barcodes have previously shown their utility for discriminating between the closely related species of other Anagrus complexes [40]. Genetic variation among the sequences was estimated by calculating uncorrected p-distances between all possible sequence pairs, using MEGA version 12 [41]. A neighbor-joining (NJ) tree based on the p-distances was constructed and branch support was estimated using a bootstrap procedure with 1000 replicates. The outgroup A. atomus was used to root the tree.
3. Results
3.1. Taxonomic Identification of the Two Anagrus Species from Florida
Anagrus vulneratus Triapitsyn (Figure 1).
Anagrus vulneratus Triapitsyn in Triapitsyn et al. [32]: 9–12. Type locality: 39°02′31″ N 108°27′58″ W, 1450 m, Colorado State University Western Colorado Research Center—Orchard Mesa, Grand Junction, Mesa Co., CO, USA, from eggs of Erasmoneura vulnerata (Fitch) (Hemiptera: Cicadellidae) on cultivated grapes.
Material examined. USA: Florida, Miami-Dade Co., Homestead, UF/IFAS Tropical Research and Education Center, 25°30′36″ N 80°30′22″ W, 2 m, Y. Velazquez-Hernandez, on okra leaves infested with Am. biguttula: 23.ix.2025 (1 female, TREC); x.2025 (4 females, UCRC [PR25-795–798; UCRC ENT 573131, 283307, 573459, 573323, respectively]).
Diagnosis: Molecularly, four of the five sequenced specimens from Florida match those of A. vulneratus from Colorado (Figure 2). This species belongs to the incarnatus species group of the nominate subgenus of Anagrus, as defined by Triapitsyn [31], and, based on the presence of a pair of adnotaular setae on the midlobe of the mesoscutum (Figure 1d) and a bare area on the widest part of the fore wing (Figure 1c), to the A. epos species complex. Triapitsyn et al. [32] keyed the species in this complex, in which they included the Nearctic A. daanei Triapitsyn, A. epos Girault, A. tretiakovae Triapitsyn, and A. vulneratus Triapitsyn. Triapitsyn et al. [32] also indicated that the Nearctic and Neotropical species, A. empoascae Dozier, is morphologically somewhat similar to these taxa in the complex, but molecular data for A. empoascae was not available. Ecologically, among Cicadellidae A. empoascae is known from eggs of Empoasca spp. on weeds, crops, and other herbaceous vegetation [29,30,31,32], so okra fits the profile of its host plants.
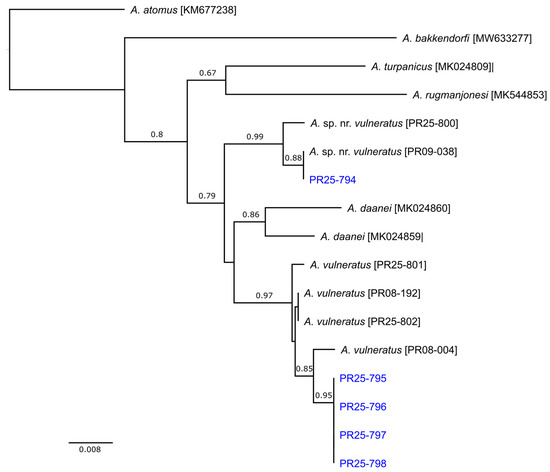
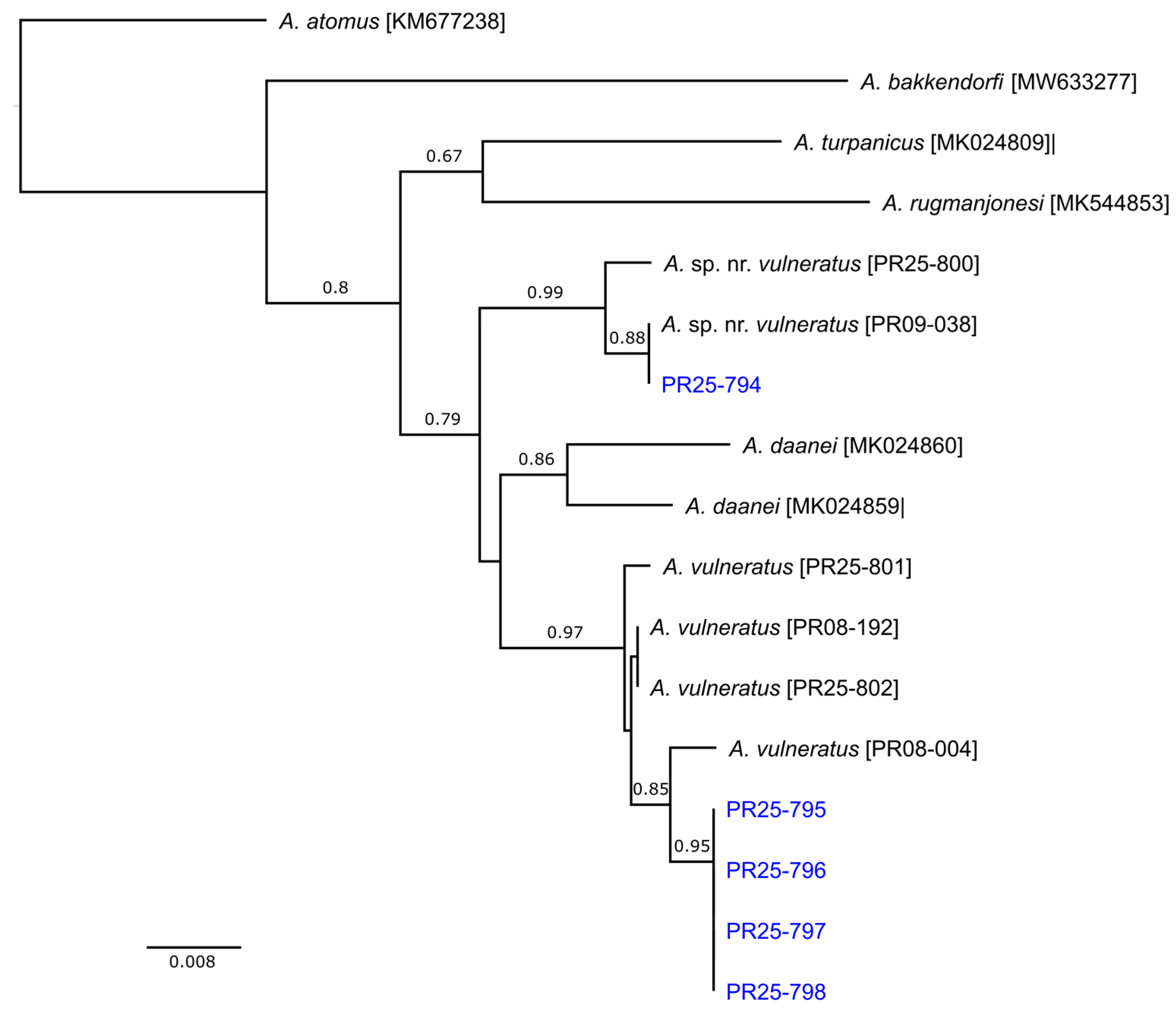
Figure 2.
Relationships among the sequences of the cytochrome c oxidase subunit I (COI) gene from Anagrus species. Optimal unrooted NJ tree with the sum of branch length = 0.247. The percentage of replicate trees in which the associated taxa clustered together in the bootstrap test (1000 replicates) are shown next to the branches and the tree is drawn to scale, with branch lengths indicating uncorrected p-distance. Specimens collected from Florida are in blue.
The five slide-mounted specimens of A. vulneratus from Florida differ, at least slightly, in one or more key morphological diagnostic features from all members of the A. epos species complex and A. empoascae (Table 1), as detailed below. They differ from A. empoascae in having fewer (2 or 3) rows of discal setae (Figure 1c) on the widest part of the fore wing (3 to 5 in A. empoascae); from A. daanei, in having 1 mps on F3 of the female antenna (Figure 1b), 3 setae on the external plate of the ovipositor (Figure 1e), and a higher (2.4–2.7) ovipositor length to fore tibia length ratio (usually no mps, almost always 2 setae, and 1.8–2.2 such ratio, respectively, in A. daanei); from A. epos, in having a relatively shorter ovipositor, with ovipositor length to fore tibia length ratio of at most 2.7 (at least 2.8 in A. epos); from A. tretiakovae, in having one longitudinal row of discal setae (Figure 1c) on the basal one-third of fore wing (two well-defined, complete such rows in A. tretiakovae); and from A. vulneratus from Colorado, in having 1 mps on F3 of the female antenna and a narrower fore wing, with length to width ratio at least 7.4 (usually no mps and at most 6.7 such ratio, respectively, in A. vulneratus). Thus, A. tretiakovae can be definitely excluded as a potentially conspecific taxon to our specimens from Florida. Among the other aforementioned species, they most closely resemble A. empoascae and A. epos, differing from them just slightly in one or more diagnostic features that are known to be prone to intraspecific variability, such as fore wing chaetotaxy and the relative length of the ovipositor [31]. Based on morphology and ecology, however, A. vulneratus from Florida could be conspecific with A. empoascae. However, a possible synonymy of A. vulneratus under A. empoascae can only be established after DNA is successfully extracted and analyzed following rearing of the latter nominal species from eggs of Empoasca spp. on herbaceous plants in Florida, Puerto Rico, or Haiti, within its native range.

Table 1.
Key diagnostic characteristics in the Anagrus epos species complex (excluding Anagrus tretiakovae, after Triapitsyn et al. [32] and Triapitsyn [31]), including Anagrus empoascae (after Triapitsyn [31]) and Anagrus sp. near vulneratus from southern Florida. Rare occurrences are indicated in parentheses.
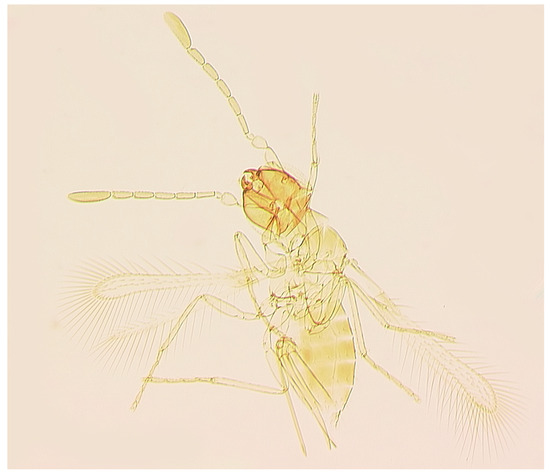
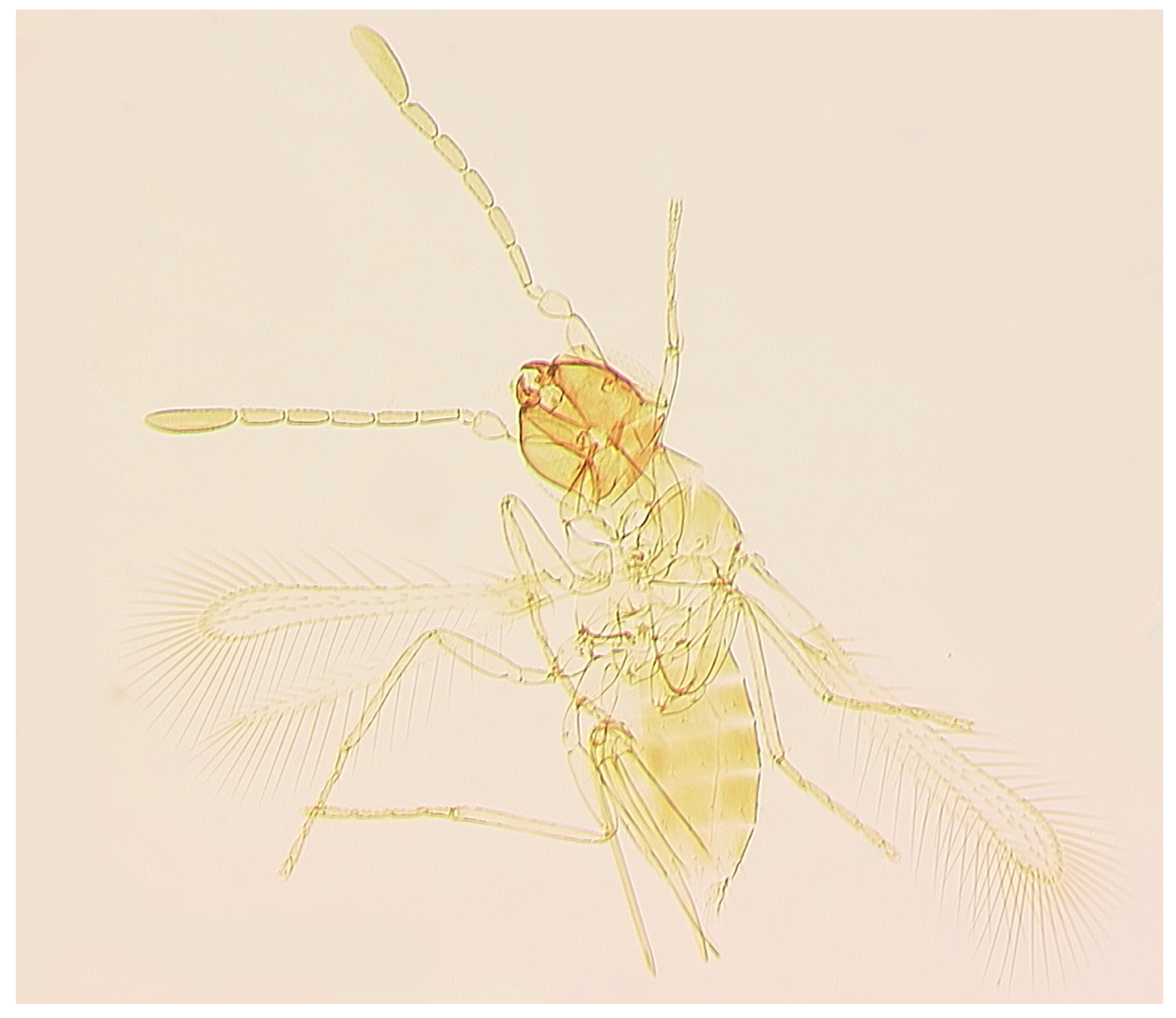
Figure 3.
Anagrus sp. near vulneratus from Homestead, FL, USA (female): habitus (body length: 615 μm).
Anagrus sp. near vulneratus Triapitsyn in Triapitsyn et al. [32]: 13–14 [known from Sonora and Baja California, Mexico from eggs of Erasmoneura variabilis (Beamer) (Hemiptera: Cicadellidae) on cultivated grapes].
Material examined. USA: Florida, Miami-Dade Co., Homestead, UF/IFAS Tropical Research and Education Center, 25°30′36″ N 80°30′22″ W, 2 m, x.2025, Y. Velazquez-Hernandez, on okra leaves infested with Am. biguttula (1 female, UCRC [PR25-794; UCRC ENT 496718]).
Diagnosis: Both molecularly (Figure 2) and morphologically (Table 1), this specimen from Florida matches those of the undescribed species, Anagrus sp. near vulneratus, from Sonora, Mexico [32]. Once more specimens are collected in Florida and its molecular comparison with A. empoascae becomes available, a description of a new species would be warranted.
3.2. Molecular Identification
Variation in the COI sequences of the five Florida specimens indicated that two species were in fact present, with one specimen (PR25-794) being 3.5% different from the other four (PR25-795, PR25-796, PR25-797, and PR25-798). The same split was evident in the ITS2 sequences. Despite this divergence, initial comparison of COI against BOLD suggested that all five specimens were Anagrus daanei Triapitsyn, each having a similarity of >99.8% to BIN (Barcode Index Number) AAN8044. This was somewhat surprising but closer inspection of the summary data for BIN:AAN0084 (https://portal.boldsystems.org/bin/BOLD:AAN8044; accessed February 27, 2026) suggested that it may contain multiple misidentified and/or cryptic species; among almost 3000 records, average pairwise distance (a key indicator of genetic variability) was 4%, with a maximum distance in excess of 8%. In contrast, comparison of ITS2 sequences with GenBank indicated that PR25-794 was a 99.43% match for an undetermined species [FJ861063], collected from Sonora, Mexico, that Triapitsyn et al. [32] previously referred to as A. sp. near vulneratus based on its similar morphology to A. vulneratus. The closest matches for the remaining specimens (94.69–95.88%) were in fact a series of cloned ITS2 sequences from two A. vulneratus specimens [FJ861041-045], collected from Mesa Co., Colorado, and sequenced as part of the same study by Triapitsyn et al. [32].
Since the COI gene had not been sequenced by Triapitsyn et al. [32], we subsequently generated sequences for A. sp. near vulneratus and A. vulneratus. These were combined with the Florida sequences along with those of several other species, including two published sequences from specimens collected from grapevines in California and identified as A. daanei by SVT [42]. Examination of genealogical relationships among these COI sequences provided clear evidence that one was indeed A. sp. near vulneratus (uncorrected p-distance = 0.0038), and the others were almost certainly A. vulneratus (uncorrected p-distance = 0.0082; Figure 2). Both species were >3% different from Californian A. daanei (uncorrected p-distances were 0.0316 and 0.0354, respectively).
4. Discussion
The recent establishment of Am. biguttula in parts of southeastern USA and some Caribbean islands as well as in parts of Central America has increased concerns about its potential impact on staple American crops [9,10,11,43]. Additionally, since this pest is regulated and known to have resistance to pesticides [14,15,16], the findings of this study provide a promising avenue for more environmentally friendly strategies. The presented data unambiguously indicate the local origin of the two Anagrus species reared in southern Florida from eggs of Am. biguttula. Both identified members of the Anagrus epos species complex of the nominate subgenus of this genus, A. vulneratus and A. sp. near vulneratus, are not known to occur in the Old World, the origin of this leafhopper pest. Rather, they are native to North America [31,32,36] while A. empoascae is known throughout the New World and the Hawaiian Islands [29,30,31]. Thus, it is possible that these two species have the ability to switch from unknown local hosts in southern Florida to parasitize abundant eggs of the invasive Am. biguttula. We can speculate that Empoasca spp. could very well be the native hosts of these two parasitoid species since they and Am. biguttula belong to the same tribe Empoascini of the leafhopper subfamily Typhlocybinae.
Due to the presence of at least slight morphological differences from all the named members of the A. epos species complex (Table 1) and limited knowledge of their intraspecific variability, we must rely on unambiguous molecular data to provide these identifications. Therefore, in the meantime, it is prudent to refer to one as A. vulneratus and the other as A. sp. near vulneratus, until comparative COI barcode and ITS2 sequences are obtained from freshly collected specimens that fit the current morphological concept of A. empoascae and its type specimens from Haiti, as redescribed by Triapitsyn [29], preferably reared from Empoasca spp. eggs on herbaceous vegetation. Thus, additional research would be necessary to reveal the true identity of these parasitoid wasps that may be important for providing at least partial biological control of the invasive Am. biguttula in Florida and other southeastern states in the USA and elsewhere in the New World.
Currently, molecular identification of Mymaridae to species level using available sequences in public DNA reference databases, such as BOLD Systems for COI barcodes, needs to be approached with great caution because of a very limited accuracy and unreliability, with less than 1% of species matching in Québec, Canada [44]. Moreover, out of the two of their species that were presumed to be correctly identified based on DNA barcoding results available from BOLD Systems [44], one (A. daanei) was actually also misidentified, as it is not conspecific with the true A. daanei reared from Western grape leafhopper (Erythroneura elegantula Osborn, Cicadellidae; PR16-029) and Virginia creeper leafhopper (Erythroneura ziczac Walsh; PR16-041) eggs in California, of which we have obtained COI barcodes (Figure 2), and elsewhere. However, some of their specimens from Québec, identified as Anagrus sp., seem to belong to A. daanei based on our COI barcode data (Figure 2). Thus, an integrative taxonomy approach to the identification of these minute and speciose parasitoid wasps is essential, when feasible, in combination with building reliable genetic reference databases for as many positively recognized species as possible.
Mymarid egg parasitoids can play a pivotal role in the regulation of Am. biguttula populations because they can attack concealed Am. biguttula eggs, hence preventing leafhopper population buildup [21,22,23]. We currently lack information on the parasitism rate of Am. biguttula eggs by A. vulneratus and A. sp. near vulneratus. Future research can evaluate their efficiency as natural enemies of this leafhopper pest and shed more light on their compatibility with other IPM practices, such as chemical and cultural control. Regardless, the presence of two natural enemies that have been found responding to the invasive Am. biguttula can pave the way for a more sustainable management, which may also include efforts to implement a classical biological control program.
Author Contributions
Conceptualization, A.M.R., S.V.T., and P.F.R.-J.; methodology, Y.V.-H., S.V.T., and P.F.R.-J.; software, P.F.R.-J.; validation, A.M.R., S.V.T., and P.F.R.-J.; formal analysis, S.V.T. and P.F.R.-J.; investigation, Y.V.-H., S.V.T., and P.F.R.-J.; resources, A.M.R., S.V.T., and P.F.R.-J.; data curation, S.V.T. and P.F.R.-J.; writing—original draft preparation, A.M.R., S.V.T., and P.F.R.-J.; writing—review and editing, Y.V.-H.; visualization, S.V.T. and P.F.R.-J.; project administration, A.M.R.; funding acquisition, A.M.R. All authors have read and agreed to the published version of the manuscript.
Funding
This research was supported by the USDA-NIFA Research Capacity Fund (Hatch) project (Accession Number 7000779). The mention of trade names or commercial products in this publication is solely for the purpose of providing specific information and does not imply recommendation or endorsement by the USDA; the USDA is an equal-opportunity provider and employer.
Data Availability Statement
The original contributions presented in this study are included in the article. Further inquiries can be directed to the corresponding author.
Acknowledgments
The authors thank Marcello De Giosa for assisting with the sampling of okra leaves and Xingbo Wu for allowing us to sample the okra leaves. The authors also thank Daniel Carrillo for providing feedback on previous versions of the manuscript.
Conflicts of Interest
The authors declare no conflicts of interest.
Abbreviations
The following abbreviations are used in this manuscript:
| COI | Cytochrome c oxidase subunit I |
| ITS2 | Internal transcribed spacer 2 |
| rRNA | Ribosomal RNA |
| F | Funicle segment of the female antenna or flagellomere of the male antenna |
| mps | Multiporous plate sensillum or sensilla on the antennal flagellar segments |
References
- Clement, R.A.; Gilligan, T.M. Screening Aid Two-spot Cotton Leafhopper (Cotton Jassid) Amrasca biguttula (Ishida, 1913); Pest Identification Technology Laboratory (PITL), USDA-APHISPPQ-S&T: Fort Collins, CO, USA, 2025; p. 4.
- Cabrera-Asencio, I.; Dietrich, C.H.; Zahniser, J.N. A new invasive pest in the Western Hemisphere: Amrasca biguttula (Hemiptera: Cicadellidae). Florida Entomol. 2023, 106, 263–266. [Google Scholar] [CrossRef]
- Rao, N.V.; Prasad, V.R.; Reddy, D.D.R. Economic thresholds and pest management in cotton. Indian J. Entomol. 1996, 58, 45–50. [Google Scholar]
- Ahmad, Z.; Attique, M.R.; Rashid, A. An estimate of the loss in cotton yield in Pakistan attributable to the jassid Amrasca devastans Dist. Crop Prot. 1985, 5, 105–108. [Google Scholar] [CrossRef]
- Ahmed, M. Evaluation of yield losses in brinjal (Solanum melongena) by Amrasca devastans. Pakistan J. Agric. Res. 1982, 3, 277–280. [Google Scholar]
- CABI. Amrasca biguttula (Indian cotton jassid). CABI Compend. 2022. [Google Scholar] [CrossRef]
- Sagarbarria, M.G.S.; Taylo, L.D.; Hautea, D.M. Morphological and molecular analyses of Leafhopper, Amrasca biguttula (Ishida) (Hemiptera: Cicadellidae) infesting eggplant (Solanum melongena L.) in Luzon Island, Philippines. J. Asia. Pac. Entomol. 2020, 23, 260–268. [Google Scholar] [CrossRef]
- Dmitriev, D.A.; Angelov, R.; Anufriev, G.A.; Bartlett, C.R.; Blanco-Rodríguez, E.; Borodin, O.I.; Al, E. TaxonPages: World Auchenorrhyncha Database—Amrasca (Sundapteryx) biguttula (Ishida, 1913). Available online: https://hoppers.speciesfile.org/otus/41773/overview (accessed on 3 August 2025).
- Michel, M.; Zamorano, E.; Orozco, H.J. First record of an Asian leafhopper, Amrasca biguttula (Ishida) (Hemiptera: Cicadellidae), in Central America. Insecta Mundi 2025, 1147, 1–8. [Google Scholar] [CrossRef]
- Liburd, O.E.; Halbert, S.E.; Samuel, N.; Dreves, A.J. Pest Alert—Two-Spot Cotton Leafhopper, Hemiptera: Cicadellidae, Typhlocybinae, Empoascini; Amrasca biguttula (Ishida)—A Serious Pest of Cotton, Okra and Eggplant That Has Become Established in the Caribbean Basin; Florida Department of Agriculture and Consumer Services, Division of Plant Industry: Gainesville, FL, USA, 2024; pp. 1–5.
- Esquivel, I.L.; Bryant, T.; Malone, S.; Jacobson, A.L.; Graham, S.H.; Gimenez-Cremonez, P.S.; Roberts, P.; Paula-Moreas, S.; Reisig, D.; Huseth, A.; et al. First Report of Two-Spot Cotton Leafhopper (Amrasca biguttula Ishida) (Hemiptera: Cicadellidae) on Commercial Cotton in the Southeastern United States. Insects 2025, 16, 966. [Google Scholar] [CrossRef] [PubMed]
- Azrag, A.A.G.; Niassy, S.; Bloukounon-Goubalan, A.Y.; Abdel-Rahman, E.M.; Tonnang, H.E.Z.; Mohamed, S.A. Cotton production areas are at high risk of invasion by Amrasca biguttula (Ishida) (Cicadellidae: Hemiptera): Potential distribution under climate change. Pest Manag. Sci. 2025, 81, 2910–2921. [Google Scholar] [CrossRef]
- Amrasca spp. Archives|Insecticide Resistance Action Committee. Available online: https://irac-online.org/pests/amrasca_biguttula_biguttula/ (accessed on 4 September 2025).
- Khalid, I.; Mohsin, M.; Abubakar, M.; Ali Shad, S.; Binyameen, M. Polygenic and autosomally inherited dimethoate resistance in Amrasca biguttula biguttula with no cross-resistance to bifenthrin, imidacloprid, and chlorfenapyr. Crop Prot. 2023, 163, 106099. [Google Scholar] [CrossRef]
- Halappa, B.; Patil, R.K. Detoxifying enzyme studies on cotton leafhopper, Amrasca biguttula biguttula (Ishida), resistance to neonicotinoid insecticides in field populations in Karnataka, India. J. Plant Prot. Res. 2016, 56, 346–352. [Google Scholar] [CrossRef]
- Saeed, R.; Razaq, M.; Abbas, N.; Jan, M.T.; Naveed, M. Toxicity and resistance of the cotton leaf hopper, Amrasca devastans (Distant) to neonicotinoid insecticides in Punjab, Pakistan. Crop Prot. 2017, 93, 143–147. [Google Scholar] [CrossRef]
- Kodandaram, M.H.; Kumar, Y.B.; Rai, A.B.; Singh, B.; Banerjee, K.; Hingmire, S. Field bioefficacy, phytotoxicity and residue dynamics of the insecticide flonicamid (50 WG) in okra [Abelmoschus esculenta (L.) Moench]. Crop Prot. 2017, 94, 13–19. [Google Scholar] [CrossRef]
- Venkateshalu, V.; Math, M. Bio-efficacy of dinotefuran 20 per cent SG against sucking insect pests of okra. Asian J. Bio Sci. 2017, 12, 8–14. [Google Scholar] [CrossRef]
- Singh, B.; Pandey, R.; Singh, A.; Mishra, M. Effectiveness of flonicamid 50 wg and flupyradifurone 200 SL against leafhopper and whitefly in okra. J. Entomol. Zool Stud. 2020, 8, 181–185. [Google Scholar]
- Malanno, K.; Norbert, B.K.K.; Houphouët, K. Efficient dose of thiamethoxam as a seed treatment product to reduce early damage of cotton leafhoppers Amrasca biguttula (Hemiptera: Cicadellidae) in Côte d’Ivoire. J. Entomol. Zool. Stud. 2025, 13, 32–35. [Google Scholar] [CrossRef]
- Sahito, H.A.; Soomro, F.; Kousar, T.; Shah, Z.H.; Mangrio, W.M. Population management of cotton Jassid, Amrasca biguttula biguttula Ishida through its biological control agent, Arescon enocki (Rao and Kaur) under field and laboratory conditions at upper Sindh, Pakistan. Int. J. Biosci. 2018, 13, 212–218. [Google Scholar] [CrossRef]
- Adachi-Hagimori, T.; Triapitsyn, S.V.; Uesato, T. Egg parasitoids (Hymenoptera: Mymaridae) of Amrasca biguttula (Ishida) (Hemiptera: Cicadellidae) on Okinawa Island, a pest of okra in Japan. J. Asia Pac. Entomol. 2020, 23, 791–796. [Google Scholar] [CrossRef]
- Triapitsyn, S.V. Egg Parasitoids as Important Natural Enemies of Auchenorrhynchan Insect Pests in East Asia. J. Taiwan Agric. Res. 2025, 74, 221–235. [Google Scholar]
- Adachi-Hagimori, T.; Triapitsyn, S.V.; Rugman-Jones, P.F. Egg parasitoids (Hymenoptera: Mymaridae and Trichogrammatidae) of okra leafhopper, Amrasca biguttula (Ishida) (Hemiptera: Cicadellidae), in southern Japan. J. Asia Pac. Entomol. 2023, 26, 102041. [Google Scholar] [CrossRef]
- Sharma, A.; Singh, R. Oviposition preference of cotton leafhopper in relation to leaf-vein morphology. J. Appl. Entomol. 2002, 126, 538–544. [Google Scholar] [CrossRef]
- Hangay, G.; Gayubo, S.F.; Hoy, M.A.; Goula, M.; Sanborn, A.; Morrill, W.L.; GÄde, G.; Marco, H.G.; Kabissa, J.C.B.; Ellis, J.; et al. Anagrus Fairyflies (Hymenoptera: Mymaridae). Encycl. Entomol. 2008, 152–154. [Google Scholar] [CrossRef]
- Adachi-Hagimori, T.; Triapitsyn, S.V.; Endou, T. Stethynium empoascae Subba Rao (Hymenoptera: Mymaridae), a newly recorded egg parasitoid of Amrasca biguttula (Ishida) (Hemiptera: Cicadellidae), a pest of okra on Ogasawara (Bonin) Islands, Japan. J. Asia Pac. Entomol. 2025, 28, 102379. [Google Scholar] [CrossRef]
- Krantz, G.W.; Walter, D.E. A Manual of Acarology-Texas Tech University Press. Available online: https://www.ttupress.org/9780896726208/a-manual-of-acarology/ (accessed on 5 December 2025).
- Triapitsyn, S.V. The genus Anagrus (Hymenoptera: Mymaridae) in America south of the United States: A review. Ceiba 1997, 38, 1–12. [Google Scholar]
- Triapitsyn, S.V. Descriptive notes on a new and other little known species of Anagrus Haliday, 1833 (Hymenoptera: Mymaridae) from the New World tropics and subtropics. Entomotropica 2002, 17, 213–223. [Google Scholar]
- Triapitsyn, S.V. Taxonomy of the genus Anagrus Haliday (Hymenoptera: Mymaridae) of the world: An annotated key to the described species, discussion of the remaining problems, and a checklist.—Univ. of Florida. Acta Zool Lilloana 2015, 59, 3–50. [Google Scholar]
- Triapitsyn, S.V.; Rugman-Jones, P.F.; Jeong, G.; Morse, J.G.; Stouthamer, R. Morphological and molecular differentiation of the Anagrus epos species complex (Hymenoptera: Mymaridae), egg parasitoids of leafhoppers (Hemiptera: Cicadellidae) in North America. Zootaxa 2010, 2428, 1–21. [Google Scholar] [CrossRef]
- Truett, G.E.; Heeger, P.; Mynatt, R.L.; Truett, A.A.; Walker, J.A.; Warman, M.L. Preparation of PCR-quality mouse genomic dna with hot sodium hydroxide and tris (HotSHOT). Biotechniques 2000, 29, 52–54. [Google Scholar] [CrossRef]
- Triapitsyn, S.V.; Rugman-Jones, P.F.; Adachi-Hagimori, T. Molecular and morphological differentiation within the Gonatocerus fuscicornis species complex (Hymenoptera Mymaridae). Bull. Insectol. 2021, 74, 181–200. [Google Scholar]
- Triapitsyn, S.V.; Adachi-Hagimori, T.; Rugman-Jones, P.F.; Barry, A.; Abe, A.; Matsuo, K.; Ohno, K. Egg parasitoids of the tea green leafhopper Empoasca onukii (Hemiptera, Cicadellidae) in Japan, with description of a new species of Anagrus (Hymenoptera, Mymaridae). ZooKeys 2019, 836, 93–112. [Google Scholar] [CrossRef]
- Triapitsyn, S.V. Anagrus (Hymenoptera: Mymaridae) Egg Parasitoids of Erythroneura spp. and Other Leafhoppers (Homoptera: Cicadellidae) in North American Vineyards and Orchards: A Taxonomic Review on JSTOR. Trans. Am. Entomol. Soc. 1998, 124, 77–112. [Google Scholar]
- Triapitsyn, S.V.; Rugman-Jones, P.F.; Tretiakov, P.S.; Luft Albarracin, E.; Moya-Raygoza, G.; Querino, R.B. Molecular, Morphological, and Biological Differentiation between Anagrus virlai sp. n., an Egg Parasitoid of the Corn Leafhopper Dalbulus maidis (Hemiptera: Cicadellidae) in the New World, and Anagrus incarnatus from the Palaearctic Region (Hymenoptera: Mymaridae). Neotrop. Entomol. 2018, 48, 87–97. [Google Scholar] [CrossRef]
- Triapitsyn, S.V.; Baquero, E.; Rugman-Jones, P.F. Anagrus avalae Soyka, 1956, a new synonym of A. bakkendorfi Soyka, 1946 (Hymenoptera: Mymaridae). Zootaxa 2021, 4941, 594–600. [Google Scholar] [CrossRef]
- Katoh, K.; Standley, D.M. MAFFT Multiple Sequence Alignment Software Version 7: Improvements in Performance and Usability. Mol. Biol. Evol. 2013, 30, 772–780. [Google Scholar] [CrossRef]
- Triapitsyn, S.V.; Rugman-Jones, P.F.; Tretiakov, P.S.; Daane, K.M.; Wilson, H. Reassessment of molecular and morphological variation within the Anagrus atomus species complex (Hymenoptera: Mymaridae): Egg parasitoids of leafhoppers (Hemiptera: Cicadellidae) in Europe and North America. J. Nat. Hist. 2020, 54, 1735–1758. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Suleski, M.; Sanderford, M.; Sharma, S.; Tamura, K. MEGA12: Molecular Evolutionary Genetic Analysis Version 12 for Adaptive and Green Computing. Mol. Biol. Evol. 2024, 41, msae263. [Google Scholar] [CrossRef]
- Triapitsyn, S.V.; Rugman-Jones, P.F.; Tretiakov, P.S.; Shih, H.T.; Huang, S.H. New synonymies in the Anagrus incarnatus Haliday ‘species complex’ (Hymenoptera: Mymaridae) including a common parasitoid of economically important planthopper (Hemiptera: Delphacidae) pests of rice in Asia. J. Nat. Hist. 2018, 52, 2795–2822. [Google Scholar] [CrossRef]
- Ahmed, M.Z.; Yadav, N.; Rustgi, S.; Saripalli, G.; Esquivel, I.L.; Bryant, T.B.; Graham, S.; Jacobson, A.L.; Gireesh, M.; Joseph, S.V.; et al. Global Phylogeography of Amrasca biguttula (Hemiptera: Cicadellidae) Across Eight Countries Reveals a Single-Haplotype Incursion Into the United States Beyond Its Putative Native Range. J. Appl. Entomol. 2026, 1–16. [Google Scholar] [CrossRef]
- Hébert, C.; Favret, C. Large-Scale Integrative Taxonomy of the Smallest Insects Reveals Astonishing Temperate Diversity (Hymenoptera: Chalcidoidea: Mymaridae). Mol. Ecol. 2025, 34, e70197. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.