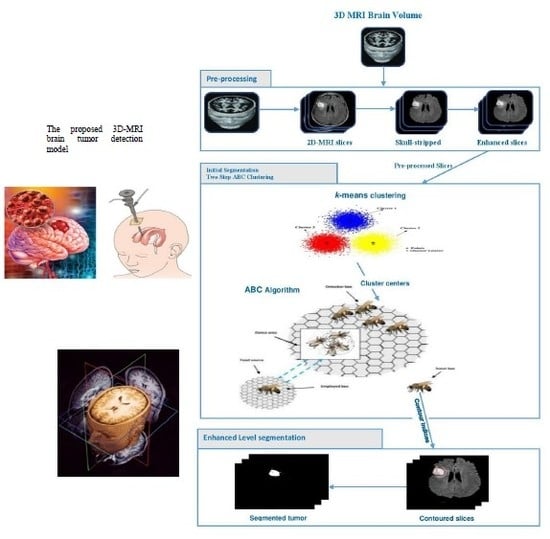
3D-MRI Brain Tumor Detection Model Using Modified Version of Level Set Segmentation Based on Dragonfly Algorithm
Abstract
1. Introduction
1.1. Problem Statement and Motivation
1.2. Contribution and Methodology
2. Related Work
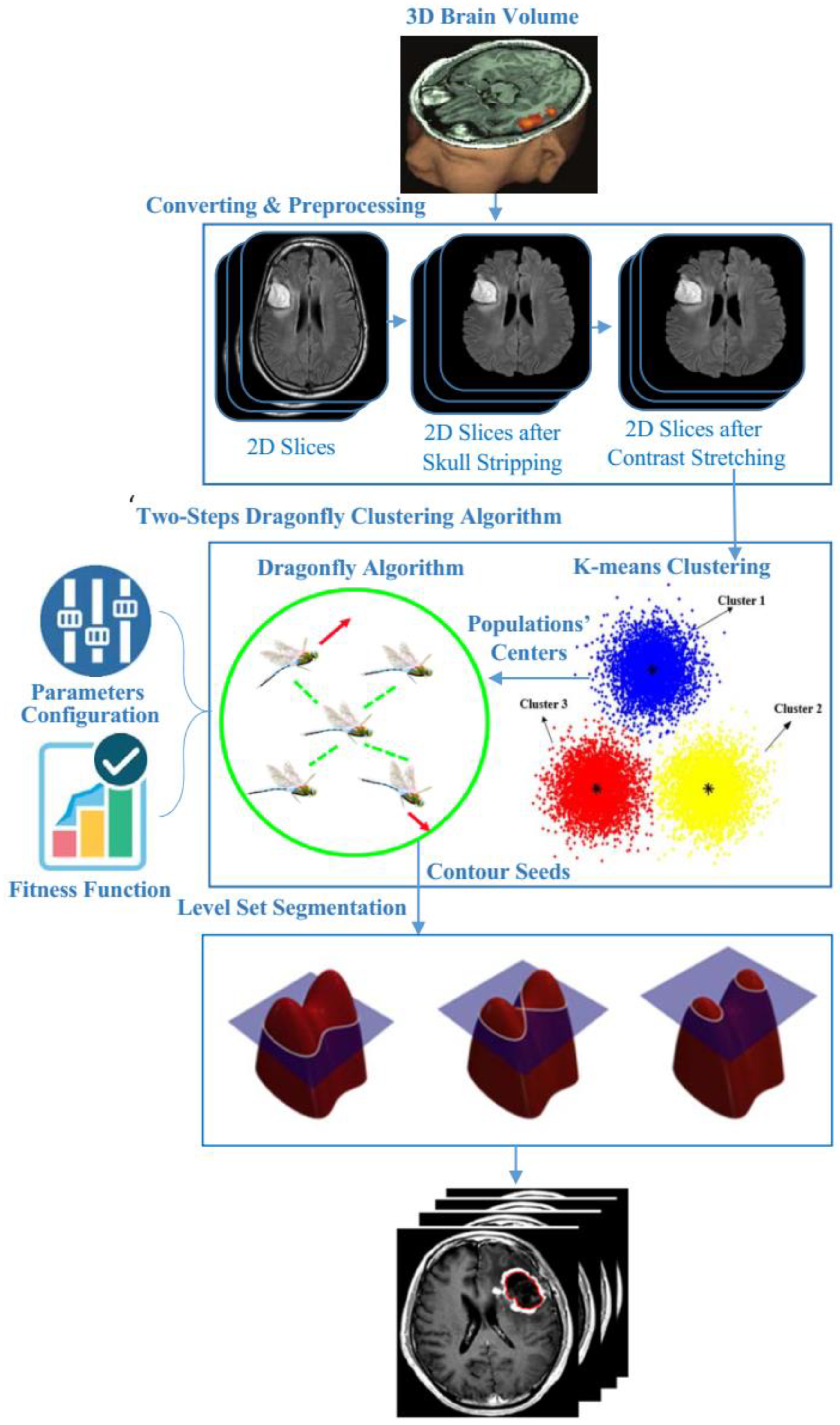
3. Materials and Methods
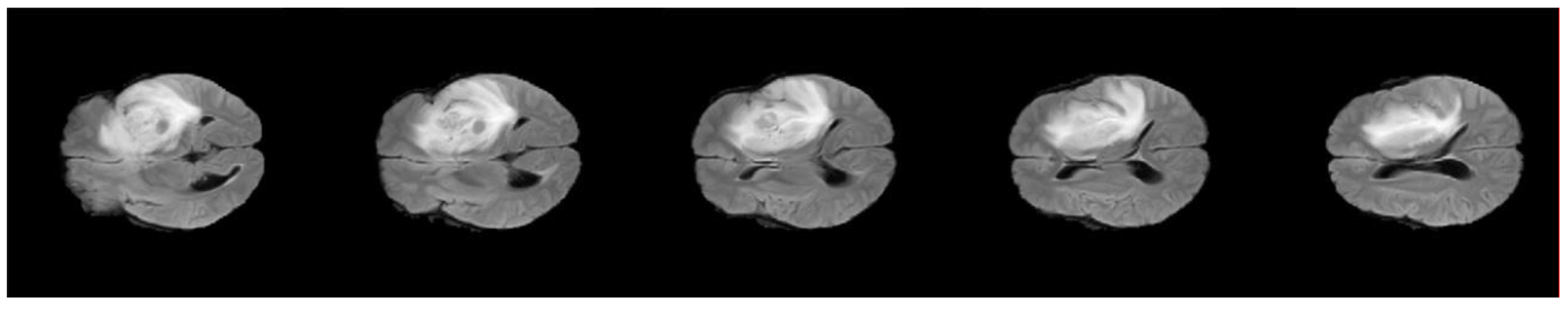
3.1. Preprocessing Phase
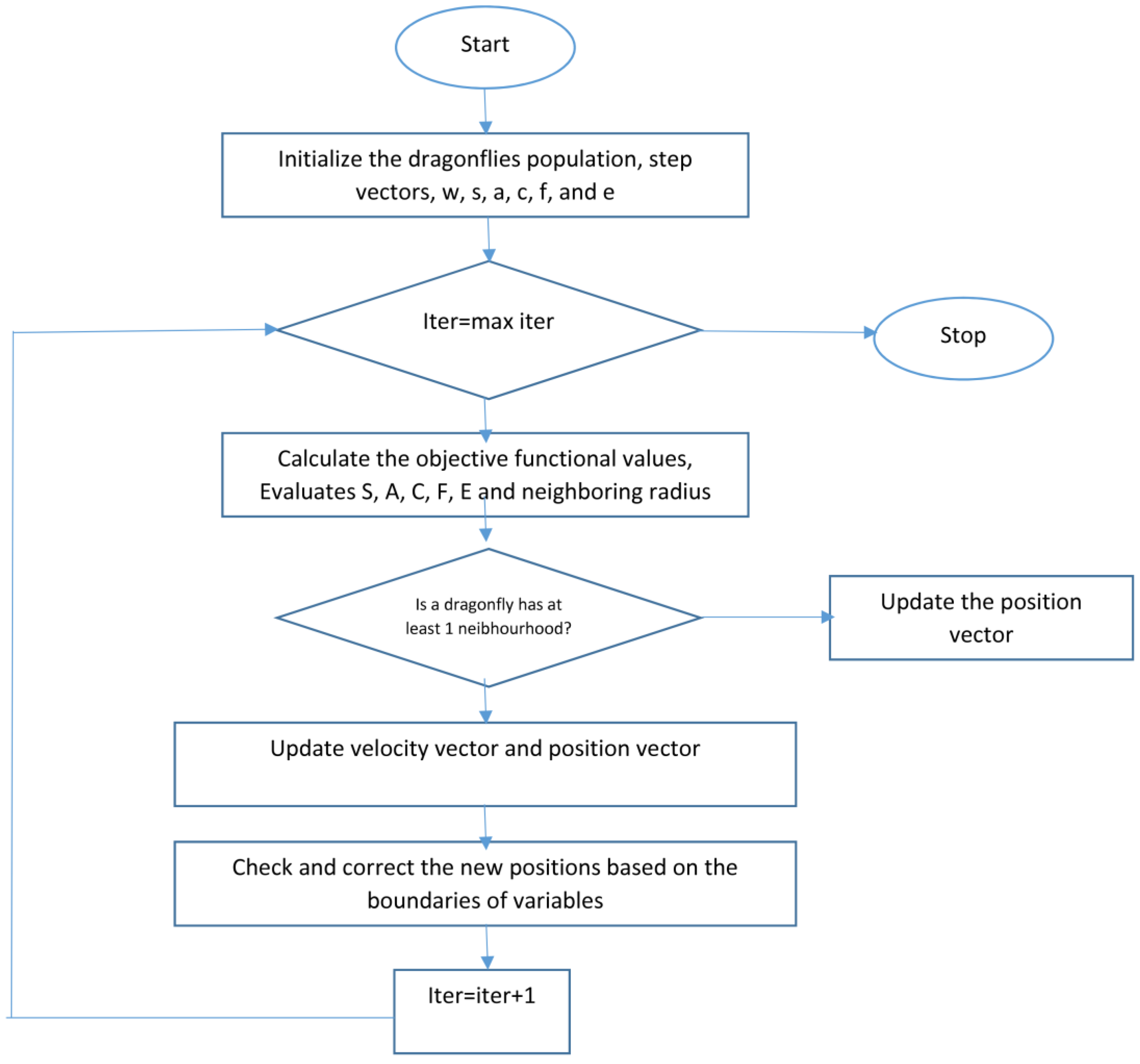
3.2. Two Steps Dragonfly-Based Clustering Phase
| Algorithm 1: Two-step Dragonfly Clustering. |
| Input: dataset contains MRI brain images Output: Best solution of final cluster center () Begin Initialization phase Initialize the position of dragonfly population Xi (i = 1 2, ..., n). Initialize step vectors Δ Xi For /* is the total number of food sources (number of clusters) */ Initialize the food source within the boundary of given dataset in random order; Evaluate the better potions of food sources by applying the k-means algorithm / *Algorithm 2*/ Send the dragonflies to the food sources; / * Computed centers */ End For Dragonfly algorithm Phase Iteration = 0; Do While (the end condition is not satisfied) For i = 1:n Calculate the fitness of each dragonfly Update the food source and enemy Update w, s, a, c, f, and e Calculate S, A, C, F, and E using Equations (4) to (8) Update neighboring radius If (a dragonfly has at least one neighboring dragonfly) Update step vector (ΔX) using Equation (9) Update position vector X using Equation (10) Else Update position vector using Equation (11) End if Check and correct the new positions based on the boundaries of variables End For For Compute the probability. /* Calculate the probability for each one */ End For For If (rand ( ) < ) /* denotes the probability associated with food source */ Calculate the new fitness of the new food source using Equation (14); Select the best food source by using a greedy selection between the old and new food source; Else ; End If End For End While Output: Final clusters‘ centers. End |
| Algorithm 2: K-means clustering [42]. |
| Input: . // the number of clusters; dataset contains MRI brain images (2D slices). Begin Arbitrary choose objects from as the initial cluster centers; Repeat - (re) group the most similar objects into a cluster, based on the Euclidian distance between the object and the cluster centroid (mean); - Update the cluster centroid, i.e., calculate the mean value of the objects for each cluster. Until no change. |
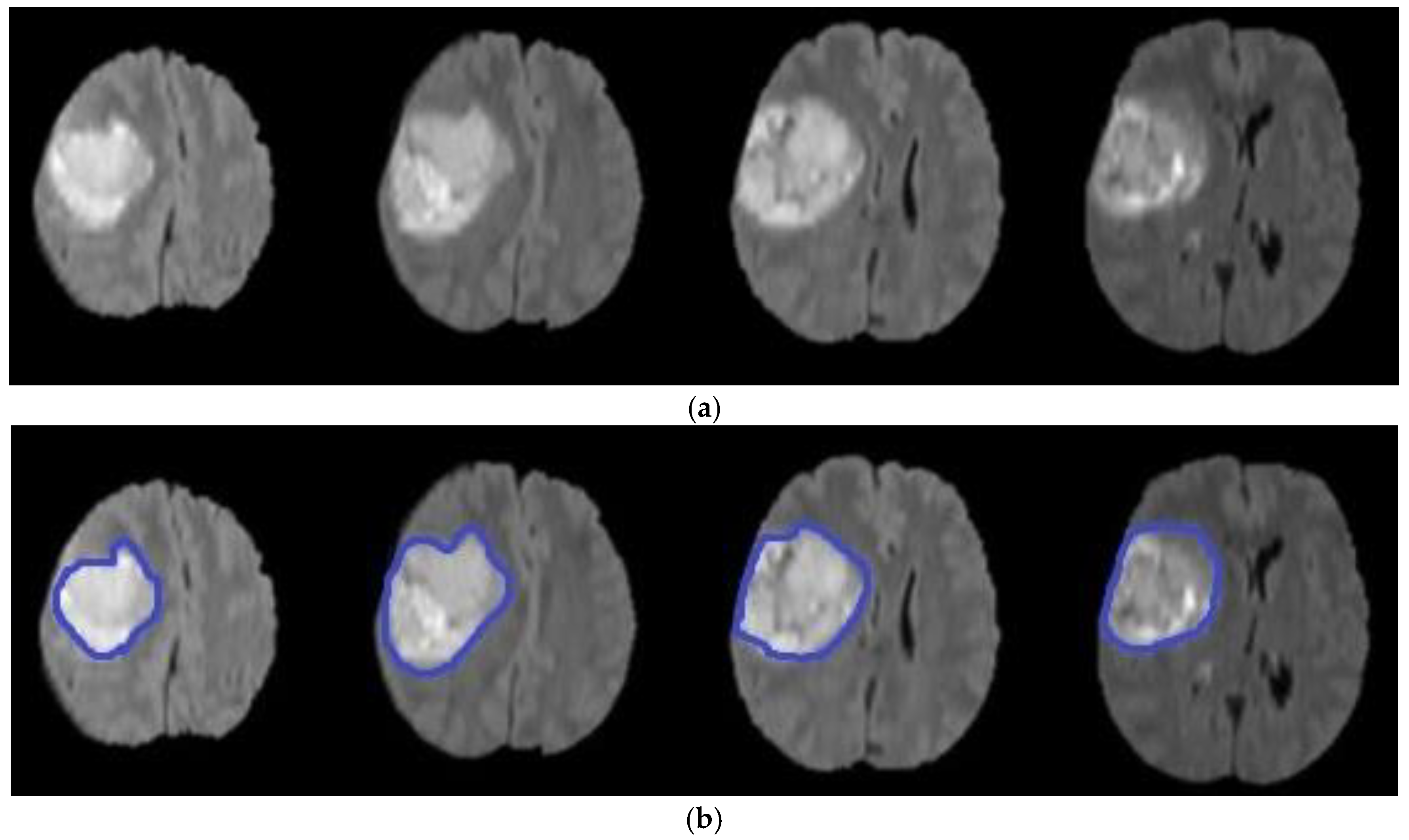
3.3. Level Set Segmentation
| Algorithm 3: Level set segmentation. |
| 1: Insert initial contour points using two-step DA clustering output (ROI indexes). 2: Construct a signed distance function. 3: Calculate feature image using Gaussian filter and gradient. 4: Obtain the curve’s narrow band. 5: Obtain curvature and use gradient descent to minimize energy. 6: Evolve the curve. 7: Repeat step number two and stop after obtaining the segmented region. |
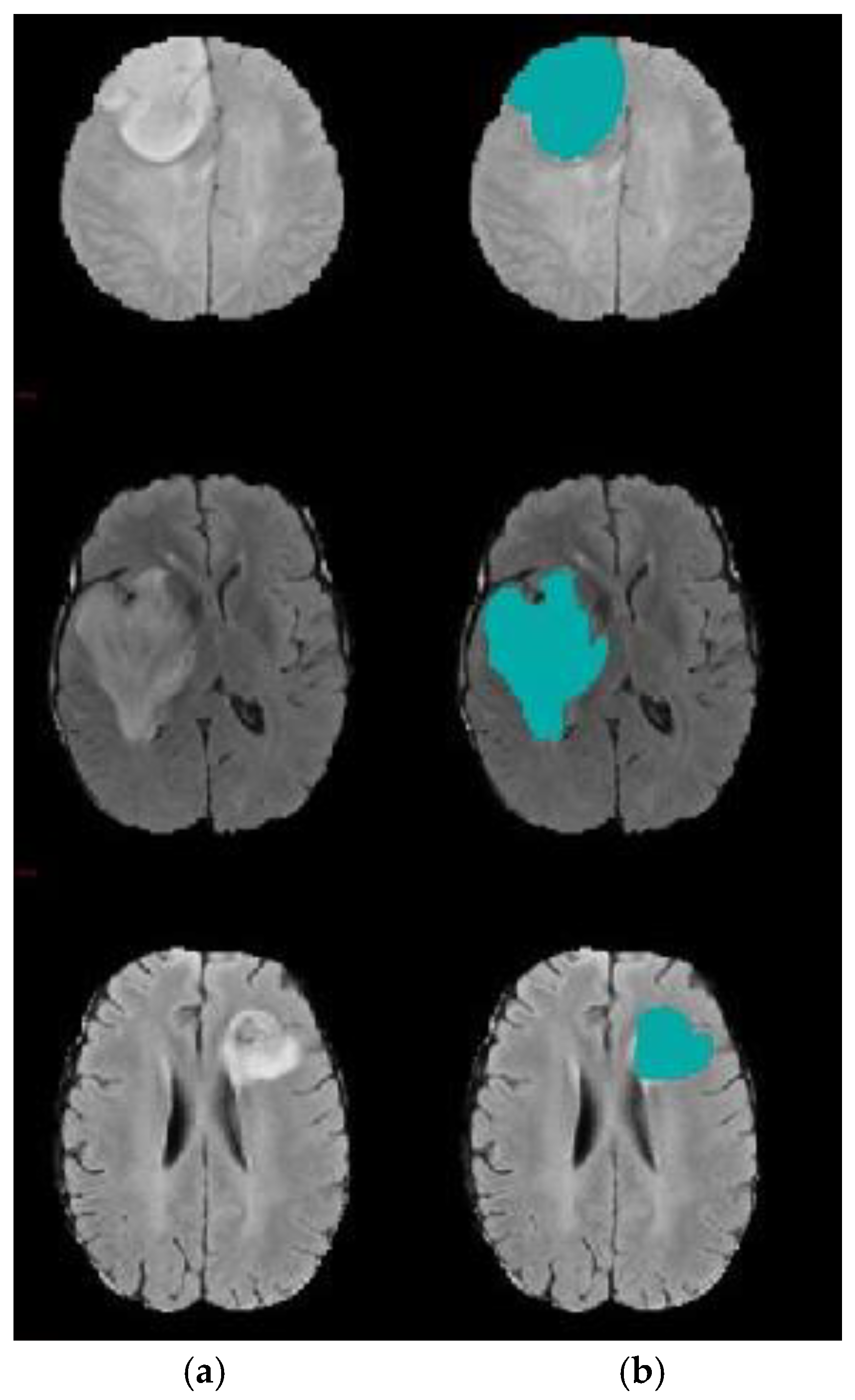
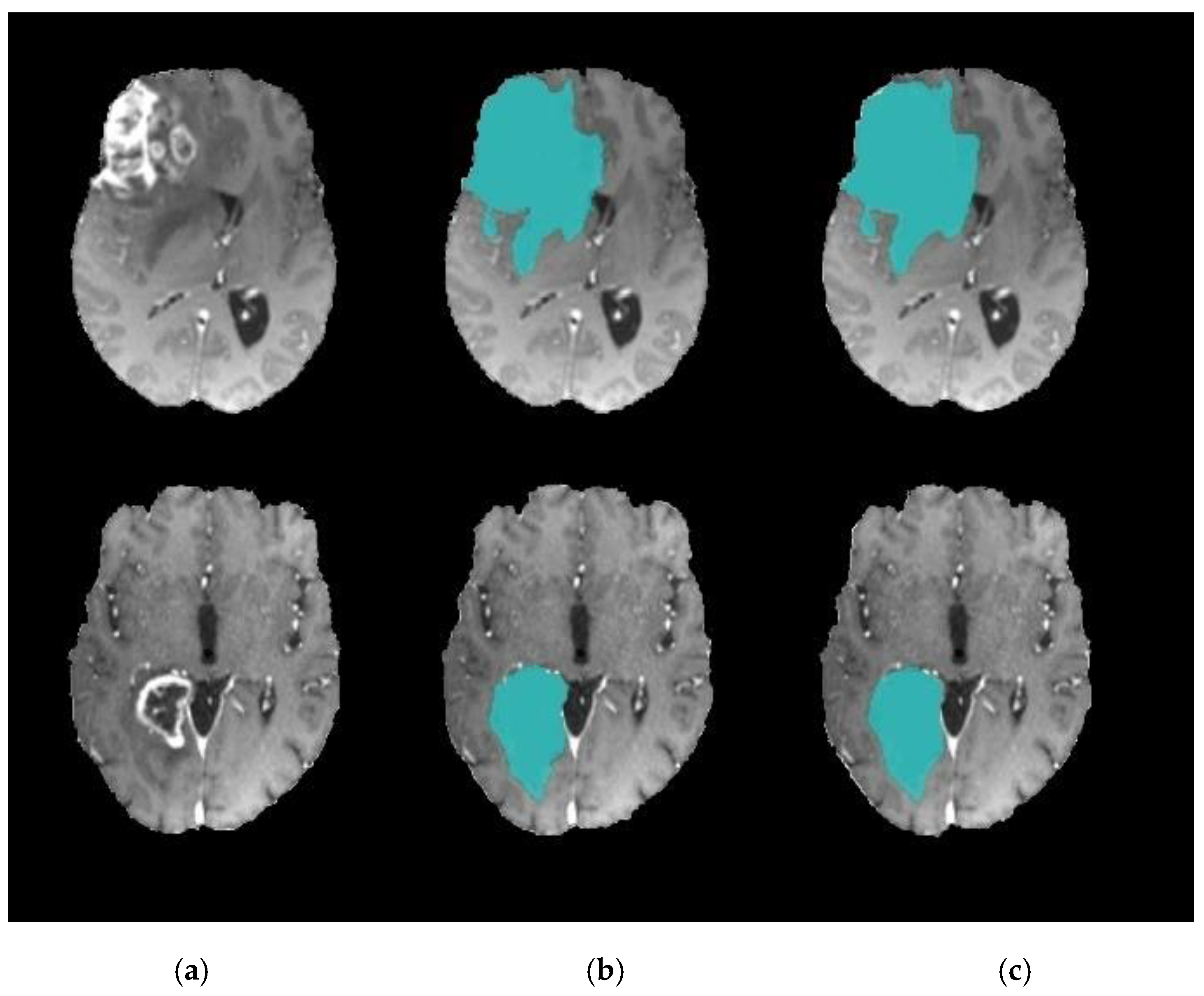
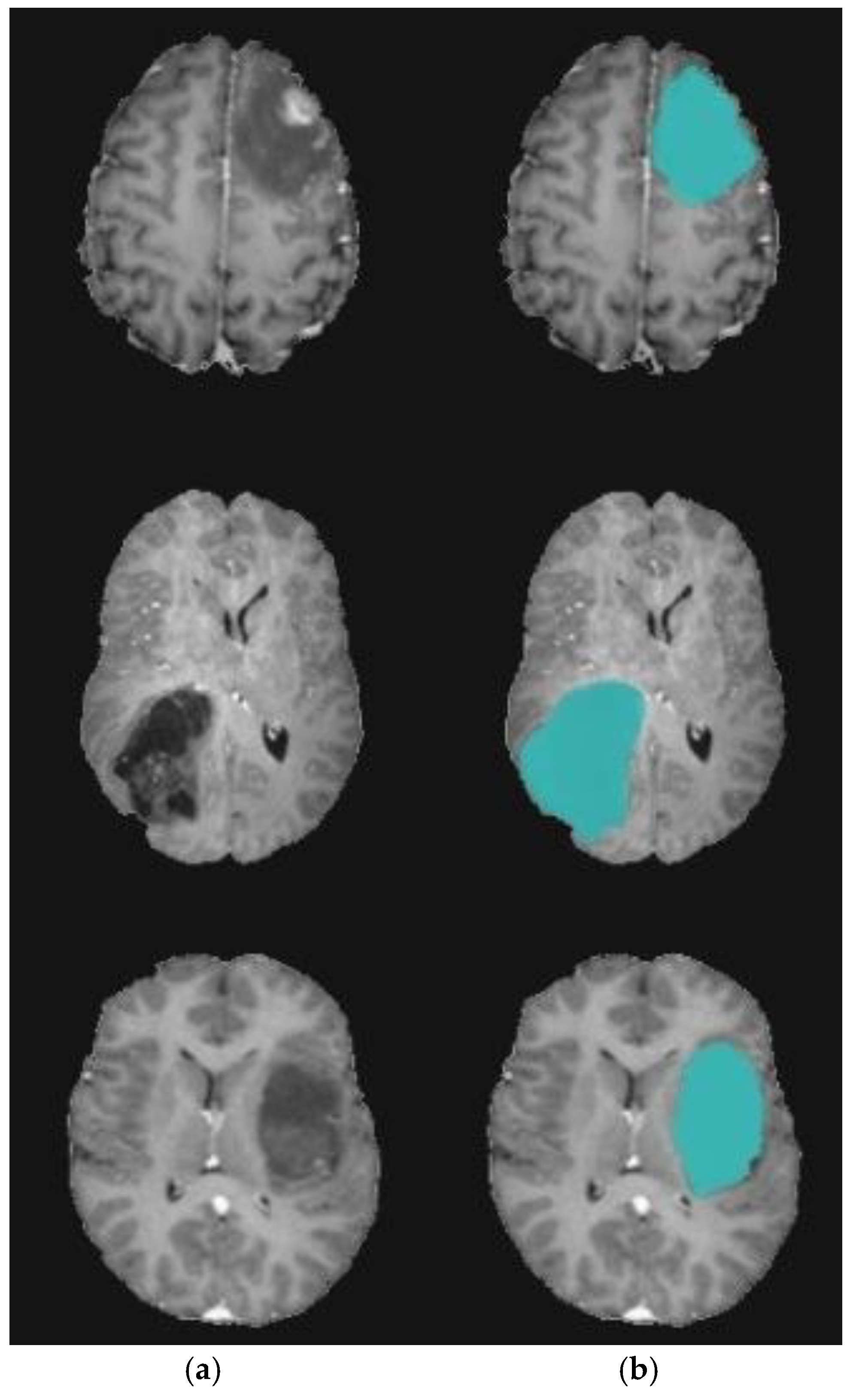
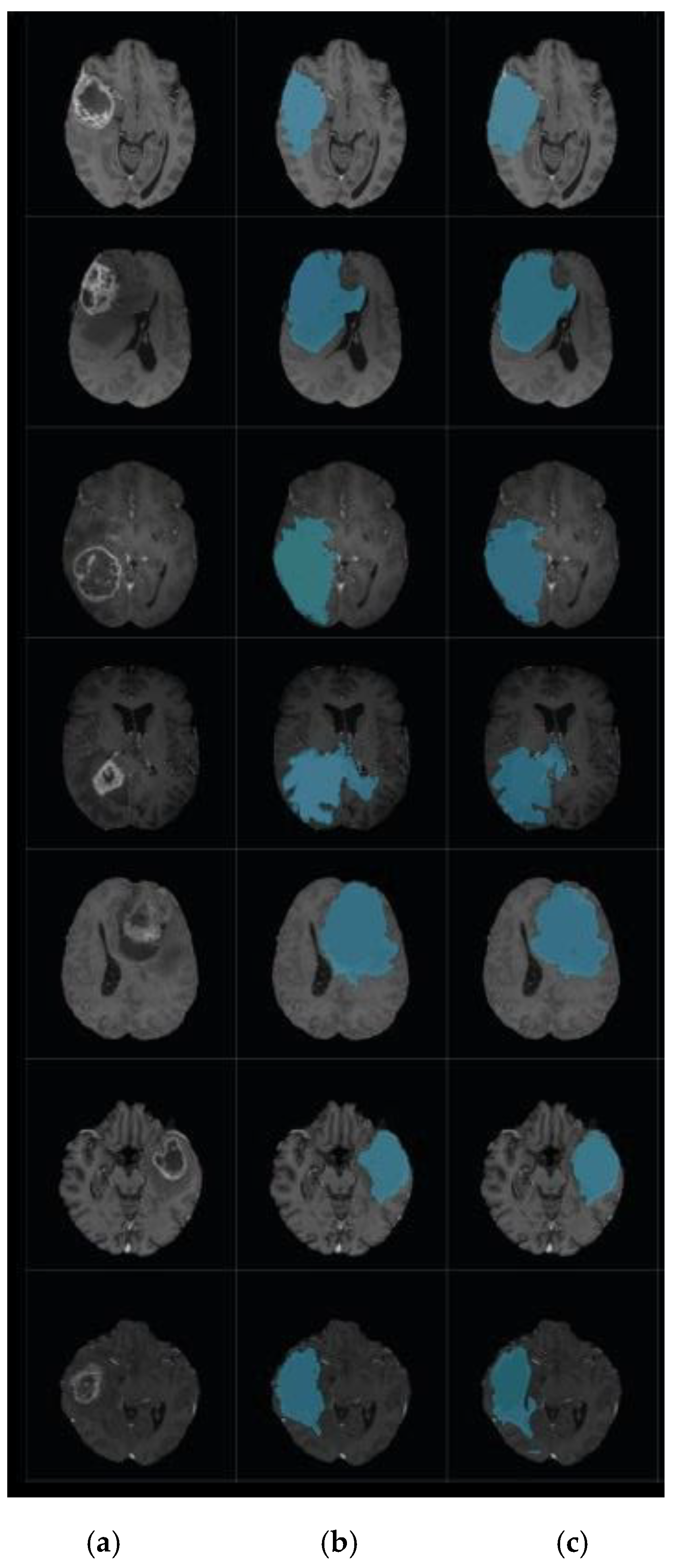
4. Experimental Results
4.1. Experiment 1: Comparison with Existing Methods
4.2. Experiment 2: Model Accuracy with and without k-Means
4.3. Experiment 3: Role of DA to Reduce Level Set Iteration
5. Conclusions and Future Work
Author Contributions
Funding
Conflicts of Interest
References
- El-Melegy, M.T.; El-Magd, K.M.A.; Ali, S.A.; Hussain, K.F.; Mahdy, Y.B. Ensemble of Multiple Classifiers for Automatic Multimodal Brain Tumor Segmentation. In Proceedings of the International Conference on Innovative Trends in Computer Engineering (ITCE), Aswan, Egypt, 2–4 February 2019; pp. 58–63. [Google Scholar]
- Aparna, R.M.; Shanmugavadivu, P. A Survey of Medical Imaging, Storage and Transfer Techniques. In Proceedings of the International Conference on ISMAC in Computational Vision and Bio-Engineering, Coimbatore, India, 16–17 May 2018; pp. 17–29. [Google Scholar]
- Lu, C.; Xu, Z.; Ye, X. Evaluation of Intraoperative MRI-Assisted Stereotactic Brain Tissue Biopsy: A Single-Center Experience in China. Chin. Neurosurg. J. 2019, 5, 1–10. [Google Scholar] [CrossRef]
- Bauer, S.; Wiest, R.; Nolte-P, L.; Reyes, M. A Survey of MRI-Based Medical Image Analysis For Brain Tumor Studies. Phys. Med. Biol. 2013, 58, 1–44. [Google Scholar] [CrossRef] [PubMed]
- Mild, K.H.; Lundström, R.H.; Wilén, J.H. Non-ionizing Radiation in Swedish Health Care Exposure and Safety Aspects. Int. J. Environ. Res. Public Health 2019, 16, 1186. [Google Scholar] [CrossRef]
- Zhuge, Y.; Krauze, A.V.; Ning, H.; Cheng, J.Y.; Arora, B.C.; Camphausen, K.; Miller, R.W. Brain tumor segmentation using holistically nested neural networks in MRI images. Med. Phys. 2017, 44, 5234–5243. [Google Scholar] [CrossRef]
- Banerjee, S.; Mitra, S. Novel Volumetric Sub-region Segmentation in Brain Tumors. Front. Comput. Neurosci. 2020, 14, 1–13. [Google Scholar] [CrossRef]
- Popoola, J.J.; Godson, T.E.; Olasoji, Y.O.; Adu, M.R. Study on Capabilities of Different Segmentation Algorithms in Detecting and Reducing Brain Tumor Size in Magnetic Resonance Imaging for Effective Telemedicine Services. Eur. J. Eng. Res. Sci. 2019, 4, 23–29. [Google Scholar] [CrossRef]
- Angulakshmi, M.; Priya, G.L. Automated Brain Tumor Segmentation Techniques—A Review. Int. J. Imaging Syst. Technol. 2017, 27, 66–77. [Google Scholar] [CrossRef]
- Shirly, S.; Ramesh, K. Review on 2D and 3D MRI Image Segmentation Techniques. Curr. Med. Imaging Former. Curr. Med. Imaging Rev. 2019, 15, 150–160. [Google Scholar] [CrossRef]
- Sajid, S.; Hussain, S.; Sarwar, A. Brain Tumor Detection and Segmentation in MR Images Using Deep Learning. Arab. J. Sci. Eng. 2019, 44, 9249–9261. [Google Scholar] [CrossRef]
- Wang, D. Efficient Level-Set Segmentation Model Driven by The Local GMM and Split Bregman Method. IET Image Process. 2019, 13, 761–770. [Google Scholar] [CrossRef]
- Rahman, C.; Rashid, T. Dragonfly Algorithm and Its Applications in Applied Science Survey. Comput. Intell. Neurosci. 2019, 2019, 11–21. [Google Scholar] [CrossRef]
- Seyedali, M. Dragonfly Algorithm: A New Meta-Heuristic Optimization Technique for Solving Single-Objective, Discrete, and Multi-Objective Problems. Neural Comput. Appl. 2016, 27, 1053–1073. [Google Scholar] [CrossRef]
- Ranjini, K.; Murugan, S. Memory Based Hybrid Dragonfly Algorithm for Numerical Optimization Problems. Expert Syst. Appl. 2017, 83, 63–78. [Google Scholar] [CrossRef]
- Saman, S.; Narayanan, S.J. Survey on brain tumor segmentation and feature extraction of MR images. Int. J. Multimed. Inf. Retr. 2018, 8, 79–99. [Google Scholar] [CrossRef]
- Tiwari, A.; Srivastava, S.; Pant, M. Brain tumor segmentation and classification from magnetic resonance images: Review of selected methods from 2014 to 2019. Pattern Recognit. Lett. 2020, 131, 244–260. [Google Scholar] [CrossRef]
- El-Baz, A.; Suri, J.S. Level Set Method in Medical Imaging Segmentation; CRC Press: Boca Raton, FL, USA, 2019. [Google Scholar]
- Al-Rifaie, M.; Aber, A.; Hemanth, J. Deploying Swarm Intelligence in Medical Imaging; Identifying Metastasis, Micro-Calcifications and Brain Image Segmentation. IET Syst. Biol. 2015, 9, 234–244. [Google Scholar] [CrossRef]
- Rupika, N.; Menon, H.; Vikram, K. A Survey on Advanced Segmentation Techniques for Brain MRI Image Segmentation. Int. J. Adv. Sci. Eng. Inf. Technol. 2017, 7, 1448–1456. [Google Scholar] [CrossRef]
- Kermi, A.; Andjouh, K.; Zidane, F. Fully automated brain tumor segmentation system in 3D-MRI using symmetry analysis of brain and level-sets. IET Image Process. 2018, 12, 1964–1971. [Google Scholar] [CrossRef]
- Bal, A.; Banerjee, M.; Chakrabarti, A.; Sharma, P. MRI Brain Tumor Segmentation and Analysis using Rough-Fuzzy C-Means and Shape Based Properties. J. King Saud Univ. Comput. Inf. Sci. 2018. [Google Scholar] [CrossRef]
- Anitha, V.; Murugavalli, S. Brain Tumor Classification using Two-Tier Classifier with Adaptive Segmentation Technique. IET Comput. Vis. 2016, 10, 9–17. [Google Scholar] [CrossRef]
- Mahalakshmi, A.; Krishnappa, H.; Jayadevappa, D. A Hybrid Approach for the Segmentation of Brain Tumor using K-Means Clustering and Variational Level Set. J. Adv. Res. Dyn. Control. Syst. 2018, 10, 258–264. [Google Scholar]
- Thapaliya, K.; Pyun, J.-Y.; Park, C.-S.; Kwon, G.-R. Level Set Method with Automatic Selective Local Statistics for Brain Tumor Segmentation in MR Images. Comput. Med. Imaging Graph. 2013, 37, 522–537. [Google Scholar] [CrossRef] [PubMed]
- Le, T.; Gummadi, R.; Savvides, M. Deep Recurrent Level Set for Segmenting Brain Tumors. Lect. Notes Comput. Sci. 2018, 11072, 646–653. [Google Scholar] [CrossRef]
- Qin, P.; Zhang, J.; Zeng, J.; Liu, H.; Cui, Y. A Framework Combining DNN and Level-Set Method to Segment Brain Tumor in Multi-Modalities MR Image. Soft Comput. 2019, 19, 9237–9251. [Google Scholar] [CrossRef]
- Rajan, P.G.; Sundar, C. Brain Tumor Detection and Segmentation by Intensity Adjustment. J. Med. Syst. 2019, 43, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Abdel-Maksoud, E.; Elmogy, M.; Al-Awadi, R. Brain Tumor Segmentation Based on A Hybrid Clustering Technique. Egypt. Inform. J. 2015, 16, 71–81. [Google Scholar] [CrossRef]
- Ural, B. A Computer-Based Brain Tumor Detection Approach with Advanced Image Processing and Probabilistic Neural Network Methods. J. Med. Biol. Eng. 2017, 38, 867–879. [Google Scholar] [CrossRef]
- Ma, C.; Luo, G.; Wang, K. Concatenated and Connected Random Forests with Multiscale Patch Driven Active Contour Model for Automated Brain Tumor Segmentation of MR Images. IEEE Trans. Med. Imaging 2018, 37, 1943–1954. [Google Scholar] [CrossRef]
- Hancer, E.; Ozturk, C.; Karaboga, D. Extraction of Brain Tumors from MRI Images with Artificial Bee Colony-Based Segmentation Methodology. In Proceedings of the 8th International Conference on Electrical and Electronics Engineering (ELECO), Bursa, Turkey, 28–30 November 2019; pp. 516–520. [Google Scholar]
- Christ, J.; Subramanian, R. Clown Fish Queuing and Switching Optimization Algorithm for Brain Tumor Segmentation. Biomed. Res. 2016, 27, 65–69. [Google Scholar]
- Narayanan, B.; Hardie, R. A Computationally Efficient U-Net Architecture for Lung Segmentation in Chest Radiographs. In Proceedings of the IEEE National Aerospace and Electronics Conference (NAECON), Dayton, OH, USA, 20–24 July 2020; pp. 279–284. [Google Scholar]
- Jin, L.; Pan, Y.; Li, M.; Chen, Z.; Tang, L.; Lu, C.; Wang, J. Applications of Deep Learning to MRI Images: A Survey. Big Data Min. Anal. 2018, 1, 1–18. [Google Scholar] [CrossRef]
- Mohammad, H.; Davy, A.; Warde-Farley, D.; Biard, A.; Courville, A.; Bengio, Y.; Pal, C.; Jodoin, P.-M.; Larochelle, H. Brain Tumor Segmentation with Deep Neural Networks. Med. Image Anal. 2017, 35, 18–31. [Google Scholar] [CrossRef]
- Iglesias, J.E.; Liu, C.-Y.; Thompson, P.; Tu, Z. Robust brain extraction across datasets and comparison with publicly available methods. IEEE Trans. Med. Imaging 2011, 30, 1617–1634. [Google Scholar] [CrossRef] [PubMed]
- Nair, R.; David, E.; Rajagopal, S. A robust anisotropic diffusion filter with low arithmetic complexity for images. EURASIP J. Image Video Process. 2019, 48, 1–14. [Google Scholar] [CrossRef]
- Chen, J.; Gong, Y. Particle swarm optimization for two-echelon location-routing problem. J. Comput. Appl. 2013, 33, 2261–2264. [Google Scholar] [CrossRef]
- Kumar, Y.; Sahoo, G. A two-step artificial bee colony algorithm for clustering. Neural Comput. Appl. 2015, 28, 537–551. [Google Scholar] [CrossRef]
- Zhang, C.; Shen, X.; Cheng, H.; Qian, Q. Brain tumor segmentation based on hybrid clustering and morphological operations. Int. J. Biomed. Imaging 2019, 2019, 1–111. [Google Scholar] [CrossRef]
- Armano, G.; Farmani, M.R. Clustering analysis with combination of artificial bee colony algorithm and k-means technique. Int. J. Comput. Theory Eng. 2014, 6, 141–145. [Google Scholar] [CrossRef]
- Arunprasath, S. Internet of medical things-load optimization of power flow based on hybrid enhanced grey wolf optimization and dragonfly algorithm. Future Gener. Comput. Syst. 2019, 98, 319–330. [Google Scholar] [CrossRef]
- Bao, X.; Jia, H.; Lang, C. Dragonfly algorithm with opposition-based learning for multilevel thresholding Color Image Segmentation. Symmetry 2019, 11, 716. [Google Scholar] [CrossRef]
- Li, C.; Xu, C.; Gui, C.; Fox, M.D. Distance regularized level set evolution and its application to image segmentation. IEEE Trans. Image Process. 2010, 19, 3243–3254. [Google Scholar] [CrossRef]
- Belaid, A.; Boukerroui, D.; Maingourd, Y.; Lerallut, J.F. Phase-based level set segmentation of ultrasound images. IEEE Trans. Inf. Technol. Biomed. 2010, 15, 138–147. [Google Scholar] [CrossRef] [PubMed]
- Menze, B.; Jakab, A.; Bauer, S.; Cramer, J.K.; Farahani, K.; Kirby, J.; Burren, Y.; Porz, N.; Slotboom, J.; Wiest, R.; et al. The multimodal brain tumor image segmentation benchmark (BRATS). IEEE Trans. Med. Imaging 2015, 34, 1993–2024. [Google Scholar] [CrossRef] [PubMed]
- James, J.A.; Dasarathy, B. Medical image fusion: A survey of the state of the art. Int. J. Inf. Fusion 2014, 19, 4–19. [Google Scholar] [CrossRef]
- Lefkovits, L.; Lefkovits, S.; Vaida, M. Brain tumor segmentation based on random forest. Mem. Sci. Sect. Rom. Acad. 2016, 39, 83–93. [Google Scholar]
- Ayachi, R.; Amor, N.B. Brain tumor segmentation using support vector machines. Lect. Notes Comput. Sci. Book Ser. (LNCS) 2009, 5590, 736–747. [Google Scholar] [CrossRef]
- Zhang, K.; Zhang, L.; Song, H.; Zhou, W. Active contours with selective local or global segmentation: A new formulation and level set method. Image Vis. Comput. 2010, 28, 668–676. [Google Scholar] [CrossRef]
| Methods | Accuracy | Recall | Precision |
|---|---|---|---|
| Proposed Model (Two-step DA, Level Set) | 98.20 | 95.13 | 93.21 |
| Symmetry Analysis, Level Set [21] | 93.63 | 89.10 | 90.45 |
| Fuzzy C-Means [22] | 85.7 | 87.6 | 72.3 |
| Rough Fuzzy C-Means [22] | 91.50 | 90 | 92 |
| K-means, Level Set [24] | 89.30 | 92.7 | 75.8 |
| Random Forest [49] | 85.60 | 91.85 | 78.3 |
| Support Vector Machine (SVM) [50] | 94.25 | 92.15 | 91.21 |
| DNN Methods | Accuracy | Recall | Precision |
|---|---|---|---|
| Proposed Model (Two-step DA, Level Set) | 98.15 | 95.40 | 93.57 |
| Two-pathway CNN [36] | 96.24 | 89.67 | 82.56 |
| DNN, level set [26] | 91.58 | 96.40 | 93.23 |
| Nature-Inspired Metaheuristic | Accuracy | Recall | Precision |
|---|---|---|---|
| DA, Level Set | 98.15 | 95.40 | 93.57 |
| ABC, Level Set | 95.90 | 92.13 | 91.40 |
| PSO, Level Set | 93.58 | 92.40 | 89.23 |
| CF, Level Set | 96.85 | 94.32 | 92.55 |
| Methods | Accuracy | Mean | Standard Deviation |
|---|---|---|---|
| k-means, DA and level set | 98.10 | 95.67 | 0.02 |
| DA, level set | 85.67 | 82.56 | 0.04 |
| Methods | Patient No.1 | Patient No.2 | Patient No.3 | Patient No.4 | Patient No.5 |
|---|---|---|---|---|---|
| Level set with DA clustering | 15 | 18 | 16 | 15 | 20 |
| Level set without DA clustering | 252 | 330 | 371 | 266 | 407 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Khalil, H.A.; Darwish, S.; Ibrahim, Y.M.; Hassan, O.F. 3D-MRI Brain Tumor Detection Model Using Modified Version of Level Set Segmentation Based on Dragonfly Algorithm. Symmetry 2020, 12, 1256. https://doi.org/10.3390/sym12081256
Khalil HA, Darwish S, Ibrahim YM, Hassan OF. 3D-MRI Brain Tumor Detection Model Using Modified Version of Level Set Segmentation Based on Dragonfly Algorithm. Symmetry. 2020; 12(8):1256. https://doi.org/10.3390/sym12081256
Chicago/Turabian StyleKhalil, Hassan A., Saad Darwish, Yasmine M. Ibrahim, and Osama F. Hassan. 2020. "3D-MRI Brain Tumor Detection Model Using Modified Version of Level Set Segmentation Based on Dragonfly Algorithm" Symmetry 12, no. 8: 1256. https://doi.org/10.3390/sym12081256
APA StyleKhalil, H. A., Darwish, S., Ibrahim, Y. M., & Hassan, O. F. (2020). 3D-MRI Brain Tumor Detection Model Using Modified Version of Level Set Segmentation Based on Dragonfly Algorithm. Symmetry, 12(8), 1256. https://doi.org/10.3390/sym12081256