Abstract
Protein interaction with polymers layers is a keystone in designing bio-nano devices. Polyamidoamines (PAMAMs) are well-known polymers. Zero aromatic core dendrimers (ZAC) are molecules with no proven toxic effect in cultured cells. When coating nanodevices with enzymatic systems, active sites are disturbed by an interaction with the biosystem surface. Computational methods were used in order to simulate, characterize, and quantify protein–polymer interaction. Protein corona, i.e., surface proteins disposed on a viral membrane or nanodevice outer surface, are crucial in interactions with a potential pharmacological target or receptor. Corona symmetry has been observed in the Middle East respiratory syndrome-related coronavirus (MERS-CoV), severe acute respiratory syndrome coronavirus (SARS-CoV), and severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2). As a protein alpha 1 antitrypsin’s a crystallographic structure was chosen. Protein–mono dendrimer layer systems were generated using in silico methods in order to simulate their interaction. Interactions were quantified using topological and quantum mechanical strategies. Results showed that PAMAM and ZAC interact differently with alpha 1 antitrypsin. Energy and topological surfaces of protein vary accordingly with the dendrimer monolayer. Topological surfaces have a higher sensibility in describing the interactions.
1. Introduction
Circulating blood proteins interact with polymers, especially with nanostructures coated by a polymer mesh [1]. Proteins tend to form a protein corona around the molecule [2]. Such coronas were observed in liposomes, gold, and fullerenes coated with polymers. Proteins tend to fix at the polymer layer and to circulate along with the coated structure interfering with its function [3]. Protein stability in the environment can be influenced by the polymer system resulting in protein precipitation [4].
Proteins interact with molecular surfaces by their functional groups present at the medium interface. In the case of enzymes, these interactions conduct enzyme inactivation as described in studies of enzymes fixed to the graphene surface [5,6]. Proteins, in aqueous solution, form a protein “corona” around the circulating coated polymer nanostructure (graphene, fullerene, gold clusters) [7]. The reactive residues of the protein surface can participate in reactions such as deamination, oxidation, glycosylation, and proteolysis. These reactions are protein dependent on the Aa sequence, but also solvent exposure and also B-factor dependent [8]. Interaction surface (dendrimers, a polymer layer, graphene) may favorize specific reactions like Schiff-base formation, SN2-nucleophilic aliphatic substitution, or act as Michael acceptor, or acyl transfer agents, respectively. Protein stability in a colloidal solution is described by the Zeta potential [9]. A Zeta potential domain between −60 and +60 mV ensures protein stability in solution. Zeta potential, when computed “in silico,” depends only on solution type, concentration (molar), pH, and temperature and is independent of the substrate [10].
Creating a computer simulation of protein folding is challenging. The large numbers of conformations cause biases in the methodology and consecutively interpretation of such simulations. In order to avoid the computationally expensive number of conformations, some methods use a minimum energy principle. Those simulations are based on Monte Carlo simulated annealing and genetic algorithms, respectively. Useful folding of a protein conformation needs an adjustment between speed, stability, and specificity, respectively. Computation can be used to identify the kinetic that cause the degradation of the protein. Also, protein folding can be used in order to solve the opposite problem, i.e., establishing a configuration of an amino acid sequence. Even here in computational design of proteins, size, stability, and folding speed are crucial.
Recent studies show that the folding of a protein is the best view as the property of a complex system. In this complex system, natures allow a limited number of possible conformations.
Dendrimers coating of the nanostructured surface is an accepted method for avoiding protein interactions. Although dendrimer coating is a practical approach in avoiding the toxic effects, the enzymatic system often presents in such assembly deteriorates. For instance, glucose oxidase (GOx) is present in a large number of nano-bio assemblies due to its property in producing H2O2. The struggle in designing a GOx-based system is avoiding interactions of dendrimers composing the coating layer with the amino acid chains of the enzyme, which are interactions that direct to enzyme inactivation or diminished activity.
In this study, alpha 1 antitrypsin (A1AT) was considered due to its trypsin inhibition properties. A1AT is used alone as a concentrate that is administrated intravenously. Aerosolized augmented A1AT therapy is under study. However, A1AT may not reach elastin fibers in the lung where elastase injures the tissue. Finding suitable transport media, i.e., as nanoparticles, might improve its drug effect. In this instance, A1AT interaction with the dendrimer layer coating the nanosystem surface is crucial. The methodology used here applied to A1AT–dendrimer monolayer system may be used to other protein/enzyme–polymer systems.
Poly(amidoamine) (PAMAMs) considered in this study is a class of dendrimers made of repetitive branch subunits of amide and amine. PAMAMs have a sphere-like shape and tree-like branching. A distinct property of these dendrimers is their high density of surface topological groups. This aspect allows many alterations to be made to the surface of each molecule, making this class of compounds a valuable component of a nanodevice.
Aspects of the interaction mechanisms of biomolecules and inorganic surfaces are unclear even if significant experimental research on protein adsorption onto solid substrates has been eventually described. For exploring protein–surface binding mechanisms, as well as the thermodynamics and kinetics of adsorption analytical modeling and simulations techniques, have been applied. The models and force fields (FFs) describing the interactions are different. Notably, FF and water models designed for use in biomolecular simulations are sometimes not precisely transferable to the simulation of surface and backward. The adsorption phenomena show a complex range of time and length that fluctuate from nanoseconds to days, and from nanometers to micrometers. Additionally, changes in the atomic structure of the surface can lead to surface rearrangement that can result in complete denaturation of the adsorbed molecules. These structural changes can generate intermediate structural and energetic states that complicate sampling.
Lastly, this study aims to comprehend the protein–organic surface interactions in the context of the rational design of new systems in nanomedicine [11]. Computational modeling supplies extra information regarding the structure and intrinsic mechanism of protein organic surface interaction.
2. Materials and Methods
Protein 1ATU, with a molecular weight of 41.97 kDa, was chosen for this study. The protein interaction was studied against a PAMAM dendrimer monolayer, of various generations: C22H48N10O4 (G0), C62H128N26O12 (G1), C110H192N26O44 (G1.5), C142H288N58O28 (G2), C302H512N122O60 (G3) and a series of several “zero-generation aromatic” core dendrimers (ZAC) —C16H28N2O4, C20H36N2O4, C22H44N2O4, C24H45N3O6, C28H52N2O4. A total of 10 protein-PAMAM complexes were computed. Also free, denaturated 1ATU, and 3D 1ATU were taken into account and compared with the resulting denaturated proteins string. Systems were optimized for normal intravascular conditions (NaCl 0.15 M, temperature 310.15 K, a dielectric solvent constant of 80) with respect to published data. The water effect was taken into account by protonating the system at physiological intravascular conditions. The force field used for all computations was AMBER10 [12].
Interaction between the two components, the protein and dendrimer layer, is viewed as an inverse receptor–ligand interaction, were the ligand models the receptor. The result of this technique is a protein “string” having primary, secondary, but no tertiary or quaternary structure. In essence, this particular folding is a result of in silicon increase in protein flexibility. Strings are stable conformations resulting in an interaction with the dendrimer layer; interactions with were amplified dramatically by heightening protein flexibility. By altering the quaternary and tertiary structures, these are irreversible states. These conformations were computationally checked to see if they are biologically accessible to the protein by computing protein conformation energy and Ramachandram plots. The strings, as a result of an interaction between 1ATU and dendrimer layers, were further computed. The analysis included energy calculations, topological descriptors, and some physicochemical properties (partial charges, surface area, volume, log P, refractivity, polarizability, mass) [13]. Biochemical data regarding reactions that could induce a protein binding-absorption and further toxicity, from the polymers point of view, were computed [14]. Free 1ATU polymer string and the 3D structure of 1ATU were also taken into study. Graphs of 3D coordinates (x,y,z) for all the 12 structures, were represented in order to show the differences in string topology, i.e., different shapes for each 1ATU–polymer pair and to determine the topological site of the properties [15].
Protein aggregation surface was studied on all the protein strings resulted, also in the “native” 1ATU structure and the “free PAMAM string.” The interactive biological properties and their topology (in terms of biochemical reactions, aggregation) were described and compared [16]. The global process of aggregation was also characterized by computing Zeta potential for a domain of temperatures from 273 °K to 320 °K, and a pH domain around 7.4 [17]. Studies were performed using the Schrodinger software package [18].
To compute the interaction between the protein and the dendrimer layer (e.g., PAMAM, ZAC), the protein–nanostructure interactions were simulated. A system of proteins and different dendrimers monolayers was generated. A procedure to distort the protein from the lavage of the quaternary structure in the secondary structure was used. In this procedure, the mobility of the quaternary and tertiary arrangements was allowed. At the end of these calculations, protein strings were obtained. The protein (1ATU) was subjected to the free process (not in the presence of the dendrimer layer), and 1ATU was also characterized without being distorted.
For the topological description for each string of proteins—three logarithmic functions (Table 1) were unified, integrated and and as such reported (virtually representing a graph of 3D points). An example is given in case of PAMAMG0 crude protein string. The expression used to construct the points on the 3D graph, i.e., the manifold, is as follows:
ʃ[(2.372log(x) + 71.411 U (0.01076log(x) + 28.956 U (2.8141log(x) + 29.356)]dx.

Table 1.
Protein string conformational potential energy (kcal/mol) and globularity.
Further, a potential energy surface map for each protein string was developed. Algorithm design for generating this map is based on the formula: Etotal = Ebond + Eangle + Etorsion + Eelectro + Evd, where Etotal is the molecule potential energy, Ebond- bond stretch energy, Eangle – angle bend energy, Etorsion- torsion energy, Eelectro- electrostatic energy, Evd- Van der Waals energy, all expressed in kcal/mol. Right equation terms for all protein strings were computed. A polynomial fourth-degree fitting function was used to generate a manifold. Two types of manifold surfaces were generated: a surface base on the representation of the fitting function and a second one based on the integration of the fitting fourth-degree polynomial function. As an example of PAMAM G0 protein string, the expressions used were, respectively:
277.730 x4 − 3790.600 x3 + 18236 x2 − 35433 x + 23590,
∫(277.730x4 − 3790.600x3 + 18236x2 − 35433x + 23590)dx
As in the instance of topological surfaces, the results were compared with surfaces generated for 3D protein structure where tertiary and quaternary arrangements are intact.
Surface areas for all protein strings were computed and represented. The aggregation score and aggregation surfaces for all protein strings were represented using the Schrodinger software package. Zeta potential for different pH and temperature domains was computed. The aggregation score used is based on AGGRESCAN methodology. The algorithm identifies aggregation-prone segments in proteins [18].
3. Results
An example of the molecular system obtained is illustrated in Figure 1 for G0; G1.5; G2; G3; C16H28N2O4; C20; C20H36N2o4; C22H40N2O4; C24H45N3O6 and C28H52N2O4, respectively (see Supplementary Materials).
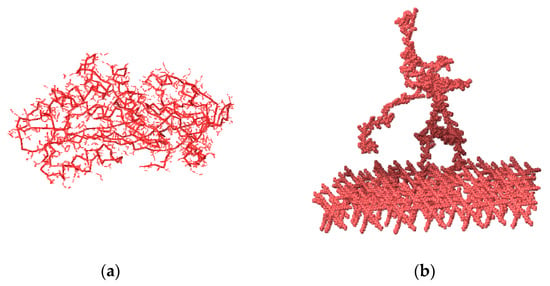
Figure 1.
1ATU system and G1.5 PAMAM. When arbitrary valences are allowed, a protein, i.e., “the string’’ is formed as the expression of forces of the PAMAM layer applied to the protein. (a) Protein 1ATU in its 3D crystallographic determine conformation. (b) Protein string fixed on the G1.5 PAMAM layer.
Cartesian coordinates of the protein string showed distinct patterns for each string of the corresponding protein-dendrimer system. Two major types of patterns were distinguished: a pattern corresponding to the free protein with intact tertiary structure and a second pattern closed to generation 1 dendrimer.
Nonintersection of Cartesian coordinates at the terminal end (the last 500 topological points) correlates with a lower rate of interaction of the protein with the polymer layer (there are the same patterns as the free protein string and the free protein with tertiary structure).
Ramachandran plot for 1ATU free, 1ATUG0, 1ATUG1.5, and 1ATU, respectively, are represented in Figure 2.
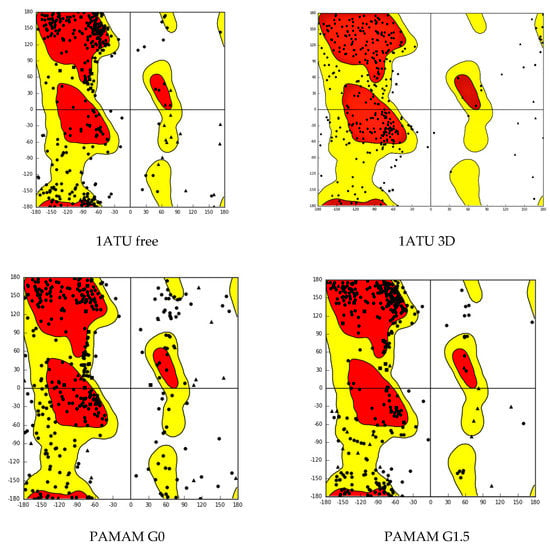
Figure 2.
Ramachandran plots for 1ATU free, 1ATUG0, 1ATUG1.5, and 1ATU. Vertical psi (degrees), horizontal phi (degrees). Red core, yellow allowed region, and points outside the contours represent the outlier (see Supplementary Material-S1).
The Cartesian coordinate obtained for all the structures are represented by a scatter plot (Figure 3). For each diagram, three logarithmic equations for the x, y, and z coordinates were calculated. Such a scatter plot is represented here for the PAMAM G1.5 string (see the rest of the graphs in Supplementary Materials).
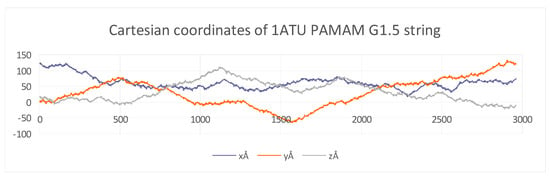
Figure 3.
Cartesian coordinates of 1ATU after interaction with PAMAM G0 showing variations for each string: X-axis (no. atoms: 1 ATU~3000 atoms); Y-axis (coordinates, in Å); (see Supplementary Materials-S2).
Some protein strings, obtained by the above-described methodology, are shown in Figure 4. All resulting “strings” are microstates energy minima at a stable conformation representative of the phase space.
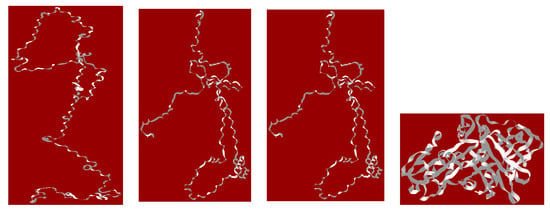
Figure 4.
1ATU protein strings after interaction with PAMAM dendrimers are represented alone (without the dendrimer monolayer) and named according to their interaction with the type of dendrimer monolayer (from left to right): 1ATUfree, 1ATU G0, 1ATU G1.5, 1ATU 3D, with the dendrimer layer (for the all protein strings, see Supplementary Materials-S2).
The resulted protein strings represented according to their surface area and volume resulted from the interaction with the polymer layer (Figure 5).
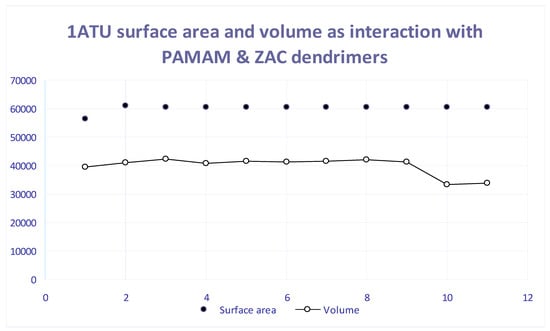
Figure 5.
Surface area and volume of 1ATU protein string in the presence of different dendrimers layers. Ox axis, according with the entries on Table 1: 1—PAMAM generation 0, 2—PAMAM generation 1, 3—PAMAM generation 1.5, 4—PAMAM generation 2, 5—PAMAM generation 3, 6—C16H28N2O4, 7—C20H36N2O4, 8—C22H40N2O4, 9—C24H45N3O6, 10—C28H52N2O4, 11—1ATU Free, 12—1ATU 3D. Oy axis: Surface area (Å2) and Volume (Å3).
Topological properties analysis showed a double value of the cluster count of protein string generated by the G1.5 layer. Some topological descriptors are unmodified, and some are sensible to polymer layer action: Balaban index, molecular topological index, and the topological diameter, respectively (see Supplementary Materials).
The aggregation surface of protein strings was computed. Results show that 1ATU and resulted in protein strings have outer regions susceptible to aggregation (Figure 6).
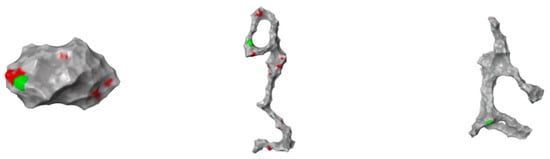
Figure 6.
Structure of 1ATU (from top to bottom): “native” (top); “free” (with no protein interaction—middle) and G1.5 (with G1.5 PMAM interaction—bottom). Aggregation surface was calculated; all of the Aa domains that may contribute to an aggregation process are represented in red; Aa with the highest score, the chance to produce an aggregation is represented in green. For each protein string, a distinct Aa set is identified accordingly to the string’s geometry.
Aggregation scores of the protein strings were represented. The lowest score has the string generated by the G1.5 layer. Comparison among the native protein score, “free” string score, and the G1.5 score is shown in Figure 7.
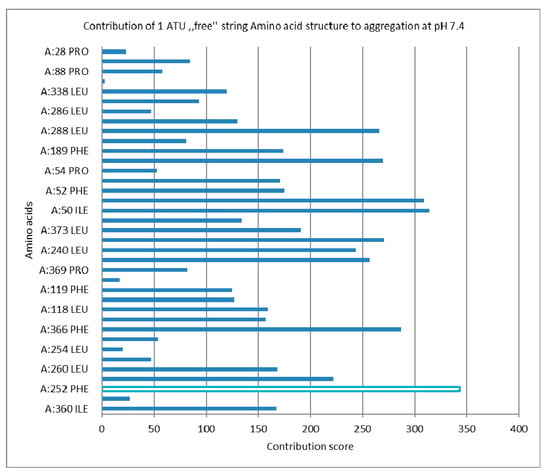
Figure 7.
Aggregation score of the native protein, 1 ATU free string, most significant Aa contribution is highlighted (see Supplementary Materials-S3).
Zeta potential, computationally calculated, for the 1ATU structures at different pH domains shows excellent stability of the protein in solution (Figure 8); see Supplementary Materials-S4.
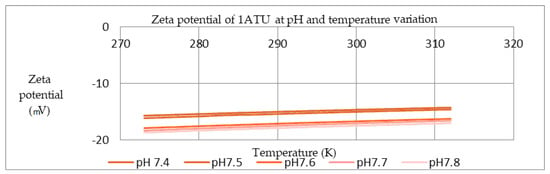
Figure 8.
Zeta potential of 1ATU at different pH and temperature domains.
In Table 2, Equations obtained for all protein strings using strings topological coordinates are represented.

Table 2.
Cartesian coordinate string equations.
The resulting protein string-based manifolds are shown in Figure 9 for 1ATU free, 1ATUG0, 1 ATU G1, and 1atu 3D, respectively.
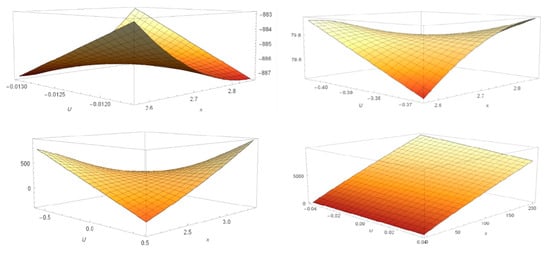
Figure 9.
Manifolds obtained using 1ATU string coordinates equation: (from up to bottom) 1ATU string free, 1ATU G0 (top); 1ATU G1, 1ATU 3D (bottom) (see Supplementary Materials-S5).
The protein strings, resulted from the interaction with dendrimers, show a distinct topology. Their representation envisaged approximately the same morphology (same curvature, angle, orientation). Differences are depending on the type of dendrimer used. When the protein is denatured computationally, without the gradient of the dendrimer layer, there is an opposite bending compared to the dendrimer-protein system. The graphical representation obtained by placing coordinates of the integrated logarithmic function for 3D 1ATU, without being denatured, shows a relatively flat shape with some small folded regions corresponding to the functional protein regions.
Potential energy manifolds obtained show notable differences between the two classes of dendrimers. In Figure 10, manifolds obtained for 1ATU string free, 1ATU G0 (top), 1ATU G1, 1ATU 3D (bottom) are represented using both computation algorithms.
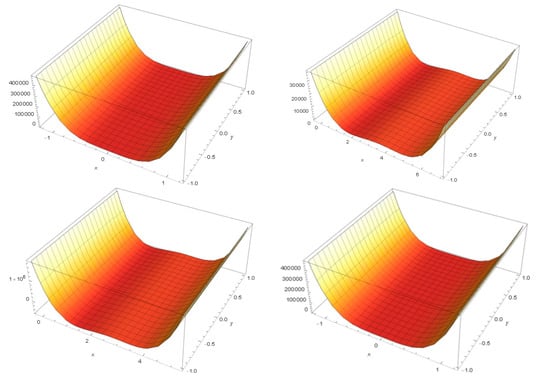
Figure 10.
Manifolds obtained using 1ATU string coordinates equation: (from up to bottom) 1ATU string free, 1ATU G0 (top); 1ATU G1, 1ATU 3D (bottom); see Supplementary Materials-S6.
For the rest of protein strings, the manifold obtained had the following aspect, as shown below for PAMAM G2 manifold and C22H4N2O4, respectively (Figure 11).
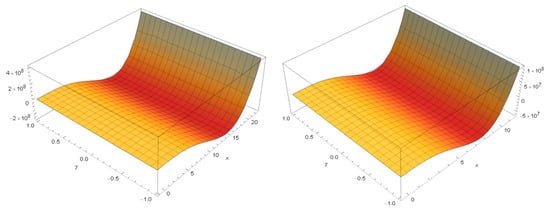
Figure 11.
Manifolds for PAMAM G2 and C22H4N2O4.
Manifolds obtained using equation integration had the same behavior (see Supplementary Materials).
Topological descriptors computed for protein strings are represented in Figure 12. Descriptors that are sensitive to string morphology were the Wiener index, molecular topological index, and Balaban index, respectively.
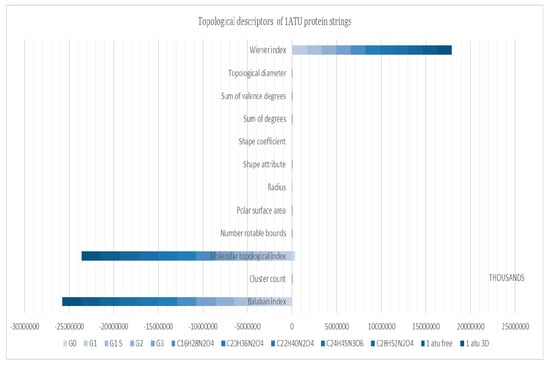
Figure 12.
Topological descriptors of protein strings (see Supplementary Materials-S7).
Solvation energy represented in Figure 13 suggests a high tendency of aggregation of proteins after interaction with dendrimer layers (see Supplementary Materials).
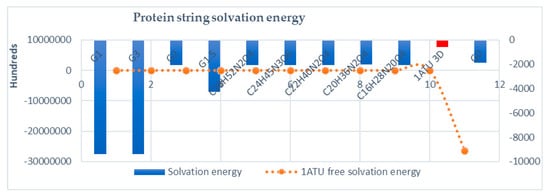
Figure 13.
Solvation energy of protein strings. In red, the solvation energy for 3D 1ATU crystallographic structure (kcal/mol). Free, computationally denaturated 1ATU energy is represented in orange as a combo graph due to higher dimensionality of data (−26,370,109 kcal/mol).
In Figure 12 and Figure 13, higher dimensionality data is presented, using some molecular descriptors and solvation energy.
Figure 12 shows some distinct molecular descriptors computed for the studied compounds. Molecular descriptors are designed to retrieve a distinct number for each structure that is computed, which means that even for small structures, molecular descriptors are composed of 3–4 digits. When computed for a structure (i.e., 1ATU) that contains 2962 atoms, such big numbers are retrieved.
The more complex the structure is, the ‘’bigger’’ the certain molecular descriptor. Those descriptors were used in order to emphasize that the protein strings, even if they have the same molecular formula, they contain distinct information due to different geometry, charge, topology, and so on. Between the descriptors computed that are not degenerated (meaning that they are distinct for each structure), the Winner index, molecular topological index, and Balaban index are distinct, and are able to contain a significant amount of structural information judging by their significantly higher value (Balaban index~25.000.000) in comparison to the rest of the descriptors. As it is observed, 1ATU has the ‘’biggest’’ value in terms of high dimensional data followed in every case by the same sequence (1ATU free, ZAC dendrimers C28 decreasing to C16, and finally by PAMAM G3, G2, G1.5, G1, G0), showing that the G0 protein string “carries” less structural information in comparison with the rest of the series.
In Figure 13, solvation energy is represented. While the protein string is of interest, solvation energies are computed in a closed system. In this case, the computations show high numbers in order to detect, even the smaller difference in protein string energy, while the same structure (same molecular formula) is computed. As expected, the 3D structure (1ATU) has the smallest solvation energy, i.e., is more solvable that the rest of the members. In the rest, protein strings resulted as the interaction of 1ATU with dendrimers monolayer have disproportionately higher solvation energies, i.e., hundreds of times more susceptible to protein denaturation and aggregation in the solvent while in contact with dendrimers monolayer.
4. Discussion
Protein or nucleic acid folding/unfolding is a critical process in biology [19]. Denaturation, the process in which the quaternary structure is altered, is relevant to the disclosure of the secondary and primary protein structure, included in the native protein state [20]. The denatured proteins illustrate a wide range of properties, a representative for the loss of aggregation [21,22,23]. When proteins are “fixed” on covalent or noncovalent surfaces, a phenomenon similar to denaturation occurs. There is no doubt that one of the most commonly observed consequences of attaching proteins to the surface of nanomaterials of carbon and plausible nanopolymers is the appearance of conformational modifications of their structures, which can lead to exposure of new epitopes, changes in functionality and avidity of proteins. therefore, to induce a robust immune response or unintended toxicity. The phenomenon of protein interaction with the surface of nanomaterials is a very complicated process, not only because of the dynamics of the observed changes, a similar effect to the effect of Vroman, which can never reach an equilibrium in vivo, but especially the complexity of the parameters of the tested system that determines the processes of adsorption of the course and thus forming the layer of biomolecular composition [24].
In short, the "corona protein" as studied in our paper, is a multi-layered structure, in contact with the protein–nanomaterial interface. The “hard” part of this structure is formed by the surface recognition of nanomaterials, with a high affinity for surface adsorption and slow exchange, and a secondary dynamic side of the short “soft” shift time of proteins and reversibly bound to the active compound (enzyme, drug molecule). A consequence of the processes of creating the “corona protein” will be a unique surface characteristic of the nanosystem and protein that determines the binding affinity to the plane of adsorption with interactions of the type: protein–protein, nanomaterial–protein, protein–water, water–nanomaterial, based on Van der Waals forces, hydrogen bonds, electrostatic, hydrophobic and π–π interactions. The first three forces mentioned above have frequently been observed in the creation and stabilization of protein complexes with carbon nanomaterials. Protein interactions with polymeric nanomaterial surfaces depend on size, composition (amino acid sequence, conformational flexibility of surface charge), isoelectric point, solution concentration, and structural stability [25]. Proteins with low internal stability are readily adsorbed on surfaces of all types, even though electrostatic repulsions exist; however, intrinsically stable proteins undergo substantial adsorption on hydrophilic surfaces only in the presence of electrostatic interactions. In general, the stronger the bond with the surface of nanomaterial large molecules, the broader the contact surface and flexible structure, with high shape complementarity of the adsorption surface [26,27,28].
When denatured, the majority of proteins lose their biological function. Enzymes fail in their role because the substrates no longer bind to the active site, and the amino acid residues involved in stabilizing the transition states of the substrates are no longer positioned properly. The denaturation process and the associated loss of activity can be measured using techniques such as double polarization interferometry, circular dichroism (CD), quartz crystal microbalance with dissipative monitoring (QCM-D), and multi-parametric surface plasmon resonance (MP-SPR). Protein denaturation involves the quaternary, tertiary, and secondary structure. In the quaternary structure, the denaturated proteins are dissociated, and the particular arrangement of the protein subunits is disrupted. If the tertiary structure is involved, the covalent interactions between amino acid side chains are disrupted, and dipole-dipole interactions between polar amino acids and Van der Waals interactions between nonpolar amino acid chains are also broken. In the secondary structure, denaturated proteins lose all repeating periodic patterns, such as alpha-helix and beta structure, and a random coil configuration is adopted. Protein strings resulted (see Supplementary Materials-S8) areas discussed above coils with primary and secondary structure. However, due to the methodology used in the process of string formation, the results were not random. When applied, the methods describe above similar outcomes were retrieved.
Cartesian coordinates represented for all antitrypsin (1ATU) strings show a particular pattern concerning dendrimer monolayer interaction. Thus Cartesian coordinates of 3D protein structure with tertiary and quaternary architecture have a distinct pattern: x, y, and z coordinates have most the same arrangement roughly parallel to one another. One exception is noticed, however: x intersects z coordinate in one point. Coordinates domain in the native protein varies with approximately 60 Å as the upper border and approximately −130 Å as a lower border.
On 1ATU computationally denaturated in the absence of dendrimer monolayer, various intersections between y and z coordinates are observed. Also, the coordinates parallelism is mold. However, x and y coordinates do not intersect. Coordinates domain is expanded between approximately 110 Å to approximately −190 Å.
Coordinates of ZAC dendrimers (see Supplementary Material-S9 for 2D structure) present roughly the same pattern with x,y,z interposition at the end of the protein string atoms. The domain of coordinates is preserved through the entire series between approximately 200 Å and approximately −60 Å.
Protein strings resulting from PAMAM interaction coordinates show a pattern in resemblance with ZAC dendrimers with constant domains observed through the entire series. The limited intersection of y with x and z coordinates are observed in strings PAMAM G1 and Pamam G2. In PAMAM G1.5 string coordinates lay intersection through the entire string but has the same domain in absolute value (although inverted) between approximately 130 Å and approximately −60 Å.
When represented as manifolds, as shown in results, the strings have topological surfaces distinct with respect to each series. In respect to the plane surface observed in the topological resulting manifold of 1ATU 3D crystallographic surface, PAMAM strings G3, G2, and respectively, G1.5 have a plane-symmetric surface.
The potential energy generated manifold shows similar behavior. When generated a manifold by integrating the functions used to obtain the energy manifold, it is observed that PAMAM G1.5 has the same energy profile as 1ATU3D string and 1ATU free string (Figure 14).
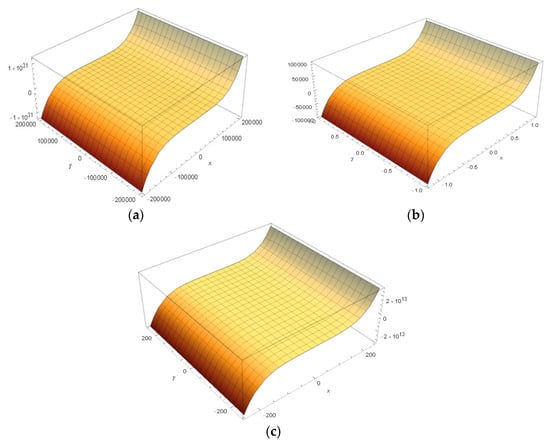
Figure 14.
Energy surfaces obtained by the integration of manifold coordinates equations. (a) PAMAM G1.5; (b) 1ATU3D; (c) 1ATU free string.
Solvation energy computed for all strings shows that the PAMAM G0 layer has the minimum effect on solvation energy compare to all other dendrimers.
In detail, angle bend energy has the biggest value in the PAMAM G0 layer, followed by the free protein (1ATU3D) and with obviously smaller values in the rest of the series. The results demonstrate that in comparison with the rest of the dendrimer monolayers, PAMAM G0 exercises fewer changes in conformation regarding the binding in any two consecutive atoms.
Electrostatic energy has negative values in PAMAM G0 derives string and the free protein (1ATU3D) and positive close to zero values in the rest of the string series. In this case, charge distribution changes when the protein (1ATU) interacts with the dendrimers monolayer. In terms of energy, the PAMAM G0 shows less disruptive effects in comparison with the other dendrimer layers.
Bond stretch energy has positive values all over the series, with the biggest value observed in the case of PAMAM G0. Its variation shows that during the interaction with the dendrimer monolayer, the resulting string has a significantly smaller tension that has the protein in its tertiary and quaternary structure. Namely, when a protein denatures, the bond stretch energy decreases significantly, while the protein is “free” from its tertiary and quaternary restraints.
Torsion energy-related closely to bond stretch energy has the same pattern with significant values in the case of the PAMAM G0 layer and the free protein (1ATU 3D) and values close to zero for the rest of the series.
Van der Waals energy demonstrates a massive increase. Van der Waals forces include attraction and repulsions between atoms, molecules, and surfaces, as well as other intermolecular forces. They contrast from covalent and ionic bonding in that they are produced by correlations in the fluctuating polarizations of neighboring particles. This is the result of a transient shift in electron density.
Even if the computed independent PAMAM G0 dendrimer monolayer seems to have a less disruptive effect on alpha 1 antitrypsin (1ATU) when put together, the results show that the PAMA G1.5 dendrimer monolayer is less disruptive in its interactions with 1ATU.
5. Conclusions
Polymer layers have a quantifiable effect on proteins and favor different reactions. This effect is demonstrated by the variation of topological coordinates. The variation of topological properties determines variations in biochemical properties that favor different types of reactions. In the case of 1ATU, the PAMAM G1.5 has shown the lowest level of interaction with the protein and lowest toxic effects. This property is reflected in the analysis of the protein sequence-cluster counting and biochemical analysis of G1.5, which have shown no potential binding and biochemical reactions to proteins. The Zeta potential suggested a less stable colloidal solution of the 1ATU protein.
The protein structure is influenced by the immobilization environment. The technique used is appropriate in calculating protein interactions with different media (carbon nanostructures, dendrimers).
Protein interaction with a surface is crucial in protein function. This interaction is crucial in enzyme surface interaction, nanostructure enzyme decoration, and in viral protein corona symmetry, respectively.
Furthermore, by creating small changes in the structure of the protein (receptor), the ligand bioactivity space can be accessed. When the whole protein is denatured, i.e., reduced to its primary and secondary structure, the bioactivity space cannot be approached. Thus, the bioactivity space can be accessed only by a structure with tertiary and quaternary architecture.
Supplementary Materials
The following are available online at https://www.mdpi.com/2073-8994/12/4/641/s1, S1-Cartesian coordinates, S2-Protein string pictures, S3-Aggregation scores, S4-Topological descriptors, S5-Zeta potential, S6-ZAC structure, S7-Energy manifolds, S8-String surface topology, S9-Protein strings.
Author Contributions
C.N.L., M.E.F. established the conceptual framework, produced the results, discussion and conclusions, and assembled the paper; I.P.G., G.O., and M.V.P. performed the literature screening of concerned compounds and methods and contributed the references and discussions from the results; all authors refined and agreed on the final manuscript. All authors have read and agree to the published version of the manuscript.
Funding
This work was financially supported by the GEMNS project granted in the European Union’s Seventh Framework Programme, under the framework of the ERA-NET EuroNanoMed II (European Innovative Research and Technological Development Projects in Nanomedicine). Mihai V. Putz acknowledges his contribution to this work within the Nucleus Programme under the project PN-19-22-01-02 and of its 2020 renewing fund by the Romanian National Authority for Scientific Research and Innovation (ANCSI).
Conflicts of Interest
The authors declare no conflicts of interest.
References
- Aggarwal, P.; Hall, J.; McLeland, C.; Dobrovolskaia, M.; Mc Neil, S. Nanoparticle interaction with plasma proteins as it relates to particle biodistribution, biocompatibility, and therapeutic efficacy. Adv. Drug Deliv. Rev. 2009, 61, 428–437. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, V.H.; Lee, B.J. Protein corona: A new approach for nanomedicine design. Int. J. Nanomed. 2017, 12, 3137–3151. [Google Scholar] [CrossRef] [PubMed]
- Lungu, C.N.; Bratanovici, B.I.; Mirabela, G.M.; Antoci, V.; Mangalagiu, I.I. Hybrid imidazole-pyridine derivatives: An approach to novel anticancer DNA intercalators. Curr. Med. Chem. 2020, 27, 154–169. [Google Scholar] [CrossRef] [PubMed]
- Kramar, R.; Shende, V.; Motl, N.; Pace, N.; Scholtz, M. Toward a molecular understanding of protein solubility: Increased negative surface change correlates with increased solubility. Biophys. J. 2012, 102, 1907–1915. [Google Scholar] [CrossRef]
- Wand, A.J.; Englander, S.W. Protein complexes studied by NMR spectroscopy. Curr. Opin. Biotechnol. 1996, 7, 403–408. [Google Scholar] [CrossRef]
- Robinson, P. Enzymes: Principles and biotechnological applications. Essays Biochem. 2015, 59, 1–41. [Google Scholar] [CrossRef]
- An, F.F.; Zhang, X.H. Strategies for preparing albumin-based nanoparticles for functional bioimaging and drug delivery. Theranostics 2017, 7, 3667–3689. [Google Scholar] [CrossRef]
- Sjöberg, B.; Foley, S.; Staicu, A.; Pascu, A.; Pascu, M.; Enescu, M. Protein reactivity with singlet oxygen: Influence of the solvent exposure of the reactive amino acid residues. J. Photochem. Photobiol. B 2016, 159, 106–110. [Google Scholar] [CrossRef]
- Uskoković, V.; Odsinada, R.; Djordjevic, S.; Habelitz, S. Dynamic light scattering of colloidal mixtures of amelogenin and hydroxyapatite in calcium and phosphate-rich iron milieus. Arch. Oral Biol. 2011, 56, 521–532. [Google Scholar] [CrossRef]
- Xu, X.; Wu, J.; Liu, Y.; Saw, P.; Tao, W.; Yu, M.; Zope, H.; Si, M.; Victorious, A.; Rasmussen, J.; et al. Multifunctional envelope-type siRNA delivery nanoparticle platform for prostate cancer therapy. ACS Nano 2017, 11, 2618–2627. [Google Scholar] [CrossRef] [PubMed]
- Ozboyaci, M.; Kokh, D.B.; Corni, S.; Wade, R.C. Modeling and simulation of protein-surface interactions: Achievements and challenges. Q. Rev. Biophys. 2016, 49, e4. [Google Scholar] [CrossRef] [PubMed]
- Salomon-Ferrer, R.; Case, D.A.; Walker, R.C. An overview of the Amber biomolecular simulation package. WIREs Comput. Mol. Sci. 2013, 3, 198–210. [Google Scholar] [CrossRef]
- Lungu, C.N.; Diudea, M.V. Binding site and potency prediction of Teixobactin and other Lipid II ligands by statistical base scoring of conformational space maps. Curr. Comput. Aided Drug Des. 2018, 14, 29–34. [Google Scholar] [CrossRef] [PubMed]
- István, J.; Lajos, S.; Zoltán, S. Resolvability and monotone normality. Israel J. Math. 2008, 166, 1–16. [Google Scholar]
- Rawlings, C.J.L.; Taylor, W.R.; Nyakairu, J.; Fox, J.; Sternberg, M.J.E. Reasoning about protein topology using the logic programming language PROLOG. J. Mol. Graph. 1985, 3, 151–157. [Google Scholar] [CrossRef]
- Majumdar, S.; Basak, S.C.; Lungu, C.N.; Diudea, M.V.; Grunwald, G.D. Mathematical structural descriptors and mutagenicity assessment: A study with congeneric and diverse data sets. SAR QSAR Environ. Res. 2018, 29, 579–590. [Google Scholar] [CrossRef]
- Lyklema, J. Fundamentals of Interface and Colloid Science; Academic Press: Cambridge, MA, USA, 1995; Volume 2, ISBN 0-12-460529-X. [Google Scholar]
- Conchillo-Solé, O.; de Groot, N.S.; Avilés, F.X.; Vendrell, J.; Daura, X.; Ventura, S. AGGRESCAN: A server for the prediction and evaluation of “hot spots” of aggregation in polypeptides. BMC Bioinform. 2007, 27, 65. [Google Scholar] [CrossRef]
- Greenwood, J.R.; Calkins, D.; Sullivan, A.P.; Shelley, J.C. Towards the comprehensive, rapid, and accurate prediction of the favorable tautomeric states of drug-like molecules in aqueous solution. J. Comput. Aided Mol. Des. 2010, 24, 591–604. [Google Scholar] [CrossRef]
- O’Rourkr, K.; Gorman, S.; Boehr, D. Biophysical and computational methods to analyze amino acid interaction networks in proteins. Comput. Struct. Biotechnol. J. 2016, 14, 245–254. [Google Scholar]
- Lungu, C.N.; Diudea, M.V.; Putz, M.V. Ligand shaping in induced fit docking of Mra Y inhibitors. Polynomial discriminant and Laplacian operator as biologically activity descriptors. Int. J. Mol. Sci. 2017, 18, 1377. [Google Scholar] [CrossRef]
- Aguzzi, A.; O’Connor, T. Protein aggregation diseases: Pathogenicity and therapeutic perspectives. Nat. Rev. Drug Discov. 2010, 9, 237–248. [Google Scholar] [CrossRef] [PubMed]
- Stefani, M.; Dobson, C.M. Protein aggregation and aggregate toxicity: New insights into protein folding, misfolding diseases, and biological evolution. J. Mol. Med. 2003, 81, 678–699. [Google Scholar] [CrossRef] [PubMed]
- Roberts, C.J. Non-native protein aggregation kinetics. Biotechnol. Bioeng. 2007, 98, 927–938. [Google Scholar] [CrossRef] [PubMed]
- Diaz-Villamevo, J.F.; Diaz-Molina, R.; Garcia Gouzález, V. Protein folding and mechanisms of proteostasis. Int. J. Mol. Sci. 2015, 16, 17193–17230. [Google Scholar] [CrossRef]
- Jahn, T.; Radford, S. Folding versus aggregation: Popypeptide conformational and competing pathways. Arch. Biochem. Biophys. 2008, 469, 100–117. [Google Scholar] [CrossRef]
- Lungu, C.N.; Ersali, S.; Szefler, B.; Pirvan-Moldovan, A.; Basak, S.; Diudea, M.V. Dimensionality of big data sets explored by Cluj descriptors. Studia UBB Chem. 2017, 62, 197–204. [Google Scholar] [CrossRef]
- Szefler, B.; Czeleń, P.; Diudea, M.V. Understanding the action of indolizines as biologically active moieties: A molecular dynamics study. Curr. Comput. Aided Drug Des. 2017, 13, 22–29. [Google Scholar] [CrossRef]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).