Integrating Casein Complex SNPs Additive, Dominance and Epistatic Effects on Genetic Parameters and Breeding Values Estimation for Murciano-Granadina Goat Milk Yield and Components
Abstract
1. Introduction
2. Materials and Methods
2.1. Milk Yield Standardization and Composition Analysis
2.2. Animals
2.3. Genotyping
2.4. Single-Nucleotide Polymorphisms (SNPs) Additive and Dominance Genetic Effects Identification and Codification and Dimensionality Reduction Using Linkage Disequilibrium (LD) and Categorical Principal Component Analysis (CATPCA)
2.5. Study of OVERALs/Nonlinear Canonical Correlations (NLCC) to Identify and Encode Epistatic Effects
2.6. Preliminary Statistical Assumption Testing
2.7. Non-Genetic and Genetic Fixed Effect Statistical Analysis
2.8. Genetic Model Comparison, Phenotypic and Genetic Parameter Estimation
2.9. Non-Genetic Best Linear Unbiased Estimators (BLUE) for Fixed Effects and Covariates and Best Linear Unbiased Predictors/Breeding Value Prediction (BLUP, PBVs)
2.10. Predicted Breeding Values (PBV), Standard Error of Prediction (SEP), Accuracies (RTi), and Reliability (Rap) Comparison
2.11. Ethics Approval and Consent to Participate
3. Results
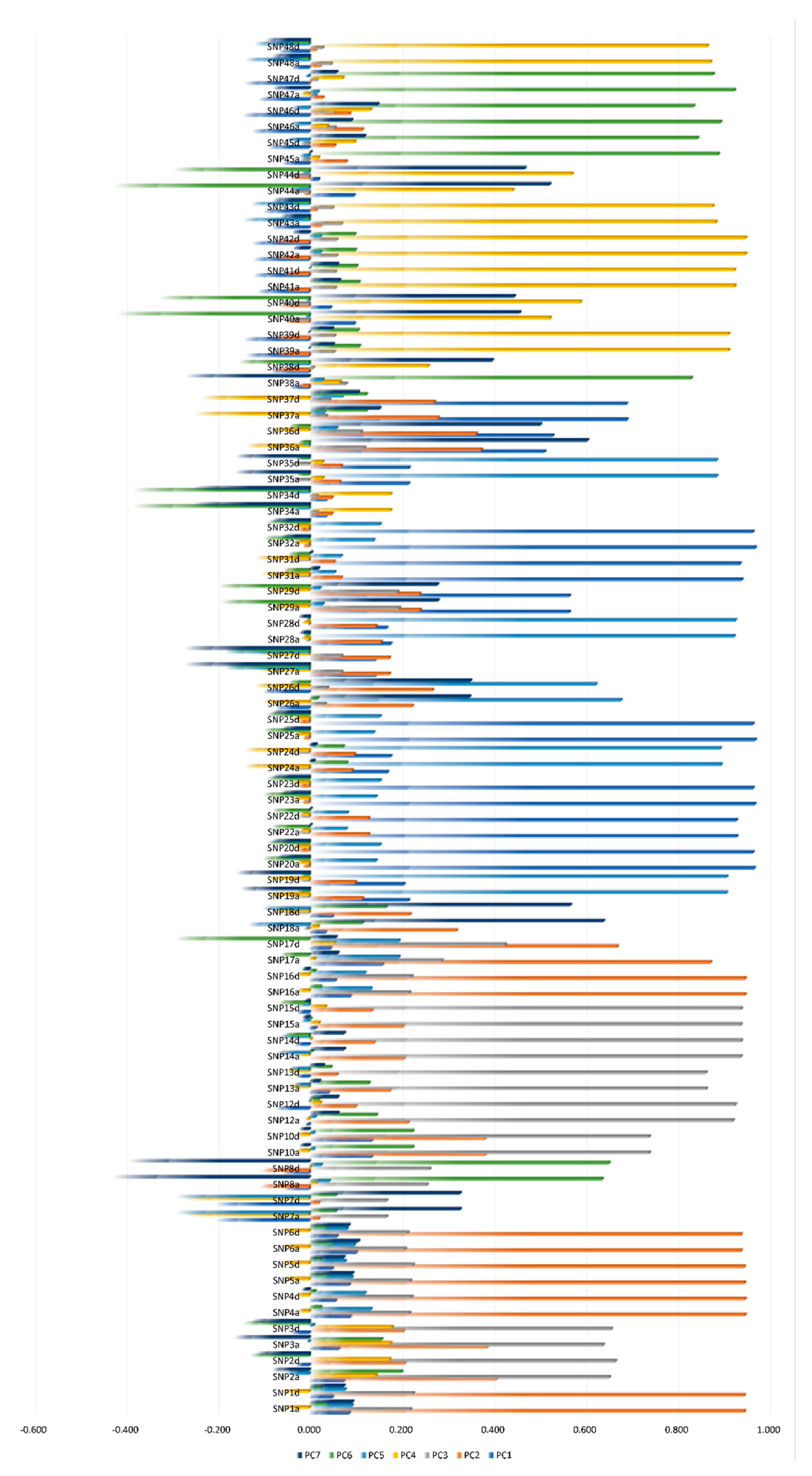
3.1. SNPs Dimensionality Reduction Using Linkage Disequilibrium and CATPCA
3.2. Study of OVERALs/Nonlinear Canonical Linear Correlations (NLCC) to Identify and Encode Epistatic Effects
3.3. Non-Genetic and Genetic Fixed Effect Analysis
3.4. Genetic Model Comparison, Phenotypic and Genetic Parameters Estimation
3.5. Predicted Breeding Values (PBV), Standard Error of Prediction (SEP), Accuracies (RTi), and Reliability (Rap) Comparison
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Delgado, J.V.; Landi, V.; Barba, C.J.; Fernández, J.; Gómez, M.M.; Camacho, M.E.; Martínez, M.A.; Navas, F.J.; León, J.M. Murciano-Granadina goat: A spanish local breed ready for the challenges of the twenty-first century. In Sustainable Goat Production in Adverse Environments: Volume II; Springer: New York, NY, USA, 2017; pp. 205–219. [Google Scholar]
- Hristov, J.P.; Teofanova, D.; Georgieva, A.; Radoslavov, G. Effect of genetic polymorphism of αS1-casein gene on qualitative and quantitative milk traits in native Bulgarian Rhodopean cattle breed. Genet. Mol. Res. 2018, 17. [Google Scholar] [CrossRef]
- Montaldo, H.; Valencia-Posadas, M.; Wiggans, G.; Shepard, L.; Torres-Vázquez, J. Genetic and environmental relationships between milk yield and kidding interval in dairy goats. J. Dairy Sci. 2010, 93, 370–372. [Google Scholar] [CrossRef] [PubMed]
- Andreazza, J.; Rorato, P.R.N.; El Faro, L.; Boligon, A.A.; Weber, T.; Kippert, C.J.; Lopes, J.S. Parâmetros genéticos e eficiência relativa de seleção para a produção de leite no dia do controle para vacas da raça Holandesa. Ciência Rural 2008, 38, 451–456. [Google Scholar] [CrossRef]
- Torres-Vázquez, J.; Valencia-Posadas, M.; Castillo-Juárez, H.; Montaldo, H. Genetic and phenotypic parameters of milk yield, milk composition and age at first kidding in Saanen goats from Mexico. Livest. Sci. 2009, 126, 147–153. [Google Scholar] [CrossRef]
- VanRaden, P.; Dematawewa, C.; Pearson, R.; Tooker, M. Productive life including all lactations and longer lactations with diminishing credits. J. Dairy Sci. 2006, 89, 3213–3220. [Google Scholar] [CrossRef]
- Pizarro Inostroza, M.; Landi, V.; Navas González, F.; León Jurado, J.; Martínez Martínez, A.; Fernández Álvarez, J.; Delgado Bermejo, J. Non-parametric association analysis of additive and dominance effects of casein complex SNPs on milk content and quality in Murciano-Granadina goats. J. Anim. Breed. Genet. 2019. [Google Scholar] [CrossRef] [PubMed]
- Buzanskas, M.E.; Grossi, D.A.; Ventura, R.V.; Schenkel, F.S.; Sargolzaei, M.; Meirelles, S.L.; Mokry, F.B.; Higa, R.H.; Mudadu, M.A.; da Silva, M.V.B. Genome-wide association for growth traits in Canchim beef cattle. PLoS ONE 2014, 9, e94802. [Google Scholar] [CrossRef]
- Jolliffe, I.T.; Cadima, J. Principal component analysis: A review and recent developments. Philos. Trans. R. Soc. A 2016, 374, 20150202. [Google Scholar] [CrossRef]
- Lopes, F.B.; da Silva, M.C.; Magnabosco, C.U.; Narciso, M.G.; Sainz, R.D. Selection indices and multivariate analysis show similar results in the evaluation of growth and carcass traits in beef cattle. PLoS ONE 2016, 11, e0147180. [Google Scholar]
- Lopes, F.B.; Magnabosco, C.U.; Mamede, M.M.; Da Silva, M.C.; Myiage, E.S.; Paulini, F.; Lôbo, R.B. Multivariate approach for young bull selection from a performance test using multiple traits of economic importance. Trop. Anim. Health Prod. 2013, 45, 1375–1381. [Google Scholar] [CrossRef]
- Pizarro, G.; Landi, V.; León Jurado, J.M.; Navas González, F.J.; Delgado Bermejo, J.V. Non-parametric analysis of the effects of αS1-casein genotype and parturition nongenetic factors on milk yield and composition in Murciano-Granadina goats. Ital. J. Anim. Sci. 2019, 18, 1021–1034. [Google Scholar] [CrossRef]
- Van Vleck, L.D. Selection Index and Introduction to Mixed Model Methods, 1st ed.; CRC Press: Boca Ratón, FL, USA, 1993. [Google Scholar]
- The MathWorks, Inc. MATLAB; release R2015a ed.; The MathWorks, Inc.: Natick, MA, USA, 2015. [Google Scholar]
- Miller, S.; Dykes, D.; Polesky, H. A simple salting out procedure for extracting DNA from human nucleated cells. Nucleic Acids Res. 1988, 16, 1215. [Google Scholar] [CrossRef] [PubMed]
- Hubbard, T.; Barker, D.; Birney, E.; Cameron, G.; Chen, Y.; Clark, L.; Cox, T.; Cuff, J.; Curwen, V.; Down, T. The Ensembl genome database project. Nucleic Acids Res. 2002, 30, 38–41. [Google Scholar] [CrossRef] [PubMed]
- Hao, K.; Li, C.; Rosenow, C.; Wong, W.H. Detect and adjust for population stratification in population-based association study using genomic control markers: An application of Affymetrix Genechip® Human Mapping 10K array. Eur. J. Hum. Genet. 2004, 12, 1001–1006. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Purcell, S.; Neale, B.; Todd-Brown, K.; Thomas, L.; Ferreira, M.A.; Bender, D.; Maller, J.; Sklar, P.; De Bakker, P.I.; Daly, M.J. PLINK: A tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 2007, 81, 559–575. [Google Scholar] [CrossRef] [PubMed]
- Price, A.L.; Patterson, N.J.; Plenge, R.M.; Weinblatt, M.E.; Shadick, N.A.; Reich, D. Principal components analysis corrects for stratification in genome-wide association studies. Nat. Genet. 2006, 38, 904–909. [Google Scholar] [CrossRef] [PubMed]
- Dagnachew, B.S.; Thaller, G.; Lien, S.; Ådnøy, T. Casein SNP in Norwegian goats: Additive and dominance effects on milk composition and quality. Genet. Sel. Evol. 2011, 43, 31. [Google Scholar] [CrossRef]
- Patterson, N.; Price, A.L.; Reich, D. Population structure and eigenanalysis. PLoS Genet. 2006, 2, e190. [Google Scholar] [CrossRef]
- Novembre, J.; Johnson, T.; Bryc, K.; Kutalik, Z.; Boyko, A.R.; Auton, A.; Indap, A.; King, K.S.; Bergmann, S.; Nelson, M.R. Genes mirror geography within Europe. Nature 2008, 456, 98–101. [Google Scholar] [CrossRef]
- Horne, B.D.; Camp, N.J. Principal component analysis for selection of optimal SNP—Sets that capture intragenic genetic variation. Genet. Epidemiol. 2004, 26, 11–21. [Google Scholar] [CrossRef]
- Zhang, F.; Wagener, D. An approach to incorporate linkage disequilibrium structure into genomic association analysis. J. Genet. Genom. 2008, 35, 381–385. [Google Scholar] [CrossRef][Green Version]
- Song, Y.; Schreier, P.J.; Roseveare, N.J. Determining the number of correlated signals between two data sets using PCA-CCA when sample support is extremely small. In Proceedings of the 2015 IEEE International Conference on Acoustics, Speech and Signal Processing (ICASSP), South Brisbane, Queensland, Australia, 19–24 April 2015; pp. 3452–3456. [Google Scholar]
- Hsieh, W.W. Nonlinear multivariate and time series analysis by neural network methods. Rev. Geophys. 2004, 42. [Google Scholar] [CrossRef]
- Paré, G.; Cook, N.R.; Ridker, P.M.; Chasman, D.I. On the use of variance per genotype as a tool to identify quantitative trait interaction effects: A report from the Women’s Genome Health Study. PLoS Genet. 2010, 6, e1000981. [Google Scholar] [CrossRef] [PubMed]
- Rönnegård, L.; Valdar, W. Detecting major genetic loci controlling phenotypic variability in experimental crosses. Genetics 2011, 188, 435–447. [Google Scholar] [CrossRef] [PubMed]
- Bárcenas, P.; Elortondo, F.P.; Albisu, M. Comparison of free choice profiling, direct similarity measurements and hedonic data for ewes’ milk cheeses sensory evaluation. Int. Dairy J. 2003, 13, 67–77. [Google Scholar] [CrossRef]
- Greenacre, M.; Hastie, T. The geometric interpretation of correspondence analysis. J. Am. Stat. Assoc. 1987, 82, 437–447. [Google Scholar] [CrossRef]
- Young, G.; Dey, S.; Rogers, A.; Exton, D. Correction: Cost and time-effective method for multi-scale measures of rugosity, fractal dimension, and vector dispersion from coral reef 3D models. PLoS ONE 2018, 13, e0201847. [Google Scholar] [CrossRef]
- Pizarro, M.I.; Landi, V.; Navas, F.G.; León, J.J.; Martínez, A.M.; Fernández, J.Á.; Delgado, J.B. Does the Acknowledgement of αS1-casein genotype affect the estimation of genetic parameters and prediction of breeding values for milk yield and composition quality-related traits in Murciano-Granadina? Animals 2019, 9, 679. [Google Scholar] [CrossRef] [PubMed]
- Bidanel, J. Benefits and limits of increasingly sophisticated models for genetic evaluation: The example of pig breeding. In Proceedings of the 6th World Congress on Genetics Applied to Livestock Production, Armidale, Australia, 11–16 January 1998; pp. 11–16. [Google Scholar]
- Andonov, S.; Ødegård, J.; Svendsen, M.; Ådnøy, T.; Vegara, M.; Klemetsdal, G. Comparison of random regression and repeatability models to predict breeding values from test-day records of Norwegian goats. J. Dairy Sci. 2013, 96, 1834–1843. [Google Scholar] [CrossRef] [PubMed]
- Meyer, K. Restricted maximum likelihood to estimate variance components for animal models with several random effects using a derivative-free algorithm. Genet. Sel. Evol. 1989, 21, 317. [Google Scholar] [CrossRef]
- Brown, J.D. Effect size and eta squared. JALT Test. Eval. SIG News 2008, 12, 38–43. [Google Scholar]
- Boldman, K.G.; Kriese, L.A.; Vleck, L.D.; Kachman, S.D. A Manual for Use of MTDFREML. A Set of Programs to Obtain Estimates of Variances and Covariances, 1st ed.; US Department of Agriculture, Agricultural Research Service: Washington, DC, USA, 1995; p. 114.
- Dagnachew, B.S.; Ådnøy, T. Additive and dominance effects of casein haplotypes on milk composition and quality in Norwegian dairy goats. Small Rumin. Res. 2014, 122, 59–69. [Google Scholar] [CrossRef]
- Murphy, K.R.; Myors, B.; Wolach, A. Statistical Power Analysis: A Simple and General Model for Traditional and Modern Hypothesis Tests; Routledge: London, UK, 2014. [Google Scholar]
- Schmitz, S.; Cherny, S.S.; Fulker, D.W. Increase in power through multivariate analyses. Behav. Genet. 1998, 28, 357–363. [Google Scholar] [CrossRef] [PubMed]
- Agudelo-Gómez, D.; Pineda-Sierra, S.; Cerón-Muñoz, M.F. Genetic evaluation of dual-purpose buffaloes (Bubalus bubalis) in Colombia using principal component analysis. PLoS ONE 2015, 10, e0132811. [Google Scholar]
- Domínguez-Viveros, J.; Rodríguez-Almeida, F.; Burrola-Barraza, M.; Callejas-Juárez, N.; Ortega-Gutiérrez, J. Análisis con componentes principales y estimación de parámetros genéticos para medidas zoométricas en caballo pura raza española de México. Arch. Zootec. 2019, 68, 434–439. [Google Scholar] [CrossRef]
- Bignardi, A.; Santana, M., Jr.; Eler, J.; Ferraz, J. Models for genetic evaluation of growth of Brazilian Bonsmara cattle. Livest. Sci. 2014, 162, 50–58. [Google Scholar] [CrossRef]
- Bignardi, A.; El Faro, L.; Rosa, G.; Cardoso, V.; Machado, P.F.; Albuquerque, L.G.d. Principal components and factor analytic models for test-day milk yield in Brazilian Holstein cattle. J. Dairy Sci. 2012, 95, 2157–2164. [Google Scholar] [CrossRef] [PubMed]
- Val, J.; Ferraudo, A.; Bezerra, L.A.F.; Corrado, M.P.; Lôbo, R.B.; Freitas, M.; Paneto, J. Alternativas para seleção de touros da raça Nelore considerando características múltiplas de importância econômica. Arq. Bras. Med. Vet. Zootec. 2008, 60, 705–712. [Google Scholar] [CrossRef]
- Buzanskas, M.; Savegnago, R.; Grossi, D.; Venturini, G.; Queiroz, S.; Silva, L.; Júnior, R.T.; Munari, D.; Alencar, M. Genetic parameter estimates and principal component analysis of breeding values of reproduction and growth traits in female Canchim cattle. Reprod. Fertil. Dev. 2013, 25, 775–781. [Google Scholar] [CrossRef]
- Mucha, S.; Mrode, R.; Coffey, M.; Conington, J. Estimation of genetic parameters for milk yield across lactations in mixed-breed dairy goats. J. Dairy Sci. 2014, 97, 2455–2461. [Google Scholar] [CrossRef]
- Bagnicka, E.; Lukaszewicz, M.; Ådnøy, T. Genetic parameters of somatic cell score and lactose content in goat s milk. J. Anim. Feed Sci. 2016, 25, 210–215. [Google Scholar] [CrossRef]
- Rupp, R.; Clément, V.; Piacere, A.; Robert-Granié, C.; Manfredi, E. Genetic parameters for milk somatic cell score and relationship with production and udder type traits in dairy Alpine and Saanen primiparous goats. J. Dairy Sci. 2011, 94, 3629–3634. [Google Scholar] [CrossRef] [PubMed]
- Benradi, Z. Genetic Analysis of Milk Composition, Cheese Yield and Reological Traits and the Malagueña and Murciano-Granadina Goat Breeds. Master’s Thesis, International Centre for Advanced Mediterranean Agronomic Studies, Zaragoza, Spain, 2007. [Google Scholar]
- Morris, C.; Wheeler, M.; Foote, B. Relationships between live weight and herd-test traits in a Saanen goat herd in New Zealand. N. Z. J. Agric. Res. 2011, 54, 315–320. [Google Scholar] [CrossRef]
- Luengo, C.; Sánchez, A.; Corrales, J.C.; Fernández, C.; Contreras, A. Influence of intramammary infection and non-infection factors on somatic cell counts in dairy goats. J. Dairy Res. 2004, 71, 169–174. [Google Scholar] [CrossRef] [PubMed]
- Moroni, P.; Pisoni, G.; Ruffo, G.; Boettcher, P. Risk factors for intramammary infections and relationship with somatic-cell counts in Italian dairy goats. Prev. Vet. Med. 2005, 69, 163–173. [Google Scholar] [CrossRef] [PubMed]
- Koop, G.; Dik, N.; Nielen, M.; Lipman, L. Repeatability of differential goat bulk milk culture and associations with somatic cell count, total bacterial count, and standard plate count. J. Dairy Sci. 2010, 93, 2569–2573. [Google Scholar] [CrossRef] [PubMed]
- Castañeda-Bustos, V.; Montaldo, H.; Torres-Hernández, G.; Pérez-Elizalde, S.; Valencia-Posadas, M.; Hernández-Mendo, O.; Shepard, L. Estimation of genetic parameters for productive life, reproduction, and milk-production traits in US dairy goats. J. Dairy Sci. 2014, 97, 2462–2473. [Google Scholar] [CrossRef]
- Mohammed, B.; Aynalem, H.; Hailu, D.; Alemu, T. Estimates of genetic and phenotypic parameters for milk traits in Arsi-Bale goat in Ethiopia. Livest. Res. Rural Dev. 2012, 24, 1–20. [Google Scholar]
- Škorput, D.; Špehar, M.; Luković, Z. Connectedness between contemporary groups in Black Slavonian pig. Livest. Sci. 2018, 216, 6–8. [Google Scholar] [CrossRef]
- Gervais, O.; Pong-Wong, R.; Navarro, P.; Haley, C.S.; Nagamine, Y. Antagonistic genetic correlations for milking traits within the genome of dairy cattle. PLoS ONE 2017, 12, e0175105. [Google Scholar] [CrossRef]
- McDougall, S.; Lopez-Villalobos, N.; Prosser, C. Relationship between estimated breeding value for somatic cell count and prevalence of intramammary infection in dairy goats. N. Z. J. Agric. Res. 2011, 59, 300–304. [Google Scholar] [CrossRef] [PubMed]
- Mucha, S.; Mrode, R.; MacLaren-Lee, I.; Coffey, M.; Conington, J. Estimation of genomic breeding values for milk yield in UK dairy goats. J. Dairy Sci. 2015, 98, 8201–8208. [Google Scholar] [CrossRef] [PubMed]
- Sant’Ana, A.; Bezerril, F.; Madruga, M.; Batista, A.; Magnani, M.; Souza, E.; Queiroga, R. Nutritional and sensory characteristics of Minas fresh cheese made with goat milk, cow milk, or a mixture of both. J. Dairy Sci. 2013, 96, 7442–7453. [Google Scholar] [CrossRef] [PubMed]
- Valencia, M.; Dobler, J.; Montaldo, H. Genetic and phenotypic parameters for lactation traits in a flock of Saanen goats in Mexico. Small Rumin. Res. 2007, 68, 318–322. [Google Scholar] [CrossRef]
- Arnal, M.; Robert-Granié, C.; Larroque, H. Diversity of dairy goat lactation curves in France. J. Dairy Sci. 2018, 101, 11040–11051. [Google Scholar] [CrossRef] [PubMed]
- Sneddon, N.; Lopez-Villalobos, N.; Davis, S.; Hickson, R.; Shalloo, L. Genetic parameters for milk components including lactose from test day records in the New Zealand dairy herd. N. Z. J. Agric. Res. 2015, 58, 97–107. [Google Scholar] [CrossRef]
- Haile-Mariam, M.; Pryce, J. Genetic parameters for lactose and its correlation with other milk production traits and fitness traits in pasture-based production systems. J. Dairy Sci. 2017, 100, 3754–3766. [Google Scholar] [CrossRef]
- Verdier-Metz, I.; Coulon, J.-B.; Pradel, P. Relationship between milk fat and protein contents and cheese yield. Anim. Res. 2001, 50, 365–371. [Google Scholar] [CrossRef]
- Alais, C. Ciencia de la Leche: Principios de Técnica Lechera; Compañía Editorial Continental, Ed.; Reverté, S.A.: Barcelona, España, 1998. [Google Scholar]
- Cheverud, J.M.; Routman, E.J. Epistasis and its contribution to genetic variance components. Genetics 1995, 139, 1455–1461. [Google Scholar]
- Carter, A.J.; Hermisson, J.; Hansen, T.F. The role of epistatic gene interactions in the response to selection and the evolution of evolvability. Theor. Popul. Biol. 2005, 68, 179–196. [Google Scholar] [CrossRef]
- Phillips, P.C. Epistasis—The essential role of gene interactions in the structure and evolution of genetic systems. Nat. Rev. Genet. 2008, 9, 855–867. [Google Scholar] [CrossRef] [PubMed]
- Clark, S.A.; Hickey, J.M.; Daetwyler, H.D.; van der Werf, J.H. The importance of information on relatives for the prediction of genomic breeding values and the implications for the makeup of reference data sets in livestock breeding schemes. Genet. Sel. Evol. 2012, 44, 4. [Google Scholar] [CrossRef] [PubMed]
| Model/Genetic Effects as a Fixed Effect | Trait (Kg) | h2±SE | ||||
|---|---|---|---|---|---|---|
| Including genetic effects as a fixed effect. | Milk yield | 0.75450 | 1.63632 | 0.140011 | 0.74180 | 0.46±0.05 |
| Fat | 0.37663 | 1.72151 | 0.204217 | 1.14066 | 0.22±0.01 | |
| Protein | 0.06216 | 0.24909 | 0.0276599 | 0.15927 | 0.25±0.01 | |
| Solids | 0.53164 | 2.58194 | 0.285996 | 1.76430 | 0.21±0.05 | |
| Lactose | 0.03361 | 0.11198 | 0.0213750 | 0.05699 | 0.30±0.01 | |
| Somatic cells | 1,450,503.8674 | 3,373,095.25 | 36,251.9 | 1,886,339.4833 | 0.43±0.07 | |
| Excluding genetic effects as a fixed effect. | Milk yield | 0.34930 | 1.65896 | 0.172186 | 1.13747 | 0.21±0.01 |
| Fat | 0.42176 | 1.72965 | 0.177331 | 1.13056 | 0.24±0.01 | |
| Protein | 0.06541 | 0.26935 | 0.0286661 | 0.17527 | 0.24±0.02 | |
| Solids | 0.67591 | 2.80513 | 0.291377 | 1.83785 | 0.24±0.01 | |
| Lactose | 0.02505 | 0.11168 | 0.0114023 | 0.07523 | 0.22±0.01 | |
| Somatic cells | 483,509.3208 | 2,368,170.87 | 244,535. | 1,640,126.5503 | 0.20±0.05 |
| Model/Genotype as a Fixed Effect | Trait | Milk Yield | Fat | Protein | Solids | Lactose | Somatic Cells Count |
|---|---|---|---|---|---|---|---|
| Including genotype as a fixed effect | Milk yield | 0.46 | −0.41 | −0.48 | −0.46 | 0.12 | −0.24 |
| Fat | −0.42 | 0.22 | 0.56 | 0.96 | −0.10 | 0.16 | |
| Protein | 0.09 | 0.43 | 0.25 | 0.71 | −0.29 | 0.29 | |
| Dry mater | 0.08 | 0.85 | 0.60 | 0.21 | −0.03 | 0.15 | |
| Lactose | −0.07 | 0.05 | −0.20 | 0.03 | 0.30 | −0.38 | |
| Cells somatic | −0.27 | 0.18 | 0.18 | 0.16 | −0.22 | 0.43 | |
| Excluding genotype as a fixed effect | Milk yield | 0.21 | −0.33 | −0.41 | −0.38 | 0.09 | −0.25 |
| Fat | −0.29 | 0.24 | 0.46 | 0.95 | −0.10 | 0.16 | |
| Protein | −0.33 | 0.39 | 0.24 | 0.66 | −0.28 | 0.28 | |
| Dry mater | −0.34 | 0.73 | 0.51 | 0.24 | 0.03 | 0.14 | |
| Lactose | 0.08 | −0.08 | −0.16 | 0.02 | 0.22 | −0.36 | |
| Cells somatic | −0.18 | 0.13 | 0.24 | 0.12 | −0.32 | 0.20 |
| Model | Including Additive, Dominance and Epistatic Genetic Effects | Excluding Additive, Dominance and Epistatic Genetic Effects | |||||||
|---|---|---|---|---|---|---|---|---|---|
| Descriptive | Min | Max | Mean | SD | Min | Max | Mean | SD | |
| Parameters | |||||||||
| Milk yield (Kg) | SEP | 0.59 | 0.69 | 0.59 | 0.01 | 0.87 | 1.02 | 0.87 | 0.01 |
| Rti | 0.00 | 0.66 | 0.01 | 0.03 | 0.00 | 0.02 | 0.00 | 0.00 | |
| Fat (Kg) | SEP | 0.00 | 0.76 | 0.65 | 0.01 | 0.61 | 0.72 | 0.61 | 0.01 |
| Rti | 0.00 | 0.96 | 0.01 | 0.05 | 0.00 | 0.02 | 0.00 | 0.00 | |
| Protein (Kg) | SEP | 0.17 | 0.30 | 0.26 | 0.00 | 0.25 | 0.29 | 0.25 | 0.00 |
| Rti | 0.00 | 0.76 | 0.01 | 0.04 | 0.00 | 0.02 | 0.00 | 0.00 | |
| Solids (Kg) | SEP | 0.00 | 0.96 | 0.82 | 0.02 | 0.73 | 0.85 | 0.73 | 0.01 |
| Rti | 0.00 | 0.96 | 0.02 | 0.06 | 0.00 | 0.02 | 0.00 | 0.00 | |
| Lactose (Kg) | SEP | 0.13 | 0.19 | 0.16 | 0.00 | 0.18 | 0.21 | 0.18 | 0.00 |
| Rti | 0.00 | 0.61 | 0.01 | 0.03 | 0.00 | 0.02 | 0.00 | 0.00 | |
| Somatic cells count (cs/mL) | SEP | 554.46 | 815.37 | 696.41 | 8.81 | 1204.37 | 1412.25 | 1206.80 | 14.82 |
| Rti | 0.00 | 0.60 | 0.01 | 0.03 | 0.00 | 0.02 | 0.00 | 0.00 | |
| PBV Parameters | Pearson Product Moment Correlation |
|---|---|
| SEP Milk yield (Kg) | 0.994 ** |
| RTi Milk yield (Kg) | 0.103 ** |
| SEP Fat (Kg) | 0.674 ** |
| RTi Fat (Kg) | 0.097 ** |
| SEP Protein (Kg) | 0.671 ** |
| RTi Protein (Kg) | 0.099 ** |
| SEP Solids (Kg) | 0.022 ** |
| RTi Solids (Kg) | −0.009 |
| SEP Lactose (Kg) | 0.045 ** |
| RTi Lactose (Kg) | 0.012 * |
| SEP Somatic cells (cs/mL) | 0.036 ** |
| RTi Somatic cells count (cs/mL) | −0.010 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pizarro Inostroza, M.G.; Landi, V.; Navas González, F.J.; León Jurado, J.M.; Delgado Bermejo, J.V.; Fernández Álvarez, J.; Martínez Martínez, M.d.A. Integrating Casein Complex SNPs Additive, Dominance and Epistatic Effects on Genetic Parameters and Breeding Values Estimation for Murciano-Granadina Goat Milk Yield and Components. Genes 2020, 11, 309. https://doi.org/10.3390/genes11030309
Pizarro Inostroza MG, Landi V, Navas González FJ, León Jurado JM, Delgado Bermejo JV, Fernández Álvarez J, Martínez Martínez MdA. Integrating Casein Complex SNPs Additive, Dominance and Epistatic Effects on Genetic Parameters and Breeding Values Estimation for Murciano-Granadina Goat Milk Yield and Components. Genes. 2020; 11(3):309. https://doi.org/10.3390/genes11030309
Chicago/Turabian StylePizarro Inostroza, María Gabriela, Vincenzo Landi, Francisco Javier Navas González, Jose Manuel León Jurado, Juan Vicente Delgado Bermejo, Javier Fernández Álvarez, and María del Amparo Martínez Martínez. 2020. "Integrating Casein Complex SNPs Additive, Dominance and Epistatic Effects on Genetic Parameters and Breeding Values Estimation for Murciano-Granadina Goat Milk Yield and Components" Genes 11, no. 3: 309. https://doi.org/10.3390/genes11030309
APA StylePizarro Inostroza, M. G., Landi, V., Navas González, F. J., León Jurado, J. M., Delgado Bermejo, J. V., Fernández Álvarez, J., & Martínez Martínez, M. d. A. (2020). Integrating Casein Complex SNPs Additive, Dominance and Epistatic Effects on Genetic Parameters and Breeding Values Estimation for Murciano-Granadina Goat Milk Yield and Components. Genes, 11(3), 309. https://doi.org/10.3390/genes11030309






