Abstract
The simultaneous response of one transcriptional regulator to different effectors remains largely unexplored. Nevertheless, such interactions can substantially impact gene expression by rapidly integrating cellular signals and by expanding the range of transcriptional responses. In this study, similarities between paralogs were exploited to engineer novel responses in CatM, a regulator that controls benzoate degradation in Acinetobacter baylyi ADP1. One goal was to improve understanding of how its paralog, BenM, activates transcription in response to two compounds (cis,cis-muconate and benzoate) at levels significantly greater than with either alone. Despite the overlapping functions of BenM and CatM, which regulate many of the same ben and cat genes, CatM normally responds only to cis,cis-muconate. Using domain swapping and site-directed amino acid replacements, CatM variants were generated and assessed for the ability to activate transcription. To create a variant that responds synergistically to both effectors required alteration of both the effector-binding region and the DNA-binding domain. These studies help define the interconnected roles of protein domains and extend understanding of LysR-type proteins, the largest family of transcriptional regulators in bacteria. Additionally, renewed interest in the modular functionality of transcription factors stems from their potential use as biosensors.
1. Introduction
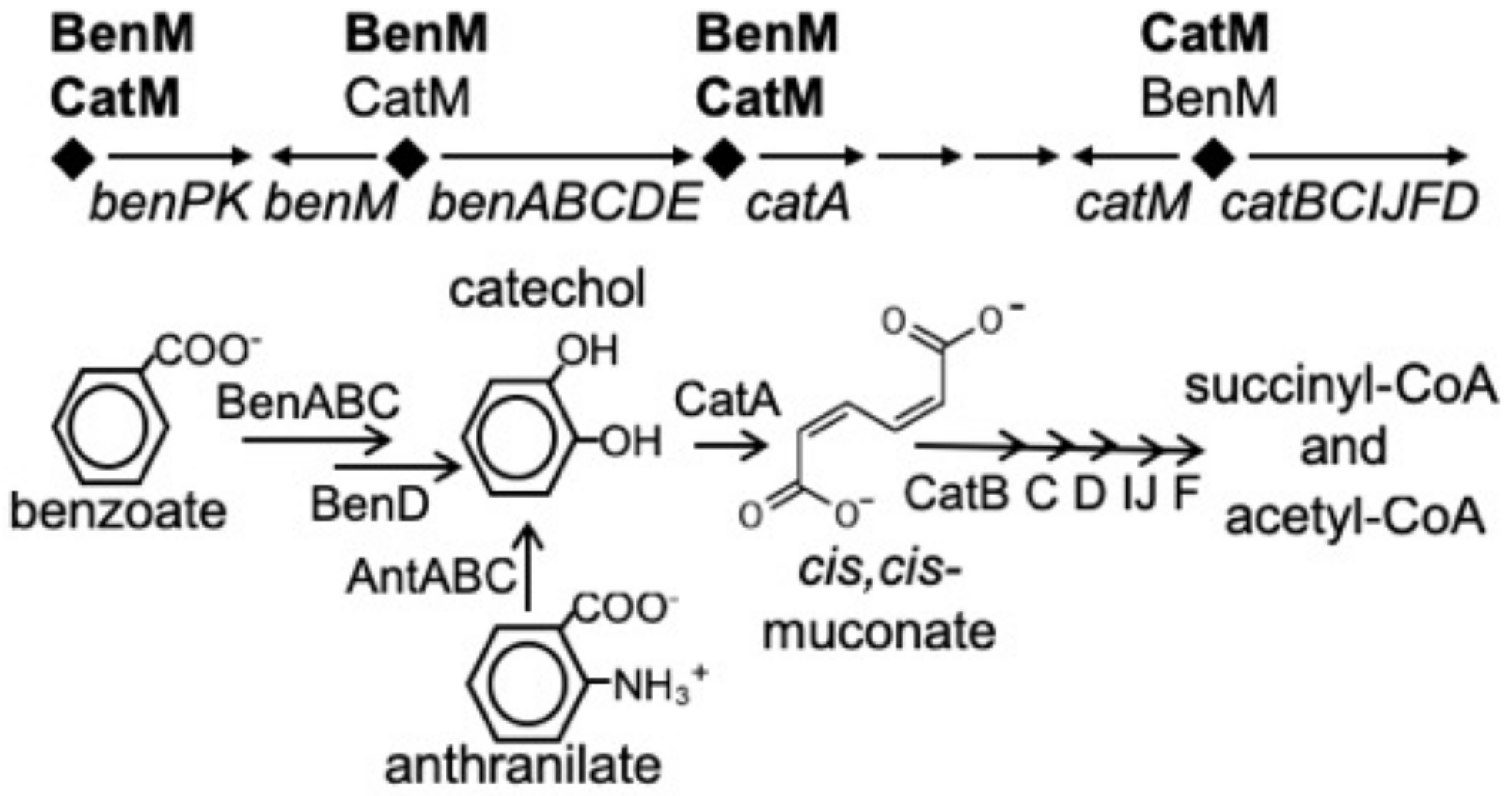
Two LysR-type transcriptional regulators (LTTRs) control benzoate degradation by a soil bacterium, Acinetobacter baylyi ADP1 [1]. These paralogs, BenM and CatM, have overlapping but distinct functions (Figure 1). BenM controls the initial steps in benzoate consumption by activating transcription of the benABCDE operon in response to benzoate and one of its metabolites, cis,cis-muconate, hereafter designated muconate [2,3]. CatM, which responds solely to muconate, activates low-level expression from this promoter, PbenA [4]. We sought to create a benzoate-responsive CatM that mimics an unusual characteristic of BenM, namely the ability to activate transcription synergistically in response to two effectors. These studies should improve understanding of the molecular basis of this type of transcriptional activation and, in general, facilitate the engineering of LTTRs to respond to novel effectors when designed for varied biotechnology applications.
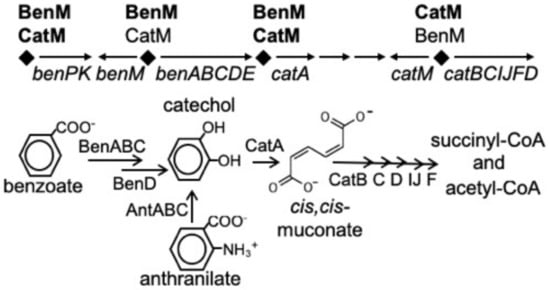
Figure 1.
BenM and CatM regulate the ben and cat genes from four promoter regions (diamonds). The benABCDE operon is primarily regulated by BenM from PbenA, and the catBCIJFD operon is primarily regulated by CatM from PcatB. The encoded enzymes are used for the degradation of benzoate.
BenM and CatM have similar N-terminal DNA-binding domains (DBDs, Figure 2), and they bind to the same regions of PbenA [2,4]. Whereas LTTR-DNA interactions repress basal transcription, conformational changes occur in response to effectors to activate transcription (depicted in Figure S1). Although CatM alone does not activate sufficient transcription for growth on benzoate as the carbon source, mutations can increase CatM-regulated PbenA transcription by augmenting its response to muconate or by enabling transcription without an effector [3,4]. Two amino acid replacements in the effector-binding domain (EBD) of CatM each enable BenM-independent growth on benzoate, yet neither of these, nor any other known CatM variant, responds to benzoate.
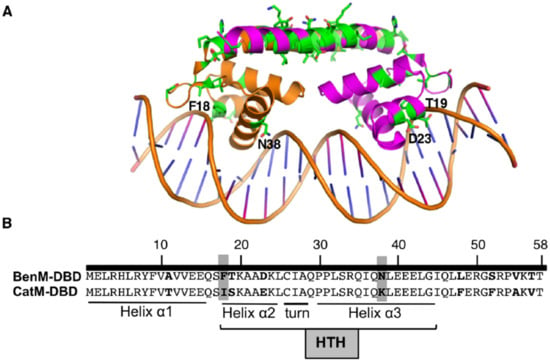
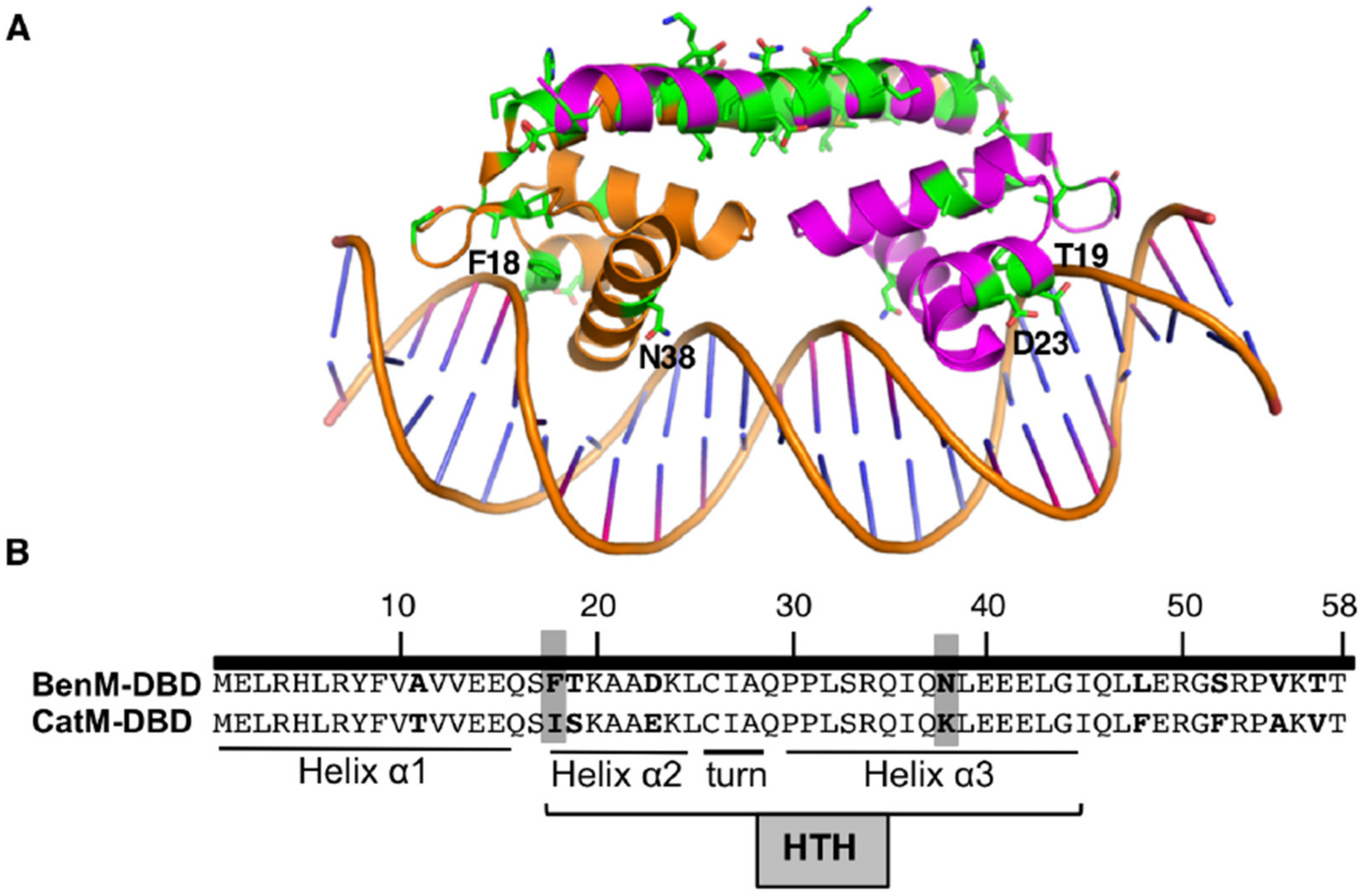
Figure 2.
(A) Ribbon representation of BenM-DBD and the adjacent “linker” helix bound to DNA (PDB ID 4IHS). Green side chains show differences between BenM and CatM. Labeled residues are discussed in the text. (B) Nine amino acids differences in the DBDs (bold), including those at positions 18 and 38 (highlighted). The Helix (α2)-Turn-Helix (α3) (HTH) motif is involved in DNA recognition.
The structures of BenM-EBD and CatM-EBD are highly conserved (Figure S2) [5,6]. Although benzoate binds in a hydrophobic pocket of BenM, this compound has not been detected in structures of the corresponding region of CatM [5,6]. This pocket is distinct from an inter-domain cleft in BenM and CatM that binds muconate and serves as the typical effector-binding site in LTTRs [6]. The EBDs of LTTRs usually assume the conformation of a periplasmic-binding protein [7,8,9]. However, BenM is the only LTTR known to have a secondary effector-binding site that enables synergistic activation of transcription with different metabolites [2]. In an effort to understand this synergism, we engineered amino acid replacements to make CatM more similar to BenM.
Two residues in the hydrophobic binding pocket of BenM are critical for benzoate-activated transcription, R160 and Y293 [5,6]. When these amino acids are replaced with those at the comparable positions of CatM, H160 and F293, BenM fails to activate transcription in response to benzoate as a sole effector or in combination with muconate [5]. Moreover, benzoate inhibits muconate-activated gene expression in these BenM variants [3,5]. As described here, the converse changes in CatM (H160R and F293Y) did not initially generate a benzoate-responsive CatM. However, efforts to alter CatM were expanded to increase understanding of these representative members of the LTTR family, the largest group of homologous transcriptional regulators in bacteria [10]. After multiple attempts, CatM variants were isolated that respond to benzoate and that activate increased transcription in response to both effectors. As discussed below, changes were required in both the DBD and EBD regions of CatM for this new functionality.
2. Materials and Methods
2.1. Bacterial Strains and Growth Conditions
A. baylyi strains, derived from the wild-type ADP1 [11,12] are listed in Table S1. Escherichia coli DH5α (Thermo Fisher Scientific, Waltham, MA, USA) and XL-1 blue (Agilent Technologies, Santa Clara, CA, USA) were used as plasmid hosts. Bacteria were grown on Luria-Bertani (LB) medium at 37 °C [13]. In some cases, A. baylyi strains were grown in minimal medium [14,15] with succinate (10 mM), pyruvate (20 mM), benzoate (2 mM), muconate, (2.5 mM), or anthranilate (1.5 mM) as the carbon source. Antibiotics were added when needed at the following final concentrations: ampicillin, 150 μg/mL, kanamycin, 25 μg/mL, spectinomycin, 13 μg/mL, and streptomycin, 13 μg/mL. Growth was monitored by turbidity (OD600).
2.2. Site-Directed Mutagenesis and Strain Construction by Allelic Replacement
Site-directed mutagenesis of plasmid DNA was conducted with mutagenic primers and methods based on the QuickChange II protocol (Agilent Technologies, Santa Clara, CA, USA [5]). The primers, and the mutations they introduce, are listed in Tables S2 and S3. In some cases, plasmids were constructed using splicing by overlap extension PCR (SOEing) [16]. Linearized plasmid-borne alleles were used to replace chromosomal genes in A. baylyi recipient strains by homologous recombination [17,18]. Transformants were identified by phenotypic changes in antibiotic resistance, carbon source utilization or loss of the sacB marker (in the presence of 10% sucrose and no NaCl in the medium) [5,18]. The genotypes of mutant strains were confirmed by PCR analysis and DNA sequencing (Genewiz laboratories, South Plainfield, NJ, USA) of the chromosomal regions where changes were introduced.
2.3. Selection for BenM-Independent Growth on Benzoate
Strains that form colonies on plates with benzoate as the sole carbon source were defined as Ben+. Spontaneous Ben+ mutants arising from strains lacking BenM were isolated as described [3,4]. Chromosomal catM DNA was recovered from Ben+ strains using the gap-repair method [18,19]. Briefly, cells were grown on benzoate medium to mid-log phase, mixed with linearized pBAC184 (Table S2) and plated on LB medium. Transformants with circularized plasmids, resulting from homologous recombination, were selected in medium with ampicillin. Drug-resistant cells were pooled, and plasmid DNA was extracted and used to transform E. coli. Recovered A. baylyi DNA was tested for the ability to confer a Ben+ phenotype to recipient strains (without BenM) by allelic replacement. Mutations were identified by DNA sequencing (Genewiz laboratories, South Plainfield, NJ, USA).
2.4. β-Galactosidase (LacZ) Assays
Transcriptional lacZ fusions were constructed as described [3,4]. For cultures grown on LB, effectors were added at final concentrations of 500 μM of benzoate or muconate, or when added together, 250 μM of each. Some cultures were grown with pyruvate (20 mM) or muconate (3 mM) as the carbon source. Effectors added to pyruvate-grown cultures were added at the following concentrations: 65 μM benzoate or muconate, or 32.5 μM of each when added together. Growth was measured by optical density (OD600), and assays were done when cultures reached late-exponential phase as described [4,5]. Directions from the FlourAce β-galactosidase reporter kit (BioRad, Hercules, CA, USA) were followed. The hydrolysis of the substrate, 4 methylumbelliferyl-galactopyranoside (MUG) to the product 4-methyllumbelliferone (4MU) was detected with a TD-360 miniflourometer (Turner Designs, San Jose, CA, USA). A standard curve was used to quantify 4MU.
2.5. Purification of BenM and CatM and Variant Proteins
Plasmids, pBAC433 and pBAC430, were used to express full-length regulators with C-terminal histidine tags, BenM-His and CatM-His, respectively [2]. Plasmids were made to encode variants, pBAC1027 (BenM-DBDCatM-His), pBAC1045 (CatM(I18F,K38N)-His), and pBAC1086 (Ben-DBDCatM(H160R,F293Y)-His). BenM-His was purified as described [20]. CatM-His and CatM variants were purified similarly but were eluted in a different buffer (30 mM Tris, 500 mM NaCl, 30% glycerol (v/v), 500 mM imidazole, and 10 mM β-mercaptoethanol (pH 7.9)). Fractions with pure CatM were pooled and dialyzed against a buffer (20 mM Tris-HCl, 250 imidazole (pH 7.9), 500 NaCl, 10% (v/v) glycerol) to increase solubility, and then were concentrated to 2–10 mg·mL−1. Protein concentrations were determined by the method of Bradford with bovine serum albumin as the standard [21]. Proteins fractions were frozen with liquid nitrogen and stored at −70 °C until use.
2.6. Electrophoretic Mobility Shift Assays (EMSAs)
Operator-promoter DNA (PbenA and PcatB) was PCR amplified with 5’-6-carboxyfluorescein (6-FAM) labeled primers (Table S3). PCR products, approximately 150–250 bp, were extracted with a gel DNA recovery kit (Zymo Research, Irvine, CA, USA). For EMSAs, 1 nM DNA was incubated with different concentrations of protein (0, 2.5 nM, 5 nM, 10 nM, 20 nM, 40 nM, 80 nM, 160 nM, 320 nM, 640 nM and 1.28 μM) for 1 h at 37 °C with or without muconate, benzoate or both. Effectors in the reaction were present at a total concentration of 1.6 mM individually or at a concentration of 800 μM each when combined. DNA-protein samples were resolved in 6% polyacrylamide gels. Electrophoresis was performed in buffer (40 mM Tris (pH 7.6), 20 mM acetic acid, 1 mM EDTA) for 1 h at 185 volts at 4 °C. When indicated, effectors were added to this buffer at the same concentrations described above. Fluorescently labeled bands were detected using the Amersham Typhoon PhosphorImager system (GE Healthcare Bio-Sciences, Pittsburgh, PA, USA) at 526 nm using the short-pass emission filter. The bound DNA relative to the unbound DNA was quantified by Gel-Pro analyzer (Media Cybernetics, Rockville, MD, USA). Values were fitted into a saturation curve using the equation for one site binding with total accounting for ligand depletion to determine the equilibrium constant (Kd) (Prism 8 software, GraphPad, San Diego, CA, USA) [22].
2.7. Modeling the Transcription Activation Complex at PbenA and PcatB
An atomic model of PbenA was created by merging the structures of BenM-DBD bound to DNA [23] and a transcription initiation complex of E. coli RNA polymerase (RNAP) [24] using the X3DNA suite [25] with the DNA atoms as alignment guides. Since the Site 1 DNA of PcatB has the most sequence identity with benA Site 2, the structure of BenM-DBD complexed with PcatB Site 1 (PDB ID 4IHS, chains A, B, E, and F) was used to model atoms locally at benA Site 2. The structure of BenM-DBD complexed with PbenA Site 1 (PDB ID 4IHT, chains A, B, E, and F) was used to model benA Site 1. The model for RNAP (PDB ID 4YLN) was positioned assuming that the TTGAAC downstream from PbenA Site 2 corresponds to the E. coli σ70 binding site with the sequence TTGACA in the 4IHT complex. DNA residues in the three structures were changed and renumbered to match the PbenA sequence with the program mutate_bases. A composite DNA backbone was generated by calculating the helical parameter sets (shear, stretch, stagger, buckle, prop-tw, opening, shift, slide, rise, tilt, roll, and twist) with program find_pair from the three individual DNA double helical segments and then merging the parameters into one file. The two helical parameter sets at the transition regions were averaged. A DNA model was generated from the combined helical parameters using the program rebuild. The three atomic structures containing both the DNA and protein residues were aligned on the composite DNA backbone using align command of PyMOL [26]. Similar structures of the PcatB and PbenA5146 transcriptional activation complexes were generated by mutating the benA DNA sequences in the composite structure. The EBD domains were not fitted to the model because the close proximity of the two binding sites is not consistent with any full-length structure of an LTTR at this time.
3. Results
3.1. Engineered CatM Variants: Amino Acid Replacements at Positions 160 and 293
In an effort to make CatM respond to benzoate, A. baylyi strains were engineered to encode CatM (H160R), CatM (F293Y), or CatM (H160R, F293Y) (designated ACN662, ACN682, and ACN685, Table S1). These amino acid replacements were designed to match residues in BenM-EBD that interact with benzoate (Figure S2). To ensure that transcriptional regulation could be attributable to the CatM variants, benM was also disrupted. The engineered strains all grew on anthranilate, catechol, and muconate as sole carbon sources. Such growth requires transcription from the cat-operon promoter, PcatB (Figure 1), indicating that the variants are functional as cat-gene regulators. To assess PbenA regulation, growth on benzoate was evaluated. Without BenM, neither wild-type CatM, in strain ISA36 [3] nor CatM (F293Y) in ACN682 supported growth on benzoate as the carbon source. In contrast, CatM (H160R) in ACN662, and CatM (H160R, F293Y), in ACN685, conferred growth on benzoate, suggesting that H160R enables higher than normal levels of PbenA transcription.
3.2. Transcriptional Regulation of PbenA by CatM Variants
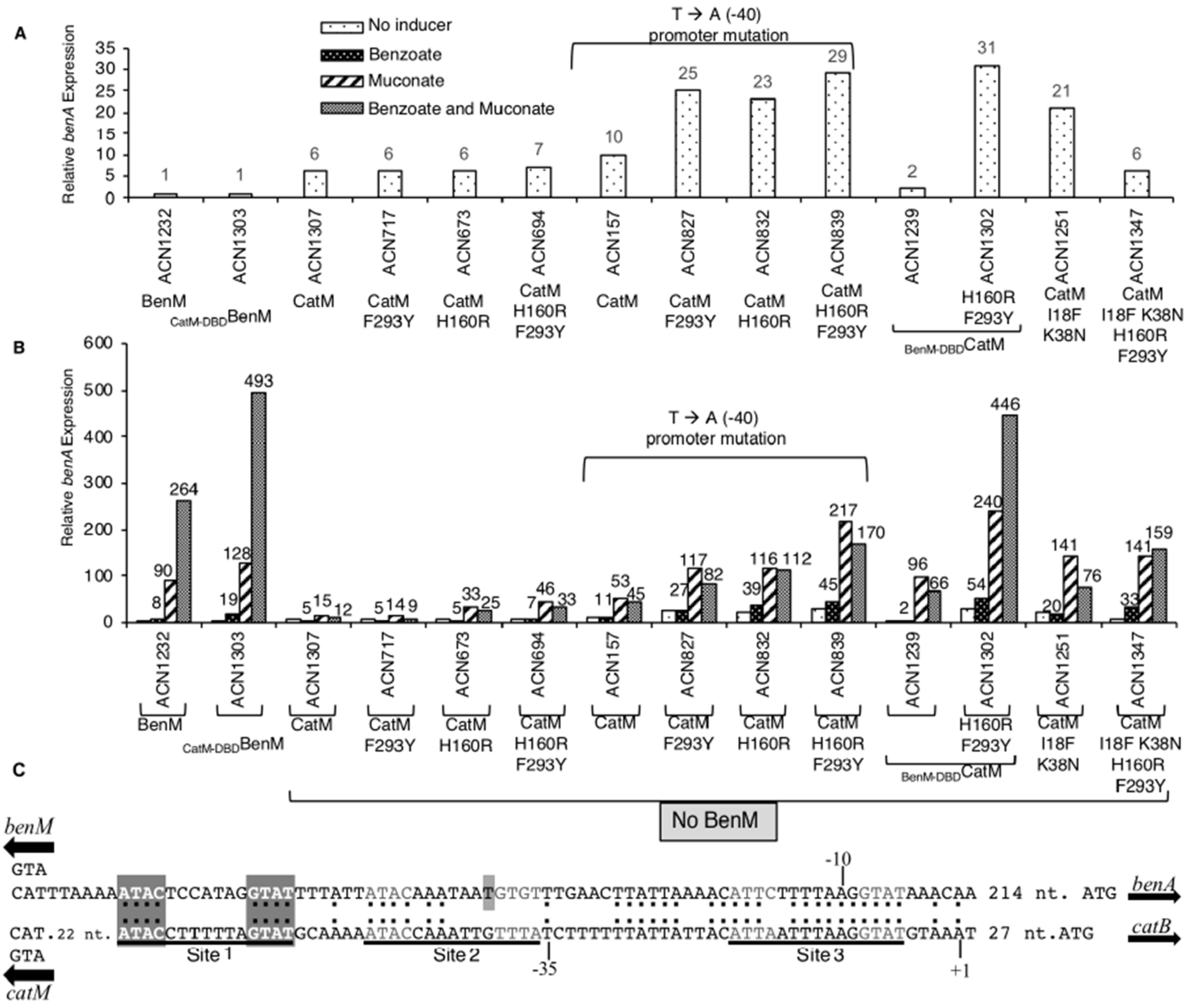
A benA::lacZ fusion was used to replace benA, thereby preventing growth on benzoate as the carbon source and enabling benzoate to be tested as a non-metabolized effector [3]. β-Galactosidase (LacZ) activity reflects PbenA regulation by CatM(F293Y), CatM(H160R), CatM(H160R,F293Y), or wild-type CatM in strains ACN717, ACN673, ACN694, and ACN1307, respectively (Figure 3). BenM is the major regulator of PbenA. Without effectors, a tetramer of BenM or CatM, can bind Site 1 and Site 3 to repress transcription of benA or catB (Figure 3C). Effectors cause a shift in which protein binding to Sites 1 and 2 improves RNAP access to Site 3 [2]. Despite possible cross-regulation, the maximum level of CatM-activated transcription was 18-fold lower than for BenM (ACN1307 versus ACN1232). However, the H160R replacement increased this low-level CatM-mediated response to muconate (ACN673 and ACN694). The elevated transcription levels remained significantly below that for BenM (ACN1232), yet they are comparable to those of other CatM variants that enable BenM-independent growth on benzoate [4]. Regulation by CatM(F293Y), which did not permit growth on benzoate, was comparable to wild-type CatM, which also fails to allow such growth.
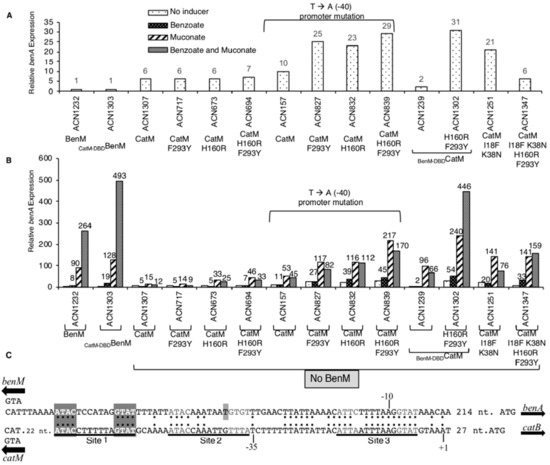
Figure 3.
Relative expression of LacZ. Transcription was controlled by PbenA, or, PbenA5146, which differs by a mutation at −40 relative to the benA transcriptional start site. (A) Enlarged scale displays basal activity (no added effectors). (B) Effectors were added (or not) as indicated. ACN1232 encodes BenM but not CatM, and ACN1307 encodes CatM but not BenM. Other strains encode a CatM variant. Cultures were grown in LB. LacZ activity is reported relative to uninduced ACN1232 (2.6 ± 0.51 nmol/min/mL/OD600). Activities are averages of at least four repetitions; standard deviations were <20% of the average value. (C) Identical nucleotides in aligned PbenA and PcatB regions are indicated (:). The transcriptional start site (+1) and promoter (−10 and −35 regions) are shown for catB. For both regions, Site 1 matches the consensus LTTR-binding motif (T-N11-A, within dyad symmetry, ATAC-N7-GTAT). Site 2 and Site 3 differ slightly from this consensus. The PbenA5146 mutation (T) is marked.
While the catM changes failed to recapitulate the effects of benzoate with BenM, this failure might be due to problems interacting with PbenA DNA as opposed to problems with binding benzoate. In previous studies, CatM activated higher levels of benA transcription when there was a transversion (T-to-A) at position −40 relative to the transcriptional start site [3,4]. Here we tested regulation of this promoter (PbenA5146) by CatM and its variants CatM(F293Y), CatM(H160R), and CatM(H160R,F293Y) using the lacZ fusion (in ACN157, ACN827, ACN832, and ACN839, respectively). CatM yielded higher levels of transcription from PbenA5146 than PbenA under all conditions (ACN157 versus ACN1307, Figure 3). At this promoter, the variants all increased transcription even further than wild-type CatM. Benzoate led to expression levels 170% or 155% of the non-induced levels for CatM(H160R) or CatM(H160R,F293Y), respectively (Figure 3). Despite this suggestion of a minor response to benzoate, muconate-mediated transcription was inhibited by benzoate. In contrast, for BenM, benzoate works synergistically with muconate (ACN1232) [2].
3.3. Spontaneous Mutant with Increased ben-Gene Expression: Changesne in the CatM-DBD
A different approach was used in another attempt to isolate a benzoate-responsive CatM. While ACN682, encoding CatM(F293Y), does not grow on benzoate, its catM mutation might facilitate the ability of additional mutations to confer this trait. Based on this rationale, Ben+ colonies were directly selected from ACN682, as was done previously for wild-type CatM [3,27]. One ACN682-derived mutant encoded two changes, F293Y and I18F. To test the role of I18F, a strain was made to encode only this change which matches residue 18 of BenM-DBD (Figure 2). CatM(I18F) conferred Ben+ growth (ACN1095), indicating increased transcription of PbenA. When tested with the benA::lacZ fusion in ACN1111, there was no response to benzoate as a sole effector, and benzoate decreased the muconate-inducible expression (data not shown). It appears that the Ben+ phenotype arises from an increase in muconate-induced PbenA expression (ACN1111 compared to ACN1307; Figure 4).
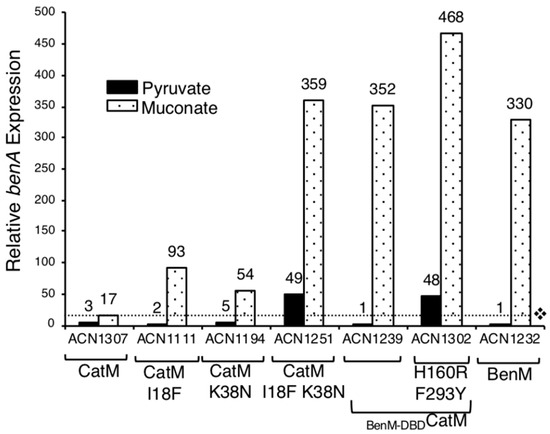
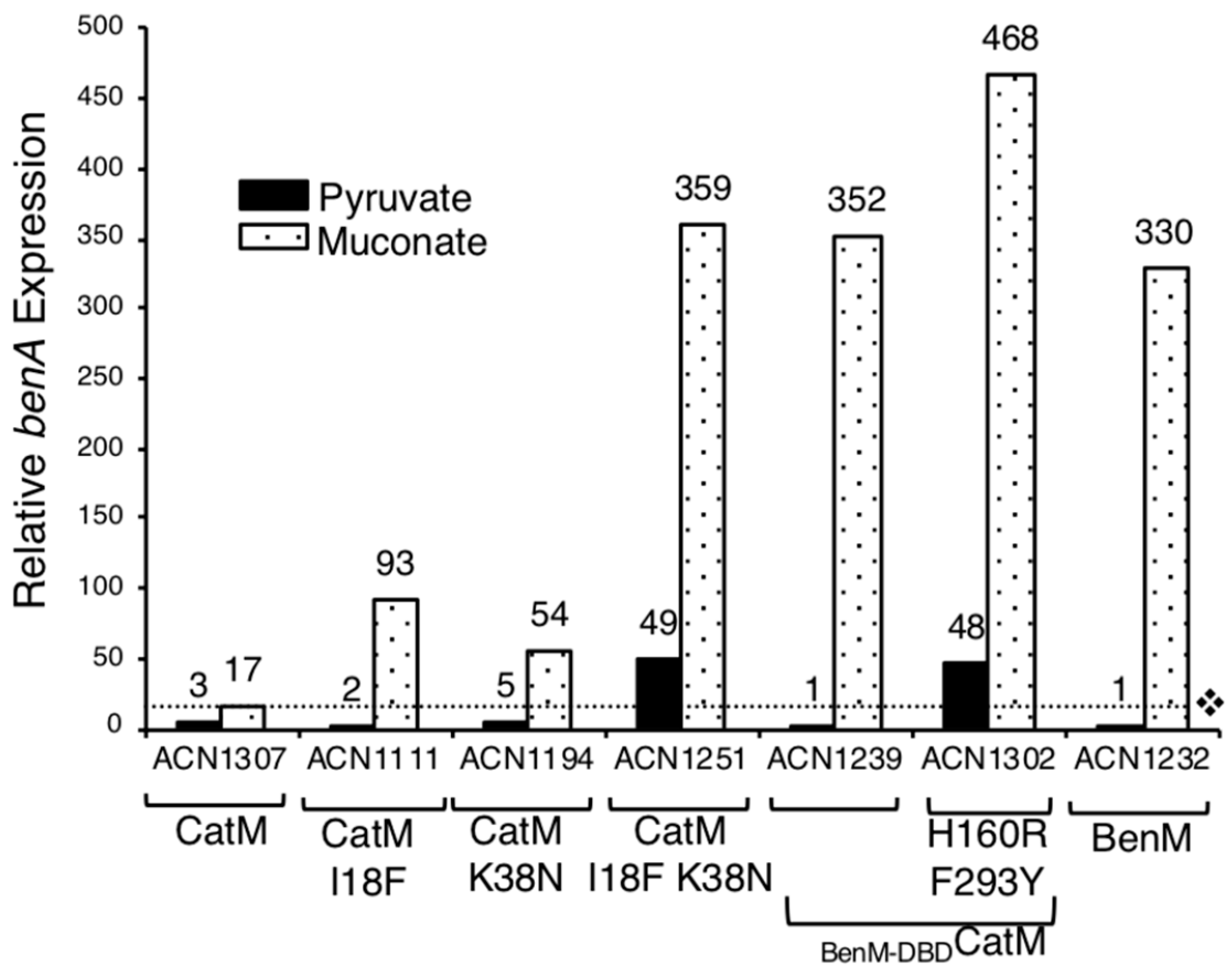
Figure 4.
Relative LacZ activity of a chromosomal benA::lacZ fusion. The only strain encoding BenM is ACN1232, which has no CatM. The dotted line shows the level of the CatM response to muconate (ACN1307). Cultures were grown on pyruvate or muconate as the sole carbon source. Expression is reported relative to the basal level of ACN1232 (6 ± 2 nmol/min/mL/OD600). Values represent the average of at least three independent replicates. Standard deviations were <20% of the average value.
Since I18F makes the variant more like BenM, all nine DBD differences were considered (Figure 2). Residue 38, in the recognition helix of the helix-turn-helix (HTH) DNA-binding motif, is implicated in BenM-DNA interactions [23]. Therefore, strains were made to encode CatM(K38N) and CatM(I18F, K38N) (ACN1193 and ACN1249, respectively). Both strains grew on anthranilate, muconate, and benzoate, indicating that the CatM variants are functional. Muconate enabled CatM(K38N) and CatM(I18F, K38N) to activate higher levels of PbenA transcription than CatM (ACN1194, ACN1251 versus ACN1307, Figure 4). For the double-replacement variant, gene expression was comparable to that mediated by BenM (in ACN1232). CatM(I18F, K38N) also resulted in high PbenA basal expression (Figure 3 and Figure 4). However, there was no transcriptional activation in response to benzoate. Moreover, for both variants with K38N, benzoate decreased the ability of muconate to activate transcription (ACN1251 in Figure 3).
3.4. Further Investigation of DBD Residues in BenM and CatM
The ability of BenM to regulate PbenA might be weakened by having the residues of CatM at positions 18 and 38. To test this possibility, strains were made to encode BenM(F18I,N38K). When tested with a benA::lacZ reporter, regulation by this variant was significantly decreased under all conditions relative to BenM. Nevertheless, BenM(F18I,N38K) remained capable of activating transcription synergistically in response to benzoate and muconate (Supplementary Figure S3).
The entire DBD of CatM was replaced with that of BenM by changing nine amino acids. This variant (BenM-DBDCatM) enabled growth on muconate, anthranilate, and benzoate as sole carbon sources (ACN1234). When regulating a benA::lacZ reporter, this variant activated transcription similarly to BenM with muconate (ACN1239 versus ACN1232). However, there was no response to benzoate as the sole effector, and benzoate decreased muconate-inducible expression as observed for the previously studied CatM variants (Figure 3).
3.5. Benzoate-Responsive CatM Variants Obtained with Combinations of DBD and EBD Changes
Starting with BenM-DBDCatM, we altered the EBD in further attempts to create a response to benzoate. BenM-DBDCatM (H160R, F293Y) enabled growth on benzoate (ACN1301, Table 1). This variant (in ACN1302) also enabled muconate or benzoate to activate high-level transcription (Figure 3 and Figure 4). Notably, there was a BenM-like regulatory pattern when benzoate and muconate were added together. This pattern indicated that 11 amino acid changes are sufficient for a benzoate-responsive CatM capable of synergistic transcriptional activation.

Table 1.
Growth on Benzoate as the Sole Carbon Source a.
Another combination of changes, CatM(I18F,K38N,H160R,F293Y), generated a Ben+ strain (ACN1345, Table 1). While two DBD changes enabled muconate-inducible expression, the two additional EBD changes (in ACN1347) were required for induction by benzoate (a 5.5-fold increase in expression, Figure 3). Moreover, when benzoate and muconate were both provided, gene expression increased for CatM(I18F,K38N,H160R,F293Y). In contrast, for CatM(I18F,K38N), benzoate inhibited the response to muconate, Figure 3.
3.6. Promoter Specificity and Regulator-DNA Binding Affinities
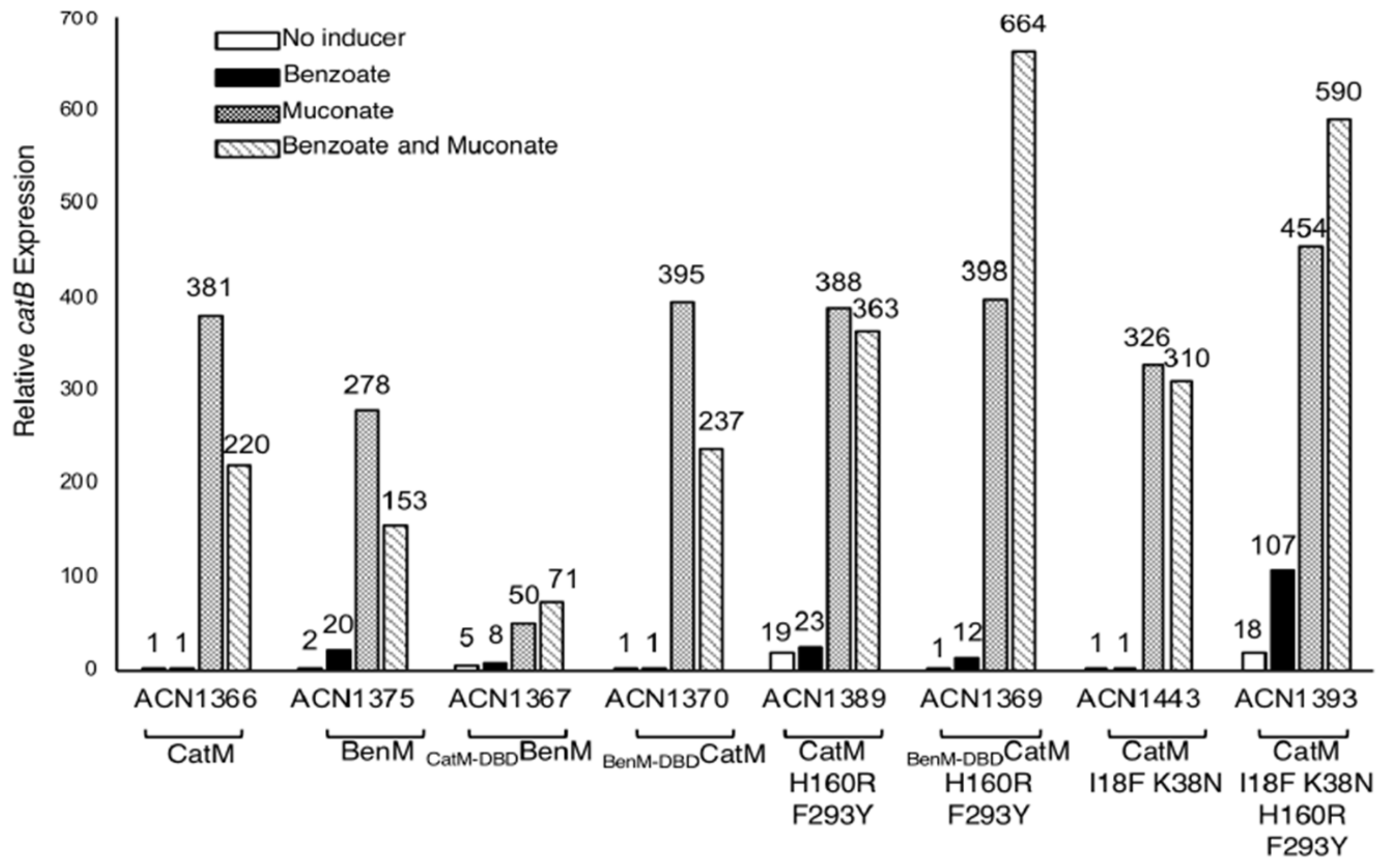
We also studied PcatB. While CatM-DBD should recognize this region, CatM(H160R,F293Y) did not activate transcription of a catB::lacZ fusion in response to benzoate (ACN1389, Figure 5). With BenM, PcatB transcription was high, but the pattern of effector responses differed from what occurred at PbenA. Notably, benzoate failed to enhance transcriptional activation in response to muconate (ACN1375, Figure 5). When the BenM-EBD was combined with the CatM-DBD, the overall response to effectors was lessened and the response pattern was altered (ACN1367 compared to ACN1375, Figure 5). As observed at PbenA, a combination of DBD and EBD changes enabled a response to benzoate. Two CatM variants increased gene expression from PcatB in response to benzoate, BenM-DBDCatM(F293Y,H160R) in ACN1369 and CatM(I18F,K38N,H160R,F293Y) in ACN1393, Figure 5. With these variants, benzoate enhanced transcriptional activation by muconate.
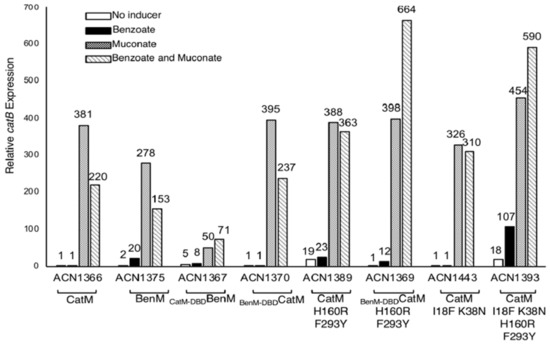
Figure 5.
Expression from a chromosomal catB::lacZ transcriptional reporter. ACN1375 encodes BenM but not CatM. All other strains lack BenM. All have a benA disruption to prevent benzoate catabolism. Cultures were grown on 20 mM pyruvate with or without added effectors. β-Galactosidase (LacZ) activity is reported relative to uninduced ACN1366 (0.71 ± 0.5 nmol/min/mL/OD600). Activities are the average of at least three repetitions, and standard deviations were <10% of the average value.
To evaluate binding to the operator–promoter regions, an electrophoretic mobility gel shift assay (EMSA) was used. However, because the DNA in the assay includes all three regulatory binding sites (Figure 3), the measured Kd coefficients do not distinguish between repression and activation. Thus, the affinity of BenM for PbenA, appears to be of the same order of magnitude regardless of the presence of effectors (Table 2). CatM appeared to have a slightly higher Kd, which corresponds to a lower affinity, for this DNA. Interestingly, the CatM variants that carry the DBD of BenM have a higher affinity for PbenA than either CatM or BenM. This increased affinity appears to require more than the two amino acid changes in the HTH region of the DBD (I18F and K38N). Moreover, increased affinity does not correlate with benzoate responsiveness. These results suggest the hybrid proteins have a conformation that differs from those of the wild-type CatM or BenM.

Table 2.
Estimated Binding Affinities of Regulators to DNA With and Without Effectors a.
In experiments with PcatB, CatM had a higher affinity than BenM for this DNA (Table 2). As for the PbenA region, the CatM variants with the entire BenM-DBD region had a higher affinity for the promoter DNA than did the variant with two replacements in the DBD. While it is difficult to infer the significance of the variations in Kd values, there appears to be no correlation between the ability of a CatM variant to respond to benzoate and its binding affinity (high or low) for PbenA or PcatB.
3.7. Regulator-DNA Interactions at PbenA: A Structural Model with RNAP
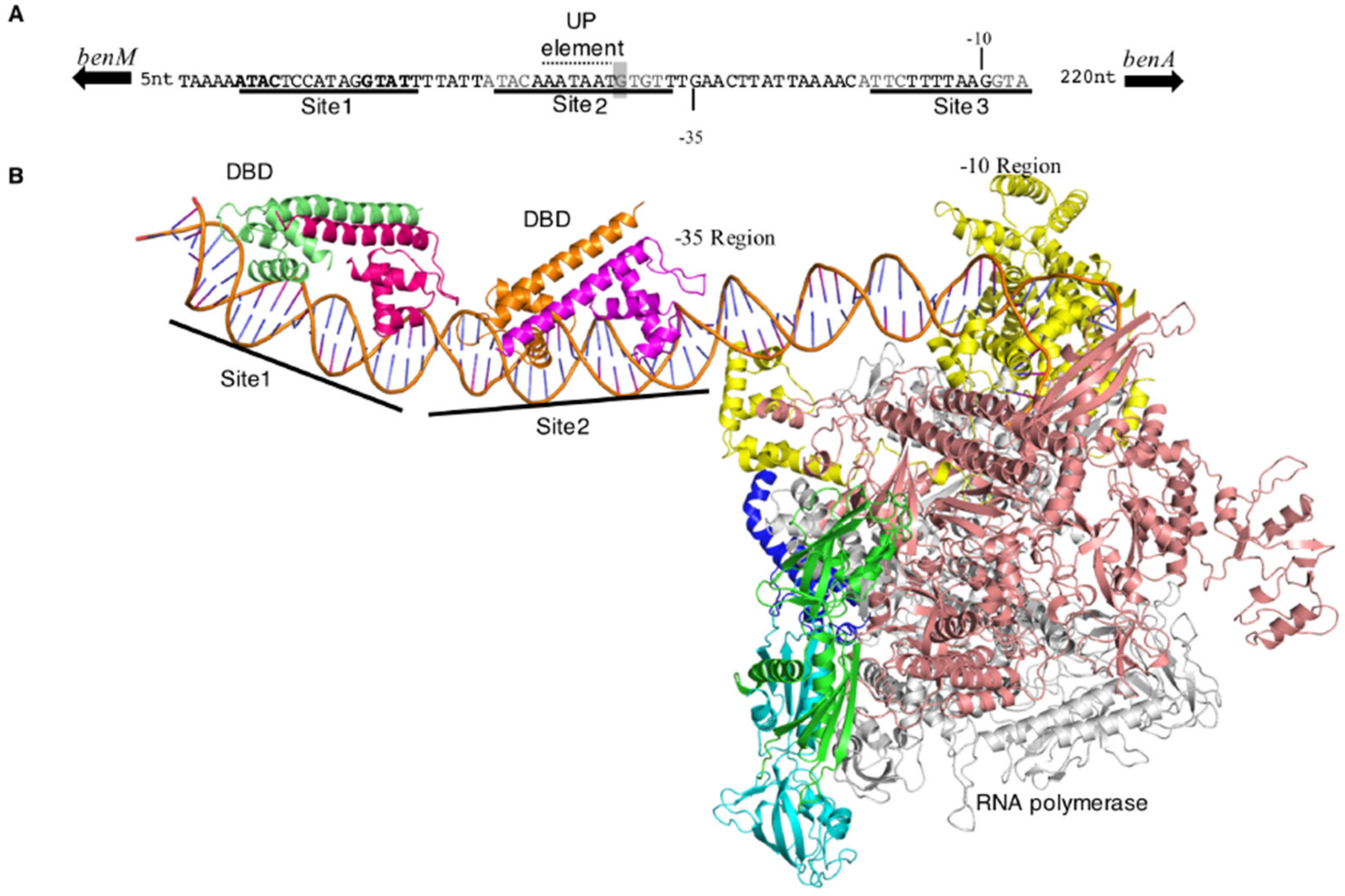
To understand protein-DNA interactions better, a model was built with available structures of BenM-DBD bound to PbenA and PcatB and E. coli RNAP (Figure 6) [23,24]. In this model, Site 2 overlaps the −35 region of the promoter. A run of adenosine nucleotides within Site 2, with a T nucleotide in the middle, appears to create a promoter feature that may increase transcription by interacting with the RNAP α-subunits, an UP element. The relative position of BenM-DBD to the promoter is typical of a class II σ70 promoter in which a regulator directly contacts domain 4 of the sigma factor of bound RNAP to stabilize the initiation complex [28]. However, the model suggests that the BenM-DBDs would not directly contact σ70. The EBD (not in the model) could make contacts with the sigma factor, but the bulk of the EBD units must sit on the opposite face of the RNAP if BenM is a tetramer. In contrast, for class I σ70 promoters, regulators do contact the α-subunit C-terminal domains (α-CTD), but typically the regulatory proteins bind further upstream of bound RNAP. Protein surface residues of the DBD-dimers that flank the UP-element, such as F31 of GcvA (C26 of BenM), have been implicated in α-CTD interactions [29]. The conserved −35 σ70 recognition-sequence in PbenA (TTGAAC vs. consensus TTGACA) suggests that de-repression by conformational change from Site 1 and Site 3 to Site 1 and Site 2 may be a significant aspect of transcriptional activation along with UP-element interactions.
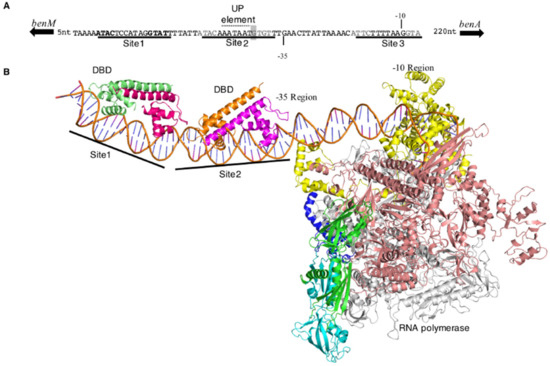
Figure 6.
Structural model of BenM-PbenA interactions. (A). PbenA with binding sites for BenM (underlined). A potential UP element [30] is indicated and discussed in the text. The boxed nucleotide (T) corresponds to the mutation in PbenA5146. (B). Model of the initiation complex at PbenA rendered as a ribbon representation. BenM-DBD subunits are colored pale green, dark magenta, gold, and magenta going from the 5’ (left) to 3’ of the benA promoter; RNA polymerase (RNAP) subunits are green and cyan (α), salmon (β), grey (β’), blue (ω) and yellow (σ).
4. Discussion
4.1. Comparisons of BenM and CatM Provide a Framework to Engineer Effector-Binding Changes in CatM
Residues R160 and Y293 in BenM-EBD, which are critical for benzoate-induced transcriptional activation of PbenA, were introduced into CatM. These replacements were designed to create a hydrophobic binding pocket resembling one in BenM that binds benzoate, termed the secondary effector-binding site (Figure S2) [5,6]. Since benzoate decreased the muconate-activated gene expression for these CatM variants (Figure 3), it is likely that benzoate competes with muconate for binding in the primary effector-binding site of this LTTR rather than binding to a secondary site. In CatM(H160R,F293Y) and CatM(H160R), R160 increased the muconate-activated transcription of PbenA. In BenM, responses to effectors in the primary and secondary binding sites appear to connect through charge-based interactions among three residues that separate muconate and benzoate in the protein structure (R146, E162 and R160) [1].Without benzoate in the secondary binding site of CatM variants, it is unclear how the replacement of H160 with R160 increases the response to muconate. Changes in the local environment may mimic what occurs in BenM upon binding benzoate. Regardless of the mechanism, the increased muconate-responsive transcription due to R160 is consistent with our previous conclusion that this residue in BenM can impact the nearby primary binding site.
The importance of the DBD in PbenA regulation was revealed by a spontaneous Ben+ mutant. In this mutant, one amino acid change in the HTH region altered transcriptional activation in response to muconate. CatM(I18F), in strain ACN1111, increased muconate-dependent PbenA expression more than five-fold relative to CatM, in ACN1307 (Figure 4). A more BenM-like DBD may improve promoter recognition at PbenA and thus enable the CatM-EBD to display its typical effector specificity. Some data support the interpretation that DBDs and EBDs follow expected patterns, whereby promoter recognition is governed by the DBD and effector specificity, is controlled by the EBD. For example, replacing the entire DBD of CatM with that of BenM significantly improves regulation of PbenA (Figure 3 and Figure 4, ACN1239 compared to ACN1307).
Most results suggest a more complicated relationship among the domains. For example, if the low level of CatM-mediated transcription from PbenA were due solely to poor interaction with CatM-DBD, then CatM-DBDBenM would be expected to display a typical BenM pattern of effector response but yield low levels of transcription. Instead, the transcriptional levels mediated by this chimeric protein at PbenA were higher than for BenM (ACN1303 compared to ACN1232, Figure 3). At PcatB this hybrid variant (CatM-DBDBenM) might be expected to increase transcriptional activation compared to BenM because CatM is the cognate regulator of catB. Instead, the overall expression levels were lowered, and the BenM-mediated pattern of response was altered (ACN1367 compared to ACN1375, Figure 5). Another example that is counter to simple predictions resulted from replacing CatM-DBD with BenM-DBD. This alteration to CatM had little effect at PcatB (ACN1366 versus ACN1370, Figure 5). Experimental data are lacking to show if the variant proteins are produced at comparable levels in vivo or whether these proteins are equally stable. Nevertheless, the regulatory patterns, in most cases, are complex. An additional layer of variability results from differences in the flexible helix (the linker helix) that connects the DBD and EBD. In the 30 amino acid residues corresponding to this helix, BenM and CatM are 50% identical and 83% similar in sequence.
4.2. Complex Patterns of Regulation
Even without effectors, DBD alterations impacted transcription. Since BenM and CatM repress transcription in the absence of effectors, a several-fold increase in PbenA basal expression, which results from the loss of BenM, is interpreted as de-repression [2,3]. Consistent with this model, basal expression in most strains encoding CatM or a CatM variant was approximately 6-fold higher than for the strain encoding BenM (e.g., ACN1307, ACN717, ACN673, or ACN694 compared to ACN1232, Figure 3A). In one notable exception, two replacements in the CatM-DBD, I18F and K38N, caused a 21-fold expression increase in basal expression relative to BenM. This high increase suggests that this variant activates transcription without effectors (ACN1251 compared to ACN1232, Figure 3A). This variant may cause a change in protein structure that relieves binding to Site 3 and helps recruit RNAP. When these two DBD changes were accompanied by changes in the EBD or the DBD, high-level basal expression was no longer observed (ACN1347 and ACN1239, respectively, Figure 3A). Yet when all these EBD and DBD changes were combined in a single variant, BenM-DBDCatM(H160R,F293Y), transcriptional activation in the absence of effectors resulted again (ACN1302, Figure 3A). These transcriptional patterns did not correlate meaningfully with changes in the affinity of these proteins for the operator–promoter DNA (Table 2). Better interpretation awaits additional crystal structures and experiments with RNAP. Until then, the crystal structures of BenM-DBD with the Site 1 region of PbenA or PcatB can provide a framework for considering LTTR-DNA interactions.
4.3. Interactions Between DBDs and Operator–Promoter Regions of PbenA and PbenA5146
The I18F replacement in CatM-DBD alters transcription sufficiently to enable Ben+ growth. Crystal structures of BenM-DBD-PbenA-Site 1 and a similar structure, CbnR-DBD-PcbnA-Site indicate that this residue is in helix α2, where it is involved in indirect readout [23,31]. Indirect readout refers to the effects of local nucleotides that cause sequence-dependent deformations of the phosphate backbone to control binding. In BenM, F18 is grouped with residues whose main chain amide N atoms form hydrogen bonds and van der Waals contacts with DNA. The equivalent residue in CbnR, M18, is involved in sugar–phosphate recognition and contributes to DNA-binding strength rather than promoter specificity [31]. CatM binding to PbenA is weaker than for BenM (Table 2), which may be due in part from I18 distorting the packing of the helix against the DNA. While binding affinity was not tested for CatM(I18F), the CatM(I18F,K38N) was evaluated (Table 2). Consistent with our interpretation, this variant had an affinity for PbenA that was intermediate between that of CatM and BenM. The entire BenM-DBD not only further increased the affinity of CatM variants for PbenA, but this affinity actually surpassed that of the native BenM (Table 2). These results indicate that binding affinity is not determined solely by the DBD and also suggest there are important variations in the oligomeric conformation of these regulators.
Residue 38 in BenM and CatM is in the recognition helix (α3) of the HTH motif. N38 in BenM can form a hydrogen bond with the DNA phosphate backbone, although this type of non-specific interaction is unlikely to confer specificity for PbenA. Instead, the importance of this residue may derive from a dynamic interaction with R34, which interacts directly with DNA and provides specificity in DNA recognition. The surrounding residues orient the side chain of R34. An interaction network between these residues and others, including E41 and Q37, might contribute to a conformational switch controlling promoter recognition or RNAP activation at PbenA [23]. K38 in CatM would form an electrostatic interaction with the phosphate backbone of PcatB and would project deeper into the major grove if a purine were positioned at the interaction point, e.g., nucleotide A36 in the BenM-DBD-catB DNA structure. However, the methyl-group of thymine, as found in this nucleotide position of PbenA, would impede this direct interaction. Consistent with this possibility, CatM(K38N) increased muconate-activated transcription of PbenA better than the wild-type CatM (Figure 4). Similar interactions are observed in other LTTRs, such as CbnR (A38, R34, and D42) [31], MetR (S38, S34, H35, and Q42) [32], DntR (R43, N39, T46 and A47) [33] and Tsar (Q38, D42, and S34) [34].
DNA interactions with residue 38 in the DBD may influence effects at PbenA5146. Binding at Site 2 of this promoter (which has a T-to-A transversion) by CatM will be enhanced compared to PbenA due to K38 interactions with the partial negative charge on the N7 atom of the adenine base. Interactions of BenM with PbenMA5146 may not be negatively impacted because a water molecule can bridge to the N7. Thus, the interaction of CatM with PbenMA5146 may be stabilized at Site 2 but not Site 3, thereby disrupting the equilibrium between the protein binding to Site 2 (stabilized by effectors) and Site 3 (stabilized by lack of effectors). This alteration could result in the de-repression of basal activity observed at this promoter for CatM and help explain transcriptional activation in the absence of effectors for the CatM variants with F293Y and/or H160R replacements in the CatM-EBD (Figure 3A).
4.4. Interactions Between DBDs and Operator–Promoter Regions of PcatB
Factors affecting K38 interactions with PbenA5146 may also affect recognition at PcatB. Between the two half sites of dyad symmetry in PcatB Site 2, a G corresponds to the position of the T-to-A mutation of PbenMA5146 (Figure 3C). In general, the sequence ATAC-pyrimidine-N5-purine-GTAT will favor binding at PcatB by CatM. This pattern is observed for Sites 1 and 2. At PbenA, Site 2 swaps the arrangement of the purine/pyrimidine pair, a situation that may partially explain why wild-type CatM shows significantly reduced responses at PbenA.
BenM activated transcription relatively strongly at PcatB despite a different pattern of response to benzoate compared to its activity at PbenA (Figure 5). As measured by EMSAs, the affinity of BenM for PbenA and PcatB is comparable. While the native CatM has a higher affinity for its cognate PcatB promoter than for PbenA, all proteins with a native BenM-DBD have a comparable affinity for both promoters (Table 2). It is not yet possible to discern the features that control such variation in binding and transcriptional control, and a better understanding of LTTR interactions with RNAP is needed.
4.5. A Model of PbenA and PcatB Promoters with RNAP and the DBDs of BenM and CatM
Aspects of the model described above for protein interactions with PbenA also apply to PcatB. However, some important differences stem from DNA sequence variation. PcatB differs in the region of the 3’ half-site of symmetry in Site 2 from that of PbenA. The sequence of PcatB in this region (TCTTTT, Figure 3) does not match the −35 consensus sequence of σ70 promoters. Further, the 3’ half-site of Site 2 (TTTA) lacks similarity to the canonical binding 3’ half-site shared by BenM and CatM, GTAT (PbenA, GTGT). This sequence divergence suggests that the regulators at PcatB may play a more direct role in transcription initiation by replacing the -35 region DNA interactions with σ70 with compensating LTTR-RNAP contact. Analysis of the electrostatic surfaces of σ70 and BenM-DBD at the benA promoter show a strongly negative surface created by D23 in BenM (E23 in CatM) and a strongly positive surface defined by residues K593, R596, and K597 of σ70 which are part of the conserved regulatory region 4.2 of the sigma factor. Clearly, rotation of the RNAP holoenzyme toward BenM by moving forward a few base pairs along PcatB will position the RNAP against BenM or CatM in a complementary fashion. The amino acids in this region 4 of sigma have been mapped as binding residues in other sigma regulatory molecules [35]. Between the half-sites of symmetry of Site 2 in PcatB, there is a mix of A and T nucleotides that could act as an UP-element to stabilize αCTD interactions, as observed in PbenA. This combination of σ70 and αCTD interactions would make the PcatB a mixed class I–class II promoter. However, the UP-element sequence differs from that of PbenA because of the flanking C/G nucleotide pair at the 5’ half-site of Site 2. Alternatively, if CatM and BenM bind only to the 5’ half-site sequence, the 3’ area of Site 2 might be occupied by σ70 (perhaps at the intervening TTGTT) and allow some conformational flexibility for the previously mentioned electrostatic interactions to be favored.
4.6. Broader Implications and Conclusions
Although LTTRs have been studied for more than 30 years, many aspects of their structure and function remain unclear [8]. Because of their similarities and the overlap in their control of a complex regulon, BenM and CatM provide unique opportunities for comparative investigation [1]. Furthermore, their role in aromatic compound catabolism holds promise for biotechnology applications, including lignin valorization [18]. Because of such applications, these and related regulators are receiving renewed attention. For example, transcriptional regulators that respond to aromatic compounds are useful as biosensors [36]. In a recent study, high throughput methods were developed to combine protein domains for DNA binding and those for effector responses with the aim of creating benzoate-responsive biosensors [37]. This approach builds on the modular nature of bacterial transcriptional regulators.
Our studies demonstrate that regulatory alterations can be engineered to integrate multiple signals by enabling the simultaneous response to more than one effector molecule. Although it took repeated efforts, we obtained CatM variants that recognize and respond to benzoate. In some variants, benzoate increased the transcriptional response to muconate, thereby approximating the synergistic transcriptional regulation mediated by BenM. This type of rapid signal amplification has potential uses in metabolic engineering and synthetic biology. While it is not yet possible to interpret some characteristics of the variant regulators that were studied, our results lay the foundation for continued investigation. LTTRs are more than the sum of their parts; both the DBDs and the EBDs affected promoter specificity as well as transcriptional responses to effectors. The abundance of this family of regulators and the importance of LTTR-regulated processes underscore the value of continued research.
Supplementary Materials
The following are available online at https://www.mdpi.com/2073-4425/10/6/421/s1, Figure S1. Regulatory model for PbenA. Figure S2. Structural comparisons of BenM-EBD and CatM-EBD. Figure S3. Changes in BenM-DBD affect relative benA expression. Table S1. Acinetobacter baylyi strains. Table S2. Plasmids. Table S3. Primers.
Author Contributions
Conception and design: M.P.T.-V., C.M., and E.L.N.; Data collection and analysis: M.P.T.-V., C.M., E.L.N; Plasmid and strain construction: M.P.T.-V. and N.S.L.; Lab experiments: M.P.T.-V.; Manuscript writing: M.P.T.-V., E.L.N and C.M.
Funding
This research was supported by NSF grants MCB1024108 (to C.M. and E.L.N) and MCB1615365 (to E.L.N.).
Acknowledgments
We thank Ajchareeya Ruangprasert and Melesse Nune for assistance in protein purification and electromobility shift assays. We also thank Curtis Bacon for help with plasmid and strain construction. Jenifer Morgan, Chelsea Kline, James Valle and Walker Whitley assisted with the selection of A. baylyi strains.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Craven, S.H.; Ezezika, O.C.; Momany, C.; Neidle, E.L. LysR homologs in Acinetobacter: Insights into a diverse and prevalent family of transcriptional regulators. In Acinetobacter Molecular Biology; Gerischer, U., Ed.; Caister Academic Press: Norfolk, UK, 2008; pp. 163–202. [Google Scholar]
- Bundy, B.M.; Collier, L.S.; Hoover, T.R.; Neidle, E.L. Synergistic transcriptional activation by one regulatory protein in response to two metabolites. Proc. Natl. Acad. Sci. USA 2002, 99, 7693–7698. [Google Scholar] [CrossRef]
- Collier, L.S.; Gaines, G.L., 3rd; Neidle, E.L. Regulation of benzoate degradation in Acinetobacter sp. strain ADP1 by BenM, a LysR-type transcriptional activator. J. Bacteriol. 1998, 180, 2493–2501. [Google Scholar]
- Ezezika, O.C.; Collier-Hyams, L.S.; Dale, H.A.; Burk, A.C.; Neidle, E.L. CatM regulation of the benABCDE operon: functional divergence of two LysR-type paralogs in Acinetobacter baylyi ADP1. Appl. Environ. Microbiol. 2006, 72, 1749–1758. [Google Scholar] [CrossRef]
- Craven, S.H.; Ezezika, O.C.; Haddad, S.; Hall, R.A.; Momany, C.; Neidle, E.L. Inducer responses of BenM, a LysR-type transcriptional regulator from Acinetobacter baylyi ADP1. Mol. Microbiol. 2009, 72, 881–894. [Google Scholar] [CrossRef] [PubMed]
- Ezezika, O.; Haddad, S.; Clark, T.; Neidle, E.; Momany, C. Distinct effector-binding sites enable synergistic transcriptional activation by BenM, a LysR-type regulator. J. Mol. Biol. 2007, 367, 616–629. [Google Scholar] [CrossRef] [PubMed]
- Maddocks, S.E.; Oyston, P.C. Structure and function of the LysR-type transcriptional regulator (LTTR) family proteins. Microbiology 2008, 154, 3609–3623. [Google Scholar] [CrossRef]
- Momany, C.; Neidle, E.L. Defying stereotypes: the elusive search for a universal model of LysR-type regulation. Mol. Microbiol. 2012, 83, 453–456. [Google Scholar] [CrossRef] [PubMed]
- Quiocho, F.A.; Ledvina, P.S. Atomic structure and specificity of bacterial periplasmic receptors for active transport and chemotaxis: Variation of common themes. Mol. Microbiol. 1996, 20, 17–25. [Google Scholar] [CrossRef] [PubMed]
- Schell, M.A. Molecular biology of the LysR family of transcriptional regulators. Annu. Rev. Microbiol. 1993, 47, 597–626. [Google Scholar] [CrossRef]
- Juni, E.; Janik, A. Transformation of Acinetobacter calco-aceticus (Bacterium anitratum). J. Bacteriol. 1969, 98, 281–288. [Google Scholar]
- Vaneechoutte, M.; Young, D.M.; Ornston, L.N.; De Baere, T.; Nemec, A.; Van Der Reijden, T.; Carr, E.; Tjernberg, I.; Dijkshoorn, L. Naturally transformable Acinetobacter sp. strain ADP1 belongs to the newly described species Acinetobacter baylyi. Appl. Environ. Microbiol. 2006, 72, 932–936. [Google Scholar] [CrossRef]
- Sambrook, J.; Fritsch, F.; Maniatis, T. Molecular cloning: A laboratory manual, 2nd ed.; Cold Spring Harbor Laboratory Press: Cold Spring Harbor, NY, USA, 1989. [Google Scholar]
- Shanley, M.S.; Neidle, E.L.; Parales, R.E.; Ornston, L.N. Cloning and expression of Acinetobacter calcoaceticus catBCDE genes in Pseudomonas putida and Escherichia coli. J. Bacteriol. 1986, 165, 557–563. [Google Scholar] [CrossRef] [PubMed]
- Singh, A.; Bedore, S.R.; Sharma, N.K.; Lee, S.A.; Eiteman, M.A.; Neidle, E.L. Removal of aromatic inhibitors produced from lignocellulosic hydrolysates by Acinetobacter baylyi ADP1 with formation of ethanol by Kluyveromyces marxianus. Biotechnol. Biofuels. 2019, 12. [Google Scholar] [CrossRef] [PubMed]
- Horton, R.M.; Cai, Z.L.; Ho, S.N.; Pease, L.R. Gene splicing by overlap extension: tailor-made genes using the polymerase chain reaction. Biotechniques 1990, 8, 528–535. [Google Scholar] [CrossRef] [PubMed]
- Neidle, E.L.; Hartnett, C.; Ornston, L.N. Characterization of Acinetobacter calcoaceticus catM, a repressor gene homologous in sequence to transcriptional activator genes. J. Bacteriol. 1989, 171, 5410–5421. [Google Scholar] [CrossRef] [PubMed]
- Tumen-Velasquez, M.; Johnson, C.W.; Ahmed, A.; Dominick, G.; Fulk, E.M.; Khanna, P.; Lee, S.A.; Schmidt, A.L.; Linger, J.G.; Eiteman, M.A.; et al. Accelerating pathway evolution by increasing the gene dosage of chromosomal segments. Proc. Natl. Acad. Sci. USA 2018, 115, 7105–7110. [Google Scholar] [CrossRef]
- Gregg-Jolly, L.A.; Ornston, L.N. Recovery of DNA from the Acinetobacter calcoaceticus chromosome by gap repair. J. Bacteriol. 1990, 172, 6169–6172. [Google Scholar] [CrossRef] [PubMed]
- Ruangprasert, A.; Craven, S.H.; Neidle, E.L.; Momany, C. Full-length structures of BenM and two variants reveal different oligomerization schemes for LysR-type transcriptional regulators. J. Mol. Biol. 2010, 404, 568–586. [Google Scholar] [CrossRef] [PubMed]
- Bradford, M.M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 1976, 72, 248–254. [Google Scholar] [CrossRef]
- Swillens, S. Interpretation of binding curves obtained with high receptor concentrations: Practical aid for computer analysis. Mol. Pharmacol. 1995, 47, 1197–1203. [Google Scholar]
- Alanazi, A.M.; Neidle, E.L.; Momany, C. The DNA-binding domain of BenM reveals the structural basis for the recognition of a T-N11-A sequence motif by LysR-type transcriptional regulators. Acta Crystallogr. D. Biol. Crystallogr. 2013, 69, 1995–2007. [Google Scholar] [CrossRef] [PubMed]
- Zuo, Y.; Steitz, T.A. Crystal structures of the E. coli transcription initiation complexes with a complete bubble. Mol. Cell. 2015, 58, 534–540. [Google Scholar] [CrossRef]
- Lu, X.J.; Olson, W.K. 3DNA: a software package for the analysis, rebuilding and visualization of three-dimensional nucleic acid structures. Nucleic. Acids. Res. 2003, 31, 5108–5121. [Google Scholar] [CrossRef] [PubMed]
- DeLano, W.L. The Pymol User’s Manual. Delano Scientific: San Carlos, CA, USA, 2002. [Google Scholar]
- Cosper, N.J.; Collier, L.S.; Clark, T.J.; Scott, R.A.; Neidle, E.L. Mutations in catB, the gene encoding muconate cycloisomerase, activate transcription of the distal ben genes and contribute to a complex regulatory circuit in Acinetobacter sp. strain ADP1. J. Bacteriol. 2000, 182, 7044–7052. [Google Scholar] [CrossRef] [PubMed]
- Busby, S.; Ebright, R.H. Transcription activation at class II CAP-dependent promoters. Mol. Microbiol. 1997, 23, 853–859. [Google Scholar] [CrossRef] [PubMed]
- Stauffer, L.T.; Stauffer, G.V. GcvA interacts with both the alpha and sigma subunits of RNA polymerase to activate the Escherichia coli gcvB gene and the gcvTHP operon. FEMS Microbiol. Lett. 2005, 242, 333–338. [Google Scholar] [CrossRef]
- Ross, W.; Gosink, K.K.; Salomon, J.; Igarashi, K.; Zou, C.; Ishihama, A.; Severinov, K.; Gourse, R.L. A third recognition element in bacterial promoters: DNA binding by the alpha subunit of RNA polymerase. Science 1993, 262, 1407–1413. [Google Scholar] [CrossRef] [PubMed]
- Koentjoro, M.P.; Adachi, N.; Senda, M.; Ogawa, N.; Senda, T. Crystal structure of the DNA-binding domain of the LysR-type transcriptional regulator CbnR in complex with a DNA fragment of the recognition-binding site in the promoter region. FEBS J. 2018, 285, 977–989. [Google Scholar] [CrossRef]
- Punekar, A.S.; Porter, J.; Carr, S.B.; Phillips, S.E. Structural basis for DNA recognition by the transcription regulator MetR. Acta Crystallogr. F. Struct. Biol. Commun. 2016, 72, 417–426. [Google Scholar] [CrossRef] [PubMed]
- Lerche, M.; Dian, C.; Round, A.; Lonneborg, R.; Brzezinski, P.; Leonard, G.A. The solution configurations of inactive and activated DntR have implications for the sliding dimer mechanism of LysR transcription factors. Sci. Rep. 2016, 6, 19988. [Google Scholar] [CrossRef]
- Monferrer, D.; Tralau, T.; Kertesz, M.A.; Dix, I.; Sola, M.; Uson, I. Structural studies on the full-length LysR-type regulator TsaR from Comamonas testosteroni T-2 reveal a novel open conformation of the tetrameric LTTR fold. Mol. Microbiol. 2010, 75, 1199–1214. [Google Scholar] [CrossRef]
- Lonetto, M.A.; Rhodius, V.; Lamberg, K.; Kiley, P.; Busby, S.; Gross, C. Identification of a contact site for different transcription activators in region 4 of the Escherichia coli RNA polymerase sigma70 subunit. J. Mol. Biol. 1998, 284, 1353–1365. [Google Scholar] [CrossRef] [PubMed]
- Jha, R.K.; Bingen, J.M.; Johnson, C.W.; Kern, T.L.; Khanna, P.; Trettel, D.S.; Strauss, C.E.; Beckham, G.T.; Dale, T. A protocatechuate biosensor for Pseudomonas putida KT2440 via promoter and protein evolution. Metab. Eng. Commun. 2018, 6, 33–38. [Google Scholar] [CrossRef] [PubMed]
- Juárez, J.F.; Lecube-Azpeitia, B.; Brown, S.L.; Johnston, C.D.; Church, G.M. Biosensor libraries harness large classes of binding domains for construction of allosteric transcriptional regulators. Nat. Commun. 2018, 9. [Google Scholar] [CrossRef] [PubMed]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).