Computational Model Reveals a Stochastic Mechanism behind Germinal Center Clonal Bursts
Abstract
1. Introduction
2. Methods: A Probabilistic Model of GC Dynamics
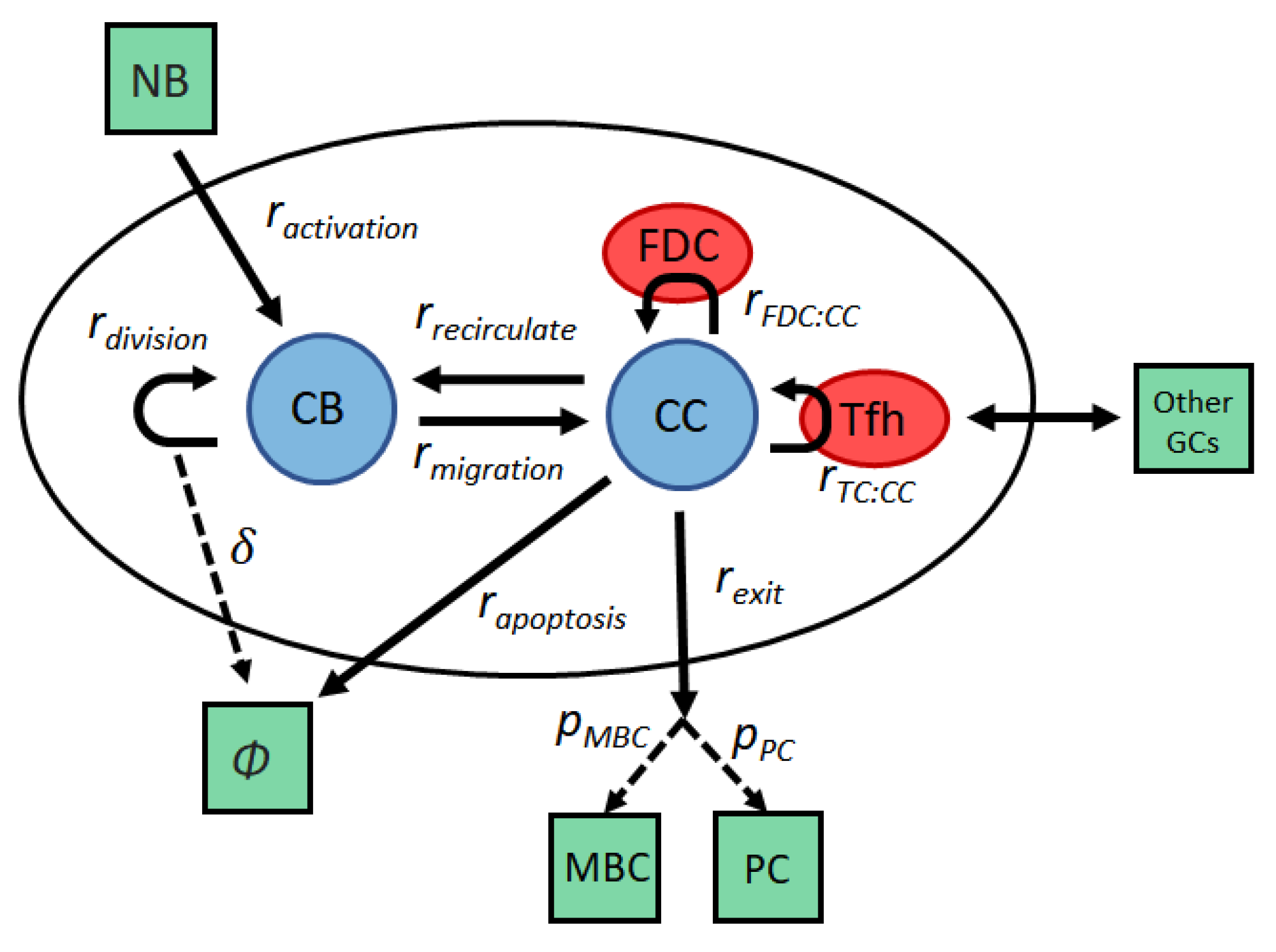
2.1. Stochastic Model of the GC
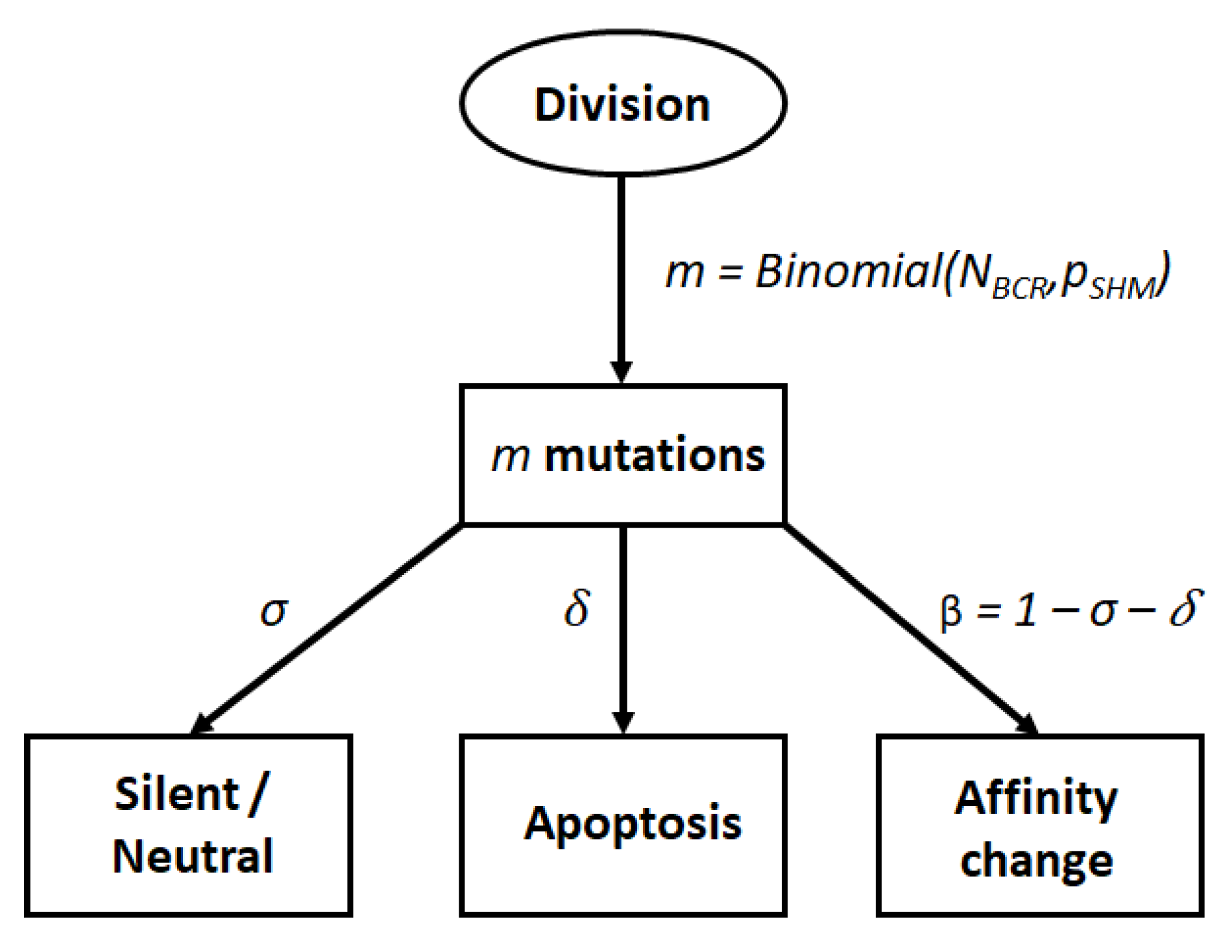
2.2. Cell Fate after Somatic Hypermutation
2.3. B Cell Receptor Affinity to Epitote
2.4. T Dynamics
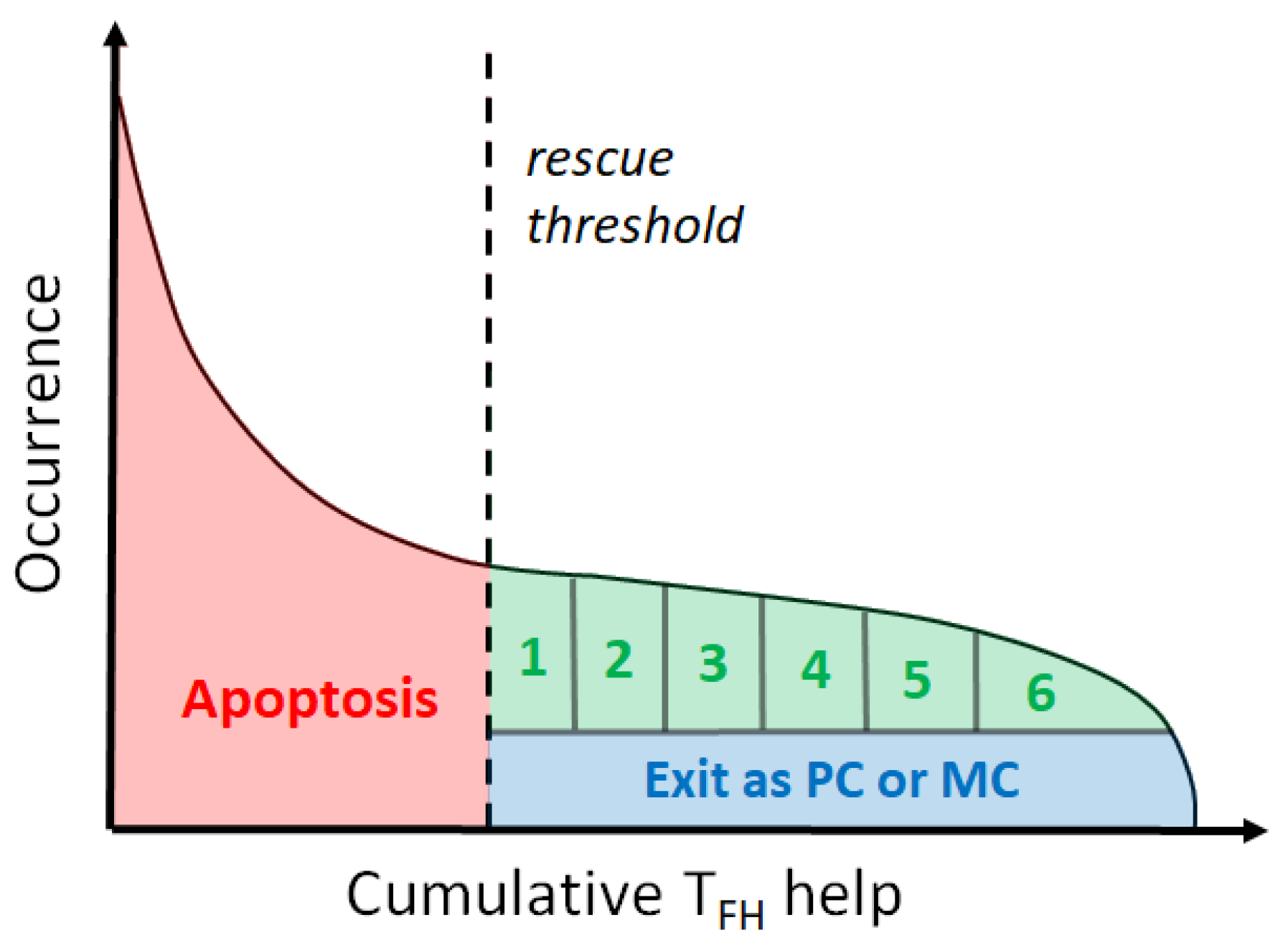
2.5. T Survival Signals
2.6. Regulation of the Cell Cycle
2.7. Plasma Cell and Memory B Cell Differentiation
- Antigen-driven differentiation model (model 1): In this first scenario, we consider that affinity determines B cell fate in a deterministic manner. Namely, there is a threshold pMHC above which a CC becomes a PC. Below that threshold, a cell becomes a MBC.
- Dynamic antigen-driven differentiation model (model 2): A more sophisticated hypothesis is that T cells progressively become less sensitive to the pMHC complex and, as the average affinity of B cells increases in the GC, a larger amount of pMHC is needed to induce the same amount of IRF4. In the second scenario, the choice between PC and MBC is also deterministic, although according to a dynamic threshold that increases linearly with time, i.e., pMHC = affinity + a.
- Stochastic differentiation model (model 3): Finally, we also investigate a probabilistic scenario where, once a B cell has been selected for differentiation, it becomes a PC or MBC according to some fixed probabilities that depend on the amount of expressed pMHC as follows: and .
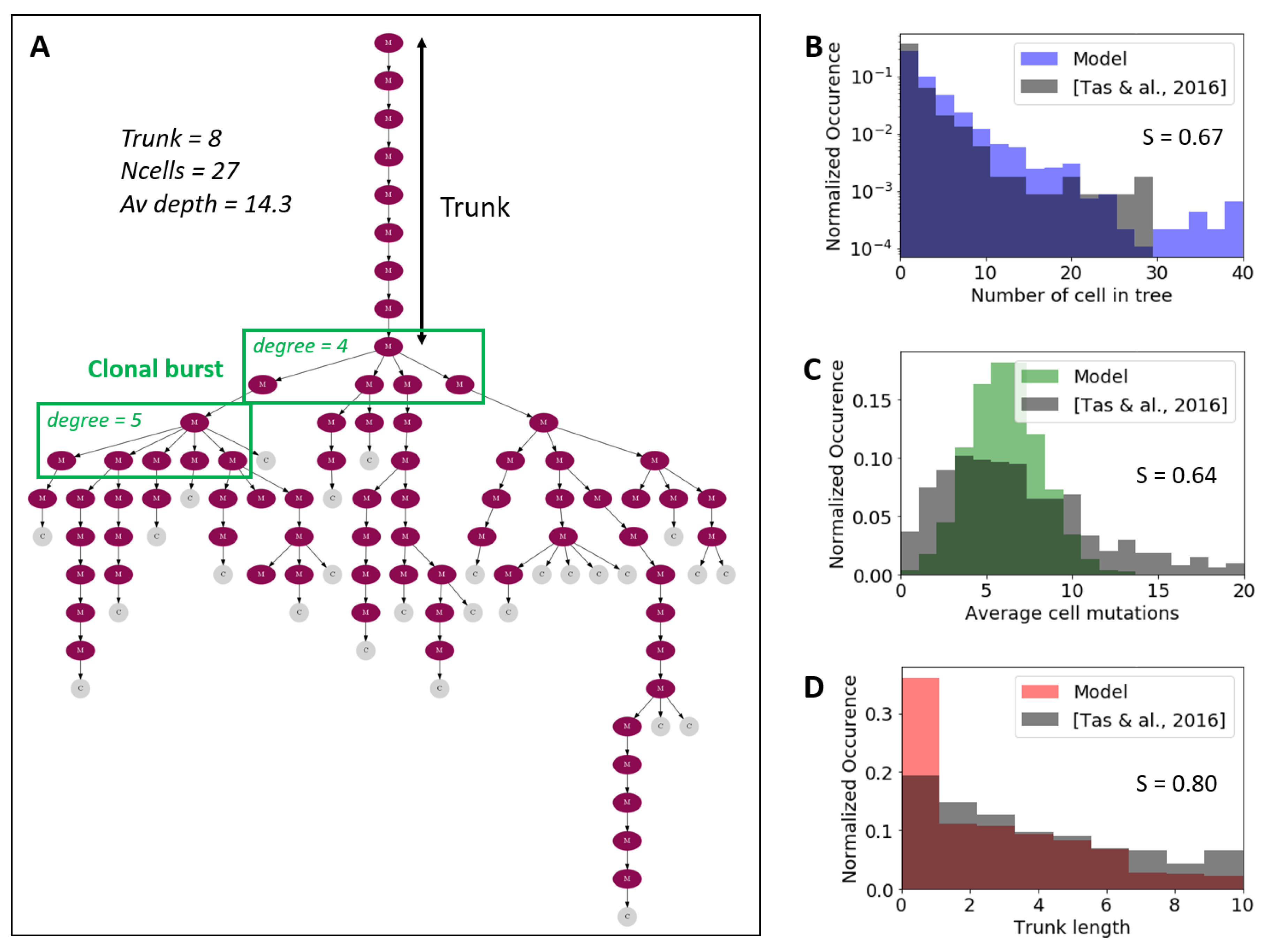
2.8. Phylogenetic Trees Representation
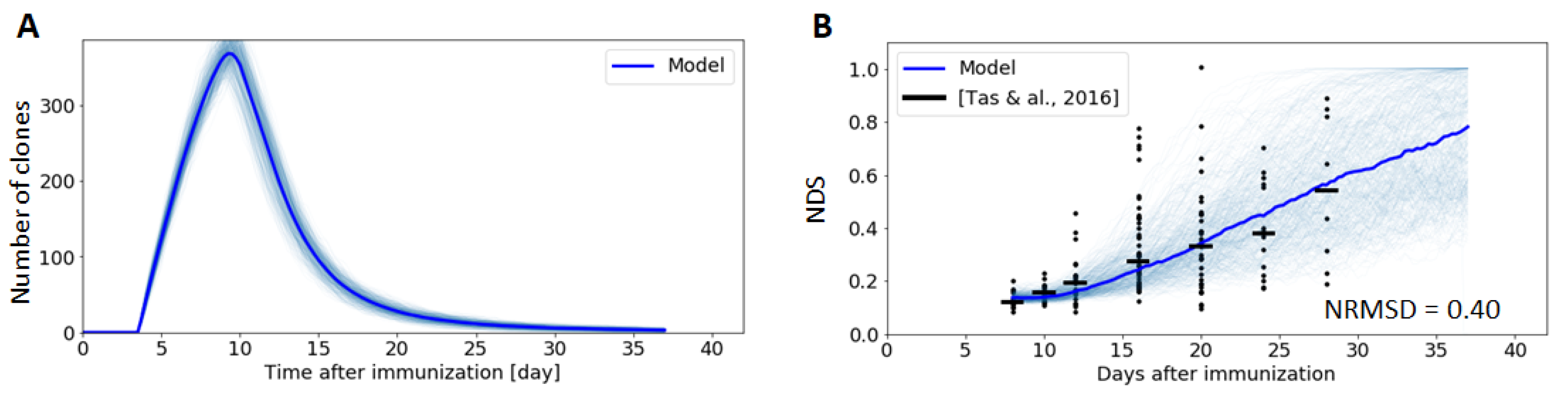
2.9. Clonal Diversity
2.10. Parameter Optimization
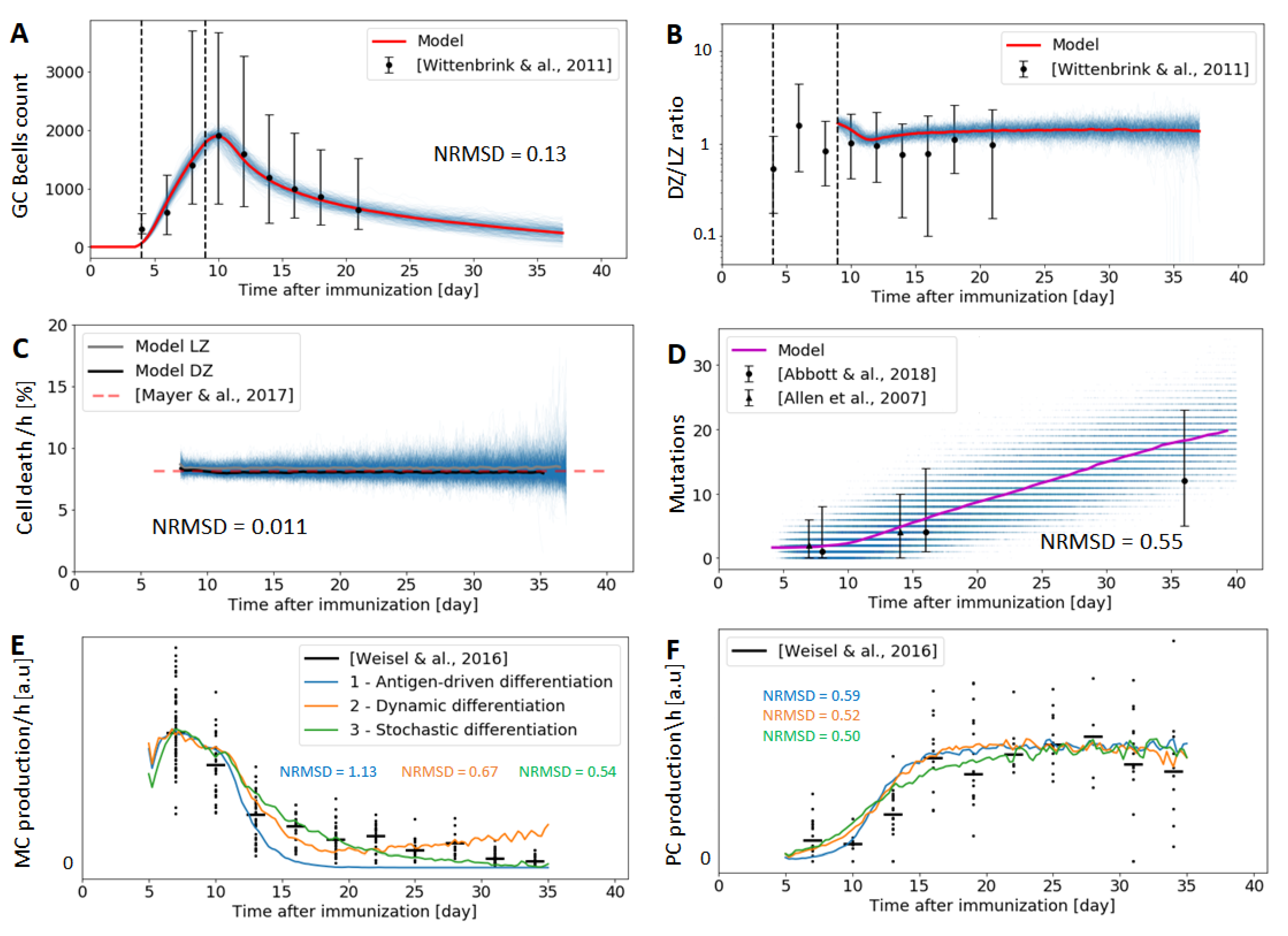
- Number of GC B cells: A time-resolved study of 3457 GC sections from mice provided information about the GC area throughout the GC reaction [18]. Assuming that GC are spherical, then , from where the radius of the GC, , can be estimated. If we further assume that B cells are the predominant cellular type in the GC, we can estimate the number of B cells in the GC as:where we have considered cells to be spherical. Taking [64] and (day 9) [18], we get an average of at the GC peak. As most histological sections do not go cut GCs through the centre, estimations of individual GC volumes are only approximated. Nevertheless, it has been shown that cross-sectional area distributions of spleen sections reflect accurately the broad real-size distributions of GCs [65]. Moreover, as there is significant variability in GC B cell sizes, notably during the cell division cycle [66,67]), the previous number has to be considered only an order of magnitude approximation to the total number of cells. Interestingly, the same study found that GC undergo a fast expansion phase from day 4 to 9, and then progressively decrease in volume by a factor of 2, roughly 10 days after the initial expansion [18].
- DZ/LZ ratio: Despite high variability in size, a typical cellular composition with relatively stable cell ratios of resident T, macrophages, proliferating cells, and apoptotic nuclei seems to be maintained during the established phase of the response [18]. Time-resolved data about the relative size of DZ to LZ is also available from the same study. While the variability in the relative volumes of DZ and LZ is considerably high at all sampled time points (Figure 4B), it stabilizes after the phase of monoclonal expansion to an average of during the entire GC reaction [18]. The high variability is consistent with some other studies that found a ratio of 2 [9]. In our simulation, the DZ/LZ ratio is defined as the number of CBs over the number of CCs; however, we only estimate this ratio after day 9 after immunization, hence avoiding the early days of the GC establishment, where the DZ and LZ are still not spatially separated.
- Cell death: Apoptosis is prevalent in the GC, with up to half of all GC B cells dying every 6 h [55]. We use a relatively constant cell death rate of 8%/h in both the DZ and LZ through the entire GC response.
- GC cellular output: Finally, we use time-resolve data of the GC cellular output [50] to constrain our model. Because the experiment only measured relative abundances rather than direct cell counts, we scale both the model predictions and experimental measures by the maximum value.
3. Results
3.1. Germinal Center Dynamics
3.2. Comparison of Simulated and Experimentally Determined BCR Sequences
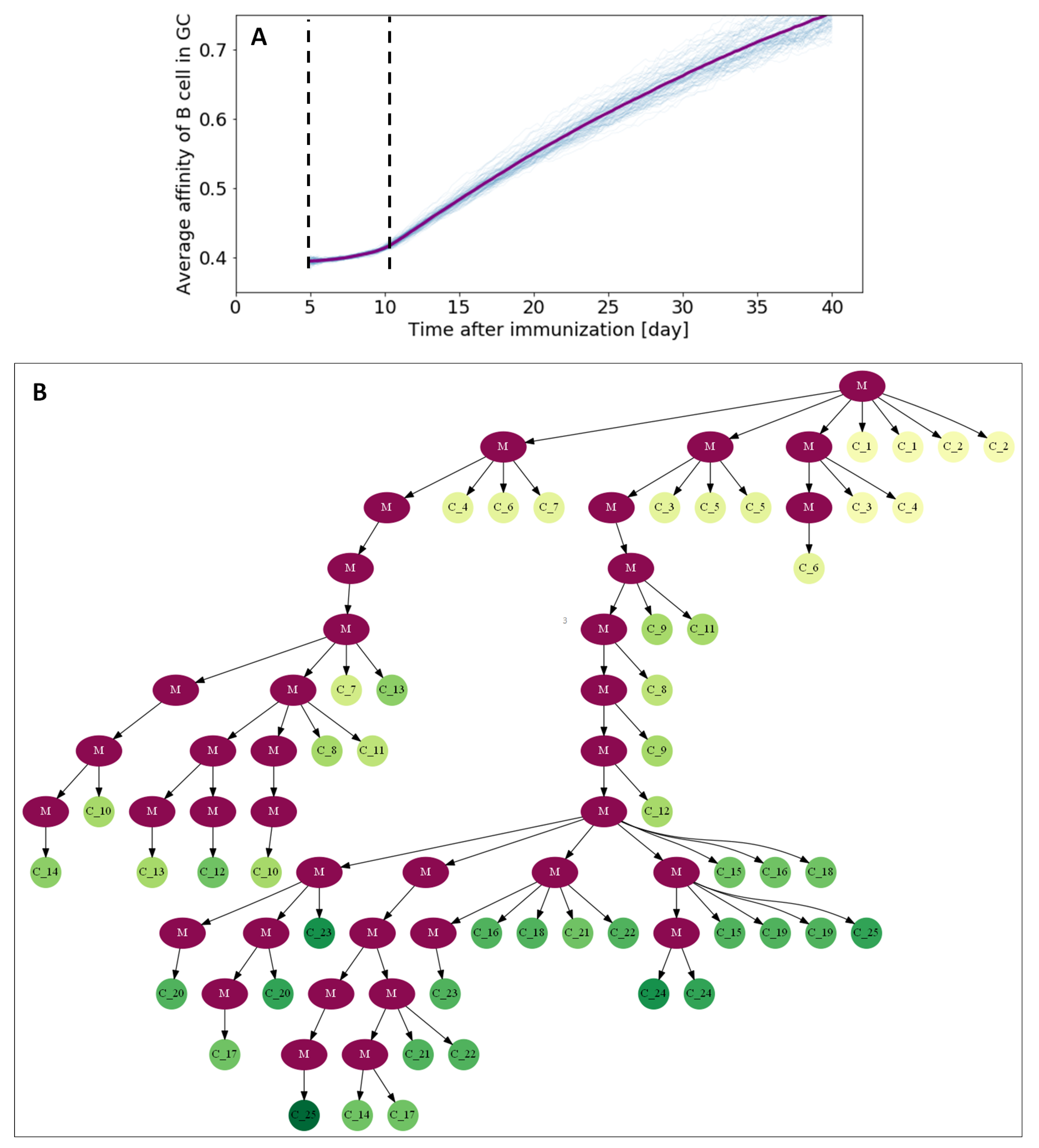
3.3. Visualizing Affinity Maturation
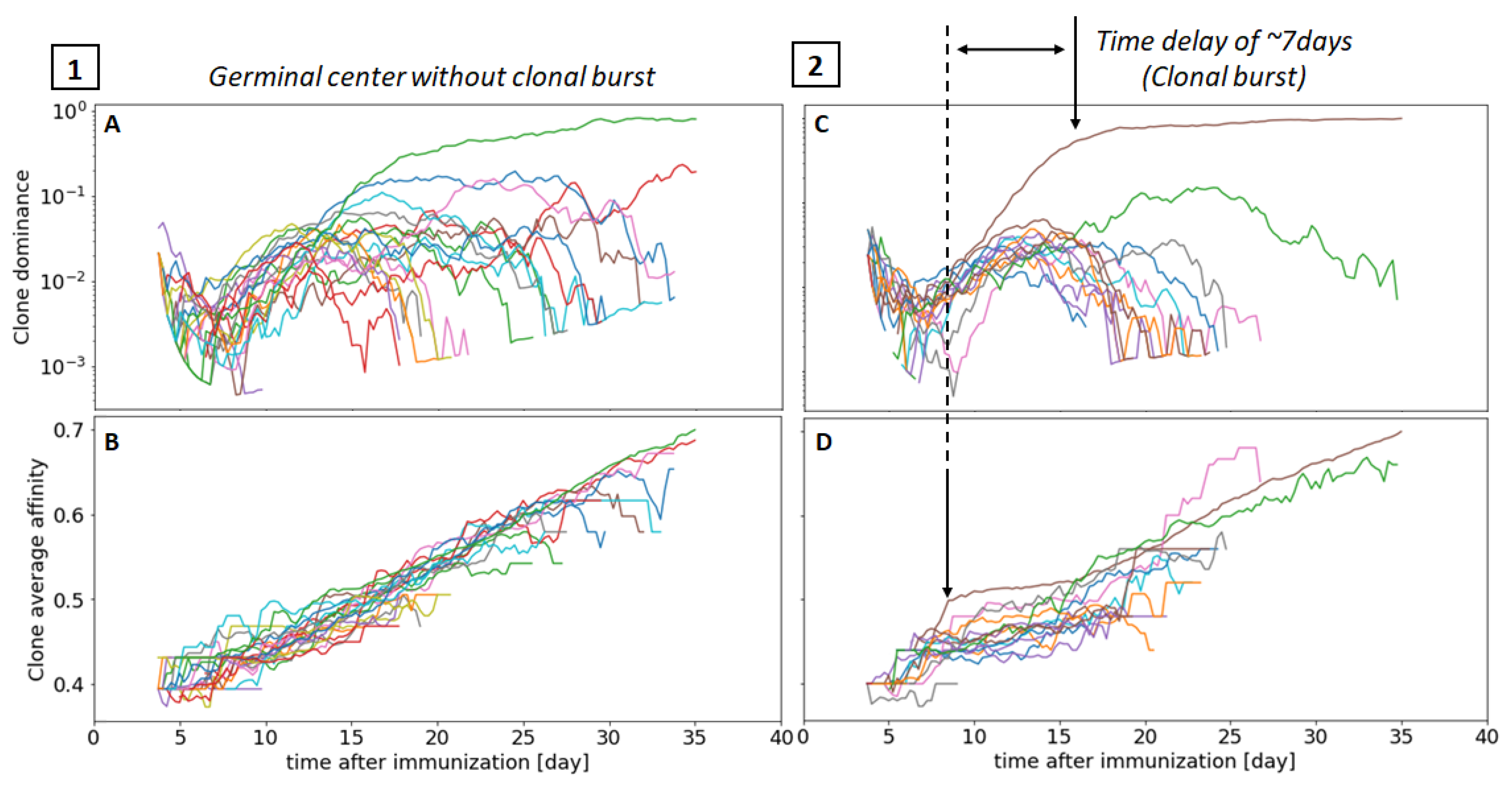
3.4. Visualizing Clonal Competition and Clonal Burst
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Mesin, L.; Ersching, J.; Victora, G.D. Germinal center B cell dynamics. Immunity 2016, 45, 471–482. [Google Scholar] [CrossRef]
- De Silva, N.S.; Klein, U. Dynamics of B cells in germinal centres. Nat. Rev. Immunol. 2015, 15, 137–148. [Google Scholar] [CrossRef] [PubMed]
- Victora, G.D.; Nussenzweig, M.C. Germinal centers. Annu. Rev. Immunol. 2012, 30, 429–457. [Google Scholar] [CrossRef] [PubMed]
- Vinuesa, C.G.; Sanz, I.; Cook, M.C. Dysregulation of germinal centres in autoimmune disease. Nat. Rev. Immunol. 2009, 9, 845–857. [Google Scholar] [CrossRef] [PubMed]
- Pasqualucci, L. Molecular pathogenesis of germinal center-derived B cell lymphomas. Immunol. Rev. 2019, 288, 240–261. [Google Scholar] [CrossRef]
- Allen, C.D.; Okada, T.; Tang, H.L.; Cyster, J.G. Imaging of germinal center selection events during affinity maturation. Science 2007, 315, 528–531. [Google Scholar] [CrossRef]
- Jacob, J.; Kelsoe, G.; Rajewsky, K.; Weiss, U. Intraclonal generation of antibody mutants in germinal centres. Nature 1991, 354, 389–392. [Google Scholar] [CrossRef]
- Berek, C.; Berger, A.; Apel, M. Maturation of the immune response in germinal centers. Cell 1991, 67, 1121–1129. [Google Scholar] [CrossRef]
- Victora, G.D.; Schwickert, T.A.; Fooksman, D.R.; Kamphorst, A.O.; Meyer-Hermann, M.; Dustin, M.L.; Nussenzweig, M.C. Germinal center dynamics revealed by multiphoton microscopy with a photoactivatable fluorescent reporter. Cell 2010, 143, 592–605. [Google Scholar] [CrossRef]
- Gitlin, A.D.; Mayer, C.T.; Oliveira, T.Y.; Shulman, Z.; Jones, M.J.; Koren, A.; Nussenzweig, M.C. T cell help controls the speed of the cell cycle in germinal center B cells. Science 2015, 349, 643–646. [Google Scholar] [CrossRef]
- Heinzel, S.; Giang, T.B.; Kan, A.; Marchingo, J.M.; Lye, B.K.; Corcoran, L.M.; Hodgkin, P.D. A Myc-dependent division timer complements a cell-death timer to regulate T cell and B cell responses. Nat. Immunol. 2017, 18, 96–103. [Google Scholar] [CrossRef] [PubMed]
- Dominguez-Sola, D.; Victora, G.D.; Ying, C.Y.; Phan, R.T.; Saito, M.; Nussenzweig, M.C.; Dalla-Favera, R. The proto-oncogene MYC is required for selection in the germinal center and cyclic reentry. Nat. Immunol. 2012, 13, 1083–1091. [Google Scholar] [CrossRef] [PubMed]
- Calado, D.P.; Sasaki, Y.; Godinho, S.A.; Pellerin, A.; Köchert, K.; Sleckman, B.P.; De Alborán, I.M.; Janz, M.; Rodig, S.; Rajewsky, K. The cell-cycle regulator c-Myc is essential for the formation and maintenance of germinal centers. Nat. Immunol. 2012, 13, 1092–1100. [Google Scholar] [CrossRef] [PubMed]
- Gitlin, A.D.; Shulman, Z.; Nussenzweig, M.C. Clonal selection in the germinal centre by regulated proliferation and hypermutation. Nature 2014, 509, 637–640. [Google Scholar] [CrossRef] [PubMed]
- Tas, J.M.; Mesin, L.; Pasqual, G.; Targ, S.; Jacobsen, J.T.; Mano, Y.M.; Chen, C.S.; Weill, J.C.; Reynaud, C.A.; Browne, E.P.; et al. Visualizing antibody affinity maturation in germinal centers. Science 2016, 351, 1048–1054. [Google Scholar] [CrossRef] [PubMed]
- Song, S.; Matthias, P.D. The transcriptional regulation of germinal center formation. Front. Immunol. 2018, 9, 2026. [Google Scholar] [CrossRef]
- Martínez, M.R.; Corradin, A.; Klein, U.; Álvarez, M.J.; Toffolo, G.M.; di Camillo, B.; Califano, A.; Stolovitzky, G.A. Quantitative modeling of the terminal differentiation of B cells and mechanisms of lymphomagenesis. Proc. Natl. Acad. Sci. USA 2012, 109, 2672–2677. [Google Scholar] [CrossRef]
- Wittenbrink, N.; Klein, A.; Weiser, A.A.; Schuchhardt, J.; Or-Guil, M. Is there a typical germinal center? A large-scale immunohistological study on the cellular composition of germinal centers during the hapten-carrier–driven primary immune response in mice. J. Immunol. 2011, 187, 6185–6196. [Google Scholar] [CrossRef]
- Buchauer, L.; Wardemann, H. Calculating Germinal Centre Reactions. Curr. Opin. Syst. Biol. 2019. [Google Scholar] [CrossRef]
- Thomas, M.J.; Klein, U.; Lygeros, J.; Rodríguez Martínez, M. A probabilistic model of the germinal center reaction. Front. Immunol. 2019, 10, 689. [Google Scholar] [CrossRef]
- Arulraj, T.; Meyer-Hermann, M.; Binder, S.C.; Robert, P.A. Synchronous germinal centre onset impacts the efficiency of antibody responses. Front. Immunol. 2019, 10, 2116. [Google Scholar] [CrossRef]
- Meyer-Hermann, M.; Mohr, E.; Pelletier, N.; Zhang, Y.; Victora, G.D.; Toellner, K.M. A theory of germinal center B cell selection, division, and exit. Cell Rep. 2012, 2, 162–174. [Google Scholar] [CrossRef] [PubMed]
- Wang, P.; Shih, C.M.; Qi, H.; Lan, Y.H. A stochastic model of the germinal center integrating local antigen competition, individualistic T–B Interactions, and B cell receptor signaling. J. Immunol. 2016, 197, 1169–1182. [Google Scholar] [CrossRef]
- Reshetova, P.; van Schaik, B.D.; Klarenbeek, P.L.; Doorenspleet, M.E.; Esveldt, R.E.; Tak, P.P.; Guikema, J.E.; de Vries, N.; van Kampen, A.H. Computational model reveals limited correlation between germinal center B-cell subclone abundancy and affinity: Implications for repertoire sequencing. Front. Immunol. 2017, 8, 221. [Google Scholar] [CrossRef] [PubMed]
- Amitai, A.; Mesin, L.; Victora, G.D.; Kardar, M.; Chakraborty, A.K. A population dynamics model for clonal diversity in a germinal center. Front. Microbial. 2017, 8, 1693. [Google Scholar] [CrossRef] [PubMed]
- Ellebedy, A.H.; Jackson, K.J.L.; Kissick, H.T.; Nakaya, H.I.; Davis, C.W.; Roskin, K.M.; McElroy, A.K.; Oshansky, C.M.; Elbein, R.; Thomas, S.; et al. Defining antigen-specific plasmablast and memory B cell subsets in human blood after viral infection or vaccination. Nat. Immunol. 2016, 17, 1226–1234. [Google Scholar] [CrossRef]
- Peng, H.P.; Lee, K.H.; Jian, J.W.; Yang, A.S. Origins of specificity and affinity in antibody–protein interactions. Proc. Natl. Acad. Sci. USA 2014, 111, E2656–E2665. [Google Scholar] [CrossRef]
- Robert, P.A.; Meyer-Hermann, M. A 3D Structural Affinity Model for Multi-Epitope in Silico Germinal Center Simulations. bioRxiv 2019. [Google Scholar] [CrossRef]
- Dunn-Walters, D.K.; Belelovsky, A.; Edelman, H.; Banerjee, M.; Mehr, R. The dynamics of germinal centre selection as measured by graph-theoretical analysis of mutational lineage trees. Clin. Dev. Immunol. 2002, 9, 233–243. [Google Scholar] [CrossRef]
- Gillespie, D.T. Exact stochastic simulation of coupled chemical reactions. J. Phys. Chem. 1977, 81, 2340–2361. [Google Scholar] [CrossRef]
- McAdams, H.H.; Arkin, A. Stochastic mechanisms in gene expression. Proc. Natl. Acad. Sci. USA 1997, 94, 814–819. [Google Scholar] [CrossRef] [PubMed]
- Arkin, A.; Ross, J.; McAdams, H.H. Stochastic Kinetic Analysis of Developmental Pathway Bifurcation in Phage lambda-Infected Escherichia coli Cells. Genetics 1998, 149, 1633–1648. [Google Scholar] [PubMed]
- Rodríguez Martínez, M.; Soriano, J.; Tlusty, T.; Pilpel, Y.; Furman, I. Messenger RNA fluctuations and regulatory RNAs shape the dynamics of a negative feedback loop. Phys. Rev. E 2010, 81, 031924. [Google Scholar] [CrossRef] [PubMed]
- Schwickert, T.A.; Victora, G.D.; Fooksman, D.R.; Kamphorst, A.O.; Mugnier, M.R.; Gitlin, A.D.; Dustin, M.L.; Nussenzweig, M.C. A dynamic T cell–limited checkpoint regulates affinity-dependent B cell entry into the germinal center. J. Exp. Med. 2011, 208, 1243–1252. [Google Scholar] [CrossRef]
- Liu, Y.J.; Zhang, J.; Lane, P.J.; Chan, E.Y.T.; Maclennan, I.C. Sites of specific B cell activation in primary and secondary responses to T cell-dependent and T cell-independent antigens. Eur. J. Immunol. 1991, 21, 2951–2962. [Google Scholar] [CrossRef]
- Bannard, O.; McGowan, S.J.; Ersching, J.; Ishido, S.; Victora, G.D.; Shin, J.S.; Cyster, J.G. Ubiquitin-mediated fluctuations in MHC class II facilitate efficient germinal center B cell responses. J. Exp. Med. 2016, 213, 993–1009. [Google Scholar] [CrossRef]
- Fiala, G.J.; Kaschek, D.; Blumenthal, B.; Reth, M.; Timmer, J.; Schamel, W.W. Pre-clustering of the B cell antigen receptor demonstrated by mathematically extended electron microscopy. Front. Immunol. 2013, 4, 427. [Google Scholar] [CrossRef][Green Version]
- El Shikh, M.E.M.; El Sayed, R.M.; Sukumar, S.; Szakal, A.K.; Tew, J.G. Activation of B cells by antigens on follicular dendritic cells. Trends Immunol. 2010, 31, 205–211. [Google Scholar] [CrossRef]
- Nowosad, C.R.; Spillane, K.M.; Tolar, P. Germinal center B cells recognize antigen through a specialized immune synapse architecture. Nat. Immunol. 2016, 17, 870–877. [Google Scholar] [CrossRef]
- Shulman, Z.; Gitlin, A.D.; Targ, S.; Jankovic, M.; Pasqual, G.; Nussenzweig, M.C.; Victora, G.D. T follicular helper cell dynamics in germinal centers. Science 2013, 341, 673–677. [Google Scholar] [CrossRef]
- Papa, I.; Saliba, D.; Ponzoni, M.; Bustamante, S.; Canete, P.F.; Gonzalez-Figueroa, P.; McNamara, H.A.; Valvo, S.; Grimbaldeston, M.; Sweet, R.A.; et al. T FH-derived dopamine accelerates productive synapses in germinal centres. Nature 2017, 547, 318–323. [Google Scholar] [CrossRef] [PubMed]
- Janeway, C.A., Jr.; Travers, P.; Walport, M.; Shlomchik, M.J. The structure of a typical antibody molecule. In Immunobiology: The Immune System in Health and Disease, 5th ed.; Garland Science: New York, NY, USA, 2001. [Google Scholar]
- Hoehn, K.B.; Fowler, A.; Lunter, G.; Pybus, O.G. The diversity and molecular evolution of B-cell receptors during infection. Mol. Biol. Evol. 2016, 33, 1147–1157. [Google Scholar] [CrossRef]
- Abbott, R.K.; Lee, J.H.; Menis, S.; Skog, P.; Rossi, M.; Ota, T.; Kulp, D.W.; Bhullar, D.; Kalyuzhniy, O.; Havenar-Daughton, C.; et al. Precursor frequency and affinity determine B cell competitive fitness in germinal centers, tested with germline-targeting HIV vaccine immunogens. Immunity 2018, 48, 133–146. [Google Scholar] [CrossRef] [PubMed]
- Kovaltsuk, A.; Krawczyk, K.; Galson, J.D.; Kelly, D.F.; Deane, C.M.; Trück, J. How B-cell receptor repertoire sequencing can be enriched with structural antibody data. Front. Immunol. 2017, 8, 1753. [Google Scholar] [CrossRef] [PubMed]
- Kleinstein, S.H.; Louzoun, Y.; Shlomchik, M.J. Estimating hypermutation rates from clonal tree data. J. Immunol. 2003, 171, 4639–4649. [Google Scholar] [CrossRef]
- Perelson, A.S.; Oster, G.F. Theoretical studies of clonal selection: Minimal antibody repertoire size and reliability of self-non-self discrimination. J. Theor. Biol. 1979, 81, 645–670. [Google Scholar] [CrossRef]
- Meyer-Hermann, M.; Deutsch, A.; Or-Guil, M. Recycling probability and dynamical properties of germinal center reactions. J. Theor. Biol. 2001, 210, 265–285. [Google Scholar] [CrossRef]
- Liu, D.; Xu, H.; Shih, C.; Wan, Z.; Ma, X.; Ma, W.; Luo, D.; Qi, H. T–B-cell entanglement and ICOSL-driven feed-forward regulation of germinal centre reaction. Nature 2015, 517, 214–218. [Google Scholar] [CrossRef]
- Weisel, F.J.; Zuccarino-Catania, G.V.; Chikina, M.; Shlomchik, M.J. A temporal switch in the germinal center determines differential output of memory B and plasma cells. Immunity 2016, 44, 116–130. [Google Scholar] [CrossRef]
- Ise, W.; Kurosaki, T. Plasma cell differentiation during the germinal center reaction. Immunol. Rev. 2019, 288, 64–74. [Google Scholar] [CrossRef]
- Ise, W.; Fujii, K.; Shiroguchi, K.; Ito, A.; Kometani, K.; Takeda, K.; Kawakami, E.; Yamashita, K.; Suzuki, K.; Okada, T.; et al. T Follicular Helper Cell-Germinal Center B Cell Interaction Strength Regulates Entry into Plasma Cell or Recycling Germinal Center Cell Fate. Immunity 2018, 48, 702–715. [Google Scholar] [CrossRef]
- Shinnakasu, R.; Inoue, T.; Kometani, K.; Moriyama, S.; Adachi, Y.; Nakayama, M.; Takahashi, Y.; Fukuyama, H.; Okada, T.; Kurosaki, T. Regulated selection of germinal-center cells into the memory B cell compartment. Nat. Immunol. 2016, 17, 861–869. [Google Scholar] [CrossRef]
- Suan, D.; Kräutler, N.J.; Maag, J.L.V.; Butt, D.; Bourne, K.; Hermes, J.R.; Avery, D.T.; Young, C.; Statham, A.; Elliott, M.; et al. CCR6 Defines Memory B Cell Precursors in Mouse and Human Germinal Centers, Revealing Light-Zone Location and Predominant Low Antigen Affinity. Immunity 2017, 47, 1142–1153. [Google Scholar] [CrossRef] [PubMed]
- Mayer, C.T.; Gazumyan, A.; Kara, E.E.; Gitlin, A.D.; Golijanin, J.; Viant, C.; Pai, J.; Oliveira, T.Y.; Wang, Q.; Escolano, A.; et al. The microanatomic segregation of selection by apoptosis in the germinal center. Science 2017, 358, eaao2602. [Google Scholar] [CrossRef] [PubMed]
- Shahaf, G.; Barak, M.; Zuckerman, N.S.; Swerdlin, N.; Gorfine, M.; Mehr, R. Antigen-driven selection in germinal centers as reflected by the shape characteristics of immunoglobulin gene lineage trees: A large-scale simulation study. J. Theor. Biol. 2008, 255, 210–222. [Google Scholar] [CrossRef]
- Lefranc, M.P.; Giudicelli, V.; Ginestoux, C.; Jabado-Michaloud, J.; Folch, G.; Bellahcene, F.; Wu, Y.; Gemrot, E.; Brochet, X.; Lane, J.; et al. IMGT®, the international ImMunoGeneTics information system®. Nucleic Acids Res. 2008, 37, D1006–D1012. [Google Scholar] [CrossRef] [PubMed]
- Retter, I.; Althaus, H.H.; Münch, R.; Müller, W. VBASE2, an integrative V gene database. Nucleic Acids Res. 2005, 33, D671–D674. [Google Scholar] [CrossRef] [PubMed]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Thompson, J.D.; Higgins, D.G.; Gibson, T.J. CLUSTAL W: Improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 1994, 22, 4673–4680. [Google Scholar] [CrossRef] [PubMed]
- Jahn, K.; Kuipers, J.; Beerenwinkel, N. Tree inference for single-cell data. Genome Biol. 2016, 17, 86. [Google Scholar] [CrossRef] [PubMed]
- Talevich, E.; Invergo, B.M.; Cock, P.J.; Chapman, B.A. Bio. Phylo: A unified toolkit for processing, analyzing and visualizing phylogenetic trees in Biopython. BMC Bioinform. 2012, 13, 209. [Google Scholar] [CrossRef] [PubMed]
- Meyer-Hermann, M.; Binder, S.C.; Mesin, L.; Victora, G.D. Computer simulation of multi-color brainbow staining and clonal evolution of B cells in germinal centers. Front. Immunol. 2018, 9, 2020. [Google Scholar] [CrossRef] [PubMed]
- Schwickert, T.A.; Lindquist, R.L.; Shakhar, G.; Livshits, G.; Skokos, D.; Kosco-Vilbois, M.H.; Dustin, M.L.; Nussenzweig, M.C. In vivo imaging of germinal centres reveals a dynamic open structure. Nature 2007, 446, 83–87. [Google Scholar] [CrossRef] [PubMed]
- Wittenbrink, N.; Weber, T.S.; Klein, A.; Weiser, A.A.; Zuschratter, W.; Sibila, M.; Schuchhardt, J.; Or-Guil, M. Broad volume distributions indicate nonsynchronized growth and suggest sudden collapses of germinal center B cell populations. J. Immunol. 2010, 184, 1339–1347. [Google Scholar] [CrossRef] [PubMed]
- Monroe, J.G.; Cambier, J.C. Sorting of B lymphoblasts based upon cell diameter provides cell populations enriched in different stages of cell cycle. J. Immunol. Methods 1983, 63, 45–56. [Google Scholar] [CrossRef]
- Ersching, J.; Efeyan, A.; Mesin, L.; Jacobsen, J.T.; Pasqual, G.; Grabiner, B.C.; Dominguez-Sola, D.; Sabatini, D.M.; Victora, G.D. Germinal center selection and affinity maturation require dynamic regulation of mTORC1 kinase. Immunity 2017, 46, 1045–1058. [Google Scholar] [CrossRef]
- Davis, K. A Global Optimization Algorithm Worth Using dlib C++ Library. 2017. Available online: http://blog.dlib.net/2017/12/a-global-optimization-algorithm-worth.html (accessed on 1 March 2020).
- Snoek, J.; Larochelle, H.; Adams, R.P. Practical bayesian optimization of machine learning algorithms. In Proceedings of the Advances in Neural Information Processing Systems 25 (NIPS 2012), Lake Tahoe, NV, USA, 3–6 December 2012; pp. 2951–2959. [Google Scholar]
- Shen, C. Analysis of detrended time-lagged cross-correlation between two nonstationary time series. Phys. Lett. A 2015, 379, 680–687. [Google Scholar] [CrossRef]
- Zhang, Y.; Meyer-Hermann, M.; George, L.A.; Figge, M.T.; Khan, M.; Goodall, M.; Young, S.P.; Reynolds, A.; Falciani, F.; Waisman, A.; et al. Germinal center B cells govern their own fate via antibody feedback. J. Exp. Med. 2013, 210, 457–464. [Google Scholar] [CrossRef]
- Koch, A.L. The logarithm in biology 1. Mechanisms generating the log-normal distribution exactly. J. Theor. Biol. 1966, 12, 276–290. [Google Scholar] [CrossRef]
- Dowling, M.R.; Milutinović, D.; Hodgkin, P.D. Modelling cell lifespan and proliferation: Is likelihood to die or to divide independent of age? J. R. Soc. Interface 2005, 2, 517–526. [Google Scholar] [CrossRef][Green Version]
- Chung, Y.; Tanaka, S.; Chu, F.; Nurieva, R.I.; Martinez, G.J.; Rawal, S.; Wang, Y.H.; Lim, H.; Reynolds, J.M.; Zhou, X.H.; et al. Follicular regulatory T cells expressing Foxp3 and Bcl-6 suppress germinal center reactions. Nat. Med. 2011, 17, 983–988. [Google Scholar] [CrossRef] [PubMed]
- Maceiras, A.R.; Almeida, S.C.P.; Mariotti-Ferrandiz, E.; Chaara, W.; Jebbawi, F.; Six, A.; Hori, S.; Klatzmann, D.; Faro, J.; Graca, L. T follicular helper and T follicular regulatory cells have different TCR specificity. Nat. Commun. 2017, 8, 15067. [Google Scholar] [CrossRef] [PubMed]
- Sage, P.T.; Sharpe, A.H. T follicular regulatory cells. Immunol. Rev. 2016, 271, 246–259. [Google Scholar] [CrossRef]
- Oskooei, A.; Manica, M.; Mathis, R.; Martínez, M.R. Network-based Biased Tree Ensembles (NetBiTE) for Drug Sensitivity Prediction and Drug Sensitivity Biomarker Identification in Cancer. Sci. Rep. 2019, 9, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Manica, M.; Oskooei, A.; Born, J.; Subramanian, V.; Sáez-Rodríguez, J.; Rodríguez Martínez, M. Toward Explainable Anticancer Compound Sensitivity Prediction via Multimodal Attention-Based Convolutional Encoders. Mol. Pharm. 2019, 16. [Google Scholar] [CrossRef] [PubMed]
- Born, J.; Manica, M.; Oskooei, A.; Cadow, J.; Rodríguez Martínez, M. PaccMannRL: Designing Anticancer Drugs From Transcriptomic Data via Reinforcement Learning. In Research in Computational Molecular Biology; Lecture Notes in Computer Science; Schwartz, R., Ed.; Springer International Publishing: Cham, Switzerland, 2020; pp. 231–233. [Google Scholar] [CrossRef]
Code Availibility: The experimental data and python code used to fit our model, are provided freely on Github under MIT license at https://github.ibm.com/SysBio/Germinal-Center. The GC kinetic data of [18] were derived from the online database at http://sysimmtools.eu, and GC B cell sequences used to construct trees were made publicly available by [15]. The normalized dominance score [15] and MBC/PC output with time [50] were kindly provided by the corresponding authors of the respective studies.
|
| Reaction | Description | Parameter Value |
|---|---|---|
| B cell activation | ||
where each daughter CB undergoes apoptosis () with probability per acquired mutation. | Centroblast division | |
| Centroblast migration to LZ | ||
| Centrocyte apoptosis | ||
| Centrocyte antigen uptake | ||
| Centrocyte binding to T | ||
| Centrocyte T switch | ||
| Centrocyte spontaneous unbinding | ||
| Centrocyte recirculation to DZ | ||
with probability , with probability . | Centrocyte exit | see Section 2.7 for descriptions of and |
| Parameter | Description | Lower Bound | Upper Bound | Fitted | Sensitivity | Mainly Affects | |
|---|---|---|---|---|---|---|---|
| Intercellular | [] | B cell activation rate | 1 | 10 | 3.94 | high | Value of GC peak |
| [] | CB division rate | 0.08 | 0.16 | 0.134 | moderate | DZ/LZ ratio | |
| [] | CB migration rate | 0.15 | 3.75 | 3.75 | low | DZ/LZ ratio | |
| [] | CC apoptosis rate | 0.06 | 0.16 | 0.084 | high | GC decay | |
| [] | CC exit rate | 0.41 | 3.75 | 1.6 | high | GC decay | |
| [] | CC recirculation rate | fixed to 3.75 | high | GC decay | |||
| [] | FDC encounter rate | fixed to | low | Clonal competition | |||
| [] | T encounter rate | fixed to | moderate | Clonal competition | |||
| [] | T unbinding rate | fixed to 2 | high | GC decay | |||
| T to CC ratio | 1/100 | 1/7 | 1/46 | high | GC decay | ||
| Number of FDCs | fixed to 250 | low | Clonal competition | ||||
| Intracellular | Lethal mutation probability | 0.1 | 0.9 | 0.52 | high | GC decay | |
| Silent mutation probability | fixed to 0.28 | low | Affinity maturation | ||||
| Mutation rate | fixed to | high | GC decay | ||||
| CDR length (nucleotides) | fixed to 25 | low | Affinity maturation | ||||
| PC/MBC | Model 1 parameter | 0 | 1 | 0.46 | moderate | PC/MBC production | |
| Model 2 parameter | 0 | ∞ | 3 days | moderate | PC/MBC production | ||
| Model 2 parameter | 0 | 1 | 0.05 | moderate | PC/MBC production | ||
| k | Model 3 parameter | 0 | ∞ | 14 | moderate | PC/MBC production | |
| Model 3 parameter | 0 | 1 | 0.4 | moderate | PC/MBC production | ||
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pélissier, A.; Akrout, Y.; Jahn , K.; Kuipers , J.; Klein , U.; Beerenwinkel, N.; Rodríguez Martínez , M. Computational Model Reveals a Stochastic Mechanism behind Germinal Center Clonal Bursts. Cells 2020, 9, 1448. https://doi.org/10.3390/cells9061448
Pélissier A, Akrout Y, Jahn K, Kuipers J, Klein U, Beerenwinkel N, Rodríguez Martínez M. Computational Model Reveals a Stochastic Mechanism behind Germinal Center Clonal Bursts. Cells. 2020; 9(6):1448. https://doi.org/10.3390/cells9061448
Chicago/Turabian StylePélissier, Aurélien, Youcef Akrout, Katharina Jahn , Jack Kuipers , Ulf Klein , Niko Beerenwinkel, and María Rodríguez Martínez . 2020. "Computational Model Reveals a Stochastic Mechanism behind Germinal Center Clonal Bursts" Cells 9, no. 6: 1448. https://doi.org/10.3390/cells9061448
APA StylePélissier, A., Akrout, Y., Jahn , K., Kuipers , J., Klein , U., Beerenwinkel, N., & Rodríguez Martínez , M. (2020). Computational Model Reveals a Stochastic Mechanism behind Germinal Center Clonal Bursts. Cells, 9(6), 1448. https://doi.org/10.3390/cells9061448





