Genome-Wide Identification of the BZR Gene Family and Expression Validation of VvBZR7 in Grape (Vitis vinifera L.)
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Materials
2.2. Identification and Analysis of Physicochemical Properties of the Grape BZR Gene
2.3. Phylogenetic Tree Construction, Conserved and Gene Structure Analysis
2.4. Chromosome Localization and Colinearity Analysis
2.5. Cis-Acting Element Analysis
2.6. Grape RNA Extraction and qRT-PCR
2.7. Transcriptome Analysis
2.8. Subcellular Localization Analysis
2.9. Construction of VvBZR7 Overexpression Vectors and Transient Transformation of Grapevine Berry
3. Results
3.1. Identification and Characterization of the BZR Gene Family in Grape
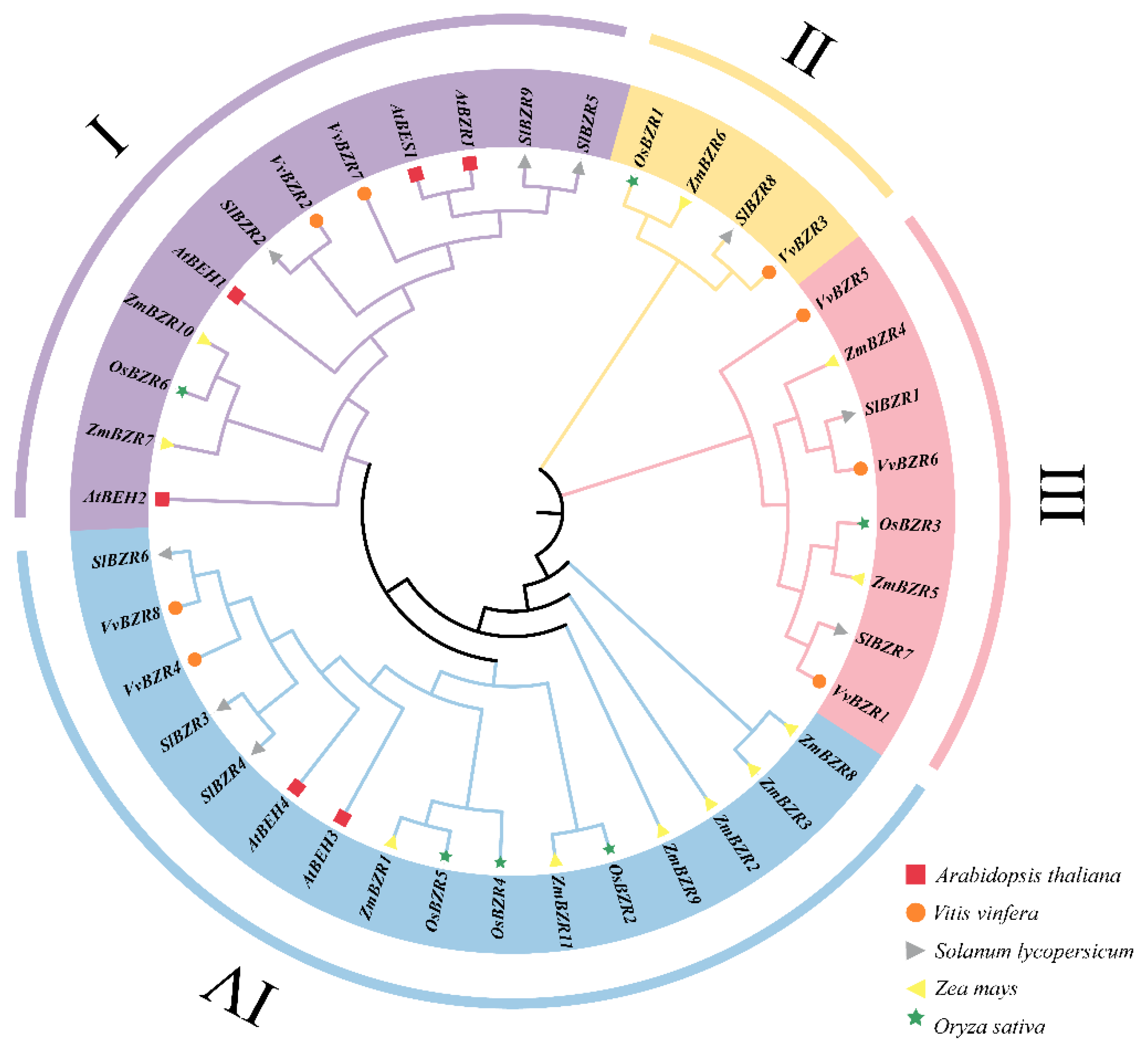
3.2. Phylogenetic Analysis of VvBZR Genes
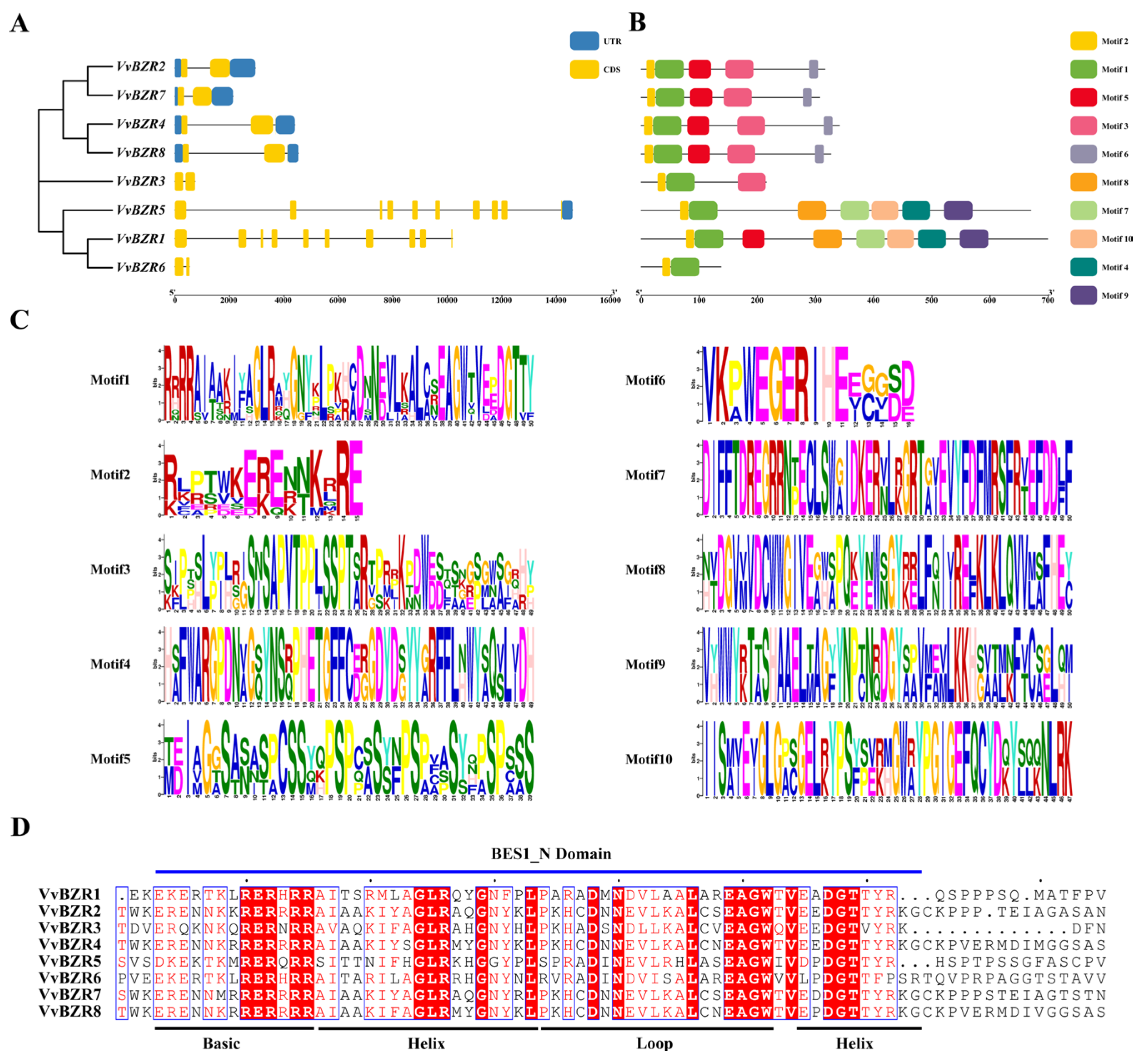
3.3. Gene Structure and Conserved Motif Analysis of VvBZR Genes
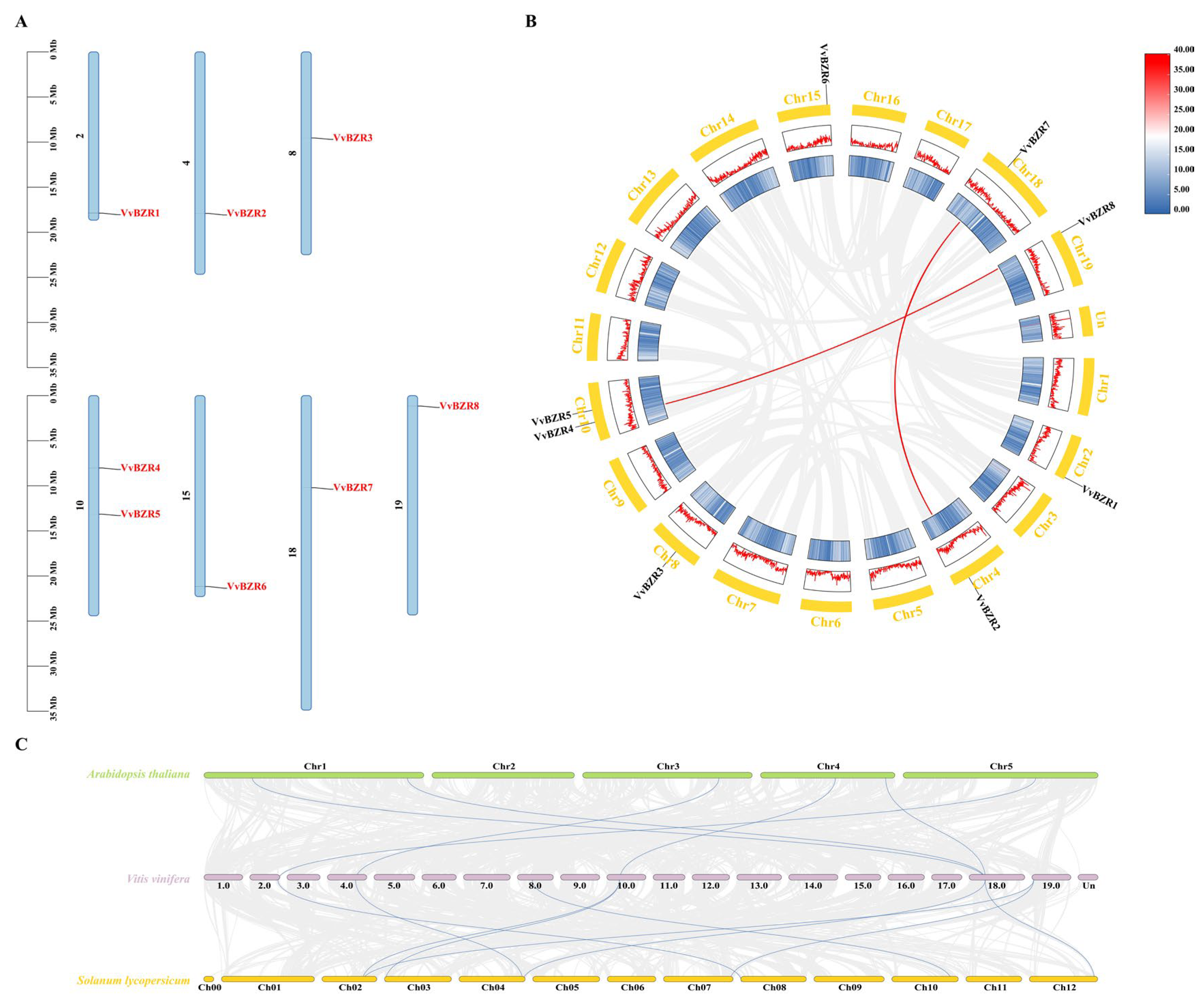
3.4. Distribution on Chromosomes and Collinear Analysis of VvBZR Proteins
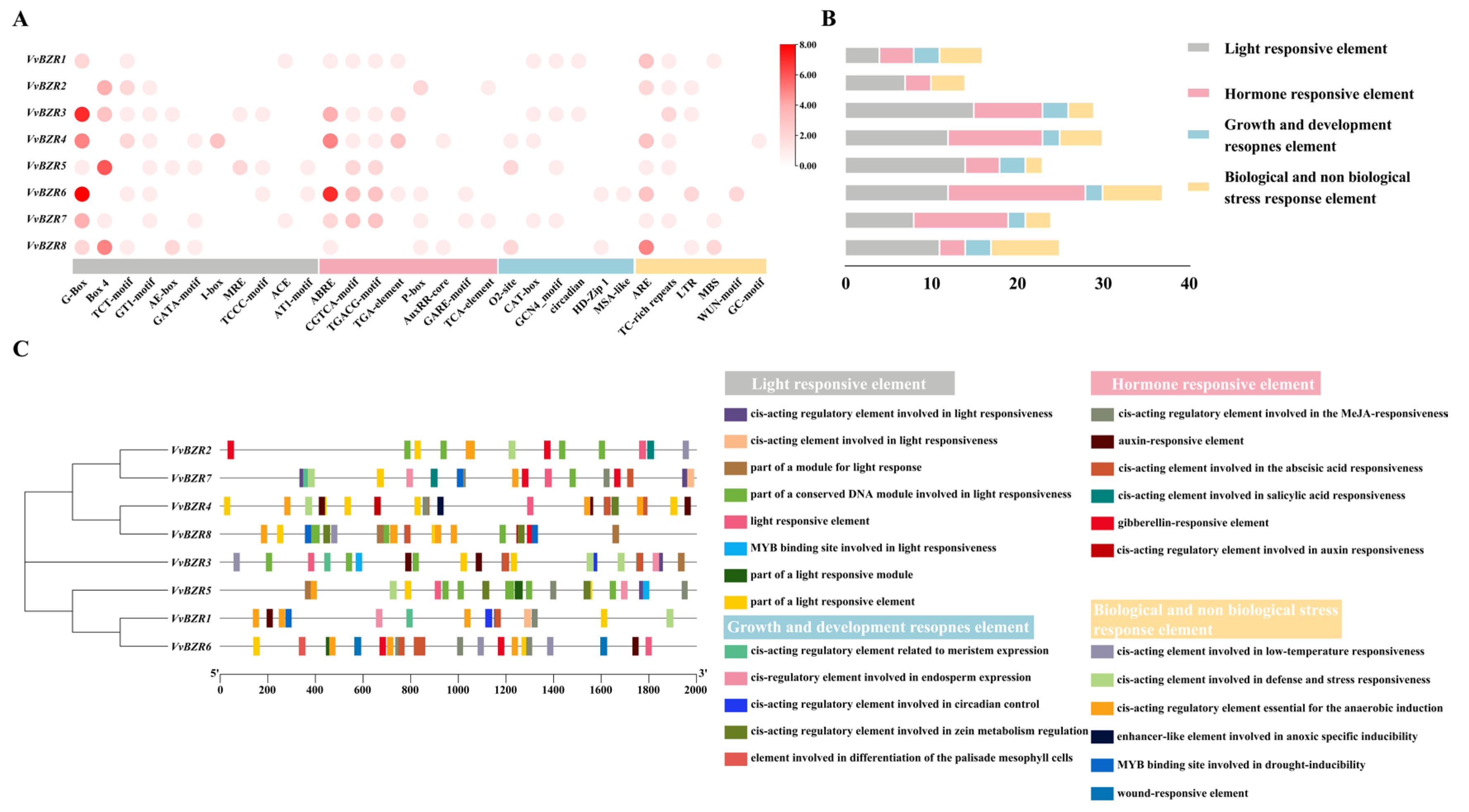
3.5. Analysis of Cis-Regulatory Element in the Promoter Region of VvBZRs
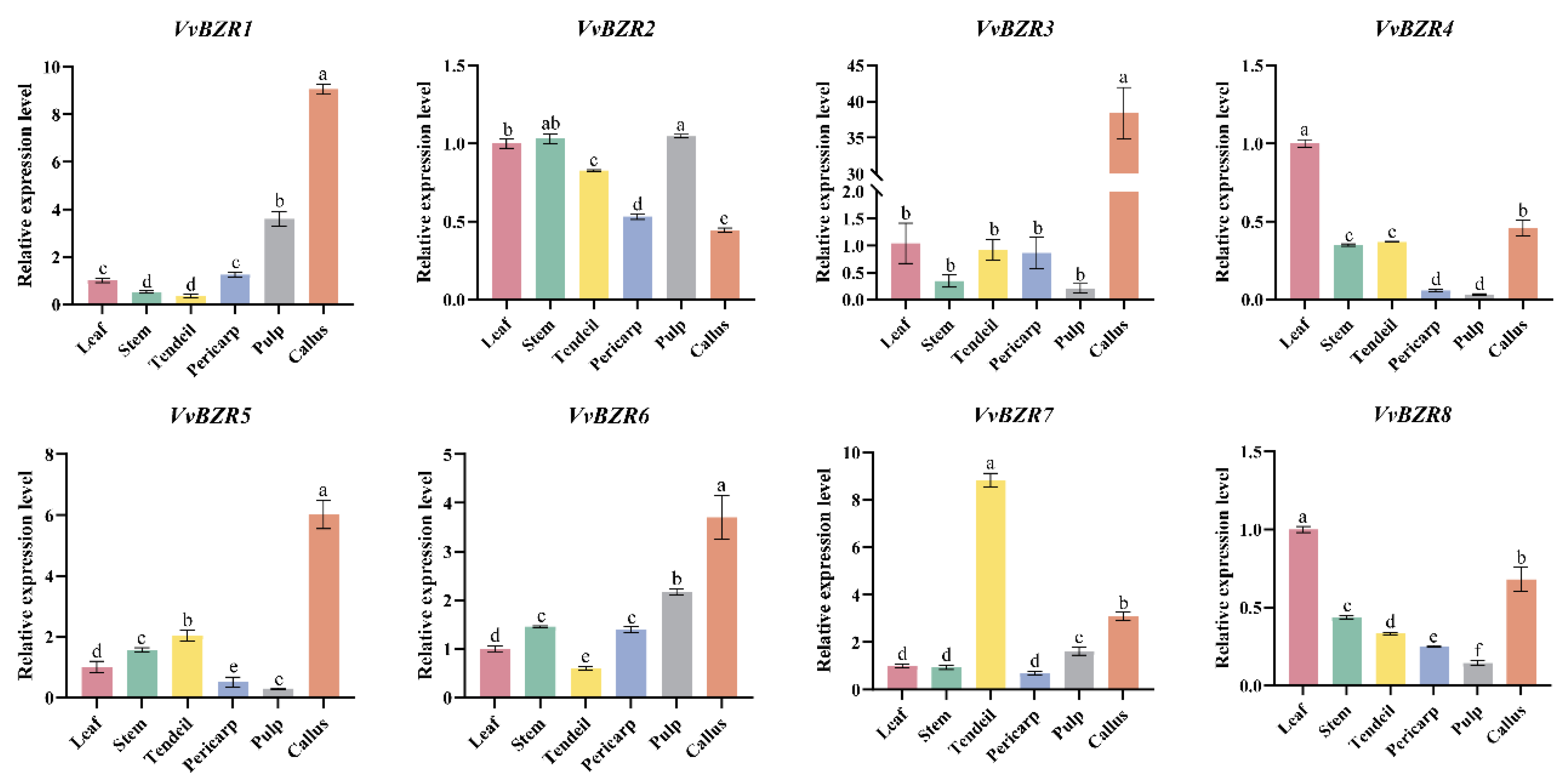
3.6. Analysis of Relative Gene Expression of VvBZRs Genes in Various Tissues
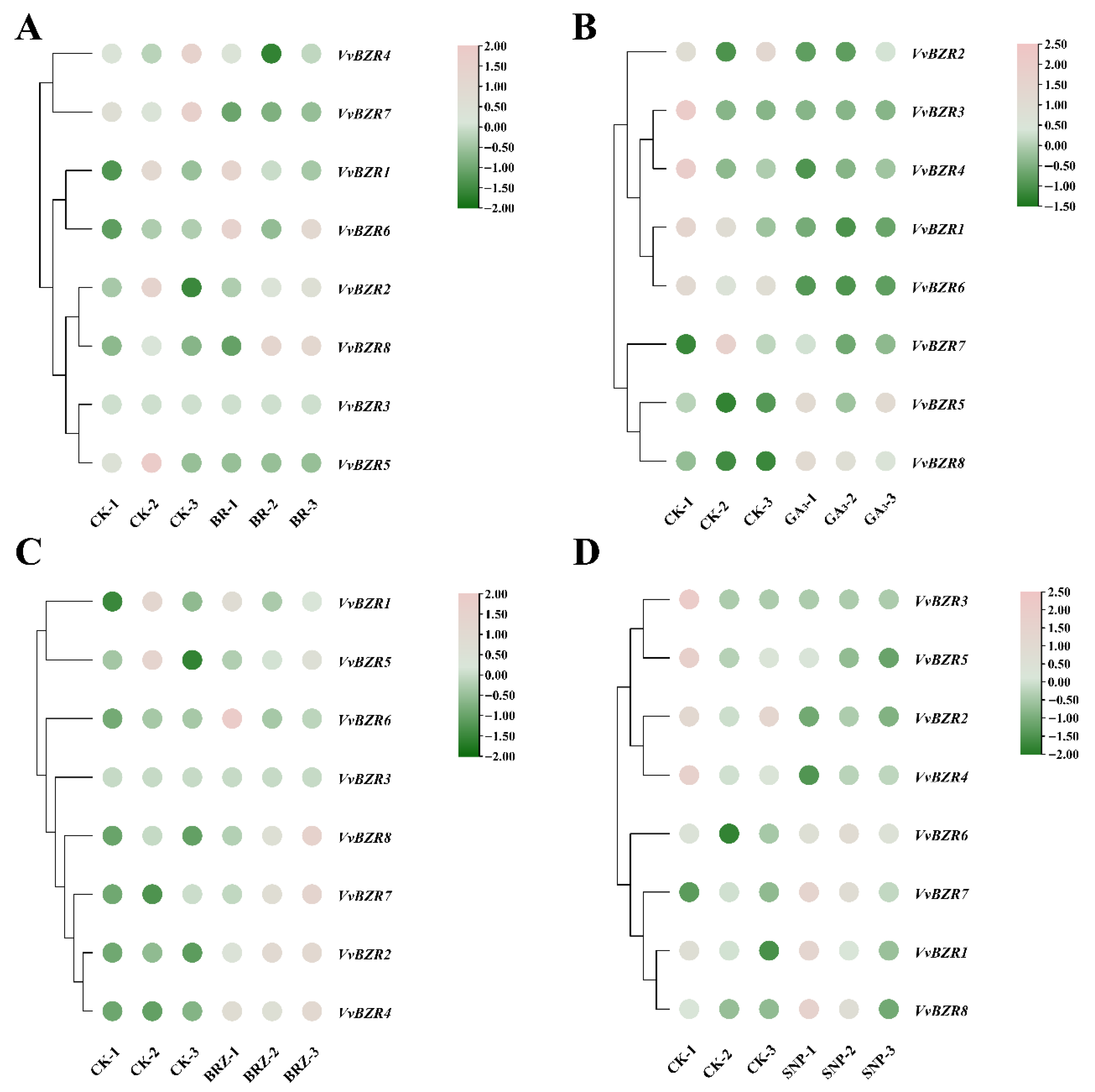
3.7. Expression Pattern of VvBZR Genes Under Hormone Treatment
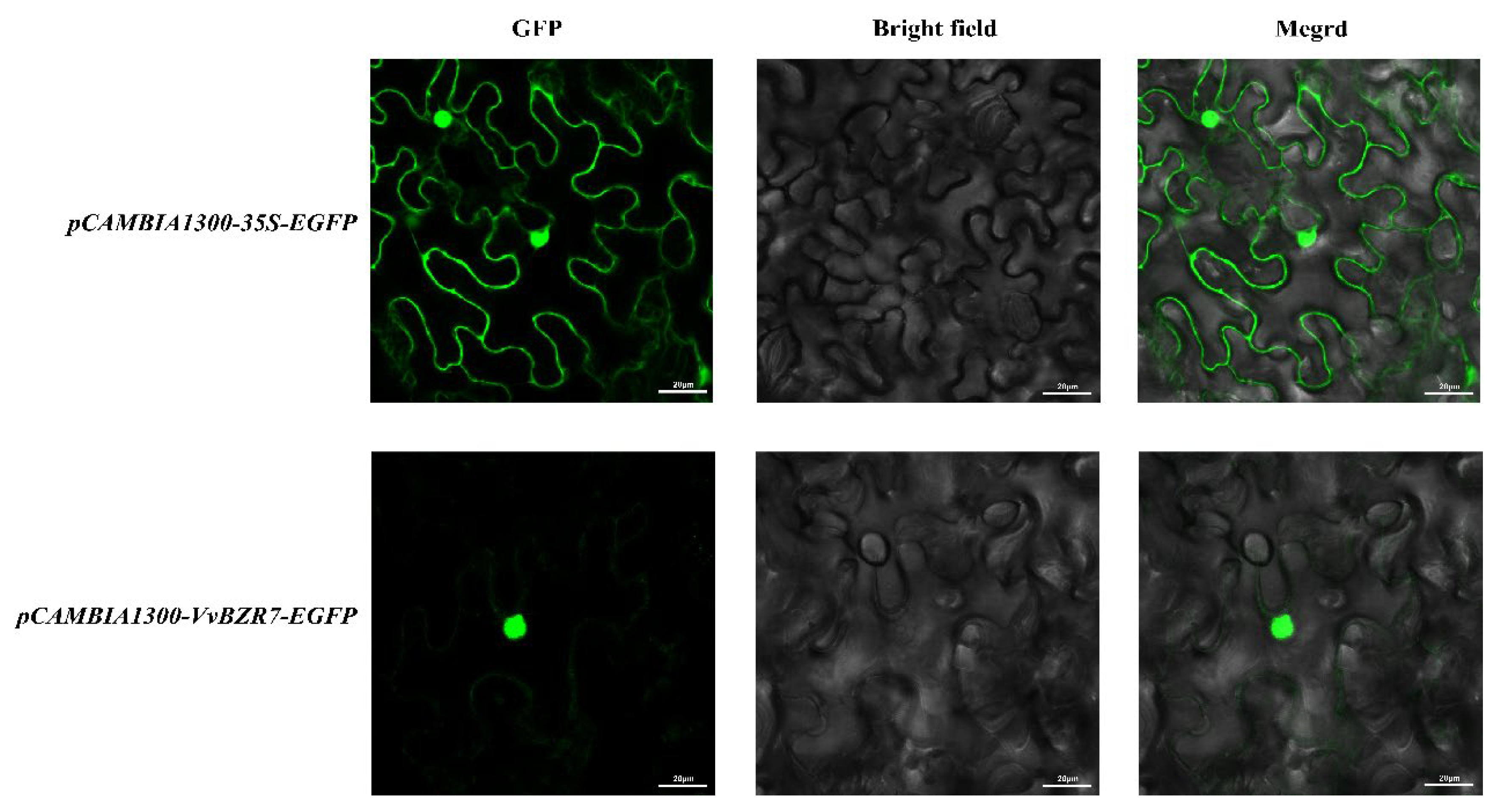
3.8. Subcellular Localization of the VvBZR7 Protein
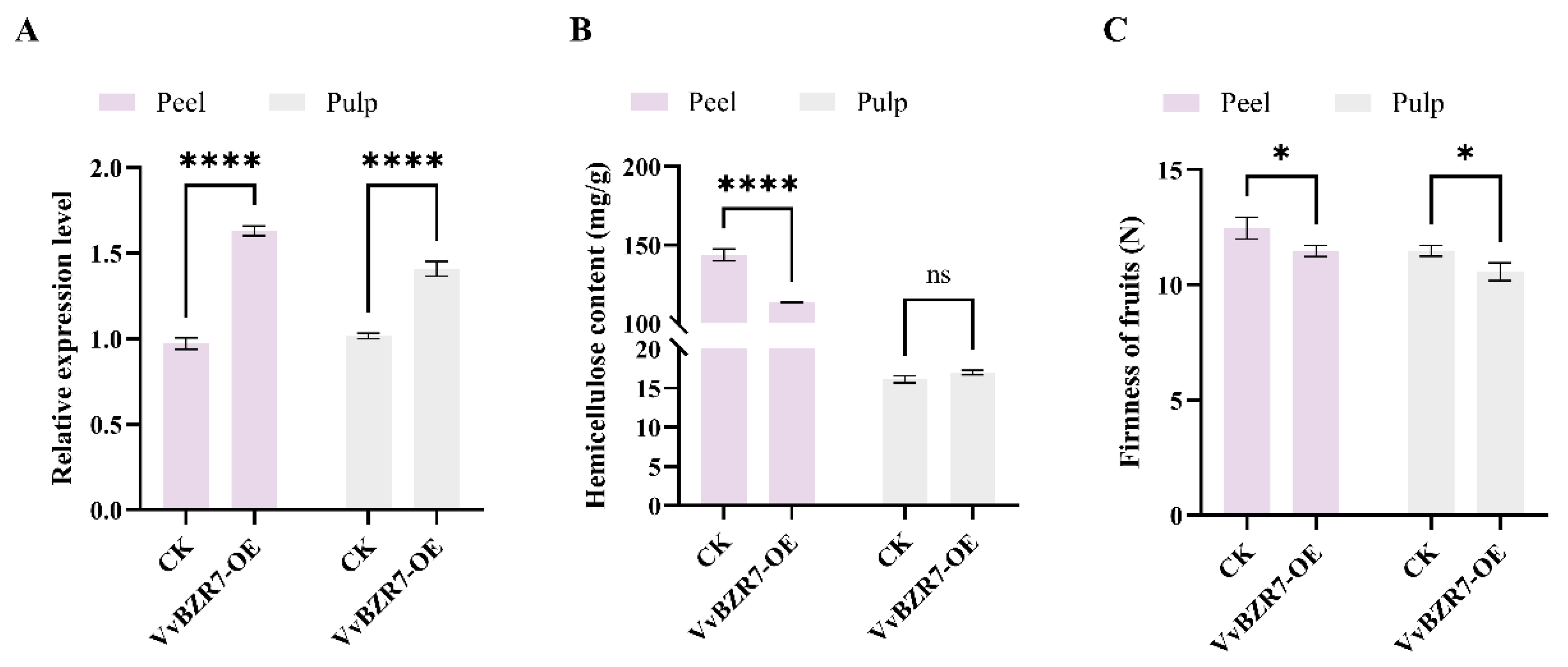
3.9. Transient Overexpression of VvBZR7 in Grapes
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Qiao, Z.; Li, J.; Zhang, X.; Guo, H.; He, C.; Zong, D. Genome-Wide Identification, Expression Analysis, and Subcellular Localization of DET2 Gene Family in Populus yunnanensis. Genes 2024, 15, 148. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Quan, Y.; Wang, L.; Wang, S. Brassinosteroid Promotes Grape Berry Quality-Focus on Physicochemical Qualities and Their Coordination with Enzymatic and Molecular Processes: A Review. Int. J. Mol. Sci. 2023, 24, 445. [Google Scholar] [CrossRef] [PubMed]
- Luo, S.; Zhang, G.; Zhang, Z.; Wan, Z.; Liu, Z.; Lv, J.; Yu, J. Genome-wide identification and expression analysis of BZR gene family and associated responses to abiotic stresses in cucumber (Cucumis sativus L.). BMC Plant Biol. 2023, 23, 214. [Google Scholar] [CrossRef] [PubMed]
- Zhao, N.; Zhao, M.; Tian, Y.; Wang, Y.; Han, C.; Fan, M.; Guo, H.; Bai, M.-Y. Interaction between BZR1 and EIN3 mediates signalling crosstalk between brassinosteroids and ethylene. New Phytol. 2021, 232, 2308–2323. [Google Scholar] [CrossRef]
- Yang, R.; Liu, P.; Zhang, T.; Dong, H.; Jing, Y.; Yang, Z.; Tang, S.; Zhang, Y.; Lv, M.; Liu, J.; et al. Plant-specific BLISTER interacts with kinase BIN2 and BRASSINAZOLE RESISTANT1 during skotomorphogenesis. Plant Physiol. 2023, 193, 1580–1596. [Google Scholar] [CrossRef]
- Khan, M.; Luo, B.; Hu, M.; Fu, S.; Liu, J.; Jiang, M.; Zhao, Y.; Huang, S.; Wang, S.; Wang, X. Brassinosteroid Signaling Downstream Suppressor BIN2 Interacts with SLFRIGIDA-LIKE to Induce Early Flowering in Tomato. Int. J. Mol. Sci. 2022, 23, 11264. [Google Scholar] [CrossRef]
- Huang, H.-Y.; Jiang, W.-B.; Hu, Y.-W.; Wu, P.; Zhu, J.-Y.; Liang, W.-Q.; Wang, Z.-Y.; Lin, W.-H. BR Signal Influences Arabidopsis Ovule and Seed Number through Regulating Related Genes Expression by BZR1. Mol. Plant 2013, 6, 456–469. [Google Scholar] [CrossRef]
- Zhou, P.; Jiang, H.; Li, J.; Jin, Q.; Wang, Y.; Xu, Y. Genome-Wide Identification Reveals That BZR1 Family Transcription Factors Involved in Hormones and Abiotic Stresses Response of Lotus (Nelumbo). Horticulturae 2023, 9, 882. [Google Scholar] [CrossRef]
- Talaat, N.B.; Ibrahim, A.S.; Shawky, B.T. Enhancement of the Expression of ZmBZR1 and ZmBES1 Regulatory Genes and Antioxidant Defense Genes Triggers Water Stress Mitigation in Maize (Zea mays L.) Plants Treated with 24-Epibrassinolide in Combination with Spermine. Agronomy 2022, 12, 2517. [Google Scholar] [CrossRef]
- Gao, A.; Weng, W.; Yao, X.; Wu, W.; Bai, Q.; Xiong, R.; Ma, C.; Cheng, J.; Ruan, J. Genome-Wide Identification, Structural Characterization, and Gene Expression Analysis of BES1 Transcription Factor Family in Tartary Buckwheat (Fagopyrum tataricum). Agronomy 2022, 12, 2729. [Google Scholar] [CrossRef]
- Kim, T.-W.; Guan, S.; Sun, Y.; Deng, Z.; Tang, W.; Shang, J.-X.; Sun, Y.; Burlingame, A.L.; Wang, Z.-Y. Brassinosteroid signal transduction from cell-surface receptor kinases to nuclear transcription factors. Nat. Cell Biol. 2009, 11, 1254–1260. [Google Scholar] [CrossRef]
- Clouse, S.D. Brassinosteroid Signal Transduction: From Receptor Kinase Activation to Transcriptional Networks Regulating Plant Development. Plant Cell 2011, 23, 1219–1230. [Google Scholar] [CrossRef]
- Qi, G.; Chen, H.; Wang, D.; Zheng, H.; Tang, X.; Guo, Z.; Cheng, J.; Chen, J.; Wang, Y.; Bai, M.-Y.; et al. The BZR1-EDS1 module regulates plant growth-defense coordination. Mol. Plant 2021, 14, 2072–2087. [Google Scholar] [CrossRef]
- Li, L.; Mu, T.; Zhang, R.; Zhang, G.; Lyu, J.; Liu, Z.; Luo, S.; Yu, J. The BES1/BZR1 family transcription factor as critical regulator of plant stress resilience. Plant Stress 2025, 15, 100730. [Google Scholar] [CrossRef]
- Zhao, N.; Zhao, M.; Wang, L.; Han, C.; Bai, M.; Fan, M. EBF1 Negatively Regulates Brassinosteroid-Induced Apical Hook Development and Cell Elongation through Promoting BZR1 Degradation. Int. J. Mol. Sci. 2022, 23, 15889. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.-H.; Lee, S.-H.; Park, T.-K.; Tian, Y.; Yu, K.; Lee, B.-H.; Bai, M.-Y.; Cho, S.-J.; Kim, T.-W. Comparative analysis of BZR1/BES1 family transcription factors in Arabidopsis. Plant J. 2024, 117, 747–765. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.-H.; Kim, S.-H.; Park, T.-K.; Kim, Y.-P.; Lee, J.-W.; Kim, T.-W. Transcription factors BZR1 and PAP1 cooperate to promote anthocyanin biosynthesis in Arabidopsis shoots. Plant Cell 2024, 36, 3654–3673. [Google Scholar] [CrossRef] [PubMed]
- Ji, Y.; Qu, Y.; Jiang, Z.; Yan, J.; Chu, J.; Xu, M.; Su, X.; Yuan, H.; Wang, A. The mechanism for brassinosteroids suppressing climacteric fruit ripening. Plant Physiol. 2021, 185, 1875–1893. [Google Scholar] [CrossRef]
- Sun, F.; Ding, L.; Feng, W.; Cao, Y.; Lu, F.; Yang, Q.; Li, W.; Lu, Y.; Shabek, N.; Fu, F.; et al. Maize transcription factor ZmBES1/BZR1-5 positively regulates kernel size. J. Exp. Bot. 2021, 72, 1714–1726. [Google Scholar] [CrossRef]
- Yu, T.; Ai, G.; Xie, Q.; Wang, W.; Song, J.; Wang, J.; Tao, J.; Zhang, X.; Hong, Z.; Lu, Y.; et al. Regulation of tomato fruit elongation by transcription factor BZR1.7 through promotion of SUN gene expression. Hortic. Res. 2022, 9, uhac121. [Google Scholar]
- Zeng, J.; Li, Z.; Ding, X.; Liang, H.; Wu, K.; Jiang, Y.; Duan, X.; Jiang, G. The H3R2me2a demethylase JMJ10 regulates tomato fruit size through its interaction with the transcription factor BZR1.3. Plant Cell 2025, 37, koaf251. [Google Scholar] [CrossRef]
- He, Y.; Liu, H.; Li, H.; Jin, M.; Wang, X.; Yin, X.; Zhu, Q.; Rao, J. Transcription factors DkBZR1/2 regulate cell wall degradation genes and ethylene biosynthesis genes during persimmon fruit ripening. J. Exp. Bot. 2021, 72, 6437–6446. [Google Scholar] [CrossRef]
- Su, W.; Shao, Z.; Wang, M.; Gan, X.; Yang, X.; Lin, S. EjBZR1 represses fruit enlargement by binding to the EjCYP90 promoter in loquat. Hortic. Res. 2021, 8, 152. [Google Scholar] [CrossRef]
- Hu, L.; Zheng, Q.; Chen, Z.; Qin, Y.; Si, H.; Ji, J.; Li, Q.; Yang, Z.; Wu, Y. Pre-harvest treatment with gibberellin (GA3) and nitric oxide donor (SNP) enhances post-harvest firmness of grape berries. Food Chem. Mol. Sci. 2025, 10, 100235. [Google Scholar] [CrossRef] [PubMed]
- Ullah, U.; Shalmani, A.; Ilyas, M.; Raza, A.; Ahmad, S.; Shah, A.Z.; Khan, F.U.; Azizud, D.; Bibi, A.; Rehman, S.U.; et al. BZR proteins: Identification, evolutionary and expression analysis under various exogenous growth regulators in plants. Mol. Biol. Rep. 2022, 49, 12039–12053. [Google Scholar] [CrossRef] [PubMed]
- Wang, B.; Zhu, X.; Wei, X. Genome-wide identification, structural analysis, and expression profiles of the BZR gene family in tomato. J. Plant Biochem. Biotechnol. 2022, 31, 739–750. [Google Scholar] [CrossRef]
- Manoli, A.; Trevisan, S.; Quaggiotti, S.; Varotto, S. Identification and characterization of the BZR transcription factor family and its expression in response to abiotic stresses in Zea mays L. Plant Growth Regul. 2018, 84, 423–436. [Google Scholar] [CrossRef]
- Hu, L.; Xu, T.; Cai, Y.; Qin, Y.; Zheng, Q.; Chen, T.; Gong, L.; Yang, J.; Zhao, Y.; Chen, J.; et al. Identifying Candidate Genes for Grape (Vitis vinifera L.) Fruit Firmness through Genome-Wide Association Studies. J. Agric. Food Chem. 2025, 73, 8413–8425. [Google Scholar] [CrossRef]
- Chen, J.; Guo, Y.; Hu, H.; Fang, C.; Wang, L.; Hu, L.; Lin, Z.; Zhang, D.; Yang, Z.; Wu, Y. Regulation of Cell Metabolism and Changes in Berry Shape of Shine Muscat Grapevines Under the Influence of Different Treatments with the Plant Growth Regulators Gibberellin A3 and N-(2-Chloro-4-Pyridyl)-N′-Phenylurea. Horticulturae 2025, 11, 1160. [Google Scholar] [CrossRef]
- Zhao, R.; Cheng, J.; Yu, Y. Genome-Wide Identification and Characterization of the BZR Transcription Factor Gene Family in Leymus chinensis. Genes 2025, 16, 155. [Google Scholar] [CrossRef]
- Zuo, C.; Zhang, L.; Yan, X.; Guo, X.; Zhang, Q.; Li, S.; Li, Y.; Xu, W.; Song, X.; Wang, J.; et al. Evolutionary analysis and functional characterization of BZR1 gene family in celery revealed their conserved roles in brassinosteroid signaling. BMC Genom. 2022, 23, 568. [Google Scholar] [CrossRef] [PubMed]
- Lachowiec, J.; Mason, G.A.; Schultz, K.; Queitsch, C. Redundancy, Feedback, and Robustness in the Arabidopsis thaliana BZR/BEH Gene Family. Front. Genet. 2018, 9, 523. [Google Scholar] [CrossRef] [PubMed]
- Ullah, M.A.; Hamza, M.; Gull, R.; Shafiq, M.; Wahid, A.; Ahmad, S.; Ahmadi, T.; Rahimi, M. Genome-wide analysis of the BoBZR1 family genes and transcriptome analysis in Brassica oleracea. Sci. Rep. 2025, 15, 15475. [Google Scholar] [CrossRef] [PubMed]
- Kesawat, M.S.; Kherawat, B.S.; Singh, A.; Dey, P.; Kabi, M.; Debnath, D.; Saha, D.; Khandual, A.; Rout, S.; Manorama Ali, A.; et al. Genome-Wide Identification and Characterization of the Brassinazole-resistant (BZR) Gene Family and Its Expression in the Various Developmental Stage and Stress Conditions in Wheat (Triticum aestivum L.). Int. J. Mol. Sci. 2021, 22, 8743. [Google Scholar] [CrossRef]
- Sarwar, R.; Geng, R.; Li, L.; Shan, Y.; Zhu, K.-M.; Wang, J.; Tan, X.-L. Genome-Wide Prediction, Functional Divergence, and Characterization of Stress-Responsive BZR Transcription Factors in B. napus. Front. Plant Sci. 2022, 12, 790655. [Google Scholar] [CrossRef]
- Wei, X.; Tang, Y.; Liu, Y.; Shen, S.; Xu, J.; Chen, L.; Li, M.; Zhao, H.; Zhang, T.; Du, H.; et al. Genome-Wide Identification of the Glycosyl Hydrolase Family 1 Genes in Brassica napus L. and Functional Characterization of BnBGLU77. Plants 2025, 14, 2686. [Google Scholar] [CrossRef]
- Moon, J.; Park, C.-H.; Son, S.-H.; Youn, J.-H.; Kim, S.-K. Endogenous level of abscisic acid down-regulated by brassinosteroids signaling via BZR1 to control the growth of Arabidopsis thaliana. Plant Signal. Behav. 2021, 16, 1926130. [Google Scholar] [CrossRef]
- Yang, X.; Bai, Y.; Shang, J.; Xin, R.; Tang, W. The antagonistic regulation of abscisic acid-inhibited root growth by brassinosteroids is partially mediated via direct suppression of ABSCISIC ACID INSENSITIVE 5 expression by BRASSINAZOLE RESISTANT 1. Plant Cell Environ. 2016, 39, 1994–2003. [Google Scholar]
- Zhang, J.; Chen, W.; Li, X.; Shi, H.; Lv, M.; He, L.; Bai, W.; Cheng, S.; Chu, J.; He, K.; et al. Jasmonates regulate apical hook development by repressing brassinosteroid biosynthesis and signaling. Plant Physiol. 2023, 193, 1561–1579. [Google Scholar] [CrossRef]
- Wang, X.; Wei, J.; Wu, J.; Shi, B.; Wang, P.; Alabd, A.; Wang, D.; Gao, Y.; Ni, J.; Bai, S.; et al. Transcription factors BZR2/MYC2 modulate brassinosteroid and jasmonic acid crosstalk during pear dormancy. Plant Physiol. 2024, 194, 1794–1814. [Google Scholar]
- Xiong, M.; Yu, J.; Wang, J.; Gao, Q.; Huang, L.; Chen, C.; Zhang, C.; Fan, X.; Zhao, D.; Liu, Q.-Q.; et al. Brassinosteroids regulate rice seed germination through the BZR1-RAmy3D transcriptional module. Plant Physiol. 2022, 189, 402–418. [Google Scholar] [CrossRef]
- Yang, D.; Shin, H.-Y.; Kang, H.K.; Shang, Y.; Park, S.Y.; Jeong, D.-H.; Nam, K.H. Reciprocal inhibition of expression between RAV1 and BES1 modulates plant growth and development in Arabidopsis. J. Integr. Plant Biol. 2023, 65, 1226–1240. [Google Scholar] [CrossRef]
- Yan, T.; Shu, X.; Ning, C.; Li, Y.; Wang, Z.; Wang, T.; Zhuang, W. Functions and Regulatory Mechanisms of bHLH Transcription Factors during the Responses to Biotic and Abiotic Stresses in Woody Plants. Plants 2024, 13, 2315. [Google Scholar] [CrossRef]
- Yang, J.; Wu, Y.; Li, L.; Li, C. Comprehensive analysis of the BES1 gene family and its expression under abiotic stress and hormone treatment in Populus trichocarpa. Plant Physiol. Biochem. 2022, 173, 1–13. [Google Scholar] [CrossRef]
- Cao, X.; Wei, Y.; Shen, B.; Liu, L.; Mao, J. Interaction of the Transcription Factors BES1/BZR1 in Plant Growth and Stress Response. Int. J. Mol. Sci. 2024, 25, 6836. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; Jia, C.; Zhang, M.; Chen, D.; Chen, S.; Guo, R.; Guo, D.; Wang, Q. Ectopic expression of a BZR1-1D transcription factor in brassinosteroid signalling enhances carotenoid accumulation and fruit quality attributes in tomato. Plant Biotechnol. J. 2014, 12, 105–115. [Google Scholar] [CrossRef]
- He, S.; Xia, X.; Yang, J.; Xin, J.; Chen, S.; Jia, C. Overexpression of AtBES1D in tomato enhances BR response and accelerates fruit ripening. J. Plant Physiol. 2025, 312, 154563. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.; Yang, Y.; Yu, W.; Liu, L.; Hu, Q.; Wei, W.; Liu, J. Dissecting the Genetic Mechanisms of Hemicellulose Content in Rapeseed Stalk. Agronomy 2022, 12, 2886. [Google Scholar] [CrossRef]
- Liu, L.; Feng, Y.; Han, Z.; Song, Y.; Guo, J.; Yu, J.; Wang, Z.; Wang, H.; Gao, H.; Yang, Y.; et al. Functional analysis of the xyloglucan endotransglycosylase/hydrolase gene MdXTH2 in apple fruit firmness formation. J. Integr. Agric. 2025, 24, 3418–3434. [Google Scholar] [CrossRef]
- Khodayari, A.; Thielemans, W.; Hirn, U.; Van Vuure, A.W.; Seveno, D. Cellulose-hemicellulose interactions—A nanoscale view. Carbohydr. Polym. 2021, 270, 118364. [Google Scholar] [CrossRef]
| Gene ID | Gene Name | Protein Length (aa) | Instability Index | MW (Da) | pI | Aliphatic Index | GRAVY | Subcellular Location |
|---|---|---|---|---|---|---|---|---|
| Vitvi02g01232 | VvBZR1 | 670 | 41.51 | 75.30 | 5.59 | 67.00 | −0.528 | Nucleus |
| Vitvi04g01234 | VvBZR2 | 316 | 70.80 | 33.93 | 9.04 | 57.50 | −0.631 | Nucleus |
| Vitvi08g00772 | VvBZR3 | 137 | 21.19 | 15.54 | 9.15 | 56.20 | −0.992 | Nucleus |
| Vitvi10g00636 | VvBZR4 | 341 | 55.31 | 36.29 | 8.53 | 49.91 | −0.611 | Nucleus |
| Vitvi10g01901 | VvBZR5 | 215 | 60.31 | 22.41 | 9.44 | 56.74 | −0.523 | Nucleus |
| Vitvi15g01128 | VvBZR6 | 699 | 41.35 | 78.78 | 5.58 | 74.88 | −0.444 | Nucleus |
| Vitvi18g00924 | VvBZR7 | 307 | 60.97 | 33.65 | 9.13 | 60.81 | −0.669 | Nucleus |
| Vitvi19g00061 | VvBZR8 | 326 | 58.4 | 34.88 | 8.39 | 56.38 | −0.548 | Nucleus |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zheng, Q.; Hu, L.; Hu, H.; Fang, C.; Wang, L.; Zhou, Y.; Xu, Y.; Chen, Z.; Wu, Y.; Yang, Z. Genome-Wide Identification of the BZR Gene Family and Expression Validation of VvBZR7 in Grape (Vitis vinifera L.). Agronomy 2025, 15, 2749. https://doi.org/10.3390/agronomy15122749
Zheng Q, Hu L, Hu H, Fang C, Wang L, Zhou Y, Xu Y, Chen Z, Wu Y, Yang Z. Genome-Wide Identification of the BZR Gene Family and Expression Validation of VvBZR7 in Grape (Vitis vinifera L.). Agronomy. 2025; 15(12):2749. https://doi.org/10.3390/agronomy15122749
Chicago/Turabian StyleZheng, Qianqian, Lingling Hu, Haichao Hu, Congling Fang, Liru Wang, Yuxuan Zhou, Yangshengkai Xu, Zhihui Chen, Yueyan Wu, and Zhongyi Yang. 2025. "Genome-Wide Identification of the BZR Gene Family and Expression Validation of VvBZR7 in Grape (Vitis vinifera L.)" Agronomy 15, no. 12: 2749. https://doi.org/10.3390/agronomy15122749
APA StyleZheng, Q., Hu, L., Hu, H., Fang, C., Wang, L., Zhou, Y., Xu, Y., Chen, Z., Wu, Y., & Yang, Z. (2025). Genome-Wide Identification of the BZR Gene Family and Expression Validation of VvBZR7 in Grape (Vitis vinifera L.). Agronomy, 15(12), 2749. https://doi.org/10.3390/agronomy15122749






