Abstract
Nontyphoidal Salmonella (NTS) is a zoonotic pathogen that threatens public health worldwide. This study investigated the prevalence, serotype, virulence, and antimicrobial resistance of NTS isolated from chicken meat in Fukuoka, Japan. Of 50 samples, 64% were positive for Salmonella spp., and 32 NTS strains were isolated from positive samples. Serotyping identified three serotypes: S. enterica ser. Schwarzengrund (78.1%), S. enterica ser. Thompson (15.6%), and S. enterica ser. Oranienburg (6.3%). Multilocus sequence typing revealed three sequence types (STs), and MALDI-TOF MS analysis revealed six distinct clusters, reflecting heterogeneity in protein expression among isolates with the same STs. All isolates harbored the virulence genes hilA, spiC, and ssrB, but not spvC. Microplate assays showed that all S. enterica ser. Schwarzengrund and S. enterica ser. Thompson strains formed biofilms with varying strengths. Antimicrobial susceptibility tests demonstrated that S. enterica ser. Thompson and S. enterica ser. Oranienburg strains were sensitive to all the antimicrobials tested. However, S. enterica ser. Schwarzengrund strains showed resistance to multiple antibiotic classes, and 36% of the isolates were multidrug resistant. These findings suggest a potential public health concern, particularly from S. enterica ser. Schwarzengrund, and underscore the importance of continuous surveillance that integrates both genotypic and phenotypic methods.
1. Introduction
The threat of illness and death due to diseases associated with contaminated food poses a persistent danger to human health and significantly hinders global socioeconomic progress. Foodborne disease outbreaks are prevalent and often result in substantial morbidity and mortality [1]. Salmonella is a key foodborne pathogen that causes acute gastroenteritis in humans. It comprises two species, S. enterica and S. bongori, with over 2500 serotypes identified to date [2]. Based on the clinical symptoms observed in humans, Salmonella is classified into typhoidal and nontyphoidal Salmonella (NTS). Various animals serve as hosts for NTS, which are associated with severe and invasive illnesses in immunocompromised adults [3]. However, the primary source of infections caused by NTS is chicken products such as chicken meat, eggs, and confectionery containing eggs [4]. In Japan, retail chicken meat is reported to be more contaminated with Salmonella than beef or pork [5]. Because the consumption of raw chicken breast sashimi, known as Torisashi in Japanese, is common in the Kyushu Islands, including Fukuoka [6], it may pose a zoonotic threat to humans. Furthermore, in Japan, the average number of patients per Salmonella outbreak is 38, although annual cases for foodborne salmonellosis have declined to less than 50 since 2011. This average is approximately five times higher than that of the Campylobacter outbreak [7]. Therefore, monitoring the prevalence of NTS in chicken meat is essential for understanding its public health risks.
Serotyping has been vital for monitoring human health concerning salmonellosis for over 50 years [8]. According to the Kauffmann–White classification, conventional serological typing identifies somatic O and flagellar H antigens by observing bacterial clumping in the presence of specific antisera. However, traditional serotyping has several drawbacks: it is time-consuming and labor-intensive, requires numerous antisera, and fails to distinguish between rough and mucoid strains [9]. Molecular serotyping using multiplex PCR has recently advanced Salmonella serotyping, facilitating the confirmation and rapid typing of important serotypes [10]. Furthermore, multilocus sequence typing (MLST), which is based on the sequencing of seven housekeeping genes, is a robust tool for Salmonella surveillance. It can immediately predict the Salmonella serotype [11,12]. Matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI-TOF MS) is widely used for bacterial typing at genus, species, and strain levels. In addition to identification, MALDI-TOF biotyper software enables the direct creation of dendrograms that illustrate the evolutionary relationships between bacterial strains using protein fingerprint data [13].
Determinants of bacterial virulence are crucial for systemic infection. Pathogenic Salmonella strains are associated with various virulence genes located on Salmonella pathogenicity islands (SPIs) [14]. To date, 23 SPIs have been identified. SPI-1 and SPI-2 are critical for Salmonella virulence and encode a type III secretion system (T3SS) that is essential for invasion into a host cell and intracellular survival of the bacterium by delivering the effector proteins [15]. Genes such as invA and hilA in SPI-1 enhance epithelial cell invasion, with invA also serving as a detection biomarker [16]. In SPI-2, spiC and ssrB aid in intracellular survival and infection [17,18]. Additionally, virulence plasmids containing spvRABCD promote rapid proliferation and survival within host cells, which are crucial for systemic infection [19].
Antimicrobial resistance (AMR) is a rising concern in Salmonella infections that affect the health of both animals and humans [20]. Recently, there has been a growing prevalence of multidrug-resistant (MDR) Salmonella and resistance to essential antimicrobials, such as third-generation cephalosporins and fluoroquinolones, globally, including Japan [21,22,23]. This drastically affects the use of antibiotics in the treatment of human salmonellosis. Therefore, the continuous surveillance of antibiotic resistance in Salmonella is critical for understanding resistance trends and implementing effective control measures and medical interventions. Furthermore, Salmonella is classified as a high-priority pathogen on the WHO list of antibiotic-resistant bacteria urgently requiring new antimicrobials [24].
However, only a few studies have reported on antimicrobial-resistant Salmonella from chickens in Fukuoka, Japan, before 2015 [21,22]. Moreover, recent reports have documented outbreaks linked to uncommon serotypes in the southern Kyushu area [25], which is on the same island as Fukuoka. Therefore, due to the public health importance of antibiotic-resistant Salmonella, we aimed to investigate the prevalence of NTS in chicken meat from Fukuoka, Japan, identify the serotype distribution, evaluate virulence potential, assess biofilm formation, and examine the antimicrobial resistance of these isolates.
2. Materials and Methods
2.1. Sampling
The study was conducted between December 2023 and August 2024. Fifty chicken samples (skin, liver, heart, wing, breast, gizzard, thigh, breast tenderloin, and minced meat) from different brands were randomly purchased as regular consumer products from ten retail shops in Fukuoka, Japan. The identities of retailers were not recorded, and all samples were examined anonymously for research purposes. Fresh chicken samples refer to raw, chilled (not frozen) retail products that are maintained refrigerated in the shops. At least three fresh raw chicken samples were collected from each retail shop. All samples were transported on ice to the laboratory for Salmonella detection within 24 h.
2.2. Detection of Salmonella spp.
Salmonella was isolated using the standard protocols outlined in ISO 6579:2002 [26]. Briefly, each sample (25 g) was placed in Stomacher® bags with 225 mL of Buffer Peptone Water (Nissui Pharmaceutical Co., Ltd., Tokyo, Japan). The sample was then homogenized using a stomacher (Masticator, IUL, Barcelona, Spain) for 60 s and incubated at 37 °C for 18 h. Each enriched sample (0.1 mL) was pipetted into 10 mL of GranuCult™ Rappaport-Vassiliadis-Soya (RVS) broth (Merck KGaA, Darmstadt, Germany) and incubated at 42 °C for 24 h. A loopful of RVS broth culture was streaked onto xylose-lysine deoxycholate (XLD) agar (CM0469B; Oxoid Ltd., Basingstoke, UK) and CHROMagar™ Salmonella (CHROMagar, Paris, France). Following the incubation at 37 °C for 24 h, the presumptive colonies (red with black centers on XLD and mauve on CHROMagar) were selected, sub-cultured onto tryptic soy agar (TSA, Becton Dickinson and Co., Franklin Lakes, NJ, USA), and subjected to molecular confirmation.
2.3. Molecular Characterization of Salmonella spp.
2.3.1. DNA Template Preparation
A portion of the single colony on TSA was suspended in 40 µL of double-sterilized water (dsH2O). The suspension (1 µL) was used as a DNA template for colony PCR. Salmonella Typhimurium NBRC 12529, obtained from the Biological Resource Center (Chiba, Japan), was used as a positive control for colony PCR.
2.3.2. PCR Amplification
Salmonella spp. were confirmed using colony PCR, which amplified the invA gene using specific primers (Table S1). The PCR was performed in a 10 μL reaction, consisting of 5 µL of 2× GoTaq® Green Master Mix (Promega Corp., Madison, WI, USA), 1 μL of invA-F and invA-R primers (10 pmol each), and 2 μL of dsH2O. The PCR conditions for amplification included initial denaturation at 95 °C for 3 min, followed by 35 cycles of denaturation at 95 °C for 30 s, annealing at 60 °C for 30 s, and extension at 72 °C for 30 s, with a final extension at 72 °C for 5 min. Colonies of Salmonella enterica serovar Typhimurium (S. enterica ser. Typhimurium) NBRC 12529 and distilled water were used as positive and negative controls, respectively.
2.3.3. Gel Electrophoresis
The PCR products were separated on a 1.5% agarose gel containing Midori Green Advance DNA stain (Nippon Genetics Europe GmbH, Düren, Germany) at 100 V for 23 min. The bands were visualized using WSE-6100H LuminoGraph I (ATTO, Tokyo, Japan). A 100 bp DNA ladder (SMOBIO, Inc., Hsinchu City, Taiwan) was used as a size standard to estimate the size of the PCR product.
2.4. Serotyping of Salmonella by Traditional and Molecular Methods
2.4.1. Traditional Serotyping
Salmonella isolates were serotyped by slide agglutination using commercial monovalent Salmonella O antisera (Denka Seikan Co., Ltd., Tokyo, Japan) in accordance with the manufacturer’s instructions. All the strains were cultured overnight on TSA at 37 °C. The somatic O antigen was confirmed using antisera. S. enterica ser. Typhimurium NBRC 12529 and S. enterica ser. Schwarzengrund (Lab. strain) [27] were used as positive controls, and normal saline served as the negative control.
2.4.2. Molecular Serotyping
Molecular serotyping was performed on all Salmonella isolates using multiplex colony PCR with primer sets targeting the common O and H serogroups (Table S1). The DNA template was prepared as described in Section 2.3.1. The somatic O-serogrouping assay was conducted in a 12.5 μL reaction mixture, containing 6.25 μL GoTaq® Green Master Mix (Promega Corp, Madison, WI, USA), 0.25 μL each of 6 forward and reverse primers, 1 μL DNA, and 2.25 μL dsH2O. For the flagellar H phase 1 assay, the 12.5 μL reaction consisted of 6.25 μL GoTaq® Green Master Mix, 0.5 μL of 3 forward and 0.25 μL of 7 reverse primers, 1 μL DNA, and 2 μL dsH2O. The flagellar H phase 2 assay was also conducted in a 12.5 μL volume with similar components: 6.25 μL GoTaq® Green Master Mix, 0.5 μL 3 forward and 6 reverse primers, 1 μL DNA, and 0.75 μL dsH2O. Table S2 lists the PCR conditions used for somatic O and flagellar H antigen assays.
All assays included positive controls (DNA templates prepared from laboratory stocks of S. enterica ser. Schwarzengrund and/or S. enterica ser. Thompson) and negative controls (distilled water). The PCR products were analyzed as described in Section 2.3.3, using a 50 bp DNA ladder (SMOBIO, Inc.) as a size standard. Serotypes were identified based on the banding patterns of each Salmonella isolate. The antigenic formula of each isolate was determined by the WHO Collaborating Center for Reference and Research on Salmonella, and serovars were assigned based on the White–Kauffmann–Le Minor scheme [9].
Serogroup O:7 was identified using a serotype-specific primer set for S. Thompson (F-TATTGCAACAATGAGGCCCTCT, R-GTCGCGATTCTGAACGTGTC). The PCR mix and gel electrophoresis followed the protocol in Section 2.3.2 and Section 2.3.3. PCR conditions were as follows: 95 °C for 5 min, followed by 35 cycles of 95 °C for 5 s, 60 °C for 5 s, and 72 °C for 5 s.
2.5. MLST of the Isolates
MLST was performed by amplifying and sequencing seven housekeeping genes (aroC, dnaN, hemD, hisD, purE, sucA, and thrA), as described by a previous study [28]. Briefly, 32 Salmonella isolates were cultured on TSA at 37 °C for 24 h. Bacterial DNA was extracted using a Cica Geneus DNA Extraction Kit (Kanto Chemical Co., Inc., Tokyo, Japan). PCR was done in 50 μL consisting of 25 μL GoTaq® Green Master Mix, 5 μL forward and reverse primers, 1 μL DNA, and 14 μL dsH2O. PCR amplification of the seven housekeeping genes was performed using the primer sets under the cycling conditions outlined in Table S2. The PCR products were purified using the FastGene™ Gel/PCR extraction kit (Nippon Genetics Co., Ltd., Tokyo, Japan) and sequenced through the Sanger method (GeneWiz, Azenta Life Sciences, Tokyo, Japan). Each obtained DNA sequence was aligned and compared to those in the MLST database (https://pubmlst.org/organisms/salmonella-spp) (accessed on 5 June 2025), and then assigned to a specific allele. The sequence type (ST) of the isolate was determined using allelic combinations of the seven genes. Allelic profiles were displayed in the following format: aroC-dnaN-hemD-hisD-purE-sucA-thrA.
2.6. MALDI-TOF MS Analysis
Thirty-two Salmonella isolates (S. enterica ser. Schwarzengrund [n = 25], S. enterica ser. Thompson [n = 5], and S. enterica ser. Oranienburg [n = 2]) were identified using a MALDI Biotyper (Bruker Daltonics GmbH, Bremen, Germany). The isolates were incubated on TSA at 37 °C for 24 h. Subsequently, single colonies were suspended in 300 μL of pure water and vortexed for 30 s, after which 900 μL of ethanol was added to the mixture. The samples were kept at −20 °C until analysis by MALDI-TOF MS at Kyushu Sangyo University (Fukuoka, Japan).
After receiving the MALDI-TOF MS data, the MALDI Biotyper classifications were interpreted as follows: based on the criteria suggested by the manufacturer, scores above 2.000 indicate accurate species identification, scores between 1.700 and 1.999 indicate genus-level identification, and scores below 1.700 indicate no reliable identification [13]. MALDI-TOF MS analysis was performed on one sample from each isolate in triplicate. Dendrograms were created using strains with scores greater than 2.000. A principal component analysis (PCA) dendrogram was generated using Biotyper software (version 4.0) to assess the similarity level of the MALDI-TOF MS profiles among the tested isolates.
2.7. Detection of Virulence Genes by PCR
A total of 32 Salmonella isolates were tested for the presence of virulence factors by targeting the hilA, spiC, ssrB, and spvC genes using simplex colony PCR. Each PCR reaction had a final volume of 10 μL, including 5 μL GoTaq® Green Master Mix (Promega Corp., Madison, WI, USA), 1 μL forward and reverse primers (10 pmol each), 1 μL DNA template prepared as described in Section 2.3.1, and 2 μL dsH2O. The PCR products were analyzed by agarose gel electrophoresis, as described in Section 2.3.3. Tables S2 and S3 show the primers and PCR conditions used to detect each virulence gene.
2.8. Detection of Biofilm Formation Ability
Biofilm formation was assessed using a microtiter plate assay as described by Miyamoto et al. [29]. Briefly, overnight bacterial cultures were diluted in LB broth supplemented with 0.2% BactoTM Soytone (LB-S) (Difco Laboratories, Detroit, MI, USA) to an OD of 0.7 at 660 nm. A 200 μL cell suspension was pipetted into individual wells of a 96-well flat-bottom polystyrene microplate (Violamo, Osaka, Japan). Sterile LB-S broth was also added to the wells as a negative control to check for sterility and nonspecific binding of the media. Following incubation at 30 °C for 24 h, the liquid in the wells was gently aspirated and washed twice with phosphate-buffered saline (PBS; pH 7.2) to remove buoyant cells before being air-dried at room temperature (RT). The plates were stained with 200 μL of 0.1% crystal violet (CV) for 30 min at RT and then rinsed four times with PBS to eliminate any stain residue. Subsequently, the biofilm-bound CV was solubilized in 200 μL of 99% ethanol for 20 min. To evaluate the ability to form biofilms, 100 μL of the ethanol extract was taken and added to a fresh microtiter plate, and the OD was recorded at 595 nm using an Infinite F50/Robotic microplate reader (Tecan Trading AG, Männedorf, Switzerland). Each strain was tested in triplicate in two independent experiments. The OD value of the negative control was used as the cut-off value (ODn). The average absorbance from the triplicate readings was utilized to assess each strain’s final optical density (ODf), which was then compared to the negative control (ODn). Isolates were defined as non-biofilm-forming if ODf ≤ ODn, weak biofilm-forming if ODn < ODf ≤ 2× ODn, moderate biofilm-forming if 2× ODn < ODf ≤ 4× ODn, or strong biofilm-forming if ODf > 4× ODn [30].
2.9. Antimicrobial Susceptibility Testing of Planktonic Salmonella Isolates
The susceptibilities of 32 Salmonella isolates (S. enterica ser. Schwarzengrund [n = 25], S. enterica ser. Thompson [n = 5], and S. enterica ser. Oranienburg [n =2]) were evaluated using broth microdilution and disk diffusion methods in accordance with the guidelines from M100 of the Clinical and Laboratory Standards Institute [31]. Microdilution was conducted in a 96-well DP 31 dry plate (Eiken Chemicals, Tokyo, Japan) containing 18 antimicrobials at various concentrations: piperacillin (PIPC, 0.5–65 µg/mL), cefazolin (CEZ, 0.5–16 µg/mL), cefotiam (CTM, 0.5–16 µg/mL), cefotaxime (CTX, 0.5–32 µg/mL), ceftazidime (CAZ, 0.5–16 µg/mL), cefepime (CFPM, 0.5–16 µg/mL), flomoxef (FMOX, 0.5–16 µg/mL), cefpodoxime (CPDX, 1–4 µg/mL), sulbactam/ampicillin (SA, 2/4–8/16 µg/mL), aztreonam (AZT, 0.5–16 µg/mL), imipenem (IPM, 0.25–8 µg/mL), meropenem (MEPM, 0.25–8 µg/mL), gentamicin (GM, 0.25–8 µg/mL), amikacin (AMK, 1–32 µg/mL), minocycline (MINO, 0.25–8 µg/mL), fosfomycin (FOM, 32–128 µg/mL), sulfamethoxazole/trimethoprim (ST, 9.5/0.5–38/2 µg/mL), and levofloxacin (LVFX, 0.12–2 µg/mL). Ciprofloxacin (CIP, 0.06–2 µg/mL), which was not included in DP31, was tested separately using the microdilution method. The isolates were cultured on TSA at 37 °C overnight. The bacterial inoculum in 0.85% sodium chloride was adjusted to approximately 0.5 McFarland turbidity standard, with OD600 nm of ~0.1. The bacterial cell suspension (25 μL) was mixed with 12 mL of Mueller–Hinton broth to obtain an inoculum concentration of 5 × 104 CFU/mL. After a 15 min interval, the cell suspension (100 μL) was transferred to each well of the 96-well plate and incubated at 37 °C for 24 h. A bacterial suspension was added to the wells without antibiotics as a control. After incubation, the absorbance was recorded using a microplate reader. Antibiotic susceptibility was determined by interpreting the MIC values in accordance with the CLSI guidelines [31].
The isolates were evaluated for resistance to antimicrobial agents that were not included in the DP 31 dry plate. Antibiotic disks (Eiken Chemicals, Tokyo, Japan), including ampicillin (ABP, 10 µg), kanamycin (KM, 30 µg), streptomycin (S, 10 µg), tetracycline (TC, 30 µg), nalidixic acid (NA, 30 µg), ciprofloxacin (CIP, 5 µg), and chloramphenicol (C, 30 µg), were used for disc diffusion. The bacterial inoculum was prepared using a microdilution method. The bacterial suspension was streaked onto Mueller–Hinton agar (Betcon Dickinson and Co., Franklin Lakes, NJ, USA) using a sterile cotton swab. After allowing the plates to dry for a few minutes, antimicrobial disks were placed and then incubated at 37 °C for 24 h. Following incubation, the diameter of the inhibition zone was measured and then interpreted as susceptible, intermediate, or resistant by the breakpoints for Enterobacteriaceae [30]. Isolates that exhibited resistance to at least one antimicrobial agent in three or more antimicrobial categories were defined as MDR [13]. Staphylococcus aureus JCM 2413, derived from ATCC 25923, was used as a quality control strain.
2.10. Antimicrobial Susceptibility Testing of Biofilm-Forming Salmonella Isolates
The susceptibilities of bacteria grown in biofilm conditions were assessed according to the method as described by do Canto Canabarro et al. [32], with minor modifications. A total of 18 isolates (S. enterica ser. Schwarzengrund [n = 16] and S. enterica ser. Thompson [n = 2]) representing diverse antimicrobial resistance profiles and varying biofilm-forming abilities (weak, moderate, and strong) were selected. Antibiotics from different classes and mechanisms of action were selected from those tested in the planktonic microbroth dilution assay.
Biofilms of all selected strains were established in 96-well microtiter plates as described above in Section 2.8. After biofilm formation, planktonic cells were removed, and the residual biofilms were gently washed three times with PBS and then air-dried. Next, 200 μL of two-fold serial dilutions of antimicrobials in cation adjusted Mueller–Hinton broth (CAMHB) was added to each well with the preformed biofilm: piperacillin (PIPC, 0.5–512 µg/mL) (Sigma-Aldrich, Inc., St. Louis, MO, USA), cefotaxime (CTX, 0.5–512 µg/mL) (FUJIFILM Wako Pure Chemical Corporation, Osaka, Japan), aztreonam (AZT, 0.5–512 µg/mL) (Tokyo Chemical Industry Co., Ltd., Tokyo, Japan), meropenem (MEPM, 0.5–512 µg/mL) (FUJIFILM Wako Pure Chemical Corporation, Osaka, Japan), gentamicin (GM, 0.5–512 µg/mL) (Nacalai Tesque, Kyoto, Japan), minocycline (MINO, 0.5–512 µg/mL) (FUJIFILM Wako Pure Chemical Corporation, Osaka, Japan), fosfomycin (FOM, 0.5–512 µg/mL) (FUJIFILM Wako Pure Chemical Corporation, Osaka, Japan), sulfamethoxazole/trimethoprim (ST, 9.5/0.5–608/32 µg/mL) (FUJIFILM Wako Pure Chemical Corporation, Osaka, Japan) and ciprofloxacin (CIP, 0.5–512 µg/mL) (Tokyo Chemical Industry Co., Ltd., Tokyo, Japan). CAMHB without antibiotics served as the positive control, while CAMHB with the corresponding antibiotics at each concentration was used as the negative control. The plates were incubated at 30 °C for 24 h. To determine the viability of the biofilm after antimicrobial exposure, 10 μL of 20 μM resazurin (FUJIFILM Wako Pure Chemical Corporation, Osaka, Japan) was added to each well, and the mixture was incubated at 37 °C in the dark for 1 h. Fluorescence intensity was measured at an excitation/emission wavelength of 540/590 using a Nano Infinite microplate reader (Tecan, Männedorf, Switzerland) [33]. The minimal biofilm-inhibition concentration (MBIC) was defined as the lowest antimicrobial concentration that causes at least an 80% reduction in biofilm metabolic activity relative to the positive control, after subtracting background fluorescence from the negative control [32]. This experiment was performed in triplicate.
2.11. Detection of Genes Associated with Antimicrobial Resistance
The major genes associated with intermediate resistance and resistance to tetracycline (tetA), sulfamethoxazole/trimethoprim (sul1 and sul2), streptomycin (strA/strB and aadA1), kanamycin (aphA1 and aadB), and the plasmid-mediated quinolone resistant (PMQR) genes (qnrA, qnrB, and qnrS) were screened by PCR. The primer pairs used to detect the resistance genes are listed in Table S4. Based on the phenotypic resistance profile, the isolates were selected and cultured overnight on TSA at 37 °C. Following this, the DNA template and the PCR reaction (10 μL) were prepared as outlined in Section 2.3.2. Table S5 lists the PCR conditions used for the detection of each resistance gene. The PCR products were analyzed by agarose gel electrophoresis, as described in Section 2.3.3. Additionally, mutations in the quinolone resistance-determining region (QRDR) of the gyrA gene were identified by sequencing the PCR product. PCR products were purified using the FastGene™ Gel/PCR extraction kit (Nippon Genetics Co., Ltd., Tokyo, Japan), followed by Sanger sequencing (GeneWiz, Azenta Life Sciences, Tokyo, Japan).
3. Results
3.1. Prevalence of Salmonella spp. and Serotype Distribution in Chicken Meat in Fukuoka
Of the 50 chicken samples collected, 32 (64%) were positive for Salmonella spp., resulting in 32 isolates. At least one sample from each of the ten retail shops in Fukuoka tested positive for Salmonella. Among these, 30 were successfully identified as S. enterica ser. Schwarzengrund (n = 25) or S. enterica ser. Thompson (n = 5). Although the remaining two isolates were serotyped as the O6,7 serogroup and G complex H1 serotype by PCR, they could not be assigned to a specific serotype. Thus, the predominant serotype was S. enterica ser. Schwarzengrund (25/32, 78.1%), followed by S. enterica ser. Thompson (5/32, 15.6%) and an unidentified serotype (2/32, 6.3%) (Table 1).

Table 1.
Nontyphoidal Salmonella strains isolated from chicken meats identified by PCR and serotyping.
3.2. Sequence Type (ST) of the Isolates
Thirty-two strains isolated from chicken meat were categorized into three STs. ST241 (25/32, 78.1%) was the most predominant, followed by ST26 (5/32, 15.6%) and ST23 (2/32, 6.3%) (Table 2). All the S. enterica ser. Schwarzengrund isolates belonged to ST241, and S. enterica ser. Thompson to ST26. Two unidentified serotypes belonged to ST23. These isolates were putatively assigned to S. enterica ser. Oranienburg based on ST23 association in the PubMLST database (https://pubmlst.org/organisms/salmonella-spp) (accessed on 5 June 2025).

Table 2.
MLST results of the isolates.
3.3. MALDI-TOF MS Analysis of the Isolates
All 32 Salmonella isolates were accurately identified to the species level, and the best matching scores ranged from 2.20 to 2.39. Table S9 presents the best-matched score values of the isolates identified using MALDI-TOF MS.
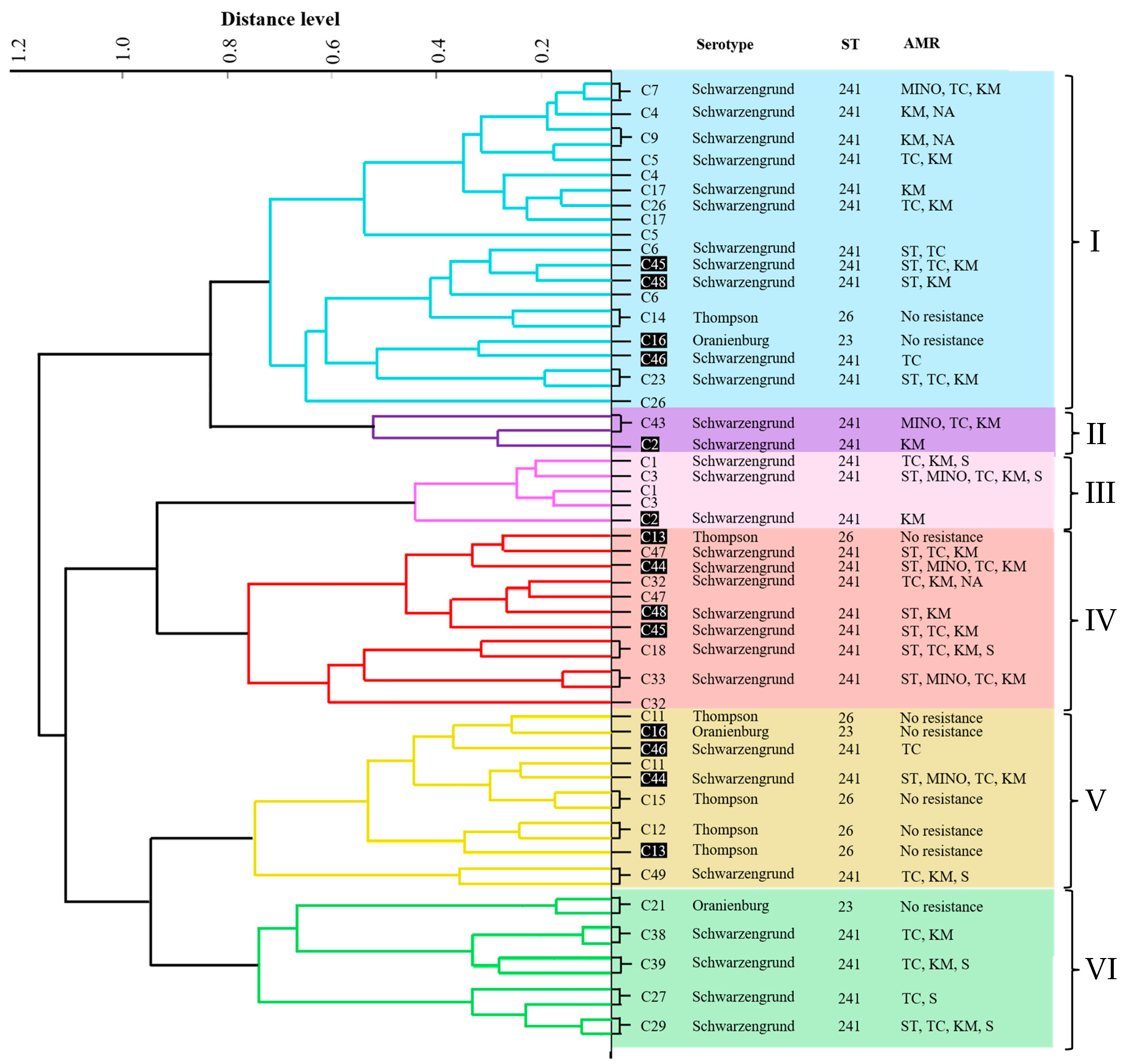
Figure 1 shows the PCA dendrogram of all the MALDI-TOF MS profiles of the isolates. The isolates were divided into six clusters (I–VI). Cluster I contained nine isolates (28.1%). Isolates C4, C9, C5, and C26 exhibited identical serotypes, STs, AMR patterns, virulence genes, and biofilm formation capabilities. Cluster II consisted of a single isolate (3.1%). C1 and C3 isolates shared the same serotype, ST, virulence gene, and biofilm-forming ability, and were grouped in cluster III (6.3%). Cluster IV consisted of four isolates (12.5%), all of which had the same serotype, ST, and virulence genes. Cluster V contained four isolates (12.5%). The isolates C11, C12, and C15 shared similar serotypes, STs, and virulence genes. Cluster VI comprised five isolates (15.6%), with isolates C27, C29, C38, and C39 showing identical serotypes, ST, and virulence genes. However, the two MALDI-TOF MS profiles of each isolate (C2, C13, C16, C44, C45, C46, and C48) were distributed into separate clusters in the PCA dendrogram.
Figure 1.
PCA dendrogram of 32 NTS isolates created from MALDI-TOF MS patterns and the characteristics of the isolates. The clusters are labeled I–VI. Isolates not grouped into the same cluster by PCA are displayed in white letters on a black background. Distance level is displayed in arbitrary units (a.u.).
3.4. Genes Encoding Virulence Factors in the Isolates
All isolates harbored hilA, spiC, and ssrB genes, except for one S. enterica ser. Schwarzengrund isolate that lacked the spiC gene (Table 3). In contrast, the spvC gene was not detected in any isolates (Table 3).

Table 3.
Prevalence of virulence gene among nontyphoidal Salmonella isolates.
3.5. Biofilm Formation Ability of Salmonella Isolates
Biofilm formation of 32 isolates (S. enterica ser. Schwarzengrund [n = 25], S. enterica ser. Thompson [n = 5], and S. enterica ser. Oranienburg [n = 2]) was quantitatively evaluated after 1 day of incubation. Among the 25 S. enterica ser. Schwarzengrund isolates, 22 (88%) showed strong biofilm formation, while three (12%) exhibited moderate biofilm formation. In contrast, one (20%) of the five S. enterica ser. Thompson isolates showed moderate biofilm formation, and four (80%) displayed weak biofilm formation. No biofilm formation was observed in any of the S. enterica ser. Oranienburg isolates. The indices for biofilm formation (BF) and related categories of each isolate are presented in Table 4. Representative crystal violet-stained image showing biofilm formation by Salmonella isolates is shown in Figure S1.

Table 4.
Biofilm formation ability of nontyphoidal Salmonella isolates.
3.6. Antimicrobial Susceptibility of Planktonic Salmonella Isolates
The antimicrobial susceptibilities of planktonic Salmonella isolates are presented in Table 5, and corresponding MIC ranges for antimicrobial agents determined by broth microdilution are shown in Table S10. In this study, all S. enterica ser. Thompson and S. enterica ser. Oranienburg isolates were susceptible to all the tested antimicrobials, with intermediate resistance to minocycline (1/5, 20%) and ciprofloxacin (3/5, 60%) in S. enterica ser. Thompson. In contrast, 22/25 (88%) of S. enterica ser. Schwarzengrund were resistant to kanamycin, 20/25 (80%) to tetracycline, 10/25 (40%) to sulfamethoxazole/trimethoprim, 7/25 (28%) to streptomycin, 5/25 (20%) to minocycline, and 3/25 (12%) to nalidixic acid. In addition, 16 of the 25 (64%) S. enterica ser. Schwarzengrund strains showed intermediate resistance to ciprofloxacin, which is recommended as a critical therapeutic agent for salmonellosis.

Table 5.
Antimicrobial susceptibility of planktonic nontyphoidal Salmonella isolates from chicken meat by microbroth dilution and disc diffusion methods.
Table 6 presents the MDR profiles of S. enterica ser. Schwarzengrund isolates. Of the 25 S. enterica ser. Schwarzengrund isolates, nine (36%) were identified as MDR, showing resistance to at least three to five antibiotics with different mechanisms and exhibiting six distinct resistance patterns.

Table 6.
MDR profile of S. enterica ser. Schwarzengrund isolates from chicken meat.
3.7. Antimicrobial Susceptibility of Biofilm Salmonella Isolates
Table S11 presents the MIC and MBIC values for tested antimicrobial agents against biofilm-forming isolates. In general, biofilm cells exhibited markedly reduced susceptibility compared to their planktonic cells. Most isolates showed very high MBIC values (>512 µg/mL) for piperacillin, cefotaxime, aztreonam, meropenem, gentamicin, and fosfomycin, as well as for sulfamethoxazole/trimethoprim (>608/32 µg/mL). However, most isolates exhibited the lowest MBIC values for minocycline (16–64 µg/mL) and ciprofloxacin (0.5–2 µg/mL) among the tested agents, although these values remained higher than the corresponding MICs (Table S11). A representative image of biofilm cell viability after antimicrobial treatment, determined by resazurin assay, is shown in Figure S2.
3.8. Detection of Genes Associated with Antimicrobial Resistance by PCR
The prevalence of genes involved in antimicrobial resistance is shown in Table 7. None of the intermediate-resistant S. enterica ser. Thompson isolates to minocycline and ciprofloxacin possessed any of the resistance genes tested.

Table 7.
Detection of genes involved in antimicrobial resistance by PCR.
All S. enterica ser. Schwarzengrund isolates exhibited resistance to or intermediate resistance to tetracycline and sulfamethoxazole/trimethoprim and tested positive for the tetA and sul1 genes (100%). The aphA1 gene was positive in 21/22 (95.5%) kanamycin-resistant isolates, and the aadA1 gene was positive in 19/20 (95%) isolates showing resistance or intermediate resistance to streptomycin. Three isolates that were resistant to nalidixic acid and showed intermediate resistance to ciprofloxacin had distinct mutations in the QRDR of the gyrA gene: D87G, S83E, and D87A. In addition, one isolate with intermediate resistance to ciprofloxacin harbored an A67S mutation in the gyrA gene. Notably, the PMQR gene, qnrA, was identified in 9/16 (56.3%) isolates exhibiting intermediate resistance to ciprofloxacin and susceptibility or intermediate resistance to nalidixic acid.
4. Discussion
This study revealed Salmonella contamination in 32 (64%) of 50 chicken samples collected from retail shops in Fukuoka (Table 1). The rate of NTS contamination was higher than that reported for the same area in 2015 (48.6%, 17/35) [22], although a statistical comparison was not conducted due to the limited sample size. In Japan, domestic broiler chicken production has increased by 26%, reaching 2.5 million tons over the past decade (2009–2019) [34]. This increase may have contributed to the higher rate of NTS carriage in chickens. In 2021, the prevalence of Salmonella infection in broiler flocks in Japan exceeded 80%. This could increase the isolation rate of NTS in chicken meat. Our findings are consistent with those of previous studies, which also indicated a significant increase in the prevalence of NTS in chicken meat during various sampling periods [35,36]. The detection of Salmonella at the retail level does not necessarily imply that contamination originated solely from poultry, as contamination can occur at various stages of the food supply chain, including slaughter, processing, distribution, and retail handling. Further farm-to-table tracing studies are needed to identify the primary source of contamination. Nevertheless, our findings highlight poultry as one of the most significant sources of human salmonellosis in Fukuoka, Japan.
The most common serotype among the 32 NTS strains was S. enterica ser. Schwarzengrund, followed by S. enterica ser. Thompson and S. enterica ser. Oranienburg (Table 1). Despite the multiyear gap between this and previous studies, our findings are consistent with those of previous reports. Prior research conducted in Fukuoka in 2015 indicated that S. enterica ser. Schwarzengrund is common in chicken meat. Our results showed that this serovar remains the most prevalent. However, the proportion of S. enterica ser. Schwarzengrund was 78.1%, which was slightly higher than that reported in earlier studies in Fukuoka (70.6%) [22]. A survey by Konishi et al. [37] demonstrated that the prevalence of S. enterica ser. Schwarzengrund in food has increased from 13.3% (2012–2014) to 51.3% (2018–2020) in Tokyo, Japan. Furthermore, the isolation rate of S. enterica ser. Schwarzengrund has significantly increased, making it the primary serovar of Salmonella isolated from food in Japan since 2016. This increase may be partly attributed to the emergence of the S. enterica ser. Schwarzengrund ST241 clone, which evolved from the globally recognized ST96 sequence type found in chickens in Japan [12,34]. Additionally, the presence of a pESI-like plasmid that enhances the ability of Schwarzengrund to colonize and spread may contribute to this trend [38]. Similarly, S. enterica ser. Schwarzengrund has been reported in countries such as Asia, Denmark, the USA, and Brazil as an emerging bacterial pathogen [39]. Moreover, previous studies in Fukuoka have shown that S. enterica ser. Infantis and S. enterica ser. Manhattan are the second and third most frequently observed serovars, respectively [22]. However, this study found S. enterica ser. Thompson and S. enterica ser. Oranienburg were the second and third most common serovars, respectively, suggesting that the prevalence of certain serovars can change over time.
MLST analysis of the 32 NTS strains identified three STs (ST241, ST26, and ST23) (Table 2), indicating low genetic variability among the strains. Therefore, a correlation was observed between the ST profile and distribution of virulence genes in our study. Additionally, a strong association was observed between serotypes and STs, such as ST241 with Schwarzengrund and ST26 with Thompson, which is consistent with previous findings [12,40]. Several studies have indicated that MLST offers a better discriminatory power than serotyping [41]. Multiple studies have shown that ST23 is strongly associated with the Oranienburg serovar [11,42]. No other known serovars in the serogroups C1 and serotype G complex H1 showed this association. Previous research in Japan has shown that ST241 is the only ST consistently linked to Schwarzengrund in chickens, which is consistent with our findings [12]; however, it differs from the globally reported ST96 [43]. According to the PubMLST database, ST26 and ST23 are common in poultry samples from various countries, although their prevalence in Japan remains remarkably low. Notably, all the identified STs pose a significant public health risk because they can cause human salmonellosis [44,45], highlighting the need for effective preventive measures.
The PCA dendrogram created from the MALDI-TOF MS data showed six clusters, indicating considerable protein diversity among the isolates. This clustering demonstrated that the three NTS serovars were not distinct (Figure 1). Thus, PCA clustering of MALDI-TOF MS profiles could not distinguish NTS serovars, which is consistent with previous reports [46]. However, PCA clustering helps distinguish relationships among closely related strains and identify their associations [47]. We found that some isolates with the same serotypes, STs, AMR profiles, virulence genes, and biofilm formation patterns clustered together. Nonetheless, there was significant variability in antimicrobial resistance among isolates within the same cluster (Figure 1). Factors such as horizontal gene transfer, mutations, and genetic recombination cause variations in antimicrobial resistance genes, leading to different resistance profiles [48]. Although MALDI-TOF MS does not identify individual proteins or their specific functions, mass spectral patterns obtained from cellular proteins provide high-resolution molecular signatures of diversity that can represent strain persistence, adaptability, and virulence. These results highlight that the use of proteomic profiling in routine surveillance has great value in providing epidemiologically relevant information to guide public health strategies and risk assessments, and is crucial for tracking NTS in the poultry food chain.
The pathogenicity of bacteria is governed by not only the presence of virulence genes but also by their ability to form biofilms and resist antimicrobials. Our study showed that all NTS serovars harbored several virulence genes (Table 3), highlighting their zoonotic potential and the emergence of antimicrobial resistance. Salmonella serotypes can form biofilms in food and processing environments, which is problematic because these biofilms protect the bacteria from antibiotics, sanitizers, and various environmental factors [49]. In the present study, S. enterica ser. Schwarzengrund and S. enterica ser. Thompson produced biofilms of varying degrees (Table 4), complicating the inhibition or removal of these pathogens. The persistence of biofilms in food and food-processing environments poses significant risks to public health. Therefore, continuous monitoring is essential because of the risks posed by these serovars in food, which are associated with enteric and systemic diseases in humans [50].
Phenotypic planktonic susceptibility analysis revealed that none of the S. enterica ser. Thompson or S. enterica ser. Oranienburg strains was resistant to any of the tested antimicrobials. However, S. enterica ser. Thompson showed intermediate resistance to minocycline and ciprofloxacin. Resistance to one or more antimicrobials was detected in S. enterica ser. Schwarzengrund isolates (Table 5). In this study, most S. enterica ser. Schwarzengrund isolates demonstrated resistance to various antibiotics, including aminoglycosides (kanamycin and streptomycin), tetracyclines (tetracycline and minocycline), sulfonamides (sulfamethoxazole/trimethoprim), and fluoroquinolones (nalidixic acid). Our findings in Fukuoka differ from those of a recent study in Kagoshima, Japan [39], which reported high levels of resistance to streptomycin, tetracycline, and sulfonamides in S. enterica ser. Schwarzengrund. The resistance rates of S. enterica ser. Schwarzengrund to kanamycin and nalidixic acid in our study were comparable to those in the Kyushu region [51], but higher than those in Kagoshima. Therefore, regional differences in antimicrobial resistance patterns of S. enterica ser. Schwarzengrund in retail chickens were observed for certain antimicrobials. Differences in the type and frequency of antimicrobial use in poultry farming may have contributed to this outcome. Nevertheless, compared to earlier studies in Japan, a lower incidence of resistance to various antimicrobials was observed in our study. These results can be attributed to the fact that farmers are using antimicrobials properly, adhering to the guidelines of the National Action Plan on Antimicrobial Resistance (2023–2027), which aims to reduce antimicrobial usage in the livestock sector by 15% from 2020 to 2027, from the Government of Japan [52].
Biofilm susceptibility testing showed that all strains were less susceptible to the tested antimicrobials in the biofilm state than in the planktonic state (Table S11). Bacterial biofilms are reported to tolerate 100- to 1000-fold higher antimicrobial concentrations than their planktonic counterparts [32]. Consistent with this, all tested beta-lactams—piperacillin, cefotaxime, aztreonam, and meropenem—as well as gentamicin and fosfomycin, exhibited very high MBIC values (>512 µg/mL) in most of the isolates (Table S11), indicating poor activity against established biofilms [32,33,53]. This reduced activity of beta-lactams is partly due to the presence of beta-lactamases within the biofilm matrix [33]. MBICs of minocycline (4–64 µg/mL) were higher than its planktonic MICs (2 -> 8 µg/mL) but lower than those of the other antibiotics. The higher lipophilicity of minocycline may enhance its penetration into the biofilm matrix [54], contributing to its comparatively better activity than that of traditional tetracycline. Ciprofloxacin also exhibited relatively low MBIC values (1–4 µg/mL) across the isolates, which may be due to its excellent penetration rate into exopolysaccharides [55]. Overall, our findings indicate that the tested biofilm-forming isolates displayed increased tolerance to commonly used antibiotics and underscore the need for alternative strategies to control biofilm-producing Salmonella.
Our results demonstrate a correlation between the phenotypic and genotypic resistance profiles of the resistant strains. Phenotypically resistant isolates lacking detectable resistant genes may result from alternative resistance mechanisms or mutations in genes used for PCR detection. We found that most S. enterica ser. Schwarzengrund isolates were resistant to kanamycin, streptomycin, tetracycline, and sulfamethoxazole/trimethoprim and carried the genes aphA1, aadA1, tetA, and sul1 (Table 7). This suggests that these genes are responsible for resistance to these antimicrobials in S. enterica ser. Schwarzengrund, which is consistent with the results of previous studies [56,57]. In addition, a single mutation was identified in codons 67, 83, and 87 of the gyrA gene in four S. enterica ser. Schwarzengrund isolates (25%) that exhibited resistance or intermediate resistance to nalidixic acid and ciprofloxacin (Table 7). Previous studies indicated that low-level resistance to quinolones and fluoroquinolones is associated with a single mutation in the gyrA gene [58]. Numerous studies have demonstrated that variations within a specific domain of the gyrA N-terminus, particularly between amino acids 67 (Ala) and 106 (Gln), are responsible for quinolone and fluoroquinolone resistance [59], which supports our findings. However, PMQR genes typically result in decreased susceptibility to ciprofloxacin while maintaining susceptibility to nalidixic acid [60]. In this study, most S. enterica ser. Schwarzengrund isolates with low susceptibility to ciprofloxacin but susceptibility or intermediate resistance to nalidixic acid had the qnrA gene but lacked mutations in the QRDR (Table 7). This indicated that the presence of the qnrA gene may contribute to reduced susceptibility to ciprofloxacin in these isolates. However, other mechanisms, such as mutations in parC or parE, the presence of additional PMQR genes, and efflux-related mechanisms [61], were not investigated in this study and therefore cannot be excluded. Based on our current literature review, this is the first report of the qnrA gene detected in Salmonella in Japan. While the qnrS1 gene has been sporadically reported in Salmonella strains isolated from domestic animals [62], the detection of qnrA represents a potentially important change in the local quinolone resistance profile. Since qnr genes are located on plasmids [60], their presence in NTS isolates from animals poses health risks to humans and animals by enabling spread to other Salmonella serovars or genera and promoting high-level fluoroquinolone resistance under selective pressure. This is especially concerning given recent invasive NTS clusters in the Kyushu area in Japan [25]. Additionally, since the Kyushu area plays a key role in poultry production and distribution across Japan, the detection of qnrA in this region is not only locally significant but may also have wider implications for the national food supply. These findings underscore the need for enhanced region-specific genomic surveillance to enable early detection and improved risk assessment.
This study is limited by its small sample size, which may affect the accuracy in representing the overall prevalence and distribution of NTS across Fukuoka, Japan. As a result, the findings should be interpreted with caution. To gain a more comprehensive understanding of regional prevalence and public health implications, future research should involve a larger, more geographically diverse sample population.
5. Conclusions
This study detected Salmonella contamination in chicken meat samples in Fukuoka, Japan, similar to findings in other regions. Compared with previous studies conducted in Japan, the prevalence of S. enterica ser. Schwarzengrund continues to increase, although the rate of antimicrobial resistance has decreased. Despite its low genetic diversity, MDR S. enterica ser. Schwarzengrund exhibiting considerable protein diversity isolated in this study presents a potential risk to food safety and public health. Fluoroquinolones are the first-line antimicrobial agents used to treat salmonellosis in Japan. The presence of the qnrA gene in S. enterica ser. Schwarzengrund isolates raises public health concerns because it can rapidly spread and increase quinolone resistance, causing difficult-to-treat infections. This highlights the need for ongoing surveillance and control efforts on broiler farms.
Supplementary Materials
The following supporting information can be downloaded at: https://www.mdpi.com/article/10.3390/microbiolres17030063/s1. Table S1: Primers used for Salmonella detection, O serogrouping, and H serotyping; Table S2: PCR conditions for assays of O serogrouping, and H serotyping; Table S3: Primers used for virulence gene detection; Table S4: PCR conditions for virulence gene detection; Table S5: Primers used for antimicrobial-resistant gene detection; Table S6: PCR conditions for antimicrobial-resistant gene detection; Table S7: Primers used for multi-locus sequence typing (MLST); Table S8: PCR conditions for MLST; Table S9: MALDI-TOF MS score values of Salmonella isolates; Table S10: MIC ranges for antimicrobial agents determined by broth microdilution for Salmonella serovars; Table S11: Minimal Inhibitory Concentration (MIC) and Minimal Biofilm Inhibitory Concentration (MBIC) of selected strains. Figure S1: Representative image of the crystal violet staining assay used to evaluate the biofilm formation by Salmonella isolates in a 96-well microtiter plate; Figure S2: Representative image of the resazurin-based cell viability assay used to evaluate antimicrobial susceptibility of Salmonella biofilms in a 96-well microtiter plate. References [28,56,63,64,65,66,67,68,69,70,71,72,73] are cited in the Supplementary Materials.
Author Contributions
Conceptualization, T.M.; methodology, K.Z.L., S.Z.C.L., A.T.M., M.N.S.A., Y.M., K.-i.H., and T.M.; validation, T.M., K.-i.H., Y.M., and K.Z.L.; investigation, K.Z.L., S.Z.C.L., A.T.M., M.N.S.A., C.D.H., Y.L., and H.Y.; data curation, K.Z.L.; writing—original draft preparation, K.Z.L.; writing—review and editing, T.M., K.-i.H., and Y.M.; visualization, K.Z.L., S.Z.C.L., A.T.M., and M.N.S.A.; supervision, T.M. All authors have read and agreed to the published version of the manuscript.
Funding
This research received no external funding.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
The original contributions presented in this study are included in the article/Supplementary Material. Further inquiries can be directed to the corresponding author.
Conflicts of Interest
The authors declare no conflicts of interest.
References
- Havelaar, A.H.; Kirk, M.D.; Torgerson, P.R.; Gibb, H.J.; Hald, T.; Lake, R.J.; Praet, N.; Bellinger, D.C.; de Silva, N.R.; Gargouri, N.; et al. World health organization global estimates and regional comparisons of the burden of foodborne disease in 2010. PLoS Med. 2015, 12, e1001923. [Google Scholar] [CrossRef]
- World Health Organization (WHO). Salmonella (Non-Typhoidal). 2018. Available online: https://www.who.int/news-room/fact-sheets/detail/salmonella-(non-typhoidal) (accessed on 26 June 2024).
- Wang, M.; Qazi, I.H.; Wang, L.; Zhou, G.; Han, H. Salmonella virulence and immune escape. Microorganisms 2020, 8, 407. [Google Scholar] [CrossRef]
- Ćwiek, K.; Korzekwa, K.; Tabiś, A.; Bania, J.; Bugla-Płoskońska, G.; Wieliczko, A. Antimicrobial resistance and biofilm formation capacity of Salmonella enterica serovar Enteritidis strains isolated from poultry and humans in Poland. Pathogens 2020, 9, 643. [Google Scholar] [CrossRef]
- Shimojima, Y.; Nishino, Y.; Fukui, R. Salmonella serovars isolated from retail meats in Tokyo, Japan and their antimicrobial susceptibility. J. Food Hyg. Soc. Jpn. 2020, 61, 211–217. [Google Scholar] [CrossRef] [PubMed]
- Moore, J.E.; Matsuda, M. Consumption of raw chicken sashimi, Kyushu Island, Japan-risk of campylobacteriosis or not? Travel Med. Infect. Dis. 2007, 5, 64–65. [Google Scholar] [CrossRef] [PubMed]
- Sasaki, Y.; Ikeda, T.; Momose, Y.; Yonemitsu, K.; Uema, M.; Asai, T. Geographical variation of antimicrobial resistance of Salmonella in Japanese chicken. Food Saf. 2024, 12, 59–66. [Google Scholar] [CrossRef]
- Centers for Disease Control and Prevention (CDC). Serotypes and the Importance of Serotyping Salmonella. 2022. Available online: http://medbox.iiab.me/modules/en-cdc/www.cdc.gov/salmonella/reportspubs/salmonella-atlas/serotyping-importance.html (accessed on 3 January 2025).
- Grimont, P.; Weill, F.X. Antigenic formulae of the Salmonella serovars. In WHO Collaborating Centre for Reference and Research on Salmonella, 9th ed.; WHO: Geneva, Switzerland, 2007; pp. 1–166. [Google Scholar]
- Santos, P.D.M.; Widmer, K.W.; Rivera, W.L. PCR-based detection and serovar identification of Salmonella in retail meat collected from wet markets in Metro Manila, Philippines. PLoS ONE 2020, 15, e0239457. [Google Scholar] [CrossRef] [PubMed]
- Gómez-Baltazar, A.; Godínez-Oviedo, A.; Vázquez-Marrufo, G.; Vázquez-Garcidueñas, M.S.; Hernández-Iturriaga, M. Genomic analysis of the MLST population structure and antimicrobial resistance genes associated with Salmonella enterica in Mexico. Genome 2023, 66, 319–332. [Google Scholar] [CrossRef]
- Ikeuchi, S.; Okumura, M.; Niwa, T.; Sasaki, Y.; Hara-Kudo, Y.; Hatashidani, H. Molecular epidemiological analysis of Salmonella Schwarzengrund isolated in Japan by newly developed multi-locus variable-number tandem repeat analysis method. LWT 2024, 207, 116593, Corrigendum in LWT 2024, 209, 116742. https://doi.org/10.1016/j.lwt.2024.116742. [Google Scholar] [CrossRef]
- Linn, K.Z.; Furuta, M.; Nakayama, M.; Masuda, Y.; Honjoh, K.; Miyamoto, T. Characterization and antimicrobial resistance of Campylobacter jejuni and Campylobacter coli isolated from chicken and pork. Int. J. Food Microbiol. 2021, 360, 109440. [Google Scholar] [CrossRef]
- Nayak, R.; Stewart, T.; Wang, R.F.; Lin, J.; Cerniglia, C.E.; Kenney, P.B. Genetic diversity and virulence gene determinants of antibiotic-resistant Salmonella isolated from preharvest turkey production sources. Int. J. Food Microbiol. 2004, 91, 51–62. [Google Scholar] [CrossRef] [PubMed]
- Mthembu, T.P.; Zishiri, O.T.; El Zowalaty, M.E. Detection and molecular identification of Salmonella virulence genes in livestock production systems in South Africa. Pathogens 2019, 8, 124. [Google Scholar] [CrossRef]
- Kadry, M.; Nader, S.M.; Dorgham, S.M.; Kandil, M.M. Molecular diversity of the invA gene obtained from human and egg samples. Vet. World 2019, 12, 1033–1038. [Google Scholar] [CrossRef] [PubMed]
- Uchiya, K.I.; Nikai, T. Salmonella virulence factor SpiC is involved in expression of flagellin protein and mediates activation of the signal transduction pathways in macrophages. Microbiol. Soc. 2008, 154, 3491–3502. [Google Scholar] [CrossRef]
- Carroll, R.K.; Liao, X.; Morgan, L.K.; Cicirelli, E.M.; Li, Y.; Sheng, W.; Feng, X.; Kenney, L.J. Structural and functional analysis of the C-terminal DNA binding domain of the Salmonella typhimurium SPI-2 response regulator SsrB. J. Biol. Chem. 2009, 284, 12008–12019. [Google Scholar] [CrossRef]
- Marcus, S.L.; Brumell, J.H.; Pfeifer, C.G.; Finlay, B.B. Salmonella pathogenicity islands: Big virulence in small packages. Microbes Infect. 2000, 2, 145–156. [Google Scholar] [CrossRef] [PubMed]
- Su, L.; Chiu, C.; Chu, C.; Ou, J. Antimicrobial resistance in nontyphoid Salmonella serovars: A global challenge. Clin. Infect. Dis. 2004, 39, 546–551. [Google Scholar] [CrossRef]
- Noda, T.; Murakami, K.; Etoh, Y.; Okamoto, F.; Yatsuyanagi, J.; Sera, N.; Furuta, M.; Onozuka, D.; Oda, T.; Asai, T.; et al. Increase in resistance to extended-spectrum cephalosporins in Salmonella isolated from retail chicken products in Japan. PLoS ONE 2015, 10, e0116927, Correction in PLoS ONE 2015, 10, e0121469. https://doi.org/10.1371/journal.pone.0121469; Correction in PLoS ONE 2015, 10, e0133094. https://doi.org/10.1371/journal.pone.0133094. [Google Scholar] [CrossRef]
- Shigemura, H.; Matsui, M.; Sekizuka, T.; Onozuka, D.; Noda, T.; Yamashita, A.; Kuroda, M.; Suzuki, S.; Kimura, H.; Fujimoto, S.; et al. Decrease in the prevalence of extended-spectrum cephalosporin-resistant Salmonella following cessation of ceftiofur use by the Japanese poultry industry. Int. J. Food Microbiol. 2018, 274, 45–51. [Google Scholar] [CrossRef]
- Teklemariam, A.D.; Al-Hindi, R.R.; Albiheyri, R.S.; Alharbi, M.G.; Alghamdi, M.A.; Filimban, A.A.R.; Al Mutiri, A.S.; Al-Alyani, A.M.; Alseghayer, M.S.; Almaneea, A.M. Human salmonellosis: A continuous global threat in the farm-to-fork food safety continuum. Foods 2023, 12, 1756. [Google Scholar] [CrossRef]
- Jesudason, T. WHO publishes updated list of bacterial priority pathogens. Lancet Microbe 2024, 5, 100940. [Google Scholar] [CrossRef]
- Ooka, T.; Gotoh, Y.; Hatanaka, S.; Yoshimori, M.; Nishitarumizu, K.; Kojo, K.; Kosakamoto, H.; Sameshima, K.; Kuroki, Y.; Chibana, N.; et al. A Salmonella enterica serovar Oranienburg Clone caused a cluster of bacteremia cases in persons with no recognizable underlying diseases in Japan. Open Forum Infect. Dis. 2022, 10, ofac695. [Google Scholar] [CrossRef] [PubMed]
- ISO 6579:2002; Microbiology of Food and Animal Feeding Stuffs–Horizontal Method for the Detection of Salmonella spp. ISO: Geneva, Switzerland, 2002.
- Zhao, J.; Wang, D.; Wang, C.; Lin, Y.; Ye, H.; Maung, A.T.; El-Telbany, M.; Masuda, Y.; Honjoh, K.; Miyamoto, T.; et al. Biocontrol of Salmonella Schwarzengrund and Escherichia coli O157:H7 planktonic and biofilm cells via combined treatment of polyvalent phage and sodium hexametaphosphate on foods and food contact surfaces. Food Microbiol. 2024, 126, 104680. [Google Scholar] [CrossRef] [PubMed]
- Stepan, R.M.; Sherwood, J.S.; Petermann, S.R.; Logue, C.M. Molecular and comparative analysis of Salmonella enterica Senftenberg from humans and animals using PFGE, MLST and NARMS. BMC Microbiol. 2011, 11, 153. [Google Scholar] [CrossRef] [PubMed]
- Miyamoto, T.; Kawagishi, J.; Oishi, A.; Shimotsu, S.; Mishima, T.; Kobayashi, H.; Honjoh, K.I. Inhibition of adhesion of several bacteria onto microtiter plate by selected food additives. Jpn. J. Food Microbiol. 2011, 28, 157–166. [Google Scholar] [CrossRef]
- Stepanovic, S.; Vukovic, D.; Dakic, I.; Savic, B.; Svabic-Vlahovic, M. A modified microtiter-plate test for quantification of staphylococcal biofilm formation. J. Microbiol. Methods 2000, 40, 175–179. [Google Scholar] [CrossRef]
- CLSI. Performance Standards for Antimicrobial Susceptibility Testing, 30th ed.; CLSI Supplement M100; CLSI: Wayne, PA, USA, 2020. [Google Scholar]
- do Canto Canabarro, M.; Meneghetti, K.L.; Geimba, M.P.; Corção, G. Biofilm formation and antibiotic susceptibility of Staphylococcus and Bacillus species isolated from human allogeneic skin. Braz. J. Microbiol. 2022, 53, 153–160. [Google Scholar] [CrossRef]
- Lozano, C.; López, M.; Rojo-Bezares, B.; Sáenz, Y. Antimicrobial susceptibility testing in Pseudomonas aeruginosa biofilms: One step closer to a standardized method. Antibiotics 2020, 9, 880. [Google Scholar] [CrossRef]
- Matsui, K.; Nakazawa, C.; Thiri Maung Maung Khin, S.; Iwabuchi, E.; Asai, T.; Ishihara, K. Molecular characteristics and antimicrobial resistance of Salmonella enterica serovar Schwarzengrund from chicken meat in Japan. Antibiotics 2021, 10, 1336. [Google Scholar] [CrossRef]
- Peruzy, M.F.; Proroga, Y.T.R.; Capuano, F.; Mancusi, A.; Montone, A.M.I.; Cristiano, D.; Balestrieri, A.; Murru, N. Occurrence and distribution of Salmonella serovars in carcasses and foods in southern Italy: Eleven-year monitoring (2011–2021). Front. Microbiol. 2022, 13, 1005035. [Google Scholar] [CrossRef]
- Regalado-Pineda, I.D.; Rodarte-Medina, R.; Resendiz-Nava, C.N.; Saenz-Garcia, C.E.; Castañeda-Serrano, P.; Nava, G.M. Three-year longitudinal study: Prevalence of Salmonella enterica in chicken meat is higher in supermarkets than wet markets from Mexico. Foods 2020, 9, 264. [Google Scholar] [CrossRef]
- Konishi, N.; Obata, H.; Yokoyama, K.; Sadamasu, K.; Kai, A. Comparison of the serovars and characteristics of Salmonella isolated from human feces and foods in the 1990s and 2010s in Tokyo. Jpn. J. Infect. Dis. 2023, 76, 14–19. [Google Scholar] [CrossRef]
- dos Santos, A.M.; Panzenhagen, P.; Ferrari, R.G.; Conte-Junior, C.A. Large-scale genomic analysis reveals the pESI-like megaplasmid presence in Salmonella Agona, Muenchen, Schwarzengrund, and Senftenberg. Food Microbiol. 2022, 108, 104112. [Google Scholar] [CrossRef]
- Duc, V.M.; Shin, J.; Nagamatsu, Y.; Fuhiwara, A.; Toyofuku, H.; Obi, T.; Chuma, T. Increased Salmonella Schwarzengrund prevalence and antimicrobial susceptibility of Salmonella enterica isolated from broiler chickens in Kagoshima Prefecture in Japan between 2013 and 2016. J. Vet. Med. Sci. 2020, 82, 585–589. [Google Scholar] [CrossRef]
- Gonzalez-Escalona, N.; Kase, J.A. Virulence gene profiles and phylogeny of shiga toxin-positive Escherichia coli strains isolated from FDA regulated foods during 2010–2017. PLoS ONE 2019, 14, e0214620. [Google Scholar] [CrossRef] [PubMed]
- Zhou, M.; Li, X.; Hou, W.; Wang, H.; Paoli, G.C.; Shi, X. Incidence and characterization of Salmonella isolates from raw meat products sold at small markets in Hubei province, China. Front. Microbiol. 2019, 10, 2265. [Google Scholar] [CrossRef] [PubMed]
- Kang, H.M.; Chu, J.; Yoo, I.H.; Yoo, I.Y.; Shin, I.; Seo, R.; Chung, J.; Jung, H.; Park, Y.J. Epidemiology of nontyphoidal Salmonella infections in Korean children and genetic factors associated with extra-intestinal invasion: A whole-genome sequencing analysis. Ann. Lab. Med. 2025, 45, 312–321. [Google Scholar] [CrossRef] [PubMed]
- Alikhan, N.F.; Zhou, Z.; Sergeant, M.J.; Achtman, M. A genomic overview of the population structure of Salmonella. PLoS Genet 2018, 14, e1007261. [Google Scholar] [CrossRef]
- Eun, Y.; Jeong, H.; Kim, S.; Park, W.; Ahn, B.; Kim, D.; Kim, E.; Park, E.; Park, S.; Hwang, I.; et al. A large outbreak of Salmonella enterica serovar Thompson infections associated with chocolate cake in Busan, Korea. Epidemiol. Health 2019, 41, e2019002. [Google Scholar] [CrossRef]
- Mitchell, M.R., Jr.; Kirchner, M.; Schneider, B.; McClure, M.; Neil, K.P.; Madad, A.; Jemaneh, T.; Tijerina, M.; Nolte, K.; Wellman, A.; et al. Multistate outbreak of Salmonella Oranienburg infections linked to bulb onions imported from Mexico–United States, 2021. Food Control 2024, 160, 110325. [Google Scholar] [CrossRef]
- Yang, S.M.; Kim, E.; Kim, D.; Baek, J.; Yoon, H.; Kim, H.Y. Rapid detection of Salmonella Enteritidis, Typhimurium, and Thompson by specific peak analysis using matrix-assisted laser desorption ionization time-of-flight mass spectrometry. Foods 2021, 10, 933. [Google Scholar] [CrossRef] [PubMed]
- Elabbasy, M.T.; Hussein, M.A.; Algahtani, F.D.; Abd El-Rahman, G.I.; Morshdy, A.E.; Elkafrawy, I.A.; Adeboye, A.A. MALDI-TOF MS based typing for rapid screening of multiple antibiotic resistance E. coli and virulent non-O157 shiga toxin-producing E. coli isolated from the slaughterhouse settings and beef carcasses. Foods 2021, 10, 820. [Google Scholar] [CrossRef]
- Davies, J.; Davies, D. Origins and evolution of antibiotic resistance. Microbiol. Mol. Bio. Rev. 2010, 74, 417–433. [Google Scholar] [CrossRef]
- Beshiru, A.; Igbinosa, I.H.; Igbinosa, E.O. Biofilm formation and potential virulence factors of Salmonella strains isolated from ready-to-eat shrimps. PLoS ONE 2018, 13, e0204345. [Google Scholar] [CrossRef] [PubMed]
- Tokuyama, Y.; Yamada, H.; Shinozuka, K.; Yunoki, T.; Ohtsuru, S. Pyogenic sacroiliitis caused by Salmonella Schwarzengrund in a young healthy woman: A case report and literature review. Int. J. Emerg. Med. 2023, 16, 21. [Google Scholar] [CrossRef]
- Yamamoto, T.; Toyofuku, H.; Mizote, T. Prevalence and antimicrobial resistance of Salmonella in poultry and chicken meat in Chugoku and Kyushu region in Japan. Jpn. J. Food Microbiol. 2021, 38, 78–87. [Google Scholar] [CrossRef]
- Hosoi, Y.; Matsuda, M.; Kawanishi, M.; Harada, S.; Kumakawa, M.; Sekiguchi, H.; Asai, T.; Sekiya, T. Antimicrobial use in the animal sector in Japan from 2011 to 2022. Antibiotics 2024, 13, 1204. [Google Scholar] [CrossRef]
- Wang, L.; Di Luca, M.; Tkhilaishvili, T.; Trampuz, A.; Gonzalez Moreno, M. Synergistic activity of fosfomycin, ciprofloxacin, and gentamicin against Escherichia coli and Pseudomonas aeruginosa biofilms. Front. Microbiol. 2019, 10, 252. [Google Scholar] [CrossRef]
- Asadi, A.; Abdi, M.; Kouhsari, E.; Panahi, P.; Sholeh, M.; Sadeghifard, N.; Amiriani, T.; Ahmadi, A.; Maleki, A.; Gholami, M. Minocycline, focus on mechanisms of resistance, antibacterial activity, and clinical effectiveness: Back to the future. J. Glob. Antimicrob. Resist. 2020, 22, 161–174. [Google Scholar] [CrossRef]
- Geremia, N.; Giovagnorio, F.; Colpani, A.; De Vito, A.; Botan, A.; Stroffolini, G.; Toc, D.A.; Zerbato, V.; Principe, L.; Madeddu, G.; et al. Fluoroquinolones and biofilm: A narrative review. Pharmaceuticals 2024, 17, 1673. [Google Scholar] [CrossRef] [PubMed]
- Kozak, G.K.; Pearl, D.L.; Parkman, J.; Reid-Smith, R.J.; Deckert, A.; Boerlin, P. Distribution of sulfonamide resistance genes in Escherichia coli and Salmonella isolates from swine and chickens at abattoirs in Ontario and Québec, Canada. Appl. Environ. Microbiol. 2009, 75, 5999–6001. [Google Scholar] [CrossRef]
- Zhu, J.; Wang, Y.; Song, X.; Cui, S.; Xu, H.; Yang, B.; Huang, J.; Liu, G.; Chen, Q.; Zhou, G.; et al. Prevalence and quantification of Salmonella contamination in raw chicken carcasses at the retail in China. Food Control 2014, 44, 198–202. [Google Scholar] [CrossRef]
- Minarini, L.A.; Darini, A.L. Mutations in the quinolone resistance-determining regions of gyrA and parC in Enterobacteriaceae isolates from Brazil. Braz. J. Microbiol. 2012, 43, 1309–1314. [Google Scholar] [CrossRef] [PubMed]
- Friedman, S.M.; Lu, T.; Drlica, K. Mutation in the DNA gyrase A gene of Escherichia coli that expands the quinolone resistance-determining region. Antimicrob. Agents Chemother. 2001, 45, 2378–2380. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez-Martínez, J.M.; Machuca, J.; Cano, M.E.; Calvo, J.; Martínez-Martínez, L.; Pascual, A. Plasmid-mediated quinolone resistance: Two decades on. Drug Resist. Updates 2016, 29, 13–29. [Google Scholar] [CrossRef] [PubMed]
- Cuypers, W.L.; Jacobs, J.; Wong, V.; Klemm, E.J.; Deborggraeve, S.; Van Puyvelde, S. Fluoroquinolone resistance in Salmonella: Insights by whole-genome sequencing. Microbial genomics 2018, 4, e000195. [Google Scholar] [CrossRef]
- Ahmed, A.M.; Ishida, Y.; Shimamoto, T. Molecular characterization of antimicrobial resistance in Salmonella isolated from animals in Japan. J. Appl. Microbiol. 2009, 106, 402–409. [Google Scholar] [CrossRef]
- Chiu, C.H.; Ou, J.T. Rapid identification of Salmonella serovars in feces by specific detection of virulence genes, invA and spvC, by an enrichment broth culture-multiplex PCR combination assay. J. Clin. Microbiol. 1996, 34, 2619–2622. [Google Scholar] [CrossRef]
- Hong, Y.; Liu, T.; Lee, M.D.; Hofacre, C.L.; Maier, M.; White, D.G.; Ayers, S.; Wang, L.; Berghaus, R.; Maurer, J.J. Rapid screening of Salmonella enterica serovars Enteritidis, Hadar, Heidelberg and Typhimurium using a serologically-correlative allelotyping PCR targeting the O and H antigen alleles. BMC Microbiol. 2008, 8, 178. [Google Scholar] [CrossRef]
- Hirose, K.; Itoh, K.I.; Nakajima, H.; Kurazono, T.; Yamaguchi, M.; Moriya, K.; Ezaki, T.; Kawamura, Y.; Tamura, K.; Watanabe, H. Selective amplification of tyv (rfbE), prt (rfbS), viaB, and fliC genes by multiplex PCR for identification of Salmonella enterica serovars Typhi and Paratyphi A. J. Clin. Microbiol. 2002, 40, 633–636. [Google Scholar] [CrossRef]
- Herrera-Leon, S.; McQuiston, J.R.; Usera, M.A.; Fields, P.I.; Garaizar, J.; Echeita, M.A. Multiplex PCR for distinguishing the most common phase-1 flagellar antigens of Salmonella spp. J. Clin. Microbiol. 2004, 42, 2581–2586. [Google Scholar] [CrossRef]
- Echeita, M.A.; Herrera, S.; Garaizar, J.; Usera, M.A. Multiplex PCR-based detection and identification of the most common Salmonella second-phase flagellar antigens. Res. Microbiol. 2002, 153, 107–113. [Google Scholar] [CrossRef] [PubMed]
- Cardona-Castro, N.; Restrepo-Pineda, E.; Correa-Ochoa, M. Detection of hilA gene sequences in serovars of Salmonella enterica subspecies enterica. Mem. Inst. Oswaldo Cruz. 2002, 97, 1153–1156. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Yue, M.; Li, X.; Liu, D.; Hu, X. Serotypes, antibiotic resistance, and virulence genes of Salmonella in children with diarrhea. J. Clin. Lab. Anal. 2020, 34, e23525. [Google Scholar] [CrossRef] [PubMed]
- Hughes, L.A.; Shopland, S.; Wigley, P.; Bradon, H.; Leatherbarrow, A.H.; Williams, N.J.; Bennett, M.; de Pinna, E.; Lawson, B.; Cunningham, A.A.; et al. Characterisation of Salmonella enterica serotype Typhimurium isolates from wild birds in northern England from 2005–2006. BMC Vet. Res. 2008, 4, 4. [Google Scholar] [CrossRef]
- Zhu, A.; Zhi, W.; Qiu, Y.; Wei, L.; Tian, J.; Pan, Z.; Kang, X.; Gu, W.; Duan, L. Surveillance study of the prevalence and antimicrobial resistance of Salmonella in pork from open markets in Xuzhou, China. Food Control 2019, 98, 474–480. [Google Scholar] [CrossRef]
- Randall, L.P.; Cooles, S.W.; Osborn, M.K.; Piddock, L.J.V.; Woodward, M.J. Antibiotic resistance genes, integrons and multiple antibiotic resistance in thirty-five serotypes of Salmonella enterica isolated from humans and animals in the UK. J. Antimicrob. Chemother. 2004, 53, 208–216. [Google Scholar] [CrossRef]
- Fabrega, A.; du Merle, L.; Le Bouguenec, C.; Jiménez de Anta, M.T.; Vila, J. Repression of invasion genes and decreased invasion in a high-level fluoroquinolone-resistant Salmonella Typhimurium mutant. PLoS ONE 2009, 4, e8029. [Google Scholar] [CrossRef][Green Version]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.