PLGA-PEG-ANG-2 Nanoparticles for Blood–Brain Barrier Crossing: Proof-of-Concept Study
Abstract
1. Introduction
2. Materials and Methods
2.1. Materials
2.2. Animals
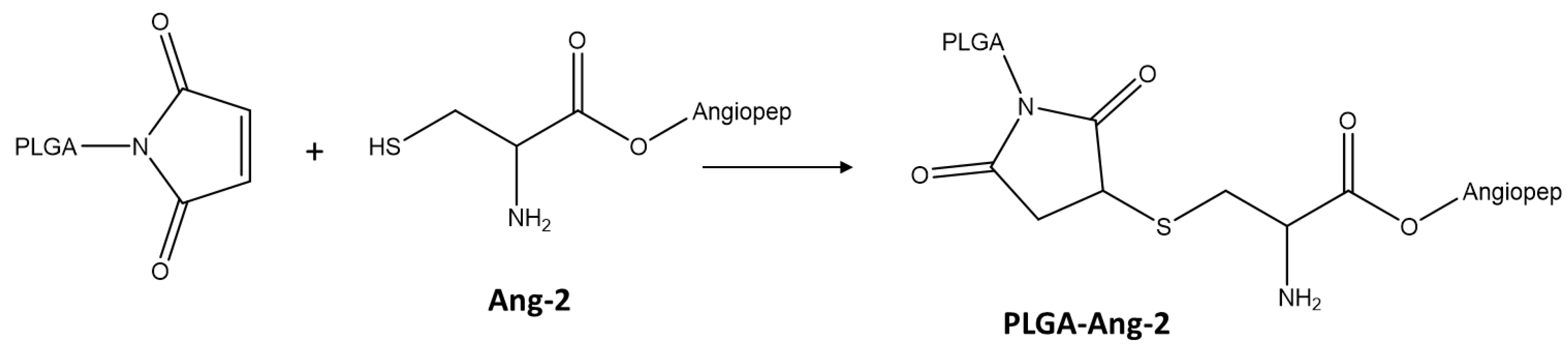
2.3. Synthesis and Characterization of PLGA Conjugate with the Ang–2
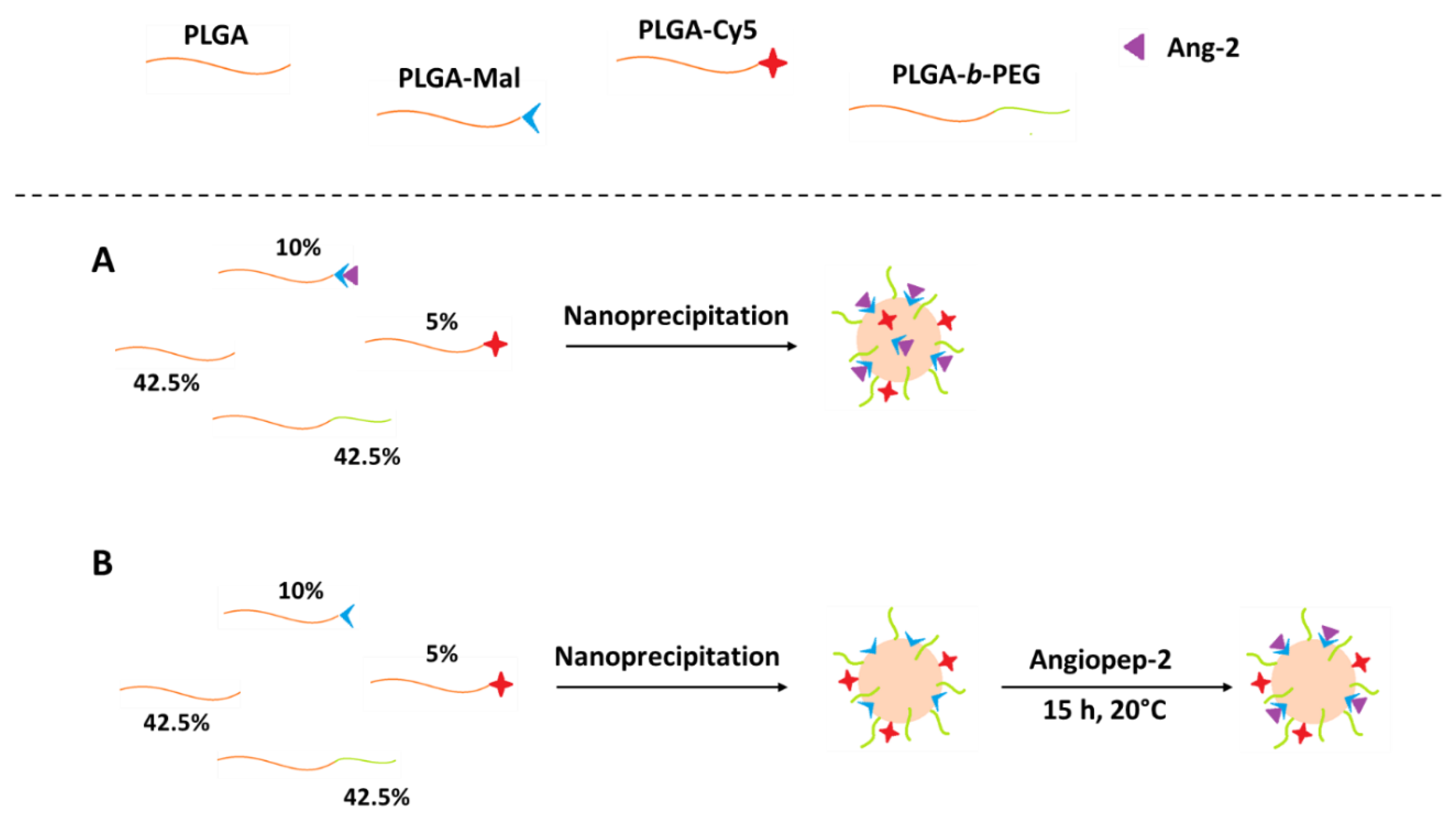
2.4. Preparation of PLGA Nanoparticles Functionalized with Ang–2 (Ang–2-NPs)
2.5. Purification of Nanoparticles
2.6. Characterization of Nanoparticles
2.6.1. Distribution of Particle Size and Zeta Potential
2.6.2. HPLC Quantification of Ang–2 in Post-Functionalized Nanoparticles
2.7. In Vivo Tests: Brain Uptake of the Nanoparticles
2.7.1. Animal Handling Protocols and Sample Preparation for Systemic Injection of Post-Functionalized Ang–2 NPs
2.7.2. Immunohistochemistry of Brain Sections
2.7.3. Confocal Analysis
3. Results and Discussion
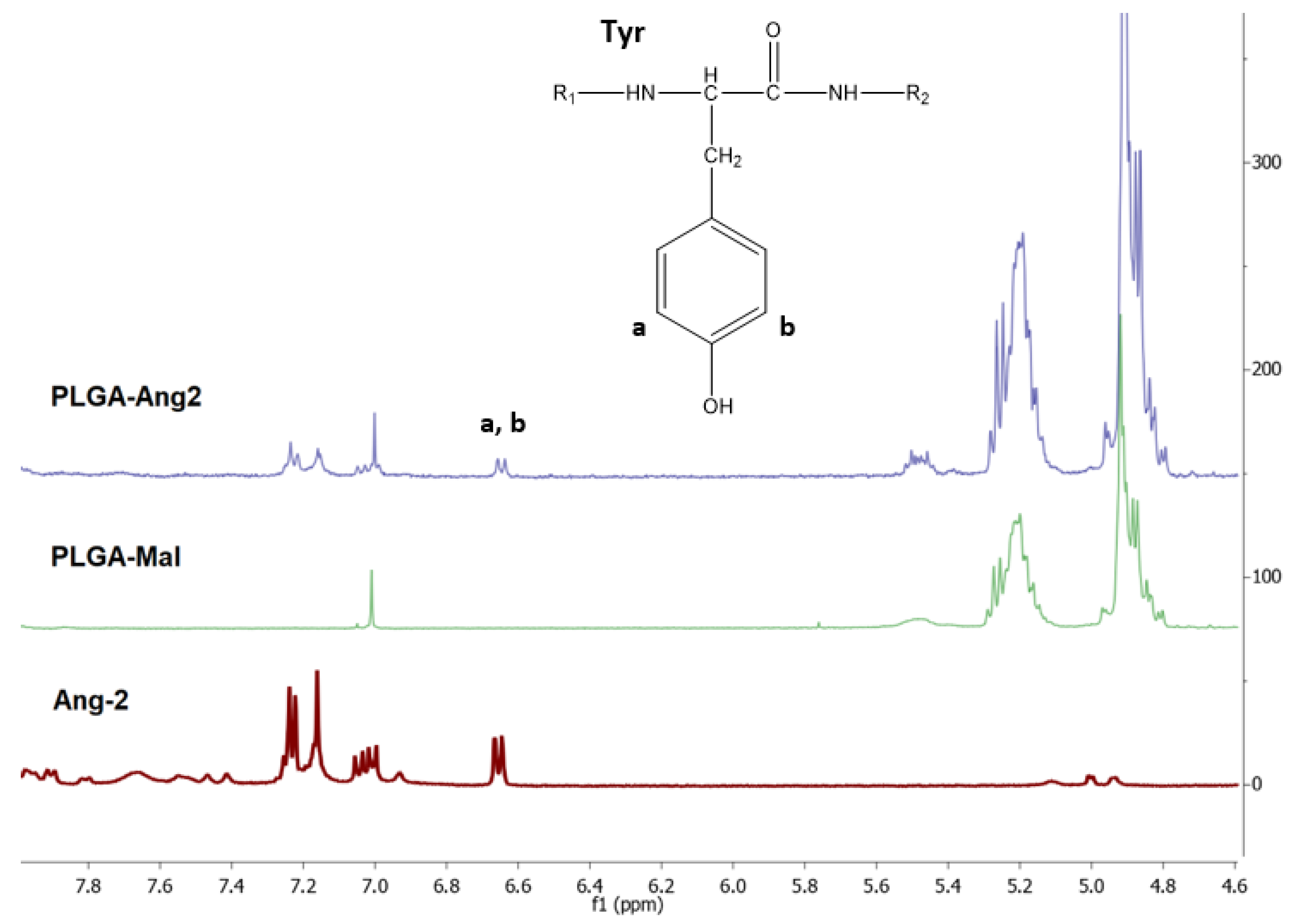
3.1. Quantification of Ang–2 on Modified PLGA by 1H-NMR
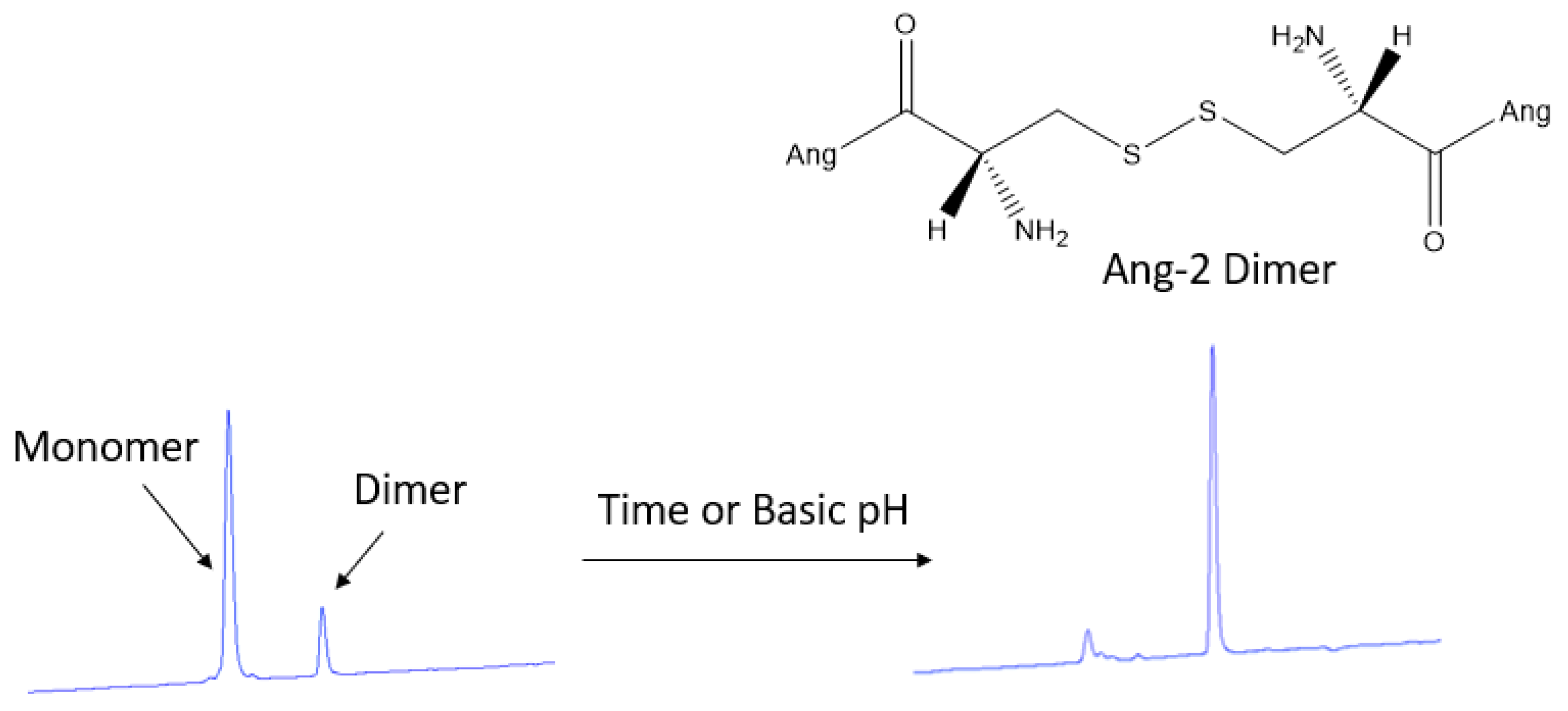
3.2. HPLC Quantification of Ang–2 in Post-Functionalized Nanoparticles
3.3. Size Distribution and Zeta Potential of Pre- and Post-Functionalized Ang2-NPs
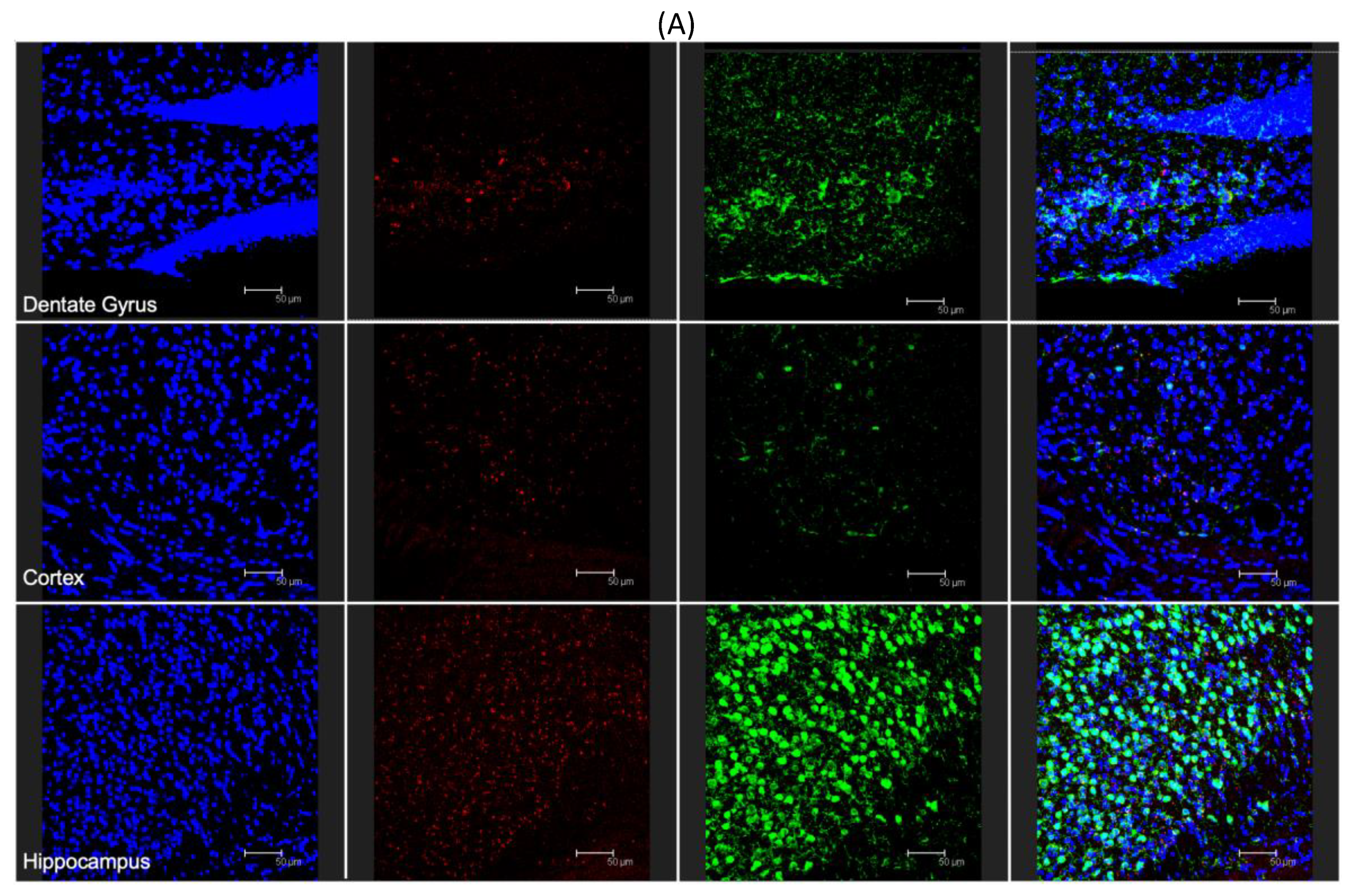
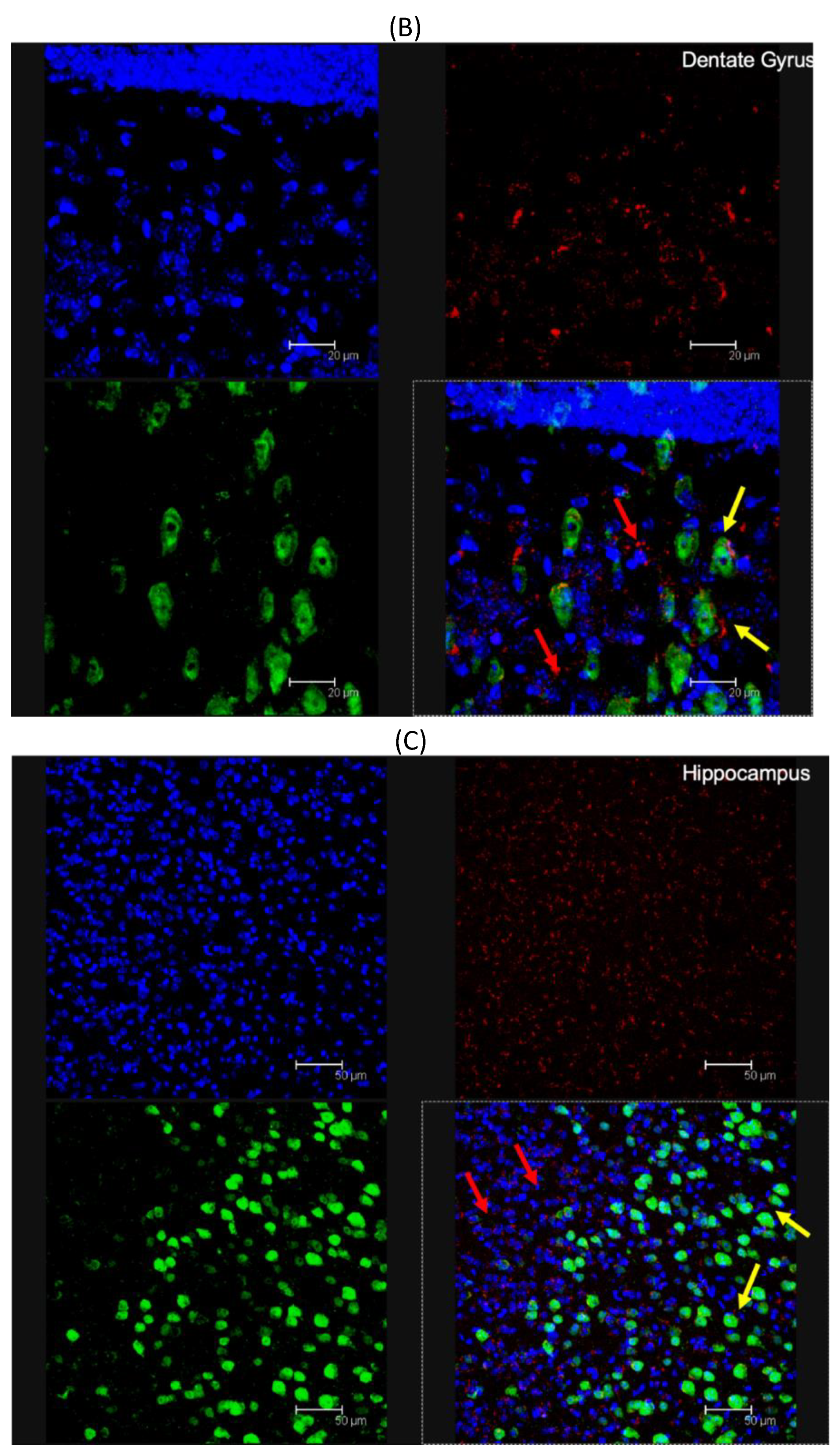
3.4. In Vivo Brain Distribution of ANG-2 NPs
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Beck-broichsitter, M.; Nicolas, J.; Couvreur, P. Design attributes of long-circulating polymeric drug delivery vehicles. Eur. J. Pharm. Biopharm. 2015, 97, 304–317. [Google Scholar] [CrossRef] [PubMed]
- Jokerst, J.; Lobovkina, T.; Zare, R.; Gambhir, S. Nanoparticle PEGylation for imaging and therapy. Nanomedicine (London) 2011, 6, 715–728. [Google Scholar] [CrossRef] [PubMed]
- Xie, J.; Shen, Z.; Anraku, Y.; Kataoka, K.; Chen, X. Biomaterials nanomaterial-based Blood-Brain-Barrier (BBB) crossing strategies. Biomaterials 2019, 224, 119491. [Google Scholar] [CrossRef] [PubMed]
- Portioli, C.; Bovi, M.; Benati, D.; Donini, M.; Perduca, M.; Romeo, A.; Dusi, S.; Monaco, H.L.; Bentivoglio, M. Novel functionalization strategies of polymeric nanoparticles as carriers for brain medications. J. Biomed. Mater. Res.-Part A 2016, 105, 847–858. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Liu, L. Modern methods for delivery of drugs across the blood-Brain barrier. Adv. Drug Deliv. Rev. 2012, 64, 640–665. [Google Scholar] [CrossRef]
- Ramalho, M.J.; Sevin, E.; Gosselet, F.; Lima, J.; Coelho, M.A.; Loureiro, J.A.; Pereira, M.C. Receptor-mediated PLGA nanoparticles for glioblastoma multiforme treatment. Int. J. Pharm. 2018, 545, 84–92. [Google Scholar] [CrossRef]
- Paka, G.D.; Ramassamy, C. Optimization of curcumin-loaded PEG-PLGA nanoparticles by GSH functionalization: Investigation of the internalization pathway in neuronal cells. Mol. Pharm. 2017, 14, 93–106. [Google Scholar] [CrossRef]
- Bors, L.A.; Erdö, F. Overcoming the Blood-Brain Barrier. Challenges and tricks for CNS drug delivery. Sci. Pharm. 2019, 87, 6. [Google Scholar] [CrossRef]
- Auderset, L.; Cullen, C.L.; Young, K.M. Low density lipoprotein-receptor related protein 1 is differentially expressed by neuronal and glial populations in the developing and mature mouse central nervous system. PLoS ONE 2016, 11, e0155878. [Google Scholar] [CrossRef]
- Tian, X.; Nyberg, S.; Sharp, P.S.; Madsen, J.; Daneshpour, N.; Armes, S.P.; Berwick, J.; Azzouz, M.; Shaw, P.; Abbott, N.J.; et al. LRP-1-Mediated intracellular antibody delivery to the central nervous system. Sci. Rep. 2015, 5, 11990. [Google Scholar] [CrossRef]
- Spuch, C.; Ortolano, S.; Navarro, C. LRP-1 and LRP-2 receptors function in the membrane neuron. trafficking mechanisms and proteolytic processing in alzheimer’s disease. Front. Physiol. 2012, 3, 269. [Google Scholar] [CrossRef] [PubMed]
- Marzolo, M.P.; Yuseff, M.I.; Retamal, C.; Donoso, M.; Ezquer, F.; Farfán, P.; Li, Y.; Bu, G. Differential distribution of low-density Lipoprotein-Receptor-Related protein (LRP) and megalin in polarized epithelial cells is determined by their cytoplasmic domains. Traffic 2003, 4, 273–288. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Xiong, Z.; Liu, Z.; Hu, X.; Jiang, X. Angiopep-2/IP10-EGFRvIIIscFv modified nanoparticles and CTL synergistically inhibit malignant glioblastoma. Sci. Rep. 2018, 8, 12827. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Zhao, C.; Liu, P.; Wang, Z.; Ding, J.; Zhou, W. Facile construction of dual-targeting delivery system by using lipid capped polymer nanoparticles. R. Soc. Chem. 2018, 8, 444–453. [Google Scholar]
- Demeule, M.; Currie, J.C.; Bertrand, Y.; Ché, C.; Nguyen, T.; Régina, A.; Gabathuler, R.; Castaigne, J.P.; Béliveau, R. Involvement of the low-density lipoprotein receptor-related protein in the transcytosis of the brain delivery vector angiopep-2. J. Neurochem. 2008, 106, 1534–1544. [Google Scholar] [CrossRef]
- Xin, H.; Jiang, X.; Gu, J.; Sha, X.; Chen, L.; Law, K.; Chen, Y.; Wang, X.; Jiang, Y.; Fang, X. Biomaterials nanoparticles as dual-targeting drug delivery system for brain glioma. Biomaterials 2011, 32, 4293–4305. [Google Scholar] [CrossRef]
- Ke, W.; Shao, K.; Huang, R.; Han, L.; Liu, Y.; Li, J.; Kuang, Y.; Ye, L.; Lou, J.; Jiang, C. Biomaterials gene delivery targeted to the brain using an angiopep-conjugated polyethyleneglycol-modified polyamidoamine dendrimer. Biomaterials 2009, 30, 6976–6985. [Google Scholar] [CrossRef]
- Wang, L.; Hao, Y.; Li, H.; Zhao, Y.; Meng, D.; Li, D.; Shi, J.; Zhang, H.; Zhang, Z.; Zhang, Y. Co-delivery of doxorubicin and sirna for glioma therapy by a brain targeting system: Angiopep-2-Modified poly(Lactic-co-Glycolic Acid) nanoparticles. J. Drug Target. 2015, 23, 832–846. [Google Scholar] [CrossRef]
- Chen, G.J.; Su, Y.Z.; Hsu, C.; Lo, Y.L.; Huang, S.J.; Ke, J.H.; Kuo, Y.C.; Wang, L.F. Angiopep-Pluronic F127-Conjugated superparamagnetic Iron Oxide nanoparticles as nanotheranostic agents for BBB targeting. J. Mater. Chem. B 2014, 2, 5666–5675. [Google Scholar] [CrossRef]
- Tosi, G.; Vilella, A.; Veratti, P.; Belletti, D.; Pederzoli, F.; Ruozi, B.; Vandelli, M.A.; Zoli, M.; Forni, F. Exploiting bacterial pathways for bbb crossing with PLGA nanoparticles modified with a mutated form of diphtheria toxin (CRM197): In vivo experiments. Mol. Pharm. 2015, 12, 3672–3684. [Google Scholar] [CrossRef]
- Bi, C.; Wang, A.; Chu, Y.; Liu, S.; Mu, H.; Liu, W.; Wu, Z.; Sun, K.; Li, Y. Intranasal delivery of rotigotine to the brain with lactoferrin-modified PEG-PLGA nanoparticles for Parkinson’s disease treatment. Int. J. Nanomed. 2016, 11, 6547–6559. [Google Scholar] [CrossRef] [PubMed]
- Hoyos-Ceballos, G.P.; Sanchez-Giraldo, V.; Mendivil-Perez, M.; Jimenez-Del-Rio, M.; Sierra-Garcia, L.; Velez-Pardo, C.; Lopez-Osorio, B.L. Design of epigallocatechin gallate loaded PLGA/PF127 nanoparticles and their effect upon an oxidative stress model. J. Drug Deliv. Sci. Technol. 2018, 48, 152–160. [Google Scholar] [CrossRef]
- Hao, Y.; Wang, L.; Zhao, Y.; Meng, D.; Li, D.; Li, H.; Zhang, B.; Shi, J.; Zhang, H.; Zhang, Z.; et al. Targeted imaging and chemo-phototherapy of brain cancer by a multifunctional drug delivery system. Macromol. Boisci. 2015, 15, 1571–1585. [Google Scholar] [CrossRef]
- Hao, Y.; Zhang, B.; Zheng, C.; Ji, R.; Ren, X.; Guo, F.; Sun, S.; Shi, J.; Zhang, H.; Zhang, Z.; et al. The tumor-targeting core—Shell structured DTX-loaded PLGA @ Au nanoparticles for chemo-photothermal therapy and X-ray imaging. J. Control. Release 2015, 220, 545–555. [Google Scholar] [CrossRef] [PubMed]
- Duskey, J.; Tosi, G.; Oddone, N.; Ottonelli, F.; Pederzoli, I.; Vilella, A.; Zoli, M.; Kovachka, S.; Spyrakis, F.; Vandelli, M.A.; et al. Novel peptide-conjugated nanomedicines for brain targeting: In vivo evidences. Unpublished work. 2020. [Google Scholar]
- Lowe, A.B. Thiol-ene ‘Click’ reactions and recent applications in polymer and materials synthesis: A. first update. R. Soc. Chem. 2014, 5, 4820–4870. [Google Scholar] [CrossRef]
- Tosi, G.; Costantino, L.; Rivasi, F.; Ruozi, B.; Leo, E.; Vergoni, A.V.; Tacchi, R.; Bertolini, A.; Vandelli, M.A.; Forni, F. Targeting the central nervous system: In vivo experiments with peptide-derivatized nanoparticles loaded with loperamide and rhodamine-123. J. Control. Release 2007, 122, 1–9. [Google Scholar] [CrossRef]
- Costantino, L.; Gandolfi, F.; Tosi, G.; Rivasi, F.; Vandelli, M.A.; Forni, F. Peptide-Derivatized biodegradable nanoparticles able to cross the blood-brain barrier. J. Control. Release 2005, 108, 84–96. [Google Scholar] [CrossRef]
- Vasconcelos, A.; Vega, E.; Pérez, Y.; Gomara, J.M.; García, M.L.; Haro, I. Conjugation of cell-penetrating peptides with poly(Lactic-co-Glycolic Acid)-polyethylene glycol nanoparticles improves ocular drug delivery. Int. J. Nanomed. 2015, 10, 609–631. [Google Scholar]
- Hu, K.; Shi, Y.; Jiang, W.; Han, J.; Huang, S.; Jiang, X. Lactoferrin Conjugated PEG-PLGA Nanoparticles for Brain Delivery: Preparation, Characterization and Efficacy in Parkinsons Disease. Int. J. Pharm. 2011, 415, 273–283. [Google Scholar] [CrossRef]
- Kastin, A.J. Handbook of Biologically Active Peptides; Academic Press: Cambridge, MA, USA, 2013. [Google Scholar]
- Huang, N.; Lu, S.; Liu, X.; Zhu, J.; Wang, Y. PLGA Nanoparticles Modified with a BBB-Penetrating Peptide co-Delivering Aβ Generation Inhibitor and Curcumin Attenuate Memory Deficits and Neuropathology in Alzheimer’ s Disease Mice. Oncotarget 2017, 8, 81001–81013. [Google Scholar] [PubMed]
- Tosi, G.; Vergoni, A.V.; Ruozi, B.; Bondioli, L.; Badiali, L.; Rivasi, F.; Costantino, L.; Forni, F.; Vandelli, M.A. Sialic acid and glycopeptides conjugated PLGA nanoparticles for central nervous system targeting: In vivo pharmacological evidence and biodistribution. J. Control. Release 2010, 145, 49–57. [Google Scholar] [CrossRef]
- Heggannavar, G.B.; Vijeth, S.; Kariduraganavar, M.Y. Development of Dual Drug Loaded PLGA Based Mesoporous Silica Nanoparticles and Their Conjugation With Angiopep-2 to Treat Glioma. J. Drug Deliv. Sci. Technol. 2019, 53, 101157. [Google Scholar] [CrossRef]
- SShen, J.; Zhan, C.; Xie, C.; Meng, Q.; Gu, B.; Li, C.; Zhang, Y.; Lu, W. Poly(Ethylene Glycol)-Block-Poly (d,l-Lactide Acid) Micelles Anchored With Angiopep-2 for Brain-Targeting Delivery. J. Drug Target. 2011, 19, 197–203. [Google Scholar] [CrossRef] [PubMed]
| Initial Amount PLGA-Mal/Ang–2 | µg Ang–2/g NPs | Final Molar Ratio Ang–2/PLGA-Mal |
|---|---|---|
| Control (3:1) | 1.65 ± 0.60 | |
| 3:1 | 3.06 ± 0.11 | 0.25 ± 0.01 |
| Control (2:1) | 2.59 ± 0.71 * | |
| 2:1 | 4.42 ± 0.74 * | 0.37 ± 0.06 |
| Control (1:1) | 4.24 ± 0.71 ** | |
| 1:1 | 8.78 ± 1.93 ** | 0.73 ± 0.23 |
| PLGA-b-PEG Formulations | Particle Size (nm) | PDI | Zeta Potential (mV) |
|---|---|---|---|
| Non-functionalized NPs | 136.3 ± 6.8 | 0.06 ± 0.01 | −27.4 ± 2.7 |
| Pre-Formulation Ang–2 NPs | 166.4 ± 2.4 | 0.08 ± 0.04 | −26.2 ± 0.9 |
| Post-Formulation Ang–2 NPs | 177.3 ± 12.7 | 0.10 ± 0.01 | −21.9 ± 3.4 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hoyos-Ceballos, G.P.; Ruozi, B.; Ottonelli, I.; Da Ros, F.; Vandelli, M.A.; Forni, F.; Daini, E.; Vilella, A.; Zoli, M.; Tosi, G.; et al. PLGA-PEG-ANG-2 Nanoparticles for Blood–Brain Barrier Crossing: Proof-of-Concept Study. Pharmaceutics 2020, 12, 72. https://doi.org/10.3390/pharmaceutics12010072
Hoyos-Ceballos GP, Ruozi B, Ottonelli I, Da Ros F, Vandelli MA, Forni F, Daini E, Vilella A, Zoli M, Tosi G, et al. PLGA-PEG-ANG-2 Nanoparticles for Blood–Brain Barrier Crossing: Proof-of-Concept Study. Pharmaceutics. 2020; 12(1):72. https://doi.org/10.3390/pharmaceutics12010072
Chicago/Turabian StyleHoyos-Ceballos, Gina P., Barbara Ruozi, Ilaria Ottonelli, Federica Da Ros, Maria Angela Vandelli, Flavio Forni, Eleonora Daini, Antonietta Vilella, Michele Zoli, Giovanni Tosi, and et al. 2020. "PLGA-PEG-ANG-2 Nanoparticles for Blood–Brain Barrier Crossing: Proof-of-Concept Study" Pharmaceutics 12, no. 1: 72. https://doi.org/10.3390/pharmaceutics12010072
APA StyleHoyos-Ceballos, G. P., Ruozi, B., Ottonelli, I., Da Ros, F., Vandelli, M. A., Forni, F., Daini, E., Vilella, A., Zoli, M., Tosi, G., Duskey, J. T., & López-Osorio, B. L. (2020). PLGA-PEG-ANG-2 Nanoparticles for Blood–Brain Barrier Crossing: Proof-of-Concept Study. Pharmaceutics, 12(1), 72. https://doi.org/10.3390/pharmaceutics12010072








