Plant-Produced Glycosylated and In Vivo Deglycosylated Receptor Binding Domain Proteins of SARS-CoV-2 Induce Potent Neutralizing Responses in Mice
Abstract
1. Introduction
2. Materials and Methods
2.1. Cloning and Expression RBD Proteins in N. benthamiana
2.2. Expression Screening of RBD Proteins Produced in N. benthamiana Plant by Western Blot Analysis
2.3. Purification of Plant-Produced gRBD and dRBD Proteins Using Anti-DYKDDDDK Affinity Gel
2.4. Gel Filtration
2.5. Glycoprotein Detection
2.6. Binding Affinity of the Plant-Produced RBD Variants to ACE2
2.7. Stability Assessment of Plant-Produced RBD Variants
2.8. Immunogenicity Studies of gRBD and dRBD Proteins in Mice
2.9. Virus Titration
2.10. Micro Neutralization Test (MNT)
2.11. Statistical Analysis
3. Results
3.1. Engineering, Cloning, Expression, Purification, and Characterization of RBD Proteins from N. benthamiana
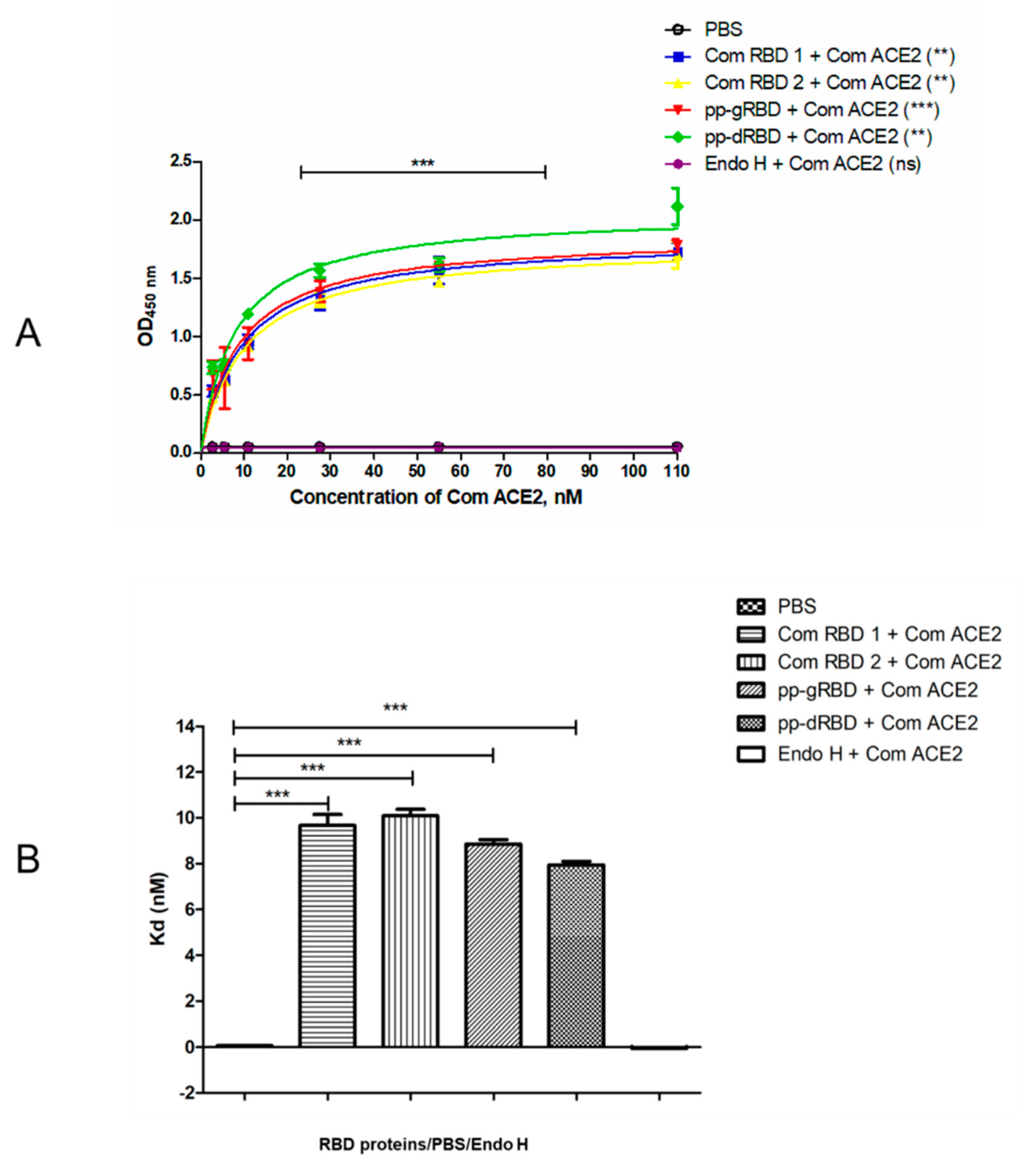
3.2. Binding of Plant-Produced RBD Variants to ACE2
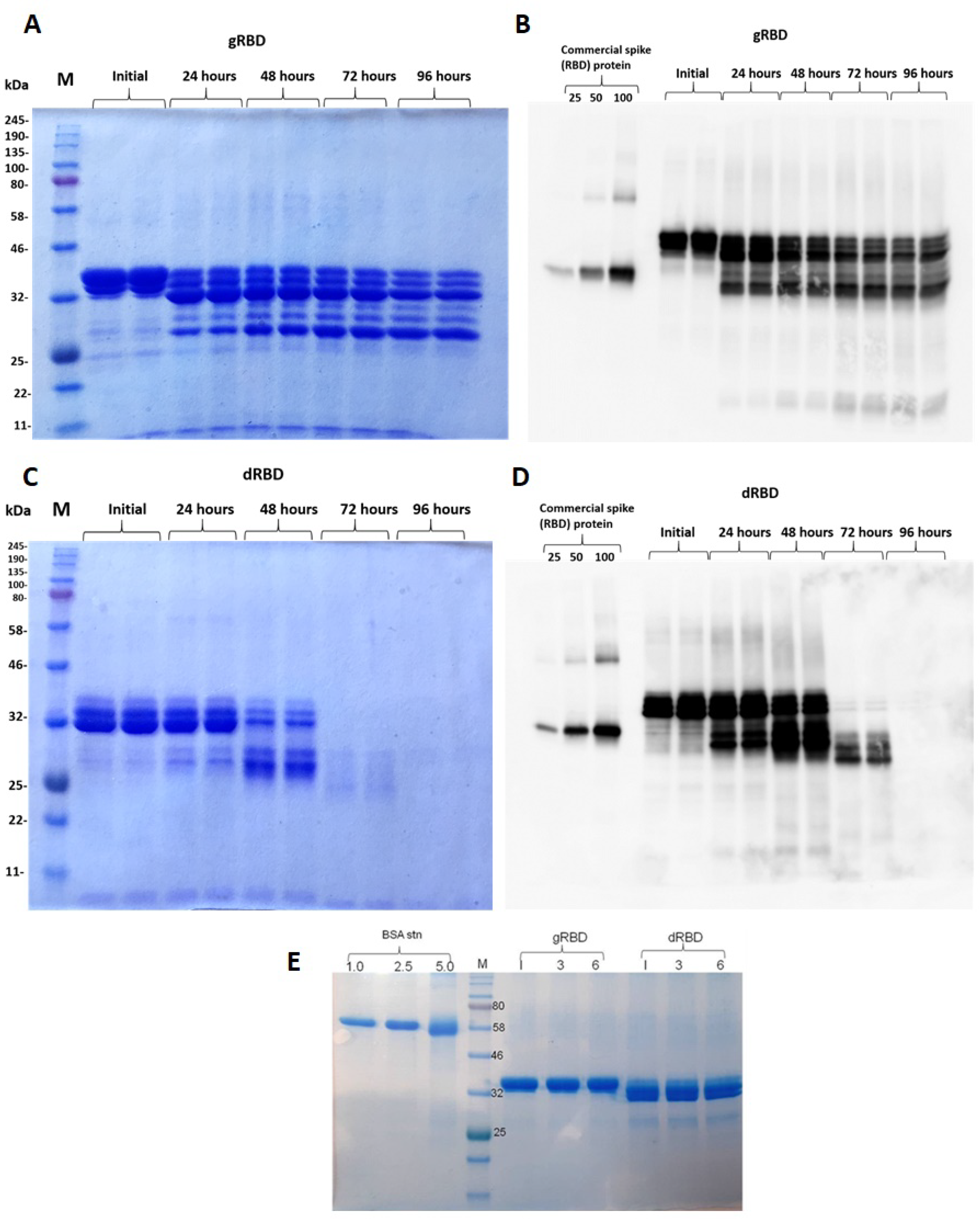
3.3. Stability Assessment of RBD Proteins
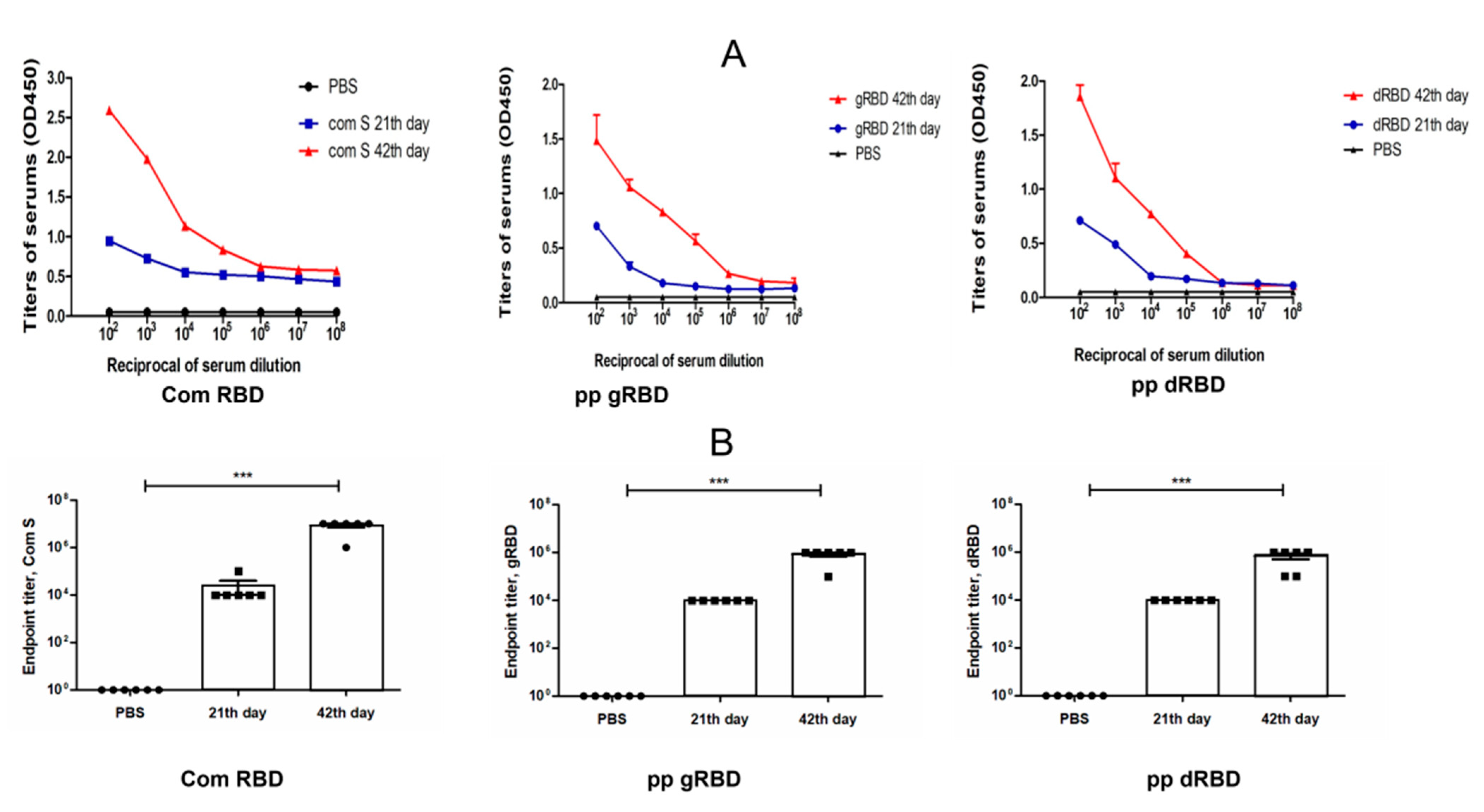
3.4. Immunogenicity Studies of gRBD and dRBD Variants in Mice
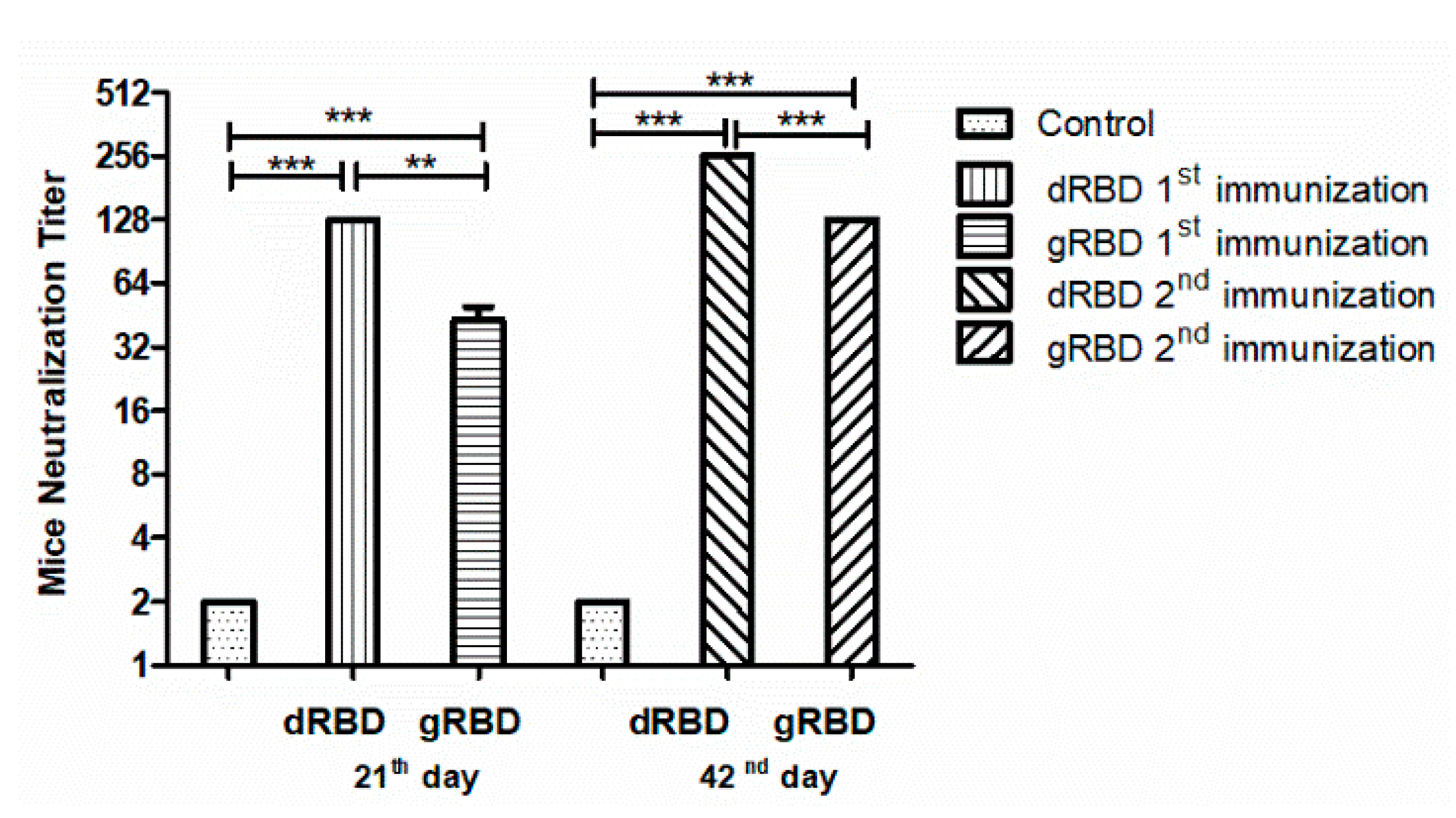
3.5. Neutralization Activity Assessment of gRBD and dRBD by Micro Neutralization Test
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Gralinski, L.E.; Menachery, V.D. Return of the Coronavirus: 2019-nCoV. Viruses 2020, 12, 135. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Wang, R.; Wang, M.; Wei, G.W. Mutations Strengthened SARS-CoV-2 Infectivity. J. Mol. Biol. 2020, 432, 5212–5226. [Google Scholar] [CrossRef] [PubMed]
- Duchene, S.; Featherstone, L.; Haritopoulou-Sinanidou, M.; Rambaut, A.; Philippe, L.; Baele, G. Temporal signal and the phylodynamic threshold of SARS-CoV-2. Virus Evol. 2020, 6, 2. [Google Scholar] [CrossRef] [PubMed]
- Worobey, M.; Pekar, J.; Larsen, B.B.; Nelson, M.I.; Hill, V.; Joy, J.B.; Rambaut, A.; Suchard, M.A.; Wertheim, J.O.; Lemey, P. The emergence of SARS-CoV-2 in Europe and North America. Science 2020, 370, 564–570. [Google Scholar] [CrossRef] [PubMed]
- Hou, Y.J.; Chiba, S.; Halfmann, P.; Ehre, C.; Kuroda, M.; Dinnon, K.H.; Leist, S.R.; Schafer, A.; Nakajima, N.; Takahashi, K.; et al. SARS-CoV-2 D614G variant exhibits efficient replication ex vivo and transmission in vivo. Science 2020, 370, 1464–1468. [Google Scholar] [CrossRef]
- Weissman, D.; Alameh, M.G.; de Silva, T.; Collini, P.; Hornsby, H.; Brown, R.; LaBranche, C.C.; Edwards, R.J.; Sutherland, L.; Santra, S.; et al. D614G Spike Mutation Increases SARS CoV-2 Susceptibility to Neutralization. Cell Host Microbe. 2021, 29, 23–31. [Google Scholar] [CrossRef] [PubMed]
- Yurkovetskiy, L.; Wang, X.; Pascal, K.E.; Tomkins-Tinch, C.; Nyalile, T.P.; Wang, Y.; Baum, A.; Diehl, W.E.; Dauphin, A.; Carbone, C.; et al. Structural and Functional Analysis of the D614G SARS-CoV-2 Spike Protein Variant. Cell 2020, 183, 739–751. [Google Scholar] [CrossRef]
- Shang, J.; Ye, G.; Shi, K.; Wan, Y.; Luo, C.; Aihara, H.; Geng, Q.; Auerbach, A.; Li, F. Structural basis of receptor recognition by SARS-CoV-2. Nature 2020, 581, 221–224. [Google Scholar] [CrossRef] [PubMed]
- Wan, Y.; Shang, J.; Graham, R.; Baric, R.S.; Li, F. Receptor Recognition by the Novel Coronavirus from Wuhan: An Analysis Based on Decade-Long Structural Studies of SARS Coronavirus. J. Virol. 2020, 94, e00127-20. [Google Scholar] [CrossRef]
- Thomson, E.C.; Rosen, L.E.; Shepherd, J.G.; Spreafico, R.; da Silva Filipe, A.; Wojcechowskyj, J.A.; Davis, C.; Piccoli, L.; Pascall, D.J.; Dillen, J.; et al. Circulating SARS-CoV-2 spike N439K variants maintain fitness while evading antibody-mediated immunity. Cell 2021, 184, 1171–1187. [Google Scholar] [CrossRef]
- Baum, A.; Fulton, B.O.; Wloga, E.; Copin, R.; Pascal, K.E.; Russo, V.; Giordano, S.; Lanza, K.; Negron, N.; Ni, M.; et al. Antibody cocktail to SARS-CoV-2 spike protein prevents rapid mutational escape seen with individual antibodies. Science 2020, 369, 1014–1018. [Google Scholar] [CrossRef] [PubMed]
- Gaebler, C.; Wang, Z.; Lorenzi, J.C.; Muecksch, F.; Finkin, S.; Tokuyama, M.; Cho, A.; Jankovic, M.; Schaefer-Babajew, D.; Oliveira, T.Y.; et al. Evolution of Antibody Immunity to SARS-CoV-2. Nature 2020, 591, 639–644. [Google Scholar] [CrossRef]
- Greaney, A.J.; Starr, T.N.; Gilchuk, P.; Zost, S.J.; Binshtein, E.; Loes, A.N.; Hilton, S.K.; Huddleston, J.; Eguia, R.; Crawford, K.H.D.; et al. Complete mapping of mutations to the SARS-CoV-2 spike receptor-binding domain that escape antibody recognition. Cell Host Microbe. 2021, 29, 44–57. [Google Scholar] [CrossRef]
- Li, Q.; Wu, J.; Nie, J.; Zhang, L.; Hao, H.; Liu, S.; Zhao, C.; Zhang, Q.; Liu, H.; Nie, L.; et al. The Impact of Mutations in SARS-CoV-2 Spike on Viral Infectivity and Antigenicity. Cell 2020, 182, 1284–1294. [Google Scholar] [CrossRef] [PubMed]
- Planas, D.; Veyer, D.; Baidaliuk, A.; Staropoli, I.; Guivel-Benhassine, F.; Rajah, M.M.; Planchais, C.; Porrot, F.; Robillard, N.; Puech, J.; et al. Reduced sensitivity of SARS-CoV-2 variant Delta to antibody neutralization. Nature 2021, 596, 276–280. [Google Scholar] [CrossRef]
- Guruprasad, L. Human SARS CoV-2 spike protein mutations. Proteins 2021, 89, 569–576. [Google Scholar] [CrossRef]
- Moreira, R.A.; Chwastyk, M.; Baker, J.L.; Guzman, H.V.; Poma, A.B. Quantitative determination of mechanical stability in the novel coronavirus spike protein. Nanoscale 2020, 12, 16409–16413. [Google Scholar] [CrossRef]
- Liu, W.; Liu, L.; Kou, G.; Zheng, Y.; Ding, Y.; Ni, W.; Wang, Q.; Tan, L.; Wu, W.; Tang, S.; et al. Evaluation of nucleocapsid and spike protein-based enzyme-linked immunosorbent assays for detecting antibodies against SARS-CoV-2. J. Clin. Microbiol. 2020, 58, e00461-20. [Google Scholar] [CrossRef]
- Zhu, F.C.; Li, Y.H.; Guan, X.H.; Hou, L.H.; Wang, W.J.; Li, J.X.; Wu, S.P.; Wang, B.S.; Wang, Z.; Wang, L.; et al. Safety, tolerability, and immunogenicity of a recombinant adenovirus type-5 vectored COVID-19 vaccine: A dose-escalation, open-label, non-randomised, first-in-human trial. Lancet 2020, 395, 1845–1854. [Google Scholar] [CrossRef]
- Jackson, L.A.; Anderson, E.J.; Rouphael, N.G.; Roberts, P.C.; Makhene, M.; Coler, R.N.; McCullough, M.P.; Chappell, J.D.; Denison, M.R.; Stevens, L.J.; et al. An mRNA vaccine against SARS-CoV-2—Preliminary report. N. Engl. J. Med. 2020, 383, 1920–1931. [Google Scholar] [CrossRef]
- Smith, T.R.; Patel, A.; Ramos, S.; Elwood, D.; Zhu, X.; Yan, J.; Gary, E.N.; Walker, S.N.; Schultheis, K.; Purwar, M.; et al. Immunogenicity of a DNA vaccine candidate for COVID-19. Nat. Commun. 2020, 11, 1–13. [Google Scholar] [CrossRef]
- Van Doremalen, N.; Lambe, T.; Spencer, A.; Belij-Rammerstorfer, S.; Purushotham, J.N.; Port, J.R.; Avanzato, V.A.; Bushmaker, T.; Flaxman, A.; Ulaszewska, M.; et al. ChAdOx1 nCoV-19 vaccination prevents SARS-CoV-2 pneumonia in rhesus macaques. Nature 2020, 586, 578–582. [Google Scholar] [CrossRef]
- Wrapp, D.; Wang, N.; Corbett, K.S.; Goldsmith, J.A.; Hsieh, C.L.; Abiona, O.; Graham, B.S.; McLellan, J.S. Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation. Science 2020, 367, 1260–1263. [Google Scholar] [CrossRef] [PubMed]
- Walls, A.C.; Park, Y.J.; Tortorici, M.A.; Wall, A.; McGuire, A.T.; Veesler, D. Structure, function, and antigenicity of the SARS-CoV-2 spike glycoprotein. Cell 2020, 181, 281–292. [Google Scholar] [CrossRef] [PubMed]
- Gui, M.; Song, W.; Zhou, H.; Xu, J.; Chen, S.; Xiang, Y.; Wang, X. Cryo-electron microscopy structures of the SARS-CoV spike glycoprotein reveal a prerequisite conformational state for receptor binding. Cell Res. 2017, 27, 119–129. [Google Scholar] [CrossRef] [PubMed]
- Fallon, L.; Belfon, K.A.A.; Raguette, L.; Wang, Y.; Corbo, C.; Stepanenko, D.; Cuomo, A.; Guerra, J.; Budhan, S.; Varghese, S.; et al. Free Energy Landscapes for RBD Opening in SARS-CoV-2 Spike Glycoprotein Simulations Suggest Key Interactions and a Potentially Druggable Allosteric Pocket. J. Am. Chem. Soc. 2021, 143, 11349–11360. [Google Scholar] [CrossRef]
- Zhou, D.; Tian, X.; Qi, R.; Peng, C.; Zhang, W. Identification of 22 N-glycosites on spike glycoprotein of SARS-CoV-2 and accessible surface glycopeptide motifs: Implications for vaccination and antibody therapeutics. Glycobiology 2020, 31, 69–80. [Google Scholar] [CrossRef]
- Casalino, L.; Gaieb, Z.; Goldsmith, J.A.; Hjorth, C.K.; Dommer, A.C.; Harbison, A.M.; Fogarty, C.A.; Barros, E.P.; Taylor, B.C.; McLellan, J.S.; et al. Beyond Shielding: The Roles of Glycans in the SARS-CoV-2 Spike Protein. ACS Cent. Sci. 2020, 28, 1722–1734. [Google Scholar] [CrossRef]
- Shin, Y.J.; König-Beihammer, J.; Vavra, U.; Schwestka, J.; Kienzl, N.F.; Klausberger, M.; Laurent, E.; Grünwald-Gruber, C.; Vierlinger, K.; Hofner, M.; et al. N-Glycosylation of the SARS-CoV-2 Receptor Binding Domain Is Important for Functional Expression in Plants. Front. Plant Sci. 2021, 12, 689104. [Google Scholar] [CrossRef]
- Cai, Y.; Zhang, J.; Xiaol, T.; Peng, H.; Sterling, S.M.; Walsh, R.M., Jr.; Rawson, S.; Rits-Volloch, S.; Chen, B. Distinct conformational states of SARS-CoV-2 spike protein. Science 2020, 369, 1586–1592. [Google Scholar] [CrossRef]
- Siriwattananon, K.; Manopwisedjaroen, S.; Shanmugaraj, B.; Rattanapisit, K.; Phumiamorn, S.; Sapsutthipas, S.; Trisiriwanich, S.; Prompetchara, E.; Ketloy, C.; Buranapraditkun, S.; et al. Plant-Produced Receptor-Binding Domain of SARS-CoV-2 Elicits Potent Neutralizing Responses in Mice and Non-human Primates. Front. Plant Sci. 2021, 13, 682953. [Google Scholar] [CrossRef] [PubMed]
- Mamedov, T.; Ghosh, A.; Jones, R.M.; Mett, V.; Farrance, C.E.; Musiychuk, K.; Horsey, A.; Yusibov, V. Production of non-glycosylated recombinant proteins in Nicotiana benthamiana plants by co-expressing bacterial PNGase F. Plant Biotechnol. J. 2012, 10, 773–782. [Google Scholar] [CrossRef] [PubMed]
- Streatfield, S.J.; Kushnir, N.; Yusibov, V. Plant-produced candidate countermeasures against emerging and reemerging infections and bioterror agents. Plant Biotechnol. J. 2015, 13, 1136–1159. [Google Scholar] [CrossRef] [PubMed]
- Prokhnevsky, A.; Mamedov, T.; Leffet, B.; Rahimova, R.; Ghosh, A.; Mett, V.; Yusibov, V. Development of a single-replicon miniBYV vector for co-expression of heterologous proteins. Mol. Biotechnol. 2015, 57, 101–110. [Google Scholar] [CrossRef] [PubMed]
- Mamedov, T.; Chichester, J.A.; Jones, R.M.; Ghosh, A.; Coffin, M.V.; Herschbach, K.; Prokhnevsky, A.I.; Streatfield, S.J.; Yusibov, V. Production of functionally active and immunogenic non-glycosylated protective antigen from Bacillus anthracis in Nicotiana benthamiana by co-expression with peptide-N-glycosidase F (PNGase F) of Flavobacterium meningosepticum. PLoS ONE 2016, 11, e0153956. [Google Scholar] [CrossRef]
- Mamedov, T.; Cicek, K.; Gulec, B.; Ungor, R.; Hasanova, G. In vivo production of non-glycosylated recombinant proteins in Nicotiana benthamiana plants by co-expression with Endo-β-N-acetylglucosaminidase H (Endo H) of Streptomyces plicatus. PLoS ONE 2017, 12, e0183589. [Google Scholar] [CrossRef]
- Mamedov, T.; Musayeva, I.; Acsora, R.; Gun, N.; Gulec, B.; Mammadova, G.; Cicek, K.; Hasanova, G. Engineering, and production of functionally active human Furin in N. benthamiana plant: In vivo post-translational processing of target proteins by Furin in plants. PLoS ONE 2019, 14, e0213438. [Google Scholar] [CrossRef]
- Mamedov, T.; Cicek, K.; Miura, K.; Gulec, B.; Akinci, E.; Mammadova, G.; Hasanova, G. A Plant-Produced in vivo deglycosylated full-length Pfs48/45 as a Transmission-Blocking Vaccine Candidate against malaria. Sci. Rep. 2019, 9, 1–12. [Google Scholar] [CrossRef]
- Montero-Morales, L.; Maresch, D.; Crescioli, S.; Castilho, A.; Ilieva, K.M.; Mele, S.; Karagiannis, S.N.; Altmann, F.; Steinkellner, H. In planta glycan engineering and functional activities of IgE antibodies. Front. Bioeng. Biotechnol. 2019, 7, 242. [Google Scholar] [CrossRef]
- Margolin, E.; Strasser, R.; Chapman, R.; Williamson, A.L.; Rybicki, E.P.; Meyers, A.E. Engineering the plant secretory pathway for the production of next-generation pharmaceuticals. Trends Biotechnol. 2020, 38, 1034–1044. [Google Scholar] [CrossRef]
- Shanmugaraj, B.; Bulaon, C.J.; Phoolcharoen, W. Plant molecular farming: A viable platform for recombinant biopharmaceutical production. Plants 2020, 9, 842. [Google Scholar] [CrossRef]
- Pogrebnyak, N.; Golovkin, M.; Andrianov, V.; Spitsin, S.; Smirnov, Y.; Egolf, R.; Koprowski, H. Severe acute respiratory syndrome (SARS) S protein production in plants: Development of recombinant vaccine. Proc. Natl. Acad. Sci. USA 2005, 102, 9062–9067. [Google Scholar] [CrossRef] [PubMed]
- Zhou, J.Y.; Wu, J.X.; Cheng, L.Q.; Zheng, X.J.; Gong, H.; Shang, S.B.; Zhou, E.M. Expression of immunogenic S1 glycoprotein of infectious bronchitis virus in transgenic potatoes. J. Virol. 2003, 77, 9090–9093. [Google Scholar] [CrossRef][Green Version]
- Bae, J.L.; Lee, J.G.; Kang, T.J.; Jang, H.S.; Jang, Y.S.; Yang, M.S. Induction of antigen-specific systemic and mucosal immune responses by feeding animals transgenic plants expressing the antigen. Vaccine 2003, 21, 4052–4058. [Google Scholar] [CrossRef]
- Lamphear, B.J.; Jilka, J.M.; Kesl, L.; Welter, M.; Howard, J.A.; Streatfield, S.J. A corn-based delivery system for animal vaccines: An oral transmissible gastroenteritis virus vaccine boosts lactogenic immunity in swine. Vaccine 2004, 22, 2420–2424. [Google Scholar] [CrossRef] [PubMed]
- Li, H.Y.; Ramalingam, S.; Chye, M.L. Accumulation of recombinant SARS-CoV spike protein in plant cytosol and chloroplasts indicate potential for development of plant-derived oral vaccines. Exp. Biol. Med. 2006, 231, 1346–1352. [Google Scholar] [CrossRef] [PubMed]
- Yusibov, V.; Mamedov, T. Plants as an alternative system for expression of vaccine antigens. Proc. ANAS Biol. Sci. 2010, 65, 195–200. [Google Scholar]
- Rattanapisit, K.; Shanmugaraj, B.; Manopwisedjaroen, S.; Purwono, P.B.; Siriwattananon, K.; Khorattanakulchai, N.; Hanittinan, O.; Boonyayothin, W.; Thitithanyanont, A.; Smith, D.R.; et al. Rapid production ofSARS-CoV-2 receptor binding domain (RBD) and spike specific monoclonal antibody CR3022 in Nicotiana Benthamiana. Sci. Rep. 2020, 10, 17698. [Google Scholar] [CrossRef] [PubMed]
- Sainsbury, F.; Thuenemann, E.C.; Lomonossoff, G.P. pEAQ: Versatile expression vectors for easy and quick transient expression of heterologous proteins in plants. Plant. Biotechnol. J. 2009, 7, 682–693. [Google Scholar] [CrossRef]
- Pavel, S.T.I.; Yetiskin, H.; Aydin, G.; Holyavkin, C.; Uygut, M.A.; Dursun, Z.B.; Celik, İ.; Cevik, C.; Ozdarendeli, A. Isolation and characterization of severe acute respiratory syndrome coronavirus 2 in Turkey. PLoS ONE 2020, 15, e0238614. [Google Scholar] [CrossRef]
- Li, F. Receptor recognition and cross-species infections of SARS coronavirus. Antivir. Res. 2013, 100, 246–254. [Google Scholar] [CrossRef]
- Lei, C.; Qian, K.; Li, T.; Zhang, S.; Fu, W.; Ding, M.; Hu, S. Neutralization of SARS-CoV-2 spike pseudotyped virus by recombinant ACE2-Ig. Nat. Commun. 2020, 11, 2070. [Google Scholar] [CrossRef]
- Quinlan, B.D.; Mou, H.; Zhang, L.; Guo, Y.; He, W.; Ojha, A.; Parcells, M.S.; Luo, G.; Li, W.; Zhong, G.; et al. The SARS-CoV-2 receptor-binding domain elicits a potent neutralizing response without antibody-dependent enhancement. SSRN Electron. J. 2020. [Google Scholar] [CrossRef]
- Rabi, F.A.; Al Zoubi, M.S.; Kasasbeh, G.A.; Salameh, D.M.; Al-Nasser, A.D. SARS-CoV-2 and Coronavirus Disease 2019: What We Know So Far. Pathogens 2020, 9, 231. [Google Scholar] [CrossRef]
- Sun, J.; Aghemo, A.; Forner, A.; Valenti, L. COVID-19 and liver disease. Liver Int. 2020, 40, 1278–1281. [Google Scholar] [CrossRef]
- Xie, X.; Xue, Q.; Zhou, Y.; Zhu, K.; Liu, Q.; Zhang, J.; Song, R. Mental Health Status among Children in Home Confinement during the Coronavirus Disease 2019 Outbreak in Hubei Province, China. JAMA Pediatr. 2020, 174, 898–900. [Google Scholar] [CrossRef] [PubMed]
- Yuki, K.; Fujiogi, M.; Koutsogiannaki, S. COVID-19 pathophysiology: A review. Clin. Immunol. 2020, 215, 108427. [Google Scholar] [CrossRef]
- Tai, W.; He, L.; Zhang, X.; Pu, J.; Voronin, D.; Jiang, S.; Zhou, Y.; Du, L. Characterization of the receptor-binding domain (RBD) of 2019 novel coronavirus: Implication for development of RBD protein as a viral attachment inhibitor and vaccine. Cell. Mol. Immunol. 2020, 17, 613–620. [Google Scholar] [CrossRef] [PubMed]
- He, Y.; Lu, H.; Siddiqui, P.; Zhou, Y.; Jiang, S. Receptor-binding domain of severe acute respiratory syndrome coronavirus spike protein contains multiple conformation-dependent epitopes that induce highly potent neutralizing antibodies. J. Immunol. 2005, 174, 4908–4915. [Google Scholar] [CrossRef]
- Zhu, X.; Liu, Q.; Du, L.; Lu, L.; Jiang, S. Receptor-binding domain as a target for developing SARS vaccines. J. Thorac. Dis. 2013, 5, 142–148. [Google Scholar]
- Lan, J.; Ge, J.; Yu, J.; Shan, S.; Zhou, H.; Fan, S.; Zhang, Q.; Shi, X.; Wang, Q.; Zhang, L.; et al. Structure of the SARS-CoV-2 spike receptor-binding domain bound to the ACE2 receptor. Nature 2020, 581, 215–220. [Google Scholar] [CrossRef]
- Wang, L.; Shi, W.; Chappell, J.D.; Joyce, M.G.; Zhang, Y.; Kanekiyo, M.; Becker, M.M.; van Doremalen, N.; Fischer, R.; Wang, N.; et al. Importance of neutralizing monoclonal antibodies targeting multiple antigenic sites on the middle east respiratory syndrome coronavirus spike glycoprotein to avoid neutralization escape. J. Virol. 2018, 92, e02002-17. [Google Scholar] [CrossRef]
- Yan, R.; Zhang, Y.; Li, Y.; Xia, L.; Guo, Y.; Zhou, Q. Structural basis for the recognition of SARS-CoV-2 by full-length human ACE2. Science 2020, 367, 1444–1448. [Google Scholar] [CrossRef]
- Shajahan, A.; Archer-Hartmann, S.; Supekar, N.T.; Gleinich, A.S.; Heiss, C.; Azadi, P. Comprehensive characterization of N- and O- glycosylation of SARS-CoV-2 human receptor angiotensin converting enzyme 2. Glycobiology 2020, 31, 410–424. [Google Scholar] [CrossRef] [PubMed]
- Watanabe, Y.; Allen, J.D.; Wrapp, D.; McLellan, J.S.; Crispin, M. Site-specific glycan analysis of the SARS-CoV-2 spike. Science 2020, 369, 330–333. [Google Scholar] [CrossRef] [PubMed]
- Zhang, N.; Jiang, S.; Du, L. Current advancements and potential strategies in the development of MERS-CoV vaccines. Expert Rev. Vaccines 2014, 13, 761–774. [Google Scholar] [CrossRef]
- Liu, D.X.; Fung, T.S.; Chong, K.K.; Shukla, A.; Hilgenfeld, R. Accessory proteins of SARS-CoV and other coronaviruses. Antivir. Res. 2014, 109, 97–109. [Google Scholar] [CrossRef] [PubMed]
- Dai, L.; Zheng, T.; Xu, K.; Han, Y.; Xu, L.; Huang, E.; An, Y.; Cheng, Y.; Li, S.; Liu, M.; et al. A Universal Design of Betacoronavirus Vaccines against COVID-19, MERS, and SARS. Cell 2020, 182, 722–733. [Google Scholar] [CrossRef]
- Mamedov, T.; Yuksel, D.; Ilgin, M.; Gurbuzaslan, I.; Gulec, B.; Mammadova, G.; Say, D.; Hasanova, G. Engineering, production and characterization of Spike and Nucleocapsid structural proteins of SARS–CoV-2 in Nicotiana benthamiana as vaccine candidates against COVID-19. bioRxiv 2020. [Google Scholar] [CrossRef]
- Grifoni, A.; Sidney, J.; Zhang, Y.; Scheuermann, R.H.; Peters, B.; Sette, A. A sequence homology and bioinformatic approach can predict candidate targets for immune responses to SARS-CoV-2. Cell Host Microbe 2020, 27, 671–680. [Google Scholar] [CrossRef]
- Siriwattananon, K.; Manopwisedjaroen, S.; Kanjanasirirat, P.; Purwono, P.B.; Rattanapisit, K.; Shanmugaraj, B.; Smith, D.R.; Borwornpinyo, S.; Thitithanyanont, A.; Phoolcharoen, W. Development of Plant-Produced Recombinant ACE2-Fc Fusion Protein as a Potential Therapeutic Agent Against SARS-CoV-2. Front. Plant Sci. 2021, 11, 604663. [Google Scholar] [CrossRef] [PubMed]
- Mamedov, T.; Gurbuzaslan, I.; Ilgin, M.; Yuksel, D.; Mammadova, G.; Ozkul, A.; Hasanova, G. High level production and characterization of truncated human angiotensin converting enzyme 2 in Nicotiana benthamiana plant as a potential therapeutic target in COVID-19. bioRxiv 2021. [Google Scholar] [CrossRef]
- Diego-Martin, B.; González, B.; Vazquez-Vilar, M.; Selma, S.; Mateos-Fernández, R.; Gianoglio, S.; Fernández-del-Carmen, A.; Orzáez, D. Pilot production of SARS-CoV-2 related proteins in plants: A proof of concept for rapid repurposing of indoor farms into biomanufacturing facilities. Front. Plant Sci. 2020, 11, 612781. [Google Scholar] [CrossRef]
- Stronge, V.S.; Saito, Y.; Ihara, Y.; Williams, D.B. Relationship between calnexin and BiP in suppressing aggregation and promoting refolding of protein and glycoprotein substrates. J. Biol. Chem. 2001, 276, 39779–39787. [Google Scholar] [CrossRef] [PubMed]
- Gomord, V.; Fitchette, A.C.; Menu-Bouaouiche, L.; Saint-Jore-Dupas, C.; Plasson, C.; Michaud, D.; Faye, L. Plant-specific glycosylation patterns in the context of therapeutic protein production. Plant Biotechnol. J. 2010, 8, 564–587. [Google Scholar] [CrossRef]
- Antonopoulos, A.; Broome, S.; Sharov, V.; Ziegenfuss, C.; Easton, R.L.; Panico, M.; Dell, A.; Morris, H.R.; Haslam, S.M. Site-specific characterization of SARS-CoV-2 spike glycoprotein receptor-binding domain. Glycobiology 2021, 31, 181–187. [Google Scholar] [CrossRef] [PubMed]



Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mamedov, T.; Yuksel, D.; Ilgın, M.; Gurbuzaslan, I.; Gulec, B.; Yetiskin, H.; Uygut, M.A.; Islam Pavel, S.T.; Ozdarendeli, A.; Mammadova, G.; et al. Plant-Produced Glycosylated and In Vivo Deglycosylated Receptor Binding Domain Proteins of SARS-CoV-2 Induce Potent Neutralizing Responses in Mice. Viruses 2021, 13, 1595. https://doi.org/10.3390/v13081595
Mamedov T, Yuksel D, Ilgın M, Gurbuzaslan I, Gulec B, Yetiskin H, Uygut MA, Islam Pavel ST, Ozdarendeli A, Mammadova G, et al. Plant-Produced Glycosylated and In Vivo Deglycosylated Receptor Binding Domain Proteins of SARS-CoV-2 Induce Potent Neutralizing Responses in Mice. Viruses. 2021; 13(8):1595. https://doi.org/10.3390/v13081595
Chicago/Turabian StyleMamedov, Tarlan, Damla Yuksel, Merve Ilgın, Irem Gurbuzaslan, Burcu Gulec, Hazel Yetiskin, Muhammet Ali Uygut, Shaikh Terkis Islam Pavel, Aykut Ozdarendeli, Gulshan Mammadova, and et al. 2021. "Plant-Produced Glycosylated and In Vivo Deglycosylated Receptor Binding Domain Proteins of SARS-CoV-2 Induce Potent Neutralizing Responses in Mice" Viruses 13, no. 8: 1595. https://doi.org/10.3390/v13081595
APA StyleMamedov, T., Yuksel, D., Ilgın, M., Gurbuzaslan, I., Gulec, B., Yetiskin, H., Uygut, M. A., Islam Pavel, S. T., Ozdarendeli, A., Mammadova, G., Say, D., & Hasanova, G. (2021). Plant-Produced Glycosylated and In Vivo Deglycosylated Receptor Binding Domain Proteins of SARS-CoV-2 Induce Potent Neutralizing Responses in Mice. Viruses, 13(8), 1595. https://doi.org/10.3390/v13081595





