Quorum Sensing Inhibition by Marine Bacteria
Abstract
1. Introduction
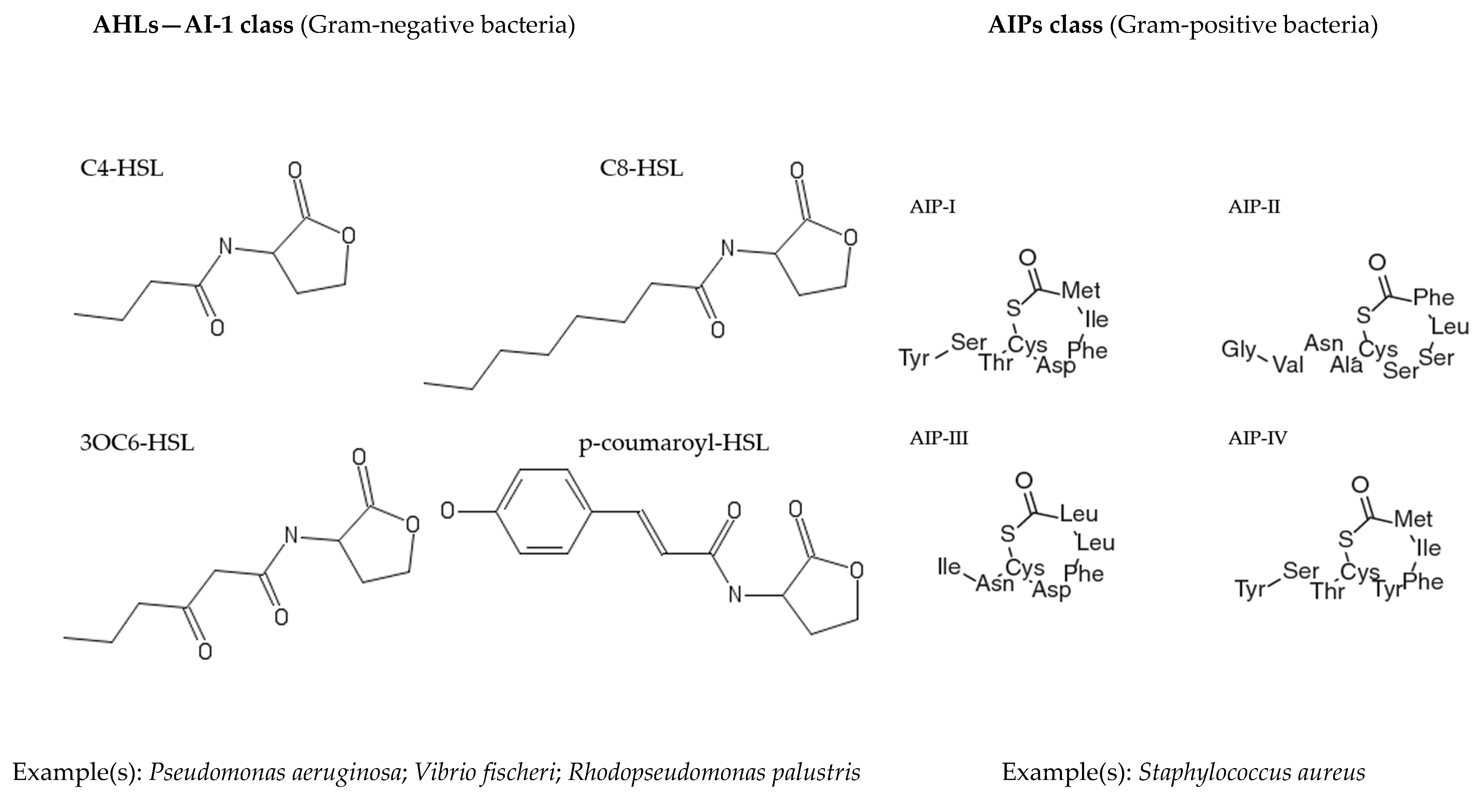
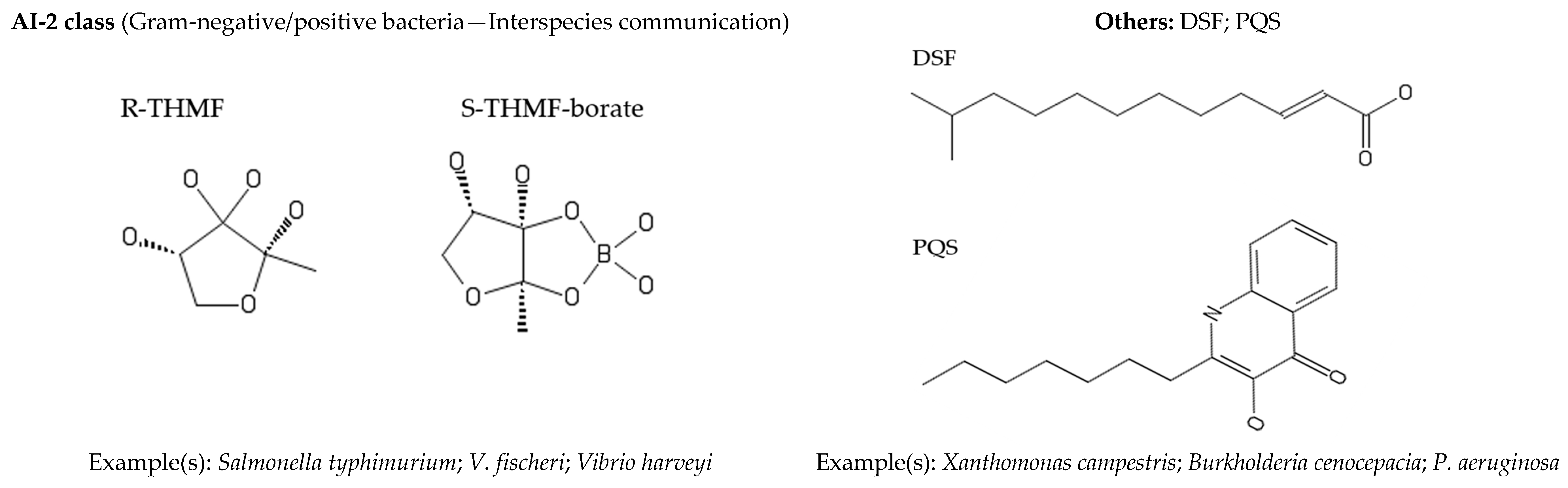
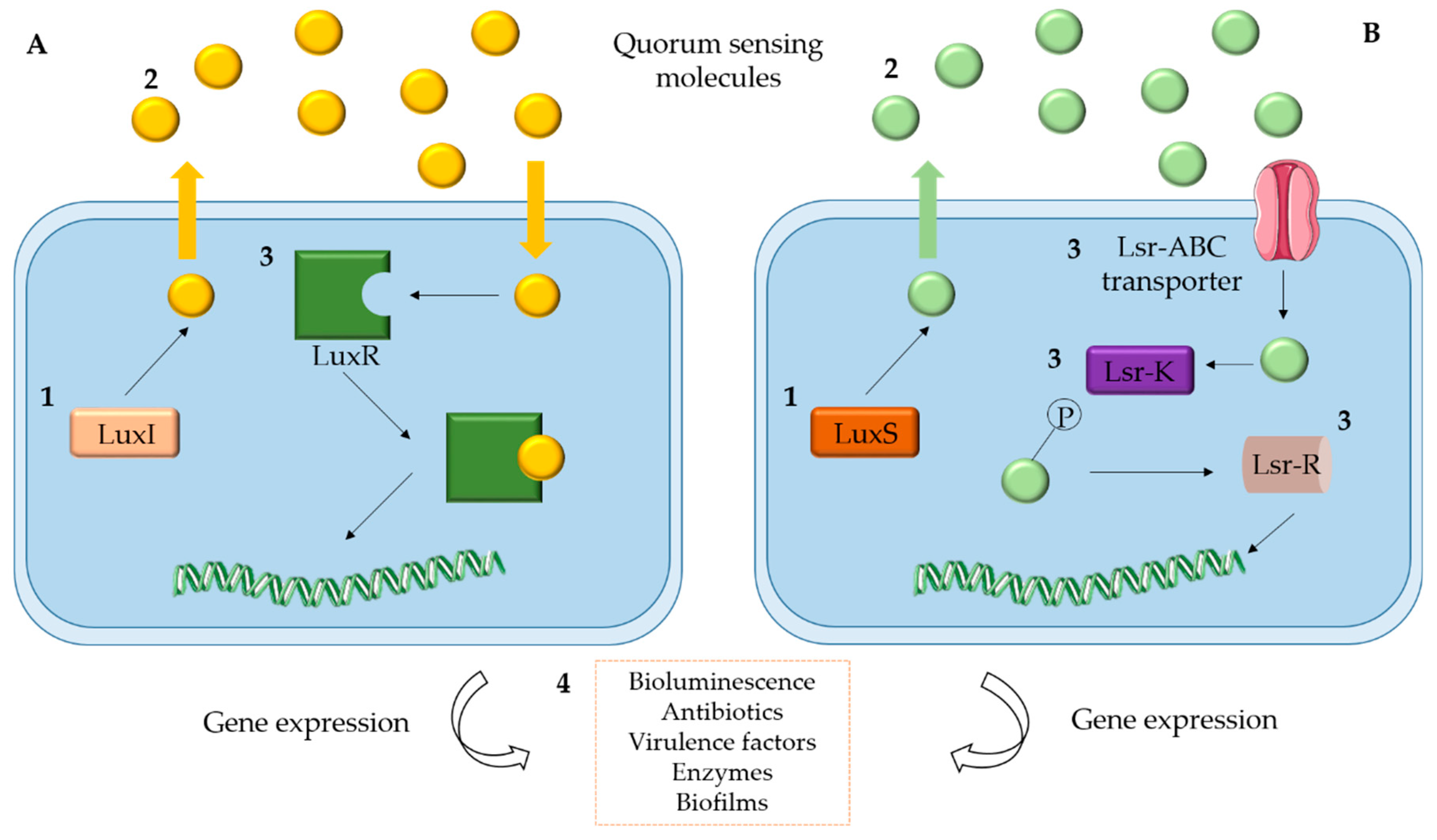
2. Quorum Sensing
Quorum Sensing in Marine Bacteria
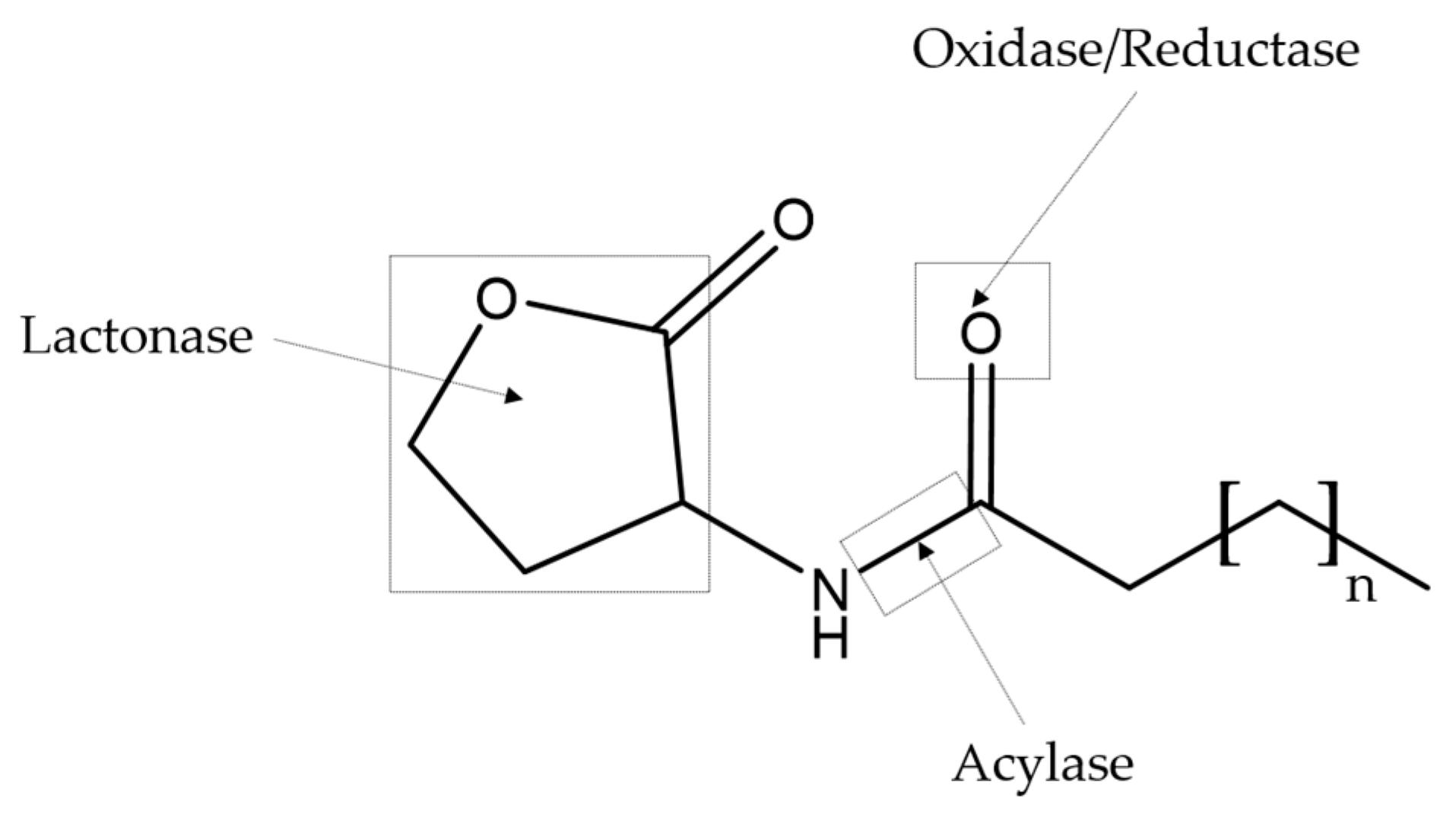
3. Quorum Sensing Inhibitors
QS Inhibitors from Marine Bacteria
4. Significance and Future Perspectives
Author Contributions
Funding
Conflicts of Interest
References
- Borges, A.; Saavedra, M.J.; Simões, M. Insights on antimicrobial resistance, biofilms and the use of phytochemicals as new antimicrobial agents. Curr. Med. Chem. 2015, 22, 2590–2614. [Google Scholar] [CrossRef] [PubMed]
- Cantas, L.; Shah, S.Q.A.; Cavaco, L.M.; Manaia, C.; Walsh, F.; Popowska, M.; Garelick, H.; Bürgmann, H.; Sørum, H. A brief multi-disciplinary review on antimicrobial resistance in medicine and its linkage to the global environmental microbiota. Front. Microbiol. 2013, 4, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Davies, J.; Davies, D. Origins and evolution of antibiotic resistance. Microbiol. Mol. Biol. Rev. 2010, 74, 417–433. [Google Scholar] [CrossRef] [PubMed]
- French, G.L. Clinical impact and relevance of antibiotic resistance. Adv. Drug Deliv. Rev. 2005, 57, 1514–1527. [Google Scholar] [CrossRef] [PubMed]
- Center for Disease Control and Prevention (CDC). Antibiotic Resistance Threats in the United States; Department of Health and Human Services: Washington, DC, USA, 2013; pp. 1–113.
- Stanton, T.B. A call for antibiotic alternatives research. Trends Microbiol. 2013, 21, 111–113. [Google Scholar] [CrossRef] [PubMed]
- Heras, B.; Scanlon, M.J.; Martin, J.L. Targeting virulence not viability in the search for future antibacterials. Br. J. Clin. Pharmacol. 2015, 79, 208–215. [Google Scholar] [CrossRef]
- LaSarre, B.; Federle, M.J. Exploiting quorum sensing to confuse bacterial pathogens. Microbiol. Mol. Biol. Rev. 2013, 77, 73–111. [Google Scholar] [CrossRef]
- Mühlen, S.; Dersch, P. Anti-virulence strategies to target bacterial infections. Curr. Top. Microbiol. Immunol. 2015, 147–183. [Google Scholar]
- Borges, A.; Abreu, A.; Dias, C.; Saavedra, M.; Borges, F.; Simões, M. New perspectives on the use of phytochemicals as an emergent strategy to control bacterial infections including biofilms. Molecules 2016, 21, 877. [Google Scholar] [CrossRef]
- Raffa, R.B.; Iannuzzo, J.R.; Levine, D.R.; Saeid, K.K.; Schwartz, R.C.; Sucic, N.T.; Terleckyj, O.D.; Young, J.M. Bacterial communication (“quorum sensing”) via ligands and receptors: A novel pharmacologic target for the design of antibiotic drugs. J. Pharmacol. Exp. Ther. 2005, 312, 417–423. [Google Scholar] [CrossRef]
- Kanagasabhapathy, M.; Yamazaki, G.; Ishida, A.; Sasaki, H.; Nagata, S. Presence of quorum-sensing inhibitor-like compounds from bacteria isolated from the brown alga Colpomenia sinuosa. Lett. Appl. Microbiol. 2009, 49, 573–579. [Google Scholar] [CrossRef] [PubMed]
- Bauer, W.D.; Robinson, J.B. Disruption of bacterial quorum sensing by other organisms. Curr. Opin. Biotechnol. 2002, 13, 234–237. [Google Scholar] [CrossRef]
- Hmelo, L.R. Quorum sensing in marine microbial environments. Ann. Rev. Mar. Sci. 2017, 9, 257–281. [Google Scholar] [CrossRef] [PubMed]
- Waters, C.M.; Bassler, B.L. Quorum sensing: Cell-to-cell communication in bacteria. Annu. Rev. Cell Dev. Biol. 2005, 21, 319–346. [Google Scholar] [CrossRef] [PubMed]
- Whitehead, N.A.; Barnard, A.M.L.; Slater, H.; Simpson, N.J.L.; Salmond, G.P.C. Quorum-sensing in Gram-negative bacteria. FEMS Microbiol. Rev. 2001, 25, 365–404. [Google Scholar] [CrossRef] [PubMed]
- Abisado, R.G.; Benomar, S.; Klaus, J.R.; Dandekar, A.A.; Chandler, J.R. Bacterial quorum sensing and microbial community interactions. mBio 2018, 9, e02331-17. [Google Scholar] [CrossRef] [PubMed]
- Vadakkan, K.; Choudhury, A.A.; Gunasekaran, R.; Hemapriya, J.; Vijayanand, S. Quorum sensing intervened bacterial signaling: Pursuit of its cognizance and repression. J. Genet. Eng. Biotechnol. 2018, 16, 239–252. [Google Scholar] [CrossRef] [PubMed]
- Papenfort, K.; Bassler, B.L. Quorum sensing signal–response systems in Gram-negative bacteria. Nat. Rev. Microbiol. 2016, 14, 576. [Google Scholar] [CrossRef]
- Jefferson, K.K. What drives bacteria to produce a biofilm? FEMS Microbiol. Lett. 2004, 236, 163–173. [Google Scholar] [CrossRef]
- Dickschat, J.S. Quorum sensing and bacterial biofilms. Nat. Prod. Rep. 2010, 27, 343–369. [Google Scholar] [CrossRef]
- Chen, J.; Wang, B.; Lu, Y.; Guo, Y.; Sun, J.; Wei, B.; Zhang, H.; Wang, H. Quorum sensing inhibitors from marine microorganisms and their synthetic derivatives. Mar. Drugs 2019, 17, 80. [Google Scholar] [CrossRef] [PubMed]
- Williams, P. Quorum sensing, communication and cross-kingdom signalling in the bacterial world. Microbiology 2007, 153, 3923–3938. [Google Scholar] [CrossRef] [PubMed]
- Uroz, S.; Dessaux, Y.; Oger, P. Quorum sensing and quorum quenching: The yin and yang of bacterial communication. Chembiochem 2009, 10, 205–216. [Google Scholar] [CrossRef] [PubMed]
- Whiteley, M.; Diggle, S.P.; Greenberg, E.P. Progress in and promise of bacterial quorum sensing research. Nature 2017, 551, 313. [Google Scholar] [CrossRef] [PubMed]
- Hense, B.A.; Schuster, M. Core principles of bacterial autoinducer systems. Microbiol. Mol. Biol. Rev. 2015, 79, 153–169. [Google Scholar] [CrossRef]
- Grandclément, C.; Tannières, M.; Moréra, S.; Dessaux, Y.; Faure, D. Quorum quenching: Role in nature and applied developments. FEMS Microbiol. Rev. 2015, 40, 86–116. [Google Scholar] [CrossRef]
- Galloway, W.R.; Hodgkinson, J.T.; Bowden, S.D.; Welch, M.; Spring, D.R. Quorum sensing in Gram-negative bacteria: Small-molecule modulation of AHL and AI-2 quorum sensing pathways. Chem. Rev. 2010, 111, 28–67. [Google Scholar] [CrossRef]
- Sturme, M.H.; Kleerebezem, M.; Nakayama, J.; Akkermans, A.D.; Vaughan, E.E.; De Vos, W.M. Cell to cell communication by autoinducing peptides in gram-positive bacteria. Antonie Van Leeuwenhoek 2002, 81, 233–243. [Google Scholar] [CrossRef]
- Thoendel, M.; Kavanaugh, J.S.; Flack, C.E.; Horswill, A.R. Peptide signaling in the staphylococci. Chem. Rev. 2010, 111, 117–151. [Google Scholar] [CrossRef]
- Lowery, C.A.; Dickerson, T.J.; Janda, K.D. Interspecies and interkingdom communication mediated by bacterial quorum sensing. Chem. Soc. Rev. 2008, 37, 1337–1346. [Google Scholar] [CrossRef]
- Hughes, D.T.; Sperandio, V. Inter-kingdom signalling: Communication between bacteria and their hosts. Nat. Rev. Micro. 2008, 6, 111–120. [Google Scholar] [CrossRef] [PubMed]
- Sperandio, V.; Torres, A.G.; Jarvis, B.; Nataro, J.P.; Kaper, J.B. Bacteria–host communication: The language of hormones. Proc. Natl. Acad. Sci. USA 2003, 100, 8951–8956. [Google Scholar] [CrossRef] [PubMed]
- Deng, Y.; Wu, J.e.; Eberl, L.; Zhang, L.-H. Structural and functional characterization of diffusible signal factor family quorum-sensing signals produced by members of the Burkholderia cepacia complex. Appl. Environ. Microbiol. 2010, 76, 4675–4683. [Google Scholar] [CrossRef] [PubMed]
- Ryan, R.P.; Dow, J.M. Communication with a growing family: Diffusible signal factor (DSF) signaling in bacteria. Trends Microbiol. 2011, 19, 145–152. [Google Scholar] [CrossRef] [PubMed]
- Lin, J.; Cheng, J.; Wang, Y.; Shen, X. The Pseudomonas quinolone signal (PQS): Not just for quorum sensing anymore. Front. Cell. Infect. Microbiol. 2018, 8, 230. [Google Scholar] [CrossRef]
- Bala, A.; Chhibber, S.; Harjai, K. Pseudomonas quinolone signalling system: A component of quorum sensing cascade is a crucial player in the acute urinary tract infection caused by Pseudomonas aeruginosa. Int. J. Med. Microbiol. 2014, 304, 1199–1208. [Google Scholar] [CrossRef]
- Zhou, J.; Lyu, Y.; Richlen, M.; Anderson, D.M.; Cai, Z. Quorum sensing is a language of chemical signals and plays an ecological role in algal-bacterial interactions. CRC Crit. Rev. Plant Sci. 2016, 35, 81–105. [Google Scholar] [CrossRef]
- Wang, Y.; Wang, Y.; Sun, L.; Grenier, D.; Yi, L. The LuxS/AI-2 system of Streptococcus suis. Appl. Microbiol. Biotechnol. 2018, 102, 7231–7238. [Google Scholar] [CrossRef]
- Miller, M.B.; Skorupski, K.; Lenz, D.H.; Taylor, R.K.; Bassler, B.L. Parallel quorum sensing systems converge to regulate virulence in Vibrio cholerae. Cell 2002, 110, 303–314. [Google Scholar] [CrossRef]
- Lee, J.; Zhang, L. The hierarchy quorum sensing network in Pseudomonas aeruginosa. Protein Cell 2015, 6, 26–41. [Google Scholar] [CrossRef]
- Dobretsov, S.; Teplitski, M.; Paul, V. Mini-review: Quorum sensing in the marine environment and its relationship to biofouling. Biofouling 2009, 25, 413–427. [Google Scholar] [CrossRef] [PubMed]
- Gram, L.; Grossart, H.-P.; Schlingloff, A.; Kiørboe, T. Possible quorum sensing in marine snow bacteria: Production of acylated homoserine lactones by Roseobacter strains isolated from marine snow. Appl. Environ. Microbiol. 2002, 68, 4111–4116. [Google Scholar] [CrossRef] [PubMed]
- Lynch, M.J.; Swift, S.; Kirke, D.F.; Keevil, C.W.; Dodd, C.E.; Williams, P. The regulation of biofilm development by quorum sensing in Aeromonas hydrophila. Environ. Microbiol. 2002, 4, 18–28. [Google Scholar] [CrossRef] [PubMed]
- Mangwani, N.; Kumari, S.; Das, S. Involvement of quorum sensing genes in biofilm development and degradation of polycyclic aromatic hydrocarbons by a marine bacterium Pseudomonas aeruginosa N6P6. Appl. Microbiol. Biotechnol. 2015, 99, 10283–10297. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Zeng, Y.; Yu, Z.; Zhang, J.; Feng, H.; Lin, X. In silico and experimental methods revealed highly diverse bacteria with quorum sensing and aromatics biodegradation systems—A potential broad application on bioremediation. Bioresour. Technol. 2013, 148, 311–316. [Google Scholar] [CrossRef] [PubMed]
- Mohamed, N.M.; Cicirelli, E.M.; Kan, J.; Chen, F.; Fuqua, C.; Hill, R.T. Diversity and quorum-sensing signal production of Proteobacteria associated with marine sponges. Environ. Microbiol. 2008, 10, 75–86. [Google Scholar] [CrossRef] [PubMed]
- Cottrell, M.T.; David, K.L. Contribution of major bacterial groups to bacterial biomass production (thymidine and leucine incorporation) in the Delaware estuary. Limnol. Oceanogr. 2003, 48, 168–178. [Google Scholar] [CrossRef]
- Zhang, L.-H.; Dong, Y.-H. Quorum sensing and signal interference: Diverse implications. Molec. Microbiol. 2004, 53, 1563–1571. [Google Scholar] [CrossRef] [PubMed]
- Defoirdt, T.; Sorgeloos, P.; Bossier, P. Alternatives to antibiotics for the control of bacterial disease in aquaculture. Curr. Opin. Microbiol. 2011, 14, 251–258. [Google Scholar] [CrossRef] [PubMed]
- Manefield, M.; de Nys, R.; Naresh, K.; Roger, R.; Givskov, M.; Peter, S.; Kjelleberg, S. Evidence that halogenated furanones from Delisea pulchra inhibit acylated homoserine lactone (AHL)-mediated gene expression by displacing the AHL signal from its receptor protein. Mycobiology 1999, 145, 283–291. [Google Scholar] [CrossRef]
- Dong, Y.-H.; Zhang, L.-H. Quorum sensing and quorum-quenching enzymes. J. Microbiol. 2005, 43, 101–109. [Google Scholar] [PubMed]
- Chun, C.K.; Ozer, E.A.; Welsh, M.J.; Zabner, J.; Greenberg, E. Inactivation of a Pseudomonas aeruginosa quorum-sensing signal by human airway epithelia. Proc. Natl. Acad. Sci. USA 2004, 101, 3587–3590. [Google Scholar] [CrossRef] [PubMed]
- Draganov, D.I.; Teiber, J.F.; Speelman, A.; Osawa, Y.; Sunahara, R.; La Du, B.N. Human paraoxonases (PON1, PON2, and PON3) are lactonases with overlapping and distinct substrate specificities. J. Lipid Res. 2005, 46, 1239–1247. [Google Scholar] [CrossRef] [PubMed]
- Xu, F.; Byun, T.; Dussen, H.-J.; Duke, K.R. Degradation of N-acylhomoserine lactones, the bacterial quorum-sensing molecules, by acylase. J. Biotechnol. 2003, 101, 89–96. [Google Scholar] [CrossRef]
- Weiland-Bräuer, N.; Fischer, M.A.; Pinnow, N.; Schmitz, R.A. Potential role of host-derived quorum quenching in modulating bacterial colonization in the moon jellyfish Aurelia aurita. Sci. Rep. 2019, 9, 34. [Google Scholar] [CrossRef] [PubMed]
- Stoltz, D.A.; Ozer, E.A.; Taft, P.J.; Barry, M.; Liu, L.; Kiss, P.J.; Moninger, T.O.; Parsek, M.R.; Zabner, J. Drosophila are protected from Pseudomonas aeruginosa lethality by transgenic expression of paraoxonase-1. J. Clin. Investig. 2008, 118, 3123–3131. [Google Scholar] [CrossRef] [PubMed]
- Pietschke, C.; Treitz, C.; Forêt, S.; Schultze, A.; Künzel, S.; Tholey, A.; Bosch, T.C.; Fraune, S. Host modification of a bacterial quorum-sensing signal induces a phenotypic switch in bacterial symbionts. Proc. Natl. Acad. Sci. USA 2017, 114, E8488–E8497. [Google Scholar] [CrossRef] [PubMed]
- Czajkowski, R.; Jafra, S. Quenching of acyl-homoserine lactone-dependent quorum sensing by enzymatic disruption of signal molecules. Acta Biochim. Pol. 2009, 56, 1–16. [Google Scholar] [CrossRef]
- Mayer, C.; Muras, A.; Romero, M.; López, M.; Tomas, M.; Otero, A. Multiple quorum quenching enzymes are active in the nosocomial pathogen Acinetobacter baumannii ATCC17978. Front. Cell. Infect. Microbiol. 2018, 8, 310. [Google Scholar] [CrossRef]
- Bzdrenga, J.; Daudé, D.; Rémy, B.; Jacquet, P.; Plener, L.; Elias, M.; Chabrière, E. Biotechnological applications of quorum quenching enzymes. Chem. Biol. Interact. 2017, 267, 104–115. [Google Scholar] [CrossRef]
- Chen, F.; Gao, Y.; Chen, X.; Yu, Z.; Li, X. Quorum quenching enzymes and their application in degrading signal molecules to block quorum sensing-dependent infection. Int. J. Mol. Sci. 2013, 14, 17477–17500. [Google Scholar] [CrossRef] [PubMed]
- Tang, K.; Su, Y.; Brackman, G.; Cui, F.; Zhang, Y.; Shi, X.; Coenye, T.; Zhang, X.-H. MomL, a novel marine-derived N-acyl homoserine lactonase from Muricauda olearia. Appl. Environ. Microbiol. 2015, 81, 774–782. [Google Scholar] [CrossRef] [PubMed]
- Roy, V.; Adams, B.L.; Bentley, W.E. Developing next generation antimicrobials by intercepting AI-2 mediated quorum sensing. Enzyme Microb. Technol. 2011, 49, 113–123. [Google Scholar] [CrossRef] [PubMed]
- Kalia, V.C.; Purohit, H.J. Quenching the quorum sensing system: Potential antibacterial drug targets. Crit. Rev. Microbiol. 2011, 37, 121–140. [Google Scholar] [CrossRef] [PubMed]
- Saurav, K.; Costantino, V.; Venturi, V.; Steindler, L. Quorum sensing inhibitors from the sea discovered using bacterial N-acyl-homoserine lactone-based biosensors. Mar. Drugs 2017, 15, 53. [Google Scholar] [CrossRef]
- Dobretsov, S.; Abed, R.M.M.; Teplitski, M. Mini-review: Inhibition of biofouling by marine microorganisms. Biofouling 2013, 29, 423–441. [Google Scholar] [CrossRef]
- Zhao, J.; Li, X.; Hou, X.; Quan, C.; Chen, M. Widespread existence of quorum sensing inhibitors in marine bacteria: Potential drugs to combat pathogens with novel strategies. Mar. Drugs 2019, 17, 275. [Google Scholar] [CrossRef]
- Newman, D.J.; Cragg, G.M. Natural products as sources of new drugs over the 30 years from 1981 to 2010. J. Nat. Prod. 2012, 75, 311–335. [Google Scholar] [CrossRef]
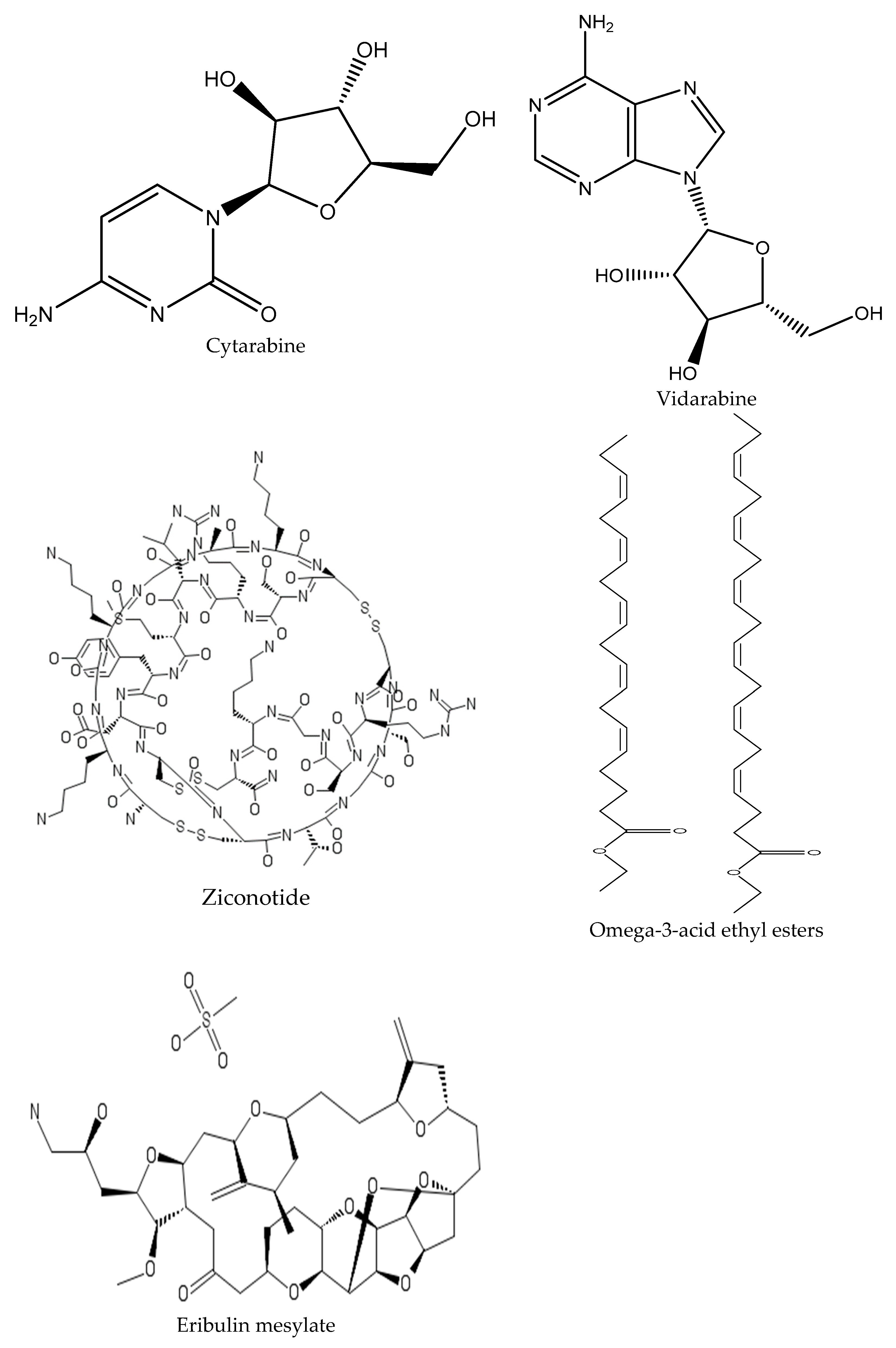
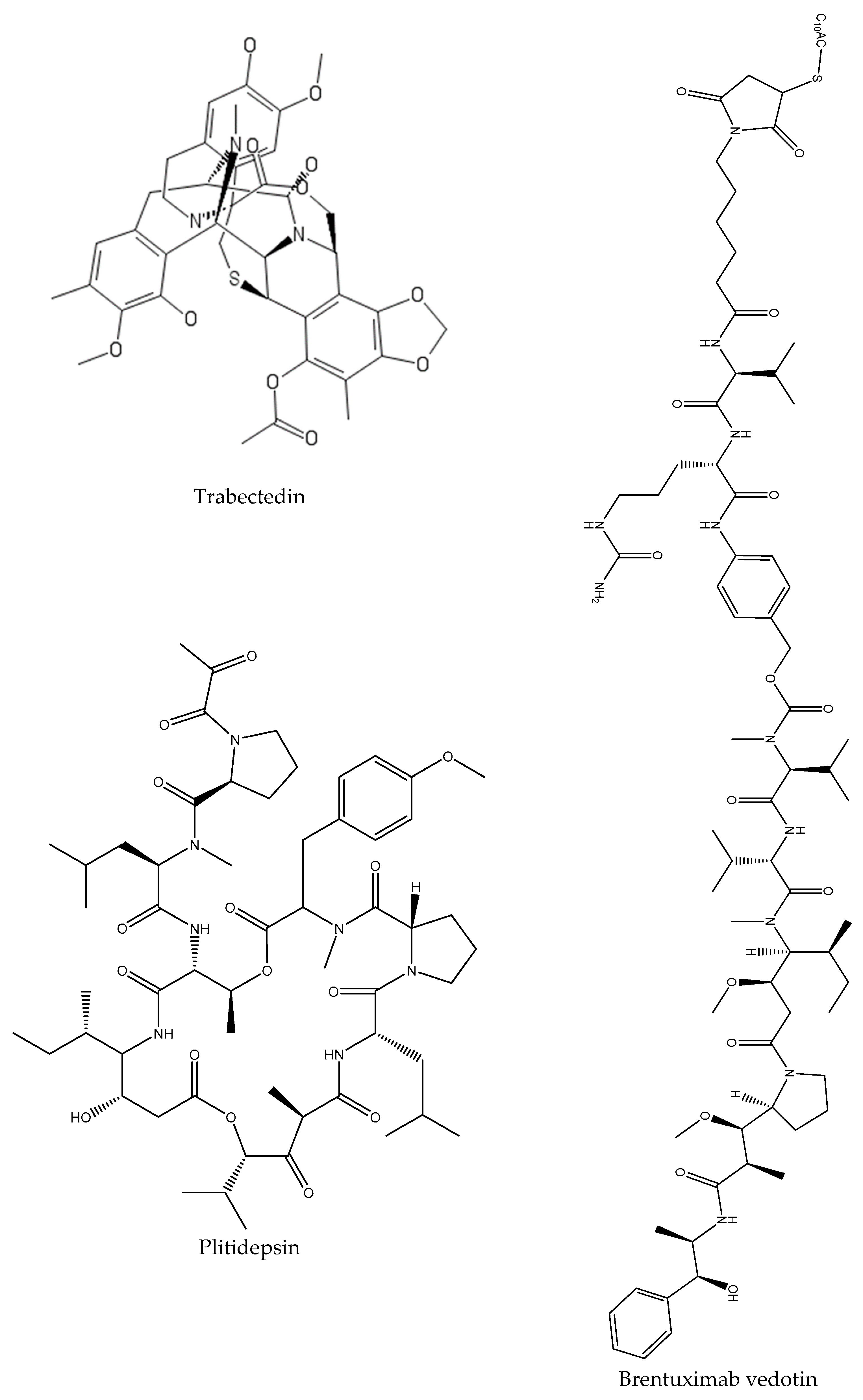
- Malve, H. Exploring the ocean for new drug developments: Marine pharmacology. J. Pharm. Bioallied. Sci. 2016, 8, 83–91. [Google Scholar] [CrossRef]
- Nair, A.; Poornachand, A.; Kodisharapu, P. Ziconotide: Indications, adverse effects, and limitations in managing refractory chronic pain. Indian J. Palliat. Care 2018, 24, 118–119. [Google Scholar]
- El-Subbagh, H.I.; Al-Badr, A.A. Chapter 2—Cytarabine. In Profiles of Drug Substances, Excipients and Related Methodology; Brittain, H.G., Ed.; Elsevier: San Diego, CA, USA; Waltham, MA, USA; London, UK, 2009; Volume 34, pp. 37–113. [Google Scholar]
- Lee, J.-Y.; Orlikova, B.; Diederich, M. Signal transducers and activators of transcription (STAT) regulatory networks in marine organisms: From physiological observations towards marine drug discovery. Mar. Drugs 2015, 13, 4967–4984. [Google Scholar] [CrossRef] [PubMed]
- Abadines, I.B.; Le, K.; Newman, D.J.; Glaser, K.B.; Mayer, A.M. The Marine Pharmacology and Pharmaceuticals Pipeline in 2018. FASEB J. 2019, 33, 504. [Google Scholar]
- Katz, J.; Janik, J.E.; Younes, A. Brentuximab Vedotin (SGN-35). Clin. Cancer Res. 2011, 17, 6428. [Google Scholar] [CrossRef] [PubMed]
- Mayer, A.M.; Glaser, K.B.; Cuevas, C.; Jacobs, R.S.; Kem, W.; Little, R.D.; McIntosh, J.M.; Newman, D.J.; Potts, B.C.; Shuster, D.E. The odyssey of marine pharmaceuticals: A current pipeline perspective. Trends Pharmacol. Sci. 2010, 31, 255–265. [Google Scholar] [CrossRef] [PubMed]
- Skindersoe, M.E.; Ettinger-Epstein, P.; Rasmussen, T.B.; Bjarnsholt, T.; de Nys, R.; Givskov, M. Quorum sensing antagonism from marine organisms. Mar. Biotechnol. 2008, 10, 56–63. [Google Scholar] [CrossRef] [PubMed]
- Debbab, A.; Aly, A.H.; Lin, W.H.; Proksch, P. Bioactive compounds from marine bacteria and fungi. Microb. Biotechnol. 2010, 3, 544–563. [Google Scholar] [CrossRef]
- Romano, G.; Costantini, M.; Sansone, C.; Lauritano, C.; Ruocco, N.; Ianora, A. Marine microorganisms as a promising and sustainable source of bioactive molecules. Mar. Environ. Res. 2017, 128, 58–69. [Google Scholar] [CrossRef] [PubMed]
- Monciardini, P.; Iorio, M.; Maffioli, S.; Sosio, M.; Donadio, S. Discovering new bioactive molecules from microbial sources. Microb. Biotechnol. 2014, 7, 209–220. [Google Scholar] [CrossRef]
- Bister, B.; Bischoff, D.; Ströbele, M.; Riedlinger, J.; Reicke, A.; Wolter, F.; Bull, A.T.; Zähner, H.; Fiedler, H.P.; Süssmuth, R.D. Abyssomicin C—A polycyclic antibiotic from a marine Verrucosispora strain as an Inhibitor of the p-aminobenzoic acid/tetrahydrofolate biosynthesis pathway. Angew. Chem. Int. Ed. Engl. 2004, 43, 2574–2576. [Google Scholar] [CrossRef]
- Oh, D.-C.; Williams, P.G.; Kauffman, C.A.; Jensen, P.R.; Fenical, W. Cyanosporasides A and B, chloro- and cyano-cyclopenta[a]indene glycosides from the marine actinomycete “Salinispora pacifica”. Org. Lett. 2006, 8, 1021–1024. [Google Scholar] [CrossRef]
- Gulder, T.A.; Moore, B.S. Salinosporamide natural products: Potent 20 S proteasome inhibitors as promising cancer chemotherapeutics. Angew. Chem. Int. Ed. Engl. 2010, 49, 9346–9367. [Google Scholar] [CrossRef] [PubMed]
- Kijjoa, A.; Sawangwong, P. Drugs and cosmetics from the sea. Mar. Drugs 2004, 2, 73–82. [Google Scholar] [CrossRef]
- Romero, M.; Martin-Cuadrado, A.-B.; Otero, A. Determination of whether quorum quenching is a common activity in marine bacteria by analysis of cultivable bacteria and metagenomic sequences. Appl. Environ. Microbiol. 2012, 78, 6345–6348. [Google Scholar] [CrossRef] [PubMed]
- Romero, M.; Martin-Cuadrado, A.-B.; Roca-Rivada, A.; Cabello, A.M.; Otero, A. Quorum quenching in cultivable bacteria from dense marine coastal microbial communities. FEMS Microbiol. Ecol. 2011, 75, 205–217. [Google Scholar] [CrossRef] [PubMed]
- Chang, H.; Zhou, J.; Zhu, X.; Yu, S.; Chen, L.; Jin, H.; Cai, Z. Strain identification and quorum sensing inhibition characterization of marine-derived Rhizobium sp. NAO1. R. Soc. Open Sci. 2017, 4, 170025. [Google Scholar] [CrossRef] [PubMed]
- Teasdale, M.E.; Liu, J.; Wallace, J.; Akhlaghi, F.; Rowley, D.C. Secondary metabolites produced by the marine bacterium Halobacillus salinus that inhibit quorum sensing-controlled phenotypes in gram-negative bacteria. Appl. Environ. Microbiol. 2009, 75, 567–572. [Google Scholar] [CrossRef] [PubMed]
- Teasdale, M.E.; Donovan, K.A.; Forschner-Dancause, S.R.; Rowley, D.C. Gram-positive marine bacteria as a potential resource for the discovery of quorum sensing inhibitors. Mar. Biotechnol. 2011, 13, 722–732. [Google Scholar] [CrossRef]
- Nithya, C.; Begum, M.F.; Pandian, S.K. Marine bacterial isolates inhibit biofilm formation and disrupt mature biofilms of Pseudomonas aeruginosa PAO1. Appl. Microbiol. Biotechnol. 2010, 88, 341–358. [Google Scholar] [CrossRef] [PubMed]
- Cho, K.W.; Lee, H.-S.; Rho, J.-R.; Kim, T.S.; Mo, S.J.; Shin, J. New lactone-containing metabolites from a marine-derived bacterium of the genus Streptomyces. J. Nat. Prod. 2001, 64, 664–667. [Google Scholar] [CrossRef]
- You, J.; Xue, X.; Cao, L.; Lu, X.; Wang, J.; Zhang, L.; Zhou, S. Inhibition of Vibrio biofilm formation by a marine actinomycete strain A66. Appl. Microbiol. Biotechnol. 2007, 76, 1137–1144. [Google Scholar] [CrossRef]
- Bakkiyaraj, D.; Karutha Pandian, S.T. In vitro and in vivo antibiofilm activity of a coral associated actinomycete against drug resistant Staphylococcus aureus biofilms. Biofouling 2010, 26, 711–717. [Google Scholar] [CrossRef] [PubMed]
- Naik, D.; Wahidullah, S.; Meena, R. Attenuation of Pseudomonas aeruginosa virulence by marine invertebrate–derived Streptomyces sp. Lett. Appl. Microbiol. 2013, 56, 197–207. [Google Scholar] [CrossRef] [PubMed]
- Miao, L.; Xu, J.; Yao, Z.; Jiang, Y.; Zhou, H.; Jiang, W.; Dong, K. The anti-quorum sensing activity and bioactive substance of a marine derived Streptomyces. Biotechnol. Biotechnol. Equip. 2017, 31, 1007–1015. [Google Scholar] [CrossRef]
- Gutiérrez-Barranquero, J.A.; Reen, F.J.; Parages, M.L.; McCarthy, R.; Dobson, A.D.; O’Gara, F. Disruption of N-acyl-homoserine lactone-specific signalling and virulence in clinical pathogens by marine sponge bacteria. Microb. Biotechnol. 2017. [Google Scholar] [CrossRef] [PubMed]
- Ding, B.-T.; Lu, Y.-Q.; Dong, K.-M.; Wang, X.-Z.; Zhou, X.-J.; Jin, C.-L.; Miao, L. Screen for the quorum sensing inhibitory activity from marine bacterial isolates. Microbiol. China 2012, 39, 0002–0008. [Google Scholar]
- Li, M.; Huiru, Z.; Biting, D.; Yun, J.; Wei, J.; Kunming, D. Study on the anti-quorum sensing activity of a marine bacterium Staphylococcus saprophyticus 108. BioTechnology 2013, 7, 480–487. [Google Scholar]
- Saurav, K.; Bar-Shalom, R.; Haber, M.; Burgsdorf, I.; Oliviero, G.; Costantino, V.; Morgenstern, D.; Steindler, L. In search of alternative antibiotic drugs: Quorum-quenching activity in sponges and their bacterial isolates. Front. Microbiol. 2016, 7, 416. [Google Scholar] [CrossRef]
- Costantino, V.; Della Sala, G.; Saurav, K.; Teta, R.; Bar-Shalom, R.; Mangoni, A.; Steindler, L. Plakofuranolactone as a quorum quenching agent from the Indonesian sponge Plakortis cf. lita. Mar. Drugs 2017, 15, 59. [Google Scholar] [CrossRef]
- Gowrishankar, S.; Pandian, S.; Balasubramaniam, B.; Balamurugan, K. Quorum quelling efficacy of marine cyclic dipeptide -cyclo(L-leucyl-L-prolyl) against the uropathogen Serratia marcescens. Food Chem. Toxicol. 2019, 123, 326–336. [Google Scholar] [CrossRef]
- Clark, B.R.; Engene, N.; Teasdale, M.E.; Rowley, D.C.; Matainaho, T.; Valeriote, F.A.; Gerwick, W.H. Natural products chemistry and taxonomy of the marine cyanobacterium Blennothrix cantharidosmum. J. Nat. Prod. 2008, 71, 1530–1537. [Google Scholar] [CrossRef]
- Dobretsov, S.; Teplitski, M.; Alagely, A.; Gunasekera, S.P.; Paul, V.J. Malyngolide from the cyanobacterium Lyngbya majuscula interferes with quorum sensing circuitry. Environ. Microbiol. Rep. 2010, 2, 739–744. [Google Scholar] [CrossRef] [PubMed]
- Romero, M.; Diggle, S.P.; Heeb, S.; Camara, M.; Otero, A. Quorum quenching activity in Anabaena sp. PCC 7120: Identification of AiiC, a novel AHL-acylase. FEMS Microbiol. Lett. 2008, 280, 73–80. [Google Scholar] [CrossRef] [PubMed]
- Gowrishankar, S.; Sivaranjani, M.; Kamaladevi, A.; Ravi, A.V.; Balamurugan, K.; Karutha Pandian, S. Cyclic dipeptide cyclo(l-leucyl-l-prolyl) from marine Bacillus amyloliquefaciens mitigates biofilm formation and virulence in Listeria monocytogenes. Pathog. Dis. 2016, 74, ftw017. [Google Scholar] [CrossRef] [PubMed]
- Gowrishankar, S.; Poornima, B.; Pandian, S.K. Inhibitory efficacy of cyclo(l-leucyl-l-prolyl) from mangrove rhizosphere bacterium–Bacillus amyloliquefaciens (MMS-50) toward cariogenic properties of Streptococcus mutans. Res. Microbiol. 2014, 165, 278–289. [Google Scholar] [CrossRef] [PubMed]
- Kalia, V.C. Quorum sensing inhibitors: An overview. Biotechnol. Adv. 2013, 31, 224–245. [Google Scholar] [CrossRef] [PubMed]
- Pan, J.; Ren, D. Quorum sensing inhibitors: A patent overview. Expert Opin. Ther. Pat. 2009, 19, 1581–1601. [Google Scholar] [CrossRef] [PubMed]
- Koul, S.; Prakash, J.; Mishra, A.; Kalia, V.C. Potential emergence of multi-quorum sensing inhibitor resistant (MQSIR) bacteria. Indian J. Microbiol. 2016, 56, 1–18. [Google Scholar] [CrossRef]
- Kalia, V.C.; Wood, T.K.; Kumar, P. Evolution of resistance to quorum-sensing inhibitors. Microb. Ecol. 2014, 68, 13–23. [Google Scholar] [CrossRef]
- Defoirdt, T.; Boon, N.; Bossier, P. Can bacteria evolve resistance to quorum sensing disruption? PLoS Pathog. 2010, 6, e1000989. [Google Scholar] [CrossRef]
| Marine Source | Bacteria and Metabolite(s) | Indicator Organism(s) | QS Inhibitory Activity and QS Related Phenotypes | Reference(s) |
|---|---|---|---|---|
| Brown algae Colpomenia sinuosa | Bacillaceae, Pseudomonadaceae, Pseudoalteromonadaceae, Vibrionaceae families | S. rubidaea | Inhibition of red pigment that can be associated with the production of QSIs or QS like compounds | [12] |
| Diatom-dominated biofilm loosely, brown seaweed Fucus vesiculosus and the sediment of an inland fish culture tank | Alpha- and Gammaproteobacteria classes; Actinobacteria, Firmicutes and Bacteroidetes phylum | C. violaceum (CV026, VIR07) and E. coli (JM109 pSB1075 | Inhibition of violacein production and light emission, which can be due to enzymatic inactivation | [86] |
| Surface water samples collected from the North Atlantic Ocean | Rhizobium sp; AHL analogues (C4-AHL) | C. violaceum ATCC12472 | Inhibition of violacein production. Interference with P. aeruginosa biofilm formation, downregulation of virulence factors production and enhanced biofilm susceptibility to antibiotics. The observed outcomes are attributed to AIs competition | [87] |
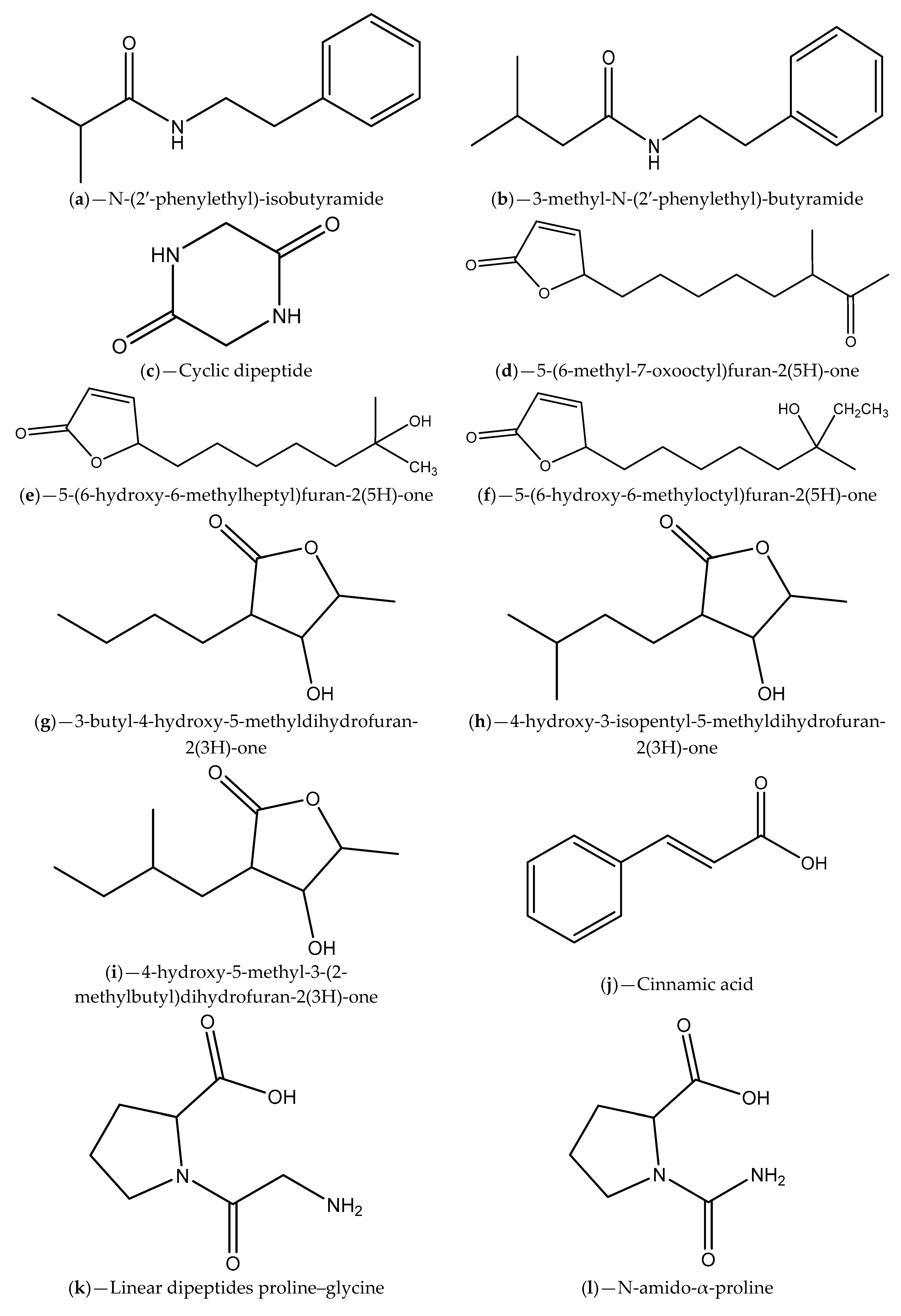
| Sea grass sample collected from a Rhode Island estuary | H. salinus; N-(2′-phenylethyl)-isobutyramide and 3-methyl-N-(2′-phenylethyl)-butyramide | V. harveyi BB120; V. harveyi BB120, C. violaceum CV026 and C. violaceum ATCC 12472 | Inhibition of violacein production and luminescence emission, which can be due to receptor binding competition | [88] |
| Marine samples (algae, invertebrates and surface sediments) collected from Rhode Island coastline | Bacillus, Halobacillus, Streptomyces and Micromonospora genera; phenethylamides and a cyclic dipeptide | V. harveyi; C. violaceum | Inhibition of bioluminescence emission and violacein production that can be related to AHL degradation | [89] |
| Sediment samples from Palk Bay region | B. pumilus, B. indicus, B. arsenicus, H. trueperi, F. balearica, and M. hydrocarbonoclasticus | C. violaceum ATCC 12472/CV026 | Inhibition of violacein production; Dispersion of P. aeruginosa biofilms | [90] |
| Shallow-water sediments from Tongyoung Bay | Streptomyces genus; Butenolides and 3-hydroxy-γ-butyrolactones | - | Competition with AHL signaling molecules | [91] |
| Marine sediments from South China | Actinomycetes (e.g., S. albus) | A. tumefaciensWCF47 (pCF372/pCF218) | Inhibition of biofilm formation and biofilm dispersion in V. vulnificus V0105, V. anguillarum AN0306, and V. harveyi H; Decrease of AHLs activity | [92] |
| Coral associated actinomycete | Actinomycetes (e.g., S. akiyoshiensis) | C. violaceum ATCC 12472 | Inhibition of violacein production; Inhibition of S. aureus (reference strains and clinical isolates) biofilm formation | [93] |
| Marine sponge-derived actinomycetes | Streptomyces; Cinnamic acid, linear dipeptides proline–glycine and N-amido-α-proline | C. violaceum CV12472 | Inhibition of violacein production; Inference with QS-regulated virulence factors in P. aeruginosa ATCC 27853 | [94] |
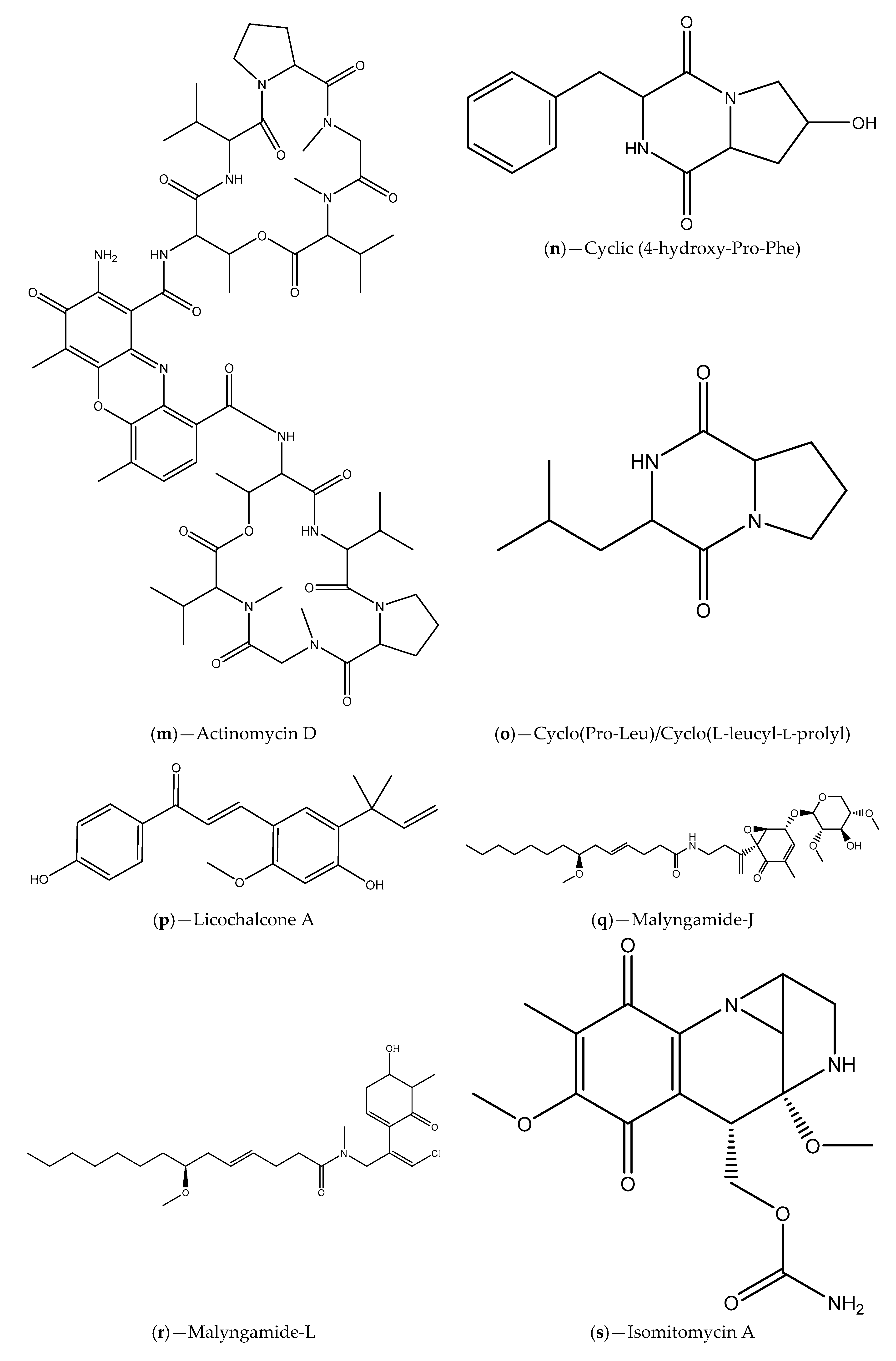
| Seawater samples from Lianyungang region | Actinomycetes (e.g., S. parvulus); Actinomycin D and cyclic (4-hydroxy-Pro-Phe) | C. violaceum ATCC 12472; S. proteamaculans | Inhibition of violacein production; Inhibition of P. aeruginosa PAO1, S. aureus 95005, M. luteus 95006 and Ruegeria sp. 01008 biofilm formation; Inhibition of prodigiosin in S. proteamaculans | [95] |
| Marine sponges | Gammaproteobacteria and Alphaproteobacteria classes; Firmicutes phylum | S.marcescens SP15, C. violaceum DSM 30191, A. tumefaciens NTL4 | Decrease of AHLs activity (hort-, medium- and long-chain AHLs); Antibiofilm activity against P. aeruginosa PA14 and B. subtilis CH8a; Inhibition of swarming/swimming motility and pyocyanin production | [96] |
| Sponge tissues | S. saprophyticus; cyclo(Pro-Leu) | C. violaceum 12472 | Inhibition of violacein production | [97,98] |
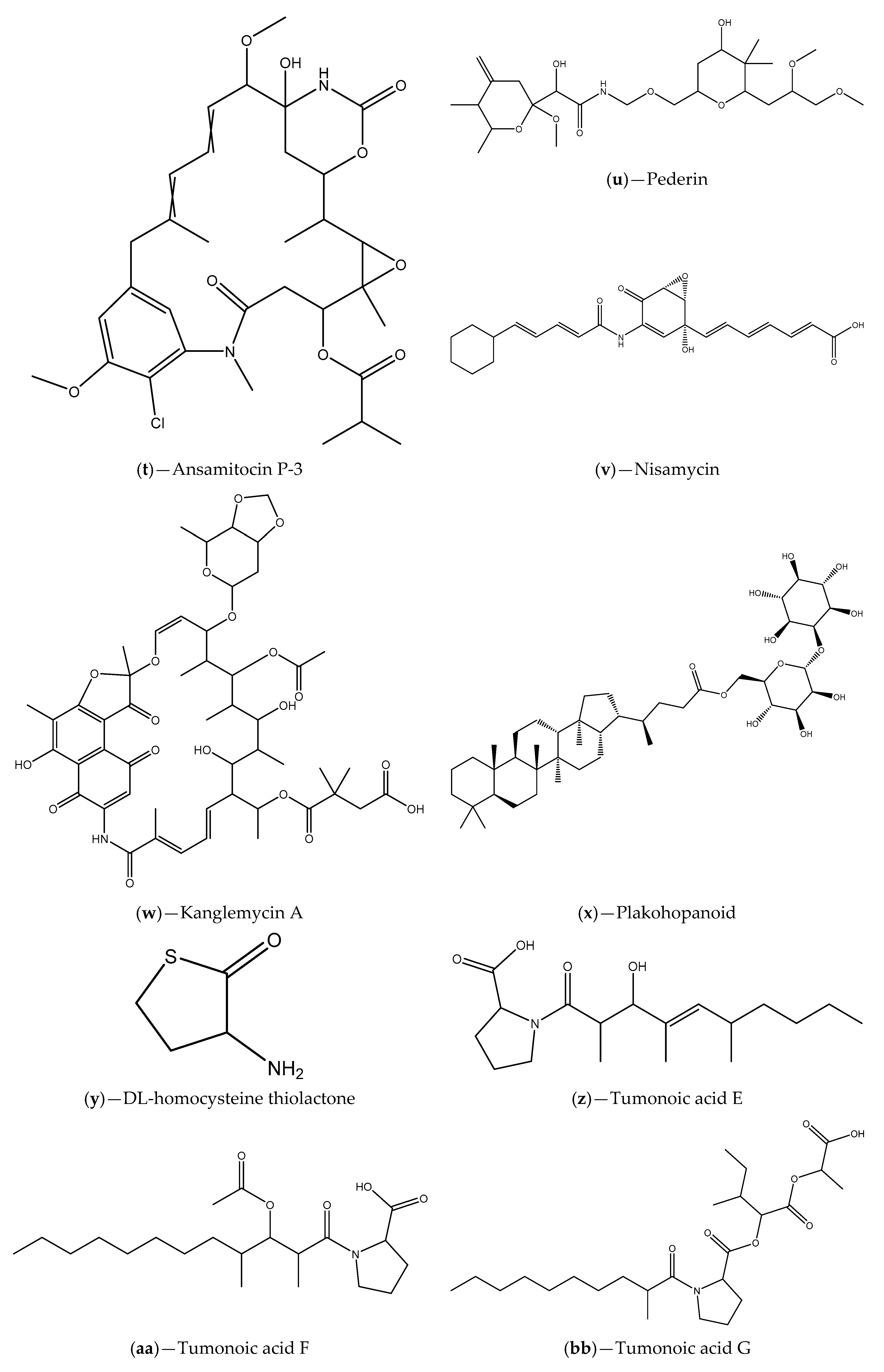
| Sponge tissues | Proteobacteria, Firmicutes, Actinobateria and Bacteroidete phylum; Licochalcone A, malyngamide-J, malyngamide-J, isomitomycin A, ansamitocin P-3, pederin, nisamycin and kanglemycin A | V. fischeri based selector system and C. violaceum CV026; A. tumefaciens NT1 | Inhibition of bioluminescence/violacein production; Interference with P. aeruginosa QS-regulated virulence factors (pyocyanin and protease production); Inhibition of P. aeruginosa biofilm formation | [99] |
| Marine sponge Plakortis cf. lita | Bacterial symbionts; plakohopanoid | E. coli pSB1075 | Inhibition of bioluminescence production; Interference with P. aeruginosa QS-regulated virulence factor (total protease activity) | [100] |
| Mangrove rhizosphere | B. amyloliquefaciens; cyclo(L-leucyl-l-prolyl) | S. marcescens | Inhibition of prodigiosin in S. marcescens; Interference with S. marcescens QS-regulated virulence factors | [101] |
| Coral Pocillopora damicornis | S. hominis; DL-homocysteine thiolactone | C. violaceum ATCC 12472 | Inhibition of violacein production; Anti-biofilm activity against P. aeruginosa PAO1; Downregulation of P. aeruginosa QS-related genes; QSI can be related with AHL competition for the receptor | [66] |
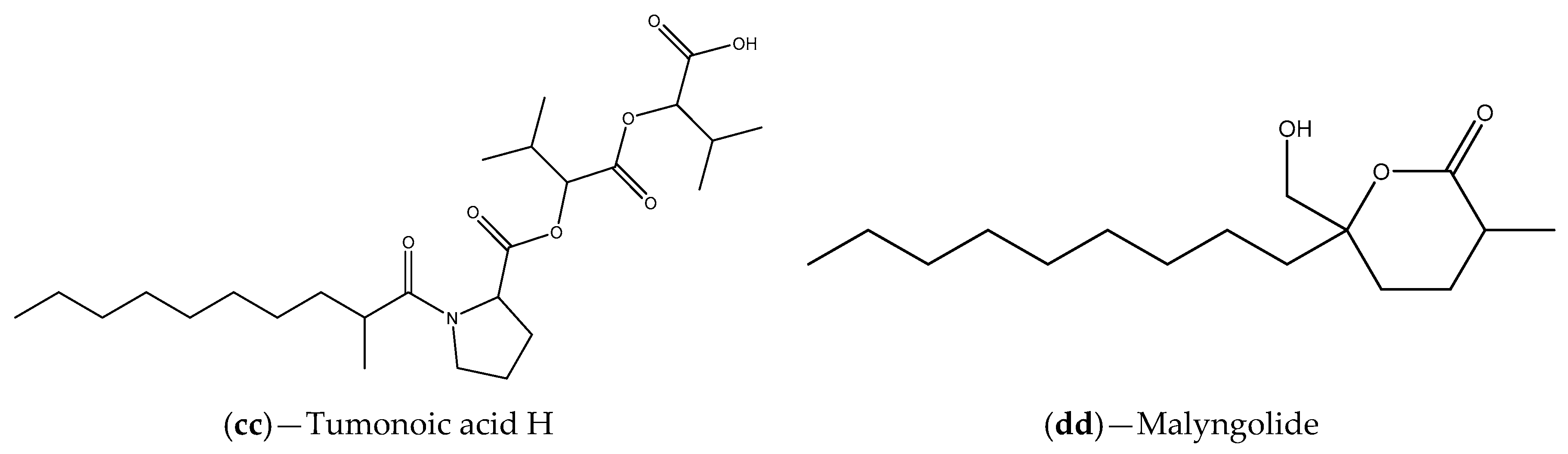
| Extracts Cyanobacterium Blennothrix cantharidosmum | Tumonoic acids (E, F, G and H) | V. harveyi | Inhibition of bioluminescence production | [102] |
| Florida waters samples | Marine cyanobacteria (e.g., S. hydnoides and L. majuscule); malyngolide | C. violaceum CV017 | Inhibition of violacein production that can be associated with competition for the binding site | [103] |
| Filamentous nitrogen-fixing cyanobacterium | Anabaena (Nostoc) sp. PCC 7120 | - | Degradation of the AI AHL by acylase enzyme | [104] |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Borges, A.; Simões, M. Quorum Sensing Inhibition by Marine Bacteria. Mar. Drugs 2019, 17, 427. https://doi.org/10.3390/md17070427
Borges A, Simões M. Quorum Sensing Inhibition by Marine Bacteria. Marine Drugs. 2019; 17(7):427. https://doi.org/10.3390/md17070427
Chicago/Turabian StyleBorges, Anabela, and Manuel Simões. 2019. "Quorum Sensing Inhibition by Marine Bacteria" Marine Drugs 17, no. 7: 427. https://doi.org/10.3390/md17070427
APA StyleBorges, A., & Simões, M. (2019). Quorum Sensing Inhibition by Marine Bacteria. Marine Drugs, 17(7), 427. https://doi.org/10.3390/md17070427






