Effect of Triclosan and Silver Nanoparticles on DNA Damage Investigated with DNA-Based Biosensor
Abstract
:1. Introduction
2. Materials and Methods
2.1. Materials
2.2. Apparatus
2.3. Preparation of the Biosensors
2.4. Methods
2.4.1. Cyclic Voltammetry (CV)
2.4.2. Square-Wave Voltammetry (SWV)
3. Results and Discussion
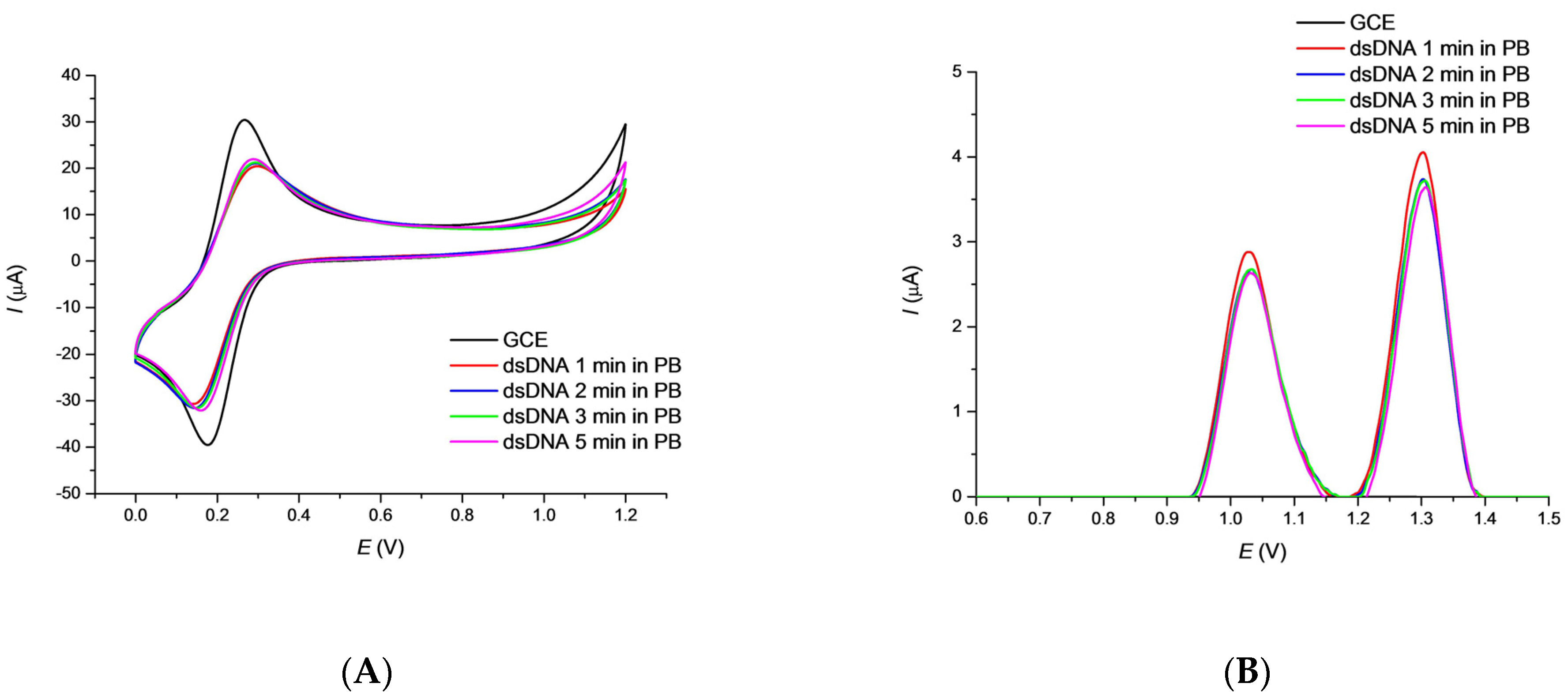
3.1. Characterization of the DNA/GCE Biosensor Stability
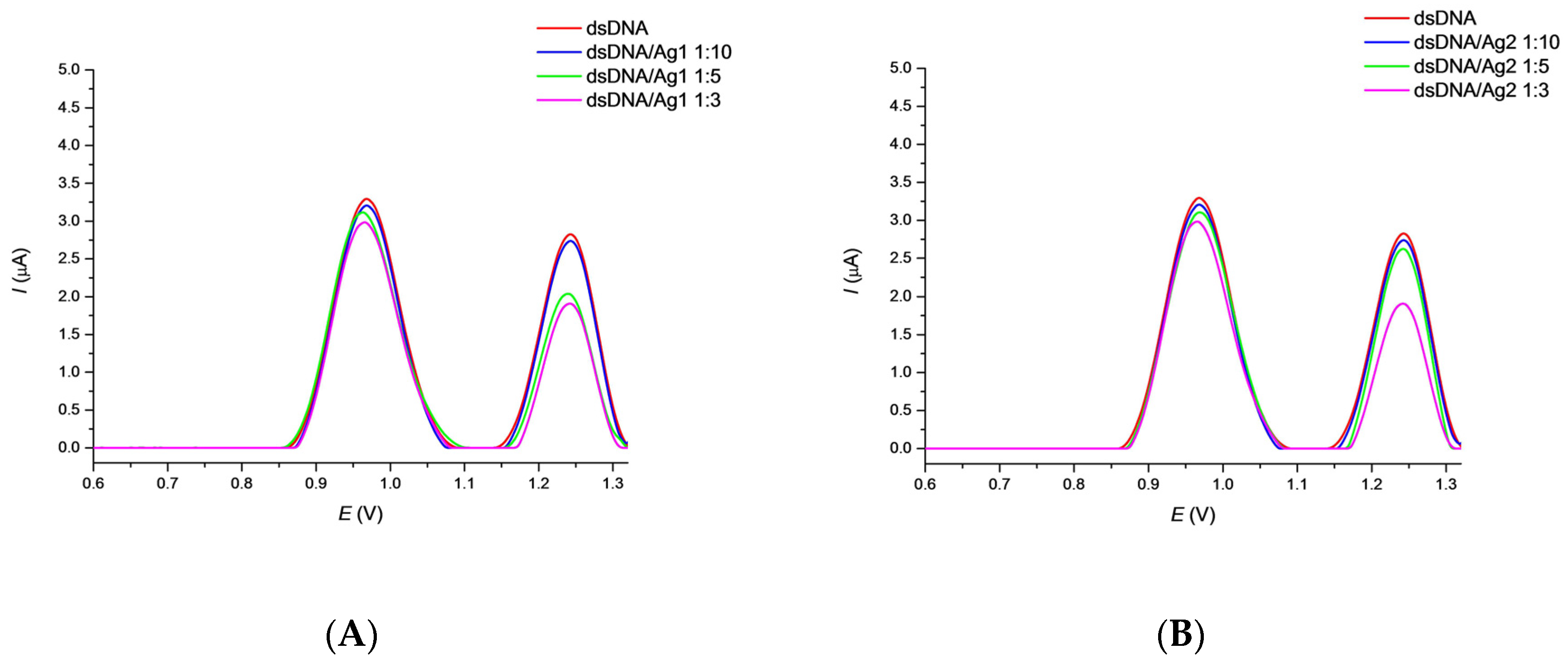
3.2. Effect of AgNPs on DNA/GCE Biosensor
3.3. Effect of TCS on DNA/GCE Biosensor
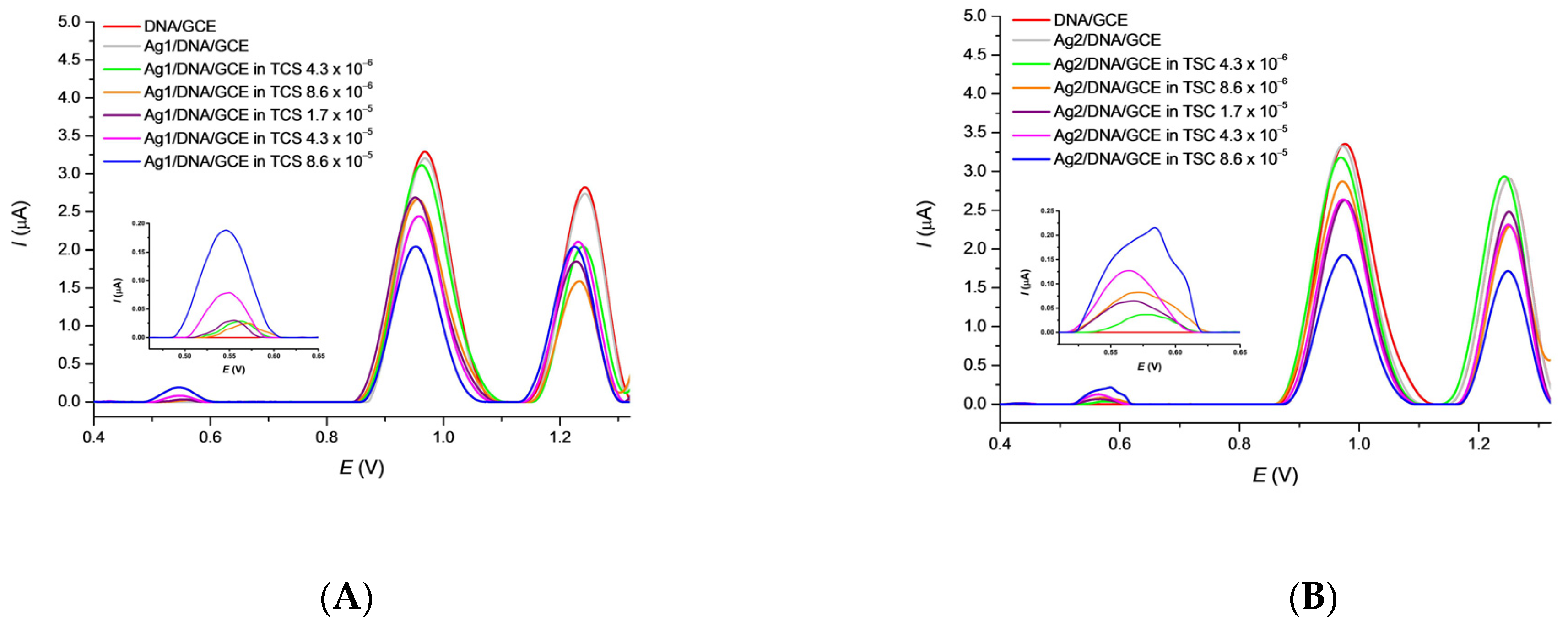
3.4. Effect of TCS on Ag/DNA/GCE Biosensor
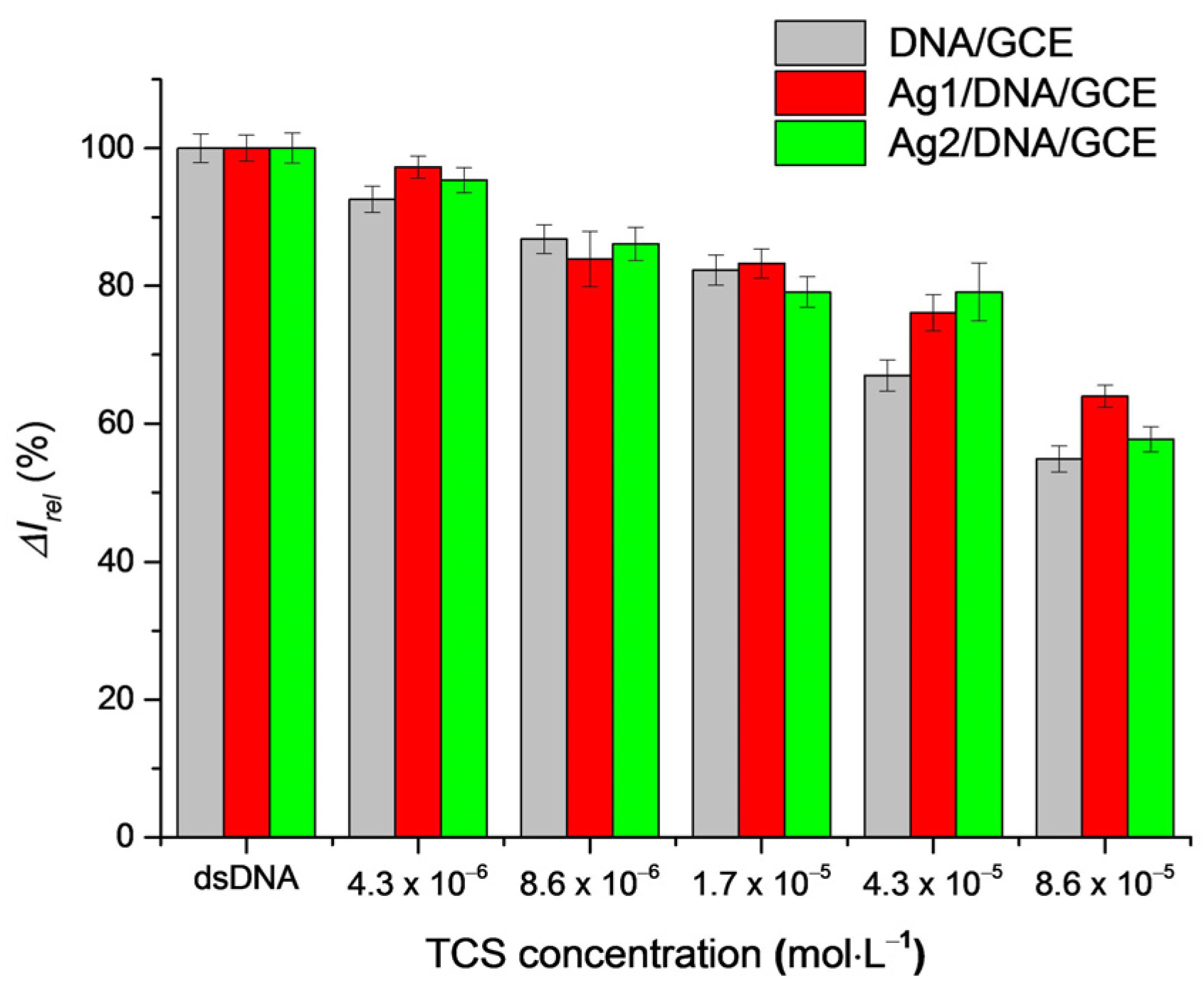
3.5. Protective Effect of AgNPs on DNA Damage by TCS
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Oliver, M.; Kudlak, B.; Wieczerzak, M.; Reis, S.; Lima, S.A.C.; Segundo, M.A.; Miro, M. Ecotoxicological equilibria of triclosan in Microtox, XenoScreen YES/YAS, Caco2, HEPG2 and liposomal systems are affected by the occurrence of other pharmaceutical and personal care emerging contaminants. Sci. Total Environ. 2020, 719, 137358. [Google Scholar] [CrossRef]
- WW-Chung, P.; Rafqah, S.; Voyard, G.; Sarakha, M. Photochemical behaviour of triclosan in aqueous solutions: Kinetic and analytical studies. J. Photochem. Photobiol. A Chem. 2007, 191, 201–208. [Google Scholar] [CrossRef]
- Thompson, A.; Griffin, P.; Stuetz, R.; Cartmell, E. The fate and removal of triclosan during wastewater treatment. Water Environ. Res. 2005, 77, 63–67. [Google Scholar] [CrossRef]
- Koeppe, E.S.; Ferguson, K.K.; Colacino, J.A.; Meeker, J.D. Relationship between urinary Triclosan and paraben concentrations and serum thyroid measures in NHANES 2007–2008. Sci. Total Environ. 2013, 445–446, 299–305. [Google Scholar] [CrossRef] [Green Version]
- Yueh, M.-F.; Tukey, R.H. Triclosan: A Widespread Environmental Toxicant with Many Biological Effects. Annu. Rev. Pharmacol. Toxicol. 2016, 56, 251–272. [Google Scholar] [CrossRef] [Green Version]
- Falfushynska, H.I.; Gnatyshyna, L.L.; Osadchuk, O.Y.; Farkas, A.; Vehovszky, A.; Carpenter, D.O.; Gyori, J.; Stoliar, O.B. Diversity of the molecular responses to separate wastewater effluents in freshwater mussels. Comp. Biochem. Physiol. Part C Toxicol. Pharmacol. 2014, 164, 51–58. [Google Scholar] [CrossRef]
- Carrouel, F.; Viennot, S.; Ottolenghi, L.; Gaillard, C.; Bourgeois, D. Nanoparticles as Anti-Microbial, Anti-Inflammatory, and Remineralizing Agents in Oral Care Cosmetics: A Review of the Current Situation. Nanomaterials 2020, 10, 140. [Google Scholar] [CrossRef] [Green Version]
- Fu, J.; Gong, Z.; Bae, S. Assessment of the Effect of Methyl-Triclosan and Its Mixture with Triclosan on Developing Zebrafish (Danio rerio) Embryos Using Mass Spectrometry-Based Metabolomics. J. Hazard. Mater. 2019, 368, 186–196. [Google Scholar] [CrossRef]
- Peng, F.J.; Hu, L.X.; Pan, C.G.; Ying, G.G.; Van den Brink, P.J. Insights into the Sediment Toxicity of Personal Care Products to Freshwater Oligochaete Worms Using Fourier Transform Infrared Spectroscopy. Ecotoxicol. Environ. Saf. 2019, 172, 296–302. [Google Scholar] [CrossRef]
- Hovander, L.; Malmberg, T.; Athanasiadou, M.; Athanassiadis, I.; Rahm, S.; Bergman, A.; Wehler, E.K. Identification of hydroxylated PCB metabolites and other phenolic halogenated pollutants in human blood plasma. Arch. Environ. Contam. Toxicol. 2002, 42, 105–117. [Google Scholar] [CrossRef]
- Calafat, A.M.; Ye, X.; Wong, L.Y.; Reidy, J.A.; Needham, L.L. Urinary Concentrations of Triclosan in the U.S. Population: 2003–2004. Environ. Health Perspect. 2008, 116, 303–307. [Google Scholar] [CrossRef]
- Fang, J.L.; Stingley, R.L.; Beland, F.A.; Harrouk, W.; Lumpkins, D.L.; Howard, P. Occurrence, efficacy, metabolism, and toxicity of triclosan. J. Environ. Sci. Health C Environ. Carcinog. Ecotoxicol. Rev. 2010, 28, 147–171. [Google Scholar] [CrossRef] [PubMed]
- Honkisz, E.; Zieba-Przybylska, D.; Wojtowicz, A.K. The effect of triclosan on hormone secretion and viability of human choriocarcinoma JEG-3 cells. Reprod. Toxicol. 2012, 34, 385–392. [Google Scholar] [CrossRef] [PubMed]
- Geens, T.; Neels, H.; Covaci, A. Distribution of bisphenol-A, triclosan and n-nonylphenol in human adipose tissue, liver and brain. Chemosphere 2012, 87, 796–802. [Google Scholar] [CrossRef] [PubMed]
- Wolff, M.S.; Teitelbaum, S.L.; Windham, G.; Pinney, S.M.; Britton, J.A.; Chelimo, C.; Godbold, J.; Biro, F.; Kushi, L.; Pfeiffer, C.M.; et al. Pilot study of urinary biomarkers of phytoestrogens, phthalates, and phenols in girls. Environ. Health Perspect. 2007, 115, 116–121. [Google Scholar] [CrossRef] [Green Version]
- Rodricks, J.V.; Swenberg, J.A.; Borzelleca, J.F.; Maronpot, R.R.; Shipp, A.M. Triclosan: A critical review of the experimental data and development of margins of safety for consumer products. Crit. Rev. Toxicol. 2010, 40, 422–484. [Google Scholar] [CrossRef]
- Yueh, M.F.; Taniguchi, K.; Chen, S.; Evans, R.M.; Hammock, B.D.; Karin, M.; Tukey, R.H. The commonly used antimicrobial additive triclosan is a liver tumor promoter. Proc. Natl. Acad. Sci. USA 2014, 111, 17200–17205. [Google Scholar] [CrossRef] [Green Version]
- Szychowski, K.A.; Wnuk, A.; Kajta, M.; Wojtowicz, A.K. Triclosan activates aryl hydrocarbon receptor (AhR)-dependent apoptosis and affects Cyp1a1 and Cyp1b1 expression in mouse neocortical neurons. Environ. Res. 2016, 151, 106–114. [Google Scholar] [CrossRef]
- Kim, J.; Oh, H.; Ryu, B.; Kim, U.; Lee, J.M.; Jung, C.R.; Kim, C.; Park, J.H. TCS affects axon formation in the neural development stages of zebrafish embryos (Danio rerio). Environ. Pollut. 2018, 236, 304–312. [Google Scholar] [CrossRef]
- Park, B.K.; Gonzales, E.L.; Yang, S.M.; Bang, M.; Choi, C.S.; Shin, C.Y. Effects of triclosan on neural stem cell viability and survival. Biol. Ther. 2016, 24, 99–107. [Google Scholar] [CrossRef] [Green Version]
- Wang, F.; Guo, X.; Chen, W.; Sun, Y.; Fan, C. Effects of triclosan on hormones and reproductive axis in female Yellow River carp (Cyprinus carpio): Potential mechanisms underlying estrogen effect. Toxicol. Appl. Pharm. 2017, 336, 49–54. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Sun, L.; Zhang, H.; Shi, M.; Dahlgren, R.A.; Wang, X.; Wang, H. Response mechanisms to joint exposure of triclosan and its chlorinated derivatives on zebrafish (Danio rerio) behavior. Chemosphere 2018, 193, 820–832. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Arya, S.; Dwivedi, A.K.; Alvarado, L.; Kupesic-Plavsic, S. Exposure of U.S. population to endocrine disruptive chemicals (parabens, benzophenone-3, bisphenol-a and triclosan) and their associations with female infertility. Environ. Pollut. 2020, 265, 114763. [Google Scholar] [CrossRef] [PubMed]
- Yee, A.L.; Gilbert, J.A. Is triclosan harming your microbiome? Science 2016, 353, 348–349. [Google Scholar] [CrossRef] [PubMed]
- Jutkina, J.; Marathe, N.P.; Flach, C.F.; Larsson, D.G.J. Antibiotics and common antibacterial biocides stimulate horizontal transfer of resistance at low concentrations. Sci. Total Environ. 2018, 616, 172–178. [Google Scholar] [CrossRef]
- Lu, J.; Wang, Y.; Zhang, S.; Bond, P.; Yuan, Z.; Guo, J. Triclosan at environmental concentrations can enhance the spread of extracellular antibiotic resistance genes through transformation. Sci. Total Environ. 2020, 713, 136621. [Google Scholar] [CrossRef]
- Sanidad, K.Z.; Xiao, H.; Zhang, G. Triclosan, a common antimicrobial ingredient, on gut microbiota and gut health. Gut. Microbes 2019, 10, 434–437. [Google Scholar] [CrossRef]
- Ruszkiewicz, J.A.; Li, S.; Rodriguez, M.B.; Aschner, M. Is triclosan a neurotoxic agent? J. Toxicol. Environ. Health B Crit. Rev. 2017, 20, 104–117. [Google Scholar] [CrossRef]
- Bertelsen, R.J.; Longnecker, M.P.; Lovik, M.; Calafat, A.M.; Carlsen, K.H.; London, S.J.; Lødrup Carlsen, K.C. Triclosan exposure and allergic sensitization in Norwegian children. Allergy 2013, 68, 84–91. [Google Scholar] [CrossRef]
- Wang, X.; Zou, L.; Mi, C.; Yu, H.; Dong, M.; Teng, Y. Characterization of binding interaction of triclosan and bovine serum albumin. J. Environ. Sci. Health A 2020, 55, 318–325. [Google Scholar] [CrossRef]
- Roszak, J.; Smok-Pieniazek, A.; Spryszynska, S.; Kowalczyk, K.; Domeradzka-Gajda, K.; Swiercz, R.; Grobelny, J.; Tomaszewska, E.; Ranoszek-Soliwoda, K.; Celichowski, G.; et al. Cytotoxic effects in transformed and non-transformed human breast cell lines after exposure to silver nanoparticles in combination with selected aluminium compounds, parabens or phthalates. J. Hazard. Mater. 2020, 392, 122442. [Google Scholar] [CrossRef] [PubMed]
- Bondarenko, O.; Juganson, K.; Ivask, A.; Kasemets, K.; Mortimer, M.; Kahru, A. Toxicity of Ag, CuO and ZnO Nanoparticles to Selected Environmentally Relevant Test Organisms and Mammalian Cells In Vitro: A Critical Review. Arch. Toxicol. 2013, 87, 1181–1200. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ahamed, M.; AlSalhi, M.S.; Siddiqui, M.K.J. Silver Nanoparticle Applications and Human Health. Clin. Chim. Acta 2010, 411, 1841–1848. [Google Scholar] [CrossRef]
- Vance, M.E.; Kuiken, T.; Vejerano, E.P.; McGinnis, S.P.; Hochella, M.F.; Rejeski, D.; Hull, M.S. Nanotechnology in the Real World: Redeveloping the Nanomaterial Consumer Products Inventory. Beilstein J. Nanotechnol. 2015, 6, 1769–1780. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, M.; Zhang, C.H. Are silver nanoparticles better than triclosan as a daily antimicrobial? Answers from the perspectives of gut microbiome disruption and pathogenicity. Sci. Total Environ. 2021, 756, 143983. [Google Scholar] [CrossRef] [PubMed]
- Prabhu, S.; Poulose, E.K. Silver nanoparticles: Mechanism of antimicrobial action, synthesis, medical applications, and toxicity efects. Int. Nano Lett. 2012, 2, 32. [Google Scholar] [CrossRef] [Green Version]
- Gaillet, S.; Rouanet, J.-M. Silver nanoparticles: Their potential toxic effects after oral exposure and underlying mechanisms—A review. Food Chem. Toxicol. 2015, 77, 58–63. [Google Scholar] [CrossRef] [PubMed]
- Do Nascimento, C.; Paulo, D.F.; Pita, M.S.; Pedrazzi, V.; de Albuquerque Junior, R.F. Microbial diversity of the supra- and subgingival biofilm of healthy individuals after brushing with chlorhexidine- or silver-coated toothbrush bristles. Can. J. Microbiol. 2015, 61, 112–123. [Google Scholar] [CrossRef]
- Mackevica, A.; Olsson, M.E.; Hansen, S.F. The release of silver nanoparticles from commercial toothbrushes. J. Hazard. Mater. 2017, 322, 270–275. [Google Scholar] [CrossRef] [Green Version]
- Li, Y.; Cummins, E. Hazard characterization of silver nanoparticles for human exposure routes. J. Environ. Sci. Health A 2020, 55, 704–725. [Google Scholar] [CrossRef]
- Shang, L.; Nienhaus, K.; Nienhaus, G.U. Engineered nanoparticles interacting with cells: Size matter. J. Nanobiotechnol. 2014, 12, 5. [Google Scholar] [CrossRef] [Green Version]
- Park, E.J.; Bae, E.; Yi, J.; Kim, Y.; Choi, K.; Lee, S.H.; Yoon, J.; Lee, B.C.; Park, K. Repeated-Dose Toxicity and Inflammatory Responses in Mice by Oral Administration of Silver Nanoparticles. Environ. Toxicol. Pharmacol. 2010, 30, 162–168. [Google Scholar] [CrossRef]
- Anderson, D.S.; Silva, R.M.; Lee, D.; Edwards, P.C.; Sharmah, A.; Guo, T.; Pinkerton, K.E.; Van Winkle, L.S. Persistence of Silver Nanoparticles in the Rat Lung: Influence of Dose, Size, and Chemical Composition. Nanotoxicology 2015, 9, 591–602. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, J.; Tang, M.; Xue, Y. Review of the effects of silver nanoparticle exposure on gut bacteria. J. Appl. Toxicol. 2019, 39, 27–37. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Van den Brule, S.; Ambroise, J.; Lecloux, H.; Levard, C.; Soulas, R.; De Temmerman, P.-J.; Palmai-Pallag, M.; Marbaix, E.; Lison, D. Dietary silver nanoparticles can disturb the gut microbiota in mice. Part. Fibre Toxicol. 2016, 13, 38. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Simko, M.; Fiedeler, U.; Gazsó, A.; Nentwich, M. The impact of nanoparticles on cellular functions. Nano Trust-Dossier 2011, 7, 151–164. [Google Scholar]
- Panacek, A.; Kvítek, L.; Prucek, R.; Kolar, M.; Vecerova, R.; Pizúrova, N.; Sharma, V.K.; Nevecna, T.; Zboril, R. Silver colloid nanoparticles: Synthesis, characterization, and their antibacterial activity. J. Phys. Chem. B 2006, 110, 16248–16253. [Google Scholar] [CrossRef]
- Morones, J.R.; Elechiguerra, J.L.; Camacho, A.; Holt, K.; Kouri, J.B.; Ramírez, J.T.; Yacaman, M.J. The bactericidal efect of silver nanoparticles. Nanotechnology 2005, 16, 2346–2353. [Google Scholar] [CrossRef] [Green Version]
- Qasim, M.; Udomluck, N.; Chang, J.; Park, H.; Kim, K. Antimicrobial activity of silver nanoparticles encapsulated in poly-N-isopropylacrylamide-based polymeric nanoparticles. Int. J. Nanomed. 2018, 13, 235–249. [Google Scholar] [CrossRef] [Green Version]
- Sotiriou, G.A.; Pratsinis, S.E. Antibacterial activity of nanosilver ions and particles. Environ. Sci. Technol. 2010, 44, 5649–5654. [Google Scholar] [CrossRef]
- Nateghi, M.R.; Hajimirzababa, H. Efect of silver nanoparticles morphologies on antimicrobial properties of cotton fabrics. J. Text. Inst. 2014, 105, 806–813. [Google Scholar] [CrossRef]
- Lkhagvajav, N.; Koizhaiganova, M.; Yasa, I.; Çelik, E.; Sari, Ö. Characterization and antimicrobial performance of nano silver coatings on leather materials. Braz. J. Microbiol. 2015, 46, 41–48. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Davenport, L.L.; Hsieh, H.; Eppert, B.L.; Carreira, V.S.; Krishan, M.; Ingle, T.; Howard, P.C.; Williams, M.T.; Vorhees, C.V.; Genter, M.B. Systemic and Behavioral Effects of Intranasal Administration of Silver Nanoparticles. Neurotoxicol. Teratol. 2015, 51, 68–76. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ghaderi, S.; Tabatabaei, S.R.F.; Varzi, H.N.; Rashno, M. Induced Adverse Effects of Prenatal Exposure to Silver Nanoparticles on Neurobehavioral Development of Offspring of Mice. J. Toxicol. Sci. 2015, 40, 263–275. [Google Scholar] [CrossRef] [Green Version]
- Tak, Y.K.; Pal, S.; Naoghare, P.K.; Rangasamy, S.; Song, J.M. Shape-Dependent Skin Penetration of Silver Nanoparticles: Does It Really Matter? Sci. Rep. 2015, 5, 16908. [Google Scholar] [CrossRef] [Green Version]
- Queckenberg, C.; Meins, J.; Wachall, B.; Doroshyenko, O.; Tomalik-Scharte, D.; Bastian, B.; Abdel-Tawab, M.; Fuhr, U. Absorption, pharmacokinetics, and safety of triclosan after dermal administration. Antimicrob. Agents Chemother. 2010, 54, 570–572. [Google Scholar] [CrossRef] [Green Version]
- Sandborgh-Englund, G.; Adolfsson-Erici, M.; Odham, G.; Ekstrand, J. Pharmacokinetics of triclosan following oral ingestion in humans. J. Toxicol. Environ. Health Part A 2006, 69, 1861–1873. [Google Scholar] [CrossRef]
- Weatherly, L.M.; Nelson, A.J.; Shim, J.; Riitano, A.M.; Gerson, E.D.; Hart, A.J.; de Juan-Sanz, J.; Ryan, T.A.; Sher, R.; Hess, S.T.; et al. Antimicrobial agent triclosan disrupts mitochondrial structure, revealed by super-resolution microscopy, and inhibits mast cell signaling via calcium modulation. Toxicol. Appl. Pharmacol. 2018, 349, 39–54. [Google Scholar] [CrossRef]
- Alfhili, M.A.; Hindawi, M.-H.L. Review Article Triclosan: An Update on Biochemical and Molecular Mechanisms. Oxid. Med. Cell. Longev. 2019, 2019, 1607304. [Google Scholar] [CrossRef]
- Ma, H.; Zheng, L.; Li, Y.; Pan, S.; Hu, J.; Yu, Z.; Zhang, G.; Sheng, G.; Fu, J. Triclosan reduces the levels of global DNA methylation in HepG2 cells. Chemosphere 2013, 90, 1023–1029. [Google Scholar] [CrossRef]
- Binelli, A.; Cogni, D.; Parolini, M.; Riva, C.; Provini, A. Cytotoxic and genotoxic effects of in vitro exposure to triclosan and trimethoprim on zebra mussel (Dreissena polymorpha) hemocytes. Comp. Biochem. Physiol. Part C Toxicol. Pharmacol. 2009, 150, 50–56. [Google Scholar] [CrossRef] [PubMed]
- Lovecka, P.; Macurkova, A.; Zaruba, K.; Hubacek, T.; Siegel, J.; Valentova, O. Genomic Damage Induced in Nicotiana tabacum L. Plants by Colloidal Solution with Silver and Gold Nanoparticles. Plants 2021, 10, 1260. [Google Scholar] [CrossRef] [PubMed]
- Kurbanoglu, S.; Dogan-Topal, B.; Hlavata, L.; Labuda, J.; Ozkan, S.A.; Uslu, B. Electrochemical investigation of an interaction of the antidepressant drug aripiprazole with original and damaged calf thymus dsDNA. Electrochim. Acta 2015, 169, 233–240. [Google Scholar] [CrossRef]
- Blaskovicova, J.; Sochr, J.; Koutsogiannis, A.; Diamantidou, D.; Kopel, P.; Adam, V.; Labuda, J. Detection of ROS Generated by UV-C Irradiation of CdS quantum Dots and their Effect on Damage to Chromosomal and Plasmid DNA. Electroanalysis 2018, 30, 698–704. [Google Scholar] [CrossRef]
- Silva, E.H.C.; Lopes, I.C.; Bruzaca, E.E.S.; de Carvalho, P.A.V.; Tanaka, A.A. Triclosan: Electrochemistry, Spontaneous Degradation and Effects on Double-Stranded DNA. Braz. J. Anal. Chem. 2021, 8, 89–102. [Google Scholar] [CrossRef]
- Talebpour, Z.; Haghighi, F.; Taheri, M.; Hosseinzadeh, M.; Gharavi, S.; Habibi, F.; Aliahmadi, A.; Sadr, A.S.; Azad, J. Binding interaction of spherical silver nanoparticles and calf thymus DNA: Comprehensive multispectroscopic, molecular docking, and RAPD PCR studies. J. Mol. Liquids 2019, 289, 111185. [Google Scholar] [CrossRef]
- Martinez-Castanon, G.A.; Niño-Martínez, N.; Martínez-Gutierrez, F.; Martínez-Mendoza, J.R.; Ruiz, F. Synthesis and antibacterial activity of silver nanoparticles with different sizes. J. Nanopart. Res. 2008, 10, 1343–1348. [Google Scholar] [CrossRef]
- Nemcekova, K.; Svitkova, V.; Sochr, J.; Gemeiner, P.; Labuda, J. Gallic acid-coated silver nanoparticles as perspective drug nanocarriers: Bioanalytical study. Anal. Bioanal. Chem. 2022, 1–13. [Google Scholar] [CrossRef]
- Yola, M.L.; Atar, N.; Eren, T.; Karimi-Malehc, H.; Wang, S. Sensitive and selective determination of aqueous triclosan based on gold nanoparticles on polyoxometalate/reduced graphene oxide nanohybrid. RSC Adv. 2015, 5, 65953. [Google Scholar] [CrossRef] [Green Version]
- Peropadre, A.; Blanco, L.; Freire, P.F.; Repetto, G.; Hazen, M. Exposure to low doses of triclosan induces DNA-damage and increased proliferation in human keratinocytes. Toxicol. Lett. 2016, 258, S248. [Google Scholar] [CrossRef]
- Stamatis, N.; Antonopoulou, M.; Hela, D.; Konstantinou, I. Photocatalytic degradation kinetics and mechanisms of antibacterial triclosan in aqueous TiO2 suspensions under simulated solar irradiation. J. Chem. Technol. Biotechnol. 2014, 89, 1145–1154. [Google Scholar] [CrossRef]

Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Blaškovičová, J.; Labuda, J. Effect of Triclosan and Silver Nanoparticles on DNA Damage Investigated with DNA-Based Biosensor. Sensors 2022, 22, 4332. https://doi.org/10.3390/s22124332
Blaškovičová J, Labuda J. Effect of Triclosan and Silver Nanoparticles on DNA Damage Investigated with DNA-Based Biosensor. Sensors. 2022; 22(12):4332. https://doi.org/10.3390/s22124332
Chicago/Turabian StyleBlaškovičová, Jana, and Ján Labuda. 2022. "Effect of Triclosan and Silver Nanoparticles on DNA Damage Investigated with DNA-Based Biosensor" Sensors 22, no. 12: 4332. https://doi.org/10.3390/s22124332
APA StyleBlaškovičová, J., & Labuda, J. (2022). Effect of Triclosan and Silver Nanoparticles on DNA Damage Investigated with DNA-Based Biosensor. Sensors, 22(12), 4332. https://doi.org/10.3390/s22124332





