In Silico and In Vitro Inhibition of SARS-CoV-2 PLpro with Gramicidin D
Abstract
1. Introduction
2. Results
2.1. Molecular Docking
2.2. Purification of PLpro
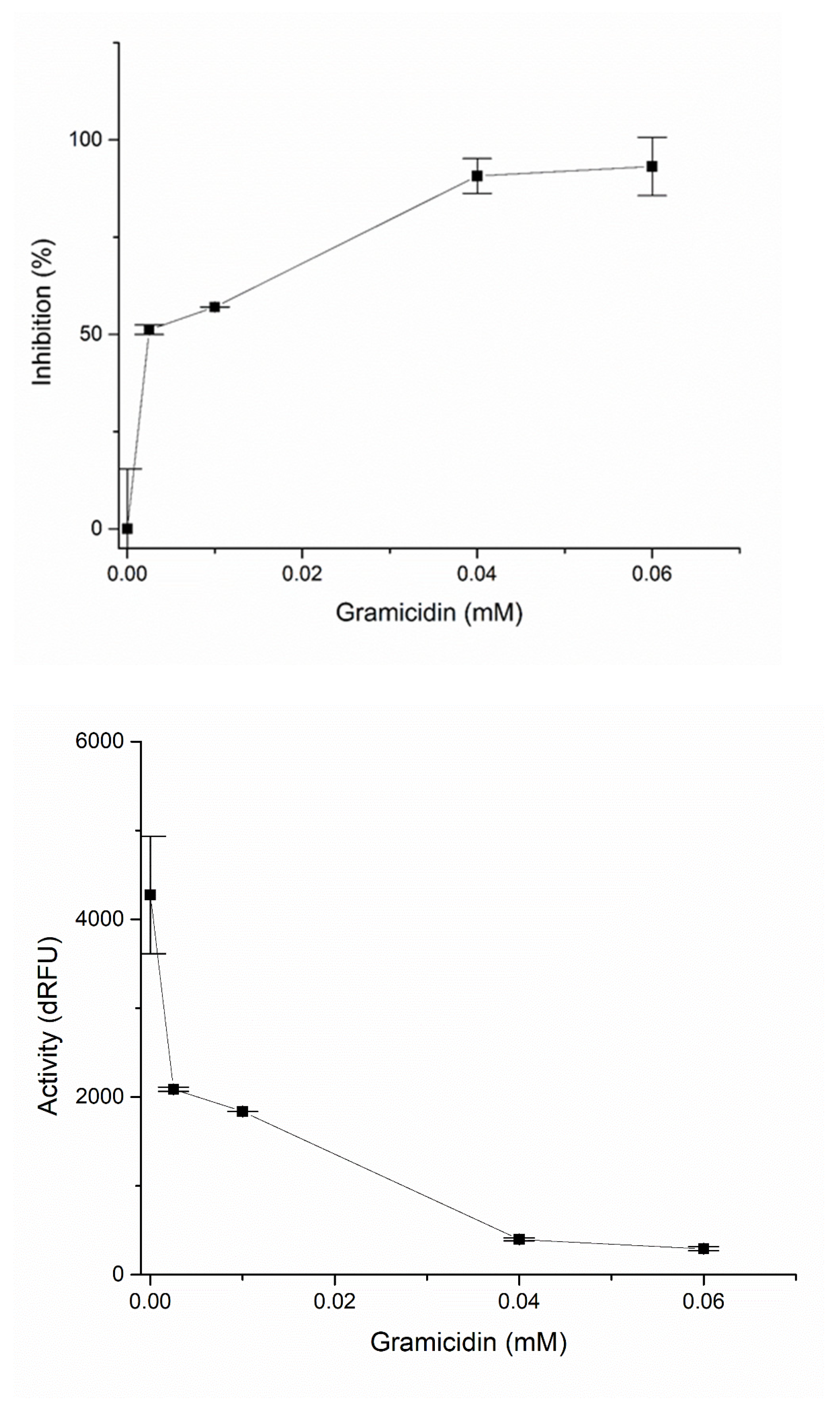
2.3. PLpro Inhibition with Gramicidin D
3. Materials and Methods
3.1. Data Preparation
3.2. ISM
3.3. Molecular Docking
3.4. Electrostatic Potential (ESP) Surface Calculations
3.5. Equipment
3.6. Chemicals
3.7. Gene for PLpro
3.8. Amino Acid Sequence of PLpro
3.9. Gramicidin D and S IUPAC Entry
3.10. Isolation and Purification
3.10.1. Cell Lysis
3.10.2. Purification
3.10.3. PLpro Assay
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Ghanbari, R.; Teimoori, A.; Sadeghi, A.; Mohamadkhani, A.; Rezasoltani, S.; Asadi, E.; Jouyban, A.; Sumner, S.C. Existing antiviral options against SARS-CoV-2 replication in COVID-19 patients. Future Microbiol. 2020, 15, 1747–1758. [Google Scholar] [CrossRef]
- Shin, D.; Mukherjee, R.; Grewe, D.; Bojkova, D.; Baek, K.; Bhattacharya, A.; Schulz, L.; Widera, M.; Mehdipour, A.R.; Tascher, G.; et al. Papain-like protease regulates SARS-CoV-2 viral spread and innate immunity. Nature 2020, 587, 657–662. [Google Scholar] [CrossRef] [PubMed]
- Devaraj, S.G.; Wang, N.; Chen, Z.; Chen, Z.; Tseng, M.; Barretto, N.; Lin, R.; Peters, C.J.; Tseng, C.T.; Baker, S.C.; et al. Regulation of IRF-3-dependent innate immunity by the papain-like protease domain of the severe acute respiratory syndrome coronavirus. J. Biol. Chem. 2007, 282, 32208–32221. [Google Scholar] [CrossRef]
- McClain, C.B.; Vabret, N. SARS-CoV-2: The many pros of targeting PLpro. Signal Transduct. Target. Ther. 2020, 5, 223. [Google Scholar] [CrossRef] [PubMed]
- Maiti, B.K. Can papain-like protease inhibitors halt SARS-CoV-2 replication? ACS Pharmacol. Transl. Sci. 2020, 3, 1017–1019. [Google Scholar] [CrossRef] [PubMed]
- Poreba, M.; Salvesen, G.S.; Drag, M. Synthesis of a HyCoSuL peptide substrate library to dissect protease substrate specificity. Nat. Protoc. 2017, 12, 2189–2214. [Google Scholar] [CrossRef]
- Rut, W.; Lv, Z.; Zmudzinski, M.; Patchett, S.; Nayak, D.; Snipas, S.J.; Oualid, F.E.; Huang, T.T.; Bekes, M.; Drag, M.; et al. Activity profiling and structures of inhibitor-bound SARS-CoV-2-PLpro protease provides a framework for anti-COVID-19 drug design. bioRxiv 2020. [Google Scholar] [CrossRef]
- Huan, Y.; Kong, Q.; Mou, H.; Yi, H. Antimicrobial Peptides: Classification, Design, Application and Research Progress in Multiple Fields. Front. Microbiol. 2020, 11, 582779. [Google Scholar] [CrossRef]
- Talapko, J.; Meštrović, T.; Juzbašić, M.; Tomas, M.; Erić, S.; Horvat, A.L.; Bekić, S.; Schwarz, D.; Matić, S.; Neuberg, M.; et al. Antimicrobial Peptides-Mechanisms of Action, Antimicrobial Effects and Clinical Applications. Antibiotics 2022, 11, 1417. [Google Scholar] [CrossRef]
- Mousavi Maleki, M.S.; Rostamian, M.; Madanchi, H. Antimicrobial peptides and other peptide-like therapeutics as promising candidates to combat SARS-CoV-2. Expert Rev. Anti-Infect. Ther. 2021, 19, 1205–1217. [Google Scholar] [CrossRef]
- Pérez de la Lastra, J.M.; Anand, U.; González-Acosta, S.; López, M.R.; Dey, A.; Bontempi, E.; Morales de la Nuez, A. Antimicrobial Resistance in the COVID-19 Landscape: Is There an Opportunity for Anti-Infective Antibodies and Antimicrobial Peptides? Front. Immunol. 2022, 13, 921483. [Google Scholar] [CrossRef] [PubMed]
- Thakur, A.; Sharma, A.; Alajangi, H.K.; Jaiswal, P.K.; Lim, Y.-B.; Singh, G.; Barnwal, R.P. In pursuit of next-generation therapeutics: Antimicrobial peptides against superbugs, their sources, mechanism of action, nanotechnology-based delivery, and clinical applications. Int. J. Biol. Macromol. 2022, 218, 135–156. [Google Scholar] [CrossRef] [PubMed]
- Bourinbaiar, A.S.; Krasinski, K.; Borkowsky, W. Anti-HIV effect of gramicidin in vitro: Potential for spermicide use. Life Sci. 1994, 54, PL5–PL9. [Google Scholar] [CrossRef] [PubMed]
- Bourinbaiar, A.S.; Coleman, C.F. The effect of gramicidin, a topical contraceptive and antimicrobial agent with anti-HIV activity, against herpes simplex viruses type 1 and 2 in vitro. Arch. Virol. 1997, 142, 2225–2235. [Google Scholar] [CrossRef]
- Prenner, E.J.; Lewis, R.N.A.H.; McElhaney, R.N. The interaction of the antimicrobial peptide gramicidin S with lipidbilayer model and biological membranes. Biochem. Biophys. Acta 1999, 1462, 201–221. [Google Scholar] [CrossRef]
- Enayathullah, M.G.; Parekh, Y.; Banu, S.; Ram, S.; Nagaraj, R.; Kumar, B.K.; Idris, M.M. Gramicidin S and melittin: Potential anti-viral therapeutic peptides to treat SARS-CoV-2 infection. Sci. Rep. 2022, 12, 3446. [Google Scholar] [CrossRef]
- Bansal, P.; Kumar, R.; Singh, J.; Dhanda, S. In silico molecular docking of SARS-CoV-2 surface proteins with microbial nonribosomal peptides: Identification of potential drugs. J. Proteins Proteom. 2021, 1, 1–8. [Google Scholar]
- Sencanski, M.; Perovic, V.; Milicevic, J.; Todorovic, T.; Prodanovic, R.; Veljkovic, V.; Paessler, S.; Glisic, S. Identification of SARS-CoV-2 Papain-like Protease (PLpro) Inhibitors Using Combined Computational Approach. ChemistryOpen 2022, 11, e202100248. [Google Scholar] [CrossRef]
- Wishart, D.S.; Feunang, Y.D.; Guo, A.C.; Lo, E.J.; Marcu, A.; Grant, J.R.; Sajed, T.; Johnson, D.; Li, C.; Sayeeda, Z.; et al. DrugBank 5.0: A major update to the DrugBank database for 2018. Nucleic Acids Res. 2018, 46, D1074–D1082. [Google Scholar] [CrossRef]
- Fu, Z.; Huang, B.; Tang, J.; Liu, S.; Liu, M.; Ye, Y.; Liu, Z.; Xiong, Y.; Zhu, W.; Cao, D.; et al. The complex structure of GRL0617 and SARS-CoV-2 PLpro reveals a hot spot for antiviral drug discovery. Nat. Commun. 2021, 12, 488. [Google Scholar] [CrossRef]
- Skwarecki, A.S.; Nowak, M.G.; Milewska, M.J. Amino Acid and Peptide-Based Antiviral Agents. ChemMedChem 2021, 16, 3106–3135. [Google Scholar] [CrossRef] [PubMed]
- Swierstra, J.; Kapoerchan, V.; Knijnenburg, A.; van Belkum, A.; Overhand, M. Structure, toxicity and antibiotic activity of gramicidin S and derivatives. Eur. J. Clin. Microbiol. Infect. Dis. 2016, 35, 763–769. [Google Scholar] [CrossRef] [PubMed]
- Wishart, D.S.; Knox, C.; Guo, A.C.; Cheng, D.; Shrivastava, S.; Tzur, D.; Gautam, B.; Hassanali, M. DrugBank: A knowledgebase for drugs, drug actions and drug targets. Nucleic Acids Res. 2008, 36, D901–D906. [Google Scholar] [CrossRef] [PubMed]
- Ahn, J.H.; Kim, J.; Hong, S.P.; Choi, S.Y.; Yang, M.J.; Ju, Y.S.; Kim, Y.T.; Kim, H.M.; Rahman, M.T.; Chung, M.K.; et al. Nasal ciliated cells are primary targets for SARS-CoV-2 replication in the early stage of COVID-19. J. Clin. Investig. 2021, 131, e148517. [Google Scholar] [CrossRef]
- Bustamante-Marin, X.M.; Ostrowski, L.E. Cilia and mucociliary clearance. Cold Spring Harb. Perspect. Biol. 2017, 9, 028241. [Google Scholar] [CrossRef]
- Keller, L.A.; Merkel, O.; Popp, A. Intranasal drug delivery: Opportunities and toxicologic challenges during drug development. Drug Deliv. Transl. Res. 2022, 12, 735–757. [Google Scholar] [CrossRef]
- Lewis, A.L.; Richard, J. Challenges in the delivery of peptide drugs: An industry perspective. Ther. Deliv. 2015, 6, 149–163. [Google Scholar] [CrossRef]
- Thoene, J.; Gavin, R.F.; Towne, A.; Wattay, L.; Ferrari, M.G.; Navarrete, J.; Pal, R. In vitro activity of cysteamine against SARS-CoV-2 variants. Mol. Genet. Metab. 2022, 137, 192–200. [Google Scholar] [CrossRef]
- Veljkovic, N.; Glisic, S.; Perovic, V.; Veljkovic, V. The role of long range intermolecular interactions in discovery of new drugs. Exp. Opin. Drug Discov. 2011, 6, 1263. [Google Scholar] [CrossRef]
- Veljkovic, V.; Veljkovic, N.; Este, J.; Huther, A.; Dietrich, U. Application of the EIIP/ISM bioinformatics concept in development of new drugs. Curr. Med. Chem. 2007, 14, 441. [Google Scholar] [CrossRef]
- Sencanski, M.; Sumonja, N.; Perovic, V.; Glisic, S.; Veljkovic, N.; Veljkovic, V. The Application of Information Spectrum Method on Small Molecules and Target Recognition. arXiv 2019, arXiv:1907.02713. [Google Scholar]
- James, J.P. MOPAC2016. In Stewart Computational Chemistry; Stewart Computational Chemistry: Colorado Springs, CO, USA, 2016. [Google Scholar]
- Stewart, J.J. Optimization of parameters for semiempirical methods VI: More modifications to the NDDO approximations and re-optimization of parameters. J. Mol. Model. 2013, 19, 1–32. [Google Scholar] [CrossRef] [PubMed]
- Trott, O.; Olson, A.J. AutoDock Vina: Improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 2010, 31, 455–461. [Google Scholar] [CrossRef] [PubMed]
- Frisch, M.J.; Trucks, G.W.; Schlegel, H.B.; Scuseria, G.E.; Robb, M.A.; Cheeseman, J.R.; Gonzalez, J.A.; Pople, J.A. Gaussian 09, Revision E. 01; Gaussian, Inc.: Wallingford, CT, USA, 2009; Volume 121, pp. 150–166. [Google Scholar]
- PubChem [Internet]. PubChem Compound Summary for CID 45267103, Gramicidin D; National Library of Medicine (US), National Center for Biotechnology Information: Bethesda, MD, USA, 2004. Available online: https://pubchem.ncbi.nlm.nih.gov/compound/Gramicidin-D (accessed on 13 January 2023).
- PubChem [Internet]. PubChem Compound Summary for CID 73357, Gramicidin S; National Library of Medicine (US), National Center for Biotechnology Information: Bethesda, MD, USA, 2004. Available online: https://pubchem.ncbi.nlm.nih.gov/compound/Gramicidin-S (accessed on 12 January 2023).

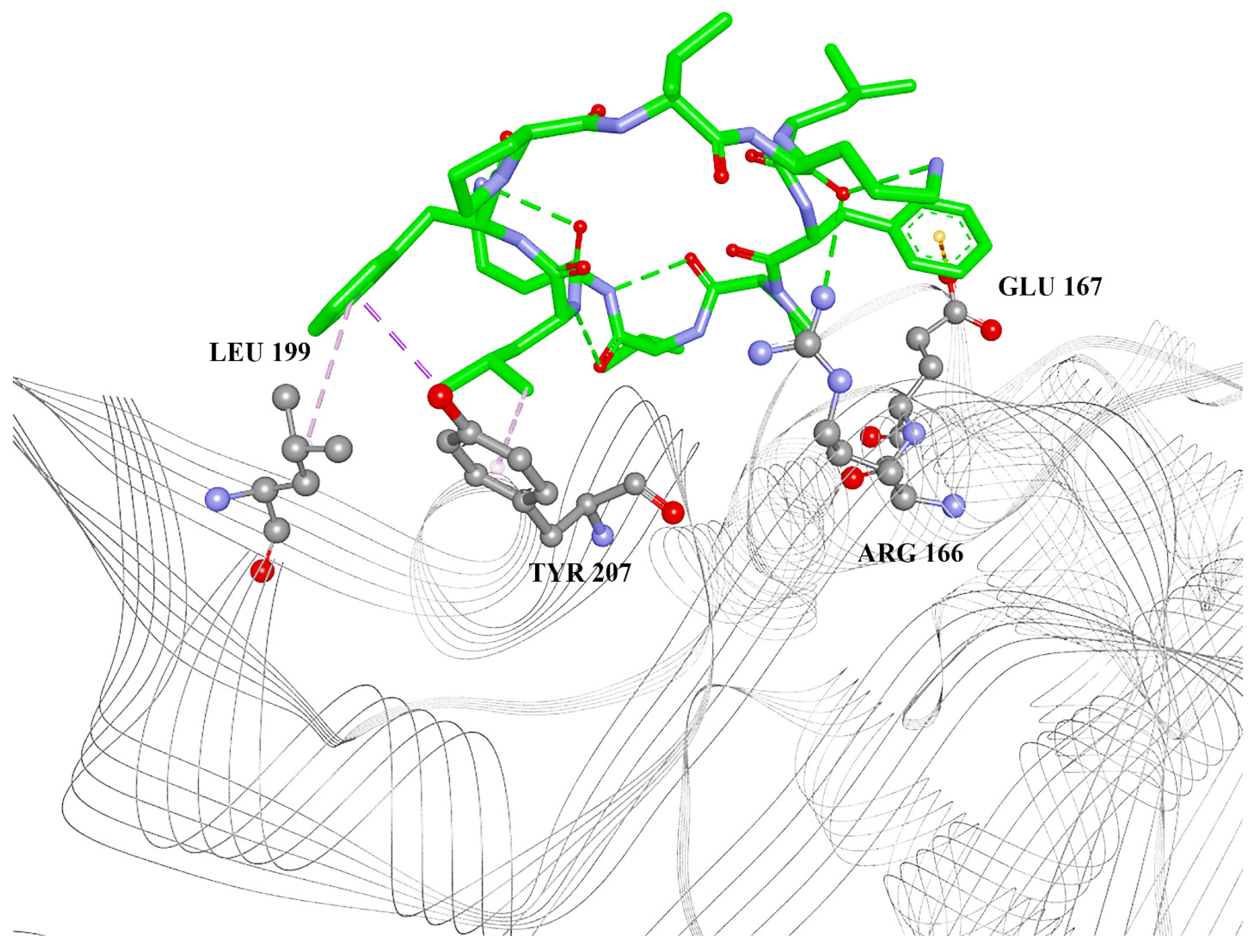

| Amino Acid Residue | Interaction Type | Gramicidin D | Gramicidin S |
|---|---|---|---|
| ASP164 | Pi-anion | Yes | No |
| ARG166 | Conventional hydrogen bond | Yes | Yes |
| GLU167 | Pi-anion | Yes | Yes |
| SER170 | Conventional hydrogen bond | Yes | No |
| TYR171 | Pi-alkyl | Yes | No |
| LEU199 | Pi-alkyl | No | Yes |
| MET206 | Conventional hydrogen bond, Pi-sulfur | Yes | No |
| TYR207 | Pi-alkyl | No | Yes |
| PRO247 | Pi-alkyl | Yes | No |
| PRO248 | Conventional hydrogen bond, Pi-alkyl | Yes | No |
| THR301 | Hydrogen bond | Yes | No |
| dRFU | Stdev | Stdev% | %Rezid. | %Inhibition | |
|---|---|---|---|---|---|
| 0 | 4274.67 | 660.27 | 15.45 | 100 | 0 |
| 0.0025 | 2087 | 25.46 | 1.22 | 48.82 | 51.18 |
| 0.01 | 1837 | 1.41 | 0.077 | 42.97 | 57.03 |
| 0.04 | 397.5 | 17.68 | 4.45 | 9.30 | 90.7 |
| 0.06 | 292.5 | 21.92 | 7.49 | 6.84 | 93.16 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Protić, S.; Kaličanin, N.; Sencanski, M.; Prodanović, O.; Milicevic, J.; Perovic, V.; Paessler, S.; Prodanović, R.; Glisic, S. In Silico and In Vitro Inhibition of SARS-CoV-2 PLpro with Gramicidin D. Int. J. Mol. Sci. 2023, 24, 1955. https://doi.org/10.3390/ijms24031955
Protić S, Kaličanin N, Sencanski M, Prodanović O, Milicevic J, Perovic V, Paessler S, Prodanović R, Glisic S. In Silico and In Vitro Inhibition of SARS-CoV-2 PLpro with Gramicidin D. International Journal of Molecular Sciences. 2023; 24(3):1955. https://doi.org/10.3390/ijms24031955
Chicago/Turabian StyleProtić, Sara, Nevena Kaličanin, Milan Sencanski, Olivera Prodanović, Jelena Milicevic, Vladimir Perovic, Slobodan Paessler, Radivoje Prodanović, and Sanja Glisic. 2023. "In Silico and In Vitro Inhibition of SARS-CoV-2 PLpro with Gramicidin D" International Journal of Molecular Sciences 24, no. 3: 1955. https://doi.org/10.3390/ijms24031955
APA StyleProtić, S., Kaličanin, N., Sencanski, M., Prodanović, O., Milicevic, J., Perovic, V., Paessler, S., Prodanović, R., & Glisic, S. (2023). In Silico and In Vitro Inhibition of SARS-CoV-2 PLpro with Gramicidin D. International Journal of Molecular Sciences, 24(3), 1955. https://doi.org/10.3390/ijms24031955








