Abstract
Soybean is an important crop that is grown worldwide. Flowering time is a critical agricultural trait determining successful reproduction and yields. For plants, light and temperature are important environmental factors that regulate flowering time. Soybean is a typical short-day (SD) plant, and many studies have elucidated the fine-scale mechanisms of how soybean responds to photoperiod. Low temperature can delay the flowering time of soybean, but little is known about the detailed mechanism of how temperature affects soybean flowering. In this study, we isolated GmFLC-like from soybean, which belongs to the FLOWERING LOCUS C clade of the MADS-box family and is intensely expressed in soybean leaves. Heterologous expression of GmFLC-like results in a delayed-flowering phenotype in Arabidopsis. Additional experiments revealed that GmFLC-like is involved in long-term low temperature-triggered late flowering by inhibiting FT gene expression. In addition, yeast one-hybrid, dual-luciferase reporter assay, and electrophoretic mobility shift assay revealed that the GmFLC-like protein could directly repress the expression of FT2a by physically interacting with its promoter region. Taken together, our results revealed that GmFLC-like functions as a floral repressor involved in flowering time during treatments with various low temperature durations. As the only the FLC gene in soybean, GmFLC-like was meaningfully retained in the soybean genome over the course of evolution, and this gene may play an important role in delaying flowering time and providing protective mechanisms against sporadic and extremely low temperatures.
1. Introduction
Understanding the molecular mechanism driving the change from vegetative to reproductive growth is crucial for maximizing the yield of seed crops in a given environment. As one of the most important traits, flowering is considered the developmental transition from juvenile to adult phase, and the flowering process is also regulated by various internal signals and environmental cues. Five major pathways controlling flowering time, which include the photoperiod, vernalization, autonomous, ageing, and gibberellin pathways, have been identified in Arabidopsis (Arabidopsis thaliana) on the basis of previous studies [1,2], and the photoperiod and vernalization pathways are triggered by environmental cues such as light and temperature [3,4].
Light, including the duration, quality, and intensity, is one of the most important environmental cues modulating flowering at multiple levels [5]. According to the day length, plants can be ranked into three categories: long-day (LD) plants, that proceed to flowering when the day length exceeds a threshold value; short-day (SD) plants, that flower in response to a shortening photoperiod; and day-neutral plants, whose flowering does not rely on the day length [6]. The change in light quality triggers multiple responses referred to as the “shade avoidance syndrome”. Plants focus on stem elongation instead of leaf expansion during the shade avoidance period and finally achieve the objective of early flowering [7]. Light intensity, one of most important parameters for light, has independent functions in the regulation of flowering time by accelerating the transition from juvenile to adult phase in Arabidopsis and other higher plants [5].
Apart from light, the temperature acts as another environmental signal affecting flowering time in two distinguishing ways [8]. The first way is that some species must undergo a prolonged cold period to obtain the capability of floral transition, and this cold-accumulation is known as vernalization [9,10]. Second, some species respond to the ambient temperature pathway, referring to non-stressful and physiological temperatures perceived by plants at the vegetative stage [8,11]. For some winter annual, biennial, and perennial plants, the vernalization requirements (at 1 to 7 °C) are absolutely necessary and determine reproductive success in spring or early summer. Previously, extensive genetic and molecular approaches exploring the underlying mechanism of vernalization in the model species, Arabidopsis thaliana, have identified that FLOWERING LOCUS C (FLC) is a major gene regulating vernalization requirements and responses. Briefly, vernalization represses the expression of FLC, a floral repressor [12,13]. Previous studies have shown that the vernalization pathway genes FRIGIDA (FRI) and VERNALIZATION INSENSITIVE 3 (VIN3) are key genes responsible for regulating FLC expression [14,15]. While the loss-of-function mutations in FRI displayed an early-flowering phenotype in Arabidopsis, this evolving phenotype has no need for the vernalization pathway [16,17]. VIN3, which is transiently induced by low temperature, is essential for FLC repression [18,19]. In addition, the epigenetic regulation is also involved in silencing FLC, including chromatin modification, COLD INDUCED LONG ANTISENSE INTRAGENIC RNA (COOLAIR), and COLD ASSISTED INTRONIC NONCOLDING RNA (COLDAIR) [20,21,22,23,24]. In Arabidopsis, the flowering genes FLOWERING LOCUS T (FT) and SOC1 are positive regulators of flowering, and the late-flowering phenotype could be due to a decrease in the expression level of FT and SUPPRESSOR OF OVEREXPRESSION OF CONSTANTS 1 (SOC1) by the FLC target [25,26,27,28].
The function of FLC in the response to vernalization was conserved in some species, such as Brassica napus, Brassica rapa, Sinapis alba, Capsella rubella, Beta vulgaris, and Cichorium intybus. These FLC-like genes, whose function is consistent with that of AtFLC, act as floral repressors and are downregulated in response to cold [29,30,31,32,33,34]. However, VERNALIZATION 1 and 2 (VRN1 and VRN2), the homologues of APETALA 1 (AP1) and FT in Arabidopsis, respectively, are responsible for the vernalization requirements instead of FLC in wheat and barley [35]. These findings indicate that different genes account for vernalization requirements in different species.
In contrast to the detailed understanding of vernalization, research in connection with the fine mechanism by which ambient temperatures affect flowering time has just started. In Arabidopsis, elevating the growth temperature from 23 to 27 °C dramatically facilitates flowering under SD conditions; this phenotype is a primary attribute of the change in FT expression levels [36]. In contrast, for some species such as Boechera stricta and Chrysanthemum morifolium, when growth temperatures moderately increase, the plants exhibit delayed flowering [37,38].
Soybean (Glycine max) is one of the most important crops worldwide for its nutritional qualities and oil content, and its flowering time is a critical agricultural trait determining successful reproduction and yields. Hence, identifying the functions of key soybean genes is of great importance for the genetic improvement of crops. Soybean is a typically SD plant whose flowering and maturation are strictly controlled by the photoperiod [39], and many studies have explored the fine-scale mechanisms of how soybean responds to the photoperiod [40,41,42,43]. In addition, although soybean is a non-vernalized plant, the temperature is also a major environmental factor affecting its flowering time. Low temperature could delay the flowering time of soybean, but little is known about the detailed mechanism of how temperature affects soybean flowering until now. In our previous study, we found that overexpression of AtDREB1A in soybean caused clearly delayed flowering [44]. qRT-PCR analyses of the expression of flowering time genes related to the vernalization pathway showed that Glyma11g13220 (GmVRN1-like) and Glyma05g28130 (designated as GmFLC-like) were strongly upregulated in the DEHYDRATION RESPONSE ELEMENT B1A (AtDREB1A)-overexpressing soybean [45]. Hence, we speculate that these genes may mainly account for the phenotype. Fortunately, we confirmed that the vernalization pathway gene Glyma11g13220 plays crucial roles in regulating flowering time [46]. In this study, we isolated Glyma05g28130 from soybean, which is intensely expressed in soybean leaves and is involved in long-term low temperature-triggered late flowering, belonging to the FLC clade of the MADS-box family. Additional experiments revealed that heterologous expression of GmFLC-like results in the phenotype of delayed flowering by inhibiting FT genes’ transcription in Arabidopsis. In addition, yeast one-hybrid (Y1H), dual-luciferase reporter assay, and electrophoretic mobility shift assay (EMSA) revealed that GmFLC-like protein could directly repress the expression of FT2a by physically interacting with its promoter region. In brief, our findings underline the importance of GmFLC-like in the soybean response to low temperature and highlight the role of this gene as a floral repressor in delaying flowering time and providing protective mechanisms against sporadic and extreme low temperatures.
2. Results
2.1. Glyma05g28130 Is a Homologue of AtFLC
Based on previous results from our laboratory, Glyma05g28130 plays crucial roles in modulating flowering time in soybean [45]. To identify the functions of Glyma05g28130 in regulating flowering time, we cloned the gene from the soybean cultivar “Huachun 5” referring to the sequence found in the Phytozome database [47]. The results showed that the cDNA sequence of Glyma05g28130 is 1513 bp in length, contains a 603 bp ORF and encodes 200 amino acid residues. Glyma05g28130 has a predicted DNA-binding MADS-domain in the N-terminus, followed by the K (keratin-like) regions, and the domains were predicted at amino acid residues 1–61 and 71–181, respectively (Figure 1A). In Arabidopsis, the ancient MIKC-type MADS-box genes were further classified into 13 distinctive subfamilies based on their phylogeny, namely, AGL2, AGL6, SQUA, AGL12, FLC, TM3, AGL17, AG, AGL15, DEF, GLO, GGM13, and STMADS11 genes [48,49,50]. Phylogenetic analyses revealed that Glyma05g28130 fell within the FLC clade of the Arabidopsis MIKC-type MADS-box family and shared a close relationship with AtFLC (AT5G10140) (Figure 1B). Hence, the gene corresponding to Glyma05g28130 was named GmFLC-like. In addition, we compared the amino acid sequence of GmFLC-like with FLC homologues of other species. The MADS-box domain sequence of GmFLC-like shared high conservation among different species, whereas conservation of the K domain was much weaker (Figure 2).
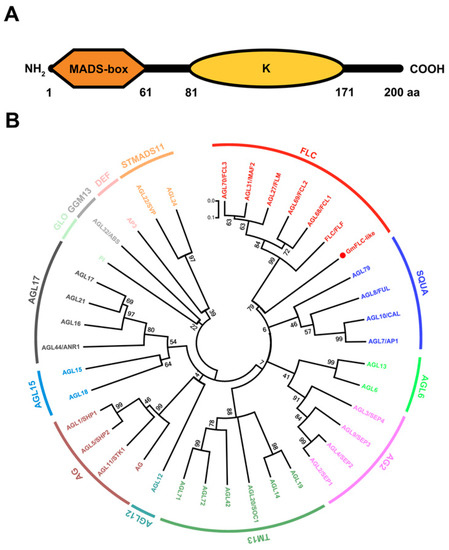
Figure 1.
Sequence analyses of GmFLC-like. (A) Structure of GmFLC-like conserved domains, MADS-box (1–61 aa) and K (keratin-like) domains (81–171 aa). (B) Phylogenetic analysis of the GmFLC-like and MIKC-type MADS-box proteins from Arabidopsis. The phylogenetic tree was constructed using the neighbor-joining method with 1000 bootstrap replicates by MEGA 7.0 software, and bootstrap values are shown at the nodes. The locus of Arabidopsis MADS-box proteins as follows: AG (AT4G18960), AGL1/SHP1 (AT3G58780), AGL2/SEP1 (AT5G15800), AGL3/SEP4 (AT2G03710), AGL4/SEP2 (AT2G21970), AGL5/SHP2 (AT2G42830), AGL6 (AT2G45650), AGL7/AP1 (AT1G69120), AGL8/FUL (AT5G60910), AGL9/SEP3 (AT1G24260), AGL10/CAL (AT1G26310), AGL11/STK1 (AT4G09960), AGL12 (AT1G71692), AGL13 (AT3G61120), AGL14 (AT4G11880), AGL15 (AT5G13790), AGL16 (AT3G57230), AGL17 (AT2G22630), AGL18 (AT3G57390), AGL19 (AT4G22950), AGL20/SOC1 (AT2G45660), AGL21 (AT4G37940), AGL22/SVP (AT2G22540), AGL24 (AT4G24540), AGL27/FLM (AT1G77080), AGL31/MAF2 (AT5G65050), AGL32/ABS (AT5G23260), AGL42 (AT5G62165), AGL44/ANR1 (AT2G14210), AGL68/FCL1 (AT5G65080), AGL69/FCL2 (AT5G65070), AGL70/FCL3 (AT5G65060), AGL71 (AT5G51870), AGL72 (AT5G51860), AGL79 (AT3G30260), PI (AT5G20240), P3 (AT3G54340), FLC/FLF (AT5G10140).
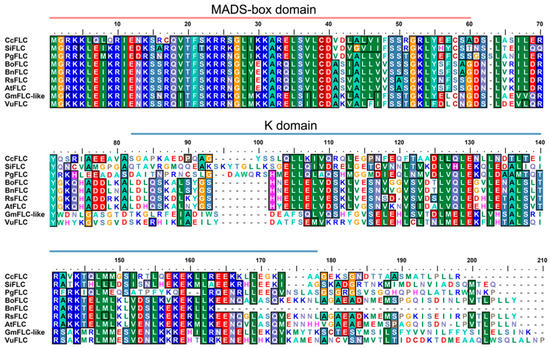
Figure 2.
Sequence alignment of the GmFLC-like protein and other FLC proteins from other species. The pink line marks the MADS-box domain (1–61 aa), and the light blue line labels the K domain (81–171 aa). GenBank accession numbers are as follows: Glycine max GmFLC-like (MK913903), Arabidopsis thaliana AtFLC (NP_196576), Vigna unguiculata VuFLC (XP_027918635), Punica granatum PgFLC (OWM76224v), Citrus clementina CcFLC (XP_024041198), Brassica oleracea BoFLC (AHH30724), Sesamum indicum SiFLC (XP_011086821), Raphanus sativus RsFLC (AJN00653), Brassica napus BnFLC (AFU61576).
2.2. Expression Profile and Biochemical Properties of GmFLC-like
In an attempt to understand whether GmFLC-like expression has tissue specificity, we analyzed GmFLC-like transcripts in multiple tissues, including shoot apexes, roots, stems, fully expanded leaves, flowers, and pods, during the soybean development process under SD conditions by qRT-PCR. The expression of GmFLC-like was greater in the leaves than in the flowers and pods and was lowest in the stems. In the shoot apex and root, the transcript of GmFLC-like was higher in the unifoliate period than in the other developmental stages (Figure 3).
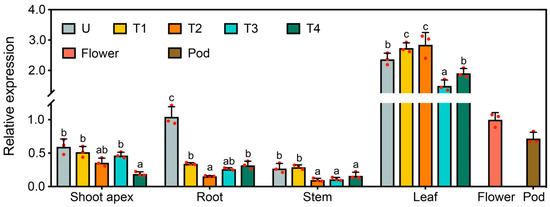
Figure 3.
Expression analysis of GmFLC-like in different organs of soybean during multiple developmental stages under short-day (SD) conditions. U, the unifoliate period; T1, the first trifoliate period; T2, the second trifoliate period; T3, the third trifoliate period; T4, the fourth trifoliate period; Shoot apex (including apical meristem and immature leaves); Pod (14 days after flowering). Gmβ-tubulin (Glyma20 g27280) was used as an internal control. Error bar represents the means of three biological replicates, and the letters indicate significant differences according to Duncan’s multiple range test (p < 0.05).
To determine the subcellular localization of GmFLC-like, we expressed a GmFLC-like-GFP protein together with a mCherry-labelled nuclear marker protein (NF-YA4-mCherry) in N. benthamiana leaves, and both proteins were driven by the 35S promoter. We observed that the GFP fluorescence signal pattern was consistent with the localization of the nuclear marker protein (mCherry) in the nucleus, indicating that GmFLC-like is a nuclear-localized protein (Figure 4). Similarly, transient expression of GmFLC-like in Arabidopsis protoplasts also confirmed that GmFLC-like is a nuclear protein (Figure S1).
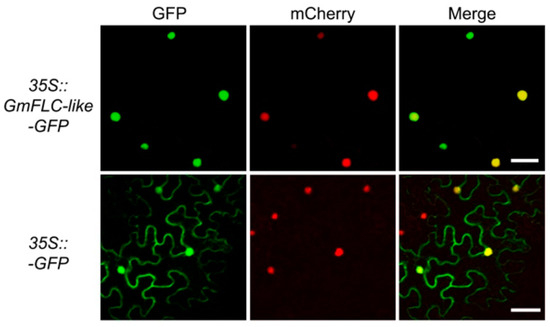
Figure 4.
Subcellular localization of GmFLC-like protein in tobacco leaves. GFP fused to the C-terminal region of GmFLC-like, and the fusion protein was driven by 35S promoter. A mCherry labeled fusion protein (NF-YA4-mCherry) was used as a nuclear marker driven by 35S, and 35S::GFP was used as negative control. At 3 days after infiltration, the fluorescence signals (GFP and mCherry) were visualized by confocal microscopy, and the excitation wavelengths for GFP and mCherry were 488 and 543 nm, respectively. Scale bar, 50 μM.
2.3. Overexpression of GmFLC-like Caused Late Flowering in Arabidopsis
To investigate the biological function of GmFLC-like in regulating flowering time, we overexpressed GmFLC-like in Arabidopsis (Col-0), and the transcript abundance of GmFLC-like in the transgenic lines was confirmed using qRT-PCR (Figure 5A). Compared with the WT plants, the plants overexpressing GmFLC-like (L46 and L48) exhibited a clear delayed-flowering phenotype (Figure 5B). At 30 days, flower buds emerged in WT plants, while flower bud emergence of L46 and L48 plants was observed at days 34 and 35, respectively (Figure 5C). In addition, we also evaluated the expression of key downstream genes that are linked to flowering time by qRT-PCR analysis. The results indicated that, compared with the levels in the WT, transcript levels of floral activators, FT, SOC1, and AP1, in transgenic Arabidopsis (L46 and L48) decreased significantly (Figure 5D). Additional evidence indicated that the transgenic lines (L1, L48, and L46) displayed lower germination rates than the WT plants (Figure S2). Overall, GmFLC-like offers a similar function to AtFLC, and both of them function as a floral repressor.
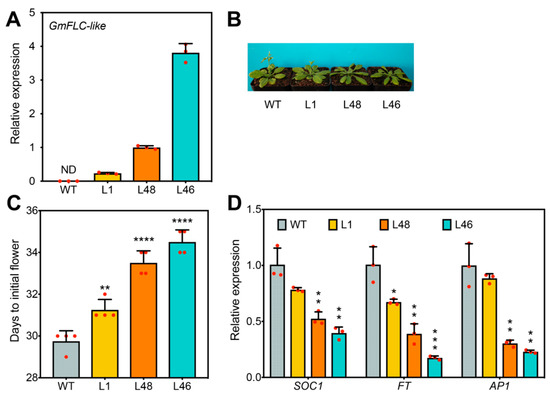
Figure 5.
Ectopic expression of GmFLC-like in Arabidopsis. (A) Expression detection of GmFLC-like in transgenic plants. ND, not detected. (B) Phenotypic display of transgenic (L1, L46, and L48) and WT (Col-0) plants. The flowering phenotype of the GmFLC-like gene overexpression lines and WT plants were photographed at 5 weeks after sowing. Plants were grown at a growth chamber at 22 ± 1 °C under long-day (LD) conditions (16 h of light/8 h of dark). (C) Statistical analysis of days until the initial flowering of transgenic and WT (Col-0) plants. (D) Expression levels of flowering time genes in Arabidopsis. Arabidopsis β-tubulin (AT5G62690) was used as a negative control. Error bar represents the means of three biological replicates. Significant differences according to the t-test are denoted as follows: * p < 0.05, ** p < 0.01, *** p < 0.001, **** p < 0.0001. WT means the wild-type of Arabidopsis; L1, L46, and L48 refer to independent transgenic lines.
2.4. GmFLC-like Is Responsive to the Photoperiod and Low Temperature
To further understand GmFLC-like potential functions, we analyzed the putative cis-acting elements in its promoter region (1500 bp sequence upstream of the start codon) by using the PlantCARE database. Different cis-acting regulatory elements involved in the light response, hormones, and development as well as abiotic stress were found (Table 1). Light response elements included AE-box, CATT-motif, G-box, TCT-motif, AT1-motif, Box-4, and so on. Hormone- and development-related elements included ABRE, GARE, TCA, and so on. HSE, MBS, ARE, and CE3, associated with abiotic stress responses, were also identified in the GmFLC-like upstream region. A variety of cis-acting regulatory elements in the GmFLC-like upstream region implies that the gene may be regulated by endogenous and external environmental signals.

Table 1.
cis-acting elements of the GmFLC-like promoter.
Based on many light-responsive cis-elements found in the GmFLC-like promoter region, we focused on the GmFLC-like function in response to the photoperiod (Table 1). Here, we chose the soybean variety “Huachun 5” as the research material, which is sensitive to the photoperiod because its flower buds emerge earlier under SD conditions [46]. Surprisingly, GmFLC-like functions as a floral inhibitor, but it displays a high transcript level under SD conditions. Further investigation concluded that GmFLC-like expression was markedly high under SD conditions at 15, 18 DAE (days after emergence), and showed the highest peak at 27 DAE, while GmFLC-like expression did not change outstandingly during the same period under the LD treatment (Figure 6). We suspect that the photoperiod pathway plays a leading role in the flowering regulation of soybean variety “Huachun 5”. Although the expression of GmFLC-like is upregulated under SD conditions, its relatively weak power cannot inhibit flowering accelerated by photoperiod.
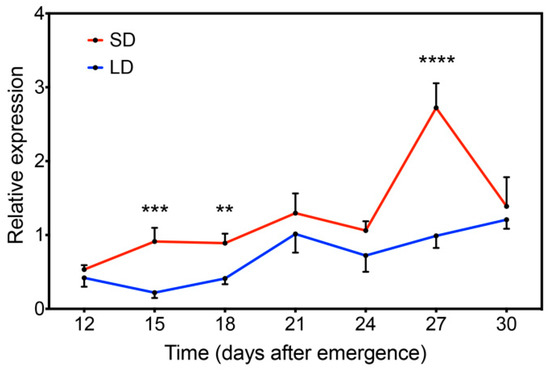
Figure 6.
Expression analysis of GmFLC-like under SD and LD conditions at 12, 15, 18, 21, 24, 27, and 30 DAE (days after emergence). All seedlings were grown under SD conditions for 10 DAE, and then a portion of the seedlings were transferred to LD conditions. Fully expanded trifoliate leaves were sampled at the appointed time from three individual plants growing under SD and LD conditions. Significant differences according to the t-test are denoted as follows: ** p < 0.01, *** p < 0.001, **** p < 0.0001.
In Arabidopsis, AtFLC functions as a floral repressor and is a key gene responding to low temperature [29,51,52]. Our previous research showed that long-term low-temperature treatment resulted in delayed flowering in soybean [46]. To investigate whether GmFLC-like is responsive to low temperature, we tested GmFLC-like expression under low-temperature treatment by qRT-PCR. With respect to a long-term low-temperature treatment, GmFLC-like expression was significantly higher in treated plants than in controls after treatment for 2, 4, 6, and 8 days. Surprisingly, compared with the untreated plants, GmFLC-like expression was decreased in the treated ones after 10 days treatment (Figure 7A).
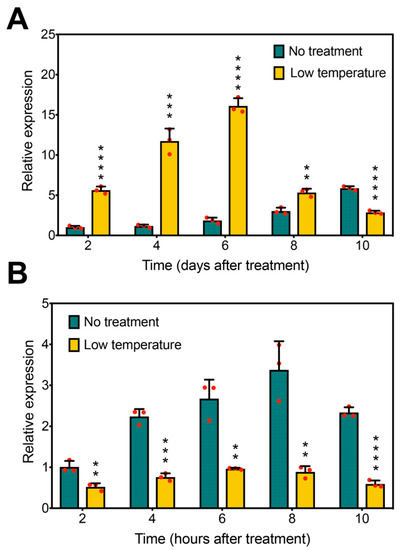
Figure 7.
GmFLC-like response to photoperiod and low temperature. (A) Expression analysis of GmFLC-like at 2, 4, 6, 8, and 10 days after the beginning of low-temperature treatments and the control. (B) Expression analysis of GmFLC-like at 2, 4, 6, 8, and 10 h after the beginning of low-temperature treatments and the control. Error bars represent the means of three biological replicates. Significant differences according to the t-test are denoted as follows: ** p < 0.01, *** p < 0.001, **** p < 0.0001.
In contrast, short-term low-temperature treatment does not trigger the phenotype of delayed flowering. GmFLC-like expression also showed a remarkable difference when subjected to short-term low-temperature treatment. Compared with the controls, the plants displayed significantly lower GmFLC-like expression when subjected to a short-term low-temperature treatment for 2, 4, 6, 8, and 10 h (Figure 7B). Together, these results suggest that the expression pattern of GmFLC-like is clearly different between long-term and short-term low-temperature treatments. In other words, GmFLC-like may play a role in long-term low temperature-triggered delayed flowering in soybean based on its expression being strongly upregulated by long-term low-temperature treatment and downregulated by short-term low-temperature treatment.
2.5. Identification of GmFT2a as a Downstream Target of GmFLC-like
In Arabidopsis, FLC is responsible for regulating floral activator FT expression. FLC suppresses flowering mainly by repressing the expression of these floral activators [25,26,27]. According to a previous study [53] and the newest information from the Phytozome database, we discovered nine FT homologues, including FT1a, FT1b, FT2a, FT2b, FT3a, FT3b, FT4, FT5a, and FT5b. The flowering-inhibiting genes FT1a and FT4 exhibited higher accumulation under LD, and represented opposite expression patterns of other FT genes [54]. To better understand whether GmFLC-like has a consistent function with AtFLC, we detected the expression profile of nine FT genes after the beginning of long-term low-temperature treatment for 8 days. QRT-PCR confirmed that, except for FT1b and FT5a, other FT homologues were downregulated after the beginning of long-term low-temperature treatment compared with the control (Figure 8A). In this study, we found that GmFLC-like expression was greatly elevated, and GmFT1a, GmFT2a, GmFT2b, and GmFT2a expression was significantly decreased after the beginning of the long-term low-temperature treatment. Previous research revealed that GmFT2a is responsible for inducing flowering under SD conditions, and the expression profiles of both GmFLC-like and GmFT2a were matched with the phenotype of delayed flowering (Figure 7A and Figure 8A). Next, we selected GmFT2a as a candidate gene for in-depth analysis to verify whether GmFT2a is a potential downstream target gene of GmFLC-like.
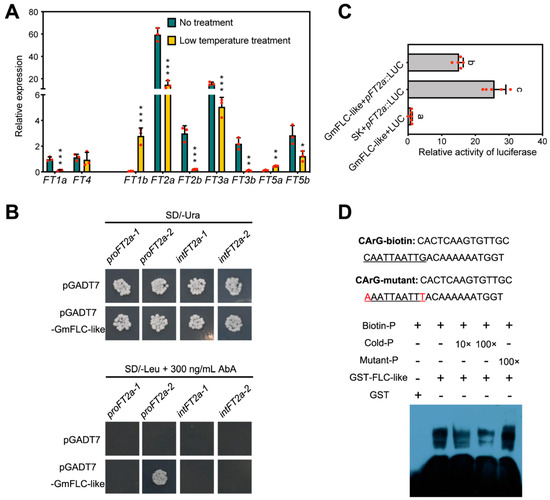
Figure 8.
GmFLC-like protein binds to the promoter region of FT2a. (A) Expression analysis of GmFTs genes in soybean after the beginning of low-temperature treatments at 8 DAE. The soybean accession numbers are as follows: GmFT1a (Glyma18g53680), GmFT4 (Glyma08g47810), GmFT1b (Glyma18g53690), GmFT2a (Glyma16g26660), GmFT2b (Glyma16g26690), GmFT3a (Glyma16g04840), GmFT3b (Glyma19g28390), GmFT5a (Glyma16g04830), and GmFT5b (Glyma19g28400). Gmβ-tubulin (Glyma20g27280) was used as an internal control. The mean values ± SD from three biological replicates are shown. Significant differences according to the t-test are denoted as follows: ** p < 0.05, ** p < 0.01, *** p < 0.001. (B) Interaction of GmFLC-like protein and CmFT2a promoter and the intron region, as revealed using a yeast one-hybrid system. The yeast transformations were plated onto SD/-Ura (upper panel) and SD/-Leu containing 300 ng/mL AbA (lower panel). pGADT7 with pAbAi-proFT2a-1 (from −1275 to −1156 bp), pAbAi-proFT2a-2 (from −671 to −552 bp), pAbAi-intFT2a-1 (from 410 to 551 bp), and pAbAi-intFT2a-2 (from 654 to 841 bp), were used as negative controls. The experiment was performed independently three times. (C) Relative reporter activity (LUC/REN) in N. benthamiana leaves. the relative luciferase activity (LUC/REN) in tobacco leaves were measured after 48 h of Agrobacterium infiltration. Experiments were repeated five times and mean value ± SD is plotted on the graph. The letters indicate significant differences according to Duncan’s multiple range test (p < 0.05). (D) Gel-shift analysis of GmFLC-like binding to the promoter region of GmFT2a. The sequence fragment from −663 to −628 of the GmFT2a promoters was used as a probe, and the core sequences are underlined. Purified protein (3 µg) was incubated with 25 picomoles biotin-labeled probe. For competition test, non-labeled probes at varying concentrations (from 10- to 100-fold excess), and mutated unlabeled CArG probe were added to the above experiment.
To better understand the regulatory mechanism between GmFLC-like and GmFT2a, a 1385-bp promoter region and a 792-bp first intron region of the GmFT2a sequence was identified. Sequence analysis using the New PLACE database (https://www.dna.affrc.go.jp/PLACE/?action=newplace) found that six CArG motifs (CWWWWWWWWG) exist in these sequences. Based on these findings, we designed the following experiments. First of all, we performed Y1H. The promoter sequence (−1275 to −1156 bp; −671 to −552 bp) and the first intron region (410 to 551 bp; 654 to 841 bp) of GmFT2a were amplified and inserted into the pAbAi vector, and the ORF sequence of GmFLC-like was inserted into the pGADT7 plasmid. The yeast one-hybrid (Y1H) assay confirmed that the GmFLC-like protein could target the promoter region (−671 to −552 bp) instead of the first intron region of GmFT2a by observing the transformants growth on SD/-Leu supplemented with 300 ng/mL AbA (Figure 8B). Additional evidence from a dual-luciferase reporter assay in N. benthamiana revealed that the activity of the promoters of GmFT2a could be inhibited by overexpression of GmFLC-like driven by the CaMV 35S promoter (Figure 8C). EMSA was also performed to verify the binding of GmFLC-like to the GmFT2a promoter. The 36 bp sequence fragment spanning positions −663 to −628 of the GmFT2a promoter was used as probe. The probe contained the predicted CArG motif (CAATTAATTG), which is the binding site for plant MADS domain protein. The fusion protein GST-GmFLC-like was purified from Escherichia coli, and then co-incubated with biotin-labeled and non-labeled probes. Finally, GmFLC-like was found to bind to the biotin-labeled proGmFT2a probe, furthermore, binding capacity was slowly weakened by increasing concentrations of non-labeled probe, while it was not affected by mutated unlabeled GCC probe (Figure 8D). These results indicated that GmFT2a was potentially one of the direct targets of GmFLC-like.
3. Discussion
For plants, temperature is a main environmental cue that has a strong influence on flowering time through different pathways, which mainly including vernalization and ambient temperature pathways [1,4,55]. In many species, FLC, encoding a MADS-box transcription factor, is the core gene in the vernalization way, which prevents flowering through inhibiting several floral activators, including FT and SOC1 [13]. In Brassica plants, FLCs have been well studied over many years [56,57,58]. The researchers found that vernalization is closely correlated with FLCs epigenetic silencing, such as antisense RNA COOLAIR. The transcripts of COOLAIR are polyadenylated at multiple sites with proximal polyadenylation promoted by components of the autonomous promotion pathway. Use of the proximal poly(A) site results in quantitative downregulation of FLC expression in a process requiring FLD, an H3K4me2 demethylase [59]. Furthermore, the functions of FLC homologues show common features among different species [31,34]. However, no functional FLC homologues have been characterized in crops thus far [60]; one possible reason for this is that some crops have lost their FLC homologues. For example, FLC homologues may be absent from the rice genome [61]. For winter cereals, a gene called VRN2, which is downregulated in vernalization and functions in inhibiting FT expression [62], plays a similar role instead of genes orthologous to FLC.
Soybean is considered a typical short-day plant that does not require vernalization. Interestingly, it still retains only one homologue of FLC, Glyma05g28130by comparative genomic analysis [63,64]. In our previous study, we found that Glyma05g28130 were strongly upregulated in delayed-flowering AtDREB1A-overexpressing transgenic soybean [45]. Hence, we speculate that Glyma05g28130 is involved in flowering time regulation. In the paper, we found Glyma05g28130 belongs to a member of the MADS-box family transcription factor, containing two conserved domains, the MADS-box domain and the K domain (Figure 1A). Phylogenetic analysis revealed that Glyma05g28130 falls within the FLC clade and is closely related to AtFLC in Arabidopsis, so named GmFLC-like (Figure 1B). In addition, the MADS-box domain between the GmFLC-like and FLC proteins from other species shared high conservation (Figure 2). These cues imply that GmFLC-like may also be a floral repressor and responsible for directly targeting and repressing the expression of floral activators FT, showing similar functions with Arabidopsis [25,26,27]. Then, heterogeneous overexpression of this gene in Arabidopsis indeed resulted in a marked late-flowering phenotype compared with the WT (Figure 5B,C). In addition, the expression of SOC1, AP1, and FT was significantly decreased in transgenic Arabidopsis (L48 and L46) (Figure 5D). Previous studies also confirmed that AtFLC homologues, such as BvFL1 and CiFL1, perform the same functions in different species [23,29,30,34,65,66]. In summary, these findings suggest that some biological functions of AtFLC homologues are conserved among different species.
For vernalization-requiring plants, the transcripts of FLC are very high before vernalization but are subsequently inhibited by epigenetic modification under cold conditions and further accelerate flowering. While for soybean, a non-vernalized plant, shows the contrary phenomena. We found that the popular variety “Huachun 5”, which is suitable for growing in south china, shows sensitivity to low temperature. Under the long-term low-temperature treatment, GmFLC-like expression was induced and attributed to the delayed-flowering phenotype (Figure 7A), As we know, the soybean originated from north china, where the temperature is low. Therefore, we speculate that the soybean genome retains the low-temperature sensitive genes in order to cope with disadvantageous environments. Afterwards, to enlarge the planting range of soybean, the breeders selected out the varieties which are available to grow in early spring of south china. Meanwhile, the low-temperature-responsive gene FLC may be retained under the selection process for fitting the relative low temperatures.
In Arabidopsis, FLC delays the flowering time by repressing the expression of the floral integrator gene FT [12,13,25,67]. In this study, we found that GmFLC-like expression was greatly increased, meanwhile the GmFT2a expression was significantly decreased under the long-term low-temperature treatment (Figure 7A and Figure 8A), the results imply that the FLC-FT model may exist in soybean as well. Afterwards, we confirm that the GmFLC-like protein binds to the promoter region of GmFT2a promoter in vivo and vitro (Figure 8B−D). The FLC-FT2a model is confirmed, revealing that the molecular mechanism of flowering time is conversed among different species once again. Besides the existing variability in the FT2a-FLC model, these results imply that GmFLC-like may play an important role in delaying flowering time and providing protective mechanisms against sporadic and extremely low temperatures.
4. Materials and Methods
4.1. Plant Materials and Growth Conditions
The soybean cultivar “Huachun 5”, which was bred by the Guangdong Sub-center of the National Center for Soybean Improvement, was used as the experimental material. Mature soybean seeds were surface sterilized and germinated in vermiculite. Uniform soybean seedlings were selected and grown in plastic pots containing turf soil and vermiculite at a ratio of 3:1 (v/v) in a growth chamber at 27 ± 2 °C, 40% relative humidity, and 100 μmol m−2 s−1 illumination with fluorescent lamps. Different regimes of day length as following: SD conditions (8 h of light/16 h of dark) and LD conditions (16 h of light/8 h of dark). The Arabidopsis wild-type (WT) and transgenic plants were of the Columbia (Col-0) ecotype. Arabidopsis seeds were surface sterilized and sown on half-strength MS medium for 2 days at 4 °C to relieve dormancy. Subsequently, the plates were transferred to a growth chamber at 22 ± 1 °C under LD conditions (16 h of light/8 h of dark) for 7 days, and then seedlings were transferred to pots containing turf soil and vermiculite (v/v 3:1).
4.2. Total RNA Isolation and qRT-PCR Analysis
The total RNA of the second trifoliate soybean leaf was isolated using TRIzol reagent (Invitrogen, Carlsbad, CA, USA). Then, 1 μg of total RNA was reverse-transcribed using HiScript II Q RT SuperMix for qPCR (R233-01, Vazyme Biotech Co., Nanjing, China). QRT-PCR analysis was carried out on a StepOne Real-Time PCR System (Applied Biosystems, Foster, CA, USA) using a Kapa SYBR Fast Universal qPCR Kit (Kapa Biosystems, Boston, MA, USA). Soybean β-tubulin (Glyma20g27280) and Arabidopsis TUB2 (AT5G62690) were used as internal controls. The experiments were performed in three biological replicates, and the data were evaluated by the 2−ΔΔCt methods [68]. All primers used in this study are listed in Table S1.
4.3. Plasmid Construction
The open reading frame (ORF) of GmFLC-like was amplified using specific primers K-FLC-F and K-FLC-R and inserted into the pCAMBIA1301 plasmid by the BamHI and KpnI restriction sites to form the expression vector 35S::GmFLC-like-GFP. Genomic DNA was isolated from leaf samples of soybean using the modified cetyltrimethyl ammonium bromide (CTAB) method [69]. The promoter sequence of GmFLC-like was amplified using specific primers designed as Pro-GmFLC-F and Pro-GmFLC-R, and genomic DNA of soybean was used as the template. The amplified PCR product was inserted into a pZeroBack/blunt vector (Tiangen Biotech Co., Beijing, China) for sequencing. All primers used are listed in Supplementary Table S1.
4.4. Sequence Analysis
The conserved domains of GmFLC-like were predicted by using InterProScan [70]. The phylogenetic tree was generated based on alignment results using the neighbor-joining algorithm with 1000 bootstrap replicates in MEGA 5.0 software [71]. Multiple amino acid sequences were aligned using ClustalW with default parameters, and the comparison result was displayed by the software BioEdit [71]. Protein sequence logos were created using WebLogo 3.3 [72]. Homologous protein sequences of GmFLC-like were searched in the Phytozome databases [47]. The cis-acting elements in the GmFLC-like promoter (1000 bp upstream of the start codon) were analyzed using the PlantCARE program [73].
4.5. Subcellular Localization of GmFLC-like Protein
The ORF sequence of GmFLC-like without the stop codon was fused to the N-terminal region of the GFP protein, and the resulting fragment was inserted into the pCAMBIA1301 plasmid to form the expression vector 35S::GmFLC-like-GFP. The fused vector was transformed into epidermal cells of Nicotiana benthamiana leaves by Agrobacterium infection and expressed for 3 days. The fluorescence signals were monitored by confocal microscope (Olympus FluoView FV1000). The GFP and mCherry protein were imaged using 488 and 543 nm excitation, respectively. The plasmids 35S::GFP and 35S::NF-YA4-mCherry were used as the negative control and nuclear marker, respectively [74].
4.6. Ectopic Expression of GmFLC-like in Arabidopsis
The expression vector 35S::GmFLC-like-GFP was transformed into the Agrobacterium tumefaciens GV3101 strain by electroporation. Arabidopsis (Col-0) was used for transgenic material and performed using the floral dip method [75]. Transgenic Arabidopsis seeds were grown on half-strength MS medium supplemented with 25 mg/L hygromycin, and three homozygous T2 transgenic lines with different GmFLC-like expression levels were chosen for further studies, including phenotype investigation and expression analyses of potential downstream genes.
4.7. Photoperiod Treatment
Soybean seeds were grown under SD conditions for 10 days, and then, part of uniform seedlings was transferred to LD conditions. Fully expanded trifoliate leaves were sampled at 12, 15, 18, 21, 24, 27, and 30 DAE from plants growing under SD and LD conditions, respectively. All samples were immediately frozen in liquid nitrogen and stored at −80 °C for further study.
4.8. Low-Temperature Treatment
Uniform soybean seedlings were grown in a growth chamber at 28 °C/26 °C (day/night) under SD conditions until the fourth trifoliate stage. Then, the seedlings were divided into two groups for the following low-temperature treatments: the long-term treatment group and the short-term treatment group. For the long-term treatment group, plants were continuously treated at 15 °C/13 °C (day/night) under SD conditions for 10 days and leaves were sampled every 2 days from three individual plants. For the short-term treatment group, plants were continuously treated at 15 °C/13 °C (day/night) for 10 h in the darkness, and leaves were sampled every 2 h from three individual plants. The control seedlings were grown at 28 °C/26 °C under the same conditions corresponding to the long- and short-term treatment groups. Leaves were sampled from three individual control plants every 2 days at the same collection time used for the low temperature-treated plants.
4.9. Yeast One-Hybrid Assay
A yeast one-hybrid assay (Y1H) assay was performed as previously described [76,77]. The promoter and the first intron sequence of GmFT2a were amplified and inserted into the pAbAi vector to form the bait plasmid, and the obtained bait plasmids were recognized as pAbAi-proFT2a-1, pAbAi-proFT2a-2, pAbAi-intFT2a-1, pAbAi-intFT2a-2, respectively. After linearization by BstBI, the bait plasmids were transformed into yeast strains Y1H Gold according to the manufacturer’s instruction of the MatchmakerTM Gold Yeast One-Hybrid Library Screening System Kit (Clontech, Mountain View, CA, USA). The GmFLC-like ORF sequence was inserted into the pGADT7 vector to form the prey plasmid. The prey plasmid was co-transformed into yeast strain Y1Hgold. Subsequently, the yeast transformants were plated onto synthetic dropout (SD) medium lacking uracil (Ura) or leucine (Leu) but supplemented with 300 ng/mL Aureobasidin A (AbA). After 3 days of incubation in a 28 °C incubator, the interactors were defined on the basis of transformants growth on SD/-Leu medium with 300 ng/mL AbA.
4.10. Dual-Luciferase Reporter Assay
For dual-luciferase assay, we introduced two vectors: pGreenII 0800-LUC and pGreenII 0029 62-SK. The GmFT2a promoter was amplified and inserted into the pGreenII 0800-LUC vector to drive the LUC reporter gene expression, and the GmFLC-like ORF sequence was inserted into the pGreenII 0029 62-SK vector. The pGreenII 0800-LUC vector carries a renilla luciferase (REN) gene driven by the 35S promoter to serve as an internal control. The two plasmids were co-expressed into tobacco leaves by Agrobacterium infiltration as previously described [72]. At 48 h after transformation, LUC and REN luciferase activities were measured using a Dual-Luciferase® Reporter Assay System (Promega, Madison, WI, USA) and a Glomax®-20/20 signal Tube Luminometer (Promega, Madison, WI, USA). The relative activity of luciferase was calculated based on the ratio of the luciferase activity of the sample to that of the Renilla luciferase control.
4.11. Electrophoretic Mobility Shift Assay
Briefly, the E. coli BL21 cells carrying the GST-GmFLC-like fusion protein was incubated in 100 mL LB medium supplemented with isopropyl IPTG to achieve the rated concentration (0.2 mM), and the cell cultures were incubated at 30 °C for 6 h. The fusion protein was extracted from E. coli cells and purified according to the manufacturer’s instructions of glutathione beads (Glutathione-Sepharose 4B, GE Healthcare, Chicago, IL, USA). EMSA was performed according to the description of the LightShift® Chemiluminescent EMSA Kits (Pierce, Milan, Italy). The biotin-labeled DNA labeled fragments (5′-CACTCAAGTGTTGCCAATTAATTGACAAAAAATGGT-3′) were synthesized, annealed, and used as probes, with unlabeled DNA of the same sequence and mutant sequence (5′-CACTCAAGTGTTGCAAATTAATTTACAAAAAATGGT-3′) used as the competitors.
4.12. Data Analysis
For data analysis and graphs generation, GraphPad Prism 8.0 (GraphPad Software Inc., San Diego, CA, USA) was used. For comparison of two groups of data, the two-sided Student’s t-test was used, thereinto, asterisks *, **, ***, and **** indicate significant differences at p < 0.05, p < 0.01, p < 0.001, and p < 0.0001, respectively. For comparison of multiple groups of data, one-way ANOVA and Bonferroni’s post hoc test was used, and p < 0.05 was considered significant.
4.13. Data Availability
The GenBank accession numbers for soybean genes used in this study are as follows: GmFLC-like (MK913903); Promoter of GmFLC-like (KY203812).
Supplementary Materials
Supplementary materials can be found at https://www.mdpi.com/1422-0067/21/4/1322/s1. Table S1. Primers used in this study; Figure S1. Subcellular localization of GmFLC-like protein in Arabidopsis protoplast cells.; Figure S2. Comparative analysis of seed germination rate among four lines of Arabidopsis.
Author Contributions
J.L. and Z.C. performed the experiments. H.N. and S.Z. designed the research. H.S., R.Y. and Y.L. provided technical support, conceptual advice, and data analysis. J.L. and S.Z. wrote the article. All authors have read and agreed to the published version of the manuscript.
Funding
We thank AJE (https://www.aje.cn/) for the careful editing of this article. This work was supported by the Projects of Science and Technology of Guangzhou (201804020015), the China Agricultural Research System (CARS-04-PS09), the Major Projects of New Varieties Cultivation of Genetically Modified Organisms (2014ZX08004-002), and the National Natural Science Foundation of China (31902054).
Conflicts of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Abbreviations
| LD | long-day |
| SD | short-day |
| FLC | flowering locus C |
| Y1H | yeast one-hybrid |
| EMSA | electrophoretic mobility shift assay |
| FT | flowering locus T |
| FRI | frigida |
| VIN3 | vernalization insensitive 3 |
| COOLAIR | cold induced long antisense intragenic RNA |
| COLDAIR | cold assisted intronic noncolding RNA |
| SOC1 | suppressor of overexpression of constants 1 |
| VRN1 | vernalization 1 |
| VRN2 | vernalization 2 |
| AP1 | apetala 1 |
| MS | murashige and skoog |
| TUB2 | tubulin beta chain 2 |
| AbA | aureobasidin A |
| IPTG | β-d-1-thiogalactopyranoside |
References
- Srikanth, A.; Schmid, M. Regulation of flowering time: All roads lead to Rome. Cell. Mol. Life Sci. 2011, 68, 2013–2037. [Google Scholar] [CrossRef]
- Wei, Q.; Ma, C.; Xu, Y.; Wang, T.; Chen, Y.; Lü, J.; Zhang, L.; Jiang, C.-Z.; Hong, B.; Gao, J. Control of chrysanthemum flowering through integration with an aging pathway. Nat. Commun. 2017, 8, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Willmann, M.R.; Poethig, R.S. Time to grow up: The temporal role of smallRNAs in plants. Curr. Opin. Plant Biol. 2005, 8, 548–552. [Google Scholar] [CrossRef] [PubMed]
- Bäurle, I.; Dean, C. The Timing of Developmental Transitions in Plants. Cell 2006, 125, 655–664. [Google Scholar] [CrossRef] [PubMed]
- Feng, P.; Guo, H.; Chi, W.; Chai, X.; Sun, X.; Xu, X.; Ma, J.; Rochaix, J.-D.; Leister, D.; Wang, H.; et al. Chloroplast retrograde signal regulates flowering. Proc. Natl. Acad. Sci. USA 2016, 113, 10708–10713. [Google Scholar] [CrossRef] [PubMed]
- Izawa, T. Daylength Measurements by Rice Plants in Photoperiodic Short-Day Flowering. In International Review of Cytology; Academic Press: Cambridge, MA, USA, 2007; Volume 256, pp. 191–222. [Google Scholar]
- Cerdán, P.D.; Chory, J. Regulation of flowering time by light quality. Nature 2003, 423, 881. [Google Scholar] [CrossRef]
- Capovilla, G.; Schmid, M.; Posé, D. Control of flowering by ambient temperature. J. Exp. Bot. 2015, 66, 59–69. [Google Scholar] [CrossRef]
- Amasino, R. Vernalization, Competence, and the Epigenetic Memory of Winter. Plant Cell 2004, 16, 2553–2559. [Google Scholar] [CrossRef]
- Schmitz, R.J.; Amasino, R.M. Vernalization: A model for investigating epigenetics and eukaryotic gene regulation in plants. Biochim. Biophys. Acta (BBA)-Gene Struct. Exp. 2007, 1769, 269–275. [Google Scholar] [CrossRef]
- Samach, A.; Wigge, P.A. Ambient temperature perception in plants. Curr. Opin. Plant Biol. 2005, 8, 483–486. [Google Scholar] [CrossRef]
- Michaels, S.D.; Amasino, R.M. FLOWERING LOCUS C Encodes a Novel MADS Domain Protein That Acts as a Repressor of Flowering. Plant Cell 1999, 11, 949–956. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.; Suh, S.-S.; Park, E.; Cho, E.; Ahn, J.H.; Kim, S.-G.; Lee, J.S.; Kwon, Y.M.; Lee, I. The AGAMOUS-LIKE 20 MADS domain protein integrates floral inductive pathways in Arabidopsis. Genes Dev. 2000, 14, 2366–2376. [Google Scholar] [CrossRef] [PubMed]
- Johanson, U.; West, J.; Lister, C.; Michaels, S.; Amasino, R.; Dean, C. Molecular Analysis of FRIGIDA, a Major Determinant of Natural Variation in Arabidopsis Flowering Time. Science 2000, 290, 344–347. [Google Scholar] [CrossRef] [PubMed]
- Le Corre, V.; Roux, F.; Reboud, X. DNA Polymorphism at the FRIGIDA Gene in Arabidopsis thaliana: Extensive Nonsynonymous Variation Is Consistent with Local Selection for Flowering Time. Mol. Biol. Evol. 2002, 19, 1261–1271. [Google Scholar] [CrossRef]
- Gazzani, S.; Gendall, A.R.; Lister, C.; Dean, C. Analysis of the Molecular Basis of Flowering Time Variation in Arabidopsis Accessions. Plant Physiol. 2003, 132, 1107–1114. [Google Scholar] [CrossRef]
- Shindo, C.; Aranzana, M.J.; Lister, C.; Baxter, C.; Nicholls, C.; Nordborg, M.; Dean, C. Role of FRIGIDA and FLOWERING LOCUS C in Determining Variation in Flowering Time of Arabidopsis. Plant Physiol. 2005, 138, 1163–1173. [Google Scholar] [CrossRef]
- Sung, S.; Amasino, R.M. Vernalization in Arabidopsis thaliana is mediated by the PHD finger protein VIN3. Nature 2004, 427, 159. [Google Scholar] [CrossRef]
- Kim, D.-H.; Sung, S. Coordination of the Vernalization Response through a VIN3 and FLC Gene Family Regulatory Network in Arabidopsis. Plant Cell 2013, 25, 454–469. [Google Scholar] [CrossRef]
- Kim, D.-H.; Doyle, M.R.; Sung, S.; Amasino, R.M. Vernalization: Winter and the Timing of Flowering in Plants. Annu. Rev. Cell Dev. Biol. 2009, 25, 277–299. [Google Scholar] [CrossRef]
- Swiezewski, S.; Liu, F.; Magusin, A.; Dean, C. Cold-induced silencing by long antisense transcripts of an Arabidopsis Polycomb target. Nature 2009, 462, 799–802. [Google Scholar] [CrossRef]
- Heo, J.B.; Sung, S. Vernalization-Mediated Epigenetic Silencing by a Long Intronic Noncoding RNA. Science 2011, 331, 76–79. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.-H.; Xi, Y.; Sung, S. Modular function of long noncoding RNA, COLDAIR, in the vernalization response. PLoS Genet. 2017, 13, e1006939. [Google Scholar] [CrossRef] [PubMed]
- Chen, M.; Penfield, S. Feedback regulation of COOLAIR expression controls seed dormancy and flowering time. Science 2018, 360, 1014–1017. [Google Scholar] [CrossRef] [PubMed]
- Hepworth, S.R.; Valverde, F.; Ravenscroft, D.; Mouradov, A.; Coupland, G. Antagonistic regulation of flowering-time gene SOC1 by CONSTANS and FLC via separate promoter motifs. EMBO J. 2002, 21, 4327–4337. [Google Scholar] [CrossRef]
- Helliwell, C.A.; Wood, C.C.; Robertson, M.; Peacock, W.J.; Dennis, E.S. The Arabidopsis FLC protein interacts directly in vivo with SOC1 and FT chromatin and is part of a high-molecular-weight protein complex. Plant J. 2006, 46, 183–192. [Google Scholar] [CrossRef]
- Searle, I.; He, Y.; Turck, F.; Vincent, C.; Fornara, F.; Kröber, S.; Amasino, R.A.; Coupland, G. The transcription factor FLC confers a flowering response to vernalization by repressing meristem competence and systemic signaling in Arabidopsis. Genes Dev. 2006, 20, 898–912. [Google Scholar] [CrossRef]
- Lai, Z.; Schluttenhofer, C.M.; Bhide, K.; Shreve, J.; Thimmapuram, J.; Lee, S.Y.; Yun, D.-J.; Mengiste, T. MED18 interaction with distinct transcription factors regulates multiple plant functions. Nat. Commun. 2014, 5, 3064. [Google Scholar] [CrossRef]
- Tadege, M.; Sheldon, C.C.; Helliwell, C.A.; Stoutjesdijk, P.; Dennis, E.S.; Peacock, W.J. Control of flowering time by FLC orthologues in Brassica napus. Plant J. 2001, 28, 545–553. [Google Scholar] [CrossRef]
- Schranz, M.E.; Quijada, P.; Sung, S.-B.; Lukens, L.; Amasino, R.; Osborn, T.C. Characterization and Effects of the Replicated Flowering Time Gene FLC in Brassica rapa. Genetics 2002, 162, 1457–1468. [Google Scholar]
- Reeves, P.A.; He, Y.; Schmitz, R.J.; Amasino, R.M.; Panella, L.W.; Richards, C.M. Evolutionary Conservation of the FLOWERING LOCUS C-Mediated Vernalization Response: Evidence from the Sugar Beet (Beta vulgaris). Genetics 2007, 176, 295–307. [Google Scholar] [CrossRef]
- D’Aloia, M.; Tocquin, P.; Périlleux, C. Vernalization-induced repression of FLOWERING LOCUS C stimulates flowering in Sinapis alba and enhances plant responsiveness to photoperiod. New Phytol. 2008, 178, 755–765. [Google Scholar] [CrossRef] [PubMed]
- Guo, Y.-L.; Todesco, M.; Hagmann, J.; Das, S.; Weigel, D. Independent FLC Mutations as Causes of Flowering-Time Variation in Arabidopsis thaliana and Capsella rubella. Genetics 2012, 192, 729–739. [Google Scholar] [CrossRef] [PubMed]
- Périlleux, C.; Pieltain, A.; Jacquemin, G.; Bouché, F.; Detry, N.; D’Aloia, M.; Thiry, L.; Aljochim, P.; Delansnay, M.; Mathieu, A.-S.; et al. A root chicory MADS box sequence and the Arabidopsis flowering repressor FLC share common features that suggest conserved function in vernalization and de-vernalization responses. Plant J. 2013, 75, 390–402. [Google Scholar] [CrossRef] [PubMed]
- Ream, T.S.; Woods, D.P.; Amasino, R.M. The Molecular Basis of Vernalization in Different Plant Groups. Cold Spring Harb. Symp. Quant. Biol. 2012, 77, 105–115. [Google Scholar] [CrossRef]
- Balasubramanian, S.; Sureshkumar, S.; Lempe, J.; Weigel, D. Potent Induction of Arabidopsis thaliana Flowering by Elevated Growth Temperature. PLoS Genet. 2006, 2, e106. [Google Scholar] [CrossRef] [PubMed]
- Anderson, J.T.; Lee, C.-R.; Mitchell-Olds, T. LIFE-HISTORY QTLS AND NATURAL SELECTION ON FLOWERING TIME IN BOECHERA STRICTA, A PERENNIAL RELATIVE OF ARABIDOPSIS. Evolution 2011, 65, 771–787. [Google Scholar] [CrossRef] [PubMed]
- Nakano, Y.; Higuchi, Y.; Sumitomo, K.; Hisamatsu, T. Flowering retardation by high temperature in chrysanthemums: Involvement of FLOWERING LOCUS T-like 3 gene repression. J. Exp. Bot. 2013, 64, 909–920. [Google Scholar] [CrossRef]
- Liu, T.; Wu, P.; Wang, Q.; Wang, W.; Zhang, C.; Sun, F.; Liu, Z.; Li, Y.; Hou, X. Comparative transcriptome discovery and elucidation of the mechanism of long noncoding RNAs during vernalization in Brassica rapa. Plant Growth Regul. 2018, 85, 27–39. [Google Scholar] [CrossRef]
- Jiang, B.; Yue, Y.; Gao, Y.; Ma, L.; Sun, S.; Wu, C.; Hou, W.; Lam, H.-M.; Han, T. GmFT2a Polymorphism and Maturity Diversity in Soybeans. PLoS ONE 2013, 8, e77474. [Google Scholar] [CrossRef]
- Nan, H.; Cao, D.; Zhang, D.; Li, Y.; Lu, S.; Tang, L.; Yuan, X.; Liu, B.; Kong, F. GmFT2a and GmFT5a Redundantly and Differentially Regulate Flowering through Interaction with and Upregulation of the bZIP Transcription Factor GmFDL19 in Soybean. PLoS ONE 2014, 9, e97669. [Google Scholar] [CrossRef]
- Wu, F.; Price, B.W.; Haider, W.; Seufferheld, G.; Nelson, R.; Hanzawa, Y. Functional and Evolutionary Characterization of the CONSTANS Gene Family in Short-Day Photoperiodic Flowering in Soybean. PLoS ONE 2014, 9, e85754. [Google Scholar] [CrossRef] [PubMed]
- Zhai, H.; Lü, S.; Liang, S.; Wu, H.; Zhang, X.; Liu, B.; Kong, F.; Yuan, X.; Li, J.; Xia, Z. GmFT4, a Homolog of FLOWERING LOCUS T, Is Positively Regulated by E1 and Functions as a Flowering Repressor in Soybean. PLoS ONE 2014, 9, e89030. [Google Scholar] [CrossRef] [PubMed]
- Suo, H.; Ma, Q.; Ye, K.; Yang, C.; Tang, Y.; Hao, J.; Zhang, Z.J.; Chen, M.; Feng, Y.; Nian, H. Overexpression of AtDREB1A Causes a Severe Dwarf Phenotype by Decreasing Endogenous Gibberellin Levels in Soybean [Glycine max (L.) Merr.]. PLoS ONE 2012, 7, e45568. [Google Scholar] [CrossRef] [PubMed]
- Suo, H.; Lü, J.; Ma, Q.; Yang, C.; Zhang, X.; Meng, X.; Huang, S.; Nian, H. The AtDREB1A transcription factor up-regulates expression of a vernalization pathway gene, GmVRN1-like, delaying flowering in soybean. Acta Physiol. Plant. 2016, 38, 137. [Google Scholar] [CrossRef][Green Version]
- Lü, J.; Suo, H.; Yi, R.; Ma, Q.; Nian, H. Glyma11g13220, a homolog of the vernalization pathway gene VERNALIZATION 1 from soybean [Glycine max (L.) Merr.], promotes flowering in Arabidopsis thaliana. BMC Plant Biol. 2015, 15, 232. [Google Scholar]
- Goodstein, D.M.; Shu, S.; Howson, R.; Neupane, R.; Hayes, R.D.; Fazo, J.; Mitros, T.; Dirks, W.; Hellsten, U.; Putnam, N.; et al. Phytozome: A comparative platform for green plant genomics. Nucleic Acids Res. 2012, 40, D1178–D1186. [Google Scholar] [CrossRef] [PubMed]
- Alvarez-Buylla, E.R.; Liljegren, S.J.; Pelaz, S.; Gold, S.E.; Burgeff, C.; Ditta, G.S.; Vergara-Silva, F.; Yanofsky, M.F. MADS-box gene evolution beyond flowers: Expression in pollen, endosperm, guard cells, roots and trichomes. Plant J. 2000, 24, 457–466. [Google Scholar] [CrossRef]
- Becker, A.; Winter, K.-U.; Meyer, B.; Saedler, H.; Theißen, G. MADS-Box Gene Diversity in Seed Plants 300 Million Years Ago. Mol. Biol. Evol. 2000, 17, 1425–1434. [Google Scholar] [CrossRef]
- Theißen, G. Shattering developments. Nature 2000, 404, 711. [Google Scholar] [CrossRef]
- Whitelaw, C.A.; Lyssenko, N.N.; Chen, L.; Zhou, D.; Mattoo, A.K.; Tucker, M.L. Delayed abscission and shorter Internodes correlate with a reduction in the ethylene receptor LeETR1 transcript in transgenic tomato. Plant Physiol. 2002, 128, 978–987. [Google Scholar] [CrossRef]
- Bastow, R.; Mylne, J.S.; Lister, C.; Lippman, Z.; Martienssen, R.A.; Dean, C. Vernalization requires epigenetic silencing of FLC by histone methylation. Nature 2004, 427, 164. [Google Scholar] [CrossRef]
- Kong, F.; Liu, B.; Xia, Z.; Sato, S.; Kim, B.M.; Watanabe, S.; Yamada, T.; Tabata, S.; Kanazawa, A.; Harada, K.; et al. Two Coordinately Regulated Homologs of FLOWERING LOCUS T Are Involved in the Control of Photoperiodic Flowering in Soybean. Plant Physiol. 2010, 154, 1220–1231. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.H.; Choi, C.W.; Kim, M.C. Identification of diversified functions of soybean FT homologs in photoperiod-dependent flowering time control. In Proceedings of the Korean Society of Crop Science Conference; The Korean Society of Crop Science: Gyeonggi-do, Korea, 2017; p. 100. [Google Scholar]
- Lutz, U.; Posé, D.; Pfeifer, M.; Gundlach, H.; Hagmann, J.; Wang, C.; Weigel, D.; Mayer, K.F.X.; Schmid, M.; Schwechheimer, C. Modulation of Ambient Temperature-Dependent Flowering in Arabidopsis thaliana by Natural Variation of FLOWERING LOCUS M. PLoS Genet. 2015, 11, e1005588. [Google Scholar] [CrossRef] [PubMed]
- Wurr, D.C.E.; Fellows, J.R.; Phelps, K.; Reader, R.J. Vernalization in calabrese (Brassica oleracea var. italica)-a model for apex development. J. Exp. Bot. 1995, 46, 1487–1496. [Google Scholar] [CrossRef]
- Liu, S.; Liu, Y.; Yang, X.; Tong, C.; Edwards, D.; Parkin, I.A.P.; Zhao, M.; Ma, J.; Yu, J.; Huang, S.; et al. The Brassica oleracea genome reveals the asymmetrical evolution of polyploid genomes. Nat. Commun. 2014, 5, 1–11. [Google Scholar] [CrossRef]
- Duan, W.; Zhang, H.; Zhang, B.; Wu, X.; Shao, S.; Li, Y.; Hou, X.; Liu, T. Role of vernalization-mediated demethylation in the floral transition of Brassica rapa. Planta 2017, 245, 227–233. [Google Scholar] [CrossRef]
- Blümel, M.; Dally, N.; Jung, C. Flowering time regulation in crops—What did we learn from Arabidopsis? Curr. Opin. Biotechnol. 2015, 32, 121–129. [Google Scholar] [CrossRef]
- Milec, Z.; Valárik, M.; Bartoš, J.; Šafář, J. Can a late bloomer become an early bird? Tools for flowering time adjustment. Biotechnol. Adv. 2014, 32, 200–214. [Google Scholar] [CrossRef]
- Arora, R.; Agarwal, P.; Ray, S.; Singh, A.K.; Singh, V.P.; Tyagi, A.K.; Kapoor, S. MADS-box gene family in rice: Genome-wide identification, organization and expression profiling during reproductive development and stress. BMC Genom. 2007, 8, 242. [Google Scholar] [CrossRef]
- Dennis, E.S.; Peacock, W.J. Vernalization in cereals. J. Biol. 2009, 8, 57. [Google Scholar] [CrossRef]
- Schmutz, J.; Cannon, S.B.; Schlueter, J.; Ma, J.; Mitros, T.; Nelson, W.; Hyten, D.L.; Song, Q.; Thelen, J.J.; Cheng, J.; et al. Genome sequence of the palaeopolyploid soybean. Nature 2010, 463, 178–183. [Google Scholar] [CrossRef] [PubMed]
- Jung, C.-H.; Wong, C.E.; Singh, M.B.; Bhalla, P.L. Comparative Genomic Analysis of Soybean Flowering Genes. PLoS ONE 2012, 7, e38250. [Google Scholar] [CrossRef] [PubMed]
- Locascio, A.; Lucchin, M.; Varotto, S. Characterization of a MADS FLOWERING LOCUS C-LIKE (MFL) sequence in Cichorium intybus: A comparative study of CiMFL and AtFLC reveals homologies and divergences in gene function. New Phytol. 2009, 182, 630–643. [Google Scholar] [CrossRef] [PubMed]
- Vogt, S.H.; Weyens, G.; Lefèbvre, M.; Bork, B.; Schechert, A.; Müller, A.E. The FLC-like gene BvFL1 is not a major regulator of vernalization response in biennial beets. Front. Plant Sci. 2014, 5, 146. [Google Scholar] [CrossRef] [PubMed]
- Sheldon, C.C.; Burn, J.E.; Perez, P.P.; Metzger, J.; Edwards, J.A.; Peacock, W.J.; Dennis, E.S. The FLF MADS Box Gene: A Repressor of Flowering in Arabidopsis Regulated by Vernalization and Methylation. Plant Cell 1999, 11, 445–458. [Google Scholar] [CrossRef] [PubMed]
- Livak, K.J.; Schmittgen, T.D. Analysis of Relative Gene Expression Data Using Real-Time Quantitative PCR and the 2−ΔΔCT Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.; Ge, X.; Sun, M. Modified CTAB Protocol Using a Silica Matrix for Isolation of Plant Genomic DNA. BioTechniques 2000, 28, 432–434. [Google Scholar] [CrossRef]
- Jones, P.; Binns, D.; Chang, H.-Y.; Fraser, M.; Li, W.; McAnulla, C.; McWilliam, H.; Maslen, J.; Mitchell, A.; Nuka, G.; et al. InterProScan 5: Genome-scale protein function classification. Bioinformatics 2014, 30, 1236–1240. [Google Scholar] [CrossRef]
- HALL, T. BioEdit: A user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp. Ser. 1999, 41, 95–98. [Google Scholar]
- Crooks, G.E.; Hon, G.; Chandonia, J.-M.; Brenner, S.E. WebLogo: A Sequence Logo Generator. Genome Res. 2004, 14, 1188–1190. [Google Scholar] [CrossRef]
- Lescot, M.; Déhais, P.; Thijs, G.; Marchal, K.; Moreau, Y.; Van de Peer, Y.; Rouzé, P.; Rombauts, S. PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res. 2002, 30, 325–327. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.; Feng, M.; Chen, W.; Zhou, X.; Lu, J.; Wang, Y.; Li, Y.; Jiang, C.-Z.; Gan, S.-S.; Ma, N.; et al. In rose, transcription factor PTM balances growth and drought survival via PIP2;1 aquaporin. Nat. Plants 2019, 5, 290–299. [Google Scholar] [CrossRef] [PubMed]
- Clough, S.J.; Bent, A.F. Floral dip: A simplified method for Agrobacterium -mediated transformation of Arabidopsis thaliana. Plant J. 1998, 16, 735–743. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Qin, R.-Y.; Li, H.; Xu, R.-F.; Yang, Y.-C.; Ni, D.-H.; Ma, H.; Li, L.; Wei, P.-C.; Yang, J.-B. Low-temperature-induced expression of rice ureidoglycolate amidohydrolase is mediated by a C-repeat/dehydration-responsive element that specifically interacts with rice C-repeat-binding factor 3. Front. Plant Sci. 2015, 6, 1011. [Google Scholar] [CrossRef]
- Ma, N.; Feng, H.; Meng, X.; Li, D.; Yang, D.; Wu, C.; Meng, Q. Overexpression of tomato SlNAC1 transcription factor alters fruit pigmentation and softening. BMC Plant Biol. 2014, 14, 351. [Google Scholar] [CrossRef]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).