Abstract
High mobility group A (HMGA) proteins are oncofoetal chromatin architectural factors that are widely involved in regulating gene expression. These proteins are unique, because they are highly expressed in embryonic and cancer cells, where they play a relevant role in cell proliferation, stemness, and the acquisition of aggressive tumour traits, i.e., motility, invasiveness, and metastatic properties. The HMGA protein expression levels and activities are controlled by a connected set of events at the transcriptional, post-transcriptional, and post-translational levels. In fact, microRNA (miRNA)-mediated RNA stability is the most-studied mechanism of HMGA protein expression modulation. In this review, we contribute to a comprehensive overview of HMGA-targeting miRNAs; we provide detailed information regarding HMGA gene structural organization and a comprehensive evaluation and description of HMGA-targeting miRNAs, while focusing on those that are widely involved in HMGA regulation; and, we aim to offer insights into HMGA-miRNA mutual cross-talk from a functional and cancer-related perspective, highlighting possible clinical implications.
1. Introduction
The high mobility group A (HMGA) family is a family of architectural nuclear proteins that are involved in the modulation of chromatin structure and regulation of gene expression. The family comprises three main members: HMGA1a and HMGA1b, which are derived from the alternative splicing of the HMGA1 gene, and HMGA2, which is derived from a different—although related—gene [1,2]. These small proteins (approximately 12 kDa) contain three DNA-binding domains, the so-called “AT-hooks”, which allow for them to bind short A/T-rich sequences through the DNA minor groove, and a highly acidic C-terminal tail [3]. These proteins have high plasticity due to their intrinsic disordered structure, enabling their interaction with a multitude of factors [4,5]. The combination of these features allows HMGA factors to organize and orchestrate the assembly of stereospecific nucleoprotein complexes at the promoter/enhancer DNA sequence level [6], thus participating in regulating the expression of numerous genes [7,8,9]. In addition, HMGA proteins can participate in chromatin relaxation and the modulation of nuclear stiffness through mechanisms that involve histone H1 competition [10] and alterations in histone H1 post-translational modifications (PTMs) [11]. HMGA proteins are involved in several cellular processes, such as cellular proliferation [12], differentiation [13], senescence [14], apoptosis [15,16], inflammation [17], metabolism [18,19], autophagy [20], DNA replication [21], DNA repair [22,23], splicing [24], and viral integration [25], given their importance within the chromatin network.
HMGA proteins are highly and widely expressed during embryonic development [26,27], where they play essential functions, as demonstrated by individual and combined knockout (KO) of Hmga1 and Hmga2 [19,26,28,29]. Conversely, HMGA expression, particularly HMGA2 expression, is generally very low or absent in adult tissues. Therefore, establishing finely regulated control of HMGA expression is very important in the correct development and the maintenance of adult cellular homoeostasis. Accordingly, aberrant expression of HMGA proteins due to the dysregulation of their expression or the expression of mutated forms causes several diseases, such as different forms of neoplasia and metabolic disorders, which have been extensively reviewed [30,31,32], and it is involved in other pathologies such as polycystic ovary syndrome, sporadic Alzheimer’s disease, myocardial infarction, obesity, ischaemia, atherosclerosis, and sepsis [24,28,33,34,35,36,37]. Therefore, precise spatiotemporal regulation of the expression of HMGA factors is crucial in the correct development and preservation of adult physiological conditions. The transcriptional regulation of both the HMGA1 and HMGA2 genes has been extensively and recently reviewed: HMGA1 is an inducible gene that is mainly regulated by transcription factors at different promoter regions and enhancers, while the HMGA2 promoter seems to be constitutively active in different cell lines, and its activity can be modulated either positively or negatively by different DNA-binding factors [38]. In addition, an R-loop-based mechanism has been demonstrated to be involved in HMGA2 gene transcription modulation, providing an open chromatin conformation for HMGA2 transcriptional cis-regulatory sequences [39]. Moreover, HMGA2 protein levels can also be modulated by stabilizing interactions with long non-coding RNA (lncRNA) molecules [40].
HMGA proteins are subjected to many PTMs that regulate their ability to bind DNA and they interact with several other factors; therefore, post-translational regulation is a relevant step contributing to the regulation of their activity [41].
Post-transcriptional regulation is a key process that regulates gene expression, and it is often altered in cancer cells [42,43]. Although both the 5′ untranslated region (5’UTR) and 3′UTR can contribute to this process, the 3′UTR in particular is more often a target of microRNAs (miRNAs), which are the most important factors that are involved in this type of regulation. Hundreds of papers have described miRNA-mediated regulation of both HMGA1 and HMGA2 mRNA in different cell types and stages (Table 1); in particular, HMGA2 regulation by let-7 is considered to be a paradigm of a miRNA action [44,45,46]. Moreover, lncRNA provides another layer of post-transcriptional regulation complexity; lncRNAs have been demonstrated to play a role in modulating HMGA expression by sponging HMGA miRNAs [47,48].

Table 1.
High mobility group A 1 (HMGA1)- and HMGA2-targeting of microRNAs (miRNAs).
A comprehensive overview and critical evaluation of the role of miRNA in regulating HMGA1 and/or HMGA2 protein expression is still missing from the literature, despite this knowledge. In this review, we specifically focus on the HMGA-targeting miRNA network. We (i) provide insights into the structural organization of HMGA genes; (ii) offer a complete overview of HMGA-targeting miRNAs, focusing on those that have been more deeply investigated; (iii) discuss the hypothesis of the mutual influence of HMGA1/HMGA2; and, (iv) highlight the role of HMGA mRNAs as competing endogenous RNA (ceRNA) molecules in the context of cancer initiation and development.
2. HMGA Genes: Structural Organization
Analysis of the organization of the HMGA1 gene [1,178] identified eight exons, several promoter regions, several transcription start sites, and numerous alternatively spliced exons, which generate different mRNAs encoding the two major protein isoforms (HMGA1a and HMGA1b) expressed in human cells. Interestingly, the three DNA-binding domains of the HMGA proteins, the AT-hooks, are encoded by three different exons, while the last exon encodes the acidic C-terminal tail and it includes the 3′UTR. The referenced papers and other papers [179,180,181] indicate that the 5′UTR of HMGA1 can be very heterogeneous, due to the use of different promoters and the occurrence of alternative splicing within the first four exons. However, no variants were described in the last four exons, which include the open reading frames (ORFs) and 3′UTR, except for the alternative splicing event in the first of these exons that generates either HMGA1a (HMGA1-201; HMGA1-205) or HMGA1b (HMGA1-204; HMGA1-202; HMGA1-203) (Figure 1) and the alternative splicing event that results from the usage of a non-canonical splice site that generates the rare HMGA1c transcript, which has only been described once and it is not present in Ensembl Genome Browser [182]. Remarkably, all of the transcripts that are described in these papers share the same 3′UTR.
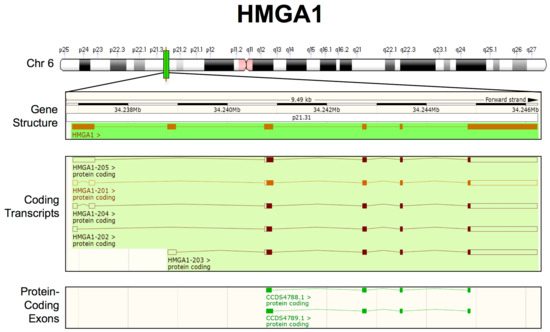
Figure 1.
Updated human HMGA1 transcripts structure from Ensembl Genome Browser (GRCh38.p12-GCF_000001405.38). From top to bottom: chromosome 6 (Chr 6) labelled with chromosome bands (white, black and grey), centromeres (pink) and the HMGA1 locus (green); the HMGA1 gene structure, showing the entire length of the region considered, the location on Chr 6 (NC_000006.12):34, 236, 485–34, 246, 445, the chromosome band (p21.31) and HMGA1 exons/introns (orange boxes/lines); and the HMGA1 coding transcripts. The empty boxes indicate UTRs, whereas the filled boxes indicate open reading frames (ORFs). Merged Ensembl/Havana and Ensembl protein-coding data are shown in orange and brown, respectively; HMGA1 protein-coding exons are highlighted in green.
Currently, whole-genome sequencing projects, together with transcriptome analysis, can provide a more complete picture of transcripts that originate from a specific locus. Starting from this consideration, we investigated whether new coding transcripts can arise from the human HMGA1 gene. We used Ensembl Genome Browser 96 for this purpose, which collects open-access, integrated genome, gene, variation, gene regulation, and comparative genomic information [183] (Figure 1). This analysis did not reveal further complexity in HMGA1 transcripts; therefore, we can conclude that all of the transcripts share the same 3′UTR and can thus be targeted by the same miRNAs.
The HMGA2 gene immediately appeared to be more complex than the HMGA1 gene, essentially due to its length and the presence of a very large intron involved in rearrangements, leading to truncated or chimeric HMGA2 transcripts especially in benign mesenchymal tumours [184,185]. Indeed, the HMGA2 gene was identified, because it was found to be rearranged in several tumours at the level of the third intron [184,185] and was only subsequently completely cloned and characterized [2,186]. The HMGA2 gene is organized into five exons that encode different protein domains. Unlike HMGA1, it contains an additional exon coding for a very short peptide of 11 amino acids (aa) that separates the final DNA-binding domain from the acidic C-terminal tail. Within the long intron, some additional exons have been described to be part of alternative splicing transcripts [187,188]. We performed analysis on publicly available transcriptomic data to better define the HMGA2 transcripts originating from the HMGA2 locus (Figure 2). In addition to the canonical HMGA2 transcript (HMGA2-204, according to Ensembl nomenclature), six splicing variants were identified, each of which ends with a different 3′UTR sequence and encodes a protein with a different C-terminal tail. Four of these variants (HMGA2-201; HMGA2-206; HMGA2-205; and, HMGA2-203) contain exons that are derived from the large third intron, while two (HMGA2-210 and HMGA2-202) contain exons derived from the fourth intron. In conclusion, unlike HMGA1, which only includes one 3′UTR, HMGA2 has several splicing variants with 3′UTRs different from the canonical 3′UTR. This difference could have implications for HMGA2-mediated miRNA regulation. In fact, it has been reported that an alternative isoform that escapes miRNA-mediated targeting (HMGA2-203) is involved in human haematopoietic stem cell development [189].
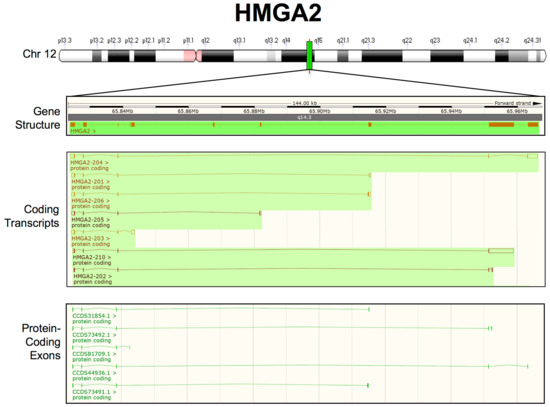
Figure 2.
Updated human HMGA2 transcripts structure from the Ensembl Genome Browser (GRCh38.p13-GCF_000001405.39). From top to bottom: chromosome 12 (Chr 12) labelled with chromosome bands (white, black, and grey), centromeres (pink), and the HMGA2 locus (green); the HMGA2 gene structure, showing the overall length of the region considered, the location on Chr 12 (NC_000012.12):65,823,216–65,968,410, the chromosome band (q14.3) and HMGA2 exons/introns (orange boxes/lines); and, HMGA2 coding transcripts. The empty boxes indicate UTRs, whereas the filled boxes indicate ORFs. Merged Ensembl/Havana and Ensembl protein-coding data are shown in orange and brown, respectively; HMGA2 protein-coding exons are highlighted in green.
3. miRNA Regulation of HMGA Transcripts
More than one hundred miRNAs are involved in the regulation of HMGA mRNA (Table 1). Among these miRNAs, we discuss, in detail those that are most widely studied, beginning with the let-7 family, which is one of the first examples of miRNA-mediated oncogene regulation [44,45].
3.1. The lin28/let–7 Axis
The description of miRNA-mediated HMGA protein regulation begins with the finding that chromosomal abnormalities in the regions 12q15 and 6p21.3 were associated with the aberrant expression of either HMGA2 or HMGA1 in several benign mesenchymal tumours [190]. Invariably, in the aberrant HMGA transcripts, the region coding for the N terminal had intact DNA-binding domains, while the region coding for the C-terminal tail, which included the 3′UTR, was deleted or substituted with other transcripts. Together with the frequent finding of a lack of correspondence between HMGA protein and mRNA levels, particularly for HMGA2, these data led to the hypothesis that regulatory elements within the 3′UTR could mediate the post-transcriptional control of HMGA protein expression. The first clue was provided by the finding that sequences in the 3′UTR of both HMGA1 and HMGA2 mRNAs could negatively control the expression of these mRNAs, whereas the deletion of these sequences—mimicking cytogenetic aberrations found in human tumours—led to protein overexpression [191]. The discovery that miRNAs of the let-7 family, which are potent regulators of larval development and adult fate specification in nematodes [192], are also expressed in human cells [193] paved the way for completing the puzzle: these ancient small molecules could also target six sites in the 3′UTR of HMGA2 [44,45], which represses its expression via mRNA degradation. The final picture was completed by the observation that translocations of the HMGA2 gene leading to the loss of 3′UTR-mediated let-7 repression could activate HMGA2 expression and lead to cell transformation, anchorage-independent growth [44], and proliferation [45]. Following translocation, the HMGA2 3′UTR was frequently found to be bound to the 3′ end of tumour suppressor genes, such as RAD51L1 and FHIT [44], thus further promoting tumorigenesis.
Unlike in C. elegans, the human let-7 miRNA family includes 13 evolutionarily conserved members (let-7a-1, 7a-2, 7a-3, 7b, 7c, 7d, 7e, 7-f1, 7f-2, 7g, 7i, mir-98, and miR-202) [194,195] that share the same seed sequence, but are located on eight different chromosomes [196]. In addition to HMGA2 truncation, a decrease in let-7 expression can be solely responsible for the increased expression of otherwise normal HMGA2 or HMGA1 [51] in several different cancers, such as breast [69], gastric [70] and non-small cell lung cancers [63], sarcomas [197,198], hepatocellular carcinomas, nasopharyngeal [72] and oesophageal squamous cell carcinomas [199], uterine leiomyomas and leiomyosarcomas [66,200], and pituitary adenomas [201], as a loss of let-7 expression is a marker for less well-differentiated cancers [46]. Moreover, a role for let-7/HMGA in epithelial-mesenchymal transition (EMT), cell migration, and metastasis has been demonstrated in several cancers and cellular systems [51,72,202,203,204,205].
In addition to its role in cancer, the let-7/HMGA2 axis was shown to regulate physiological processes, such as adipose [206], osteogenic [71], myeloerythroid [207] and gliogenic differentiation [208], post-natal proliferation and ageing [209], and glucose metabolism [210]. This role mechanistically explains the finding of let-7 downregulation in pathologies, such as diffuse lipomatosis [211] and renal fibrogenesis [212]. The let-7/HMGA2 axis affects self-renewal and stemness in stem cells of different origins, including haematopoietic, neural, breast, lung, and intestinal cancer stem cells ([69,213,214,215,216], respectively). Most importantly, the let-7b/HMGA2 axis has been shown to induce the direct conversion of adult somatic cells into induced neural stem cells [56].
The factors controlling let-7 expression form a regulatory feedback loop with downstream targets of let-7 and are pivotal regulators of stemness or differentiation. The major molecules regulating let-7 are the two RNA-binding proteins LIN28A and LIN28B, [217], which recruit terminal uridyltransferase (TUT4) and add an oligomeric U at the 3′ end, which prevents pre-let-7 from being processed into a mature miRNA [218]. Strikingly, the LIN28 proteins dramatically affect HMGA1 and HMGA2 through both let-7-dependent and let-7-independent mechanisms [195]. The components of this network are linked by entangled positive and negative reciprocal regulatory activities that each factor exerts on the other factors and on itself (Figure 3) [219].
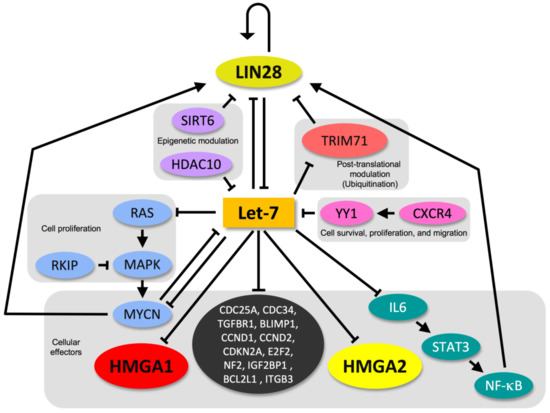
Figure 3.
The LIN28/let-7 circuitry. HMGA proteins are inserted in the connected regulatory network of the LIN28/let-7 axis, which contains several regulatory feedback loops.
3.2. hsa-miR–26a
The hsa-miR-26 family includes miR-26a and miR-26b. miR-26a is specifically located on chromosome 3p22, a region that is subjected to the loss of heterozygosity in cancer [220]. In the physiological context, miR-26a controls cell growth, development, and differentiation in processes, such as myogenesis [221]. Via the transfection of cells with vectors containing a wild-type or mutated 3′UTR of HMGA1 or HMGA2 and reporter gene assays, several studies have demonstrated that miR-26a specifically targets well-conserved regions of the 3′UTRs of HMGA1 [87,89,90,92,95] and HMGA2 [91,93,94]. miR-26a downregulates HMGA1 at both the mRNA and protein levels, as assessed by qRT-PCR and Western blot analyses [87,90,92], while HMGA2 downregulation has only been validated at the protein level [93,222]. The role of the miR-26a/HMGA1 axis in the context of cancer has been extensively studied, and this axis has been shown to act on the proliferation and migration of pancreatic cancer [96], bladder cancer [95], breast cancer [92], lung adenocarcinoma [90], and osteosarcoma [89] cells. A negative correlation between miR-26a and HMGA1 levels has also been shown in cancer tissue specimens, specifically in tissues from urothelial bladder cancer patients [223] and osteosarcoma patients- [89]. Moreover, miR-26a acts on the inflammatory process by downregulating HMGA1 and MALT1, thus impacting the TNF-α/NF-κB inflammatory genes in human bronchial epithelial cells [88]; in addition, the miR-26a/HMGA1 axis controls coronary microembolization-induced myocardial inflammation [87]. miR-26a target HMGA2 mRNA, which regulates cellular senescence [224] and EMT in idiopathic pulmonary fibrosis both in vitro and in vivo [93]. In addition, miR-26a and HMGA2 mRNA are inversely correlated in gallbladder cancer (GBC) tissues, and miR-26a reduces the proliferation of GBC cells [94] and human lung adenocarcinoma cells, which increases the sensitivity to cisplatin treatment via HMGA2 [91].
3.3. hsa-miR–33b
The intronic human hsa-miR-33b-5p belongs to the miR33 gene family and it is located in a non-coding region of the human SREBP-1 gene at the 17p11.2 genomic locus (miRbase, HGNC) [225]. miR-33b can target the HMGA2 3′UTR, which prevents the expression of the HMGA2 protein [101,102,103,104].
miR-33b is downregulated in breast tumour and melanoma tissue samples and it is inversely correlated with the clinical stage [101,102].
Ectopic expression of miR-33b, which affects the levels of HMGA2, SALL4 and Twist1, can decrease the stem cell-like properties of breast cancer cells. Moreover, miR-33b suppresses cell migration and invasion in vitro through decreases in HMGA2 and Twist1 expression [101]. miR-33b also reduces the migration and invasiveness of melanoma cell lines upon cordycepin exposure via targeting the HMGA2 3′UTR [102]. miR-33b exerts its anti-tumorigenic effects on human gastric cancer cells by affecting HMGA2 expression to inhibit cell growth and increase cellular sensitivity to docetaxel and cisplatin [104]. In addition, the upregulation of miR-33b by EF24, which is a curcumin analogue, suppresses EMT and the induction of migration in melanoma cell lines via the targeting of HMGA2 [103].
miR-33b modulates pathways controlling the levels of high-density lipoprotein (HDL) cholesterol, triglycerides, and insulin signalling, and negatively regulates adipogenesis [100,226]. miR-33b expression was found to be elevated in ovarian tissues of rats with insulin-resistant polycystic ovary syndrome and to inhibit GLUT4 expression by targeting HMGA2 3′UTR, therefore contributing to the progression of insulin resistance in PCOS/IR rats [80]. Furthermore, HMGA2 mediates the effects of miR-33b on adipogenesis in the cells from patients with Simpson-Golabi-Behmel syndrome [100].
3.4. hsa-miR–98
hsa-miR-98-5p belongs to the let-7 gene family and it is located at the Xp11.22 genomic locus (miRbase, HGNC site) [225]. Several studies have reported that miR-98 binds the 3′UTR of HMGA2 mRNA, regulating its expression [107,108,109,110,111,112]. A correlation between miR-98 and chemoresistance that is associated with the downregulation of HMGA2 was found in head and neck squamous cell carcinoma, providing the first link between miR-98 levels and HMGA2 expression [227]. Other authors have reported that miR-98 reduces tumour aggressiveness through the inhibition of HMGA2 expression [107,108,110,111]. miR-98 was shown to downregulate HMGA2 expression in glioma cells as a part of the RKIP/miR-98/HMGA2 axis, leading to a reduction in glioma cell invasion [110]. In breast cancer, retinoblastoma, and laryngeal squamous cell carcinoma, restoration of miR-98 with the consequent inhibition of HMGA2 expression reduced the proliferative and migratory abilities of cells, EMT, and metastasis formation in vivo [107,108,111]. Another work showed that miR-98 could reduce the inflammatory response and alleviate neuropathic pain progression in chronic constriction injury-induced (CCI) rats by binding to the HMGA2 3′UTR, [112]. The miR-98/HMGA2 axis is also involved in differentiation, as the expression of miR-98, with the consequent downregulation of HMGA2, promotes the expression of the osteogenic differentiation genes RUX2, BSC, and OCN in mesenchymal stem cells [109].
3.5. hsa-miR-16
miR-16 is considered to be a critical tumour suppressor miRNA downregulated in many types of cancer, among the validated miRNAs targeting both HMGA1 and HMGA2 [228]. Via a combined bioinformatic and molecular approach, HMGA1 was demonstrated to be directly targeted by miR-16 [84]. A luciferase assay that was performed on HMGA1 mutant constructs at predicted miR-16 binding sites determined that only one site is active. Moreover, the overexpression of miR-16 induced a decrease in the endogenous HMGA1 protein level due in part to a decrease in the HMGA1 mRNA level. In fact, a decrease in HMGA1 mRNA has only been found in HeLa cells, but not in MCF-7 cells, revealing a cell-dependent mechanism for HMGA1 regulation [84]. Later, another group confirmed these results on HMGA1 and demonstrated that miR-16 could also directly target HMGA2 [50]. Bioinformatic tools predicted one miR-16 binding site in the 3′UTR of HMGA2; subsequently, via a luciferase assay, miR-16 was demonstrated to target HMGA2. Consistent with previous observations, the overexpression of miR-16 reduced both HMGA1 and HMGA2 mRNA and protein levels in the GH3 rat pituitary adenoma cell line. Accordingly, another work described the relationship between miR-16 and HMGA2 in pituitary adenoma cells [83]. Interestingly, both works showed an inverse association between miR-16 and HMGA expression levels in human pituitary adenomas; in fact, they found that miR-16 was downregulated, but HMGA1 and HMGA2 were upregulated in pituitary adenomas when compared to normal pituitary tissues, consistent with the action of miR-16 in regulating the expression of HMGA1 and HMGA2 [50,83]. Recently, a new layer of miR-16/HMGA2 regulation in pituitary tumours has been added—ribosomal protein SA pseudogene 52 (RPSAP52), an antisense lncRNA targeting the HMGA2 gene was demonstrated to increase HMGA2 protein expression via a ceRNA mechanism, in which it acts as a sponge for miR-16 and miR-15 [48].
3.6. hsa-miR-142-3p
hsa-miR-142 shows pleiotropic functions in physiological processes that are connected with embryonic development and stem cell pluripotency, whereas the alteration of its expression supports the development of cardiovascular disease [229] and tumours [133,230,231,232]. Decreased levels of miR-142-3p have often been reported in cancer tissues when compared with normal tissues, supporting the data while considering miR-142-3p mainly an onco-suppressor miRNA [233,234,235]. Interestingly, the “guide strand” of miR-142-3p is negatively modulated by interleukin-6 in glioblastoma multiforme (GBM), the most aggressive and stem cell-rich primary brain tumour [133], causing the upregulation of HMGA2 protein expression. Indeed, miR-142-3p decreased HMGA2 protein levels. This decrease led to inhibited expression of SOX2, which is a target gene of HMGA2 and a master regulator of stemness features, thus inducing the suppression of cancer cell pluripotency and tumour cell growth. Therefore, in GBM, miR-142-3p downregulation is required to maintain stem cell-like properties and cell proliferation through HMGA2 upregulation. Metformin, which is a first-line drug for type 2 diabetes mellitus, was proposed to disrupt the sponge effect of MALAT1 on miR-142-3p, enabling miR-142-3p to bind the HMGA2 3′UTR in cervical cancer and inhibit cervical cell invasion and migration [132]. Moreover, the miR-142-3p response element is also shared by HMGA1. Indeed, an inverse correlation between miR-142-3p and HMGA1 expression levels was found in osteosarcoma tissues, and miR-142-3p was subsequently demonstrated to inhibit the growth, migration, and invasion of osteosarcoma cells by targeting HMGA1 [134].
4. HMGA1 and HMGA2: Common and Specific miRNAs
Among the 75 miRNAs experimentally explored for their ability to target HMGA mRNA, 26 targeted HMGA1, 64 targeted HMGA2, and 15 targeted both HMGA1 and HMGA2 (Table 1). We used TargetScan software [236] to identify predicted HMGA-targeting miRNAs and compare them with those that have been experimentally validated. We arbitrarily restricted the list of predicted HMGA-targeting miRNAs to consider both miRNA families broadly conserved among vertebrates and miRNA families conserved only among mammals. A total of 132 predicted HMGA-targeting miRNAs were identified; 41 targeted HMGA1, 115 targeted HMGA2, and 24 targeted both HMGA1 and HMGA2 (Table 1). The number of predicted miRNAs is approximately twice that of experimentally validated miRNAs; however, notably, the target distribution in the two categories is almost identical (Figure 4, panel A). The experimental and bioinformatic data both suggest that the modulation of HMGA2 protein expression is controlled by a more wide miRNA network than the modulation of HMGA1 protein expression, which could simply be because the HMGA2 3′UTR is longer than the HMGA1 3′UTR (Figure 2).
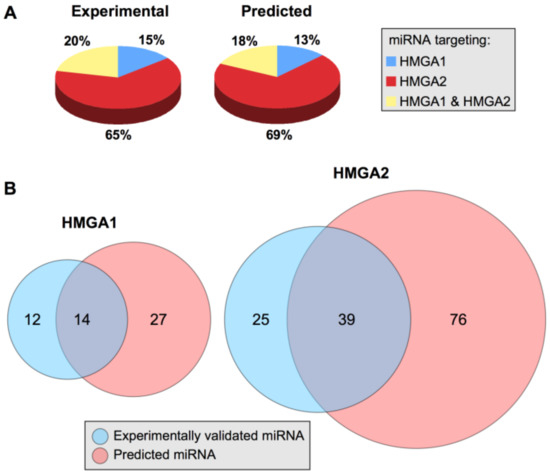
Figure 4.
An overview of HMGA1- and HMGA2-targeting miRNAs. (A) Comparison of the percentage distributions of experimentally validated and predicted HMGA1- and HMGA2-targeting miRNAs. (B) Venn diagrams of experimentally validated and predicted miRNAs targeting HMGA1 and HMGA2.
A comparison between experimentally validated and bioinformatically predicted miRNAs revealed that of the experimentally validated miRNAs targeting HMGA1 (26), 54% (14) were also predicted by TargetScan and that almost the same percentage (61%; 39 miRNAs) of the 64 miRNAs targeting HMGA2 were also predicted by TargetScan (Figure 4, panel B). miRNA target prediction programs usually generate many false positives [237]. However, while only focusing on miRNAs that have been experimentally validated, it is evident that a very high number of miRNAs are involved in modulating HMGA protein expression. In general, the quantitative effect of miRNA target repression is very limited [238,239]; therefore, it is reasonable to hypothesize that the HMGA protein expression level could be controlled by a set of cooperating miRNAs.
For each predicted miRNA, we report in Table 1 the cumulative weighted context ++ score (CWCS), which is a score that is related to the predicted efficacy of the sites. On the basis of this score, this table allowed for us to speculate that the miRNA network that is involved in regulating HMGA protein expression could be even more connected than it actually is and some miRNAs could very possibly be added to the experimentally validated list. A set of miRNAs already experimentally validated to target one HMGA mRNA could be reasonably tested for targeting of the other HMGA mRNA: miR-98-5p and miR-107 could be validated for HMGA1, miR-424-5p could be another miRNA targeting both HMGA mRNAs, and miR-497-5p could be validated for HMGA2. Moreover, a set of miRNAs with promising CWCSs (i.e., miR-17-5p, miR-20-5p, miR-93-5p, miR-132-3p, miR-137, miR-190-5p, miR-212-3p, miR-212-3p, miR-519-3p, miR-532-3p, miR-485-5p, miR-491-5p, and miR-760) could be reasonably tested.
We believe that the shared regulation of HMGA1 and HMGA2 by a set of miRNAs implies a degree of interdependence between HMGA1 and HMGA2. Studies have already demonstrated that the expression of HMGA2 could be controlled by the expression levels of the HIF1A 3′UTR via a ceRNA mechanism [240]. Therefore, a finding that the expression of the two HMGA genes might be linked through a ceRNA mechanism would not be surprising. A bioinformatic search for mRNAs, which can act as ceRNAs that target HMGA1 or HMGA2, showed that HMGA2 mRNA had the highest ceRNA score for HMGA1 (Competing Endogenous mRNA DataBase (ceRDB), https://www.oncomir.umn.edu/cefinder/index.php). [241]. Let-7 is a master regulator of HMGA2, and the HMGA2 3′UTR contains 7 predicted let-7 consensus sequences, while the HMGA1 3′UTR contains only one. Simple reasoning suggests that, at least regarding the effects of let-7, the HMGA2 3′UTR-sequestering activity is very high and, therefore, even low HMGA2 expression levels could have a relevant impact on HMGA1 expression. In contrast, only a very high expression level of HMGA1 could relieve the suppressive effects of let-7 on HMGA2. The targeting of both mRNAs by multiple miRNAs suggests a degree of interdependence between the two proteins; however, in most research that is related to HMGA proteins, only one protein (i.e., HMGA1 or HMGA2) is studied without considering the possible interdependence with the other. This aspect might be considered in future studies.
While considering that the transcription of the two HMGA genes is controlled by different pathways/stimuli [38], the post-transcriptional modulation via miRNA adds another layer to the regulation of HMGA protein expression, and the integration of transcriptional and post-transcriptional mechanisms could dictate the relative HMGA1/HMGA2 protein expression levels.
5. HMGA–Targeting miRNAs: Their Role in the Modulation of Other Relevant Pathways Involved in Cancer
The power of miRNA regulation depends, in part, on the ability of these molecules to simultaneously target several mRNAs, thus affecting different pathways and inducing a fan-shaped effect. The types of targets and the pathways affected, however, occasionally make defining a miRNA as an oncogene or onco-suppressor difficult, as one miRNA sometimes performs opposing functions. The existence of self-regulating feedback loops or the effect of different cellular backgrounds might explain these dichotomies.
An analysis of the miRTarBase database (http://mirtarbase.mbc.nctu.edu.tw) [242] allowed for us to highlight additional and strongly validated targets (assessed by reporter assays, Western blotting and qPCR) of the major HMGA-targeting miRNAs. The let-7 family, in particular, is a miRNA family with several validated targets, including cell cycle regulators (such as MYC, HRAS, KRAS, CDC25A, NKIRAS2, CDC34, TGFBR1, BLIMP1 (or PRDM1), CCND1, CCND2, CDKN2A, and E2F2), transcriptional and translational regulators (such as NF2 and IGF2BP1), apoptosis regulators (BCL2L1), stem cell regulators (TRIM71), and EMT mediators (ITGB3). A potent positive feedback loop links inflammation and cancer through the NF-κB/LIN28/let-7 axis. NF-κB/LIN28-mediated let-7 repression leads to the activation of the IL-6, STAT3, and NF-κB mediators (Figure 3), thus maintaining the epigenetic transformed state in the absence of the inducing signal [243].
On the other hand, some let-7 targets function as molecular sponges that can remove let-7 molecules from other targets, hence functioning as oncogenes. In MYCN-amplified neuroblastoma cell lines, MYCN mRNA levels are exceptionally high and they are sufficient for sponge let-7 [244]. Not only mRNAs but also lncRNAs, can sponge let-7 and regulate important physiological processes, as seen for the lncRNA H19, which harbours several let-7 binding sites and mediates muscle differentiation [245].
MirTarBase screening for strongly validated targets shows that miR-26a also affects tumorigenesis, by often acting as a tumour suppressor and targeting genes that regulate fundamental cancer hallmarks, including the histone methyltransferase EZH2 [246,247], the pro-metastatic Methaderin (MTDH) [248], and the cell cycle regulator CCNE2 [249]. miR-26a participates in differentiation and angiogenic programs by targeting SMAD1 [250], whereas, in human gliomas, it is amplified and can target PTEN [251] and RB1 [252].
Validated, relevant miR-33b targets include PIM1 [253] and XIAP, both of which are involved in apoptosis [254], and ZEB1 [102,255], and the aforementioned TWIST1 [102], whose cooperation in the miR-33b/HMGA2/Twist1/ZEB1 axis plays an important role in modulating melanoma dissemination [102].
The strongly validated miR-98-5p targets include EZH2 [247], IL-6 [256], PGRMC1 [257], HK2 [258], ITGB3 [259], E2F2 [260], and FUS1 [261].
As miR-16 is considered to be a crucial onco-suppressor, many researchers have investigated the mechanism by which it affects cancer properties and have revealed that it is involved in targeting numerous genes that are crucial in mediating different cancer hallmarks [228]. Via analysis with the MiRTarBase tool, we found 34 strongly validated targets in addition to HMGA1 and HMGA2. Among these targets were CDK6, CCND1, CCND3, and CCNE1 [262], which indicated that miR-16 is involved in a mechanism by which it represses proliferation. Moreover, miR-16 targets VEGFA [234,263], YAP1 [264], FGF2, and FGFR1 [265], indicating its critical role in regulating the signalling, angiogenic, migratory, and invasive properties of cancer cells.
miR-142-3p has been validated to target a wide range of mRNAs [229]. Among these mRNAs, some factors that are crucial in cancer progression, specifically in metastasis promotion, such as TGFBR1 [266], RAC1 [233], and ROCK2 [267,268], were highlighted in the miRTarBase tool, emphasizing the role of miR-142-3p as an onco-suppressor.
6. HMGA–Related lncRNAs: Their Role as ceRNAs for HMGA-Targeting miRNAs
The abundance of miRNA response elements (MREs) in the HMGA1 and HMGA2 sequences not only makes them susceptible to negative regulation by miRNAs, but can also influence other transcripts or non-coding genes, such as lncRNAs or pseudogenes that share the MREs. Thus, the HMGA1 and HMGA2 mRNAs can reciprocally regulate each other by acting as ceRNAs.
One of the first lines of evidence indicating that HMGA mRNAs participate in the ceRNA network came from the identification and characterization of two HMGA1 non-coding processed pseudogenes, HMGA1P6 and HMGA1P7. The conserved seed sequences in miRNAs that target the HMGA1 and HMGA2 mRNAs that are also present in the HMGA1 pseudogenes transcripts can protect HMGA mRNAs from miRNA inhibition; therefore, pseudogene overexpression can sustain cancer cell proliferation and migration and inhibit cell death [269]. Consistent with this finding, HMGA1 pseudogenes upregulate several cancer-related genes [269], and RNA sequencing (RNA-Seq) of HMGA1P7 transgenic mouse embryonic fibroblasts (MEFs) revealed that HMGA1P7 mRNA induces overexpression of H19 and Igf2 by acting as ceRNA [270]. Interestingly, the overexpression of miR-483 and miR-675, whose maturation is dependent on the transcription of H19 and Igf2, is dependent on early growth response protein 1 (Egr1), whose level is directly increased by HMGA1P7 via miRNA sponging. The consequent increase in the Egr1 level induces H19 and Igf2 transcript maturation and then generates miR-483 and miR-675, two oncogenic miRNAs (oncomiRs) that are overexpressed in several neoplasias [271].
Accumulating lines of evidence reveal that lncRNAs are also critical regulators of HMGA-dependent tumour growth. HOXC13-AS, a lncRNA that is involved in the HMGA2 ceRNA network, was recently identified. It can function as a ceRNA in nasopharyngeal carcinoma to promote the expression of HMGA2 by sponging miR-383-3p, and its silencing induced cell cycle arrest and apoptosis, consistent with the inhibition of the pro-proliferative function of HMGA2 [272].
Moreover, a very recent analysis of lncRNA expression in gonadotroph adenomas revealed the RPSAP52 gene, which is the head-to-head natural antisense transcript of HMGA2, among the most highly upregulated lncRNAs. RPSAP52 acts as an endogenous sponge, which protects HMGA1 and HMGA2 from miR-15a-, miR-15b-, and miR-16-mediated inhibition. RPSAP52 modulates HMGA expression and promotes cell proliferation by acting as a decoy for miRNAs and, most likely, redirecting them to other genes by interacting with RNA-binding proteins to alter the local DNA conformation [48]. Intriguingly, RPSAP52 can also enhance HMGA2-IGF2BP2-RAS axis activity and the balance between the LIN28B and let-7 levels [273] and can positively regulate HMGA2 transcription [39]. Table 2 shows a list of lncRNAs that are involved in the HMGA-ceRNA network.

Table 2.
List of lncRNAs acting as ceRNA towards HMGA-targeting miRNA.
7. HMGA Post-Transcriptional Regulation: Is It Only a Matter of miRNA?
As the discovery of miRNA-mediated post-transcriptional regulation of RNA in C. elegans [192], this mechanism of regulating RNA stability and translation has been described in other organisms and it is now considered to be the most prevalent example of post-transcriptional regulation, at least in mammals. The observation that the 3′UTRs of mammalian RNAs are highly conserved [278] and extensively bound by proteins [279] suggests that other mechanisms, known in part but largely unexplored, could regulate RNA stability, translation, and sub-cellular localization. For example, the Hmga2 transcript was selected as a case study, and regulatory sequences in its 3′UTR were systematically identified [280]. A high-resolution map of the 3′UTR was generated via reporter gene assays, and numerous previously unidentified regulatory sites that were well conserved across species were found. Interestingly, several sites with a positive effect on gene reporter expression have been discovered, including some candidate positive regulatory elements containing U-rich or CU-rich sequences consistent with AU-rich elements (AREs), in addition to previously identified let-7 miRNA target sites. Intriguingly, very recently, it was shown that the circular RNA circNSUN2, by interacting with a short sequence within an ARE in the 3′UTR of HMGA2 mRNA, plays a critical role in promoting the formation of a ternary complex with the RNA-binding protein IGF2B2 that stabilizes HMGA2 mRNA to promote colorectal liver metastasis [281].
In addition, the RNA-binding protein IMP3 has been found to form cytoplasmic granules that contain HMGA2 mRNA and other let-7 targets, including LIN28B, and to stabilize HMGA2 by opposing the action of let-7 on HMGA2 mRNA. This effect was not achieved by direct competition between IMP3 and let-7, but by providing HMGA2 with an IMP3-dependent, membrane-less cytoplasmic domain (i.e., IMP granules) that is devoid of RNA-induced silencing complex (RISC) pathway components [282].
8. Conclusions
Accumulating evidence indicates not only the importance of miRNAs as molecular players in tumorigenesis, but also their valuable diagnostic and therapeutic applications. The term “exo-miRNAs” has been coined to describe miRNAs that are stored in cellular exosomes and released into biological fluids, where they can be easily detected due to their high stability to provide more precise diagnoses and more personalized therapeutic strategies [283].
Among the HMGA-targeting miRNAs, the let-7 family members undoubtedly have the greatest clinical relevance, even though some of the data are contradictory and need further investigation [284]. Let-7 expression has been shown to have prognostic value in many tumours [53,70,200,201,285]. Moreover, the evaluation of let-7 family member expression in body fluids (plasma, serum, urine, and stool) has demonstrated utility in the early detection of tumours, as a prognostic tool [284] and as a chemoresistance predictor [205]. The downregulation of let-7a-5p in serum has been shown to predict lymph node metastasis and prognosis in colorectal cancer patients [286], whereas the RKIP/let-7 pathway metastasis signature has been demonstrated to predict a high risk of metastasis in breast cancer with higher accuracy than the individual genes [287].
An interesting application is in the monitoring of patients during therapy for cancers, such as lung cancer and acute promyelocytic leukaemia, since reduced let-7 plasma levels are considered to be an indicator of relapse [288].
On the other hand, the HMGA-targeting miRNAs are also attractive for their potential applications in cancer therapy, acting as drug targets or as off-the-shelf drugs.
Accordingly, some anticancer drugs have been shown to act through let-7 activation/induction, and their use has been proposed alone or in combination with other treatments. The pan-deacetylase inhibitor panobinostat, for example, affects liver cancer cell lines by inducing the let-7b-mediated downregulation of HMGA2 [289]. Similarly, phenformin has been shown to target the let-7/HMGA2 axis and has been proposed to be effective in combination with temozolomide against glioblastoma [290].
Even more fascinating is the potential administration of let-7 mimics as a next-generation treatment for cancer [291]. In this context, the administration of let-7g miRNA was shown to sensitize fluorouracil-resistant human hepatoma cells [292], whereas a locked nucleic acid-GapmeR (LNA-GapmeR) containing a let-7b mimic was shown to suppress tumour growth in vivo by regulating MYC expression in multiple myeloma and potentially in other MYC-dependent cancers [293].
In addition to let-7, which appears to be a dominant miRNA that is involved in controlling HMGA proteins, particularly HMGA2, a knotty miRNA network clearly controls the regulation of HMGA protein expression. Obviously, each miRNA has been separately studied, but it is reasonable to assume that the final effect on the modulation of HMGA protein expression is achieved by the cooperation of multiple miRNAs and it is not necessarily a simple additive effect. In addition, the inverse mechanism is intriguing, i.e., the possibility that HMGA transcripts act as ceRNAs. This aspect, together with the observation that miRNAs have multiple targets, suggests that small changes in one factor could lead to connected and amplified biological outcomes, but the quantitative prediction of these effects is currently not possible.
In conclusion, HMGA post-transcriptional regulation mechanisms appear to be much more complex than expected, and this field will certainly experience strong development in the future, with new miRNA and other ncRNA involved in HMGA regulation.
Funding
This research was funded by Associazione Italiana per la Ricerca sul Cancro (AIRC, IG18385) and Regione Friuli Venezia Giulia (TNBCneo and RiFT) to GM.
Conflicts of Interest
The authors declare no conflicts of interest.
References
- Friedmann, M.; Holth, L.T.; Zoghbi, H.Y.; Reeves, R. Organization, inducible-expression and chromosome localization of the human HMG-I(Y) nonhistone protein gene. Nucleic Acids Res. 1993, 21, 4259–4267. [Google Scholar] [CrossRef]
- Chau, K.Y.; Patel, U.A.; Lee, K.L.; Lam, H.Y.; Crane-Robinson, C. The gene for the human architectural transcription factor HMGI-C consists of five exons each coding for a distinct functional element. Nucleic Acids Res. 1995, 23, 4262–4266. [Google Scholar] [CrossRef] [PubMed]
- Reeves, R. Molecular biology of HMGA proteins: Hubs of nuclear function. Gene 2001, 277, 63–81. [Google Scholar] [CrossRef]
- Sgarra, R.; Tessari, M.A.; Di Bernardo, J.; Rustighi, A.; Zago, P.; Liberatori, S.; Armini, A.; Bini, L.; Giancotti, V.; Manfioletti, G. Discovering high mobility group A molecular partners in tumour cells. Proteomics 2005, 5, 1494–1506. [Google Scholar] [CrossRef] [PubMed]
- Sgarra, R.; Furlan, C.; Zammitti, S.; Lo Sardo, A.; Maurizio, E.; Di Bernardo, J.; Giancotti, V.; Manfioletti, G. Interaction proteomics of the HMGA chromatin architectural factors. Proteomics 2008, 8, 4721–4732. [Google Scholar] [CrossRef] [PubMed]
- Munshi, N.; Agalioti, T.; Lomvardas, S.; Merika, M.; Chen, G.; Thanos, D. Coordination of a transcriptional switch by HMGI(Y) acetylation. Science 2001, 293, 1133–1136. [Google Scholar] [CrossRef] [PubMed]
- Pegoraro, S.; Ros, G.; Piazza, S.; Sommaggio, R.; Ciani, Y.; Rosato, A.; Sgarra, R.; Del Sal, G.; Manfioletti, G. HMGA1 promotes metastatic processes in basal-like breast cancer regulating EMT and stemness. Oncotarget 2013, 4, 1293–1308. [Google Scholar] [CrossRef] [PubMed]
- Shah, S.N.; Cope, L.; Poh, W.; Belton, A.; Roy, S.; Talbot, C.C.; Sukumar, S.; Huso, D.L.; Resar, L.M.S. HMGA1: A master regulator of tumor progression in triple-negative breast cancer cells. PLoS ONE 2013, 8, e63419. [Google Scholar] [CrossRef]
- Shah, S.N.; Kerr, C.; Cope, L.; Zambidis, E.; Liu, C.; Hillion, J.; Belton, A.; Huso, D.L.; Resar, L.M.S. HMGA1 reprograms somatic cells into pluripotent stem cells by inducing stem cell transcriptional networks. PLoS ONE 2012, 7, e48533. [Google Scholar] [CrossRef]
- Catez, F.; Yang, H.; Tracey, K.J.; Reeves, R.; Misteli, T.; Bustin, M. Network of dynamic interactions between histone H1 and high-mobility-group proteins in chromatin. Mol. Cell. Biol. 2004, 24, 4321–4328. [Google Scholar] [CrossRef]
- Senigagliesi, B.; Penzo, C.; Severino, L.U.; Maraspini, R.; Petrosino, S.; Morales-Navarrete, H.; Pobega, E.; Ambrosetti, E.; Parisse, P.; Pegoraro, S.; et al. The High Mobility Group A1 (HMGA1) Chromatin Architectural Factor Modulates Nuclear Stiffness in Breast Cancer Cells. Int. J. Mol. Sci. 2019, 20, 2733. [Google Scholar] [CrossRef] [PubMed]
- Himes, S.R.; Reeves, R.; Attema, J.; Nissen, M.; Li, Y.; Shannon, M.F. The role of high-mobility group I(Y) proteins in expression of IL-2 and T cell proliferation. J. Immunol. 2000, 164, 3157–3168. [Google Scholar] [CrossRef] [PubMed]
- Battista, S.; Pentimalli, F.; Baldassarre, G.; Fedele, M.; Fidanza, V.; Croce, C.M.; Fusco, A. Loss of Hmga1 gene function affects embryonic stem cell lympho-hematopoietic differentiation. FASEB J. 2003, 17, 1496–1498. [Google Scholar] [CrossRef] [PubMed]
- Narita, M.; Narita, M.; Krizhanovsky, V.; Nuñez, S.; Chicas, A.; Hearn, S.A.; Myers, M.P.; Lowe, S.W. A novel role for high-mobility group a proteins in cellular senescence and heterochromatin formation. Cell 2006, 126, 503–514. [Google Scholar] [CrossRef] [PubMed]
- Diana, F.; Sgarra, R.; Manfioletti, G.; Rustighi, A.; Poletto, D.; Sciortino, M.T.; Mastino, A.; Giancotti, V. A link between apoptosis and degree of phosphorylation of high mobility group A1a protein in leukemic cells. J. Biol. Chem. 2001, 276, 11354–11361. [Google Scholar] [CrossRef] [PubMed]
- Sgarra, R.; Diana, F.; Rustighi, A.; Manfioletti, G.; Giancotti, V. Increase of HMGA1a protein methylation is a distinctive characteristic of leukaemic cells induced to undergo apoptosis. Cell Death Differ. 2003, 10, 386–389. [Google Scholar] [CrossRef]
- Schuldenfrei, A.; Belton, A.; Kowalski, J.; Talbot, C.C.; Di Cello, F.; Poh, W.; Tsai, H.-L.; Shah, S.N.; Huso, T.H.; Huso, D.L.; et al. HMGA1 drives stem cell, inflammatory pathway, and cell cycle progression genes during lymphoid tumorigenesis. BMC Genom. 2011, 12, 549. [Google Scholar] [CrossRef]
- Williams, M.D.; Zhang, X.; Belton, A.S.; Xian, L.; Huso, T.; Park, J.-J.; Siems, W.F.; Gang, D.R.; Resar, L.M.S.; Reeves, R.; et al. HMGA1 drives metabolic reprogramming of intestinal epithelium during hyperproliferation, polyposis, and colorectal carcinogenesis. J. Proteome Res. 2015, 14, 1420–1431. [Google Scholar] [CrossRef]
- Foti, D.; Chiefari, E.; Fedele, M.; Iuliano, R.; Brunetti, L.; Paonessa, F.; Manfioletti, G.; Barbetti, F.; Brunetti, A.; Croce, C.M.; et al. Lack of the architectural factor HMGA1 causes insulin resistance and diabetes in humans and mice. Nat. Med. 2005, 11, 765–773. [Google Scholar] [CrossRef]
- Conte, A.; Paladino, S.; Bianco, G.; Fasano, D.; Gerlini, R.; Tornincasa, M.; Renna, M.; Fusco, A.; Tramontano, D.; Pierantoni, G.M. High mobility group A1 protein modulates autophagy in cancer cells. Cell Death Differ. 2017, 24, 1948. [Google Scholar] [CrossRef]
- Thomae, A.W.; Pich, D.; Brocher, J.; Spindler, M.-P.; Berens, C.; Hock, R.; Hammerschmidt, W.; Schepers, A. Interaction between HMGA1a and the origin recognition complex creates site-specific replication origins. Proc. Natl. Acad. Sci. USA 2008, 105, 1692–1697. [Google Scholar] [CrossRef] [PubMed]
- Adair, J.E.; Kwon, Y.; Dement, G.A.; Smerdon, M.J.; Reeves, R. Inhibition of nucleotide excision repair by high mobility group protein HMGA1. J. Biol. Chem. 2005, 280, 32184–32192. [Google Scholar] [CrossRef] [PubMed]
- Li, A.Y.J.; Boo, L.M.; Wang, S.-Y.; Lin, H.H.; Wang, C.C.C.; Yen, Y.; Chen, B.P.C.; Chen, D.J.; Ann, D.K. Suppression of nonhomologous end joining repair by overexpression of HMGA2. Cancer Res. 2009, 69, 5699–5706. [Google Scholar] [CrossRef] [PubMed]
- Manabe, T.; Katayama, T.; Sato, N.; Gomi, F.; Hitomi, J.; Yanagita, T.; Kudo, T.; Honda, A.; Mori, Y.; Matsuzaki, S.; et al. Induced HMGA1a expression causes aberrant splicing of Presenilin-2 pre-mRNA in sporadic Alzheimer’s disease. Cell Death Differ. 2003, 10, 698–708. [Google Scholar] [CrossRef]
- Farnet, C.M.; Bushman, F.D. HIV-1 cDNA integration: Requirement of HMG I(Y) protein for function of preintegration complexes in vitro. Cell 1997, 88, 483–492. [Google Scholar] [CrossRef]
- Zhou, X.; Benson, K.F.; Ashar, H.R.; Chada, K. Mutation responsible for the mouse pygmy phenotype in the developmentally regulated factor HMGI-C. Nature 1995, 376, 771–774. [Google Scholar] [CrossRef]
- Chiappetta, G.; Avantaggiato, V.; Visconti, R.; Fedele, M.; Battista, S.; Trapasso, F.; Merciai, B.M.; Fidanza, V.; Giancotti, V.; Santoro, M.; et al. High level expression of the HMGI (Y) gene during embryonic development. Oncogene 1996, 13, 2439–2446. [Google Scholar]
- Anand, A.; Chada, K. In vivo modulation of Hmgic reduces obesity. Nat. Genet. 2000, 24, 377–380. [Google Scholar] [CrossRef]
- Federico, A.; Forzati, F.; Esposito, F.; Arra, C.; Palma, G.; Barbieri, A.; Palmieri, D.; Fedele, M.; Pierantoni, G.M.; De Martino, I.; et al. Hmga1/Hmga2 double knock-out mice display a “superpygmy” phenotype. Biol. Open 2014, 3, 372–378. [Google Scholar] [CrossRef]
- Fusco, A.; Fedele, M. Roles of HMGA proteins in cancer. Nat. Rev. Cancer 2007, 7, 899–910. [Google Scholar] [CrossRef]
- Sgarra, R.; Rustighi, A.; Tessari, M.A.; Di Bernardo, J.; Altamura, S.; Fusco, A.; Manfioletti, G.; Giancotti, V. Nuclear phosphoproteins HMGA and their relationship with chromatin structure and cancer. FEBS Lett. 2004, 574, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Resar, L.; Chia, L.; Xian, L. Lessons from the Crypt: HMGA1-Amping up Wnt for Stem Cells and Tumor Progression. Cancer Res. 2018, 78, 1890–1897. [Google Scholar] [CrossRef] [PubMed]
- Jones, M.R.; Goodarzi, M.O. Genetic determinants of polycystic ovary syndrome: Progress and future directions. Fertil. Steril. 2016, 106, 25–32. [Google Scholar] [CrossRef] [PubMed]
- De Rosa, S.; Chiefari, E.; Salerno, N.; Ventura, V.; D’Ascoli, G.L.; Arcidiacono, B.; Ambrosio, G.; Bilotta, F.L.; Torella, D.; Foti, D.; et al. HMGA1 is a novel candidate gene for myocardial infarction susceptibility. Int. J. Cardiol. 2017, 227, 331–334. [Google Scholar] [CrossRef] [PubMed]
- Camós, S.; Gubern, C.; Sobrado, M.; Rodríguez, R.; Romera, V.G.; Moro, M.A.; Lizasoain, I.; Serena, J.; Mallolas, J.; Castellanos, M. The high-mobility group I-Y transcription factor is involved in cerebral ischemia and modulates the expression of angiogenic proteins. Neuroscience 2014, 269, 112–130. [Google Scholar] [CrossRef] [PubMed]
- Pullinger, C.R.; Goldfine, I.D.; Tanyolaç, S.; Movsesyan, I.; Faynboym, M.; Durlach, V.; Chiefari, E.; Foti, D.P.; Frost, P.H.; Malloy, M.J.; et al. Evidence that an HMGA1 gene variant associates with type 2 diabetes, body mass index, and high-density lipoprotein cholesterol in a Hispanic-American population. Metab. Syndr. Relat. Disord. 2014, 12, 25–30. [Google Scholar] [CrossRef]
- Baron, R.M.; Kwon, M.-Y.; Castano, A.P.; Ghanta, S.; Riascos-Bernal, D.F.; Lopez-Guzman, S.; Macias, A.A.; Ith, B.; Schissel, S.L.; Lederer, J.A.; et al. Frontline Science: Targeted expression of a dominant-negative high mobility group A1 transgene improves outcome in sepsis. J. Leukoc. Biol. 2018, 104, 677–689. [Google Scholar] [CrossRef]
- Sgarra, R.; Pegoraro, S.; Ros, G.; Penzo, C.; Chiefari, E.; Foti, D.; Brunetti, A.; Manfioletti, G. High Mobility Group A (HMGA) proteins: Molecular instigators of breast cancer onset and progression. Biochim. Biophys. Acta Rev. Cancer 2018, 1869, 216–229. [Google Scholar] [CrossRef]
- Boque-Sastre, R.; Soler, M.; Oliveira-Mateos, C.; Portela, A.; Moutinho, C.; Sayols, S.; Villanueva, A.; Esteller, M.; Guil, S. Head-to-head antisense transcription and R-loop formation promotes transcriptional activation. Proc. Natl. Acad. Sci. USA 2015, 112, 5785–5790. [Google Scholar] [CrossRef]
- Tian, Y.; Zhang, N.; Chen, S.; Ma, Y.; Liu, Y. The long non-coding RNA LSINCT5 promotes malignancy in non-small cell lung cancer by stabilizing HMGA2. Cell Cycle 2018, 17, 1188–1198. [Google Scholar] [CrossRef]
- Zhang, Q.; Wang, Y. HMG modifications and nuclear function. Biochim. Biophys. Acta 2010, 1799, 28–36. [Google Scholar] [CrossRef]
- Schuster, S.L.; Hsieh, A.C. The Untranslated Regions of mRNAs in Cancer. Trends Cancer 2019, 5, 245–262. [Google Scholar] [CrossRef]
- Lujambio, A.; Lowe, S.W. The microcosmos of cancer. Nature 2012, 482, 347–355. [Google Scholar] [CrossRef]
- Mayr, C.; Hemann, M.T.; Bartel, D.P. Disrupting the pairing between let-7 and Hmga2 enhances oncogenic transformation. Science 2007, 315, 1576–1579. [Google Scholar] [CrossRef]
- Lee, Y.S.; Dutta, A. The tumor suppressor microRNA let-7 represses the HMGA2 oncogene. Genes Dev. 2007, 21, 1025–1030. [Google Scholar] [CrossRef]
- Shell, S.; Park, S.-M.; Radjabi, A.R.; Schickel, R.; Kistner, E.O.; Jewell, D.A.; Feig, C.; Lengyel, E.; Peter, M.E. Let-7 expression defines two differentiation stages of cancer. Proc. Natl. Acad. Sci. USA 2007, 104, 11400–11405. [Google Scholar] [CrossRef]
- Yan, L.; Zhou, J.; Gao, Y.; Ghazal, S.; Lu, L.; Bellone, S.; Yang, Y.; Liu, N.; Zhao, X.; Santin, A.D.; et al. Regulation of tumor cell migration and invasion by the H19/let-7 axis is antagonized by metformin-induced DNA methylation. Oncogene 2015, 34, 3076–3084. [Google Scholar] [CrossRef]
- D’Angelo, D.; Mussnich, P.; Sepe, R.; Raia, M.; Del Vecchio, L.; Cappabianca, P.; Pellecchia, S.; Petrosino, S.; Saggio, S.; Solari, D.; et al. RPSAP52 lncRNA is overexpressed in pituitary tumors and promotes cell proliferation by acting as miRNA sponge for HMGA proteins. J. Mol. Med. 2019, 97, 1019–1032. [Google Scholar] [CrossRef]
- Watanabe, S.; Ueda, Y.; Akaboshi, S.; Hino, Y.; Sekita, Y.; Nakao, M. HMGA2 maintains oncogenic RAS-induced epithelial-mesenchymal transition in human pancreatic cancer cells. Am. J. Pathol. 2009, 174, 854–868. [Google Scholar] [CrossRef]
- Palmieri, D.; D’Angelo, D.; Valentino, T.; De Martino, I.; Ferraro, A.; Wierinckx, A.; Fedele, M.; Trouillas, J.; Fusco, A. Downregulation of HMGA-targeting microRNAs has a critical role in human pituitary tumorigenesis. Oncogene 2012, 31, 3857–3865. [Google Scholar] [CrossRef]
- Liu, J.; Zhu, L.; Xie, G.; Bao, J.; Yu, Q. Let-7 miRNAs Modulate the Activation of NF-κB by Targeting TNFAIP3 and Are Involved in the Pathogenesis of Lupus Nephritis. PLoS ONE 2015, 10, e0121256. [Google Scholar] [CrossRef] [PubMed]
- Wang, T.; Zhu, H.; Yang, S.; Fei, X. Let-7a-5p may participate in the pathogenesis of diabetic nephropathy through targeting HMGA2. Mol. Med. Rep. 2019, 19, 4229–4237. [Google Scholar] [CrossRef] [PubMed]
- Schubert, M.; Spahn, M.; Kneitz, S.; Scholz, C.J.; Joniau, S.; Stroebel, P.; Riedmiller, H.; Kneitz, B. Distinct microRNA Expression Profile in Prostate Cancer Patients with Early Clinical Failure and the Impact of let-7 as Prognostic Marker in High-Risk Prostate Cancer. PLoS ONE 2013, 8, e65064. [Google Scholar] [CrossRef] [PubMed]
- Guo, L.; Chen, C.; Shi, M.; Wang, F.; Chen, X.; Diao, D.; Hu, M.; Yu, M.; Qian, L.; Guo, N. Stat3-coordinated Lin-28-let-7-HMGA2 and miR-200-ZEB1 circuits initiate and maintain oncostatin M-driven epithelial-mesenchymal transition. Oncogene 2013, 32, 5272–5282. [Google Scholar] [CrossRef]
- Alajez, N.M.; Shi, W.; Wong, D.; Lenarduzzi, M.; Waldron, J.; Weinreb, I.; Liu, F.-F. Lin28b promotes head and neck cancer progression via modulation of the insulin-like growth factor survival pathway. Oncotarget 2012, 3, 1641–1652. [Google Scholar] [CrossRef]
- Yu, K.-R.; Shin, J.-H.; Kim, J.-J.; Koog, M.G.; Lee, J.Y.; Choi, S.W.; Kim, H.-S.; Seo, Y.; Lee, S.; Shin, T.-H.; et al. Rapid and Efficient Direct Conversion of Human Adult Somatic Cells into Neural Stem Cells by HMGA2/let-7b. Cell Rep. 2015, 10, 441–452. [Google Scholar] [CrossRef] [PubMed]
- Chien, C.-S.; Wang, M.-L.; Chu, P.-Y.; Chang, Y.-L.; Liu, W.-H.; Yu, C.-C.; Lan, Y.-T.; Huang, P.-I.; Lee, Y.-Y.; Chen, Y.-W.; et al. Lin28B/Let-7 Regulates Expression of Oct4 and Sox2 and Reprograms Oral Squamous Cell Carcinoma Cells to a Stem-like State. Cancer Res. 2015, 75, 2553–2565. [Google Scholar] [CrossRef]
- Ma, J.; Li, D.; Kong, F.-F.; Yang, D.; Yang, H.; Ma, X.-X. miR-302a-5p/367-3p-HMGA2 axis regulates malignant processes during endometrial cancer development. J. Exp. Clin. Cancer Res. 2018, 37, 19. [Google Scholar] [CrossRef]
- Park, S.-M.; Shell, S.; Radjabi, A.R.; Schickel, R.; Feig, C.; Boyerinas, B.; Dinulescu, D.M.; Lengyel, E.; Peter, M.E. Let-7 prevents early cancer progression by suppressing expression of the embryonic gene HMGA2. Cell Cycle 2007, 6, 2585–2590. [Google Scholar] [CrossRef]
- Pandit, K.V.; Corcoran, D.; Yousef, H.; Yarlagadda, M.; Tzouvelekis, A.; Gibson, K.F.; Konishi, K.; Yousem, S.A.; Singh, M.; Handley, D.; et al. Inhibition and role of let-7d in idiopathic pulmonary fibrosis. Am. J. Respir. Crit. Care Med. 2010, 182, 220–229. [Google Scholar] [CrossRef]
- Chen, Y.-N.; Ren, C.-C.; Yang, L.; Nai, M.-M.; Xu, Y.-M.; Zhang, F.; Liu, Y. MicroRNA let-7d-5p rescues ovarian cancer cell apoptosis and restores chemosensitivity by regulating the p53 signaling pathway via HMGA1. Int. J. Oncol. 2019, 54, 1771–1784. [Google Scholar] [CrossRef] [PubMed]
- D’Angelo, D.; Palmieri, D.; Mussnich, P.; Roche, M.; Wierinckx, A.; Raverot, G.; Fedele, M.; Croce, C.M.; Trouillas, J.; Fusco, A. Altered microRNA expression profile in human pituitary GH adenomas: Down-regulation of miRNA targeting HMGA1, HMGA2, and E2F1. J. Clin. Endocrinol. Metab. 2012, 97, E1128–E1138. [Google Scholar] [CrossRef] [PubMed]
- Kumar, M.S.; Erkeland, S.J.; Pester, R.E.; Chen, C.Y.; Ebert, M.S.; Sharp, P.A.; Jacks, T. Suppression of non-small cell lung tumor development by the let-7 microRNA family. Proc. Natl. Acad. Sci. USA 2008, 105, 3903–3908. [Google Scholar] [CrossRef]
- Boyerinas, B.; Park, S.-M.; Shomron, N.; Hedegaard, M.M.; Vinther, J.; Andersen, J.S.; Feig, C.; Xu, J.; Burge, C.B.; Peter, M.E. Identification of let-7-regulated oncofetal genes. Cancer Res. 2008, 68, 2587–2591. [Google Scholar] [CrossRef]
- Wang, T.; Zhang, X.; Obijuru, L.; Laser, J.; Aris, V.; Lee, P.; Mittal, K.; Soteropoulos, P.; Wei, J.-J. A micro-RNA signature associated with race, tumor size, and target gene activity in human uterine leiomyomas. Genes Chromosomes Cancer 2007, 46, 336–347. [Google Scholar] [CrossRef]
- Peng, Y.; Laser, J.; Shi, G.; Mittal, K.; Melamed, J.; Lee, P.; Wei, J.-J. Antiproliferative effects by Let-7 repression of high-mobility group A2 in uterine leiomyoma. Mol. Cancer Res. 2008, 6, 663–673. [Google Scholar] [CrossRef]
- Wu, J.; Liu, Z.; Shao, C.; Gong, Y.; Hernando, E.; Lee, P.; Narita, M.; Muller, W.; Liu, J.; Wei, J.-J. HMGA2 overexpression-induced ovarian surface epithelial transformation is mediated through regulation of EMT genes. Cancer Res. 2011, 71, 349–359. [Google Scholar] [CrossRef]
- Mansoori, B.; Mohammadi, A.; Naghizadeh, S.; Gjerstorff, M.; Shanehbandi, D.; Shirjang, S.; Najafi, S.; Holmskov, U.; Khaze, V.; Duijf, P.H.G.; et al. miR-330 suppresses EMT and induces apoptosis by downregulating HMGA2 in human colorectal cancer. J. Cell. Physiol. 2019. [Google Scholar] [CrossRef]
- Yu, F.; Yao, H.; Zhu, P.; Zhang, X.; Pan, Q.; Gong, C.; Huang, Y.; Hu, X.; Su, F.; Lieberman, J.; et al. let-7 regulates self renewal and tumorigenicity of breast cancer cells. Cell 2007, 131, 1109–1123. [Google Scholar] [CrossRef]
- Motoyama, K.; Inoue, H.; Nakamura, Y.; Uetake, H.; Sugihara, K.; Mori, M. Clinical significance of high mobility group A2 in human gastric cancer and its relationship to let-7 microRNA family. Clin. Cancer Res. 2008, 14, 2334–2340. [Google Scholar] [CrossRef]
- Wei, J.; Li, H.; Wang, S.; Li, T.; Fan, J.; Liang, X.; Li, J.; Han, Q.; Zhu, L.; Fan, L.; et al. let-7 enhances osteogenesis and bone formation while repressing adipogenesis of human stromal/mesenchymal stem cells by regulating HMGA2. Stem Cells Dev. 2014, 23, 1452–1463. [Google Scholar] [CrossRef]
- Wu, A.; Wu, K.; Li, J.; Mo, Y.; Lin, Y.; Wang, Y.; Shen, X.; Li, S.; Li, L.; Yang, Z. Let-7a inhibits migration, invasion and epithelial-mesenchymal transition by targeting HMGA2 in nasopharyngeal carcinoma. J. Transl. Med. 2015, 13, 105. [Google Scholar] [CrossRef]
- Halle, B.; Marcusson, E.G.; Aaberg-Jessen, C.; Jensen, S.S.; Meyer, M.; Schulz, M.K.; Andersen, C.; Kristensen, B.W. Convection-enhanced delivery of an anti-miR is well-tolerated, preserves anti-miR stability and causes efficient target de-repression: A proof of concept. J. Neurooncol. 2016, 126, 47–55. [Google Scholar] [CrossRef]
- Wong, L.L.; Saw, E.L.; Lim, J.Y.; Zhou, Y.; Richards, A.M.; Wang, P. MicroRNA Let-7d-3p Contributes to Cardiac Protection via Targeting HMGA2. Int. J. Mol. Sci. 2019, 20, 1522. [Google Scholar] [CrossRef]
- Qin, M.-M.; Chai, X.; Huang, H.-B.; Feng, G.; Li, X.-N.; Zhang, J.; Zheng, R.; Liu, X.-C.; Pu, C. let-7i inhibits proliferation and migration of bladder cancer cells by targeting HMGA1. BMC Urol. 2019, 19, 53. [Google Scholar] [CrossRef]
- Cui, H.; Song, R.; Wu, J.; Wang, W.; Chen, X.; Yin, J. MicroRNA-337 regulates the PI3K/AKT and Wnt/β-catenin signaling pathways to inhibit hepatocellular carcinoma progression by targeting high-mobility group AT-hook 2. Am. J. Cancer Res. 2018, 8, 405–421. [Google Scholar]
- Emmrich, S.; Katsman-Kuipers, J.E.; Henke, K.; Khatib, M.E.; Jammal, R.; Engeland, F.; Dasci, F.; Zwaan, C.M.; den Boer, M.L.; Verboon, L.; et al. miR-9 is a tumor suppressor in pediatric AML with t(8;21). Leukemia 2014, 28, 1022–1032. [Google Scholar] [CrossRef]
- Xu, X.; Zou, H.; Luo, L.; Wang, X.; Wang, G. MicroRNA-9 exerts antitumor effects on hepatocellular carcinoma progression by targeting HMGA2. FEBS Open Bio 2019, 9, 1784–1797. [Google Scholar] [CrossRef]
- Zhu, S.; Deng, S.; Ma, Q.; Zhang, T.; Jia, C.; Zhuo, D.; Yang, F.; Wei, J.; Wang, L.; Dykxhoorn, D.M.; et al. MicroRNA-10A* and MicroRNA-21 modulate endothelial progenitor cell senescence via suppressing high-mobility group A2. Circ. Res. 2013, 112, 152–164. [Google Scholar] [CrossRef]
- Yang, Y.; Jiang, H.; Xiao, L.; Yang, X. MicroRNA-33b-5p is overexpressed and inhibits GLUT4 by targeting HMGA2 in polycystic ovarian syndrome: An in vivo and in vitro study. Oncol. Rep. 2018, 39, 3073–3085. [Google Scholar] [CrossRef]
- Wang, J.; Liang, H.; Ge, H.; Guo, X.; Gu, D.; Yuan, Y. MicroRNA-363-3p inhibits hepatocarcinogenesis by targeting HMGA2 and is associated with liver cancer stage. Mol. Med. Rep. 2019, 19, 935–942. [Google Scholar] [CrossRef]
- Qi, J.; Rice, S.J.; Salzberg, A.C.; Runkle, E.A.; Liao, J.; Zander, D.S.; Mu, D. MiR-365 regulates lung cancer and developmental gene thyroid transcription factor 1. Cell Cycle 2012, 11, 177–186. [Google Scholar] [CrossRef]
- Niu, Y.; Zhou, H.; Liu, Y.; Wang, Y.; Xie, J.; Feng, C.; An, N. miR-16 regulates proliferation and apoptosis of pituitary adenoma cells by inhibiting HMGA2. Oncol. Lett. 2019, 17, 2491–2497. [Google Scholar] [CrossRef]
- Kaddar, T.; Rouault, J.-P.; Chien, W.W.; Chebel, A.; Gadoux, M.; Salles, G.; Ffrench, M.; Magaud, J.-P. Two new miR-16 targets: Caprin-1 and HMGA1, proteins implicated in cell proliferation. Biol. Cell 2009, 101, 511–524. [Google Scholar] [CrossRef]
- Liu, H.; Wang, X.; Liu, S.; Li, H.; Yuan, X.; Feng, B.; Bai, H.; Zhao, B.; Chu, Y.; Li, H. Effects and mechanism of miR-23b on glucose-mediated epithelial-to-mesenchymal transition in diabetic nephropathy. Int. J. Biochem. Cell Biol. 2016, 70, 149–160. [Google Scholar] [CrossRef]
- Leone, V.; Langella, C.; D’Angelo, D.; Mussnich, P.; Wierinckx, A.; Terracciano, L.; Raverot, G.; Lachuer, J.; Rotondi, S.; Jaffrain-Rea, M.-L.; et al. Mir-23b and miR-130b expression is downregulated in pituitary adenomas. Mol. Cell. Endocrinol. 2014, 390, 1–7. [Google Scholar] [CrossRef]
- Kong, B.; Qin, Z.; Ye, Z.; Yang, X.; Li, L.; Su, Q. microRNA-26a-5p affects myocardial injury induced by coronary microembolization by modulating HMGA1. J. Cell. Biochem. 2019, 120, 10756–10766. [Google Scholar] [CrossRef]
- Chen, C.-Y.A.; Chang, J.T.; Ho, Y.-F.; Shyu, A.-B. MiR-26 down-regulates TNF-α/NF-κB signalling and IL-6 expression by silencing HMGA1 and MALT1. Nucleic Acids Res. 2016, 44, 3772–3787. [Google Scholar] [CrossRef]
- Liu, J.; Mi, B.; Wang, Y.; Shi, C.; Mi, X.; Lu, Y.; Yu, P. miR-26a suppresses osteosarcoma migration and invasion by directly targeting HMGA1. Oncol. Lett. 2018, 15, 8303–8310. [Google Scholar] [CrossRef]
- Sekimoto, N.; Suzuki, A.; Suzuki, Y.; Sugano, S. Expression of miR-26a exhibits a negative correlation with HMGA1 and regulates cancer progression by targeting HMGA1 in lung adenocarcinoma cells. Mol. Med. Rep. 2017, 15, 534–542. [Google Scholar] [CrossRef]
- Yang, Y.; Zhang, P.; Zhao, Y.; Yang, J.; Jiang, G.; Fan, J. Decreased MicroRNA-26a expression causes cisplatin resistance in human non-small cell lung cancer. Cancer Biol. Ther. 2016, 17, 515–525. [Google Scholar] [CrossRef]
- Zhao, X.-X.; Yuan, Q.-Z.; Mu, D.-P.; Sun, D.-W.; Bo, Q.-A.; Pan, G.-Z.; Li, G.-Q.; Cui, T.; Ding, P.-P.; You, F.-P.; et al. MicroRNA-26a inhibits proliferation by targeting high mobility group AT-hook 1 in breast cancer. Int. J. Clin. Exp. Pathol. 2015, 8, 368–373. [Google Scholar]
- Liang, H.; Gu, Y.; Li, T.; Zhang, Y.; Huangfu, L.; Hu, M.; Zhao, D.; Chen, Y.; Liu, S.; Dong, Y.; et al. Integrated analyses identify the involvement of microRNA-26a in epithelial-mesenchymal transition during idiopathic pulmonary fibrosis. Cell Death Dis. 2014, 5, e1238. [Google Scholar] [CrossRef]
- Zhou, H.; Guo, W.; Zhao, Y.; Wang, Y.; Zha, R.; Ding, J.; Liang, L.; Hu, J.; Shen, H.; Chen, Z.; et al. MicroRNA-26a acts as a tumor suppressor inhibiting gallbladder cancer cell proliferation by directly targeting HMGA2. Int. J. Oncol. 2014, 44, 2050–2058. [Google Scholar] [CrossRef]
- Lin, Y.; Chen, H.; Hu, Z.; Mao, Y.; Xu, X.; Zhu, Y.; Xu, X.; Wu, J.; Li, S.; Mao, Q.; et al. miR-26a inhibits proliferation and motility in bladder cancer by targeting HMGA1. FEBS Lett. 2013, 587, 2467–2473. [Google Scholar] [CrossRef]
- Li, W.; Yuan, Y.; Huang, L.; Qiao, M.; Zhang, Y. Metformin alters the expression profiles of microRNAs in human pancreatic cancer cells. Diabetes Res. Clin. Pract. 2012, 96, 187–195. [Google Scholar] [CrossRef]
- Baba, O.; Horie, T.; Nakao, T.; Hakuno, D.; Nakashima, Y.; Nishi, H.; Kuwabara, Y.; Nishiga, M.; Nishino, T.; Ide, Y.; et al. MicroRNA 33 Regulates the Population of Peripheral Inflammatory Ly6Chigh Monocytes through Dual Pathways. Mol. Cell. Biol. 2018, 38, e00604-17. [Google Scholar] [CrossRef]
- Wang, H.; Sun, Z.; Wang, Y.; Hu, Z.; Zhou, H.; Zhang, L.; Hong, B.; Zhang, S.; Cao, X. miR-33-5p, a novel mechano-sensitive microRNA promotes osteoblast differentiation by targeting Hmga2. Sci. Rep. 2016, 6, 23170. [Google Scholar] [CrossRef]
- Rice, S.J.; Lai, S.-C.; Wood, L.W.; Helsley, K.R.; Runkle, E.A.; Winslow, M.M.; Mu, D. MicroRNA-33a mediates the regulation of high mobility group AT-hook 2 gene (HMGA2) by thyroid transcription factor 1 (TTF-1/NKX2-1). J. Biol. Chem. 2013, 288, 16348–16360. [Google Scholar] [CrossRef]
- Price, N.L.; Holtrup, B.; Kwei, S.L.; Wabitsch, M.; Rodeheffer, M.; Bianchini, L.; Suárez, Y.; Fernández-Hernando, C. SREBP-1c/MicroRNA 33b Genomic Loci Control Adipocyte Differentiation. Mol. Cell. Biol. 2016, 36, 1180–1193. [Google Scholar] [CrossRef]
- Lin, Y.; Liu, A.Y.; Fan, C.; Zheng, H.; Li, Y.; Zhang, C.; Wu, S.; Yu, D.; Huang, Z.; Liu, F.; et al. MicroRNA-33b Inhibits Breast Cancer Metastasis by Targeting HMGA2, SALL4 and Twist1. Sci. Rep. 2015, 5, 9995. [Google Scholar] [CrossRef]
- Zhang, P.; Huang, C.; Fu, C.; Tian, Y.; Hu, Y.; Wang, B.; Strasner, A.; Song, Y.; Song, E. Cordycepin (3′-deoxyadenosine) suppressed HMGA2, Twist1 and ZEB1-dependent melanoma invasion and metastasis by targeting miR-33b. Oncotarget 2015, 6, 9834–9853. [Google Scholar] [CrossRef]
- Zhang, P.; Bai, H.; Liu, G.; Wang, H.; Chen, F.; Zhang, B.; Zeng, P.; Wu, C.; Peng, C.; Huang, C.; et al. MicroRNA-33b, upregulated by EF24, a curcumin analog, suppresses the epithelial-to-mesenchymal transition (EMT) and migratory potential of melanoma cells by targeting HMGA2. Toxicol. Lett. 2015, 234, 151–161. [Google Scholar] [CrossRef]
- Yang, X.; Zhao, Q.; Yin, H.; Lei, X.; Gan, R. MiR-33b-5p sensitizes gastric cancer cells to chemotherapy drugs via inhibiting HMGA2 expression. J. Drug Target. 2017, 25, 653–660. [Google Scholar] [CrossRef]
- Ma, S.; Fu, T.; Zhao, S.; Gao, M. MicroRNA-34a-5p suppresses tumorigenesis and progression of glioma and potentiates Temozolomide-induced cytotoxicity for glioma cells by targeting HMGA2. Eur. J. Pharmacol. 2019, 852, 42–50. [Google Scholar] [CrossRef]
- Ji, Q.; Hao, X.; Meng, Y.; Zhang, M.; Desano, J.; Fan, D.; Xu, L. Restoration of tumor suppressor miR-34 inhibits human p53-mutant gastric cancer tumorspheres. BMC Cancer 2008, 8, 266. [Google Scholar] [CrossRef]
- Li, W.; Wang, J.; Zhang, D.; Zhang, X.; Xu, J.; Zhao, L. MicroRNA-98 targets HMGA2 to inhibit the development of retinoblastoma through mediating Wnt/β-catenin pathway. Cancer Biomark. 2019, 25, 79–88. [Google Scholar] [CrossRef]
- Wang, M.-J.; Zhang, H.; Li, J.; Zhao, H.-D. microRNA-98 inhibits the proliferation, invasion, migration and promotes apoptosis of breast cancer cells by binding to HMGA2. Biosci. Rep. 2018, 38, BSR20180571. [Google Scholar] [CrossRef]
- Gao, X.-L.; Cao, M.-G.; Ai, G.-G.; Hu, Y.-B. Mir-98 reduces the expression of HMGA2 and promotes osteogenic differentiation of mesenchymal stem cells. Eur. Rev. Med. Pharmacol. Sci. 2018, 22, 3311–3317. [Google Scholar]
- Chen, Z.; Cheng, Q.; Ma, Z.; Xi, H.; Peng, R.; Jiang, B. Overexpression of RKIP inhibits cell invasion in glioma cell lines through upregulation of miR-98. BioMed Res. Int. 2013, 2013, 695179. [Google Scholar] [CrossRef]
- Zhu, M.; Zhang, C.; Chen, D.; Chen, S.; Zheng, H. MicroRNA-98-HMGA2-POSTN signal pathway reverses epithelial-to-mesenchymal transition in laryngeal squamous cell carcinoma. Biomed. Pharmacother. 2019, 117, 108998. [Google Scholar] [CrossRef]
- Zhang, Y.; Su, Z.; An, L.-J.; Li, L.; Wei, M.; Ge, D.-J.; Liu, H.-L. miR-98 acts as an inhibitor in chronic constriction injury-induced neuropathic pain via downregulation of high-mobility group AT-hook 2. J. Cell. Biochem. 2019, 120, 10363–10369. [Google Scholar] [CrossRef]
- Zheng, F.; Wang, F.; Xu, Z. MicroRNA-98-5p prevents bone regeneration by targeting high mobility group AT-Hook 2. Exp. Ther. Med. 2019, 18, 2660–2666. [Google Scholar] [CrossRef]
- Zhu, J.; Lin, X.; Yan, C.; Yang, S.; Zhu, Z. microRNA-98 protects sepsis mice from cardiac dysfunction, liver and lung injury by negatively regulating HMGA2 through inhibiting NF-κB signaling pathway. Cell Cycle 2019, 18, 1948–1964. [Google Scholar] [CrossRef]
- Qiu, K.; Xie, Q.; Jiang, S.; Lin, T. miR-98-5p promotes apoptosis and inhibits migration and cell growth in papillary thyroid carcinoma through Bax/Caspase-3 by HMGA2. J. Clin. Lab. Anal. 2019, e23044. [Google Scholar] [CrossRef]
- Jiang, W.; Gu, W.; Qiu, R.; He, S.; Shen, C.; Wu, Y.; Zhang, J.; Zhou, J.; Guo, Y.; Wan, D.; et al. miRNA-101 Suppresses Epithelial-to-Mesenchymal Transition by Targeting HMGA2 in Pancreatic Cancer Cells. Anticancer Agents Med. Chem. 2016, 16, 432–439. [Google Scholar] [CrossRef]
- Chen, Z.; Li, Q.; Wang, S.; Zhang, J. miR-485-5p inhibits bladder cancer metastasis by targeting HMGA2. Int. J. Mol. Med. 2015, 36, 1136–1142. [Google Scholar] [CrossRef]
- He, Q.-Y.; Wang, G.-C.; Zhang, H.; Tong, D.-K.; Ding, C.; Liu, K.; Ji, F.; Zhu, X.; Yang, S. miR-106a-5p Suppresses the Proliferation, Migration, and Invasion of Osteosarcoma Cells by Targeting HMGA2. DNA Cell Biol. 2016, 35, 506–520. [Google Scholar] [CrossRef]
- Wang, Y.; Chen, F.; Zhao, M.; Yang, Z.; Zhang, S.; Ye, L.; Gao, H.; Zhang, X. MiR-107 suppresses proliferation of hepatoma cells through targeting HMGA2 mRNA 3′UTR. Biochem. Biophys. Res. Commun. 2016, 480, 455–460. [Google Scholar] [CrossRef]
- Liu, W.; Xu, G.; Liu, H.; Li, T. MicroRNA-490-3p regulates cell proliferation and apoptosis by targeting HMGA2 in osteosarcoma. FEBS Lett. 2015, 589, 3148–3153. [Google Scholar] [CrossRef]
- Zhang, F.; Wu, A.; Wang, Y.; Liu, J. miR-490-3p functions as a tumor suppressor in glioma by inhibiting high-mobility group AT-hook 2 expression. Exp. Ther. Med. 2019, 18, 664–670. [Google Scholar] [CrossRef]
- Lau, K.-M.; Chan, Q.K.Y.; Pang, J.C.S.; Ma, F.M.-T.; Li, K.K.W.; Yeung, W.W.; Cheng, A.S.L.; Feng, H.; Chung, N.Y.F.; Li, H.-M.; et al. Overexpression of HMGA1 deregulates tumor growth via cdc25A and alters migration/invasion through a cdc25A-independent pathway in medulloblastoma. Acta Neuropathol. 2012, 123, 553–571. [Google Scholar] [CrossRef]
- Liu, Z.; Lü, Y.; Jiang, Q.; Yang, Y.; Dang, C.; Sun, R. miR-491 inhibits BGC-823 cell migration via targeting HMGA2. Int. J. Biol. Markers 2019, 34, 364–372. [Google Scholar] [CrossRef]
- Mei, L.-L.; Wang, W.-J.; Qiu, Y.-T.; Xie, X.-F.; Bai, J.; Shi, Z.-Z. miR-125b-5p functions as a tumor suppressor gene partially by regulating HMGA2 in esophageal squamous cell carcinoma. PLoS ONE 2017, 12, e0185636. [Google Scholar] [CrossRef]
- Sun, B.; Zhang, Y.; Zhou, L.; Yin, L.; Li, F.; Li, C.; Xia, J. The proliferation of cervical cancer is promoted by miRNA-125b through the regulation of the HMGA1. Onco Targets Ther. 2019, 12, 2767–2776. [Google Scholar] [CrossRef]
- Kleemann, M.; Schneider, H.; Unger, K.; Bereuther, J.; Fischer, S.; Sander, P.; Marion Schneider, E.; Fischer-Posovszky, P.; Riedel, C.U.; Handrick, R.; et al. Induction of apoptosis in ovarian cancer cells by miR-493-3p directly targeting AKT2, STK38L, HMGA2, ETS1 and E2F5. Cell. Mol. Life Sci. 2019, 76, 539–559. [Google Scholar] [CrossRef]
- Tian, Z.; Zhou, H.; Xu, Y.; Bai, J. MicroRNA-495 Inhibits New Bone Regeneration via Targeting High Mobility Group AT-Hook 2 (HMGA2). Med. Sci. Monit. 2017, 23, 4689–4698. [Google Scholar] [CrossRef]
- Wang, H.; Jiang, Z.; Chen, H.; Wu, X.; Xiang, J.; Peng, J. MicroRNA-495 Inhibits Gastric Cancer Cell Migration and Invasion Possibly via Targeting High Mobility Group AT-Hook 2 (HMGA2). Med. Sci. Monit. 2017, 23, 640–648. [Google Scholar] [CrossRef]
- Qiu, H.; Zhong, J.; Luo, L.; Tang, Z.; Liu, N.; Kang, K.; Li, L.; Gou, D. Regulatory Axis of miR-195/497 and HMGA1-Id3 Governs Muscle Cell Proliferation and Differentiation. Int. J. Biol. Sci. 2017, 13, 157–166. [Google Scholar] [CrossRef][Green Version]
- Yang, Q.; Wang, X.; Tang, C.; Chen, X.; He, J. H19 promotes the migration and invasion of colon cancer by sponging miR-138 to upregulate the expression of HMGA1. Int. J. Oncol. 2017, 50, 1801–1809. [Google Scholar] [CrossRef]
- Li, W.; Li, J.; Mu, H.; Guo, M.; Deng, H. MiR-503 suppresses cell proliferation and invasion of gastric cancer by targeting HMGA2 and inactivating WNT signaling pathway. Cancer Cell Int. 2019, 19, 164. [Google Scholar] [CrossRef]
- Xia, C.; Liang, S.; He, Z.; Zhu, X.; Chen, R.; Chen, J. Metformin, a first-line drug for type 2 diabetes mellitus, disrupts the MALAT1/miR-142-3p sponge to decrease invasion and migration in cervical cancer cells. Eur. J. Pharmacol. 2018, 830, 59–67. [Google Scholar] [CrossRef]
- Chiou, G.-Y.; Chien, C.-S.; Wang, M.-L.; Chen, M.-T.; Yang, Y.-P.; Yu, Y.-L.; Chien, Y.; Chang, Y.-C.; Shen, C.-C.; Chio, C.-C.; et al. Epigenetic regulation of the miR142-3p/interleukin-6 circuit in glioblastoma. Mol. Cell 2013, 52, 693–706. [Google Scholar] [CrossRef]
- Xu, G.; Wang, J.; Jia, Y.; Shen, F.; Han, W.; Kang, Y. MiR-142-3p functions as a potential tumor suppressor in human osteosarcoma by targeting HMGA1. Cell. Physiol. Biochem. 2014, 33, 1329–1339. [Google Scholar] [CrossRef]
- Kim, T.H.; Song, J.-Y.; Park, H.; Jeong, J.-Y.; Kwon, A.-Y.; Heo, J.H.; Kang, H.; Kim, G.; An, H.J. miR-145, targeting high-mobility group A2, is a powerful predictor of patient outcome in ovarian carcinoma. Cancer Lett. 2015, 356, 937–945. [Google Scholar] [CrossRef]
- Khanna, N.; Ge, Y.; Chen, J. MicroRNA-146b promotes myogenic differentiation and modulates multiple gene targets in muscle cells. PLoS ONE 2014, 9, e100657. [Google Scholar] [CrossRef]
- Ye, Z.-H.; Gui, D.-W. miR-539 suppresses proliferation and induces apoptosis in renal cell carcinoma by targeting high mobility group A2. Mol. Med. Rep. 2018, 17, 5611–5618. [Google Scholar] [CrossRef]
- Xu, L.; Du, B.; Lu, Q.-J.; Fan, X.-W.; Tang, K.; Yang, L.; Liao, W.-L. miR-541 suppresses proliferation and invasion of squamous cell lung carcinoma cell lines via directly targeting high-mobility group AT-hook 2. Cancer Med. 2018, 7, 2581–2591. [Google Scholar] [CrossRef]
- Zhang, Z.-C.; Wang, G.-P.; Yin, L.-M.; Li, M.; Wu, L.-L. Increasing miR-150 and lowering HMGA2 inhibit proliferation and cycle progression of colon cancer in SW480 cells. Eur. Rev. Med. Pharmacol. Sci. 2018, 22, 6793–6800. [Google Scholar]
- Tang, W.; Xu, P.; Wang, H.; Niu, Z.; Zhu, D.; Lin, Q.; Tang, L.; Ren, L. MicroRNA-150 suppresses triple-negative breast cancer metastasis through targeting HMGA2. Onco Targets Ther. 2018, 11, 2319–2332. [Google Scholar] [CrossRef]
- Dai, F.-Q.; Li, C.-R.; Fan, X.-Q.; Tan, L.; Wang, R.-T.; Jin, H. miR-150-5p Inhibits Non-Small-Cell Lung Cancer Metastasis and Recurrence by Targeting HMGA2 and β-Catenin Signaling. Mol. Ther. Nucleic Acids 2019, 16, 675–685. [Google Scholar] [CrossRef]
- Fan, C.; Lin, Y.; Mao, Y.; Huang, Z.; Liu, A.Y.; Ma, H.; Yu, D.; Maitikabili, A.; Xiao, H.; Zhang, C.; et al. MicroRNA-543 suppresses colorectal cancer growth and metastasis by targeting KRAS, MTA1 and HMGA2. Oncotarget 2016, 7, 21825–21839. [Google Scholar] [CrossRef]
- Zhu, C.; Li, J.; Cheng, G.; Zhou, H.; Tao, L.; Cai, H.; Li, P.; Cao, Q.; Ju, X.; Meng, X.; et al. miR-154 inhibits EMT by targeting HMGA2 in prostate cancer cells. Mol. Cell. Biochem. 2013, 379, 69–75. [Google Scholar] [CrossRef]
- Zou, Q.; Wu, H.; Fu, F.; Yi, W.; Pei, L.; Zhou, M. RKIP suppresses the proliferation and metastasis of breast cancer cell lines through up-regulation of miR-185 targeting HMGA2. Arch. Biochem. Biophys. 2016, 610, 25–32. [Google Scholar] [CrossRef]
- Belkaya, S.; Murray, S.E.; Eitson, J.L.; de la Morena, M.T.; Forman, J.A.; van Oers, N.S.C. Transgenic expression of microRNA-185 causes a developmental arrest of T cells by targeting multiple genes including Mzb1. J. Biol. Chem. 2013, 288, 30752–30762. [Google Scholar] [CrossRef]
- Wang, Y.; Chen, F.; Zhao, M.; Yang, Z.; Li, J.; Zhang, S.; Zhang, W.; Ye, L.; Zhang, X. The long noncoding RNA HULC promotes liver cancer by increasing the expression of the HMGA2 oncogene via sequestration of the microRNA-186. J. Biol. Chem. 2017, 292, 15395–15407. [Google Scholar] [CrossRef]
- Zhao, H.; Zhao, H.; Xia, X.; Liu, X. MicroRNA-599 targets high-mobility group AT-hook 2 to inhibit cell proliferation and invasion in clear cell renal carcinoma. Mol. Med. Rep. 2018, 17, 7451–7459. [Google Scholar] [CrossRef]
- Zhou, W.; Zhong, C.; Luo, X.; Zhang, Y.; Zhang, G.; Zhou, D.; Liu, L. miR-625 suppresses cell proliferation and migration by targeting HMGA1 in breast cancer. Biochem. Biophys. Res. Commun. 2016, 470, 838–844. [Google Scholar] [CrossRef]
- Gao, X.; Dai, M.; Li, Q.; Wang, Z.; Lu, Y.; Song, Z. HMGA2 regulates lung cancer proliferation and metastasis. Thorac. Cancer 2017, 8, 501–510. [Google Scholar] [CrossRef]
- Li, Y.; Wu, D.; Wang, P.; Li, X.; Shi, G. miR-195 Regulates Proliferation and Apoptosis through Inhibiting the mTOR/p70s6k Signaling Pathway by Targeting HMGA2 in Esophageal Carcinoma Cells. Dis. Markers 2017, 2017, 8317913. [Google Scholar] [CrossRef]
- Zhang, X.; Tao, T.; Liu, C.; Guan, H.; Huang, Y.; Xu, B.; Chen, M. Downregulation of miR-195 promotes prostate cancer progression by targeting HMGA1. Oncol. Rep. 2016, 36, 376–382. [Google Scholar] [CrossRef]
- You, X.-Y.; Huang, J.-H.; Liu, B.; Liu, S.-J.; Zhong, Y.; Liu, S.-M. HMGA1 is a new target of miR-195 involving isoprenaline-induced cardiomyocyte hypertrophy. Biochemistry (Moscow) 2014, 79, 538–544. [Google Scholar] [CrossRef]
- Hu, J.; Shen, W. microRNA-196a attenuates ischemic brain injury in rats by directly targeting high mobility group A1. Exp. Ther. Med. 2019, 17, 1579–1586. [Google Scholar] [CrossRef]
- Zhang, D.-L.; Liu, X.; Wang, Q.; Li, N.; Wu, S.-H.; Wang, C. Downregulation of microRNA-196a attenuates ischemic brain injury in rats by directly targeting HMGA1. Eur. Rev. Med. Pharmacol. Sci. 2019, 23, 740–748. [Google Scholar]
- Huang, W.; Li, J.; Guo, X.; Zhao, Y.; Yuan, X. miR-663a inhibits hepatocellular carcinoma cell proliferation and invasion by targeting HMGA2. Biomed. Pharmacother. 2016, 81, 431–438. [Google Scholar] [CrossRef]
- Wang, R.; Shen, J.; Wang, Q.; Zhang, M. Bortezomib inhibited the progression of diffuse large B-cell lymphoma via targeting miR-198. Biomed. Pharmacother. 2018, 108, 43–49. [Google Scholar] [CrossRef]
- Zhang, Y.; Liu, Y.; Wu, M.; Wang, H.; Wu, L.; Xu, B.; Zhou, W.; Fan, X.; Shao, J.; Yang, T. MicroRNA-664a-5p promotes osteogenic differentiation of human bone marrow-derived mesenchymal stem cells by directly downregulating HMGA2. Biochem. Biophys. Res. Commun. 2019. [Google Scholar] [CrossRef]
- Xi, X.; Teng, M.; Zhang, L.; Xia, L.; Chen, J.; Cui, Z. MicroRNA-204-3p represses colon cancer cells proliferation, migration, and invasion by targeting HMGA2. J. Cell. Physiol. 2019. [Google Scholar] [CrossRef]
- Ren, J.; Yang, M.; Xu, F.; Chen, J. microRNA-758 inhibits the malignant phenotype of osteosarcoma cells by directly targeting HMGA1 and deactivating the Wnt/β-catenin pathway. Am. J. Cancer Res. 2019, 9, 36–52. [Google Scholar]
- Tsai, S.-C.; Huang, S.-F.; Chiang, J.-H.; Chen, Y.-F.; Huang, C.-C.; Tsai, M.-H.; Tsai, F.-J.; Kao, M.-C.; Yang, J.-S. The differential regulation of microRNAs is associated with oral cancer. Oncol. Rep. 2017, 38, 1613–1620. [Google Scholar] [CrossRef]
- Wu, H.; Liang, Y.; Shen, L.; Shen, L. MicroRNA-204 modulates colorectal cancer cell sensitivity in response to 5-fluorouracil-based treatment by targeting high mobility group protein A2. Biol. Open 2016, 5, 563–570. [Google Scholar] [CrossRef]
- Wu, Z.Y.; Wang, S.M.; Chen, Z.H.; Huv, S.X.; Huang, K.; Huang, B.J.; Du, J.L.; Huang, C.M.; Peng, L.; Jian, Z.X.; et al. MiR-204 regulates HMGA2 expression and inhibits cell proliferation in human thyroid cancer. Cancer Biomark. 2015, 15, 535–542. [Google Scholar] [CrossRef]
- Leung, Y.-K.; Chan, Q.K.-Y.; Ng, C.-F.; Ma, F.M.-T.; Tse, H.-M.; To, K.-F.; Maranchie, J.; Ho, S.-M.; Lau, K.-M. Hsa-miRNA-765 as a key mediator for inhibiting growth, migration and invasion in fulvestrant-treated prostate cancer. PLoS ONE 2014, 9, e98037. [Google Scholar] [CrossRef]
- Li, X.; Wang, S.; Li, Z.; Long, X.; Guo, Z.; Zhang, G.; Zu, J.; Chen, Y.; Wen, L. The lncRNA NEAT1 facilitates cell growth and invasion via the miR-211/HMGA2 axis in breast cancer. Int. J. Biol. Macromol. 2017, 105, 346–353. [Google Scholar] [CrossRef]
- Li, W.; Miao, X.; Liu, L.; Zhang, Y.; Jin, X.; Luo, X.; Gao, H.; Deng, X. Methylation-mediated silencing of microRNA-211 promotes cell growth and epithelial to mesenchymal transition through activation of the AKT/β-catenin pathway in GBM. Oncotarget 2017, 8, 25167–25176. [Google Scholar] [CrossRef]
- Chandrasekaran, K.S.; Sathyanarayanan, A.; Karunagaran, D. MicroRNA-214 suppresses growth, migration and invasion through a novel target, high mobility group AT-hook 1, in human cervical and colorectal cancer cells. Br. J. Cancer 2016, 115, 741–751. [Google Scholar] [CrossRef]
- Wang, J.; Zhao, X.; Guo, Z.; Ma, X.; Song, Y.; Guo, Y. Regulation of NEAT1/miR-214-3p on the growth, migration and invasion of endometrial carcinoma cells. Arch. Gynecol. Obstet. 2017, 295, 1469–1475. [Google Scholar] [CrossRef]
- Chen, X.; Zeng, K.; Xu, M.; Liu, X.; Hu, X.; Xu, T.; He, B.; Pan, Y.; Sun, H.; Wang, S. P53-induced miR-1249 inhibits tumor growth, metastasis, and angiogenesis by targeting VEGFA and HMGA2. Cell Death Dis. 2019, 10, 131. [Google Scholar] [CrossRef]
- Sun, X.; Xu, M.; Liu, H.; Ming, K. MicroRNA-219 is downregulated in non-small cell lung cancer and inhibits cell growth and metastasis by targeting HMGA2. Mol. Med. Rep. 2017, 16, 3557–3564. [Google Scholar] [CrossRef]
- Xing, F.; Song, Z.; He, Y. MiR-219-5p inhibits growth and metastasis of ovarian cancer cells by targeting HMGA2. Biol. Res. 2018, 51, 50. [Google Scholar] [CrossRef]
- Wang, J.; Xu, X.; Mo, S.; Tian, Y.; Wu, J.; Zhang, J.; Zhao, J. Involvement of microRNA-1297, a new regulator of HMGA1, in the regulation of glioma cell growth in vivo and in vitro. Am. J. Transl. Res. 2016, 8, 2149–2158. [Google Scholar] [PubMed]
- Liu, Y.; Liang, H.; Jiang, X. MiR-1297 promotes apoptosis and inhibits the proliferation and invasion of hepatocellular carcinoma cells by targeting HMGA2. Int. J. Mol. Med. 2015, 36, 1345–1352. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.-C.; Liu, J.-S.; Tang, H.-K.; Nie, J.; Zhu, J.-X.; Wen, L.-L.; Guo, Q.-L. miR-221 targets HMGA2 to inhibit bleomycin-induced pulmonary fibrosis by regulating TGF-β1/Smad3-induced EMT. Int. J. Mol. Med. 2016, 38, 1208–1216. [Google Scholar] [CrossRef] [PubMed]
- Wei, J.-J.; Wu, X.; Peng, Y.; Shi, G.; Basturk, O.; Olca, B.; Yang, X.; Daniels, G.; Osman, I.; Ouyang, J.; et al. Regulation of HMGA1 expression by microRNA-296 affects prostate cancer growth and invasion. Clin. Cancer Res. 2011, 17, 1297–1305. [Google Scholar] [CrossRef] [PubMed]
- Lopez-Bertoni, H.; Lal, B.; Michelson, N.; Guerrero-Cázares, H.; Quiñones-Hinojosa, A.; Li, Y.; Laterra, J. Epigenetic modulation of a miR-296-5p:HMGA1 axis regulates Sox2 expression and glioblastoma stem cells. Oncogene 2016, 35, 4903–4913. [Google Scholar] [CrossRef]
- Ma, Y.; Li, X.; Chen, S.; Du, B.; Li, Y. MicroRNA-4458 suppresses migration and epithelial-mesenchymal transition via targeting HMGA1 in non-small-cell lung cancer cells. Cancer Manag. Res. 2019, 11, 637–649. [Google Scholar] [CrossRef]
- Yu, F.Y.; Tu, Y.; Deng, Y.; Guo, C.; Ning, J.; Zhu, Y.; Lv, X.; Ye, H. MiR-4500 is epigenetically downregulated in colorectal cancer and functions as a novel tumor suppressor by regulating HMGA2. Cancer Biol. Ther. 2016, 17, 1149–1157. [Google Scholar] [CrossRef]
- Ogram, S.A.; Reeves, R. Differential regulation of a multipromoter gene. Selective 12-O-tetradecanoylphorbol-13-acetate induction of a single transcription start site in the HMG-I/Y gene. J. Biol. Chem. 1995, 270, 14235–14242. [Google Scholar] [CrossRef]
- Wood, L.J.; Mukherjee, M.; Dolde, C.E.; Xu, Y.; Maher, J.F.; Bunton, T.E.; Williams, J.B.; Resar, L.M. HMG-I/Y, a new c-Myc target gene and potential oncogene. Mol. Cell. Biol. 2000, 20, 5490–5502. [Google Scholar] [CrossRef]
- Giannini, G.; Cerignoli, F.; Mellone, M.; Massimi, I.; Ambrosi, C.; Rinaldi, C.; Dominici, C.; Frati, L.; Screpanti, I.; Gulino, A. High mobility group A1 is a molecular target for MYCN in human neuroblastoma. Cancer Res. 2005, 65, 8308–8316. [Google Scholar] [CrossRef][Green Version]
- Cleynen, I.; Huysmans, C.; Sasazuki, T.; Shirasawa, S.; Van de Ven, W.; Peeters, K. Transcriptional control of the human high mobility group A1 gene: Basal and oncogenic Ras-regulated expression. Cancer Res. 2007, 67, 4620–4629. [Google Scholar] [CrossRef] [PubMed]
- Nagpal, S.; Ghosn, C.; DiSepio, D.; Molina, Y.; Sutter, M.; Klein, E.S.; Chandraratna, R.A. Retinoid-dependent recruitment of a histone H1 displacement activity by retinoic acid receptor. J. Biol. Chem. 1999, 274, 22563–22568. [Google Scholar] [CrossRef] [PubMed]
- Zerbino, D.R.; Achuthan, P.; Akanni, W.; Amode, M.R.; Barrell, D.; Bhai, J.; Billis, K.; Cummins, C.; Gall, A.; Girón, C.G.; et al. Ensembl 2018. Nucleic Acids Res. 2018, 46, D754–D761. [Google Scholar] [CrossRef]
- Ashar, H.R.; Fejzo, M.S.; Tkachenko, A.; Zhou, X.; Fletcher, J.A.; Weremowicz, S.; Morton, C.C.; Chada, K. Disruption of the architectural factor HMGI-C: DNA-binding AT hook motifs fused in lipomas to distinct transcriptional regulatory domains. Cell 1995, 82, 57–65. [Google Scholar] [CrossRef]
- Schoenmakers, E.F.; Wanschura, S.; Mols, R.; Bullerdiek, J.; Van den Berghe, H.; Van de Ven, W.J. Recurrent rearrangements in the high mobility group protein gene, HMGI-C, in benign mesenchymal tumours. Nat. Genet. 1995, 10, 436–444. [Google Scholar] [CrossRef] [PubMed]
- Ashar, H.R.; Cherath, L.; Przybysz, K.M.; Chada, K. Genomic characterization of human HMGIC, a member of the accessory transcription factor family found at translocation breakpoints in lipomas. Genomics 1996, 31, 207–214. [Google Scholar] [CrossRef] [PubMed]
- Hauke, S.; Leopold, S.; Schlueter, C.; Flohr, A.M.; Murua Escobar, H.; Rogalla, P.; Bullerdiek, J. Extensive expression studies revealed a complex alternative splicing pattern of the HMGA2 gene. Biochim. Biophys. Acta 2005, 1729, 24–31. [Google Scholar] [CrossRef]
- Hauke, S.; Flohr, A.M.; Rogalla, P.; Bullerdiek, J. Sequencing of intron 3 of HMGA2 uncovers the existence of a novel exon. Genes Chromosomes Cancer 2002, 34, 17–23. [Google Scholar] [CrossRef]
- Cesana, M.; Guo, M.H.; Cacchiarelli, D.; Wahlster, L.; Barragan, J.; Doulatov, S.; Vo, L.T.; Salvatori, B.; Trapnell, C.; Clement, K.; et al. A CLK3-HMGA2 Alternative Splicing Axis Impacts Human Hematopoietic Stem Cell Molecular Identity throughout Development. Cell Stem Cell 2018, 22, 575–588.e7. [Google Scholar] [CrossRef]
- Fedele, M.; Fusco, A. HMGA and cancer. Biochim. Biophys. Acta 2010, 1799, 48–54. [Google Scholar] [CrossRef]
- Borrmann, L.; Wilkening, S.; Bullerdiek, J. The expression of HMGA genes is regulated by their 3′UTR. Oncogene 2001, 20, 4537–4541. [Google Scholar] [CrossRef] [PubMed]
- Reinhart, B.J.; Slack, F.J.; Basson, M.; Pasquinelli, A.E.; Bettinger, J.C.; Rougvie, A.E.; Horvitz, H.R.; Ruvkun, G. The 21-nucleotide let-7 RNA regulates developmental timing in Caenorhabditis elegans. Nature 2000, 403, 901–906. [Google Scholar] [CrossRef] [PubMed]
- Pasquinelli, A.E.; Reinhart, B.J.; Slack, F.; Martindale, M.Q.; Kuroda, M.I.; Maller, B.; Hayward, D.C.; Ball, E.E.; Degnan, B.; Müller, P.; et al. Conservation of the sequence and temporal expression of let-7 heterochronic regulatory RNA. Nature 2000, 408, 86–89. [Google Scholar] [CrossRef] [PubMed]
- Ruby, J.G.; Jan, C.; Player, C.; Axtell, M.J.; Lee, W.; Nusbaum, C.; Ge, H.; Bartel, D.P. Large-scale sequencing reveals 21U-RNAs and additional microRNAs and endogenous siRNAs in C. elegans. Cell 2006, 127, 1193–1207. [Google Scholar] [CrossRef] [PubMed]
- Roush, S.; Slack, F.J. The let-7 family of microRNAs. Trends Cell Biol. 2008, 18, 505–516. [Google Scholar] [CrossRef] [PubMed]
- Boyerinas, B.; Park, S.-M.; Hau, A.; Murmann, A.E.; Peter, M.E. The role of let-7 in cell differentiation and cancer. Endocr. Relat. Cancer 2010, 17, F19–F36. [Google Scholar] [CrossRef]
- Hisaoka, M.; Matsuyama, A.; Nagao, Y.; Luan, L.; Kuroda, T.; Akiyama, H.; Kondo, S.; Hashimoto, H. Identification of altered MicroRNA expression patterns in synovial sarcoma. Genes Chromosomes Cancer 2011, 50, 137–145. [Google Scholar] [CrossRef]
- De Vito, C.; Riggi, N.; Suvà, M.-L.; Janiszewska, M.; Horlbeck, J.; Baumer, K.; Provero, P.; Stamenkovic, I. Let-7a is a direct EWS-FLI-1 target implicated in Ewing’s sarcoma development. PLoS ONE 2011, 6, e23592. [Google Scholar] [CrossRef]
- Liu, Q.; Lv, G.-D.; Qin, X.; Gen, Y.-H.; Zheng, S.-T.; Liu, T.; Lu, X.-M. Role of microRNA let-7 and effect to HMGA2 in esophageal squamous cell carcinoma. Mol. Biol. Rep. 2012, 39, 1239–1246. [Google Scholar] [CrossRef]
- Shi, G.; Perle, M.A.; Mittal, K.; Chen, H.; Zou, X.; Narita, M.; Hernando, E.; Lee, P.; Wei, J.-J. Let-7 repression leads to HMGA2 overexpression in uterine leiomyosarcoma. J. Cell. Mol. Med. 2009, 13, 3898–3905. [Google Scholar] [CrossRef]
- Qian, Z.R.; Asa, S.L.; Siomi, H.; Siomi, M.C.; Yoshimoto, K.; Yamada, S.; Wang, E.L.; Rahman, M.M.; Inoue, H.; Itakura, M.; et al. Overexpression of HMGA2 relates to reduction of the let-7 and its relationship to clinicopathological features in pituitary adenomas. Mod. Pathol. 2009, 22, 431–441. [Google Scholar] [CrossRef] [PubMed]
- Liu, Q.; Liu, T.; Zheng, S.; Gao, X.; Lu, M.; Sheyhidin, I.; Lu, X. HMGA2 is down-regulated by microRNA let-7 and associated with epithelial-mesenchymal transition in oesophageal squamous cell carcinomas of Kazakhs. Histopathology 2014, 65, 408–417. [Google Scholar] [CrossRef] [PubMed]
- Dangi-Garimella, S.; Yun, J.; Eves, E.M.; Newman, M.; Erkeland, S.J.; Hammond, S.M.; Minn, A.J.; Rosner, M.R. Raf kinase inhibitory protein suppresses a metastasis signalling cascade involving LIN28 and let-7. EMBO J. 2009, 28, 347–358. [Google Scholar] [CrossRef] [PubMed]
- Chen, K.-J.; Hou, Y.; Wang, K.; Li, J.; Xia, Y.; Yang, X.-Y.; Lv, G.; Xing, X.-L.; Shen, F. Reexpression of Let-7g microRNA inhibits the proliferation and migration via K-Ras/HMGA2/snail axis in hepatocellular carcinoma. BioMed Res. Int. 2014, 2014, 742417. [Google Scholar] [CrossRef] [PubMed]
- Xiao, G.; Wang, X.; Yu, Y. CXCR4/Let-7a Axis Regulates Metastasis and Chemoresistance of Pancreatic Cancer Cells Through Targeting HMGA2. Cell. Physiol. Biochem. 2017, 43, 840–851. [Google Scholar] [CrossRef]
- Sun, T.; Fu, M.; Bookout, A.L.; Kliewer, S.A.; Mangelsdorf, D.J. MicroRNA let-7 regulates 3T3-L1 adipogenesis. Mol. Endocrinol. 2009, 23, 925–931. [Google Scholar] [CrossRef]
- Rowe, R.G.; Wang, L.D.; Coma, S.; Han, A.; Mathieu, R.; Pearson, D.S.; Ross, S.; Sousa, P.; Nguyen, P.T.; Rodriguez, A.; et al. Developmental regulation of myeloerythroid progenitor function by the Lin28b-let-7-Hmga2 axis. J. Exp. Med. 2016, 213, 1497–1512. [Google Scholar] [CrossRef]
- Patterson, M.; Gaeta, X.; Loo, K.; Edwards, M.; Smale, S.; Cinkornpumin, J.; Xie, Y.; Listgarten, J.; Azghadi, S.; Douglass, S.M.; et al. let-7 miRNAs Can Act through Notch to Regulate Human Gliogenesis. Stem Cell Rep. 2014, 3, 758–773. [Google Scholar] [CrossRef]
- Keane, M.; de Magalhães, J.P. MYCN/LIN28B/Let-7/HMGA2 pathway implicated by meta-analysis of GWAS in suppression of post-natal proliferation thereby potentially contributing to aging. Mech. Ageing Dev. 2013, 134, 346–348. [Google Scholar] [CrossRef]
- Zhu, H.; Shyh-Chang, N.; Segrè, A.V.; Shinoda, G.; Shah, S.P.; Einhorn, W.S.; Takeuchi, A.; Engreitz, J.M.; Hagan, J.P.; Kharas, M.G.; et al. The Lin28/let-7 axis regulates glucose metabolism. Cell 2011, 147, 81–94. [Google Scholar] [CrossRef]
- Saâda, E.; Bianchini, L.; Mouroux, J.; Dupré, F.; Butori, C.; Birtwisle-Peyrottes, I.; Padovani, B.; Yver, M.; Ferrari, C.; Pedeutour, F. First description of inhibition of let-7 microRNA expression and HMGA2 overexpression in a case of deep-seated diffuse lipomatosis. Histopathology 2012, 61, 519–522. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Le, Y.; Xue, J.-Y.; Zheng, Z.-J.; Xue, Y.-M. Let-7d miRNA prevents TGF-β1-induced EMT and renal fibrogenesis through regulation of HMGA2 expression. Biochem. Biophys. Res. Commun. 2016, 479, 676–682. [Google Scholar] [CrossRef] [PubMed]
- Copley, M.R.; Babovic, S.; Benz, C.; Knapp, D.J.H.F.; Beer, P.A.; Kent, D.G.; Wohrer, S.; Treloar, D.Q.; Day, C.; Rowe, K.; et al. The Lin28b-let-7-Hmga2 axis determines the higher self-renewal potential of fetal haematopoietic stem cells. Nat. Cell Biol. 2013, 15, 916–925. [Google Scholar] [CrossRef] [PubMed]
- Mao, X.; Hütt-Cabezas, M.; Orr, B.A.; Weingart, M.; Taylor, I.; Rajan, A.K.D.; Odia, Y.; Kahlert, U.; Maciaczyk, J.; Nikkhah, G.; et al. LIN28A facilitates the transformation of human neural stem cells and promotes glioblastoma tumorigenesis through a pro-invasive genetic program. Oncotarget 2013, 4, 1050–1064. [Google Scholar] [CrossRef]
- Alam, M.; Ahmad, R.; Rajabi, H.; Kufe, D. MUC1-C Induces the LIN28B→LET-7→HMGA2 Axis to Regulate Self-Renewal in NSCLC. Mol. Cancer Res. 2015, 13, 449–460. [Google Scholar] [CrossRef]
- Madison, B.B.; Jeganathan, A.N.; Mizuno, R.; Winslow, M.M.; Castells, A.; Cuatrecasas, M.; Rustgi, A.K. Let-7 Represses Carcinogenesis and a Stem Cell Phenotype in the Intestine via Regulation of Hmga2. PLoS Genet. 2015, 11, e1005408. [Google Scholar] [CrossRef]
- Viswanathan, S.R.; Daley, G.Q.; Gregory, R.I. Selective blockade of microRNA processing by Lin28. Science 2008, 320, 97–100. [Google Scholar] [CrossRef]
- Heo, I.; Joo, C.; Cho, J.; Ha, M.; Han, J.; Kim, V.N. Lin28 mediates the terminal uridylation of let-7 precursor MicroRNA. Mol. Cell 2008, 32, 276–284. [Google Scholar] [CrossRef]
- Balzeau, J.; Menezes, M.R.; Cao, S.; Hagan, J.P. The LIN28/let-7 Pathway in Cancer. Front. Genet. 2017, 8, 31. [Google Scholar] [CrossRef]
- GAO, J.; LIU, Q.-G. The role of miR-26 in tumors and normal tissues (Review). Oncol. Lett. 2011, 2, 1019–1023. [Google Scholar] [CrossRef]
- Wong, C.F.; Tellam, R.L. MicroRNA-26a targets the histone methyltransferase Enhancer of Zeste homolog 2 during myogenesis. J. Biol. Chem. 2008, 283, 9836–9843. [Google Scholar] [CrossRef] [PubMed]
- De Freitas, R.C.C.; Bortolin, R.H.; Lopes, M.B.; Tamborlin, L.; Meneguello, L.; Silbiger, V.N.; Hirata, R.D.C.; Hirata, M.H.; Luchessi, A.D.; Luchessi, A.D. Modulation of miR-26a-5p and miR-15b-5p Exosomal Expression Associated with Clopidogrel-Induced Hepatotoxicity in HepG2 Cells. Front. Pharmacol. 2017, 8, 906. [Google Scholar] [CrossRef] [PubMed]
- Lin, R.; Shen, W.; Zhi, Y.; Zhou, Z. Prognostic value of miR-26a and HMGA1 in urothelial bladder cancer. Biomed. Pharmacother. 2014, 68, 929–934. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.; Jung, J.-W.; Park, S.-B.; Roh, K.; Lee, S.Y.; Kim, J.H.; Kang, S.-K.; Kang, K.-S. Histone deacetylase regulates high mobility group A2-targeting microRNAs in human cord blood-derived multipotent stem cell aging. Cell. Mol. Life Sci. 2011, 68, 325–336. [Google Scholar] [CrossRef] [PubMed]
- Kozomara, A.; Birgaoanu, M.; Griffiths-Jones, S. miRBase: From microRNA sequences to function. Nucleic Acids Res. 2019, 47, D155–D162. [Google Scholar] [CrossRef] [PubMed]
- Dávalos, A.; Goedeke, L.; Smibert, P.; Ramírez, C.M.; Warrier, N.P.; Andreo, U.; Cirera-Salinas, D.; Rayner, K.; Suresh, U.; Pastor-Pareja, J.C.; et al. miR-33a/b contribute to the regulation of fatty acid metabolism and insulin signaling. Proc. Natl. Acad. Sci. USA 2011, 108, 9232–9237. [Google Scholar] [CrossRef]
- Hebert, C.; Norris, K.; Scheper, M.A.; Nikitakis, N.; Sauk, J.J. High mobility group A2 is a target for miRNA-98 in head and neck squamous cell carcinoma. Mol. Cancer 2007, 6, 5. [Google Scholar] [CrossRef]
- Huang, E.; Liu, R.; Chu, Y. miRNA-15a/16: As tumor suppressors and more. Future Oncol. 2015, 11, 2351–2363. [Google Scholar] [CrossRef]
- Sharma, S.; Liu, J.; Wei, J.; Yuan, H.; Zhang, T.; Bishopric, N.H. Repression of miR-142 by p300 and MAPK is required for survival signalling via gp130 during adaptive hypertrophy. EMBO Mol. Med. 2012, 4, 617–632. [Google Scholar] [CrossRef] [PubMed]
- Isobe, T.; Hisamori, S.; Hogan, D.J.; Zabala, M.; Hendrickson, D.G.; Dalerba, P.; Cai, S.; Scheeren, F.; Kuo, A.H.; Sikandar, S.S.; et al. miR-142 regulates the tumorigenicity of human breast cancer stem cells through the canonical WNT signaling pathway. Elife 2014, 3, e01977. [Google Scholar] [CrossRef]
- Tsang, F.H.-C.; Au, S.L.-K.; Wei, L.; Fan, D.N.-Y.; Lee, J.M.-F.; Wong, C.C.-L.; Ng, I.O.-L.; Wong, C.-M. MicroRNA-142-3p and microRNA-142-5p are downregulated in hepatocellular carcinoma and exhibit synergistic effects on cell motility. Front. Med. 2015, 9, 331–343. [Google Scholar] [CrossRef]
- Shrestha, A.; Mukhametshina, R.T.; Taghizadeh, S.; Vásquez-Pacheco, E.; Cabrera-Fuentes, H.; Rizvanov, A.; Mari, B.; Carraro, G.; Bellusci, S. MicroRNA-142 is a multifaceted regulator in organogenesis, homeostasis, and disease. Dev. Dyn. 2017, 246, 285–290. [Google Scholar] [CrossRef] [PubMed]
- Wu, L.; Cai, C.; Wang, X.; Liu, M.; Li, X.; Tang, H. MicroRNA-142-3p, a new regulator of RAC1, suppresses the migration and invasion of hepatocellular carcinoma cells. FEBS Lett. 2011, 585, 1322–1330. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.; Wang, X.-S.; Yang, G.-H.; Zhai, P.-F.; Xiao, Z.; Xia, L.-Y.; Chen, L.-R.; Wang, Y.; Wang, X.-Z.; Bi, L.-X.; et al. miR-29a and miR-142-3p downregulation and diagnostic implication in human acute myeloid leukemia. Mol. Biol. Rep. 2012, 39, 2713–2722. [Google Scholar] [CrossRef] [PubMed]
- Shen, W.-W.; Zeng, Z.; Zhu, W.-X.; Fu, G.-H. MiR-142-3p functions as a tumor suppressor by targeting CD133, ABCG2, and Lgr5 in colon cancer cells. J. Mol. Med. 2013, 91, 989–1000. [Google Scholar] [CrossRef]
- Agarwal, V.; Bell, G.W.; Nam, J.-W.; Bartel, D.P. Predicting effective microRNA target sites in mammalian mRNAs. Elife 2015, 4, e05005. [Google Scholar] [CrossRef] [PubMed]
- Pinzón, N.; Li, B.; Martinez, L.; Sergeeva, A.; Presumey, J.; Apparailly, F.; Seitz, H. microRNA target prediction programs predict many false positives. Genome Res. 2017, 27, 234–245. [Google Scholar] [CrossRef]
- Selbach, M.; Schwanhäusser, B.; Thierfelder, N.; Fang, Z.; Khanin, R.; Rajewsky, N. Widespread changes in protein synthesis induced by microRNAs. Nature 2008, 455, 58–63. [Google Scholar] [CrossRef]
- Baek, D.; Villén, J.; Shin, C.; Camargo, F.D.; Gygi, S.P.; Bartel, D.P. The impact of microRNAs on protein output. Nature 2008, 455, 64–71. [Google Scholar] [CrossRef]
- Chiu, H.-S.; Martínez, M.R.; Komissarova, E.V.; Llobet-Navas, D.; Bansal, M.; Paull, E.O.; Silva, J.; Yang, X.; Sumazin, P.; Califano, A. The number of titrated microRNA species dictates ceRNA regulation. Nucleic Acids Res. 2018, 46, 4354–4369. [Google Scholar] [CrossRef]
- Sarver, A.L.; Subramanian, S. Competing endogenous RNA database. Bioinformation 2012, 8, 731–733. [Google Scholar] [CrossRef] [PubMed]
- Chou, C.-H.; Shrestha, S.; Yang, C.-D.; Chang, N.-W.; Lin, Y.-L.; Liao, K.-W.; Huang, W.-C.; Sun, T.-H.; Tu, S.-J.; Lee, W.-H.; et al. miRTarBase update 2018: A resource for experimentally validated microRNA-target interactions. Nucleic Acids Res. 2018, 46, D296–D302. [Google Scholar] [CrossRef] [PubMed]
- Iliopoulos, D.; Hirsch, H.A.; Struhl, K. An epigenetic switch involving NF-kappaB, Lin28, Let-7 MicroRNA, and IL6 links inflammation to cell transformation. Cell 2009, 139, 693–706. [Google Scholar] [CrossRef] [PubMed]
- Powers, J.T.; Tsanov, K.M.; Pearson, D.S.; Roels, F.; Spina, C.S.; Ebright, R.; Seligson, M.; de Soysa, Y.; Cahan, P.; Theiβen, J.; et al. Multiple mechanisms disrupt the let-7 microRNA family in neuroblastoma. Nature 2016, 535, 246–251. [Google Scholar] [CrossRef] [PubMed]
- Kallen, A.N.; Zhou, X.-B.; Xu, J.; Qiao, C.; Ma, J.; Yan, L.; Lu, L.; Liu, C.; Yi, J.-S.; Zhang, H.; et al. The imprinted H19 lncRNA antagonizes let-7 microRNAs. Mol. Cell 2013, 52, 101–112. [Google Scholar] [CrossRef]
- Dang, X.; Ma, A.; Yang, L.; Hu, H.; Zhu, B.; Shang, D.; Chen, T.; Luo, Y. MicroRNA-26a regulates tumorigenic properties of EZH2 in human lung carcinoma cells. Cancer Genet 2012, 205, 113–123. [Google Scholar] [CrossRef]
- Alajez, N.M.; Shi, W.; Hui, A.B.Y.; Bruce, J.; Lenarduzzi, M.; Ito, E.; Yue, S.; O’Sullivan, B.; Liu, F.-F. Enhancer of Zeste homolog 2 (EZH2) is overexpressed in recurrent nasopharyngeal carcinoma and is regulated by miR-26a, miR-101, and miR-98. Cell Death Dis. 2010, 1, e85. [Google Scholar] [CrossRef]
- Yang, C.; Zheng, S.; Liu, T.; Liu, Q.; Dai, F.; Zhou, J.; Chen, Y.; Sheyhidin, I.; Lu, X. Down-regulated miR-26a promotes proliferation, migration, and invasion via negative regulation of MTDH in esophageal squamous cell carcinoma. FASEB J. 2017, 31, 2114–2122. [Google Scholar] [CrossRef]
- Kota, J.; Chivukula, R.R.; O’Donnell, K.A.; Wentzel, E.A.; Montgomery, C.L.; Hwang, H.-W.; Chang, T.-C.; Vivekanandan, P.; Torbenson, M.; Clark, K.R.; et al. Therapeutic microRNA delivery suppresses tumorigenesis in a murine liver cancer model. Cell 2009, 137, 1005–1017. [Google Scholar] [CrossRef]
- Luzi, E.; Marini, F.; Sala, S.C.; Tognarini, I.; Galli, G.; Brandi, M.L. Osteogenic differentiation of human adipose tissue-derived stem cells is modulated by the miR-26a targeting of the SMAD1 transcription factor. J. Bone Miner. Res. 2008, 23, 287–295. [Google Scholar] [CrossRef]
- Huse, J.T.; Brennan, C.; Hambardzumyan, D.; Wee, B.; Pena, J.; Rouhanifard, S.H.; Sohn-Lee, C.; le Sage, C.; Agami, R.; Tuschl, T.; et al. The PTEN-regulating microRNA miR-26a is amplified in high-grade glioma and facilitates gliomagenesis in vivo. Genes Dev. 2009, 23, 1327–1337. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.; Huang, W.; Jiang, X.; Pennicooke, B.; Park, P.J.; Johnson, M.D. Integrative genome analysis reveals an oncomir/oncogene cluster regulating glioblastoma survivorship. Proc. Natl. Acad. Sci. USA 2010, 107, 2183–2188. [Google Scholar] [CrossRef] [PubMed]
- Tian, Z.; Zhao, J.; Tai, Y.-T.; Amin, S.B.; Hu, Y.; Berger, A.J.; Richardson, P.; Chauhan, D.; Anderson, K.C. Investigational agent MLN9708/2238 targets tumor-suppressor miR33b in MM cells. Blood 2012, 120, 3958–3967. [Google Scholar] [CrossRef] [PubMed]
- Sun, Q.; Zhang, W.; Guo, Y.; Li, Z.; Chen, X.; Wang, Y.; Du, Y.; Zang, W.; Zhao, G. Curcumin inhibits cell growth and induces cell apoptosis through upregulation of miR-33b in gastric cancer. Tumour Biol. 2016, 37, 13177–13184. [Google Scholar] [CrossRef]
- Qu, J.; Li, M.; An, J.; Zhao, B.; Zhong, W.; Gu, Q.; Cao, L.; Yang, H.; Hu, C. MicroRNA-33b inhibits lung adenocarcinoma cell growth, invasion, and epithelial-mesenchymal transition by suppressing Wnt/β-catenin/ZEB1 signaling. Int. J. Oncol. 2015, 47, 2141–2152. [Google Scholar] [CrossRef]
- Ji, M.; Lu, J.; Shi, P.; Zhang, X.; Wang, S.; Chang, Q.; Chen, H.; Wang, C. Dysregulated miR-98 Contributes to Extracellular Matrix Degradation by Targeting IL-6/STAT3 Signaling Pathway in Human Intervertebral Disc Degeneration. J. Bone Miner. Res. 2016, 31, 900–909. [Google Scholar] [CrossRef]
- Panda, H.; Chuang, T.-D.; Luo, X.; Chegini, N. Endometrial miR-181a and miR-98 expression is altered during transition from normal into cancerous state and target PGR, PGRMC1, CYP19A1, DDX3X, and TIMP3. J. Clin. Endocrinol. Metab. 2012, 97, E1316–E1326. [Google Scholar] [CrossRef]
- Zhu, W.; Huang, Y.; Pan, Q.; Xiang, P.; Xie, N.; Yu, H. MicroRNA-98 Suppress Warburg Effect by Targeting HK2 in Colon Cancer Cells. Dig. Dis. Sci. 2017, 62, 660–668. [Google Scholar] [CrossRef]
- Ni, R.; Huang, Y.; Wang, J. miR-98 targets ITGB3 to inhibit proliferation, migration, and invasion of non-small-cell lung cancer. Onco Targets Ther. 2015, 8, 2689–2697. [Google Scholar]
- Bhat-Nakshatri, P.; Wang, G.; Collins, N.R.; Thomson, M.J.; Geistlinger, T.R.; Carroll, J.S.; Brown, M.; Hammond, S.; Srour, E.F.; Liu, Y.; et al. Estradiol-regulated microRNAs control estradiol response in breast cancer cells. Nucleic Acids Res. 2009, 37, 4850–4861. [Google Scholar] [CrossRef]
- Du, L.; Schageman, J.J.; Subauste, M.C.; Saber, B.; Hammond, S.M.; Prudkin, L.; Wistuba, I.I.; Ji, L.; Roth, J.A.; Minna, J.D.; et al. miR-93, miR-98, and miR-197 regulate expression of tumor suppressor gene FUS1. Mol. Cancer Res. 2009, 7, 1234–1243. [Google Scholar] [CrossRef] [PubMed]
- Liu, Q.; Fu, H.; Sun, F.; Zhang, H.; Tie, Y.; Zhu, J.; Xing, R.; Sun, Z.; Zheng, X. miR-16 family induces cell cycle arrest by regulating multiple cell cycle genes. Nucleic Acids Res. 2008, 36, 5391–5404. [Google Scholar] [CrossRef]
- Sun, C.-Y.; She, X.-M.; Qin, Y.; Chu, Z.-B.; Chen, L.; Ai, L.-S.; Zhang, L.; Hu, Y. miR-15a and miR-16 affect the angiogenesis of multiple myeloma by targeting VEGF. Carcinogenesis 2013, 34, 426–435. [Google Scholar] [CrossRef] [PubMed]
- Kang, W.; Tong, J.H.M.; Lung, R.W.M.; Dong, Y.; Zhao, J.; Liang, Q.; Zhang, L.; Pan, Y.; Yang, W.; Pang, J.C.S.; et al. Targeting of YAP1 by microRNA-15a and microRNA-16-1 exerts tumor suppressor function in gastric adenocarcinoma. Mol. Cancer 2015, 14, 52. [Google Scholar] [CrossRef] [PubMed]
- Musumeci, M.; Coppola, V.; Addario, A.; Patrizii, M.; Maugeri-Saccà, M.; Memeo, L.; Colarossi, C.; Francescangeli, F.; Biffoni, M.; Collura, D.; et al. Control of tumor and microenvironment cross-talk by miR-15a and miR-16 in prostate cancer. Oncogene 2011, 30, 4231–4242. [Google Scholar] [CrossRef]
- Lei, Z.; Xu, G.; Wang, L.; Yang, H.; Liu, X.; Zhao, J.; Zhang, H.-T. MiR-142-3p represses TGF-β-induced growth inhibition through repression of TGFβR1 in non-small cell lung cancer. FASEB J. 2014, 28, 2696–2704. [Google Scholar] [CrossRef]
- Liu, J.; Li, W.; Wang, S.; Wu, Y.; Li, Z.; Wang, W.; Liu, R.; Ou, J.; Zhang, C.; Wang, S. MiR-142-3p attenuates the migration of CD4+ T cells through regulating actin cytoskeleton via RAC1 and ROCK2 in arteriosclerosis obliterans. PLoS ONE 2014, 9, e95514. [Google Scholar] [CrossRef]
- Riley, K.J.; Rabinowitz, G.S.; Yario, T.A.; Luna, J.M.; Darnell, R.B.; Steitz, J.A. EBV and human microRNAs co-target oncogenic and apoptotic viral and human genes during latency. EMBO J. 2012, 31, 2207–2221. [Google Scholar] [CrossRef]
- Esposito, F.; De Martino, M.; Petti, M.G.; Forzati, F.; Tornincasa, M.; Federico, A.; Arra, C.; Pierantoni, G.M.; Fusco, A. HMGA1 pseudogenes as candidate proto-oncogenic competitive endogenous RNAs. Oncotarget 2014, 5, 8341–8354. [Google Scholar] [CrossRef]
- De Martino, M.; Forzati, F.; Marfella, M.; Pellecchia, S.; Arra, C.; Terracciano, L.; Fusco, A.; Esposito, F. HMGA1P7-pseudogene regulates H19 and Igf2 expression by a competitive endogenous RNA mechanism. Sci. Rep. 2016, 6, 37622. [Google Scholar] [CrossRef]
- De Martino, M.; Palma, G.; Azzariti, A.; Arra, C.; Fusco, A.; Esposito, F. The HMGA1 Pseudogene 7 Induces miR-483 and miR-675 Upregulation by Activating Egr1 through a ceRNA Mechanism. Genes (Basel) 2017, 8, 330. [Google Scholar] [CrossRef] [PubMed]
- Gao, C.; Lu, W.; Lou, W.; Wang, L.; Xu, Q. Long noncoding RNA HOXC13-AS positively affects cell proliferation and invasion in nasopharyngeal carcinoma via modulating miR-383-3p/HMGA2 axis. J. Cell. Physiol. 2019, 234, 12809–12820. [Google Scholar] [CrossRef] [PubMed]
- Oliveira-Mateos, C.; Sánchez-Castillo, A.; Soler, M.; Obiols-Guardia, A.; Piñeyro, D.; Boque-Sastre, R.; Calleja-Cervantes, M.E.; Castro de Moura, M.; Martínez-Cardús, A.; Rubio, T.; et al. The transcribed pseudogene RPSAP52 enhances the oncofetal HMGA2-IGF2BP2-RAS axis through LIN28B-dependent and independent let-7 inhibition. Nat. Commun. 2019, 10, 3979. [Google Scholar] [CrossRef] [PubMed]
- Ma, C.; Nong, K.; Zhu, H.; Wang, W.; Huang, X.; Yuan, Z.; Ai, K. H19 promotes pancreatic cancer metastasis by derepressing let-7′s suppression on its target HMGA2-mediated EMT. Tumour Biol. 2014, 35, 9163–9169. [Google Scholar] [CrossRef] [PubMed]
- Deng, L.; Yang, S.-B.; Xu, F.-F.; Zhang, J.-H. Long noncoding RNA CCAT1 promotes hepatocellular carcinoma progression by functioning as let-7 sponge. J. Exp. Clin. Cancer Res. 2015, 34, 18. [Google Scholar] [CrossRef] [PubMed]
- Esposito, F.; De Martino, M.; D’Angelo, D.; Mussnich, P.; Raverot, G.; Jaffrain-Rea, M.-L.; Fraggetta, F.; Trouillas, J.; Fusco, A. HMGA1-pseudogene expression is induced in human pituitary tumors. Cell Cycle 2015, 14, 1471–1475. [Google Scholar] [CrossRef]
- Miao, J.-T.; Gao, J.-H.; Chen, Y.-Q.; Chen, H.; Meng, H.-Y.; Lou, G. LncRNA ANRIL affects the sensitivity of ovarian cancer to cisplatin via regulation of let-7a/HMGA2 axis. Biosci. Rep. 2019, 39, BSR20182101. [Google Scholar] [CrossRef]
- Siepel, A.; Bejerano, G.; Pedersen, J.S.; Hinrichs, A.S.; Hou, M.; Rosenbloom, K.; Clawson, H.; Spieth, J.; Hillier, L.W.; Richards, S.; et al. Evolutionarily conserved elements in vertebrate, insect, worm, and yeast genomes. Genome Res. 2005, 15, 1034–1050. [Google Scholar] [CrossRef]
- Baltz, A.G.; Munschauer, M.; Schwanhäusser, B.; Vasile, A.; Murakawa, Y.; Schueler, M.; Youngs, N.; Penfold-Brown, D.; Drew, K.; Milek, M.; et al. The mRNA-bound proteome and its global occupancy profile on protein-coding transcripts. Mol. Cell 2012, 46, 674–690. [Google Scholar] [CrossRef]
- Kristjánsdóttir, K.; Fogarty, E.A.; Grimson, A. Systematic analysis of the Hmga2 3′ UTR identifies many independent regulatory sequences and a novel interaction between distal sites. RNA 2015, 21, 1346–1360. [Google Scholar] [CrossRef]
- Chen, R.-X.; Chen, X.; Xia, L.-P.; Zhang, J.-X.; Pan, Z.-Z.; Ma, X.-D.; Han, K.; Chen, J.-W.; Judde, J.-G.; Deas, O.; et al. N6-methyladenosine modification of circNSUN2 facilitates cytoplasmic export and stabilizes HMGA2 to promote colorectal liver metastasis. Nat. Commun. 2019, 10, 4695. [Google Scholar] [CrossRef] [PubMed]
- Jønson, L.; Christiansen, J.; Hansen, T.V.O.; Vikeså, J.; Yamamoto, Y.; Nielsen, F.C. IMP3 RNP safe houses prevent miRNA-directed HMGA2 mRNA decay in cancer and development. Cell Rep. 2014, 7, 539–551. [Google Scholar] [CrossRef] [PubMed]
- Ingenito, F.; Roscigno, G.; Affinito, A.; Nuzzo, S.; Scognamiglio, I.; Quintavalle, C.; Condorelli, G. The Role of Exo-miRNAs in Cancer: A Focus on Therapeutic and Diagnostic Applications. Int. J. Mol. Sci. 2019, 20, 4687. [Google Scholar] [CrossRef] [PubMed]
- Chirshev, E.; Oberg, K.C.; Ioffe, Y.J.; Unternaehrer, J.J. Let-7 as biomarker, prognostic indicator, and therapy for precision medicine in cancer. Clin. Transl. Med. 2019, 8, 24. [Google Scholar] [CrossRef]
- Takamizawa, J.; Konishi, H.; Yanagisawa, K.; Tomida, S.; Osada, H.; Endoh, H.; Harano, T.; Yatabe, Y.; Nagino, M.; Nimura, Y.; et al. Reduced expression of the let-7 microRNAs in human lung cancers in association with shortened postoperative survival. Cancer Res. 2004, 64, 3753–3756. [Google Scholar] [CrossRef]
- Liu, T.-P.; Huang, C.-C.; Yeh, K.-T.; Ke, T.-W.; Wei, P.-L.; Yang, J.-R.; Cheng, Y.-W. Down-regulation of let-7a-5p predicts lymph node metastasis and prognosis in colorectal cancer: Implications for chemotherapy. Surg. Oncol. 2016, 25, 429–434. [Google Scholar] [CrossRef]
- Yun, J.; Frankenberger, C.A.; Kuo, W.-L.; Boelens, M.C.; Eves, E.M.; Cheng, N.; Liang, H.; Li, W.-H.; Ishwaran, H.; Minn, A.J.; et al. Signalling pathway for RKIP and Let-7 regulates and predicts metastatic breast cancer. EMBO J. 2011, 30, 4500–4514. [Google Scholar] [CrossRef]
- Careccia, S.; Mainardi, S.; Pelosi, A.; Gurtner, A.; Diverio, D.; Riccioni, R.; Testa, U.; Pelosi, E.; Piaggio, G.; Sacchi, A.; et al. A restricted signature of miRNAs distinguishes APL blasts from normal promyelocytes. Oncogene 2009, 28, 4034–4040. [Google Scholar] [CrossRef]
- Di Fazio, P.; Montalbano, R.; Neureiter, D.; Alinger, B.; Schmidt, A.; Merkel, A.L.; Quint, K.; Ocker, M. Downregulation of HMGA2 by the pan-deacetylase inhibitor panobinostat is dependent on hsa-let-7b expression in liver cancer cell lines. Exp. Cell Res. 2012, 318, 1832–1843. [Google Scholar] [CrossRef]
- Jiang, W.; Finniss, S.; Cazacu, S.; Xiang, C.; Brodie, Z.; Mikkelsen, T.; Poisson, L.; Shackelford, D.B.; Brodie, C. Repurposing phenformin for the targeting of glioma stem cells and the treatment of glioblastoma. Oncotarget 2016, 7, 56456–56470. [Google Scholar] [CrossRef]
- Barh, D.; Malhotra, R.; Ravi, B.; Sindhurani, P. MicroRNA let-7: An emerging next-generation cancer therapeutic. Curr. Oncol. 2010, 17, 70–80. [Google Scholar] [CrossRef] [PubMed]
- Tang, H.; Zhang, P.; Xiang, Q.; Yin, J.; Yu, J.; Yang, X.; Lei, X. Let-7 g microRNA sensitizes fluorouracil-resistant human hepatoma cells. Pharmazie 2014, 69, 287–292. [Google Scholar] [PubMed]
- Manier, S.; Powers, J.T.; Sacco, A.; Glavey, S.V.; Huynh, D.; Reagan, M.R.; Salem, K.Z.; Moschetta, M.; Shi, J.; Mishima, Y.; et al. The LIN28B/let-7 axis is a novel therapeutic pathway in Multiple Myeloma. Leukemia 2017, 31, 853–860. [Google Scholar] [CrossRef] [PubMed]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).