Abstract
Leaf angle (LA), defined as the angle between the plant stem and leaf adaxial side of the blade, generally shapes the plant architecture into a loosen or dense structure, and thus influences the light interception and competition between neighboring plants in natural settings, ultimately contributing to the crop yield and productivity. It has been elucidated that brassinosteroid (BR) plays a dominant role in determining LA, and other phytohormones also positively or negatively participate in regulating LA. Accumulating evidences have revealed that these phytohormones interact with each other in modulating various biological processes. However, the comprehensive discussion of how the phytohormones and their interaction involved in shaping LA is relatively lack. Here, we intend to summarize the advances in the LA regulation mediated by the phytohormones and their crosstalk in different plant species, mainly in rice and maize, hopefully providing further insights into the genetic manipulation of LA trait in crop breeding and improvement in regarding to overcoming the challenge from the continuous demands for food under limited arable land area.
1. Introduction
To overcome the challenge of ever-increasing global demands for food, feedstock, and bioenergy products, breeders have been forced to select and breed cultivars with a key feature that can be planted at higher densities in order to increase grain yield with the limited availability of arable land area [1,2]. To this end, genetic improvement of crops with ideal plant architecture is considered as one of the most powerful strategies for addressing this issue. The key components of ideal plant architecture in crop generally include plant height, grain architecture, and leaf angle (LA) [3]. Since the 1960s, genetic engineering of decreasing plant height in crops has dramatically boosted the crop yield and definitely benefited millions of people worldwide, which is remarked as the “Green Revolution”. The great achievement of Green Revolution is attributed to the utilization of two “Green Revolution” genes, mutant allelic Semi-dwarf1 (Sd1) in rice, which encodes a key enzyme in the gibberellic pathway GA20ox2, and Reduced height-1 (Rht-1) in wheat that encodes a key repressor in gibberellin signaling pathway called DELLA, are responsible for gibberellin metabolism in rice and wheat [4,5], respectively. However, excessive application of nitrogen fertilizer for ensuring the yield and productivity of the semidwarf varieties has brought severe contamination on environment. Recently, a novel gibberellin-GIBBERELLIN INSENSITIVE DWARF1 (GID1)-NITROGEN-MEDIATED TILLER GROWTH RESPONSE 5 (NGR5) signaling pathway has been stated [6], which can be effectively used to boost yield of semi-dwarf crop by simultaneously enhancing the tiller number and nitrogen use efficiency (NUE) in next-generation Green Revolution. Besides plant height, the LA trait has also been substantially selected for high-yield varieties breeding, particularly in maize and rice, due to its vital role for optimal light interception and competition between neighboring plants in natural settings.
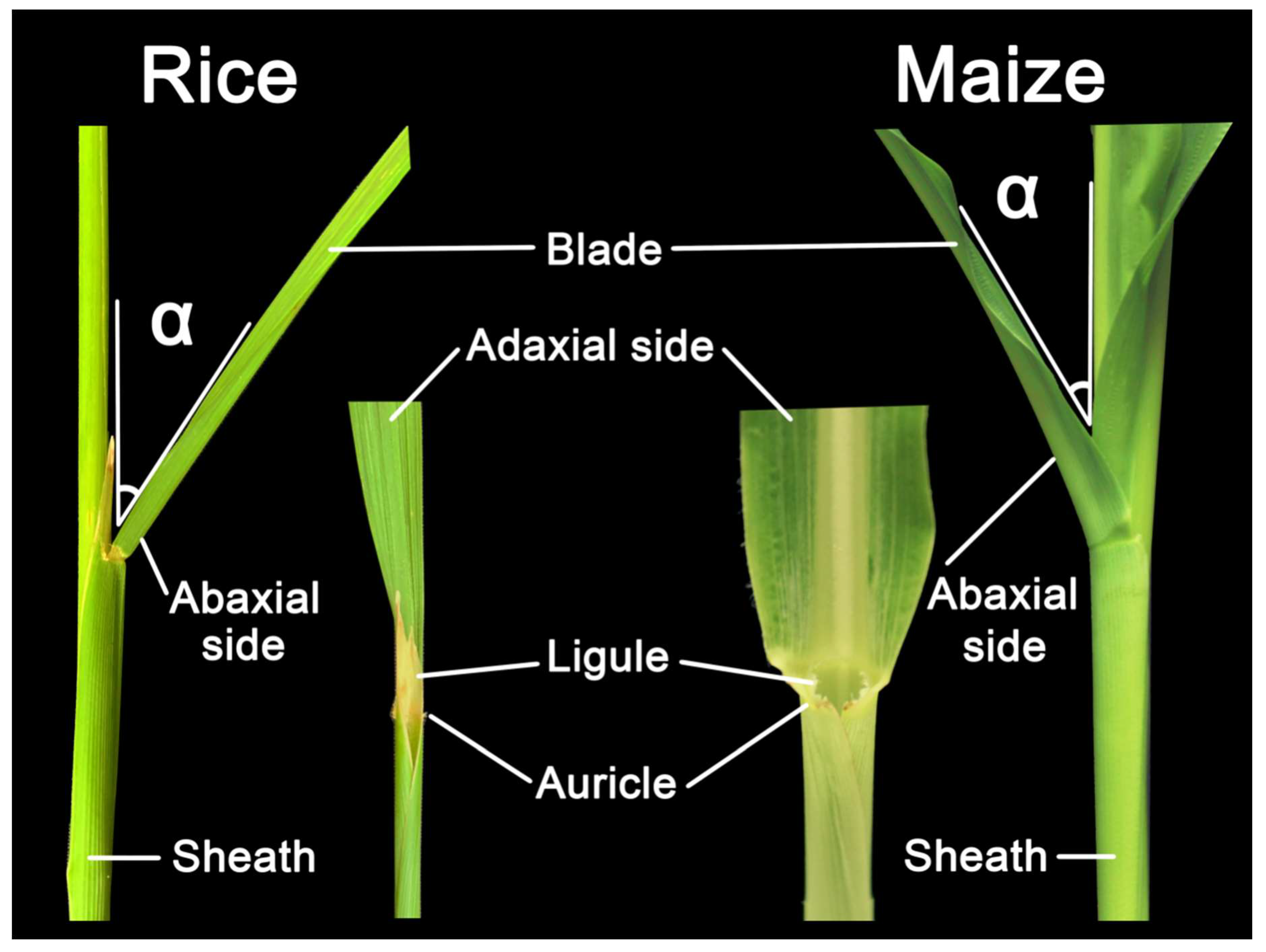
A typical grass leaf is consisted of distal blade, proximal sheath, and a boundary called ligular region (also called lamina joint) that separates blade and sheath into distinct parts, all of which contain epidermal, ground, and vascular tissues that are continuous with each other but distinct in cell types and patterns [7,8]. LA, defined as the angle between the stem and adaxial side of the blade (Figure 1), is one of the most important architecture traits selected for crop yield improvement [9]. Crops with architecture of smaller LA and more upright leaves can facilitate higher plant density and enhance the photosynthetic efficiency, thus elevating yield [10,11,12]. The ligular region, an annular structure outside the joint of the leaf blade and leaf sheath, is the pivotal structure for determining LA in grasses. Accumulating evidences have been implicated that the formation of LA is mediated by the shape of the lamina joint, differences in cell numbers/size at adaxial/abaxial region and distinctive mechanical tissue strength [9,13,14,15], and thus any influence on them could alter the LA. For example, failure of longitudinal elongation of the adaxial cells in the lamina joint may result in erect leaves [16], whereas excessive expansion of the adaxial cells would increase the LA [17,18].
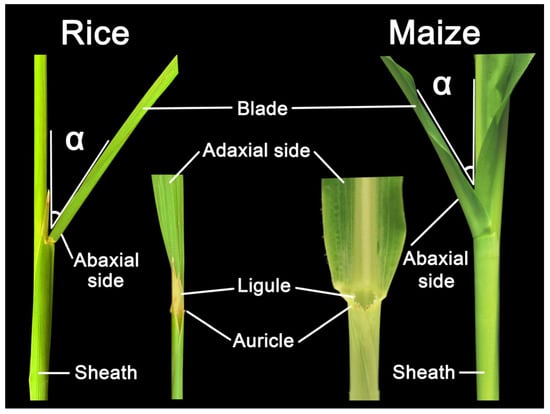
Figure 1.
Structural composition of leaf angle in rice and maize. Leaf angle is defined by the angle between the plant stem and leaf adaxial side of the blade, indicating as the α in the figure. This leaf morphology is affected by the development of ligule and auricle as well.
Given the rapid developments of plant functional genomics, a number of genes controlling LA have been cloned and the relevant regulatory network underlying the development of LA in grasses has been well characterized. These studies revealed that phytohormones, such as brassinosteroids (BRs), auxin, and gibberellins (GAs), comprehensively participate in regulating LA by orchestrating the homeostais of their biosynthesis and the expression of signaling transcription factors (TFs). Though each phytohormone and TF has been found to contribute quite similar traits/phenotypes in term of LA, how they interplay with each other is still far beyond understood. This review is an attempt to highlight the regulation mechanism of LA and the genetic interactions among phytohormones in regulating LA formation with particular emphasis on maize and rice.
2. Regulation of Lamina Joint Bending by Brassinosteroid (BR)
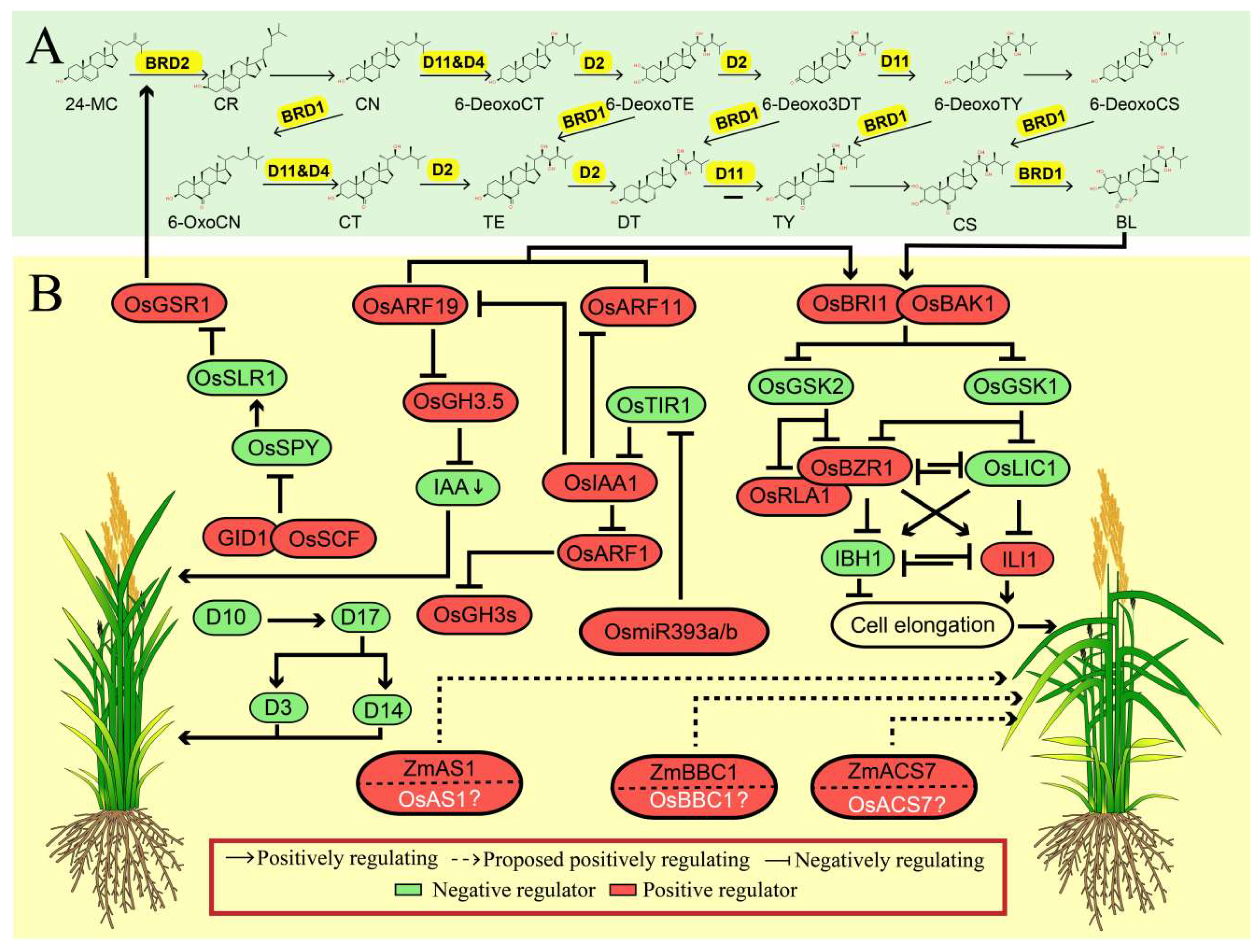
BRs are a group of steroid phytohormones involved in many important biological processes, and thus play a vital role in regulating several important agronomic traits, such as leaf angle, plant height and stress resistance. The regulatory pathways of BR biosynthesis, metabolism, and signal transduction have been well established in rice [19,20,21]. Since LA is a grass-species-specific trait, the role of BR in regulating LA has only been characterized in maize and rice rather than Arabidopsis, indicating that BR is a positive regulator of LA (Figure 2).
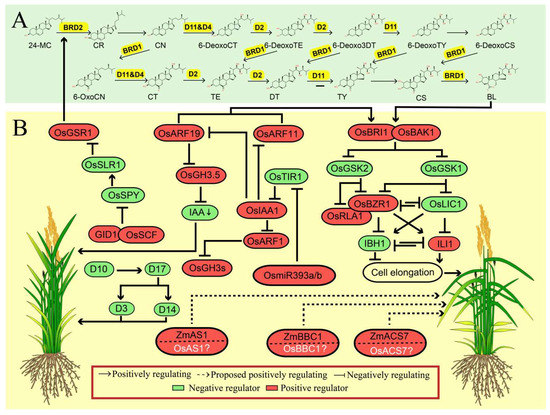
Figure 2.
The synergistic regulation mechanisms of leaf angle by phytohormones in rice. (A) Brassinosteroid biosynthesis pathway. The corresponding enzyme that catalyze each reaction in yellow color. The secondary structure of the chemical is obtained from ChemSpider (http://www.chemspider.com/Default.aspx). (B) Positive and negative regulation of leaf angle by brassinosteroid (BR) signaling pathway and its crosstalk with other regulators involved in other plant hormones pathway, and other phytohormones regulators positively participate in regulation of leaf angle. The proteins in white color and with a question mark represent that they are homologs of those reported to be involved in the regulation of leaf angle. The green arrow represents the downregulation of indoleacetic acid (IAA). The protein in red color box represents as the positive regulator while the one in green color box represents the negative regulator of leaf angle. The dashed line and solid line represent the indirect or direct evidence supporting the responsible regulation, respectively.
Up-regulation of the BR content within lamina joint region or enhanced BR signaling pathway by boosting the expression of BR related regulators resulted in increasing LA (Table 1). For instance, many researches in rice have elaborated that leaf inclination is closely associated with biosynthesis or signaling of BR [19,22]. Loss-of-function of BR biosynthetic genes, such as Dwarf 2 (D2), perturbed the endogenous level of BR, eventually resulting in erect leaf phenotype [11,23,24,25,26]. Besides, other BR biosynthetic regulators, such as the Cytochrome P450 family proteins, have also been documented to function in the development of LA. For example, BR-deficient Dwarf1 (OsBRD1) is cytochrome P450 protein and encodes a key enzyme (BR C-6 oxidase) catalyzing BR biosynthesis. Disruption of OsBRD1 causes pleiotropic effects, including severe dwarf phenotype, tortuous leaves, short panicles, small seeds, etc. [27,28]. Another two Cytochrome P450 proteins, OsDWARF4 and OsDWARF11, also catalyze the rate-limiting reaction (c-22 hydroxylation) of BR biosynthesis. OsDWARF4 is highly expressed in leaf blade and root, which is inhibited by BR but increased in BR-insensitive or -deficient mutants, indicating there is a feedback regulation on OsDWARF4 expression. Depletion of OsDWARF4 caused mild phenotype with erect leaves but not any detrimental effect on the development of leaf, inflorescence, and seed relative to wild-type plant [11,24]. Distinct from Osdwarf4 mutant, Osdwarf11 mutant showed a much more severe phenotype, including dwarfism in plants, erection of leaves, pollen abortion, and small grains. Further investigation revealed that OsDWARF4 and OsDWARF11 function redundantly in BRs biosynthesis, but OsDWARF11 performs a major role in BR biosynthesis pathway while OsDWARF4 plays the complementary role [25], which explained the different effects of them in term of plant growth and development, as well as the LA. It is worthy to mention that the Osdwarf4 mutant with erect leaf simultaneously promoted biomass production and higher yields than wild type at different planting densities, even without additional fertilization, indicating that this gene/allele is a potential candidate for sustainability increasing crop yield in limited land area [11]. Taken together, these studies have clearly illustrated that BR metabolism is responsible for the LA formation in crop.

Table 1.
Cloned genes associated with leaf angle in rice and maize.
In the BR signaling transduction pathway, BR is perceived by extracellular domain of BRASSINOSTEROID INSENSITIVE 1 (BRI1), a single transmembrane leucine-rich repeat receptor-like protein kinase (LRR-RLK), and its co-receptor BRI1-ASSOCIATED KINASE 1 (BAK1). In the absence of BR, BRI1′s kinase domain is deactivated by the negative factor BRI1 KINASE INHIBITOR 1 (BKI1); while BR is present, the BR compound binds to the extracellular domain of the BRI1 and BAK1. Subsequently, BRI1 phosphorylates BKI1, which results in the release of BKI1 from the plasma membrane, and then induce the phosphorylation of the kinase domain of BRI1 and BAK1, finally activating the initiation of BRI1-mediated signaling transduction [29]. The activation of BRI1 in turn phosphorylates downstream BR-SIGNALING KINASE1 (BSK1), CONSTITUTIVE DIFFERENTIAL GROWTH 1 (CDG1), and some of their homologs. Both BSK1 and CDG1/CDL1 phosphorylates BRI1-SUPPRESSOR 1 (BSU1) and subsequently activates BSU1. Activated BSU1 dephosphorylates and inactivates BRASSINOSTEROID INSENSITIVE 2 (BIN2) to release the suppression of BRASSINAZOLE-RESISTANT 1 (BZR1) and BRI1-EMS-SUPPRESSOR 1 (BES1), which are two key downstream transcription factors positively mediating BR responses. BZR1 and BES1 are rapidly dephosphorylated by Protein Phosphatase 2A (PP2A) family, leading to their nuclear accumulation and their regulation of thousands of BR-responsive genes expression [30]. Besides, in the absence of BR, the interaction of phosphorylated BZR1 and BES1 with 14-3-3 proteins, a group of conserved phosphopeptide-binding proteins, can lead to their cytoplasm retention and degradation, while in the presence of BR, dissociated BKI1 in cytoplasm can competitively bind to 14-3-3 proteins to inhibit this process [31]. Otherwise, the PPA2 dephosphorylates BRI1, BZR1 and BES1, relieving the binding of BZR1 and BES1 to 14-3-3 proteins, which regulates downstream genes in response to BR and induce diverse BR responses [32]. In addition to the BR biosynthetic genes mentioned above, BR signaling genes also play an essential role in regulating the development of LA in rice. For example, OsBRI1 is an ortholog of Arabidopsis BRI protein in rice. OsBRI1 participates in regulating various aspects of growth and development processes in rice, including the bending of lamina joint, the intercalary meristem formation, and the longitudinal elongation of internode cells. Therefore, depletion of OsBRI1 significantly perturbed the internode elongation and bending of the lamina joint [33]. OsBAK1 (a homologous gene to Arabidopsis BAK1) encodes SERK family receptor kinase protein and acts as co-receptor kinase for OsBRI1 to mediate BR signal transduction. Down-regulating the expression level of OsBAK1 produced a rice variety with erect leaf and normal reproduction so that considered as a promising target for improving rice grain yield [34]. Nevertheless, suppression of OsBZR1 expression by RNA interference (RNAi) in rice greatly altered the expression of BR-responsive genes and resulted in plant dwarfism and erect leaf, suggesting that OsBZR1 plays an central downstream role in BR signaling [35]. The rice 14-3-3 proteins have been proven to interact with and retain OsBZR1 in cytoplasm instead of nucleus, ultimately leading to inhibiting the function of OsBZR1. However, BR treatment can dissociate their interactions and activate OsBZR1, in turn causing the erect leaf phenotype [36]. Similarly, a rice zinc finger transcription factor, LEAF and TILLER ANGLE INCREASED CONTROLLER (OsLIC), also antagonized OsBZR1 to repress BR signaling in rice. Overexpression of OsLIC resulted in erect leaves by eliminating BR response, indicating that OsLIC1 negatively modulates leaf inclination in rice. OsLIC directly regulated the INCREASED LEAF INCLINATION 1 (ILI1), a positive regulator in lamina inclination, to oppose the action of BZR1 [36]. In addition, recent studies have identified an APETALA2 (AP2)/ERF (ethylene-responsive element binding factor) family transcription factor, Reduced Leaf Angle 1 (RLA1) that is identical to the SMALL ORGAN SIZE 1 (SMOS1), as a positive regulator of BR signaling, which physically interacted with OsBZR1 to enhance its transcriptional activity to enlarge LA in rice [37,38]. BU1 encodes a helix-loop-helix protein and acts as a novel BR positive regulator. Overexpressing BU1 in rice led to enhanced bending of the lamina joint, increased grain size, and resistance to BR, while repression of BU1 and its homologs in rice displays erect leaves [39]. Furthermore, a pair of antagonizing HLH/bHLH factors, OsILI1 that is the homology of Arabidopsis thaliana Paclobutrazol Resistance 1 (PRE1) and ILI1 binding bHLH 1 (OsIBH1), functioned as downstream factors of OsBZR1 to regulate leaf angle. OsILI1 positively regulated BR-mediated cell elongation, whereas OsIBH1 directly interacted with OsILI1 and performed an opposite role [40]. In addition, another bHLH transcription factor, BRASSINOSTEROID-RESPONSIVE LEAF ANGLE REGULATOR 1 (OsBLR1), has also been implicated to participate in LA regulation though the BR pathway in rice. Over-expressing OsBLR1 simultaneously increased leaf angle, grain length and sensitivity to BR, whereas mutation of OsBLR1 resulted in erect leaf and shorter grain [41]. Similarly, gain-of-function of OsbHLH079 also caused wide LA, longer grain and hypersensitive to BR in rice, while OsbHLH079-RNAi lines showed opposite phenotype [42]. Collectively, these findings suggested that bHLH family transcription factors may be broadly involved in BR-mediated LA regulation. Previously, a plant-specific gene family transcription factor, OsGRAS19 (GA INSENSITIVE (GAI), REPRESSOR OF GAI (RGA), and SCARECROW (SCR) 19) was suggested to be involved in regulating BR signaling, because the knockdown lines of OsGRAS19 displayed less sensitivity to the 24-epi-brassinolid (BL) treatment as compared to WT in rice. Higher expression of OsGRAS19 caused larger LA, narrow leaf and thin culm and panicle in rice [43]. Recently, a novel mutant of OsGRAS19, D26, has been identified in rice, which displayed typical BR-mediated phenotypes, including semidwarf, wider, and shorter leaf in addition to the erect leaf, as well as longer grain [44]. Taken together, these results further elaborated that the BR signaling is extensively involved in regulating LA in rice.
Until now, the molecular regulation of BR underlying LA in maize has been rarely investigated. Previous reports illustrated that suppression of ZmBRI1 or knock-out of Dwarf and Irregular Leaf 1 (ZmDIL1) displayed dwarf plant and erect leaves [45,46], suggesting a conserved function of BR for plant architecture in monocots. Recently, a SQUAMOSA-PROMOTER BINDING PROTEIN-LIKE (SPL) family protein LIGULELESS 1 (LG1), which is recognized as a conserved key factor in the formation of ligule and auricle in both rice and maize [47,48], confers an important role in controlling the leaf angle through regulating BR and auxin signaling pathway in maize and wheat. In maize, ZmLG1 activated a B3-domain containing transcription factor ZmRAVL1, which sequentially activated ZmBRD1 that was also designated as Upright Plant Architecture 1 (UPA1), eventually resulting in the alternation of leaf angle. UPA2 functioned as a distant cis element to regulate ZmRAVL1 expression, which was controlled by the Drooping Leaf 1 (DRL1). This UPA2-ZmRAV1-UPA1 module fine-tuned the BR pathway and finally determined the plant architecture and LA in maize [49]. In wheat, the LG1 ortholog, TaSPL8, directly bound to the promoter of an auxin response factor TaARF6 and the BR biosynthesis gene CYP90D2 (TaD2), and subsequently activated their expressions, leading to enhancing lamina joint development [50]. Recently, ZmILI1, an ortholog of OsILI1 that plays an important role in LA in rice, was found to directly bind to the ZmLG1 and CYP90D1 promoters to affect the BR biosynthesis and signal, eventually changing the LA [51]. Collectively, these studies supported a notion that components of BR signaling pathway play a dominant and conserved role in the formation of leaf angle in multiple crops and would be the potential targets for genetic improvement of crop architecture and yield.
3. Regulation of Lamina Joint Bending by Indoleacetic Acid (IAA)
Indoleacetic acid (IAA) is another crucial hormone, regulating leaf inclination mainly through patterning the adaxial/abaxial cell growth of leaves. In contrast to the positive role of BR in LA regulation, IAA functioned as a negative regulator since eliminating IAA content resulted in increased leaf inclination while increasing IAA content caused reduction of leaf inclination and upright leaves [52].
Auxin signal transduction is a sophisticated pathway, which is consisted of several key components, including the F-box TRANSPORT INHIBITOR RESPONSE 1/AUXIN SIGNALING F-BOX PROTEIN (TIR1/AFB) auxin co-receptors, the Auxin/INDOLE-3-ACETIC ACID (Aux/IAA) transcriptional repressors, and the AUXIN RESPONSE FACTOR (ARF) transcription factors and a ubiquitin-dependent protein degradation system. When IAA is deficient, Aux/IAA proteins form heterodimers with ARFs and block the function of ARFs to inhibit the auxin signal. Presence of auxin promotes the interaction between TIR1/AFB and Aux/IAA proteins, and then triggers a proteasome-mediated degradation of Aux/IAA by 26S Proteasome, eventually resulting in the release of ARF transcriptional activity. Subsequently, activation of ARF induces the changes of auxin-mediated gene expression pattern and growth responses [53,54,55,56]. This auxin signaling transduction model in rice has been implicated to associate with the regulation of lamina joint bending (Figure 1).
For instance, FISH BONE (FIB) encodes an orthologous of TAA protein, which plays a negative effect on leaf inclination. Loss-function of FIB caused a reduction of IAA level and altered auxin polar transport activity, thus producing small leaves with enlarged lamina joint angle [57]. LEAF INCLINATION1 (LC1) encoding an IAA amino synthetase, termed as OsGH3.1 in rice, maintained auxin homeostasis by catalyzing excess IAA binding to various amino acids. A gain-of-function mutant lc1-D in rice displayed a reducing content of free IAA and increasing leaf angle due to the promotion of cell elongation in the adaxial surface of lamina joint [58]. In addition, two OsMIR393a/b targeted auxin receptors, OsTIR1 and AUXIN SIGNALING f-box 2 (OsAFB2), have also been proven to be involved in the regulation of leaf angle. Overexpression of OsmiR393a/b repressed the expression of OsTIR1 and OsAFB2, which led to greater leaf angle in rice [59]. Further study demonstrated that BR promoted OsTIR1 and OsAFB2 to trigger the degradation of OsIAA1, which resulted in de-suppression of OsARF11 and OsARF19 proteins and consequently caused enlarge LA in rice [60]. Furthermore, a SPOC domain-containing transcription suppressor Leaf inclination 3 (LC3) regulated leaf inclination through interacting with a HIT zinc finger domain-containing protein, LC3-interacting protein 1 (LIP1). Meanwhile, LC3 could also directly bind to the promoter regions of OsIAA12 and OsGH3.2 to regulate auxin signal transduction and auxin homeostasis, finally influencing the formation of leaf inclination [61].
The maize auxin efflux carrier P-glycoprotein (ZmPGP1) was an adenosine triphosphate (ATP) binding cassette (ABC) transporter, which was involved in the polar transport of auxin and associated with the LA in maize [62]. Notably, multiple haplotypes of ZmPGP1 were present in various landraces, teosintes, and inbred accessions [63], suggesting a domesticated selection and improvement due to the demand of higher density planting.
4. Regulation of Lamina Joint Bending by Gibberellins (GA)
GA signaling pathway has also been implicated to participate in the regulation of the LA through the BR signaling dependent manner. Previously, transcriptomic profiling identified the co-expression pattern between GA and BR associated genes, however, it was less understood whether these genes were coordinated to regulate the LA. Recently, several researches have verified that certain GA genes interact with BR genes to modulate the LA. For instance, knockdown of the rice SPINDLY (OsSPY) caused the eliminated expression of BR biosynthesis genes D11, D2, and OsDWARF/BRD1, but increased the OsDWARF4 expression that functions downstream of the GA gene, SLENDER RICE 1 (SLR1), ultimately resulting in the enhanced leaf angle [64]. A recent study found that the Arabidopsis O-fucosyltransferase SPY could mono-O-fucosylate the DELLA protein, leading to higher affinity of interaction between DELLA and BZR1 [65]. From this point of view, it is supposed that OsSPY also could activate the OsSLR1 encoding a DELLA protein to interact with OsBZR1, thereby affecting LA in rice. The rice GA-stimulated transcript gene (OsGSR1) induced by GA is another downstream gene of SLR1, and it directly binds to BRD2 to enhance the BR biosynthesis, which subsequently altered the leaf angle [64,66]. In addition to these GA genes, other regulators involved in GA pathway have also been identified to participate in LA regulation. OsDCL3a encodes a Dicer-like endoribonuclease involved in generating siRNA. Down-regulation of the OsDCL3a resulted in increased leaf angle by modulating the expression of GA and BR associated genes, including OsGSR1 and BRD1 [67]. Recently, OsmiR396d has been reported to regulate the LA in rice, which was promoted by OsBZR1 and then regulated the expression of BR responsive genes by targeting the GROWTH REGULATING FACTOR 4 (OsGRF4) [68]. Alternatively, as another target of OsmiR396d, the OsGRF6 participated in GA biosynthesis and signal transduction but was not directly involved in BR signaling only modulated the plant height rather than LA [68]. Taken together, it is supposed that GA likely regulates LA dependent on the BR pathway, and thus identification of much more GA components involved in response to BR would extend our knowledge to this issue.
5. Regulation of Lamina Joint Bending by Crosstalk among Various Phytohormones
Recent studies have shown that crosstalk between IAA and BR cooperatively regulates the development of leaf angle. OsIAA1 is a key negative regulator for auxin signal transduction, which is induced by auxin and BR. Over-expression of OsIAA1 in rice resulted in dwarfism and increased leaf angle with decreased sensitivity to auxin treatment but increased sensitivity to BR treatment. OsARF1 is an auxin signal positive regulator which is inhibited by OsIAA1 in the absence of auxin. Further analysis showed that mutation of OsARF1 reduced sensitivity to BR treatment, resembling the phenotype of OsIAA1-overexpression plants, which indicated that BR may interact with auxin through the OsIAA1-OsARF1 module to regulate LA in rice [69,70]. OsARF19 acted as another coordinator of auxin and BR by positively regulating the expression of OsGH3.5 to reduce the content of free IAA on the one hand and activating OsBRI1 to stimulate BR signal cascades on the other hand, thus resulting in increased leaf angle [71]. RLA1/SMOS1 functioned downstream of the auxin signaling pathway, and enhanced the transcriptional activity of OsBZR1 by interacting with OsBZR1, suggesting that RLA1/SMOS1 integrated BR and IAA signal pathway to regulate the development of leaf angle [37,38]. In summary, the above studies showed that the auxin antagonized BR by interfering both BR metabolism and signaling to negatively regulate the leaf angle in rice.
Additionally, the rice d1 mutant with null function of an α subunit of G-protein (Gα), Rho GTPase activating protein 1 (RGA1), exhibited a dwarfism phenotype with erect leaves, and reduced sensitivity to GA and BR, indicating that RGA1 was involved in both GA and BR responses [29,72]. Further studies showed that D1/RGA1 interacted with TUD1 to induce BU1 expression, resulting in the increased leaf inclination [73,74]. BR can also act upstream of GA by modulating GA metabolism to regulate cell elongation. BR activated OsBZR1 and induced the expression of D18/GA3ox-2, one of the GA biosynthetic genes, leading to increased bioactive GA levels in rice seedlings. In contrast, GA extensively inhibited BR biosynthesis and the BR response with a feedback mechanism, so that GA treatment decreased the enlarged leaf angles in plants by attenuated BR biosynthesis or signaling [75]. These results showed that BR and GA were intertwined to regulate the leaf angle in rice.
6. Other Phytohormones Involved in Regulation of Lamina Joint
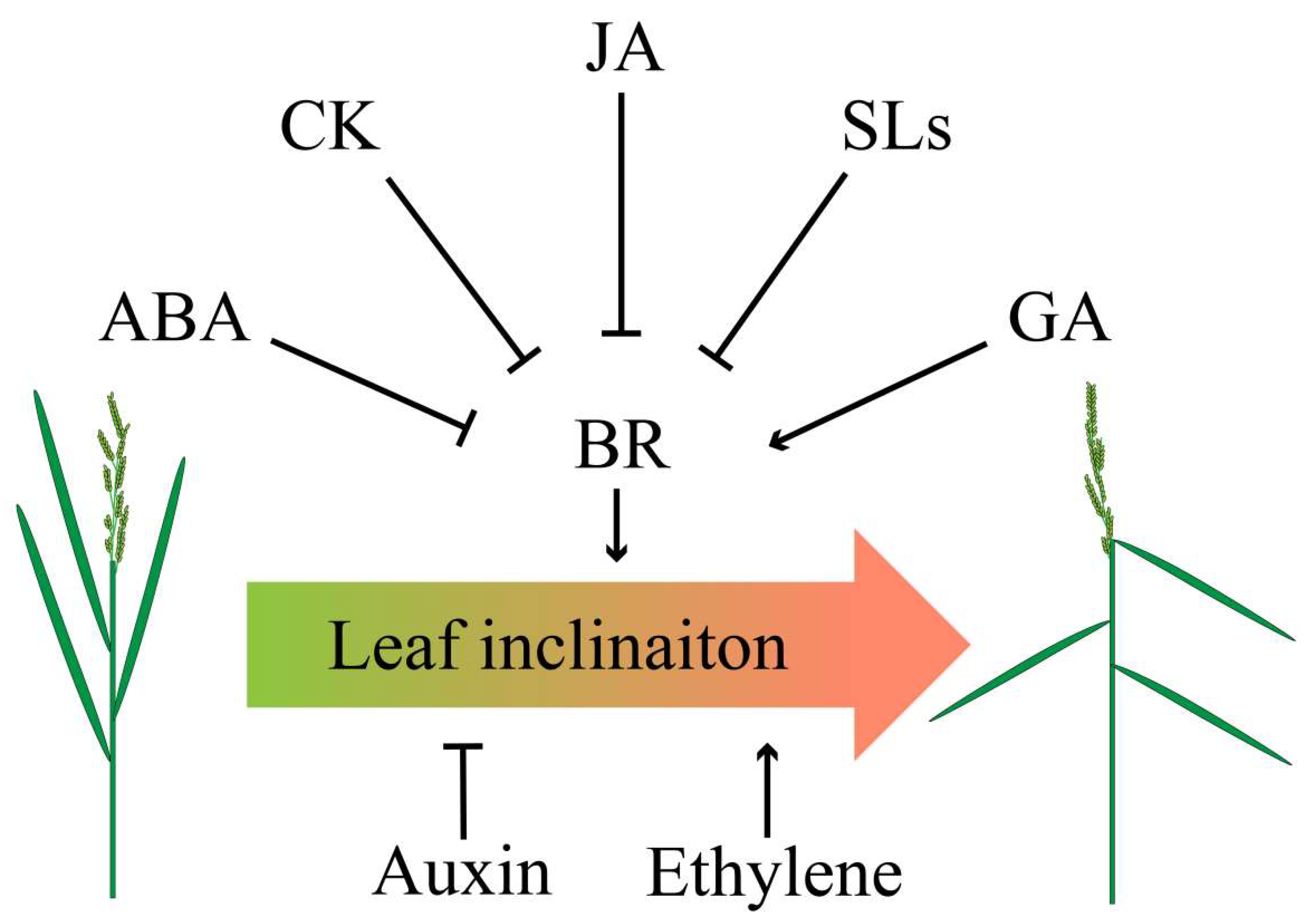
Phytohormones such as ethylene, strigolactones (SLs), jasmonic acid (JA), and abscisic acid (ABA) were also involved in regulating leaf inclination of rice (Figure 3). An early report demonstrated the interaction of ethylene and BR to regulate leaf inclination of rice, but the underlying mechanism remaining to be elusive [18]. Currently, a study has validated that altering the C terminus of 1-Aminocyclopropane-1-carboxylate (ACC) synthase 7 (ZmACS7) responsible for ethylene biosynthesis in maize led to the stability of this protein and the accumulation of ACC and ethylene contents, as well as the up-regulation of ethylene responsive genes, which finally reduced plant height and increased leaf angle [76]. Similar to ZmACS7, overexpression of its closest paralog ZmACS2 also resulted in flatter leaves [76], further suggesting that ethylene positively regulates LA.
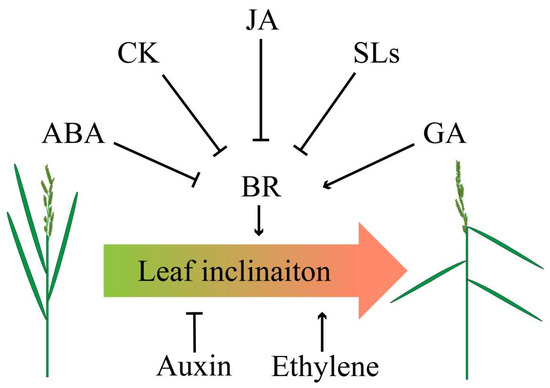
Figure 3.
Crosstalk of phytohormones in determining leaf angle. Abscisic acid (ABA), CK, jasmonic acid (JA), and strigolactones (SLs) negatively regulate BR, thereby inhibiting leaf angle, whereas gibberellins (GA) positively coordinates BR to increase leaf angle. Application of auxin results in erect leaf while ethylene leads to flat leaf.
SLs negatively regulate leaf inclination at seedling stage [77]. Interestingly, recent report showed that SLs also mediated leaf inclination in response to nutrient deficiencies in rice [78]. Similar to SLs, JA also showed a negative role in leaf inclination. JA treatment decreased lamina joint inclination by repressing the expression of BR biosynthesis-related genes which thus decreased endogenous BRs levels. Besides, inactivation of a negative regulator of BR signaling, GSK3-like kinase, partly rescued the inhibited effect of JA on lamina joint inclination, indicating that JA may disturb both BR biosynthesis and BR signaling pathway to limit lamina joint inclination [79].
Previous research has revealed that BR antagonized with ABA in regulating seed germination and hypocotyl elongation in Arabidopsis. The study showed that the ABSCISIC ACID INSENSITIVE5 (ABI5) directly interacts with BRASSINOSTEROID INSENSITIVE2 (BIN2), and then was phosphorylated and stabilized by BIN2 upon ABA treatment [80]. Additionally, the BES1 also physically interacted with ABI5 to hinder the expression of ABI5-targeted EARLY METHIONINE-LABELED 1 (EM1) and EM6 [81], eventually facilitating the seed germination in Arabidopsis. Interestingly, the ABI1 and ABI2 can interact with and dephosphorylate BIN2 to attenuate the BR signaling in Arabidopsis [82], indicating a complicated crosstalk between ABA and BR. Recently, there is research which further revealed that ABA also antagonized BR to regulate the lamina joint inclination in rice by targeting the BR biosynthesis gene D11 and BR signaling genes GSK2 and DLT [83], and it therefore raises an issue that whether the rice homologs of Arabidopsis ABI1 and ABI2 may also interfere the BR-mediated LA. Investigation of the LA phenotype of the Osabi1 and Osabi2, as well as overexpression lines of these two genes, may provide the answer for this hypothesis. Another research preprinted in bioRxiv demonstrated that a transcriptional repressor ZmCLA4 (the ortholog of LAZY1 in rice and Arabidopsis) responsible for multiple phytohormone mediated pathways negatively regulated LA by altering mRNA accumulation. Further analysis showed that ZmCLA4 could directly bind to two key components of BR signaling ZmBZR3 and 14-3-3, and two important responsive transcription factors of ABA ZmWRKY4 and ZmWRKY72 respectively, thus mediating the crosstalk between BR and ABA in LA regulation [84]. In maize, a bHLH transcription factor ZmIBH1-1 is a negative regulator of LA in maize. Transcriptome analysis suggested that the ZmIBH1-1-mediated LA in the leaf ligular region was highly correlated with cytokinin (CK), JA, and ethylene synthesis and signal transduction pathways associated genes, in particular the two CK responsive genes, GRMZM2G145280 (BBC1) and GRMZM2G149952 (ZmAS1) that were tightly correlated with cell division, implying that CK may modulate LA through the control of cell profile [85]. Notably, it has been demonstrated that CK indirectly interacted with BR through auxin pathway. For example, BR regulated the development of root primordia through increasing the PIN genes expression while the CK inhibited the root primordia by repressing PIN genes [86]. However, it was also found that CK was accumulated in the young seedling of wheat with BR treatment [87], whereas overexpression of the rice Isopentyl Transferase (IPT) driven by the promoter of stress- and maturation-inducible gene, Senescence-associated Receptor Kinase (SARK), resulted in up-regulation of BR genes, including DWF4, BRI1, and BZR1 etc., ultimately leading to higher rice yield during water-stress [88]. These studies further suggest that CK interplays with BR in regulating plant growth and stress response. However, it is still unclear whether and how they cooperated in the LA regulation. As mentioned above, the bHLH family generally integrated BR signaling to regulate the LA, such as the OsILI1 and OsBLR1, and thus we proposed that the CK may interplay with BR to regulate LA through the bHLH-BBC1/AS1 module mediated BR signaling.
In summary, almost all of the regulators modulated leaf angle through the phytohormones-dependent manner (Table 1), however, whether there are unknown components independent on phytohormones, the pathway still remains elusive.
7. Future Perspectives
Leaf angle directly influences the shape of plant architecture, consequentially affecting yield. To feed the ever-increasing global population, demand of higher crop yield has triggered the breeders to breed new cultivars that maintain sustainable productivity and yield by dense planting with limited arable lands. Therefore, how to genetically manipulating leaf angle has become one of the most important tasks to be tackled in crop genetic improvement. Until now, the regulatory mechanism underlying leaf angle has been extensively elucidated. However, a few issues are still beyond understanding, preventing the application of specific gene resource in term of crop improvement. To address these issues, the relationship among these genes still needs to be further uncovered, in particular the interaction network. On another hand, knockout or overexpression of these genes generally caused pleiotropic effects on plant growth and development in addition to the LA, such as plant dwarfism and smaller grain. Therefore, identification of interest single nucleotide polymorphisms (SNPs) and/or haplotypes of LA genes could not only be an efficient strategy to extent our knowledge about the relevant regulatory network, but also provide suitable alleles for marker selection breeding. The studies of ZmLG1 provide an excellent case for bridging the basic research and application. ZmLG1 was initially identified as a key factor for formation of ligule and auricle in maize and rice, and then regard to be an important player during rice domestication [47,90,91]. Further large-scale genetic analysis revealed that ZmLG1 was the major QTL controlling leaf angle in maize [92,93,94,95]. Based on these studies, a recent study demonstrated that genetic manipulation of ZmLG1 gene can significantly increase photosynthetic efficiency and maize yield at higher planting density [96]. Therefore, genome resequencing of larger accessions in crop indeed facilitates deciphering interest haplotypes of LA regulators. Alternatively, generation of novel alleles by CRISPR-mediated gene editing is also a powerful and efficient approach regarding this issue. Nevertheless, it would be fascinating to ask if there are novel regulators involved in LA regulation independent on the phytohormones pathways, which might only modulate LA without pleiotropic effects on other traits. On the other hand, it is also attractive to see whether and how LA coordinates or integrates with other agronomic traits in promoting yield, since yield consists of numerous components, such as plant height and tiller number. Furthermore, manipulating the spatial and temporal expression pattern of LA genes may be also important for timely shaping the plant architecture to achieve optimal photosynthesis, as well as eliminating shade avoidance and neighbor interference.
In conclusion, BR signaling pathway performs along with other plant hormones to form a complex signaling crosstalk to coordinate LA architecture under regular growth condition and in response to environmental stimuli. The well-established regulatory network for LA would provide vast promising targets to be manipulated, no matter through traditional molecular marker-assisted breeding or the gene editing technology, for crop architecture improvement to pursue high production.
Author Contributions
Writing—original draft preparation, X.L., R.S. and Q.X.; writing—review and editing, X.L., P.W., Y.L., Z.Z., R.S. and Q.X.; visualization, X.L., P.W., Y.L., Z.Z. and S.G.; supervision, Q.X.; project administration, S.G. and Q.X.; funding acquisition, R.S. and Q.X. All authors have read and agreed to the published version of the manuscript.
Funding
This work was supported by the Major Program of Guangdong Basic and Applied Research (2019B030302006), the National Natural Science Foundation of China (31971920) and the support from the “Top Young Scientist of the Pearl River Talent Plan” (No. 20170104) to Qingjun Xie, as well as the National Natural Science Foundation of China (31771739) and the Natural Science Foundation of Guangdong Province-Guangzhou City Collaborative Key Project (2019B1515120061) to Rongxin Shen.
Acknowledgments
We are grateful to the supports of experimental platform and funding from the Guangdong Provincial Key Laboratory of Plant Molecular Breeding (No. GPKLPMB201804). We apologize in advance to colleagues whose valuable work was not cited due to article length considerations.
Conflicts of Interest
The authors declare no conflict of interest.
Abbreviations
| LA | Leaf angle |
| BR | Brassinosteroid |
| IAA | Indoleacetic acid |
| GA | Gibberellin |
| SLs | Strigolactones |
| JA | Jasmonic acid |
| ABA | Abscisic acid |
References
- Duvick, D.N. The Contribution of breeding to yield advances in maize (Zea mays L.). Adv. Agron. 2005, 86, 83–145. [Google Scholar] [CrossRef]
- Duvick, D.N. Genetic progress in yield of United States maize (Zea mays L.). Maydica 2005, 50, 193–202. [Google Scholar]
- Leivar, P.; Tepperman, J.M.; Cohn, M.M.; Monte, E.; Al-Sady, B.; Erickson, E.; Quail, P.H. Dynamic antagonism between phytochromes and pif family basic helix-loop-helix factors induces selective reciprocal responses to light and shade in a rapidly responsive transcriptional network in arabidopsis. Plant Cell 2012, 24, 1398–1419. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Khush, G.S. Green revolution: Preparing for the 21st century. Genome 1999, 42, 646–655. [Google Scholar] [CrossRef] [PubMed]
- Sasaki, A.; Ashikari, M.; Ueguchi-Tanaka, M.; Itoh, H.; Nishimura, A.; Swapan, D.; Ishiyama, K.; Saito, T.; Kobayashi, M.; Khush, G.S.; et al. Green revolution: A mutant gibberellin-synthesis gene in rice. Nature 2002, 416, 701–702. [Google Scholar] [CrossRef] [PubMed]
- Wu, K.; Wang, S.; Song, W.; Zhang, J.; Wang, Y.; Liu, Q.; Yu, J.; Ye, Y.; Li, S.; Chen, J.; et al. Enhanced sustainable green revolution yield via nitrogen-responsive chromatin modulation in rice. Science 2020, 367, eaaz2046. [Google Scholar] [CrossRef] [PubMed]
- Sharman, B.C. Developmental anatomy of the shoot of Zea mays L. Ann. Bot. 1942, 6, 245–282. [Google Scholar] [CrossRef]
- Sylvester, A.W.; Cande, W.Z.; Freeling, M. Division and differentiation during normal and liguleless-1 maize leaf development. Development 1990, 110, 985–1000. [Google Scholar]
- Mantilla-Perez, M.B.; Fernandez, M.G.S. Differential manipulation of leaf angle throughout the canopy: Current status and prospects. J. Exp. Bot. 2017, 68, 5699–5717. [Google Scholar] [CrossRef] [Green Version]
- Duvick, D.N.; Cassman, K. Post-green revolution trends in yield potential of temperate maize in the north-central united states. Crop. Sci. 1999, 39, 1622–1630. [Google Scholar] [CrossRef]
- Sakamoto, T.; Morinaka, Y.; Ohnishi, T.; Sunohara, H.; Fujioka, S.; Ueguchi-Tanaka, M.; Mizutani, M.; Sakata, K.; Takatsuto, S.; Yoshida, S.; et al. Erect leaves caused by brassinosteroid deficiency increase biomass production and grain yield in rice. Nat. Biotechnol. 2005, 24, 105–109. [Google Scholar] [CrossRef] [PubMed]
- Sinclair, T.R. Erect leaves and photosynthesis in rice. Science 1999, 283, 1456. [Google Scholar] [CrossRef]
- Kong, F.; Zhang, T.; Liu, J.; Heng, S.; Shi, Q.; Zhang, H.; Wang, Z.; Ge, L.; Li, P.; Lu, X.; et al. Regulation of leaf angle by auricle development in maize. Mol. Plant 2017, 10, 516–519. [Google Scholar] [CrossRef] [Green Version]
- Luo, X.; Zheng, J.; Huang, R.; Huang, Y.; Wang, H.; Jiang, L.; Fang, X. Phytohormones signaling and crosstalk regulating leaf angle in rice. Plant Cell Rep. 2016, 35, 2423–2433. [Google Scholar] [CrossRef] [PubMed]
- Zhou, L.; Xiao, L.; Xue, H.-W. Dynamic cytology and transcriptional regulation of rice lamina joint development. Plant Physiol. 2017, 174, 1728–1746. [Google Scholar] [CrossRef] [Green Version]
- Hong, Z.; Ueguchi-Tanaka, M.; Matsuoka, M. Brassinosteroids and rice architecture. J. Pestic. Sci. 2004, 29, 184–188. [Google Scholar] [CrossRef] [Green Version]
- Duan, K.; Li, L.; Hu, P.; Xu, S.-P.; Xu, Z.-H.; Xue, H.-W. A brassinolide-suppressed rice MADS-box transcription factor, OsMDP1, has a negative regulatory role in BR signaling. Plant J. 2006, 47, 519–531. [Google Scholar] [CrossRef]
- Cao, H.; Chen, S. Brassinosteroid-induced rice lamina joint inclination and its relation to indole-3-acetic acid and ethylene. Plant Growth Regul. 1995, 16, 189–196. [Google Scholar] [CrossRef]
- Wang, Z.-Y.; Bai, M.-Y.; Oh, E.; Zhu, J.-Y. Brassinosteroid signaling network and regulation of photomorphogenesis. Annu. Rev. Genet. 2012, 46, 701–724. [Google Scholar] [CrossRef]
- Zhang, C.; Bai, M.-Y.; Chong, K. Brassinosteroid-mediated regulation of agronomic traits in rice. Plant Cell Rep. 2014, 33, 683–696. [Google Scholar] [CrossRef]
- Tong, H.; Chu, C. Functional specificities of brassinosteroid and potential utilization for crop improvement. Trends Plant Sci. 2018, 23, 1016–1028. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.; Bai, M.-Y.; Wang, Z.-Y. The brassinosteroid signaling network — a paradigm of signal integration. Curr. Opin. Plant Biol. 2014, 21, 147–153. [Google Scholar] [CrossRef] [Green Version]
- Wu, Y.; Fu, Y.; Zhao, S.; Gu, P.; Zhu, Z.; Sun, C.; Tan, L. Clustered primary branch 1, a new allele of DWARF11, controls panicle architecture and seed size in rice. Plant Biotechnol. J. 2015, 14, 377–386. [Google Scholar] [CrossRef] [PubMed]
- Morinaka, Y.; Sakamoto, T.; Inukai, Y.; Agetsuma, M.; Kitano, H.; Ashikari, M.; Matsuoka, M. Morphological alteration caused by brassinosteroid insensitivity increases the biomass and grain production of rice1. Plant Physiol. 2006, 141, 924–931. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tanabe, S.; Ashikari, M.; Fujioka, S.; Takatsuto, S.; Yoshida, S.; Yano, M.; Yoshimura, A.; Kitano, H.; Matsuoka, M.; Fujisawa, Y.; et al. A novel cytochrome p450 is implicated in brassinosteroid biosynthesis via the characterization of a rice dwarf mutant, dwarf11, with reduced seed length. Plant Cell 2005, 17, 776–790. [Google Scholar] [CrossRef] [Green Version]
- Hong, Z.; Ueguchi-Tanaka, M.; Fujioka, S.; Takatsuto, S.; Yoshida, S.; Hasegawa, Y.; Ashikari, M.; Kitano, H.; Matsuoka, M. The rice brassinosteroid-deficient dwarf2 mutant, defective in the rice homolog of arabidopsis DIMINUTO/DWARF1, is rescued by the endogenously accumulated alternative bioactive brassinosteroid, dolichosterone. Plant Cell 2005, 17, 2243–2254. [Google Scholar] [CrossRef] [Green Version]
- Mori, M.; Nomura, T.; Ooka, H.; Ishizaka, M.; Yokota, T.; Sugimoto, K.; Okabe, K.; Kajiwara, H.; Satoh, K.; Yamamoto, K.; et al. Isolation and characterization of a rice dwarf mutant with a defect in brassinosteroid biosynthesis1. Plant Physiol. 2002, 130, 1152–1161. [Google Scholar] [CrossRef] [Green Version]
- Hong, Z.; Ueguchi-Tanaka, M.; Shimizu-Sato, S.; Inukai, Y.; Fujioka, S.; Shimada, Y.; Takatsuto, S.; Agetsuma, M.; Yoshida, S.; Watanabe, Y.; et al. Loss-of-function of a rice brassinosteroid biosynthetic enzyme, C-6 oxidase, prevents the organized arrangement and polar elongation of cells in the leaves and stem. Plant J. 2002, 32, 495–508. [Google Scholar] [CrossRef]
- Wang, L.; Xu, Y.-Y.; Ma, Q.-B.; Li, D.; Xu, Z.-H.; Chong, K. Heterotrimeric G protein α subunit is involved in rice brassinosteroid response. Cell Res. 2006, 16, 916–922. [Google Scholar] [CrossRef]
- Sun, Y.; Fan, X.-Y.; Cao, D.-M.; Tang, W.; He, K.; Zhu, J.-Y.; He, J.-X.; Bai, M.-Y.; Zhu, S.; Oh, E.; et al. Integration of brassinosteroid signal transduction with the transcription network for plant growth regulation in Arabidopsis. Dev. Cell 2010, 19, 765–777. [Google Scholar] [CrossRef] [Green Version]
- Wang, H.; Yang, C.; Zhang, C.; Wang, N.; Lu, D.; Wang, J.; Zhang, S.; Wang, Z.-X.; Ma, H.; Wang, X. Dual Role of BKI1 and 14-3-3 s in Brassinosteroid Signaling to Link Receptor with Transcription Factors. Dev. Cell 2011, 21, 825–834. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tang, W.; Yuan, M.; Wang, R.; Yang, Y.; Wang, C.; Oses-Prieto, J.A.; Kim, T.-W.; Zhou, H.-W.; Deng, Z.; Gampala, S.S.; et al. PP2A activates brassinosteroid-responsive gene expression and plant growth by dephosphorylating BZR1. Nat. Cell Biol. 2011, 13, 124–131. [Google Scholar] [CrossRef] [PubMed]
- Yamamuro, C.; Ihara, Y.; Wu, X.; Noguchi, T.; Fujioka, S.; Takatsuto, S.; Ashikari, M.; Kitano, H.; Matsuoka, M. Loss of function of a rice brassinosteroid insensitive1 homolog prevents internode elongation and bending of the lamina joint. Plant Cell 2000, 12, 1591–1605. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, D.; Wang, L.; Wang, M.; Xu, Y.-Y.; Luo, W.; Liu, Y.-J.; Xu, Z.-H.; Li, J.; Chong, K. EngineeringOsBAK1gene as a molecular tool to improve rice architecture for high yield. Plant Biotechnol. J. 2009, 7, 791–806. [Google Scholar] [CrossRef] [PubMed]
- Bai, M.-Y.; Zhang, L.-Y.; Gampala, S.S.; Zhu, S.-W.; Song, W.-Y.; Chong, K.; Wang, Z.-Y. Functions of OsBZR1 and 14-3-3 proteins in brassinosteroid signaling in rice. Proc. Natl. Acad. Sci. USA 2007, 104, 13839–13844. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, C.; Xu, Y.; Guo, S.; Zhu, J.; Huan, Q.; Liu, H.; Wang, L.; Luo, G.; Wang, X.; Chong, K. Dynamics of brassinosteroid response modulated by negative regulator LIC in rice. PLoS Genet. 2012, 8, e1002686. [Google Scholar] [CrossRef] [Green Version]
- Qiao, S.; Sun, S.; Wang, L.; Wu, Z.; Li, C.; Li, X.; Wang, T.; Leng, L.; Tian, W.; Lu, T.; et al. The RLA1/SMOS1 transcription factor functions with OsBZR1 to regulate brassinosteroid signaling and rice architecture. Plant Cell 2017, 29, 292–309. [Google Scholar] [CrossRef] [Green Version]
- Hirano, K.; Yoshida, H.; Aya, K.; Kawamura, M.; Hayashi, M.; Hobo, T.; Sato-Izawa, K.; Kitano, H.; Ueguchi-Tanaka, M.; Matsuoka, M. Small organ size 1 and small organ size 2/dwarf and low-tillering form a complex to integrate auxin and brassinosteroid signaling in rice. Mol. Plant 2017, 10, 590–604. [Google Scholar] [CrossRef] [Green Version]
- Tanaka, A.; Nakagawa, H.; Tomita, C.; Shimatani, Z.; Ohtake, M.; Nomura, T.; Jiang, C.-J.; Dubouzet, J.G.; Kikuchi, S.; Sekimoto, H.; et al. Brassinosteroid upregulated1, encoding a Helix-Loop-Helix Protein, Is a novel gene involved in brassinosteroid signaling and controls bending of the lamina joint in rice1. Plant Physiol. 2009, 151, 669–680. [Google Scholar] [CrossRef] [Green Version]
- Zhang, L.-Y.; Bai, M.-Y.; Wu, J.; Zhu, J.-Y.; Wang, H.; Zhang, Z.; Wang, W.; Sun, Y.; Zhao, J.; Sun, X.; et al. Antagonistic HLH/bHLH transcription factors mediate brassinosteroid regulation of cell elongation and plant development in rice and Arabidopsis. Plant Cell 2009, 21, 3767–3780. [Google Scholar] [CrossRef] [Green Version]
- Wang, K.; Li, M.-Q.; Chang, Y.-P.; Zhang, B.; Zhao, Q.-Z.; Zhao, W.-L. The basic helix-loop-helix transcription factor OsBLR1 regulates leaf angle in rice via brassinosteroid signalling. Plant Mol. Biol. 2020, 102, 589–602. [Google Scholar] [CrossRef] [PubMed]
- Seo, H.; Kim, S.-H.; Lee, B.-D.; Lim, J.-H.; Lee, S.-J.; An, G.; Paek, N.-C. The rice basic Helix–Loop–Helix 79 (OsbHLH079) determines leaf angle and grain shape. Int. J. Mol. Sci. 2020, 21, 2090. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, L.; Xiong, G.; Cui, X.; Yan, M.; Xu, T.; Qian, Q.; Xue, Y.; Li, J.; Wang, Y. OsGRAS19 may be a novel component involved in the brassinosteroid signaling pathway in rice. Mol. Plant 2013, 6, 988–991. [Google Scholar] [CrossRef] [Green Version]
- Lin, Z.; Yan, J.; Su, J.; Liu, H.; Hu, C.; Li, G.; Wang, F.; Lin, Y. Novel OsGRAS19 mutant, D26, positively regulates grain shape in rice (Oryza sativa). Funct. Plant Biol. 2019, 46, 857. [Google Scholar] [CrossRef]
- Jiang, F.; Guo, M.; Yang, F.; Duncan, K.; Jackson, D.; Rafalski, A.; Wang, S.; Li, B. Mutations in an AP2 transcription factor-like gene affect internode length and leaf shape in maize. PLoS ONE 2012, 7, e37040. [Google Scholar] [CrossRef] [Green Version]
- Kir, G.; Ye, H.; Nelissen, H.; Neelakandan, A.; Kusnandar, A.S.; Luo, A.; Inzé, D.; Sylvester, A.W.; Yin, Y.; Becraft, P.W. RNA interference knockdown of brassinosteroid insensitive1 in maize reveals novel functions for brassinosteroid signaling in controlling plant architecture. Plant Physiol. 2015, 169, 826–839. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Moreno, M.A.; Harper, E.C.; Krueger, R.W.; Dellaporta, S.L.; Freeling, M. liguleless1 encodes a nuclear-localized protein required for induction of ligules and auricles during maize leaf organogenesis. Genes Dev. 1997, 11, 616–628. [Google Scholar] [CrossRef] [Green Version]
- Lee, J.; Park, J.; Kim, S.L.; Yim, J.; An, G. Mutations in the rice liguleless gene result in a complete loss of the auricle, ligule, and laminar joint. Plant Mol. Biol. 2007, 65, 487–499. [Google Scholar] [CrossRef]
- Tian, J.; Wang, C.; Xia, J.; Wu, L.; Xu, G.; Wu, W.; Li, D.; Qin, W.; Han, X.; Chen, Q.; et al. Teosinte ligule allele narrows plant architecture and enhances high-density maize yields. Science 2019, 365, 658–664. [Google Scholar] [CrossRef]
- Liu, K.; Cao, J.; Yu, K.; Liu, X.; Gao, Y.; Chen, Q.; Zhang, W.; Peng, H.; Du, J.; Xin, M.; et al. Wheat TaSPL8 modulates leaf angle through auxin and brassinosteroid signaling. Plant Physiol. 2019, 181, 179–194. [Google Scholar] [CrossRef] [Green Version]
- Ren, Z.; Wu, L.; Ku, L.; Wang, H.; Zeng, H.; Su, H.; Wei, L.; Dou, D.; Liu, H.; Cao, Y.; et al. ZmILI1 regulates leaf angle by directly affecting liguleless1 expression in maize. Plant Biotechnol. J. 2019, 18, 881–883. [Google Scholar] [CrossRef] [Green Version]
- Fellner, M.; Horton, L.A.; Cocke, A.E.; Stephens, N.R.; Ford, E.D.; Van Volkenburgh, E. Light interacts with auxin during leaf elongation and leaf angle development in young corn seedlings. Planta 2003, 216, 366–376. [Google Scholar] [CrossRef]
- Mockaitis, K.; Estelle, M. Auxin receptors and plant development: A new signaling paradigm. Annu. Rev. Cell Dev. Biol. 2008, 24, 55–80. [Google Scholar] [CrossRef] [Green Version]
- Hagen, G. Auxin signal transduction. Essays Biochem. 2015, 58, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Vanneste, S.; Friml, J. Auxin: A trigger for change in plant development. Cell 2009, 136, 1005–1016. [Google Scholar] [CrossRef] [PubMed]
- Lavy, M.; Estelle, M. Mechanisms of auxin signaling. Development 2016, 143, 3226–3229. [Google Scholar] [CrossRef] [Green Version]
- Yoshikawa, T.; Ito, M.; Sumikura, T.; Nakayama, A.; Nishimura, T.; Kitano, H.; Yamaguchi, I.; Koshiba, T.; Hibara, K.-I.; Nagato, Y.; et al. The rice FISH BONE gene encodes a tryptophan aminotransferase, which affects pleiotropic auxin-related processes. Plant J. 2014, 78, 927–936. [Google Scholar] [CrossRef] [PubMed]
- Zhao, S.-Q.; Xiang, J.-J.; Xue, H.-W. Studies on the rice leaf inclination1 (lc1), an iaa–amido synthetase, reveal the effects of auxin in leaf inclination control. Mol. Plant 2013, 6, 174–187. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bian, H.; Xie, Y.; Guo, F.; Han, N.; Ma, S.; Zeng, Z.; Wang, J.; Yang, Y.; Zhu, M. Distinctive expression patterns and roles of the miRNA393/TIR1 homolog module in regulating flag leaf inclination and primary and crown root growth in rice (Oryza sativa). New Phytol. 2012, 196, 149–161. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Yang, C.Y.; Miao, R.; Zhou, C.L.; Cao, P.H.; Lan, J.; Zhu, X.J.; Mou, C.L.; Huang, Y.S.; Liu, S.J.; et al. DS1/OsEMF1 interacts with OsARF11 to control rice architecture by regulation of brassinosteroid signaling. Rice 2018, 11, 46. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, S.-H.; Zhou, L.; Xu, P.; Xue, H.-W. SPOC domain-containing protein Leaf inclination3 interacts with LIP1 to regulate rice leaf inclination through auxin signaling. PLoS Genet. 2018, 14, e1007829. [Google Scholar] [CrossRef] [PubMed]
- Wei, L.; Zhang, X.; Zhang, Z.; Liu, H.; Lin, Z. A new allele of the Brachytic2 gene in maize can efficiently modify plant architecture. Heredity 2018, 121, 75–86. [Google Scholar] [CrossRef] [PubMed]
- Li, P.; Wei, J.; Wang, H.; Fang, Y.; Yin, S.; Xu, Y.; Liu, J.; Yang, Z.; Xu, C. Natural variation and domestication selection of zmpgp1 affects plant architecture and yield-related traits in maize. Genes 2019, 10, 664. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shimada, A.; Ueguchi-Tanaka, M.; Sakamoto, T.; Fujioka, S.; Takatsuto, S.; Yoshida, S.; Sazuka, T.; Ashikari, M.; Matsuoka, M. The riceSPINDLYgene functions as a negative regulator of gibberellin signaling by controlling the suppressive function of the DELLA protein, SLR1, and modulating brassinosteroid synthesis. Plant J. 2006, 48, 390–402. [Google Scholar] [CrossRef] [PubMed]
- Zentella, R.; Sui, N.; Barnhill, B.; Hsieh, W.-P.; Hu, J.; Shabanowitz, J.; Boyce, M.; Olszewski, N.E.; Zhou, P.; Hunt, N.F.; et al. The Arabidopsis O-fucosyltransferase spindly activates nuclear growth repressor DELLA. Nat. Methods 2017, 13, 479–485. [Google Scholar] [CrossRef] [Green Version]
- Wan, H.; Wang, J.; Liang, S.; Fang, H.; Xiao, Z. Estimating leaf area index by fusing MODIS and MISR data. In Proceedings of the 2006 IEEE International Symposium on Geoscience and Remote Sensing, Denver, CO, USA, 31 July–4 August 2006; Volume 29, pp. 1820–1823. [Google Scholar] [CrossRef] [Green Version]
- Wei, L.; Gu, L.; Song, X.; Cui, X.; Lu, Z.; Zhou, M.; Wang, L.; Hu, F.; Zhai, J.; Meyers, B.C.; et al. Dicer-like 3 produces transposable element-associated 24-nt siRNAs that control agricultural traits in rice. Proc. Natl. Acad. Sci. USA 2014, 111, 3877–3882. [Google Scholar] [CrossRef] [Green Version]
- Tang, Y.; Liu, H.; Guo, S.; Wang, B.; Li, Z.; Chong, K.; Xu, Y. OsmiR396d affects gibberellin and brassinosteroid signaling to regulate plant architecture in rice. Plant Physiol. 2017, 176, 946–959. [Google Scholar] [CrossRef] [Green Version]
- Attia, K.A.; Abdelkhalik, A.F.; Ammar, M.H.; Wei, C.; Yang, J.; Lightfoot, D.A.; El-Sayed, W.M.; El-Shemy, H.A. Antisense phenotypes reveal a functional expression of OsARF1, an auxin response factor, in transgenic rice. Curr. Issues Mol. Biol. 2009, 11. [Google Scholar]
- Song, Y.; You, J.; Xiong, L. Characterization of OsIAA1 gene, a member of rice Aux/IAA family involved in auxin and brassinosteroid hormone responses and plant morphogenesis. Plant Mol. Biol. 2009, 70, 297–309. [Google Scholar] [CrossRef]
- Zhang, S.; Wang, S.; Xu, Y.; Yu, C.; Shen, C.; Qian, Q.; Geißler, M.; Jiang, D.A.; Qi, Y. The auxin response factor, OsARF19, controls rice leaf angles through positively regulating OsGH 3-5 and OsBRI 1. Plant Cell Environ. 2014, 38, 638–654. [Google Scholar] [CrossRef] [Green Version]
- Ferrero-Serrano, Á.; Assmann, S.M. The α-subunit of the rice heterotrimeric G protein, RGA1, regulates drought tolerance during the vegetative phase in the dwarf rice mutant d1. J. Exp. Bot. 2016, 67, 3433–3443. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Oki, K.; Inaba, N.; Kitagawa, K.; Fujioka, S.; Kitano, H.; Fujisawa, Y.; Kato, H.; Iwasaki, Y. Function of the subunit of rice heterotrimeric g protein in brassinosteroid signaling. Plant Cell Physiol. 2008, 50, 161–172. [Google Scholar] [CrossRef]
- Jang, S.; An, G.; Li, H.-Y. Rice leaf angle and grain size are affected by the OsBUL1 transcriptional activator complex. Plant Physiol. 2016, 173, 688–702. [Google Scholar] [CrossRef] [PubMed]
- Tong, H.; Xiao, Y.; Liu, D.; Gao, S.; Liu, L.; Yin, Y.; Jin, Y.; Qian, Q.; Chu, C. Brassinosteroid regulates cell elongation by modulating gibberellin metabolism in rice. Plant Cell 2014, 26, 4376–4393. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, H.; Wang, L.; Liu, M.; Dong, Z.; Li, Q.; Fei, S.; Xiang, H.; Liu, B.; Jin, W. Maize plant architecture is regulated by the ethylene biosynthetic gene ZmACS7. Plant Physiol. 2020. [Google Scholar] [CrossRef]
- Li, X.; Sun, S.; Li, C.; Qiao, S.; Wang, T.; Leng, L.; Shen, H.; Wang, X. The Strigolactone-related mutants have enhanced lamina joint inclination phenotype at the seedling stage. J. Genet. Genom. 2014, 41, 605–608. [Google Scholar] [CrossRef]
- Shindo, M.; Yamamoto, S.; Shimomura, K.; Umehara, M. Strigolactones Decrease leaf angle in response to nutrient deficiencies in rice. Front. Plant Sci. 2020, 11, 135. [Google Scholar] [CrossRef]
- Gan, L.; Wu, H.; Wu, D.; Zhang, Z.; Guo, Z.; Yang, N.; Xia, K.; Zhou, X.; Oh, K.; Matsuoka, M.; et al. Methyl jasmonate inhibits lamina joint inclination by repressing brassinosteroid biosynthesis and signaling in rice. Plant Sci. 2015, 241, 238–245. [Google Scholar] [CrossRef]
- Hu, Y.; Yu, D. Brassinosteroid insensitive2 interacts with abscisic acid insensitive5 to mediate the antagonism of brassinosteroids to abscisic acid during seed germination in Arabidopsis. Plant Cell 2014, 26, 4394–4408. [Google Scholar] [CrossRef] [Green Version]
- Zhao, X.; Dou, L.; Gong, Z.; Wang, X.; Liu, X. BES 1 hinders abscisic acid insensitive 5 and promotes seed germination in Arabidopsis. New Phytol. 2018, 221, 908–918. [Google Scholar] [CrossRef] [Green Version]
- Wang, H.; Tang, J.; Liu, J.; Hu, J.; Liu, J.; Chen, Y.; Cai, Z.; Wang, X. Abscisic acid signaling inhibits brassinosteroid signaling through dampening the dephosphorylation of BIN2 by ABI1 and ABI2. Mol. Plant 2018, 11, 315–325. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, Q.-F.; Lu, J.; Zhou, Y.; Wu, F.; Tong, H.; Wang, J.-D.; Yu, J.-W.; Zhang, C.-Q.; Fan, X.-L.; Liu, Q. Abscisic acid represses rice lamina joint inclination by antagonizing brassinosteroid biosynthesis and signaling. Int. J. Mol. Sci. 2019, 20, 4908. [Google Scholar] [CrossRef] [Green Version]
- Dou, D.; Han, S.; Ku, L.; Liu, H.; Su, H.; Ren, Z.; Zhang, D.; Zeng, H.; Dong, Y.; Liu, Z.; et al. ZmCLA4 regulates leaf angle through multiple plant hormone-mediated signal pathways in maize 2020. bioRxiv 2020. [Google Scholar] [CrossRef] [Green Version]
- Cao, Y.; Zeng, H.; Ku, L.; Ren, Z.; Han, Y.; Su, H.; Dou, D.; Liu, H.; Dong, Y.; Zhu, F.; et al. ZmIBH1-1 regulates plant architecture in maize. J. Exp. Bot. 2020, 71, 2943–2955. [Google Scholar] [CrossRef] [PubMed]
- Bao, F.; Shen, J.; Brady, S.R.; Muday, G.K.; Asami, T.; Yang, Z. Brassinosteroids interact with auxin to promote lateral root development in Arabidopsis. Plant Physiol. 2004, 134, 1624–1631. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yuldashev, R.; Avalbaev, A.; Bezrukova, M.; Vysotskaya, L.; Khripach, V.; Shakirova, F. Cytokinin oxidase is involved in the regulation of cytokinin content by 24-epibrassinolide in wheat seedlings. Plant Physiol. Biochem. 2012, 55, 1–6. [Google Scholar] [CrossRef]
- Peleg, Z.; Reguera, M.; Tumimbang, E.; Walia, H.; Blumwald, E. Cytokinin-mediated source/sink modifications improve drought tolerance and increase grain yield in rice under water-stress. Plant Biotechnol. J. 2011, 9, 747–758. [Google Scholar] [CrossRef]
- Yoshida, H.; Tanimoto, E.; Hirai, T.; Miyanoiri, Y.; Mitani, R.; Kawamura, M.; Takeda, M.; Takehara, S.; Hirano, K.; Kainosho, M.; et al. Evolution and diversification of the plant gibberellin receptor GID1. Proc. Natl. Acad. Sci. USA 2018, 115, E7844–E7853. [Google Scholar] [CrossRef] [Green Version]
- Ishii, T.; Numaguchi, K.; Miura, K.; Yoshida, K.; Thanh, P.T.; Htun, T.M.; Yamasaki, M.; Komeda, N.; Matsumoto, T.; Terauchi, R.; et al. OsLG1 regulates a closed panicle trait in domesticated rice. Nat. Genet. 2013, 45, 462–465. [Google Scholar] [CrossRef]
- Zhu, Z.; Tan, L.; Fu, Y.; Liu, F.; Cai, H.; Xie, D.; Wu, F.; Wu, J.; Matsumoto, T.; Sun, C. Genetic control of inflorescence architecture during rice domestication. Nat. Commun. 2013, 4, 2200. [Google Scholar] [CrossRef]
- Ding, J.; Zhang, L.; Chen, J.; Li, X.; Li, Y.; Cheng, H.; Huang, R.; Zhou, B.; Li, Z.; Wang, J.; et al. Genomic dissection of leaf angle in maize (Zea mays L.) using a four-way cross mapping population. PLoS ONE 2015, 10, e0141619. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Li, Y.; Shi, Y.; Song, Y.; Zhang, D.; Buckler, E.S.; Zhang, Z.; Wang, T.; Li, Y. Genetic control of the leaf angle and leaf orientation value as revealed by ultra-high density maps in three connected maize populations. PLoS ONE 2015, 10, e0121624. [Google Scholar] [CrossRef]
- Tian, F.; Bradbury, P.J.; Brown, P.J.; Hung, H.; Sun, Q.; Flint-Garcia, S.; Rocheford, T.R.; McMullen, M.D.; Holland, J.B.; Buckler, E.S. Genome-wide association study of leaf architecture in the maize nested association mapping population. Nat. Genet. 2011, 43, 159–162. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Xu, J.; Deng, D.; Ding, H.; Bian, Y.; Yin, Z.; Wu, Y.; Zhou, B.; Zhao, Y. A comprehensive meta-analysis of plant morphology, yield, stay-green, and virus disease resistance QTL in maize (Zea mays L.). Planta 2015, 243, 459–471. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Liu, C.; Qi, X.; Wu, Y.; Fei, X.; Mao, L.; Cheng, B.; Li, X.; Xie, C. RNA-guided Cas9 as an in vivo desired-target mutator in maize. Plant Biotechnol. J. 2017, 15, 1566–1576. [Google Scholar] [CrossRef] [PubMed] [Green Version]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).