Design, Synthesis, and In Vitro and In Silico Study of New Hybrid 1-(2-(4-Arylthiazol-2-yl)hydrazineylidene)-5,6-dihydro-4H-pyrrolo[3,2,1-ij]quinolin-2-ones as Factor Xa and Factor XIa Inhibitors
Abstract
1. Introduction
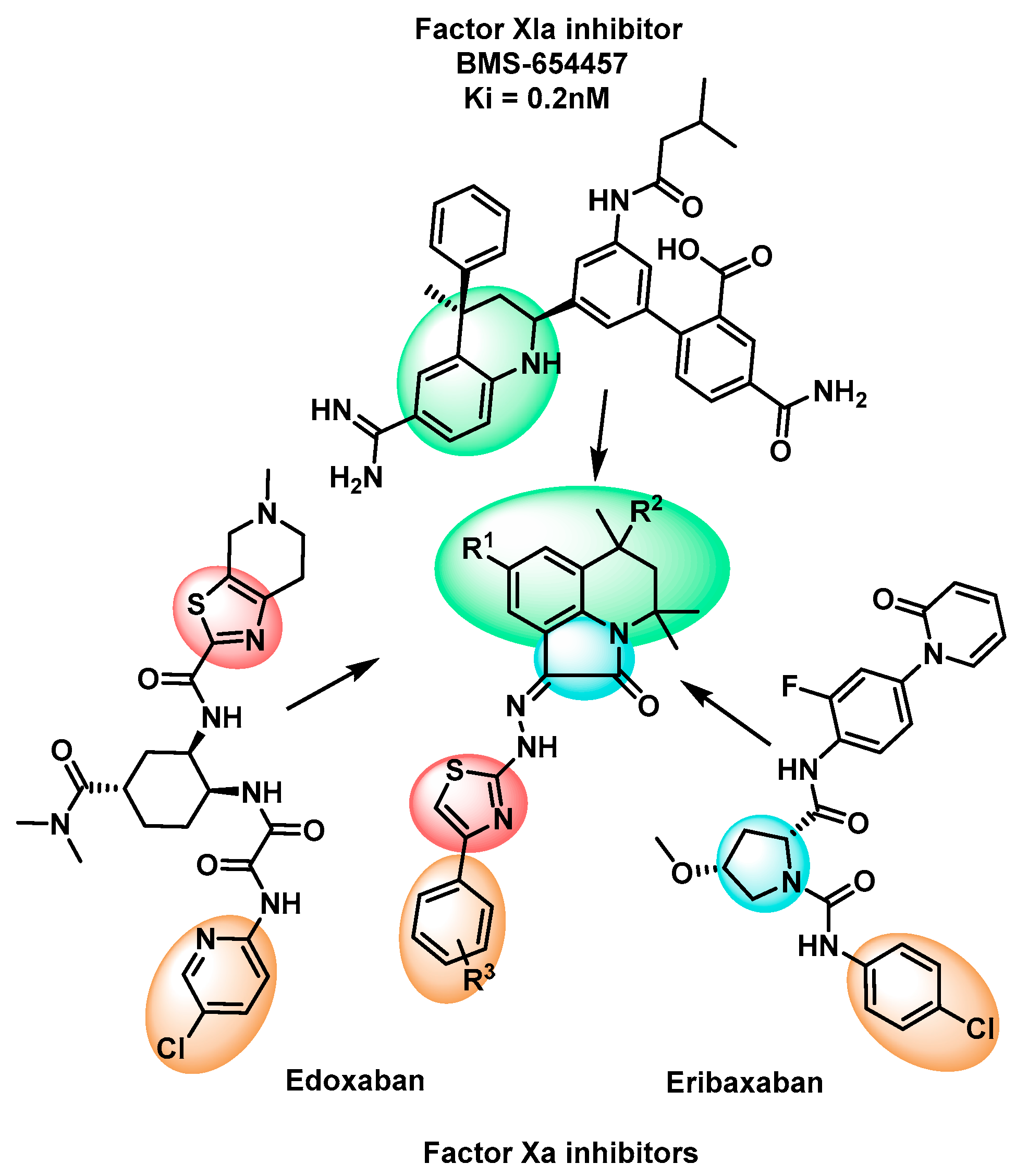
Rational Design
2. Results and Discussion
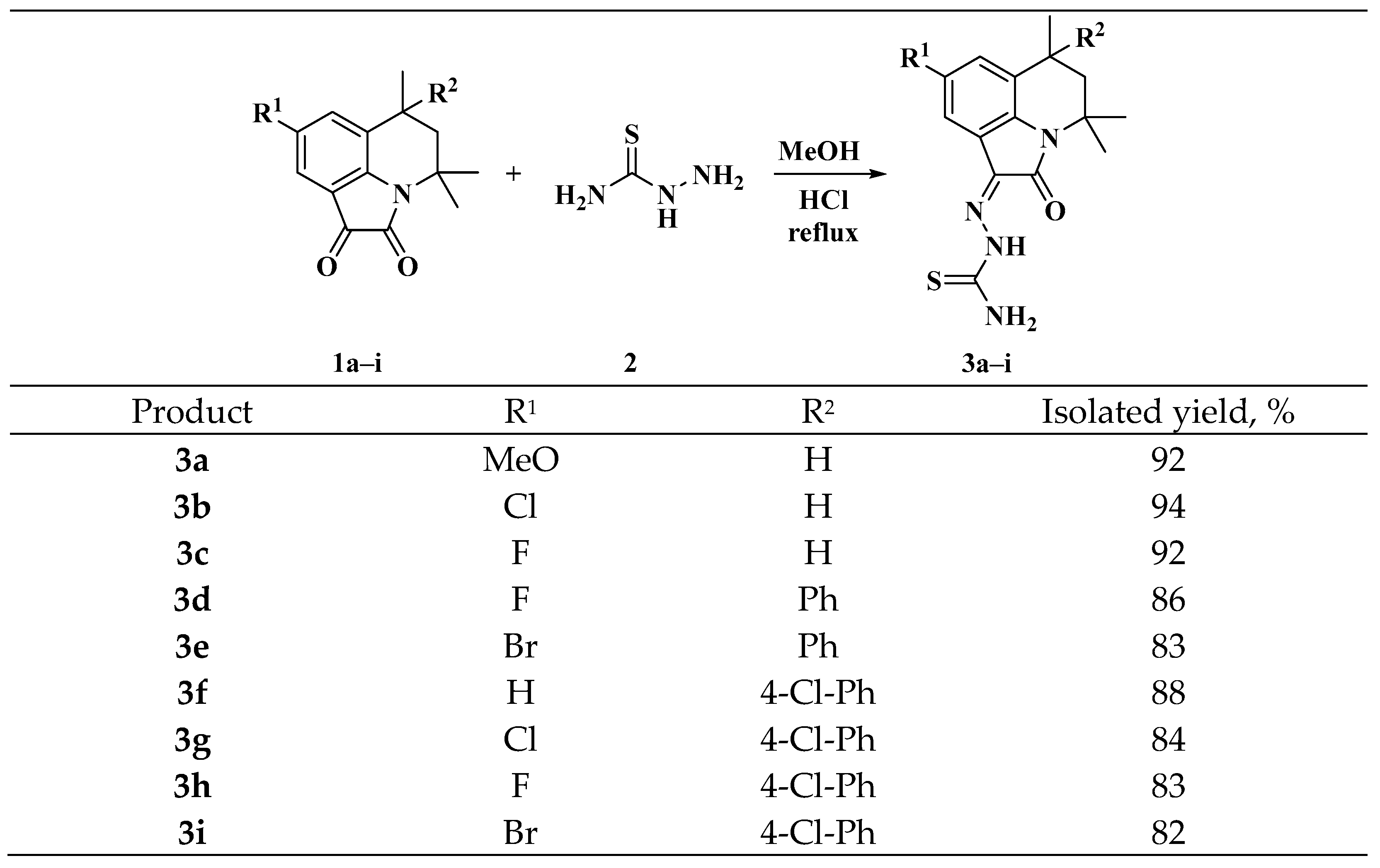
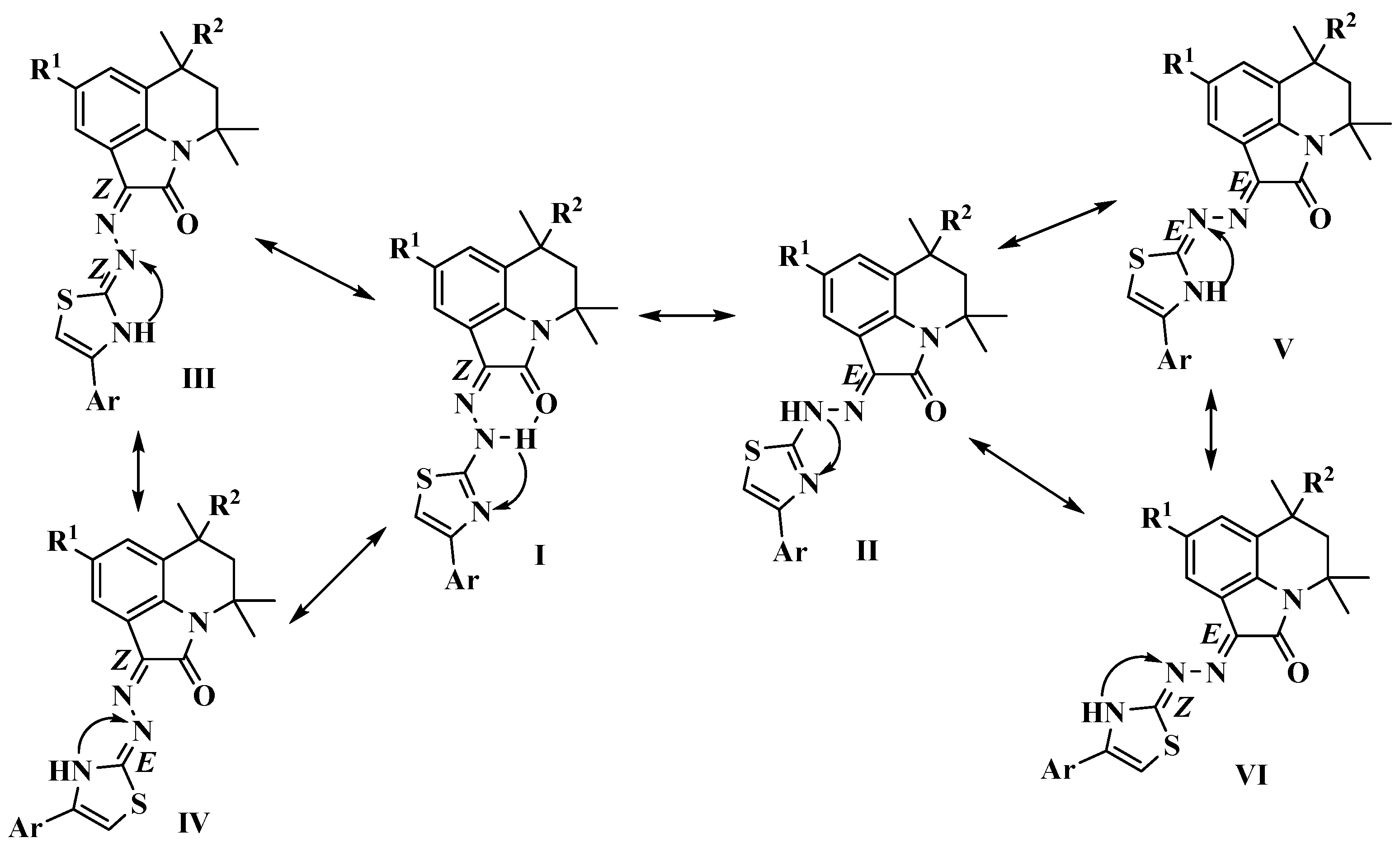
2.1. Chemistry
2.2. Anticoagulant Studies
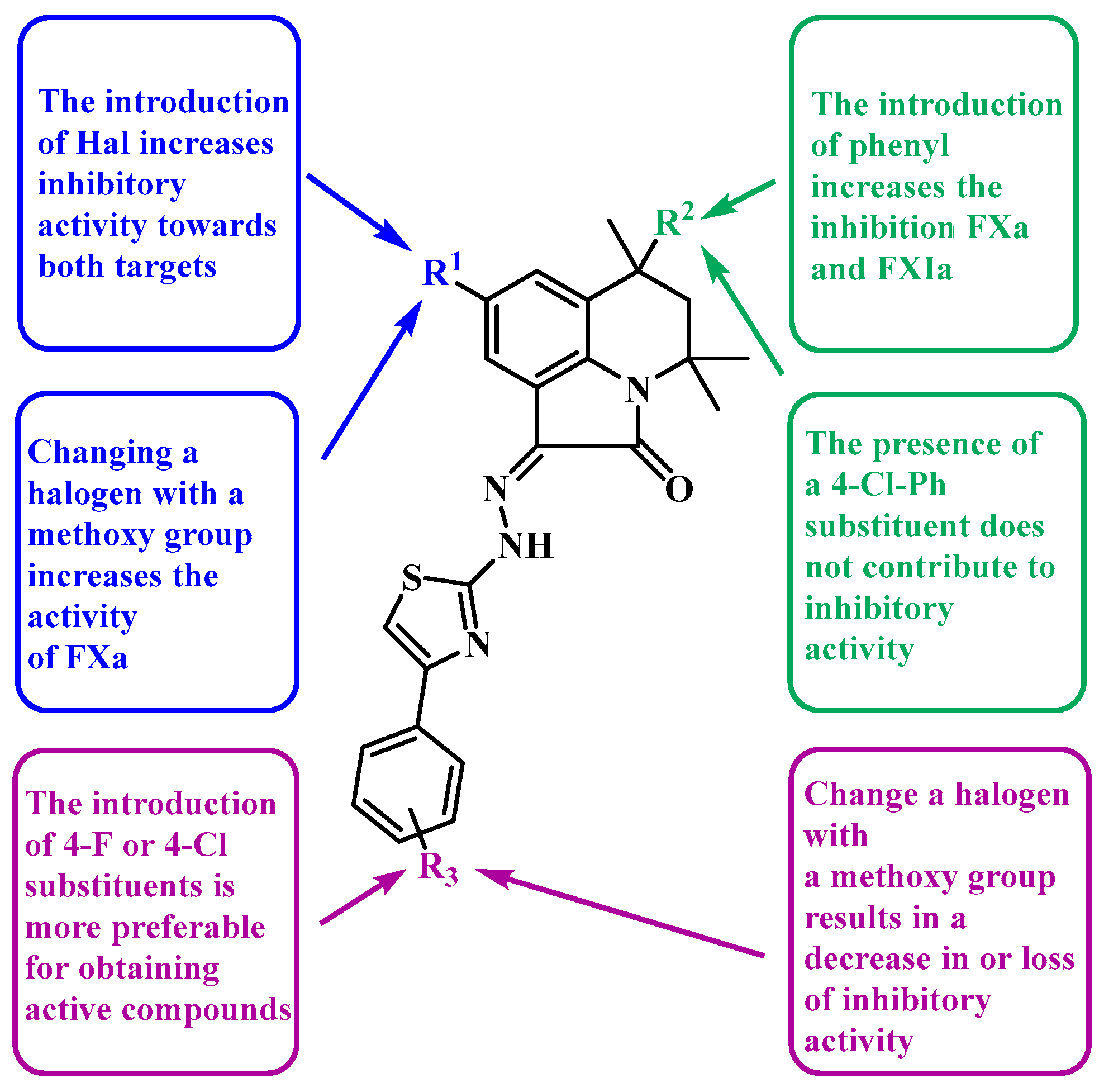
- The introduction of halogen substituents into position 8 of pyrrolo[3,2,1-ij]quinolin-2-one may contribute to the inhibition of both factors Xa and XIa (5d,g,h), thus increasing activity from −30% to 98% for Xa and from −6% to 99% for XIa.
- The presence of a methoxy group at position 8 of pyrrolo[3,2,1-ij]quinolin-2-one or in the phenyl substituent of the thiazole ring can result in a moderate potentiation of factor Xa activity by up to 40% but does not have a pronounced effect on factor XIa (5a–c,f,i).
- The introduction of F or Cl atoms into the para-position of the phenyl fragment in the thiazole ring is more preferable for the preparation of active compounds (5d,g,h,k) for XIa from −19% (5n) to 37% (5k) or from −8% and −2% (5e) to 98% and 99% (5d) for Xa and XIa, respectively.
- The appearance of a phenyl substituent in position 6 of pyrrolo[3,2,1-ij]quinoline-2-one increases the inhibitory ability (5g,h), thus increasing activity from −12% to 51% for Xa and from 37% to 76% for XIa.
2.3. Docking Studies
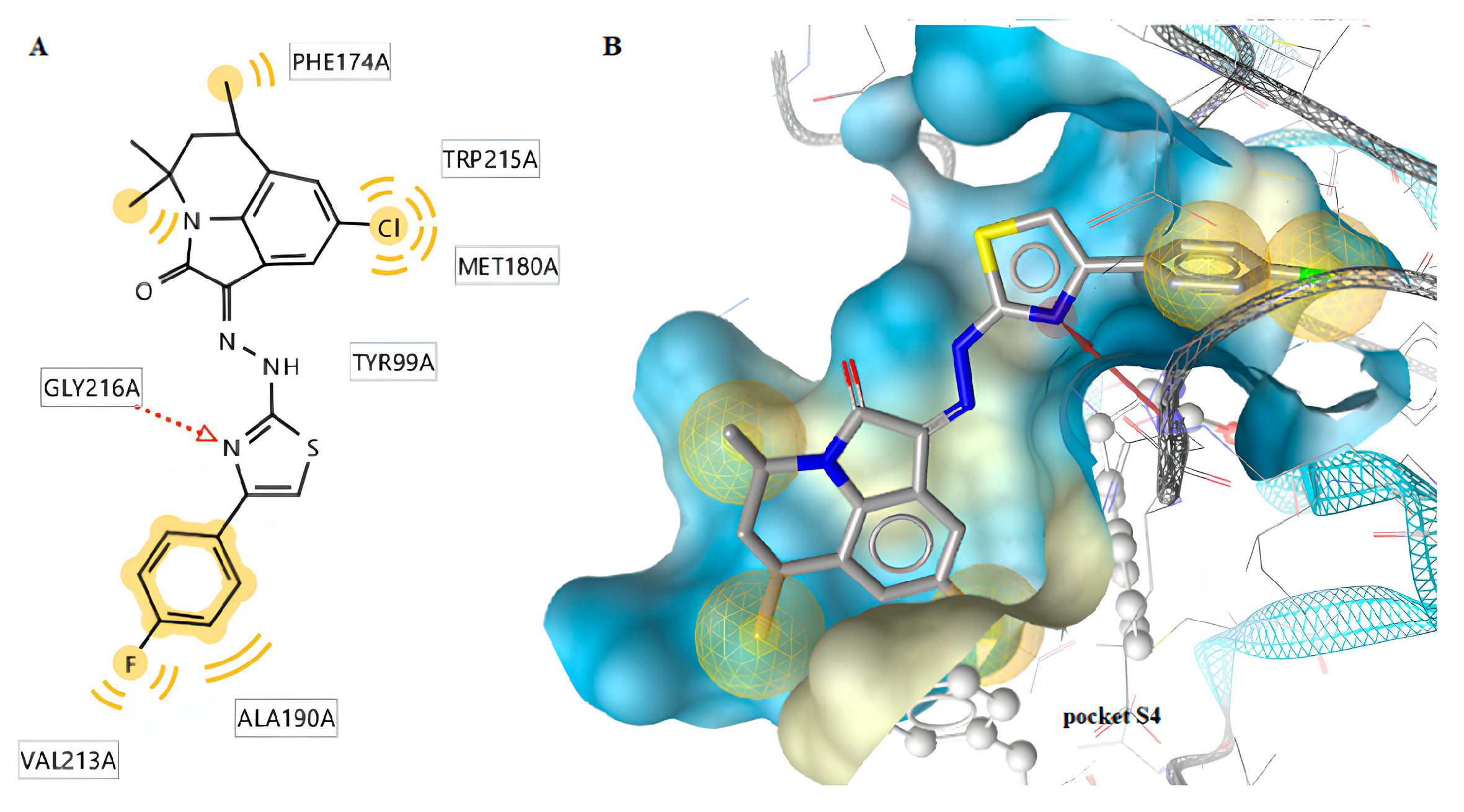
2.3.1. Docking to Factor Xa
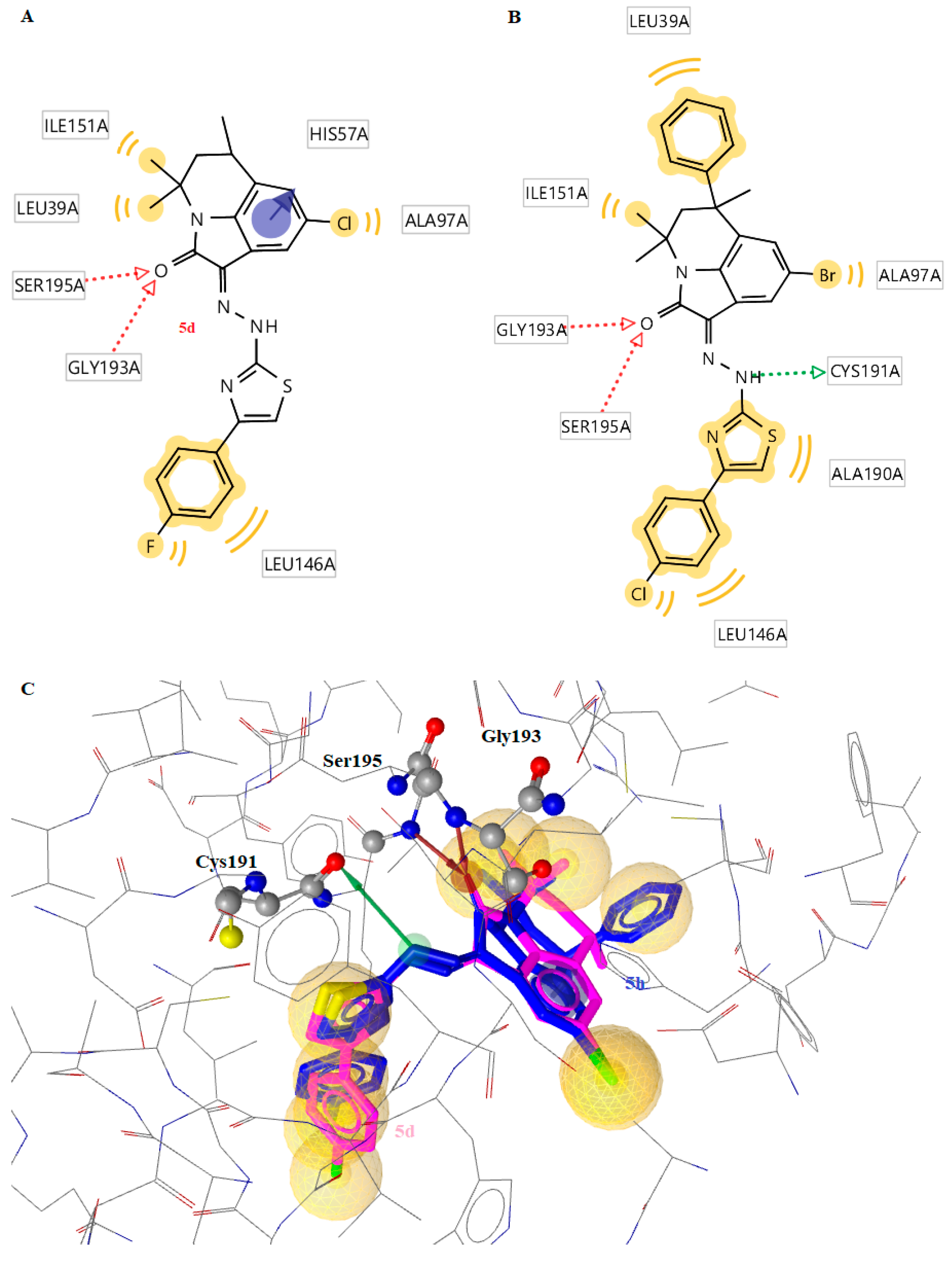
2.3.2. Docking to Factor XIa
2.4. Drug-Likeness
3. Materials and Methods
3.1. Synthesis
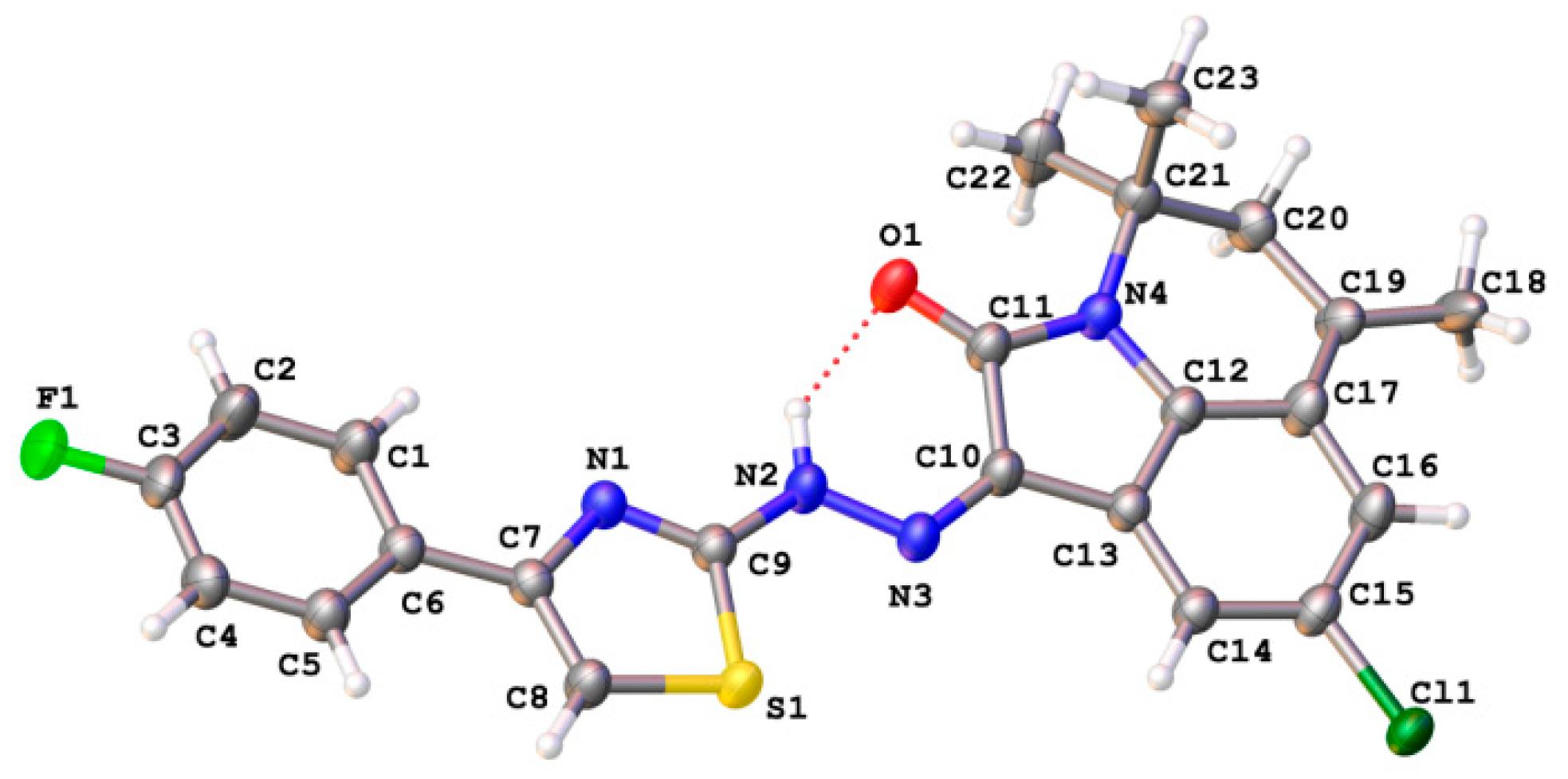
3.2. X-Ray Diffraction
3.3. In Vitro Assays
3.4. Molecular Docking Studies
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Jackson, S.P. Arterial Thrombosis—Insidious, Unpredictable and Deadly. Nat. Med. 2011, 17, 1423–1436. [Google Scholar] [CrossRef] [PubMed]
- Quan, M.L.; Glunz, P.W.; Smallheer, J.M. Advances in Anticoagulants. In Comprehensive Medicinal Chemistry III; Chackalamannil, S., Ward, S.E., Rotella, D., Eds.; Elsevier: Oxford, UK, 2017; pp. 600–627. [Google Scholar]
- Lewis, B.S.; Hasegawa, K. Factor XIa Inhibitors: Collecting the Clinical Evidence. Eur. Heart J. Cardiovasc. Pharmacother. 2024, 10, 5–6. [Google Scholar] [CrossRef]
- Patel, N.R.; Patel, D.V.; Murumkar, P.R.; Yadav, M.R. Contemporary Developments in the Discovery of Selective Factor Xa Inhibitors: A Review. Eur. J. Med. Chem. 2016, 121, 671–698. [Google Scholar] [CrossRef]
- Male, C.; Lensing, A.W.A.; Palumbo, J.S.; Kumar, R.; Nurmeev, I.; Hege, K.; Bonnet, D.; Connor, P.; Hooimeijer, H.L.; Torres, M.; et al. Rivaroxaban Compared with Standard Anticoagulants for the Treatment of Acute Venous Thromboembolism in Children: A Randomised, Controlled, Phase 3 Trial. Lancet Haematol. 2020, 7, e18–e27. [Google Scholar] [CrossRef]
- Sharma, M.; Molina, C.A.; Toyoda, K.; Bereczki, D.; Bangdiwala, S.I.; Kasner, S.E.; Lutsep, H.L.; Tsivgoulis, G.; Ntaios, G.; Czlonkowska, A.; et al. Safety and Efficacy of Factor XIa Inhibition with Milvexian for Secondary Stroke Prevention (AXIOMATIC-SSP): A Phase 2, International, Randomised, Double-Blind, Placebo-Controlled, Dose-Finding Trial. Lancet Neurol. 2024, 23, 46–59. [Google Scholar] [CrossRef]
- Al-Horani, R.A.; Afosah, D.K. Recent Advances in the Discovery and Development of Factor XI/XIa Inhibitors. Med. Res. Rev. 2018, 38, 1974–2023. [Google Scholar] [CrossRef]
- Seligsohn, U. Factor XI deficiency in humans. J. Thromb. Haemost. 2009, 7, 84–87. [Google Scholar] [CrossRef]
- Salomon, O.; Steinberg, D.M.; Dardik, R.; Rosenberg, N.; Zivelin, A.; Tamarin, I.; Ravid, B.; Berliner, S.; Seligsohn, U. Inherited factor XI deficiency confers no protection against acute myocardial infarction. J. Thromb. Haemost. 2003, 1, 658–661. [Google Scholar] [CrossRef]
- Gailani, D.; Renne, T. The Intrinsic Pathway of Coagulation: A Target for Treating Thromboembolic Disease? J. Thromb. Haemost. 2007, 5, 1106–1112. [Google Scholar] [CrossRef]
- Wichaiyo, S.; Parichatikanond, W.; Visansirikul, S.; Saengklub, N.; Rattanavipanon, W. Determination of the Potential Clinical Benefits of Small Molecule Factor XIa Inhibitors in Arterial Thrombosis. ACS Pharmacol. Transl. Sci. 2023, 6, 970–981. [Google Scholar] [CrossRef]
- Neves, A.R.; Correia-da-Silva, M.; Sousa, E.; Pinto, M. Structure–Activity Relationship Studies for Multitarget Antithrombotic Drugs. Future Med. Chem. 2016, 8, 2305–2355. [Google Scholar] [CrossRef] [PubMed]
- Huang, X.; Swanson, R.; Broze, G.J.; Olson, S.T. Kinetic Characterization of the Protein Z-Dependent Protease Inhibitor Reaction with Blood Coagulation Factor Xa. J. Biol. Chem. 2008, 283, 29770–29783. [Google Scholar] [CrossRef] [PubMed]
- Gan, W.; Deng, L.; Yang, C.; He, Q.; Hu, J.; Yin, H.; Jin, X.; Lu, C.; Wu, Y.; Peng, L. An Anticoagulant Peptide from the Human Hookworm, Ancylostoma duodenale That Inhibits Coagulation Factors Xa and XIa. FEBS Lett. 2009, 583, 1976–1980. [Google Scholar] [CrossRef] [PubMed]
- Kim, T.K.; Tirloni, L.; Radulovic, Z.; Lewis, L.; Bakshi, M.; Hill, C.; da Silva Vaz, I., Jr.; Logullo, C.; Termignoni, C.; Mulenga, A. Conserved Amblyomma americanum Tick Serpin19, an Inhibitor of Blood Clotting Factors Xa and XIa, Trypsin and Plasmin, Has Anti-Haemostatic Functions. Int. J. Parasitol. 2015, 45, 613–627. [Google Scholar] [CrossRef]
- Mishra, S.; Singh, P. Hybrid Molecules: The Privileged Scaffolds for Various Pharmaceuticals. Eur. J. Med. Chem. 2016, 124, 500–536. [Google Scholar] [CrossRef]
- Quan, M.L.; Wong, P.C.; Wang, C.; Woerner, F.; Smallheer, J.M.; Barbera, F.A.; Bozarth, J.M.; Brown, R.L.; Harrell, C.M.; Luettgen, J.M.; et al. Tetrahydroquinoline derivatives as potent and selective factor XIa inhibitors. J. Med. Chem. 2014, 57, 955–969. [Google Scholar] [CrossRef]
- Novichikhina, N.; Ilin, I.; Tashchilova, A.; Sulimov, A.; Kutov, D.; Ledenyova, I.; Krysin, M.; Shikhaliev, K.S.; Gantseva, A.; Gantseva, E.; et al. Synthesis, Docking, and In Vitro Anticoagulant Activity Assay of Hybrid Derivatives of Pyrrolo[3,2,1-ij]quinolin-2(1H)-one as New Inhibitors of Factor Xa and Factor XIa. Molecules 2020, 25, 1889. [Google Scholar] [CrossRef]
- Skoptsova, A.A.; Novichikhina, N.P.; Kosheleva, E.A.; Baranin, S.V.; Podoplelova, N.A.; Panteleev, M.A.; Shikhaliev, K.S. Synthesis and Study of New Anticoagulant Candidates Based on 6-Aryl-5,6-dihydro-4H-pyrrolo[3,2,1-ij]quinoline-1,2-diones. Russ. J. Gen. Chem. 2023, 93, S115–S123. [Google Scholar] [CrossRef]
- Novichikhina, N.P.; Pantykina, D.A.; Shestakov, A.S.; Potapov, A.Y.; Ledenyova, I.V.; Kuznetsov, M.A.; Shikhaliev, K.S. Allylic and Retro-Allylic Rearrangements upon Bromination of 8,9-Substituted 4,4,6-Trimethyl-4H-Pyrrolo[3,2,1-ij]Quinoline-1,2-Diones. New Aspects and Synthetic Applications. ChemistrySelect 2023, 8, e202203981. [Google Scholar] [CrossRef]
- Novichikhina, N.P.; Shestakov, A.S.; Potapov, A.Y.; Kosheleva, E.A.; Shatalov, G.V.; Verezhnikov, V.N.; Vandyshev, D.Y.; Ledeneva, I.V.; Shikhaliev, K.S. Synthesis of 4H-Pyrrolo[3,2,1-ij]quinoline-1,2-diones Containing a Piperazine Fragment and Study of Their Inhibitory Properties Against Protein Kinases. Russ. Chem. Bull. 2020, 69, 787–792. [Google Scholar] [CrossRef]
- Quan, M.L.; Pinto, D.J.; Smallheer, J.M.; Ewing, W.R.; Rossi, K.A.; Luettgen, J.M.; Seiffert, D.A.; Wexler, R.R. Factor XIa inhibitors as new anticoagulants. J. Med. Chem. 2018, 61, 7425–7447. [Google Scholar] [CrossRef]
- Fradera, X.; Kazemier, B.; Carswell, E.; Cooke, A.; Oubrie, A.; Hamilton, W.; Dempster, M.; Krapp, S.; Nagel, S.; Jestel, A. High-resolution crystal structures of factor XIa coagulation factor in complex with nonbasic high-affinity synthetic inhibitors. Acta Crystallogr. Sect. F Struct. Biol. Cryst. Commun. 2012, 68, 404–408. [Google Scholar] [CrossRef]
- Fjellström, O.; Akkaya, S.; Beisel, H.-G.; Eriksson, P.-O.; Erixon, K.; Gustafsson, D.; Jurva, U.; Kang, D.; Karis, D.; Knecht, W.; et al. Creating novel activated factor XI inhibitors through fragment based lead generation and structure aided drug design. PLoS ONE 2015, 10, e0113705. [Google Scholar] [CrossRef]
- Hu, Z.; Wong, P.C.; Gilligan, P.J.; Han, W.; Pabbisetty, K.B.; Bozarth, J.M.; Crain, E.J.; Harper, T.; Luettgen, J.M.; Myers, J.E.; et al. Discovery of a Potent Parenterally Administered Factor XIa Inhibitor with Hydroxyquinolin-2(1H)-one as the P2′ Moiety. ACS Med. Chem. Lett. 2015, 6, 590–595. [Google Scholar] [CrossRef]
- Al-Horani, R.A.; Desai, U.R. Factor XIa Inhibitors: A Review of the Patent Literature. Expert Opin. Ther. Pat. 2016, 26, 323–345. [Google Scholar] [CrossRef]
- Corte, J.R.; Fang, T.; Hangeland, J.J.; Friends, T.J.; Rendina, A.R.; Luettgen, J.M.; Bozarth, J.M.; Barbera, F.A.; Rossi, K.A.; Wei, A.; et al. Pyridine and pyridinone-based factor XIa inhibitors. Bioorg. Med. Chem. Lett. 2015, 25, 925–930. [Google Scholar] [CrossRef]
- Dilger, A.K.; Pabbisetty, K.B.; Corte, J.R.; De Lucca, I.; Fang, T.; Yang, W.; Pinto, D.J.P.; Wang, Y.; Zhu, Y.; Mathur, A.; et al. Discovery of milvexian, a high-affinity, orally bioavailable inhibitor of factor XIa in clinical studies for antithrombotic therapy. J. Med. Chem. 2021, 65, 1770–1785. [Google Scholar] [CrossRef]
- Yao, N.; Jia, Z.; Tian, Y.; Hou, S.; Yang, X.; Han, J.; Duan, Y.; Liao, C.; Kong, Y.; Xie, Z. Targeting the S2 Subsite Enables the Structure-Based Discovery of Novel Highly Selective Factor XIa Inhibitors. J. Med. Chem. 2022, 65, 4318–4334. [Google Scholar] [CrossRef]
- Lei, Y.; Zhang, B.; Zhang, Y.; Dai, X.; Duan, Y.; Mao, Q.; Gao, J.; Yang, Y.; Bao, Z.; Fu, X.; et al. Design, synthesis and biological evaluation of novel FXIa inhibitors with 2-phenyl-1H-imidazole-5-carboxamide moiety as P1 fragment. Eur. J. Med. Chem. 2021, 220, 113437. [Google Scholar] [CrossRef]
- Yang, W.; Wang, Y.; Lai, A.; Clark, C.G.; Corte, J.R.; Fang, T.; Gilligan, P.J.; Jeon, Y.; Pabbisetty, K.B.; Rampulla, R.A.; et al. Discovery of a high affinity, orally bioavailable macrocyclic FXIa inhibitor with antithrombotic activity in preclinical species. J. Med. Chem. 2020, 63, 7226–7242. [Google Scholar] [CrossRef]
- Pinto, D.J.P.; Smallheer, J.M.; Corte, J.R.; Austin, E.J.D.; Wang, C.; Fang, T.; Smith II, L.M.; Rossi, K.A.; Rendina, A.R.; Bozarth, J.M.; et al. Structure-based design of inhibitors of coagulation factor XIa with novel P1 moieties. Bioorg. Med. Chem. Lett. 2015, 25, 1635–1642. [Google Scholar] [CrossRef]
- Wang, Y.; Yuan, J.; Yan, S.; Liu, P.; Zheng, Z.; Zhang, S.; Meng, F.; Liu, W.; Huang, C.; Wei, Q. Design, synthesis and biological evaluation of 6-chloro-quinolin-2-one derivatives as novel FXIa inhibitors. Bioorg. Med. Chem. Lett. 2024, 99, 129610. [Google Scholar] [CrossRef]
- Leshcheva, Y.V.; Shikhaliev, K.S.; Shatalov, G.V.; Yermolova, G.I. New functional derivatives 4,4,6-trimethyl-4H-pyrrolo[3,2,1-ij]quinoline-1,2-diones. Izv. vuzov. Khimiya I khim. Tekhnologia [ChemChemTech] 2003, 46, 105. (In Russian) [Google Scholar]
- Skoptsova, A.A.; Geronikaki, A.; Novichikhina, N.P.; Sulimov, A.V.; Ilin, I.S.; Sulimov, V.B.; Bykov, G.A.; Podoplelova, N.A.; Pyankov, O.V.; Shikhaliev, K.S. Design, Synthesis, and Evaluation of New Hybrid Derivatives of 5, 6-Dihydro-4 H-Pyrrolo [3, 2, 1-ij] Quinolin-2 (1 H)-One as Potential Dual Inhibitors of Blood Coagulation Factors Xa and XIa. Molecules 2024, 29, 373. [Google Scholar] [CrossRef]
- Mahmoud, H.K.; Farghaly, T.A.; Abdulwahab, H.G.; Al-Qurashi, N.T.; Shaaban, M.R. Novel 2-indolinone thiazole hybrids as sunitinib analogues: Design, synthesis, and potent VEGFR-2 inhibition with potential anti-renal cancer activity. Eur. J. Med. Chem. 2020, 208, 112752. [Google Scholar] [CrossRef]
- Meleddu, R.; Distinto, S.; Corona, A.; Bianco, G.; Cannas, V.; Esposito, F.; Artese, A.; Alcaro, S.; Matyus, P.; Bogdan, D.; et al. (3Z)-3-(2-[4-(aryl)-1,3-thiazol-2-yl]hydrazin-1-ylidene)-2,3-dihydro-1H-indol-2-one derivatives as dual inhibitors of HIV-1 reverse transcriptase. Eur. J. Med. Chem. 2015, 93, 452–460. [Google Scholar] [CrossRef]
- Łączkowski, K.Z.; Anusiak, J.; Świtalska, M.; Dzitko, K.; Cytarska, J.; Baranowska-Łączkowska, A.; Plech, T.; Paneth, A.; Wietrzyk, J.; Białczyk, J. Synthesis, molecular docking, ctDNA interaction, DFT calculation and evaluation of antiproliferative and anti-Toxoplasma gondii activities of 2,4-diaminotriazine-thiazole derivatives. Med. Chem. Res. 2018, 27, 1131–1148. [Google Scholar] [CrossRef]
- Shaik, S.; Sirigireddy, R.M.R.; Godugu, K.; Vemula, V.; Kakarla, R.R.; Balaraman, E.; Nallagondu, C.G.R.; Aminabhavi, T.M. SiO2-supported HClO4 catalyzed synthesis of (Z)-thiazolylhydrazonoindolin-2-ones and their electrochemical properties. Chemosphere 2022, 309, 136667. [Google Scholar] [CrossRef]
- Kökbudak, Z.; Saracoglu, M.; Akkoç, S.; Çimen, Z.; Yilmazer, M.I.; Kandemirli, F. Synthesis, Cytotoxic Activity and Quantum Chemical Calculations of New 7-Thioxopyrazolo[1,5-f]Pyrimidin-2-one Derivatives. J. Mol. Struct. 2020, 1202, 127261. [Google Scholar] [CrossRef]
- Oliveira, N.J.C.; Teixeira, I.N.S.; Fernandes, P.O.; Veríssimo, G.C.; Valério, A.D.; de Souza Moreira, C.P.; Freitas, T.R.; Fonseca, A.C.V.; de Paula Sabino, A.; Johann, S.; et al. Computer-Aided Molecular Design, Synthesis and Evaluation of Antifungal Activity of Heterocyclic Compounds. J. Mol. Struct. 2022, 1267, 133573. [Google Scholar] [CrossRef]
- Solangi, M.; Kanwal; Khan, K.M.; Chigurupati, S.; Saleem, F.; Qureshi, U.; Ul-Haq, Z.; Jabeen, A.; Felemban, S.G.; Zafar, F.; et al. Isatin Thiazoles as Antidiabetic: Synthesis, In Vitro Enzyme Inhibitory Activities, Kinetics, and In Silico Studies. Arch. Pharm. 2022, 355, 2100481. [Google Scholar] [CrossRef]
- Fayed, E.A.; Ragab, A.; Eldin, R.R.E.; Bayoumi, A.H.; Ammar, Y.A. In Vivo Screening and Toxicity Studies of Indolinone Incorporated Thiosemicarbazone, Thiazole and Piperidinosulfonyl Moieties as Anticonvulsant Agents. Bioorg. Chem. 2021, 116, 105300. [Google Scholar] [CrossRef]
- Yamali, C.; Gul, M.; Gul, H.I. Current pharmaceutical research on the significant pharmacophore mannich bases in drug design. Curr. Top. Med. Chem. 2023, 23, 2590–2608. [Google Scholar] [CrossRef]
- CrysAlisPro; Version 1.171.41.106a; Rigaku Oxford Diffraction: Yarnton, UK, 2021.
- Dolomanov, O.V.; Bourhis, L.J.; Gildea, R.J.; Howard, J.A.K.; Puschmann, H. OLEX2: A Complete Structure Solution, Refinement and Analysis Program. J. Appl. Crystallogr. 2009, 42, 339–341. [Google Scholar] [CrossRef]
- Sheldrick, G.M. SHELXT—Integrated Space-Group and Crystal-Structure Determination. Acta Crystallogr. Sect. A Found. Adv. 2015, A71, 3–8. [Google Scholar] [CrossRef]
- Sheldrick, G.M. Crystal Structure Refinement with SHELXL. Acta Crystallogr. Sect. C Struct. Chem. 2015, C71, 3–8. [Google Scholar] [CrossRef]
- Berman, H.M.; Westbrook, J.; Feng, Z.; Gilliland, G.; Bhat, T.N.; Weissig, H.; Shindyalov, I.N.; Bourne, P.E. The Protein Data Bank. Nucleic Acids Res. 2000, 28, 235–242. [Google Scholar] [CrossRef]
- Morris, G.M.; Goodsell, D.S.; Halliday, R.S.; Huey, R.; Hart, W.E.; Belew, R.K.; Olson, A.J. Automated Docking Using a Lamarckian Genetic Algorithm and an Empirical Binding Free Energy Function. J. Comput. Chem. 1998, 19, 1639–1662. [Google Scholar] [CrossRef]
- Wolber, G.; Langer, T. LigandScout: 3-D Pharmacophores Derived from Protein-Bound Ligands and Their Use as Virtual Screening Filters. J. Chem. Inf. Model. 2005, 45, 160–169. [Google Scholar] [CrossRef]


| No. | Percent Inhibition at 30 μM ± SEM (n = 3) | IC50, μM ± SEM (n = 3) | |||
|---|---|---|---|---|---|
| Xa | XIa | Plasmin | Xa | XIa | |
| 5a | −30 ± 7 | −13 ± 11 | |||
| 5b | −30 ± 10 | −6 ± 10 | |||
| 5c | −40 ± 10 | −7 ± 8 | |||
| 5d | 98 ± 1 | 99 | 5 ± 1 | 5.43 ± 0.02 | 7.70 ± 0.05 |
| 5e | −8 ± 10 | −2 ± 11 | |||
| 5f | −25 ± 9 | −11 ± 9 | |||
| 5g | 51 ± 18 | 76 ± 3 | |||
| 5h | 99 ± 1 | 100 | 8 ± 2 | 5.75 ± 0.02 | 7.66 ± 0.08 |
| 5i | −21 ± 9 | −14 ± 10 | |||
| 5j | −3 ± 3 | −3 ± 6 | |||
| 5k | −12 ± 11 | 37 ± 8 | |||
| 5l | −4 ± 3 | −11 ± 7 | |||
| 5m | −17 ± 11 | −14 ± 7 | |||
| 5n | −2 ± 3 | −19 ± 8 | |||
| 5o | −4 ± 6 | −11 ± 12 | |||
| Rivaroxaban | 94 | 8 | 0.007 ± 0.001 | ||
| No. | Est. Binding Energy (kcal/mol) | No. | Est. Binding Energy (kcal/mol) | ||
|---|---|---|---|---|---|
| FXa | FXIa | FXa | FXIa | ||
| 5a | −6.62 | −4.28 | 5i | −5.14 | −4.67 |
| 5b | −6.44 | - | 5j | - | - |
| 5c | −6.48 | - | 5k | −1.56 | - |
| 5d | −11.56 | −11.60 | 5l | - | −2.56 |
| 5e | −1.28 | - | 5m | −4.63 | −3.10 |
| 5f | −2.63 | −1.34 | 5n | −1.87 | - |
| 5g | −6.59 | −7.17 | 5o | - | −3.14 |
| 5h | −12.47 | −12.80 | Rivaroxaban | −13.48 | −5.34 |
| Property | Predicted Value | Unit | |||||||
|---|---|---|---|---|---|---|---|---|---|
| 5a | 5b | 5c | 5d | 5e | 5f | 5g | 5h | ||
| Molecular weight | 466.98 | 450.53 | 511.43 | 454.95 | 466.98 | 450.53 | 531.04 | 591.95 | Numeric (g/mol) |
| Num. rotatable bonds | 4 | 4 | 4 | 3 | 4 | 4 | 4 | 4 | Numeric |
| Num. H-bond acceptors | 4 | 5 | 4 | 4 | 4 | 5 | 4 | 3 | Numeric |
| Num. H-bond donors | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | Numeric |
| TPSA | 95.06 | 95.06 | 95.06 | 85.83 | 95.06 | 95.06 | 85.83 | 85.83 | Numeric (Å2) |
| Consensus logPo/w | 5.02 | 4.81 | 5.10 | 5.32 | 4.98 | 4.76 | 6.55 | 685 | average of five methods |
| Solubility | Poor | Poor | Poor | Poor | Poor | Poor | Poor | Poor | Categorical |
| GI absorption | High | High | High | High | High | High | Low | Low | Categorical |
| P-gp substrate | Yes | Yes | Yes | Yes | Yes | Yes | No | No | Categorical |
| CYP2D6 inhibitor | No | No | No | No | No | No | No | Yes | Categorical |
| CYP3A4 inhibitor | Yes | Yes | Yes | Yes | Yes | Yes | No | No | Categorical |
| Lipinski | 0 | 0 | 1 | 1 | 0 | 0 | 2 | 2 | Categorical |
| Bioavailability Score | 0.55 | 0.55 | 0.55 | 0.55 | 0.55 | 0.55 | 0.17 | 0.17 | |
| PAINS | 1 * | 1 * | 1 * | 1 * | 1 * | 1 * | 1 * | 1 * | Categorical |
| - | 5i | 5j | 5k | 5l | 5m | 5n | 5o | ||
| Molecular weight | 543.08 | 577.52 | 565.49 | 549.03 | 565.49 | 561.07 | 591.95 | Numeric (g/mol) | |
| Num. rotatable bonds | 5 | 5 | 4 | 4 | 4 | 5 | 4 | Numeric | |
| Num. H-bond acceptors | 4 | 4 | 4 | 5 | 4 | 5 | 3 | Numeric | |
| Num. H-bond donors | 1 | 1 | 1 | 1 | 1 | 1 | 1 | Numeric | |
| TPSA | 95.06 | 95.06 | 85.83 | 85.83 | 85.83 | 95.06 | 85.83 | Numeric (Å2) | |
| Consensus logPo/w | 6.18 | 6.67 | 7.03 | 8.32 | 7.04 | 6.45 | 6.85 | average of five methods | |
| Solubility | Poor | Poor | Poor | Poor | Poor | Poor | Poor | Categorical | |
| GI absorption | Low | Low | Low | Low | Low | Low | Low | Categorical | |
| P-gp substrate | No | No | No | No | No | No | No | Categorical | |
| CYP2D6 inhibitor | Yes | Yes | Yes | No | Yes | Yes | Yes | Categorical | |
| CYP3A4 inhibitor | No | No | No | No | No | No | No | Categorical | |
| Lipinski | 2 | 2 | 2 | 2 | 2 | 2 | 2 | Categorical | |
| Bioavailability Score | 0.17 | 0.17 | 0.17 | 0.17 | 0.17 | 0.17 | 0.17 | ||
| PAINS | 1 * | 1 * | 1 * | 1 * | 1 * | 1 * | 1 * | Categorical | |
 .
.| Property | Model Name | Predicted Value | Unit | |||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Rivaroxaban | 5a | 5b | 5c | 5d | 5e | 5f | 5g | |||
| Absorption | Water solubility | −4.606 | −5.766 | −5.166 | −5.831 | −5.517 | −5.525 | −5.296 | −6.060 | Numeric (log mol/L) |
| Absorption | Caco2 permeability | 1.049 | 0.71 | 1.111 | 0.702 | 1.101 | 1.121 | 1.119 | 1.063 | Numeric (log Papp in 10−6 cm/s) |
| Absorption | Intestinal absorption (human) | 92.743 | 91.439 | 92.534 | 91.372 | 90.499 | 91.907 | 93.014 | 91.282 | Numeric (% absorbed) |
| Absorption | Skin permeability | −3.186 | −2.768 | −2.793 | −2.767 | −2.769 | −2.789 | −2.801 | −2.724 | Numeric (log Kp) |
| Absorption | P-glycoprotein substrate | No | Yes | Yes | Yes | Yes | Yes | Yes | Yes | Categorical (Yes/No) |
| Absorption | P-glycoprotein I inhibitor | Yes | Yes | Yes | Yes | Yes | Yes | Yes | Yes | Categorical (Yes/No) |
| Absorption | P-glycoprotein II inhibitor | Yes | Yes | Yes | Yes | Yes | Yes | Yes | Yes | Categorical (Yes/No) |
| Distribution | VDss (human) | −0.625 | 0.464 | 0.435 | 0484 | 0.509 | 0.581 | 0.388 | 0.471 | Numeric (log L/kg) |
| Distribution | Fraction unbound (human) | 0.002 | 0.01 | 0.065 | 0.007 | 0.062 | 0.021 | 0.022 | 0.155 | Numeric (Fu) |
| Distribution | BBB permeability | −1.022 | −0.054 | −0.180 | −0.055 | 0.075 | −0.047 | −0.201 | 0.031 | Numeric (log BB) |
| Distribution | CNS permeability | −2.644 | −1.588 | −1.78 | −1.565 | −1.494 | −1.622 | −1.773 | −1.218 | Numeric (log PS) |
| Metabolism | CYP2D6 substrate | No | No | No | No | No | No | No | No | Categorical (Yes/No) |
| Metabolism | CYP3A4 substrate | Yes | Yes | Yes | Yes | Yes | Yes | Yes | Yes | Categorical (Yes/No) |
| Metabolism | CYP2D6 inhibitor | No | No | No | No | No | No | No | No | Categorical (Yes/No) |
| Metabolism | CYP3A4 inhibitor | Yes | Yes | Yes | Yes | Yes | Yes | Yes | Yes | Categorical (Yes/No) |
| Excretion | Total clearance | 0.294 | −0.01 | −0.142 | −0.032 | −0.308 | −0.128 | −0.025 | −0.189 | Numeric (log ml/min/kg) |
| Excretion | Renal OCT2 substrate | No | No | No | No | No | No | No | No | Categorical (Yes/No) |
| Toxicity | AMES toxicity | Yes | No | No | No | No | No | No | No | Categorical (Yes/No) |
| Toxicity | Max. tolerated dose (human) | 0.495 | 0.259 | 0.168 | 0.254 | 0.198 | 0.206 | 0.173 | 0.478 | Numeric (log mg/kg/day) |
| Toxicity | Oral rat acute toxicity (LD50) | 2.704 | 3.061 | 3.128 | 3.064 | 3.105 | 3.128 | 3.043 | 2.754 | Numeric (mol/kg) |
| Toxicity | hERG I inhibitor | No | No | No | No | No | No | No | No | Categorical (Yes/No) |
| Toxicity | Hepatotoxicity | Yes | Yes | Yes | Yes | Yes | Yes | Yes | Yes | Categorical (Yes/No) |
| Toxicity | Skin Sensitization | No | No | No | No | No | No | No | No | Categorical (Yes/No) |
| Property | Model name | Predicted value | Unit | |||||||
| 5h | 5i | 5j | 5k | 5l | 5m | 5n | 5o | |||
| Absorption | Water solubility | −6.225 | −5.647 | −5.702 | −6.044 | −5.441 | −6.026 | −5.634 | −6.193 | Numeric (log mol/L) |
| Absorption | Caco2 permeability | 1.037 | 1.036 | 1.054 | 1.056 | 1.097 | 1.05 | 1.052 | 1.039 | Numeric (log Papp in 10−6 cm/s) |
| Absorption | Intestinal absorption (human) | 90.174 | 94.393 | 92.432 | 90.970 | 92.065 | 91.731 | 93.539 | 91.357 | Numeric (% Absorbed) |
| Absorption | Skin permeability | −2.723 | −2.728 | −2.728 | −2.723 | −2.731 | −2.733 | −2.728 | −2.721 | Numeric (log Kp) |
| Absorption | P-glycoprotein substrate | Yes | Yes | Yes | Yes | Yes | Yes | Yes | Yes | Categorical (Yes/No) |
| Absorption | P-glycoprotein I inhibitor | Yes | Yes | Yes | Yes | Yes | Yes | Yes | Yes | Categorical (Yes/No) |
| Absorption | P-glycoprotein II inhibitor | Yes | Yes | Yes | Yes | Yes | Yes | Yes | Yes | Categorical (Yes/No) |
| Distribution | VDss (human) | 0.618 | 0.619 | 0.684 | 0.575 | 0.467 | 0.574 | 0.535 | 0.648 | Numeric (log L/kg) |
| Distribution | Fraction unbound (human) | 0.128 | 0.165 | 0.153 | 0.164 | 0.222 | 0.152 | 0.152 | 0.128 | Numeric (Fu) |
| Distribution | BBB permeability | 0.156 | −0.075 | −0.081 | 0.042 | −0.085 | 0.05 | −0.235 | 0.215 | Numeric (log BB) |
| Distribution | CNS permeability | −1.037 | −1.392 | −1.272 | −1.111 | −1.303 | −1.113 | −1.424 | −1.046 | Numeric (log PS) |
| Metabolism | CYP2D6 substrate | No | No | No | No | No | No | No | No | Categorical (Yes/No) |
| Metabolism | CYP3A4 substrate | Yes | Yes | Yes | Yes | Yes | Yes | Yes | Yes | Categorical (Yes/No) |
| Metabolism | CYP2D6 inhibitor | No | No | No | No | No | No | No | No | Categorical (Yes/No) |
| Metabolism | CYP3A4 inhibitor | Yes | Yes | Yes | Yes | Yes | Yes | Yes | Yes | Categorical (Yes/No) |
| Excretion | Total clearance | −0.195 | −0.125 | −0.254 | −0.32 | −0.451 | −0.314 | −0.152 | −0.194 | Numeric (log ml/min/kg) |
| Excretion | Renal OCT2 substrate | No | No | No | No | No | No | No | No | Categorical (Yes/No) |
| Toxicity | AMES toxicity | Yes | No | No | Yes | No | Yes | No | Yes | Categorical (Yes/No) |
| Toxicity | Max. tolerated dose (human) | 0.448 | 0.470 | 0.468 | 0.429 | 0.562 | 0.426 | 0.474 | 0.448 | Numeric (log mg/kg/day) |
| Toxicity | Oral rat acute toxicity (LD50) | 2.705 | 2.964 | 2.952 | 2.831 | 2.941 | 2.825 | 2.962 | 2.728 | Numeric (mol/kg) |
| Toxicity | hERG I inhibitor | No | No | No | No | No | No | No | No | Categorical (Yes/No) |
| Toxicity | Hepatotoxicity | No | Yes | Yes | Yes | Yes | Yes | Yes | No | Categorical (Yes/No) |
| Toxicity | Skin sensitization | No | No | No | No | No | No | No | No | Categorical (Yes/No) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Skoptsova, A.A.; Geronikaki, A.; Petrou, A.; Novichikhina, N.P.; Podoplelova, N.A.; Bykov, G.A.; Anis’kov, A.A.; Soloveva, S.A.; Shikhaliev, K.S. Design, Synthesis, and In Vitro and In Silico Study of New Hybrid 1-(2-(4-Arylthiazol-2-yl)hydrazineylidene)-5,6-dihydro-4H-pyrrolo[3,2,1-ij]quinolin-2-ones as Factor Xa and Factor XIa Inhibitors. Molecules 2025, 30, 3544. https://doi.org/10.3390/molecules30173544
Skoptsova AA, Geronikaki A, Petrou A, Novichikhina NP, Podoplelova NA, Bykov GA, Anis’kov AA, Soloveva SA, Shikhaliev KS. Design, Synthesis, and In Vitro and In Silico Study of New Hybrid 1-(2-(4-Arylthiazol-2-yl)hydrazineylidene)-5,6-dihydro-4H-pyrrolo[3,2,1-ij]quinolin-2-ones as Factor Xa and Factor XIa Inhibitors. Molecules. 2025; 30(17):3544. https://doi.org/10.3390/molecules30173544
Chicago/Turabian StyleSkoptsova, Anna A., Athina Geronikaki, Anthi Petrou, Nadezhda P. Novichikhina, Nadezhda A. Podoplelova, Georgii A. Bykov, Aleksandr A. Anis’kov, Svetlana A. Soloveva, and Khidmet S. Shikhaliev. 2025. "Design, Synthesis, and In Vitro and In Silico Study of New Hybrid 1-(2-(4-Arylthiazol-2-yl)hydrazineylidene)-5,6-dihydro-4H-pyrrolo[3,2,1-ij]quinolin-2-ones as Factor Xa and Factor XIa Inhibitors" Molecules 30, no. 17: 3544. https://doi.org/10.3390/molecules30173544
APA StyleSkoptsova, A. A., Geronikaki, A., Petrou, A., Novichikhina, N. P., Podoplelova, N. A., Bykov, G. A., Anis’kov, A. A., Soloveva, S. A., & Shikhaliev, K. S. (2025). Design, Synthesis, and In Vitro and In Silico Study of New Hybrid 1-(2-(4-Arylthiazol-2-yl)hydrazineylidene)-5,6-dihydro-4H-pyrrolo[3,2,1-ij]quinolin-2-ones as Factor Xa and Factor XIa Inhibitors. Molecules, 30(17), 3544. https://doi.org/10.3390/molecules30173544







