South African Abietane Diterpenoids and Their Analogs as Potential Antimalarials: Novel Insights from Hybrid Computational Approaches
Abstract
1. Introduction
2. Results and Discussion
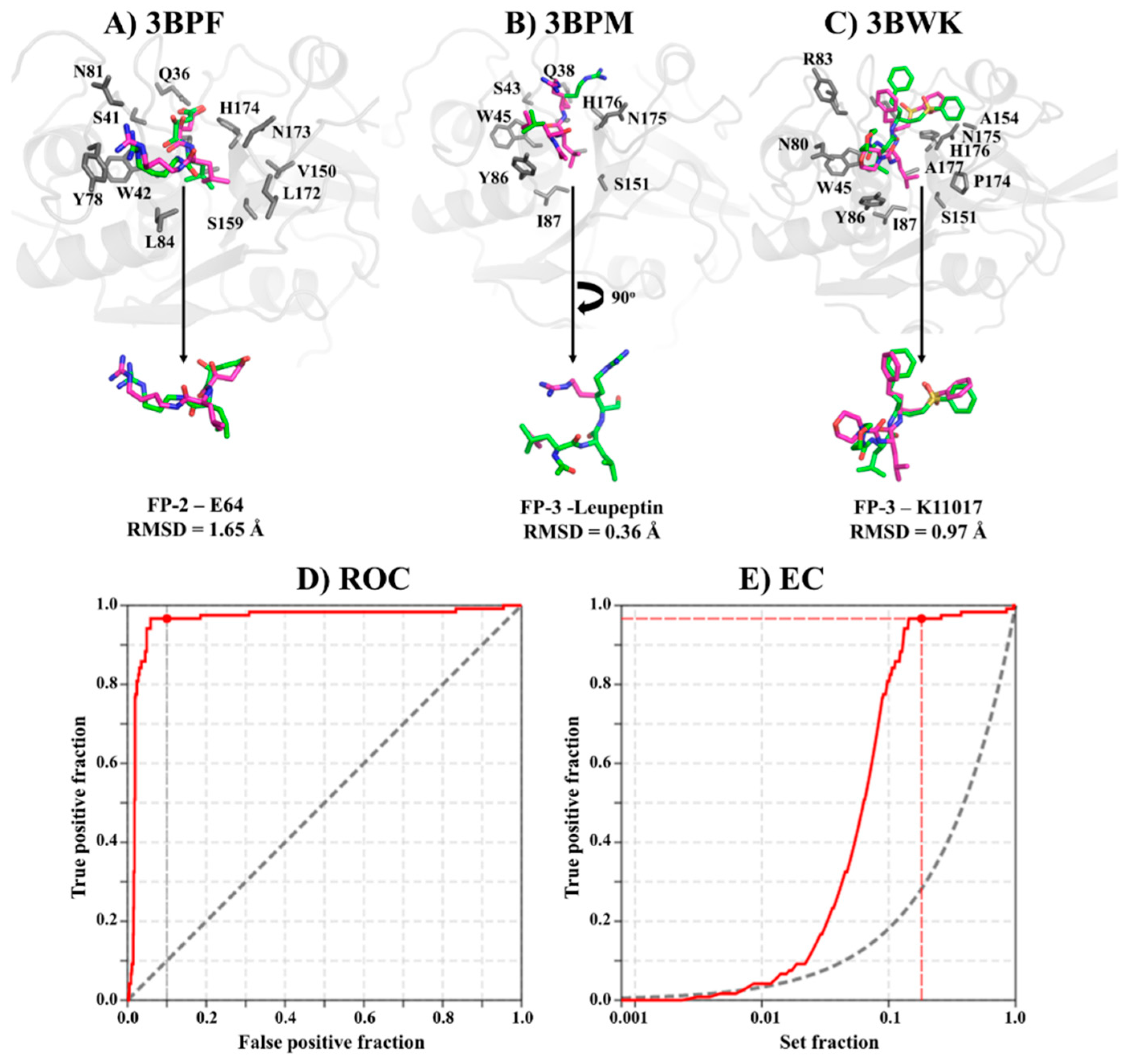
2.1. Accurate Ligand Docking Parameters are Established and Validated
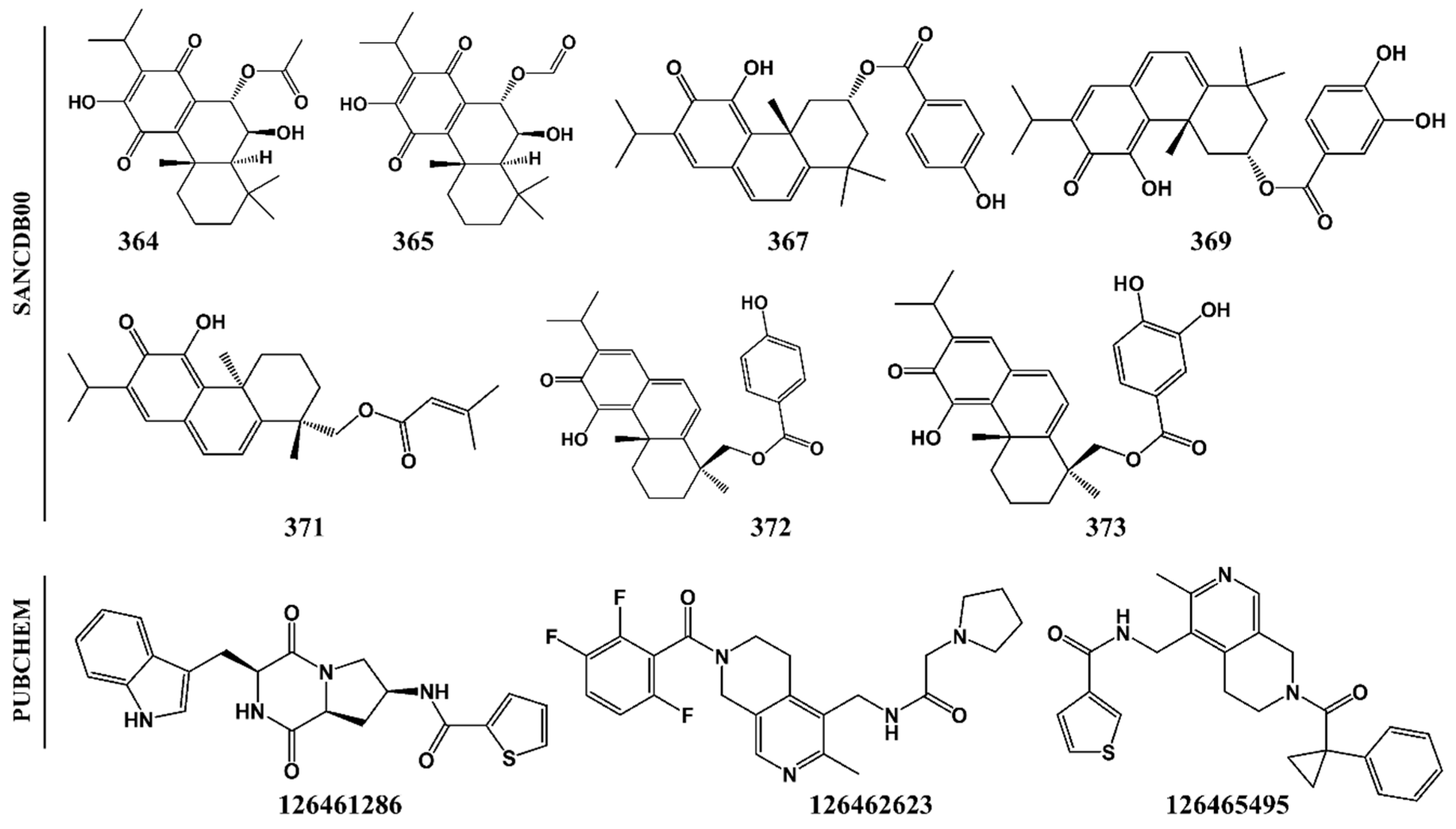
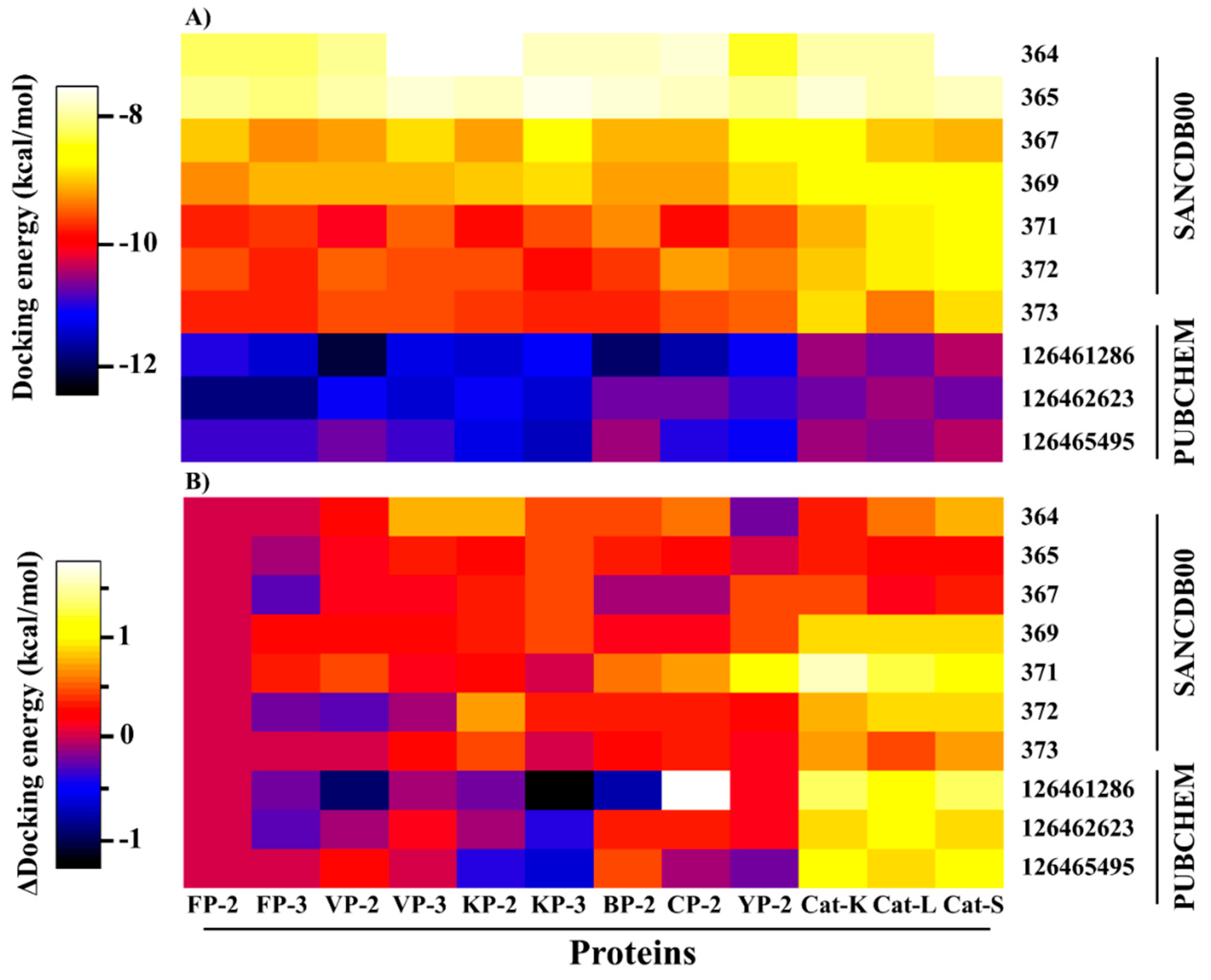
2.2. Potential Plasmodial Cysteine Protease Inhibitors Are Identified from SANCDB
2.3. Drug-Like Properties of the Identified Hits and Analogs are Studied
2.4. Protein-Compound Stability for Each System is Assessed Through Molecular Dynamic Studies
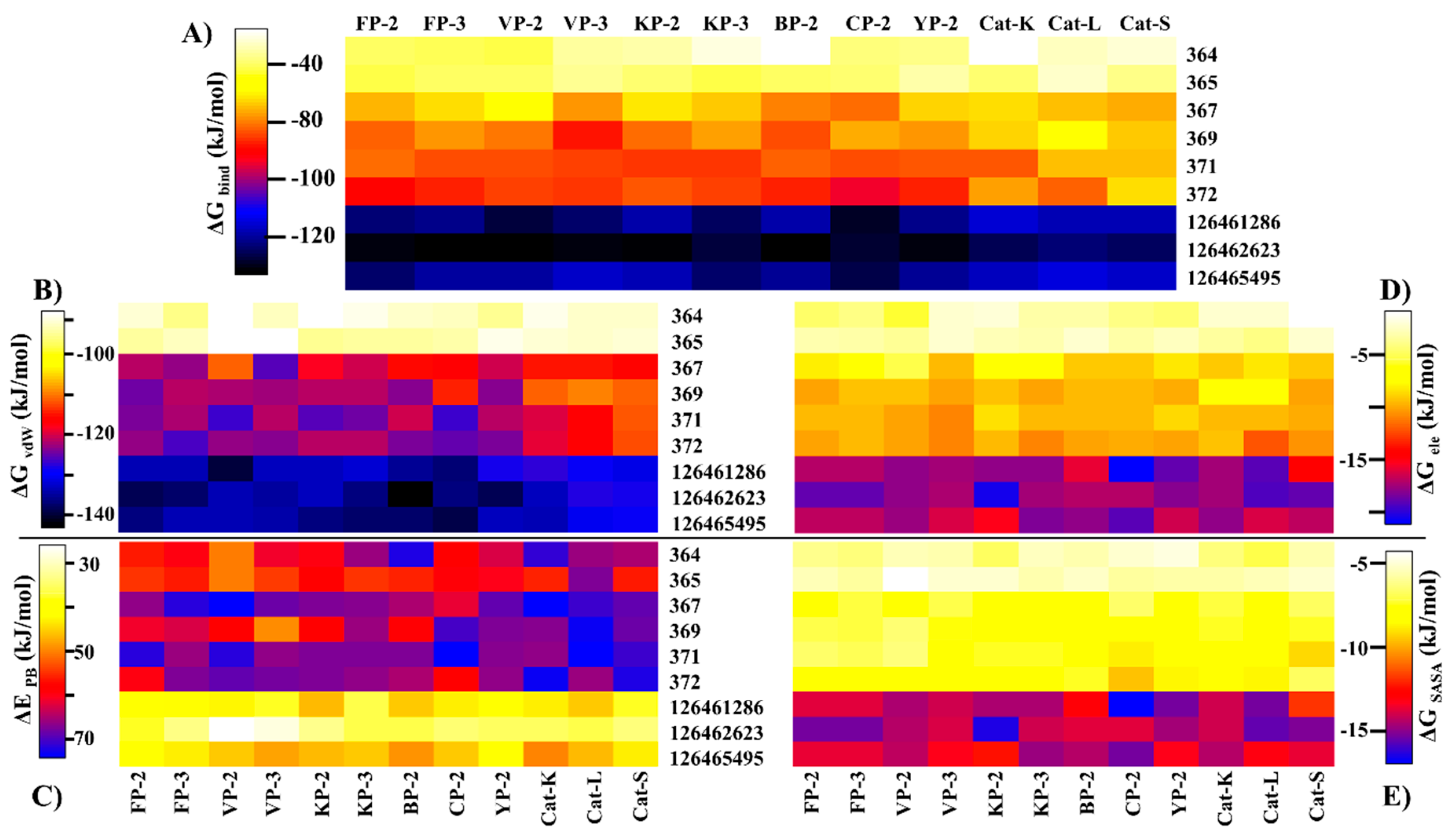
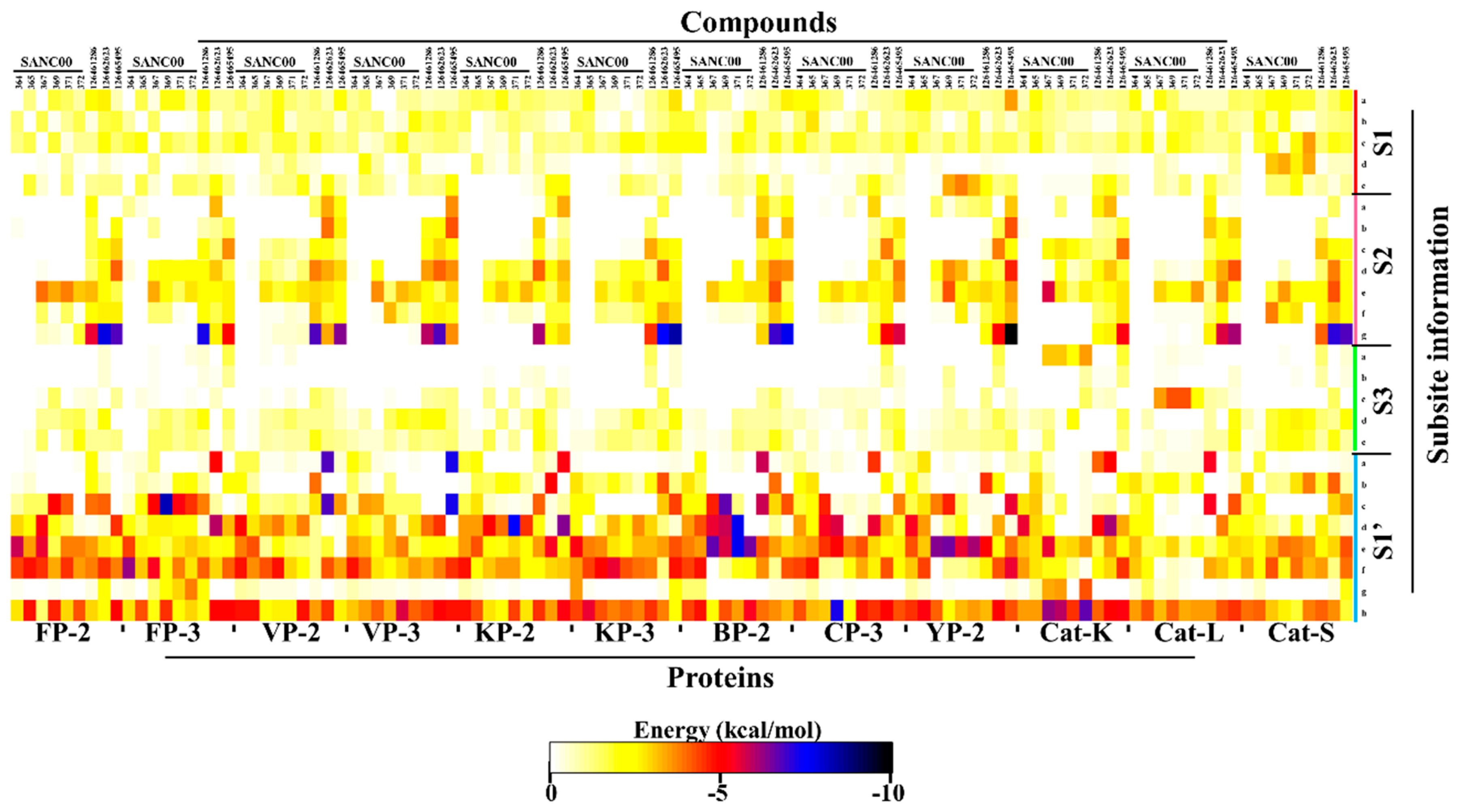
2.5. Key Residues Involved in the Protein Stabilization of Selected Compounds Are Identified via Binding Free Energy
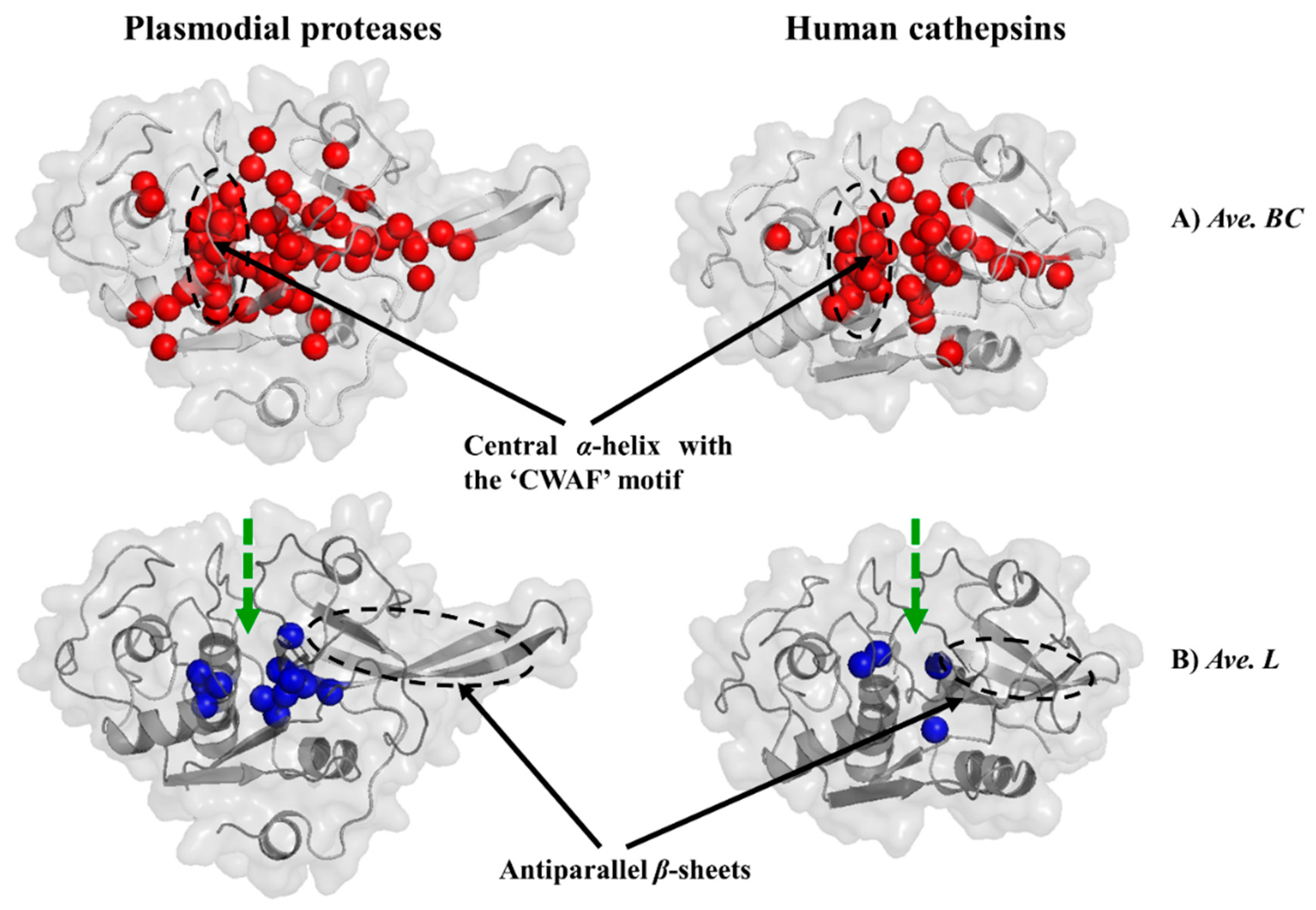
2.6. Important Residues in Protein Communication and More Evidence for Allosteric Sutes are Identified via Dynamic Residue Network Analysis
3. Materials and Methods
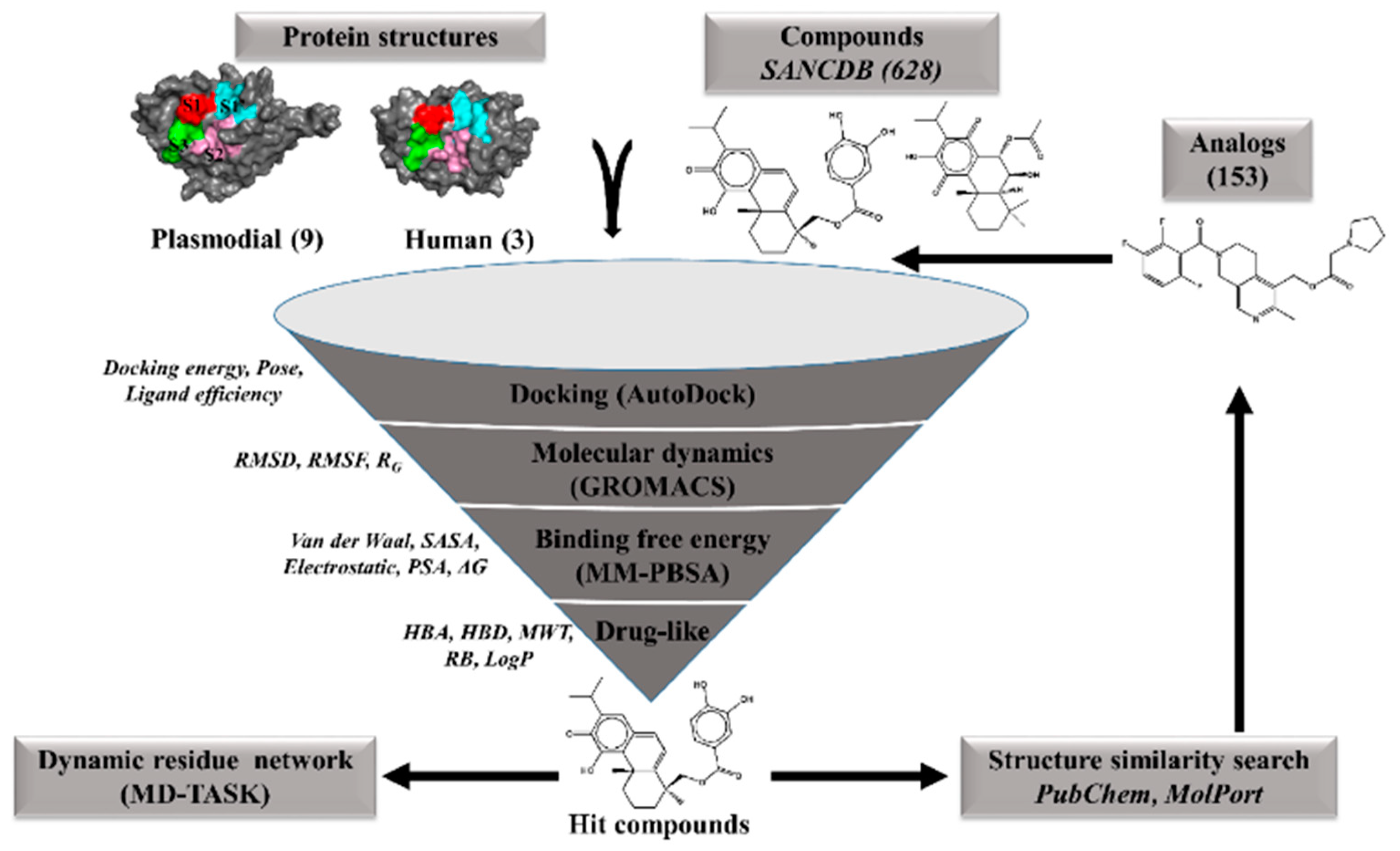
3.1. Identification of Hit Compounds from the South African Natural Compounds Database (SANCDB)
3.2. Molecular Dynamics and Trajectory Analysis
3.3. Binding Free Energy Calculations
3.4. Dynamic Residue Network Analysis
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- World Malaria Report 2018; World Health Organisation: Geneva, Switzerland, 2018.
- Tun, K.M.; Imwong, M.; Lwin, K.M.; Win, A.A.; Hlaing, T.M.; Hlaing, T.; Lin, K.; Kyaw, M.P.; Plewes, K.; Faiz, M.A.; et al. Spread of artemisinin-resistant Plasmodium falciparum in Myanmar: A cross-sectional survey of the K13 molecular marker. Lancet Infect. Dis. 2015, 15, 415–421. [Google Scholar] [CrossRef]
- Takala-Harrison, S.; Jacob, C.G.; Arze, C.; Cummings, M.P.; Silva, J.C.; Dondorp, A.M.; Fukuda, M.M.; Hien, T.T.; Mayxay, M.; Noedl, H.; et al. Independent Emergence of Artemisinin Resistance Mutations Among Plasmodium falciparum in Southeast Asia. J. Infect. Dis. 2015, 211, 670–679. [Google Scholar] [CrossRef] [PubMed]
- Ariey, F.; Witkowski, B.; Amaratunga, C.; Beghain, J.; Langlois, A.-C.; Khim, N.; Kim, S.; Duru, V.; Bouchier, C.; Ma, L.; et al. A molecular marker of artemisinin-resistant Plasmodium falciparum malaria. Nature 2014, 505, 50–55. [Google Scholar] [CrossRef] [PubMed]
- Ashley, E.A.; Dhorda, M.; Fairhurst, R.M.; Amaratunga, C.; Lim, P.; Suon, S.; Sreng, S.; Anderson, J.M.; Mao, S.; Sam, B.; et al. Spread of Artemisinin Resistance in Plasmodium falciparum Malaria. N. Engl. J. Med. 2014, 371, 411–423. [Google Scholar] [CrossRef]
- Takala-Harrison, S.; Laufer, M.K. Antimalarial drug resistance in Africa: Key lessons for the future. Ann. N. Y. Acad. Sci. 2015, 1342, 62–67. [Google Scholar] [CrossRef]
- Rosenthal, P.J.; Rathod, P.K.; Ndiaye, D.; Mharakurwa, S.; Cui, L. Antimalarial Drug Resistance: Literature Review and Activities and Findings of the ICEMR Network. Am. J. Trop. Med. Hyg. 2015, 93, 57–68. [Google Scholar] [CrossRef]
- Francis, S.E.; Sullivan, D.J.; Goldberg, D.E. Hemoglobin Metabolism in the Malaria Parasite Plasmodium Falciparum. Annu. Rev. Microbiol. 2002, 51, 97–123. [Google Scholar] [CrossRef]
- Goldberg, D.E. Plasmodial hemoglobin degradation: An ordered pathway in a specialized organelle. Infect. Agents Dis. 1992, 1, 207–211. [Google Scholar] [CrossRef]
- Deu, E. Proteases as antimalarial targets: Strategies for genetic, chemical, and therapeutic validation. FEBS J. 2017, 284, 2604–2628. [Google Scholar] [CrossRef]
- Goldberg, D.E.; Winzeler, E.A.; Istvan, E.S.; Gluzman, I.; Dharia, N.V.; Bopp, S.E. Validation of isoleucine utilization targets in Plasmodium falciparum. Proc. Natl. Acad. Sci. USA 2011, 108, 1627–1632. [Google Scholar] [CrossRef]
- Gluzman, I.Y.; Liu, J.; Goldberg, D.E.; Gross, J.; Istvan, E.S. Plasmodium falciparum ensures its amino acid supply with multiple acquisition pathways and redundant proteolytic enzyme systems. Proc. Natl. Acad. Sci. USA 2006, 103, 8840–8845. [Google Scholar] [CrossRef]
- Singh, A.; Sijwali, P.S.; Rosenthal, P.J.; Gut, J.; Shenai, B.R. Expression and characterization of the Plasmodium falciparum haemoglobinase falcipain-3. Biochem. J. 2015, 360, 481–489. [Google Scholar] [CrossRef]
- Sijwali, P.S.; Koo, J.; Singh, N.; Rosenthal, P.J. Gene disruptions demonstrate independent roles for the four falcipain cysteine proteases of Plasmodium falciparum. Mol. Biochem. Parasitol. 2006, 150, 96–106. [Google Scholar] [CrossRef] [PubMed]
- Sijwali, P.S.; Rosenthal, P.J. Gene disruption confirms a critical role for the cysteine protease falcipain-2 in hemoglobin hydrolysis by Plasmodium falciparum. Proc. Natl. Acad. Sci. USA 2004, 101, 4384–4389. [Google Scholar] [CrossRef]
- Subramanian, S.; Hardt, M.; Choe, Y.; Niles, R.K.; Johansen, E.B.; Legac, J.; Gut, J.; Kerr, I.D.; Craik, C.S.; Rosenthal, P.J. Hemoglobin cleavage site-specificity of the Plasmodium falciparum cysteine proteases falcipain-2 and falcipain-3. PLoS ONE 2009, 4, e5156. [Google Scholar] [CrossRef] [PubMed]
- Rosenthal, P.J. Falcipains and other cysteine proteases of malaria parasites. Adv. Exp. Med. Biol. 2011, 712, 30–48. [Google Scholar] [CrossRef]
- Pandey, K.C.; Sijwali, P.S.; Craik, C.S.; Shenai, B.R.; Choe, Y.; Singh, A.; Na, B.-K.; Rosenthal, P.J. Identification and biochemical characterization of vivapains, cysteine proteases of the malaria parasite Plasmodium vivax. Biochem. J. 2004, 378, 529–538. [Google Scholar] [CrossRef]
- Prasad, R.; Atul, P.; Soni, A.; Puri, S.K.; Sijwali, P.S. Expression, Characterization, and Cellular Localization of Knowpains, Papain-Like Cysteine Proteases of the Plasmodium knowlesi Malaria Parasite. PLoS ONE 2012, 7, e51619. [Google Scholar] [CrossRef]
- Vaughan, A.M.; Pei, Y.; Kappe, S.H.I.; Lindner, S.E.; Torii, M.; Miller, J.L. Plasmodium yoelii inhibitor of cysteine proteases is exported to exomembrane structures and interacts with yoelipain-2 during asexual blood-stage development. Cell. Microbiol. 2013, 15, 1508–1526. [Google Scholar] [CrossRef]
- Martins, T.M.; Domingos, A.; Gonçalves, L.M.D.; Silveira, H.; do Rosário, V.; Caldeira, R.L.; Novo, C. Plasmodium chabaudi: Expression of active recombinant chabaupain-1 and localization studies in Anopheles sp. Exp. Parasitol. 2009, 122, 97–105. [Google Scholar] [CrossRef]
- Rosenthal, P.J.; Nelson, R.G. Isolation and characterization of a cysteine proteinase gene of Plasmodium falciparum. Mol. Biochem. Parasitol. 1992, 51, 143–152. [Google Scholar] [CrossRef]
- Shenai, B.R.; Sijwali, P.S.; Singh, A.; Rosenthal, P.J. Characterization of native and recombinant falcipain-2, a principal trophozoite cysteine protease and essential hemoglobinase of Plasmodium falciparum. J. Biol. Chem. 2000, 275, 29000–29010. [Google Scholar] [CrossRef] [PubMed]
- Teixeira, C.; Gomes, J.R.B.; Gomes, P. Falcipains, Plasmodium falciparum cysteine proteases as key drug targets against malaria. Curr. Med. Chem. 2011, 18, 1555–1572. [Google Scholar] [CrossRef] [PubMed]
- Kerr, I.D.; Lee, J.H.; Pandey, K.C.; Harrison, A.; Sajid, M.; Rosenthal, P.J.; Brinen, L.S. Structures of falcipain-2 and falcipain-3 bound to small molecule inhibitors: Implications for substrate specificity. J. Med. Chem. 2009, 52, 852–857. [Google Scholar] [CrossRef]
- Musyoka, T.M.; Njuguna, J.N.; Tastan Bishop, Ö. Comparing sequence and structure of falcipains and human homologs at prodomain and catalytic active site for malarial peptide based inhibitor design. Malar. J. 2019, 18, 159. [Google Scholar] [CrossRef]
- Musyoka, T.M.; Kanzi, A.M.; Lobb, K.A.; Tastan Bishop, Ö. Analysis of non-peptidic compounds as potential malarial inhibitors against Plasmodial cysteine proteases via integrated virtual screening workflow. J. Biomol. Struct. Dyn. 2016, 34, 2084–2101. [Google Scholar] [CrossRef]
- Marco, M.; Coteron, J.M. Falcipain inhibition as a promising antimalarial target. Curr. Top. Med. Chem. 2012, 12, 408–444. [Google Scholar] [CrossRef]
- Siddiqui, F.A.; Cabrera, M.; Wang, M.; Brashear, A.; Kemirembe, K.; Wang, Z.; Miao, J.; Chookajorn, T.; Yang, Z.; Cao, Y.; et al. Plasmodium falciparum Falcipain-2a Polymorphisms in Southeast Asia and Their Association With Artemisinin Resistance. J. Infect. Dis. 2018, 218, 434–442. [Google Scholar] [CrossRef]
- Hatherley, R.; Brown, D.K.; Musyoka, T.M.; Penkler, D.L.; Faya, N.; Lobb, K.A.; Tastan Bishop, Ö. SANCDB: A South African natural compound database. J. Cheminform. 2015, 7, 29. [Google Scholar] [CrossRef]
- Musyoka, T.M.; Kanzi, A.M.; Lobb, K.A.; Tastan Bishop, Ö. Structure Based Docking and Molecular Dynamic Studies of Plasmodial Cysteine Proteases against a South African Natural Compound and its Analogs. Sci. Rep. 2016, 6, 1–12. [Google Scholar] [CrossRef]
- Shen, B. A New Golden Age of Natural Products Drug Discovery. Cell 2015, 163, 1297–3000. [Google Scholar] [CrossRef] [PubMed]
- Harvey, A.L.; Edrada-Ebel, R.; Quinn, R.J. The re-emergence of natural products for drug discovery in the genomics era. Nat. Rev. Drug Discov. 2015, 14, 111–129. [Google Scholar] [CrossRef] [PubMed]
- Li, G.; Lou, H.-X. Strategies to diversify natural products for drug discovery. Med. Res. Rev. 2018, 38, 1255–1294. [Google Scholar] [CrossRef] [PubMed]
- Chibale, K.; Guantai, E.M.; Masimirembwa, C. Extracting molecular information from African natural products to facilitate unique African-led drug-discovery efforts. Future Med. Chem. 2011, 3, 257–261. [Google Scholar] [CrossRef] [PubMed]
- Salas-Sarduy, E.; Guerra, Y.; Covaleda Cortés, G.; Avilés, F.; Chávez Planes, M.; Salas-Sarduy, E.; Guerra, Y.; Covaleda Cortés, G.; Avilés, F.X.; Chávez Planes, M.A. Identification of Tight-Binding Plasmepsin II and Falcipain 2 Inhibitors in Aqueous Extracts of Marine Invertebrates by the Combination of Enzymatic and Interaction-Based Assays. Mar. Drugs 2017, 15, 123. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Zhang, S.; Zhu, J.; Zhu, L.; Liu, X.; Shan, L.; Huang, J.; Zhang, W.; Li, H. Identification of diverse natural products as falcipain-2 inhibitors through structure-based virtual screening. Bioorg. Med. Chem. Lett. 2014, 24, 1261–1264. [Google Scholar] [CrossRef]
- van Zyl, R.L.; Khan, F.; Edwards, T.J.; Drewes, S.E. Antiplasmodial activities of some abietane diterpenes from the leaves of Plectranthus species. S. Afri. J. Sci. 2008, 104, 62–64. [Google Scholar]
- Marques, A.F.; Gomes, P.S.F.C.; Oliveira, P.L.; Rosenthal, P.J.; Pascutti, P.G.; Lima, L.M.T.R. Allosteric regulation of the Plasmodium falciparum cysteine protease falcipain-2 by heme. Arch. Biochem. Biophys. 2015, 573, 92–99. [Google Scholar] [CrossRef]
- Sliwoski, G.; Kothiwale, S.; Meiler, J.; Lowe, E.W. Computational methods in drug discovery. Pharmacol. Rev. 2014, 66, 334–395. [Google Scholar] [CrossRef]
- Hung, C.-L.; Chen, C.-C. Computational Approaches for Drug Discovery. Drug Dev. Res. 2014, 75, 412–418. [Google Scholar] [CrossRef]
- Diller, D.J.; Merz, K.M. High throughput docking for library design and library prioritization. Proteins Struct. Funct. Genet. 2001, 43, 113–124. [Google Scholar] [CrossRef]
- Westermaier, Y.; Barril, X.; Scapozza, L. Virtual screening: An in silico tool for interlacing the chemical universe with the proteome. Methods 2015, 71, 44–57. [Google Scholar] [CrossRef] [PubMed]
- Swets, J.A.; Dawes, R.M.; Monahan, J. Better Decisions through Science. Sci. Am. 2000, 283, 82–87. [Google Scholar] [CrossRef] [PubMed]
- Carregal, A.P.; Maciel, F.V.; Carregal, J.B.; dos Reis Santos, B.; da Silva, A.M.; Taranto, A.G. Docking-based virtual screening of Brazilian natural compounds using the OOMT as the pharmacological target database. J. Mol. Model. 2017, 23, 111. [Google Scholar] [CrossRef] [PubMed]
- Mugumbate, G.; Newton, A.S.; Rosenthal, P.J.; Gut, J.; Moreira, R.; Chibale, K.; Guedes, R.C. Novel anti-plasmodial hits identified by virtual screening of the ZINC database. J. Comput. Aided. Mol. Des. 2013, 27, 859–871. [Google Scholar] [CrossRef] [PubMed]
- Triballeau, N.; Acher, F.; Brabet, I.; Pin, J.-P.; Bertrand, H.-O. Virtual Screening Workflow Development Guided by the “Receiver Operating Characteristic” Curve Approach. Application to High-Throughput Docking on Metabotropic Glutamate Receptor Subtype 4. J. Med. Chem. 2004. [Google Scholar] [CrossRef] [PubMed]
- Shityakov, S.; Förster, C.; Shityakov, S. In silico structure-based screening of versatile P-glycoprotein inhibitors using polynomial empirical scoring functions. Adv. Appl. Bioinforma. Chem. 2014, 7–8. [Google Scholar] [CrossRef]
- Empereur-Mot, C.; Zagury, J.-F.O.; Montes, M. Screening Explorer−An Interactive Tool for the Analysis of Screening Results. J. Chem. Inf. Model. 2016, 56, 2281–2286. [Google Scholar] [CrossRef]
- Laskowski, R.A.; Swindells, M.B. LigPlot+: Multiple Ligand–Protein Interaction Diagrams for Drug Discovery. J. Chem. Inf. Model. 2011, 51, 2778–2786. [Google Scholar] [CrossRef]
- Wang, J.; Li, Y.; Yang, Y.; Zhang, S.; Yang, L. Profiling the Structural Determinants of Heteroarylnitrile Scaffold-Based Derivatives as Falcipain-2 Inhibitors by In Silico Methods. Curr. Med. Chem. 2013, 20, 2032–2042. [Google Scholar] [CrossRef]
- Coteron, J.M.; Catterick, D.; Castro, J.; Chaparro, M.J.; Diaz, B.; Fernandez, E.; Ferrer, S.; Gamo, F.J.; Gordo, M.; Gut, J.; et al. Falcipain inhibitors: Optimization studies of the 2-pyrimidinecarbonitrile lead series. J. Med. Chem. 2010, 53, 6129–6152. [Google Scholar] [CrossRef] [PubMed]
- Potshangbam, A.M.; Tanneeru, K.; Reddy, B.M.; Guruprasad, L. 3D-QSAR and molecular docking studies of 2-pyrimidinecarbonitrile derivatives as inhibitors against falcipain-3. Bioorg. Med. Chem. Lett. 2011, 21, 7219–7223. [Google Scholar] [CrossRef] [PubMed]
- Sharma, R.K.; Younis, Y.; Mugumbate, G.; Njoroge, M.; Gut, J.; Rosenthal, P.J.; Chibale, K. Synthesis and structure–activity-relationship studies of thiazolidinediones as antiplasmodial inhibitors of the Plasmodium falciparum cysteine protease falcipain-2. Eur. J. Med. Chem. 2015, 90, 507–518. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.; Thiessen, P.A.; Bolton, E.E.; Chen, J.; Fu, G.; Gindulyte, A.; Han, L.; He, J.; He, S.; Shoemaker, B.A.; et al. PubChem Substance and Compound databases. Nucleic Acids Res. 2016, 44, D1202-13. [Google Scholar] [CrossRef] [PubMed]
- MolPort. Available online: https://www.molport.com (accessed on 22 August 2018).
- Lipinski, C.A.; Lombardo, F.; Dominy, B.W.; Feeney, P.J. Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv. Drug Deliv. Rev. 1997, 23, 3–25. [Google Scholar] [CrossRef]
- Baell, J.B.; Holloway, G.A. New Substructure Filters for Removal of Pan Assay Interference Compounds (PAINS) from Screening Libraries and for Their Exclusion in Bioassays. J. Med. Chem. 2010, 53, 2719–2740. [Google Scholar] [CrossRef]
- Lagorce, D.; Sperandio, O.; Galons, H.; Miteva, M.A.; Villoutreix, B.O. FAF-Drugs2: Free ADME/tox filtering tool to assist drug discovery and chemical biology projects. BMC Bioinform. 2008, 9, 396. [Google Scholar] [CrossRef]
- Baell, J.B. Feeling Nature’s PAINS: Natural Products, Natural Product Drugs, and Pan Assay Interference Compounds (PAINS). J. Nat. Prod. 2016, 79, 616–628. [Google Scholar] [CrossRef]
- Baell, J.B.; Nissink, J.W.M. Seven Year Itch: Pan-Assay Interference Compounds (PAINS) in 2017-Utility and Limitations. ACS Chem. Biol. 2018, 13, 36–44. [Google Scholar] [CrossRef]
- Zhao, H.; Caflisch, A. Molecular dynamics in drug design. Eur. J. Med. Chem. 2014, 1–11. [Google Scholar] [CrossRef]
- Mortier, J.; Rakers, C.; Bermudez, M.; Murgueitio, M.S.; Riniker, S.; Wolber, G. The impact of molecular dynamics on drug design: Applications for the characterization of ligand–macromolecule complexes. Drug Discov. Today 2015, 20, 686–702. [Google Scholar] [CrossRef] [PubMed]
- De Vivo, M.; Masetti, M.; Bottegoni, G.; Cavalli, A. Role of Molecular Dynamics and Related Methods in Drug Discovery. J. Med. Chem. 2016, 59, 4035–4061. [Google Scholar] [CrossRef] [PubMed]
- Abraham, M.J.; Murtola, T.; Schulz, R.; Páll, S.; Smith, J.C.; Hess, B.; Lindahl, E. GROMACS: High performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 2015, 1–2, 19–25. [Google Scholar] [CrossRef]
- Massey, F.J. The Kolmogorov-Smirnov Test for Goodness of Fit. J. Am. Stat. Assoc. 1951, 46, 68–78. [Google Scholar] [CrossRef]
- Penkler, D.L.; Tastan Bishop, Ö. Modulation of Human Hsp90α Conformational Dynamics by Allosteric Ligand Interaction at the C-Terminal Domain. Sci. Rep. 2019, 9, 1600. [Google Scholar] [CrossRef]
- Gilson, M.K.; Zhou, H.-X. Calculation of Protein-Ligand Binding Affinities. Annu. Rev. Biophys. Biomol. Struct. 2007, 36, 21–42. [Google Scholar] [CrossRef]
- Cournia, Z.; Allen, B.; Sherman, W. Relative Binding Free Energy Calculations in Drug Discovery: Recent Advances and Practical Considerations. J. Chem. Inf. Model. 2017, 57, 2911–2937. [Google Scholar] [CrossRef]
- Chodera, J.D.; Mobley, D.L. Entropy-Enthalpy Compensation: Role and Ramifications in Biomolecular Ligand Recognition and Design. Annu. Rev. Biophys. 2013, 42, 121–142. [Google Scholar] [CrossRef]
- Chipot, C. Frontiers in free-energy calculations of biological systems. Wiley Interdiscip. Rev. Comput. Mol. Sci. 2014, 4, 71–89. [Google Scholar] [CrossRef]
- Kumari, R.; Kumar, R.; Lynn, A. g_mmpbsa-A GROMACS Tool for High-Throughput MM-PBSA Calculations. J. Chem. Inf. Model. 2014, 54, 1951–1962. [Google Scholar] [CrossRef]
- Brown, D.K.; Penkler, D.L.; Sheik Amamuddy, O.; Ross, C.; Atilgan, A.R.; Atilgan, C.; Tastan Bishop, Ö. MD-TASK: A software suite for analyzing molecular dynamics trajectories. Bioinformatics 2017, 33, 2768–2771. [Google Scholar] [CrossRef] [PubMed]
- Penkler, D.L.; Atilgan, C.; Tastan Bishop, Ö. Allosteric Modulation of Human Hsp90α Conformational Dynamics. J. Chem. Inf. Model. 2018, 58, 383–404. [Google Scholar] [CrossRef] [PubMed]
- del Sol, A.; Fujihashi, H.; Amoros, D.; Nussinov, R. Residues crucial for maintaining short paths in network communication mediate signaling in proteins. Mol. Syst. Biol. 2006, 2, 2006.0019. [Google Scholar] [CrossRef] [PubMed]
- Hernández Alvarez, L.; Barreto Gomes, D.E.; Hernández González, J.E.; Pascutti, P.G. Dissecting a novel allosteric mechanism of cruzain: A computer-aided approach. PLoS ONE 2019, 14, e0211227. [Google Scholar] [CrossRef] [PubMed]
- Novinec, M. Computational investigation of conformational variability and allostery in cathepsin K and other related peptidases. PLoS ONE 2017, 12, e0182387. [Google Scholar] [CrossRef]
- Marques, A.F.; Esser, D.; Rosenthal, P.J.; Kassack, M.U.; Lima, L.M.T.R. Falcipain-2 inhibition by suramin and suramin analogues. Bioorg. Med. Chem. 2013, 21, 3667–3673. [Google Scholar] [CrossRef]
- Bertoldo, J.B.; Chiaradia-Delatorre, L.D.; Mascarello, A.; Leal, P.C.; Cordeiro, M.N.S.; Nunes, R.J.; Sarduy, E.S.; Rosenthal, P.J.; Terenzi, H. Synthetic compounds from an in house library as inhibitors of falcipain-2 from Plasmodium falciparum. J. Enzym. Inhib. Med. Chem. 2015, 30, 299–307. [Google Scholar] [CrossRef]
- Morris, G.M.; Ruth, H.; Lindstrom, W.; Sanner, M.F.; Belew, R.K.; Goodsell, D.S.; Olson, A.J. Software news and updates AutoDock4 and AutoDockTools4: Automated docking with selective receptor flexibility. J. Comput. Chem. 2009, 30, 2785–2791. [Google Scholar] [CrossRef]
- Bernstein, F.C.; Koetzle, T.F.; Williams, G.J.; Meyer, E.F.; Brice, M.D.; Rodgers, J.R.; Kennard, O.; Shimanouchi, T.; Tasumi, M. The Protein Data Bank: A computer-based archival file for macromolecular structures. J. Mol. Biol. 1977, 112, 535–542. [Google Scholar] [CrossRef]
- Ettari, R.; Bova, F.; Zappala, M.; Grasso, S.; Micale, N.; Zappalà, M. Falcipain-2 inhibitors. Med. Res. Rev. 2010, 30, 136–167. [Google Scholar] [CrossRef]
- Schmidt, I.; Pradel, G.; Sologub, L.; Golzmann, A.; Ngwa, C.J.; Kucharski, A.; Schirmeister, T.; Holzgrabe, U. Bistacrine derivatives as new potent antimalarials. Bioorg. Med. Chem. 2016, 24, 3636–3642. [Google Scholar] [CrossRef] [PubMed]
- Weldon, D.J.; Shah, F.; Chittiboyina, A.G.; Sheri, A.; Chada, R.R.; Gut, J.; Rosenthal, P.J.; Shivakumar, D.; Sherman, W.; Desai, P.; et al. Synthesis, biological evaluation, hydration site thermodynamics, and chemical reactivity analysis of α-keto substituted peptidomimetics for the inhibition of Plasmodium falciparum. Bioorg. Med. Chem. Lett. 2014, 24, 1274–1279. [Google Scholar] [CrossRef] [PubMed]
- Ettari, R.; Pinto, A.; Tamborini, L.; Angelo, I.C.; Grasso, S.; Zappalà, M.; Capodicasa, N.; Yzeiraj, L.; Gruber, E.; Aminake, M.N.; et al. Synthesis and Biological Evaluation of Papain-Family Cathepsin L-Like Cysteine Protease Inhibitors Containing a 1,4-Benzodiazepine Scaffold as Antiprotozoal Agents. ChemMedChem 2014, 9, n/a. [Google Scholar] [CrossRef] [PubMed]
- Conroy, T.; Guo, J.T.; Elias, N.; Cergol, K.M.; Gut, J.; Legac, J.; Khatoon, L.; Liu, Y.; McGowan, S.; Rosenthal, P.J.; et al. Synthesis of Gallinamide A Analogues as Potent Falcipain Inhibitors and Antimalarials. J. Med. Chem. 2014, 57, 10557–10563. [Google Scholar] [CrossRef] [PubMed]
- Huang, H.; Lu, W.; Li, X.; Cong, X.; Ma, H.; Liu, X.; Zhang, Y.; Che, P.; Ma, R.; Li, H.; et al. Design and synthesis of small molecular dual inhibitor of falcipain-2 and dihydrofolate reductase as antimalarial agent. Bioorg. Med. Chem. Lett. 2012, 22, 958–962. [Google Scholar] [CrossRef] [PubMed]
- Mysinger, M.M.; Carchia, M.; Irwin, J.J.; Shoichet, B.K. Directory of Useful Decoys, Enhanced (DUD-E): Better Ligands and Decoys for Better Benchmarking. J. Med. Chem. 2012, 55, 6582–6594. [Google Scholar] [CrossRef] [PubMed]
- Models, M.M.; Philosophy, A.; Kollman, P.A. Advances and Continuing Challenges in Achieving Realistic and Predictive Simulations of the Properties of Organic and Biological Molecules. Acc. Chem. Res. 1996, 29, 461–469. [Google Scholar] [CrossRef]
- Sousa da Silva, A.W.; Vranken, W.F. ACPYPE-AnteChamber PYthon Parser interfacE. BMC Res. Notes 2012, 5, 367. [Google Scholar] [CrossRef]
- Parrinello, M. Polymorphic transitions in single crystals: A new molecular dynamics method. J. Appl. Phys. 1981, 52, 7182. [Google Scholar] [CrossRef]
- Hess, B.; Bekker, H.; Berendsen, H.J.C.; Fraaije, J.G.E.M. LINCS: A linear constraint solver for molecular simulations. J. Comput. Chem. 1997, 18, 1463–1472. [Google Scholar] [CrossRef]
- Petersen, H.G. Accuracy and efficiency of the particle mesh Ewald method. J. Chem. Phys. 1995, 103, 3668. [Google Scholar] [CrossRef]
- Kollman, P.A.; Massova, I.; Reyes, C.; Kuhn, B.; Huo, S.; Chong, L.; Lee, M.; Lee, T.; Duan, Y.; Wang, W.; et al. Calculating Structures and Free Energies of Complex Molecules: Combining Molecular Mechanics and Continuum Models. Acc. Chem. Res. 2000, 33, 889–897. [Google Scholar] [CrossRef] [PubMed]
- Ashley, E.A.; Phyo, A.P. Drugs in Development for Malaria. Drugs 2018, 78, 861–879. [Google Scholar] [CrossRef] [PubMed]
- Nussinov, R.; Tsai, C.-J. Allostery in Disease and in Drug Discovery. Cell 2013, 153, 293–305. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Nussinov, R. Allostery: An Overview of Its History, Concepts, Methods, and Applications. PLOS Comput. Biol. 2016, 12, e1004966. [Google Scholar] [CrossRef] [PubMed]
- Lu, S.; Li, S.; Zhang, J. Harnessing Allostery: A Novel Approach to Drug Discovery. Med. Res. Rev. 2014, 34, 1242–1285. [Google Scholar] [CrossRef]
Sample Availability: Not available. |
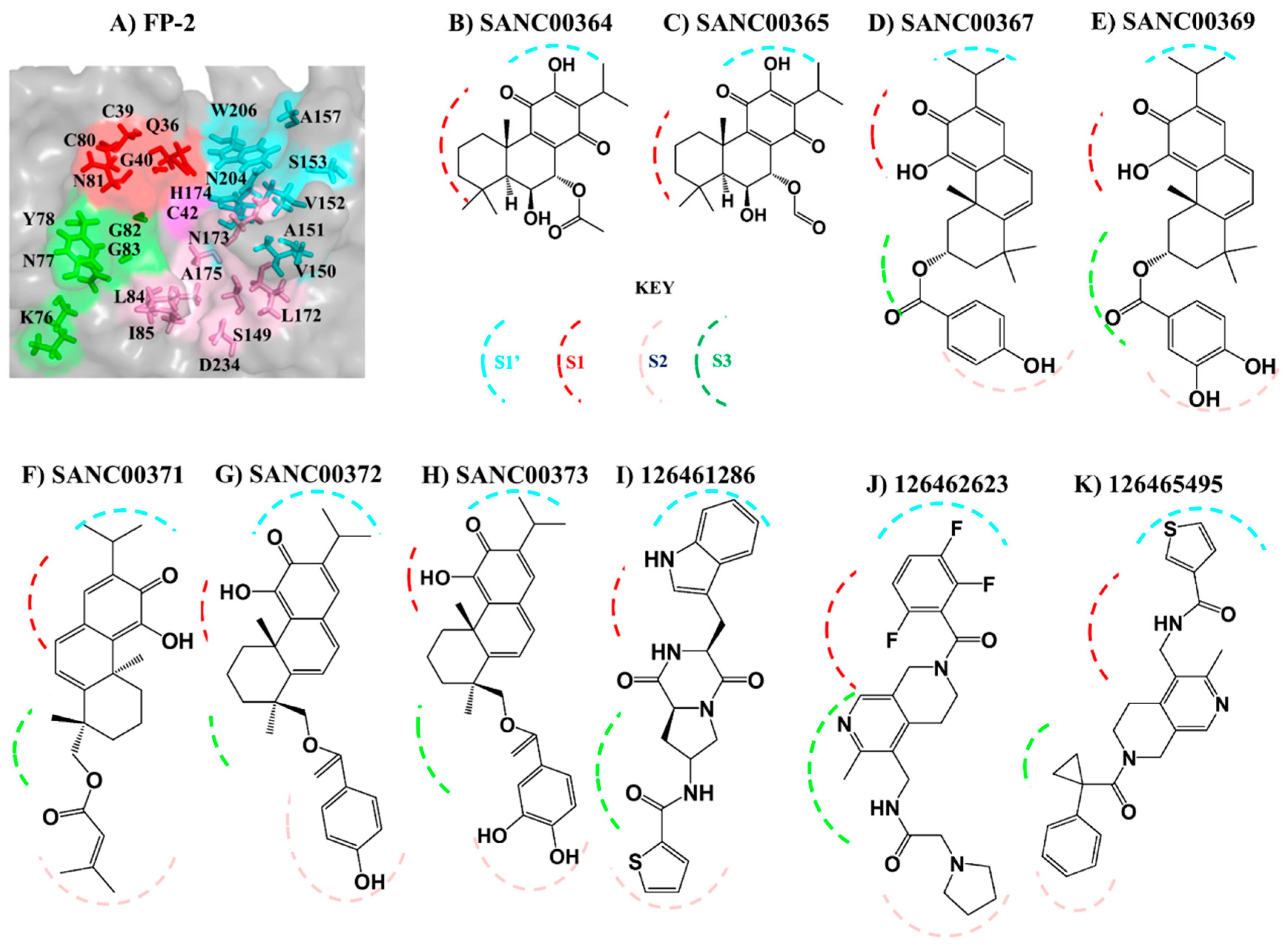
| Compound ID | Chemical Formula | Lipinski’s Rule of Five (RO5) | PAINS | ||||
|---|---|---|---|---|---|---|---|
| Mol. wt | HBA | HBD | nRB | LogP | |||
| SANC00364 | C22H30O6 | 390.2 | 6 | 2 | 3 | 3.9 | Fail |
| SANC00365 | C21H28O6 | 376.2 | 6 | 2 | 3 | 3.6 | Fail |
| SANC00367 | C27H30O5 | 434.2 | 5 | 2 | 4 | 5.0 | Pass |
| SANC00369 | C27H30O6 | 450.2 | 6 | 3 | 4 | 5.4 | Fail |
| SANC00371 | C25H32O4 | 396.2 | 4 | 1 | 5 | 5.2 | Pass |
| SANC00372 | C27H30O5 | 434.2 | 5 | 2 | 5 | 4.7 | Pass |
| SANC00373 | C27H30O6 | 450.2 | 6 | 3 | 5 | 5.0 | Fail |
| CID126461286 | C21H20N4O3S | 408.1 | 7 | 3 | 5 | 0.8 | Pass |
| CID126462623 | C23H25F3N4O2 | 446.2 | 6 | 1 | 7 | -0.4 | Pass |
| CID126465495 | C25H25N3O2S | 431.2 | 5 | 1 | 7 | 1.8 | Pass |
| Protein | Compound | Residues |
|---|---|---|
| FP-2 | SANC00364 | C42, W206 |
| SANC00367 | K37, C39, W207 | |
| SANC00371 | Q36, N38, C39, W206 | |
| 126462623 | Q36, N81, G83, N173, W206 | |
| VP-2 | SANC00364 | D36, W207 |
| SANC00367 | A38, H175, W207 | |
| SANC00371 | Q37, A38, W207 | |
| 126462623 | Q37, G84, N174, W207 | |
| BP-2 | SANC00364 | Q37, W207 |
| SANC00367 | Q37, E82, N174 | |
| SANC00371 | Q37, G84, W207 | |
| 126462623 | Q37, G84, N174, W207 | |
| Cat-K | SANC00364 | W184 |
| SANC00367 | Q19, W184 | |
| SANC00371 | Q19, W184 | |
| 126462623 | Q19, W184 | |
| Cat-L | SANC00364 | Q20 |
| SANC00367 | N19, W190 | |
| SANC00371 | G21, W190 | |
| 126462623 | Q20, G21 |
| Protein | Residues |
|---|---|
| (A) Average BC | |
| FP-2 | W24, T31, C42-Y55, S66, Q68, V71, N86, F89, E90, I93, K126, P132, R141, G144-S149, A151, S153, A175-M183, K196-W206, N217, E219, D221, I237, P238 |
| FP-3 | W26, T33, C44-Y57, S68, Q70, V73, T88, F91, D92, I95, K128, I133, L142, R143, P147, S149-S151, A176-G182, G184, K186, K198-W208, N219, E221, D223, Y227, V239, P240 |
| VP-2 | W25, T32, C43-Y56, S67, Q69, V71, P87, F90, E91, I94, I126, I132, I141, R142, G145-S150, A176-E185, K197-W207, R218, E220, D222, Y226, V238, A239 |
| VP-3 | W24, T31, C42-Y55, S66, Q68, V71, P86, F89, E90, L93, I125, I131, I140, K141, G144-S149, A175-E184, K196-W206, K217, Q219, Y223, V237, A238 |
| KP-2 | W25, T32, C43-Y56, S67, Q69, V72, P87, F90, E91, I94, E131, I141, K142, G145-S150, A176-E185, K197-W207, K218, Q220, V238, A239 |
| KP-3 | W23, T30, C41-Y54, S65, Q67, V70, P85, I88, E89, I92, E129, V139, R140, G143-N148, A174-E184, K195-W205, R216, E218, D220, V237, L238 |
| BP-2 | W25, I32, C43-Y56, S67, Q69, V72, P87, F90, E91, S127, P133, Q142, G145-V151, A176-V184, K197-W207, R218, K220, N222, A237, P238 |
| CP-2 | W25, I32, C43-Y56, S67, Q69, V72, P87, F90, E91, I127, P133, Q142, G145-V151, A176-V184, D197-W207, R218, K220, D222, V237, P238 |
| YP-2 | W25, I32, C43-Y56, S67, Q69, V72, P87, F90, E91, S127, P133, Q142, G145-V151, A176-V184, K197-W207, R218, K220, N222, A237, P238 |
| Cat-K | C25-E35, N52, Y67, N70, A124, G129-S132, A134, N161-A166, K176-W184, Y192 |
| Cat-L | C26-E36, Q52, V55, D72, F75, Q76, A128, S133-D138, G140, G165-G170, K182-W190, Y199 |
| Cat-S | C25-E35, Q51, V54, T72, F75, Q76, A129, S135-D139, R141, G165-170, Y179-186, R197 |
| (B) Average L | |
| FP-2 | S47, I48, S50, V51, I146-S149, M177-G180, K203 |
| FP-3 | S49, V50, S52, V53, I148-S151, I179-G181, K205 |
| VP-2 | T48, V49, V51, V52, P146-V149, V177-V180, K204 |
| VP-3 | T47, V48, V50, V51, I146-S149, I177-G180, K203 |
| KP-2 | T48, V49, V51, V52, I147-S150, I178-G181, R204 |
| KP-3 | T46, V47, V49, V50, I145-N148, I176-G179, K202 |
| BP-2 | T48, A49, V51, V52, I147-A150, M178-G181, R204 |
| CP-2 | T48, A49, V51, I52, L147-A150, I178-G181, R204 |
| YP-2 | T48, A49, V51, V52, I147-A150, M178-G181, R204 |
| Cat-K | A27, F28, V131, V164 |
| Cat-L | A31, T32, V135, V168 |
| Cat-S | A30, V31, V136, V168 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Musyoka, T.; Bishop, Ö.T. South African Abietane Diterpenoids and Their Analogs as Potential Antimalarials: Novel Insights from Hybrid Computational Approaches. Molecules 2019, 24, 4036. https://doi.org/10.3390/molecules24224036
Musyoka T, Bishop ÖT. South African Abietane Diterpenoids and Their Analogs as Potential Antimalarials: Novel Insights from Hybrid Computational Approaches. Molecules. 2019; 24(22):4036. https://doi.org/10.3390/molecules24224036
Chicago/Turabian StyleMusyoka, Thommas, and Özlem Tastan Bishop. 2019. "South African Abietane Diterpenoids and Their Analogs as Potential Antimalarials: Novel Insights from Hybrid Computational Approaches" Molecules 24, no. 22: 4036. https://doi.org/10.3390/molecules24224036
APA StyleMusyoka, T., & Bishop, Ö. T. (2019). South African Abietane Diterpenoids and Their Analogs as Potential Antimalarials: Novel Insights from Hybrid Computational Approaches. Molecules, 24(22), 4036. https://doi.org/10.3390/molecules24224036







