Determination of Adulteration Content in Extra Virgin Olive Oil Using FT-NIR Spectroscopy Combined with the BOSS–PLS Algorithm
Abstract
:1. Introduction
2. Results
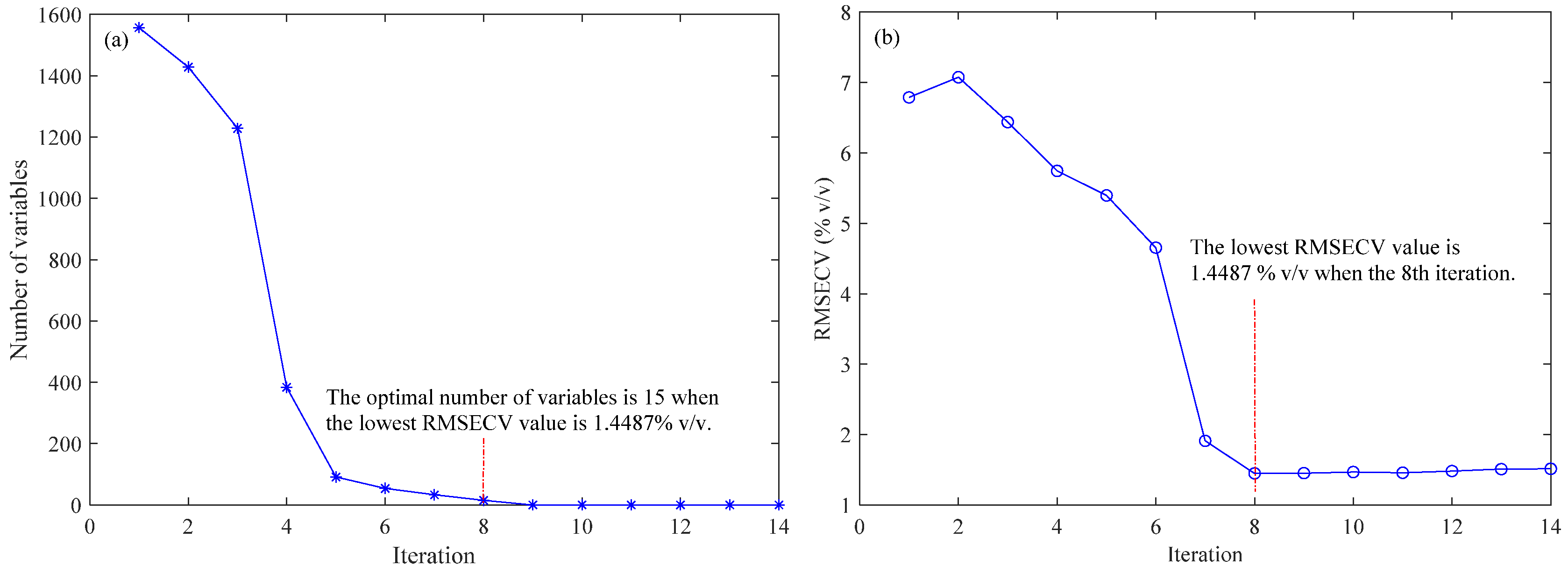
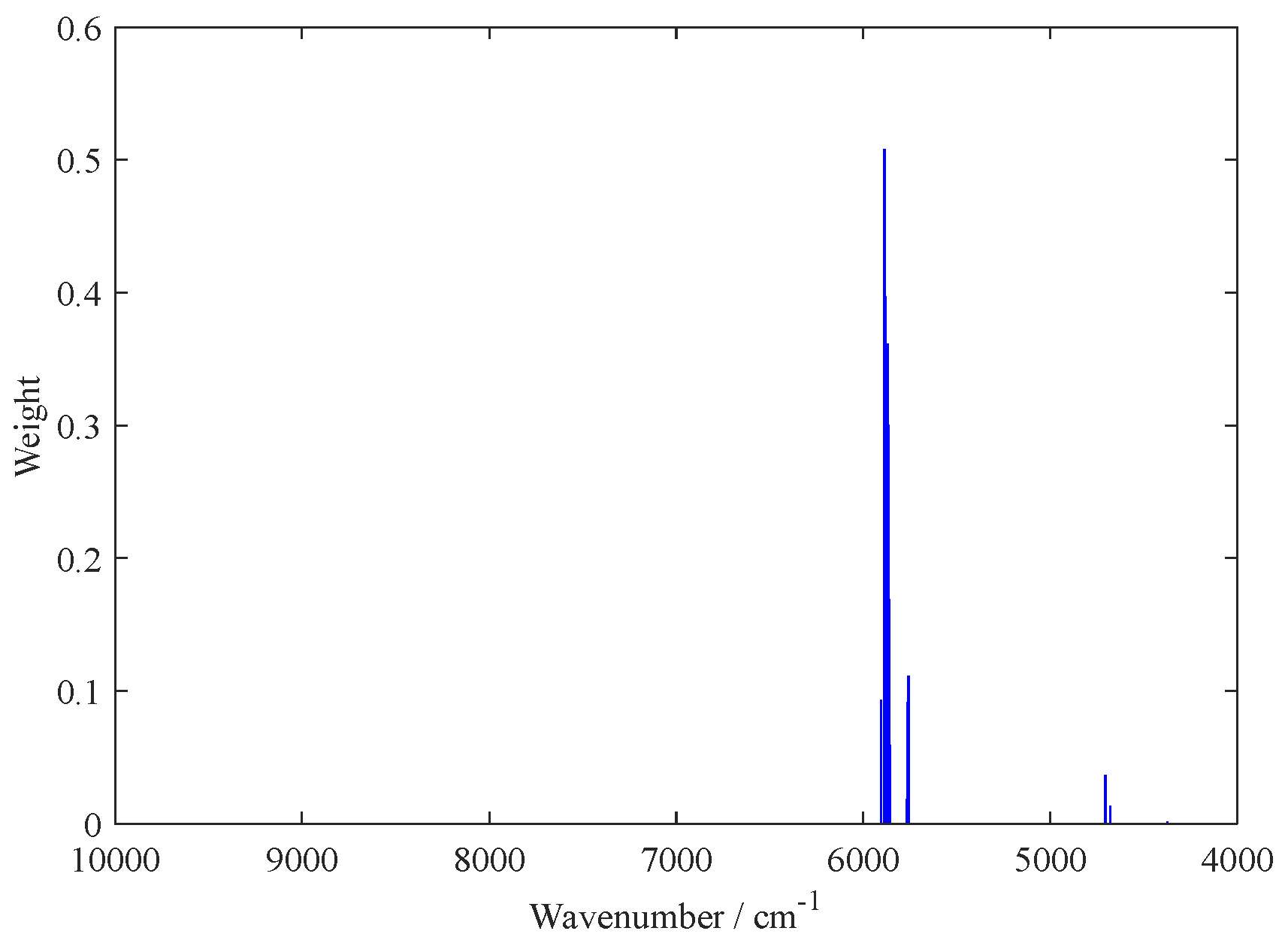
2.1. Variable Selection by the BOSS Algorithm
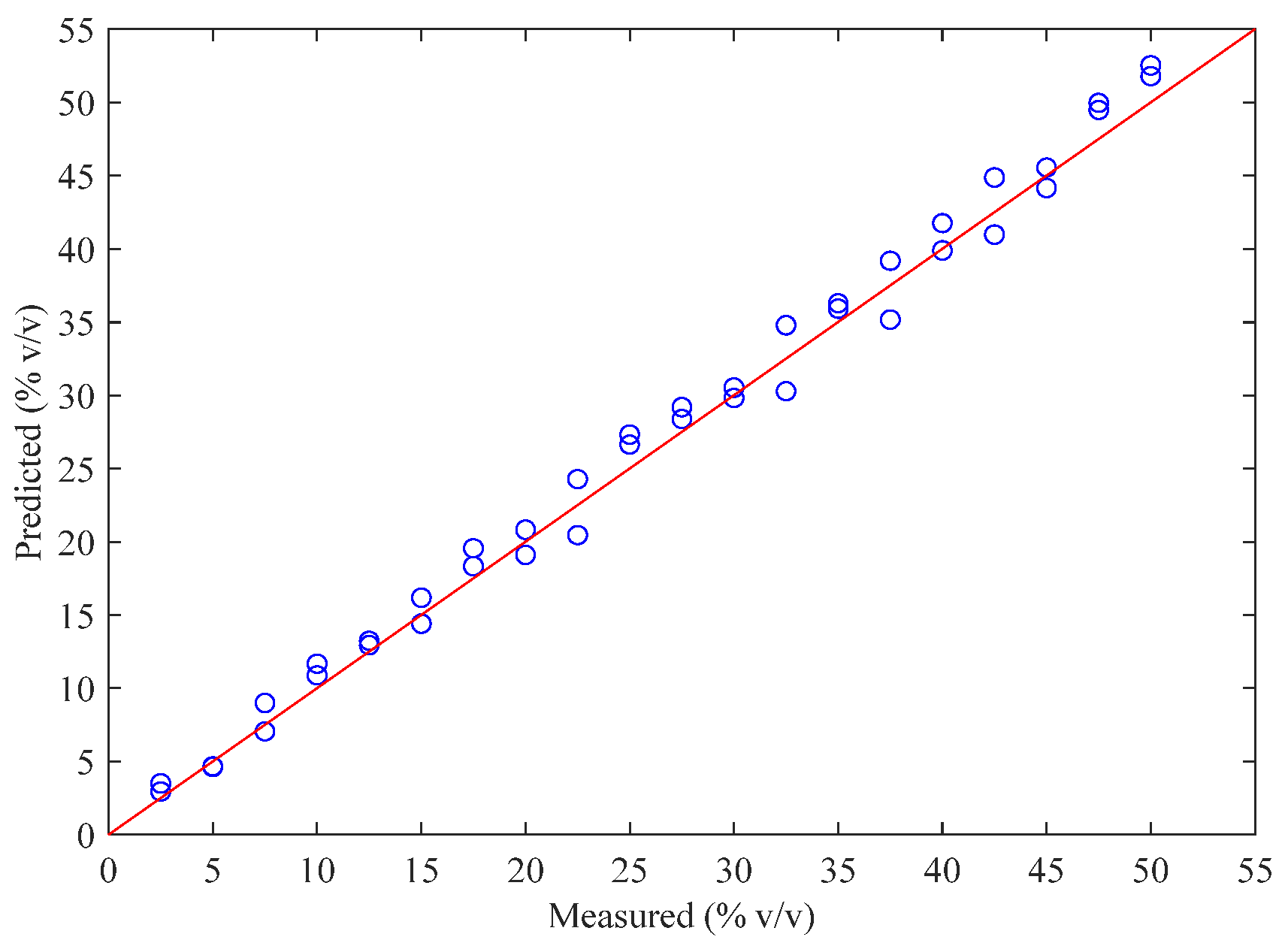
2.2. Results of the PLS Model
3. Discussion
4. Materials and Methods
4.1. Sample Preparation and Division
4.2. FT-NIR Spectra Acquisition
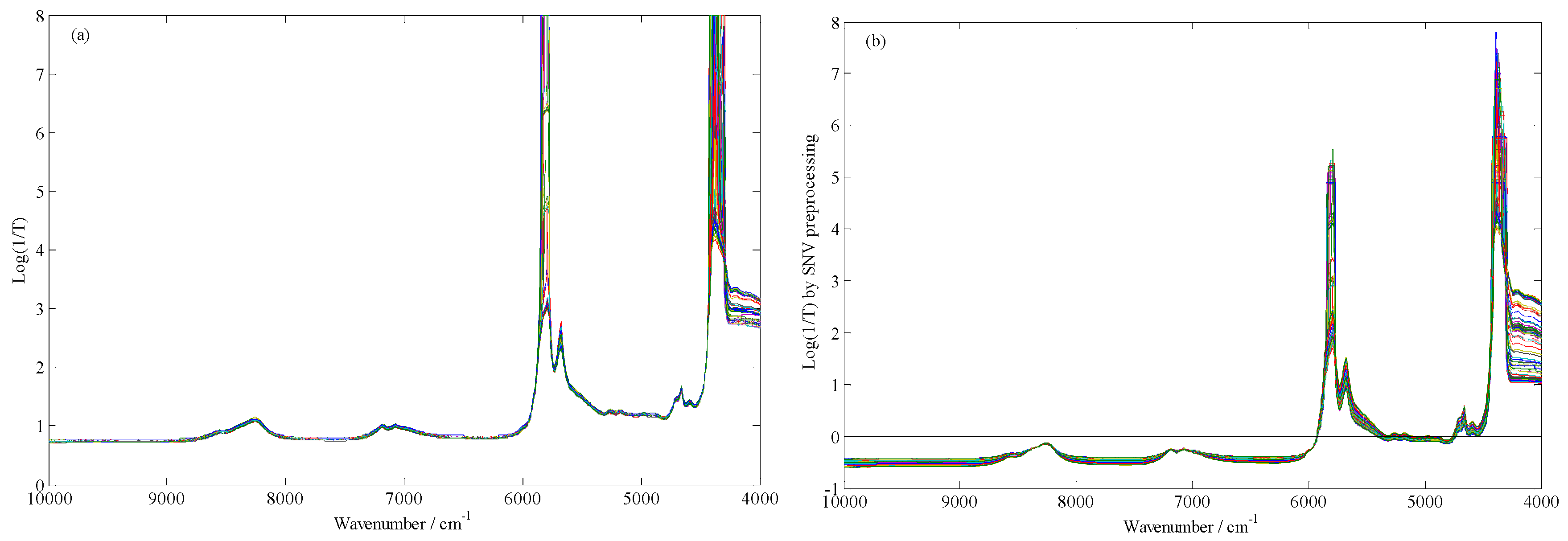
4.3. Spectra Preprocessing
4.4. Data Analyses Methods
4.5. Model Evaluation
4.6. Software
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Chen, H.; Lin, Z.; Tan, C. Fast quantitative detection of sesame oil adulteration by near-infrared spectroscopy and chemometric models. Vib. Spectrosc. 2018, 99, 178–183. [Google Scholar] [CrossRef]
- Xu, Y.; Li, H.; Chen, Q.; Zhao, J.; Ouyang, Q. Rapid detection of adulteration in extra-virgin olive oil using three-dimensional fluorescence spectra technology with selected multivariate calibrations. Int. J. Food Prop. 2015, 18, 2085–2098. [Google Scholar] [CrossRef]
- Chen, Q.; Chen, M.; Liu, Y.; Wu, J.; Wang, X.; Ouyang, Q.; Chen, X.H. Application of FT-NIR spectroscopy for simultaneous estimation of taste quality and taste-related compounds content of black tea. J. Food Sci. Tech. Mys. 2018, 55, 4363–4368. [Google Scholar] [CrossRef] [PubMed]
- Hu, W.; He, R.; Hou, F.; Ouyang, Q.; Chen, Q. Real-time monitoring of alcalase hydrolysis of egg white protein using near infrared spectroscopy technique combined with efficient modeling algorithm. Int. J. Food Prop. 2017, 20, 1488–1499. [Google Scholar] [CrossRef]
- Guo, Z.; Huang, W.; Peng, Y.; Chen, Q.; Ouyang, Q.; Zhao, J. Color compensation and comparison of shortwave near infrared and long wave near infrared spectroscopy for determination of soluble solids content of ‘Fuji’ apple. Postharvest Biol. Technol. 2016, 115, 81–90. [Google Scholar] [CrossRef]
- Zhang, H.; Jiang, H.; Liu, G.; Mei, C.; Huang, Y. Identification of Radix puerariae starch from different geographical origins by FT-NIR spectroscopy. Int. J. Food Prop. 2017, 20, 1567–1577. [Google Scholar] [CrossRef]
- Azizian, H.; Mossoba, M.M.; Fardin-Kia, A.R.; Karunathilaka, S.R.; Kramer, J.K.G. Developing FT-NIR and PLS1 methodology for predicting adulteration in representative varieties/blends of extra virgin olive oils. Lipids 2016, 51, 1309–1321. [Google Scholar] [CrossRef]
- Mossoba, M.M.; Azizian, H.; Fardin-Kia, A.R.; Karunathilaka, S.R.; Kramer, J.K.G. First application of newly developed FT-NIR spectroscopic methodology to predict authenticity of extra virgin olive oil retail products in the USA. Lipids 2017, 52, 443–455. [Google Scholar] [CrossRef]
- Ozdemir, I.S.; Dag, C.; Ozinanc, G.; Sucsoran, O.; Ertas, E.; Bekiroglu, S. Quantification of sterols and fatty acids of extra virgin olive oils by FT-NIR spectroscopy and multivariate statistical analyses. Lwt-Food Sci. Technol. 2018, 91, 125–132. [Google Scholar] [CrossRef]
- Mabood, F.; Boque, R.; Folcarelli, R.; Busto, O.; Jabeen, F.; Al-Harrasi, A.; Hussain, J. The effect of thermal treatment on the enhancement of detection of adulteration in extra virgin olive oils by synchronous fluorescence spectroscopy and chemometric analysis. Spectrochim. Acta A 2016, 161, 83–87. [Google Scholar] [CrossRef]
- Tan, J.; Li, R.; Jiang, Z.T.; Shi, M.; Xiao, Y.Q.; Jia, B.; Lu, T.X.; Wang, H. Detection of extra virgin olive oil adulteration with edible oils using front-face fluorescence and visible spectroscopies. J. Am. Oil Chem. Soc. 2018, 95, 535–546. [Google Scholar] [CrossRef]
- Tavares Melo Milanez, K.D.; Araujo Nobrega, T.C.; Nascimento, D.S.; Insausti, M.; Fernandez Band, B.S.; Coelho Pontes, M.J. Multivariate modeling for detecting adulteration of extra virgin olive oil with soybean oil using fluorescence and UV-Vis spectroscopies: A preliminary approach. Lwt-Food Sci. Technol. 2017, 85, 9–15. [Google Scholar] [CrossRef]
- Poiana, M.A.; Alexa, E.; Munteanu, M.F.; Gligor, R.; Moigradean, D.; Mateescu, C. Use of ATR-FTIR spectroscopy to detect the changes in extra virgin olive oil by adulteration with soybean oil and high temperature heat treatment. Open Chem. 2015, 13, 689–698. [Google Scholar] [CrossRef]
- Sun, X.; Lin, W.; Li, X.; Shen, Q.; Luo, H. Detection and quantification of extra virgin olive oil adulteration with edible oils by FT-IR spectroscopy and chemometrics. Anal. Methods 2015, 7, 3939–3945. [Google Scholar] [CrossRef]
- Xu, Y.; Hassan, M.M.; Kutsanedzie, F.Y.H.; Li, H.H.; Chen, Q.S. Evaluation of extra-virgin olive oil adulteration using FTIR spectroscopy combined with multivariate algorithms. Qual. Assur. Saf. Crops Foods 2018, 10, 411–421. [Google Scholar] [CrossRef]
- Dong, W.; Zhang, Y.; Zhang, B.; Wang, X. Quantitative analysis of adulteration of extra virgin olive oil using Raman spectroscopy improved by Bayesian framework least squares support vector machines. Anal. Methods 2012, 4, 2772–2777. [Google Scholar] [CrossRef]
- Philippidis, A.; Poulakis, E.; Papadaki, A.; Velegrakis, M. Comparative study using Raman and visible spectroscopy of cretan extra virgin olive oil adulteration with sunflower oil. Anal. Lett. 2017, 5, 1182–1195. [Google Scholar] [CrossRef]
- Tiryaki, G.Y.; Ayvaz, H. Quantification of soybean oil adulteration in extra virgin olive oil using portable raman spectroscopy. J. Food Meas. Charact. 2017, 11, 523–529. [Google Scholar] [CrossRef]
- Fragaki, G.; Spyros, A.; Siragakis, G.; Salivaras, E.; Dais, P. Detection of extra virgin olive oil adulteration with lampante olive oil and refined olive oil using nuclear magnetic resonance spectroscopy and multivariate statistical analysis. J. Agric. Food Chem. 2005, 53, 2810–2816. [Google Scholar] [CrossRef]
- Jiang, X.Y.; Li, C.; Chen, Q.Q.; Weng, X.C. Comparison of F-19 and H-1 NMR spectroscopy with conventional methods for the detection of extra virgin olive oil adulteration. Grasas Aceites 2018, 69. [Google Scholar] [CrossRef]
- Zhu, J.; Agyekum, A.A.; Kutsanedzie, F.Y.H.; Li, H.; Chen, Q.; Ouyang, Q.; Jiang, H. Qualitative and quantitative analysis of chlorpyrifos residues in tea by surface-enhanced Raman spectroscopy (SERS) combined with chemometric models. Lwt-Food Sci. Techol. 2018, 97, 760–769. [Google Scholar] [CrossRef]
- Ouyang, Q.; Chen, Q.; Zhao, J.; Lin, H. Determination of amino acid nitrogen in soy sauce using near infrared spectroscopy combined with characteristic variables selection and extreme learning machine. Food Bioprocess Technol. 2013, 6, 2486–2493. [Google Scholar] [CrossRef]
- Jiang, H.; Mei, C.; Li, K.; Huang, Y.; Chen, Q. Monitoring alcohol concentration and residual glucose in solid state fermentation of ethanol using FT-NIR spectroscopy and L1-PLS regression. Spectrochim. Acta A 2018, 204, 73–80. [Google Scholar] [CrossRef] [PubMed]
- Liu, G.H.; Jiang, H.; Xiao, X.H.; Zhang, D.J.; Mei, C.L.; Ding, Y.H. Determination of process variable pH in solid-state fermentation by FT-NIR spectroscopy and extreme learning machine (ELM). Spectrosc. Spect. Anal. 2012, 32, 970–973. [Google Scholar]
- Wold, S.; Sjöström, M.; Eriksson, L. PLS-regression: A basic tool of chemometrics. Chemom. Intell. Lab. Syst. 2001, 58, 109–130. [Google Scholar] [CrossRef]
- Lin, Y.W.; Deng, B.C.; Wang, L.L.; Xu, Q.S.; Liu, L.; Liang, Y.Z. Fisher optimal subspace shrinkage for block variable selection with applications to NIR spectroscopic analysis. Chemom. Intell. Lab. Syst. 2016, 159, 196–204. [Google Scholar] [CrossRef]
- Farrés, M.; Platikanov, S.; Tsakovski, S.; Tauler, R. Comparison of the variable importance in projection (VIP) and of the selectivity ratio (SR) methods for variable selection and interpretation. J. Chemom. 2015, 29, 528–536. [Google Scholar] [CrossRef]
- Li, H.D.; Zeng, M.M.; Tan, B.B.; Liang, Y.Z.; Xu, Q.S.; Cao, D.S. Recipe for revealing informative metabolites based on model population analysis. Metabolomics 2010, 6, 353–361. [Google Scholar] [CrossRef]
- Cai, W.; Li, Y.; Shao, X.A. Variable selection method based on uninformative variable elimination for multivariate calibration of near-infrared spectra. Chemom. Intell. Lab. Syst. 2008, 90, 188–194. [Google Scholar] [CrossRef]
- Rajalahti, T.; Arneberg, R.; Kroksveen, A.C.; Berle, M.; Myhr, K.M.; Kvalheim, O.M. Discriminating variable test and selectivity ratio plot: Quantitative tools for interpretation and variable (biomarker) selection in complex spectral or chromatographic profiles. Anal. Chem. 2009, 81, 2581–2590. [Google Scholar] [CrossRef]
- Deng, B.C.; Yun, Y.H.; Liang, Y.Z. Model population analysis in chemometrics. Chemom. Intell. Lab. Syst. 2015, 149, 166–176. [Google Scholar] [CrossRef]
- Yun, Y.H.; Wang, W.T.; Tan, M.L.; Liang, Y.Z.; Li, H.D.; Cao, D.S. A strategy that iteratively retains informative variables for selecting optimal variable subset in multivariate calibration. Anal. Chim. Acta 2014, 807, 36–43. [Google Scholar] [CrossRef] [PubMed]
- Deng, B.C.; Yun, Y.H.; Liang, Y.Z.; Yi, L.Z. A novel variable selection approach that iteratively optimizes variable space using weighted binary matrix sampling. Analyst 2014, 139, 4836–4845. [Google Scholar] [CrossRef] [PubMed]
- Deng, B.C.; Yun, Y.H.; Ma, P.; Lin, C.C.; Ren, D.B.; Liang, Y.Z. A new method for wavelength interval selection that intelligently optimizes the locations, widths and combinations of the intervals. Analyst 2015, 140, 1876–1885. [Google Scholar] [CrossRef] [PubMed]
- Yun, Y.H.; Wang, W.T.; Deng, B.C.; Lai, G.B.; Liu, X.B.; Ren, D.B. Using variable combination population analysis for variable selection in multivariate calibration. Anal. Chim. Acta 2015, 862, 14–23. [Google Scholar] [CrossRef] [PubMed]
- Deng, B.C.; Yun, Y.H.; Cao, D.S.; Yin, Y.L.; Wang, W.T.; Lu, H.M. A bootstrapping soft shrinkage approach for variable selection in chemical modeling. Anal. Chim. Acta 2016, 908, 63–74. [Google Scholar] [CrossRef]
- Linden, A.; Adams, J.L.; Roberts, N. Evaluating disease management program effectiveness—An introduction to the bootstrap technique. Dis. Manag. Health Out. 2005, 13, 159–167. [Google Scholar] [CrossRef]
- Ma, S.G.; Kosorok, M.R. Robust serniparametric M-estimation and the weighted bootstrap. J. Multivariate Anal. 2005, 96, 190–217. [Google Scholar] [CrossRef]
Sample Availability: Samples of the compounds are available from the authors. |
| Models | Selected Wavenumbers (cm−1) | Number of Variables | PLS Factors | Calibration Set | Validation Set | ||
|---|---|---|---|---|---|---|---|
| R2 | RMSECV | R2 | RMSEP | ||||
| PLS | 9999.10-3999.64 | 1557 | 6 | 0.9421 | 3.4618 | 0.9599 | 3.2520 |
| CARS-PLS | 4192.49; 4242.63; 4261.92; 4578.18; 4593.61; 4655.32; 4659.18; 4666.89; 4670.75; 4674.60; 4682.32; 4690.03; 5746.83; 5754.55; 5758.40; 5766.12; 5858.68; 5862.54; 5870.25; 5874.11; 5877.97; 5881.82; 5885.68; 5889.54; 5897.25; 5901.11; 5912.68; 5920.39; 5935.82; 8234.55 | 30 | 4 | 0.9617 | 2.9647 | 0.9683 | 2.7664 |
| MCUVE-PLS | 4373.76; 4412.33; 4566.61; 4593.61 4612.89; 4632.18; 4647.61 4670.75; 4690.03; 4709.32; 5750.69; 5762.26; 5777.69; 5866.40; 5885.68; 5904.97; 5924.25; 5939.68; 6001.39; 6028.39; 8238.41; 8253.84; 8261.55; 8265.41 | 24 | 3 | 0.9694 | 2.6828 | 0.9778 | 2.3232 |
| IRIV-PLS | 4373.76; 4412.33; 5750.69; 5754.55; 5758.40; 5762.26; 5769.97; 5773.83; 5777.69; 5854.83; 5858.68; 5862.54; 5866.40; 5874.11 | 14 | 2 | 0.9901 | 1.4877 | 0.9887 | 1.8471 |
| BOSS-PLS | 4373.76; 4678.46; 4705.46; 5758.40; 5762.26; 5766.12; 5777.69; 5858.68; 5862.54; 5866.40; 5870.25; 5877.97; 5881.82; 5885.68; 5904.97 | 15 | 3 | 0.9908 | 1.4487 | 0.9922 | 1.4889 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jiang, H.; Chen, Q. Determination of Adulteration Content in Extra Virgin Olive Oil Using FT-NIR Spectroscopy Combined with the BOSS–PLS Algorithm. Molecules 2019, 24, 2134. https://doi.org/10.3390/molecules24112134
Jiang H, Chen Q. Determination of Adulteration Content in Extra Virgin Olive Oil Using FT-NIR Spectroscopy Combined with the BOSS–PLS Algorithm. Molecules. 2019; 24(11):2134. https://doi.org/10.3390/molecules24112134
Chicago/Turabian StyleJiang, Hui, and Quansheng Chen. 2019. "Determination of Adulteration Content in Extra Virgin Olive Oil Using FT-NIR Spectroscopy Combined with the BOSS–PLS Algorithm" Molecules 24, no. 11: 2134. https://doi.org/10.3390/molecules24112134
APA StyleJiang, H., & Chen, Q. (2019). Determination of Adulteration Content in Extra Virgin Olive Oil Using FT-NIR Spectroscopy Combined with the BOSS–PLS Algorithm. Molecules, 24(11), 2134. https://doi.org/10.3390/molecules24112134







