Aggregating Knockoffs for False Discovery Rate Control with an Application to Gut Microbiome Data
Abstract
1. Introduction
- We introduce an aggregation scheme that provably retains the original methods’ guarantees—see Theorem 1.
- We show numerically that the aggregation can increase the original methods’ power—see Section 3.1 and Section 3.2.
- We show that the resulting pipelines for FDR control can be readily applied to empirical data and lead to new discoveries—see Section 3.3.
2. Methods and Theory
2.1. A Brief Introduction to the Knockoff Filter
2.2. Aggregating Knockoffs
2.3. Other Approaches
3. Simulations and a Real Data Analysis
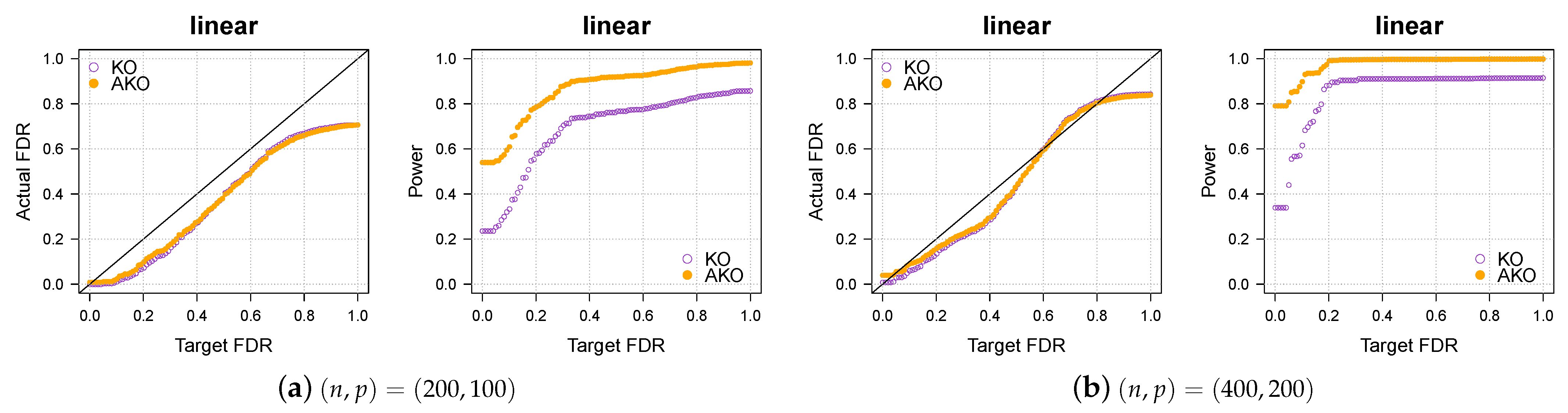
3.1. Simulation 1: Linear Regression
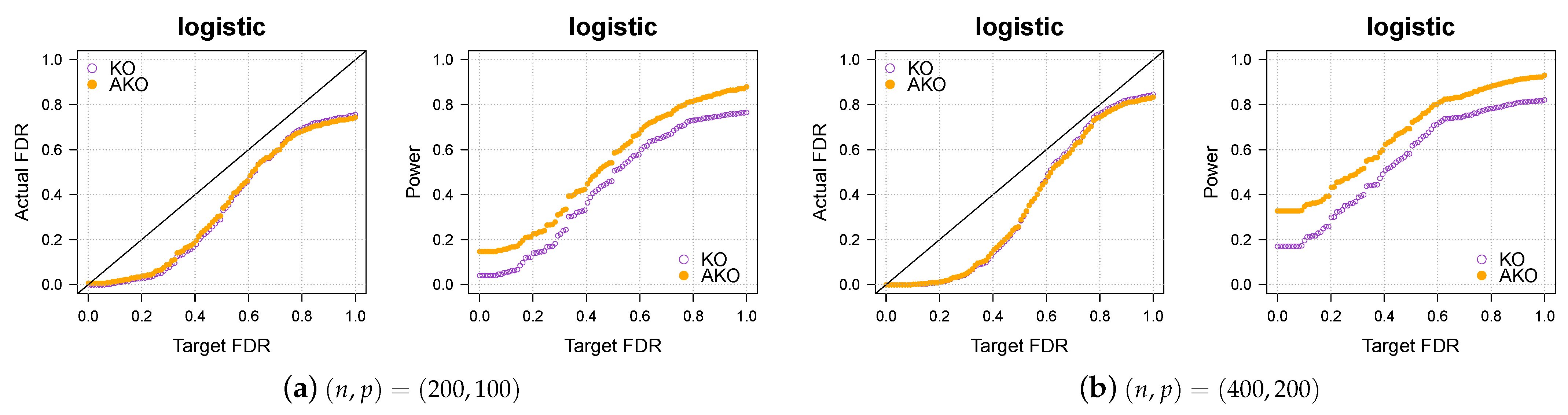
3.2. Simulation 2: Logistic Regression
3.3. Influence of the Gut Microbiome on Obesity
4. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Appendix A. Additional Explanations
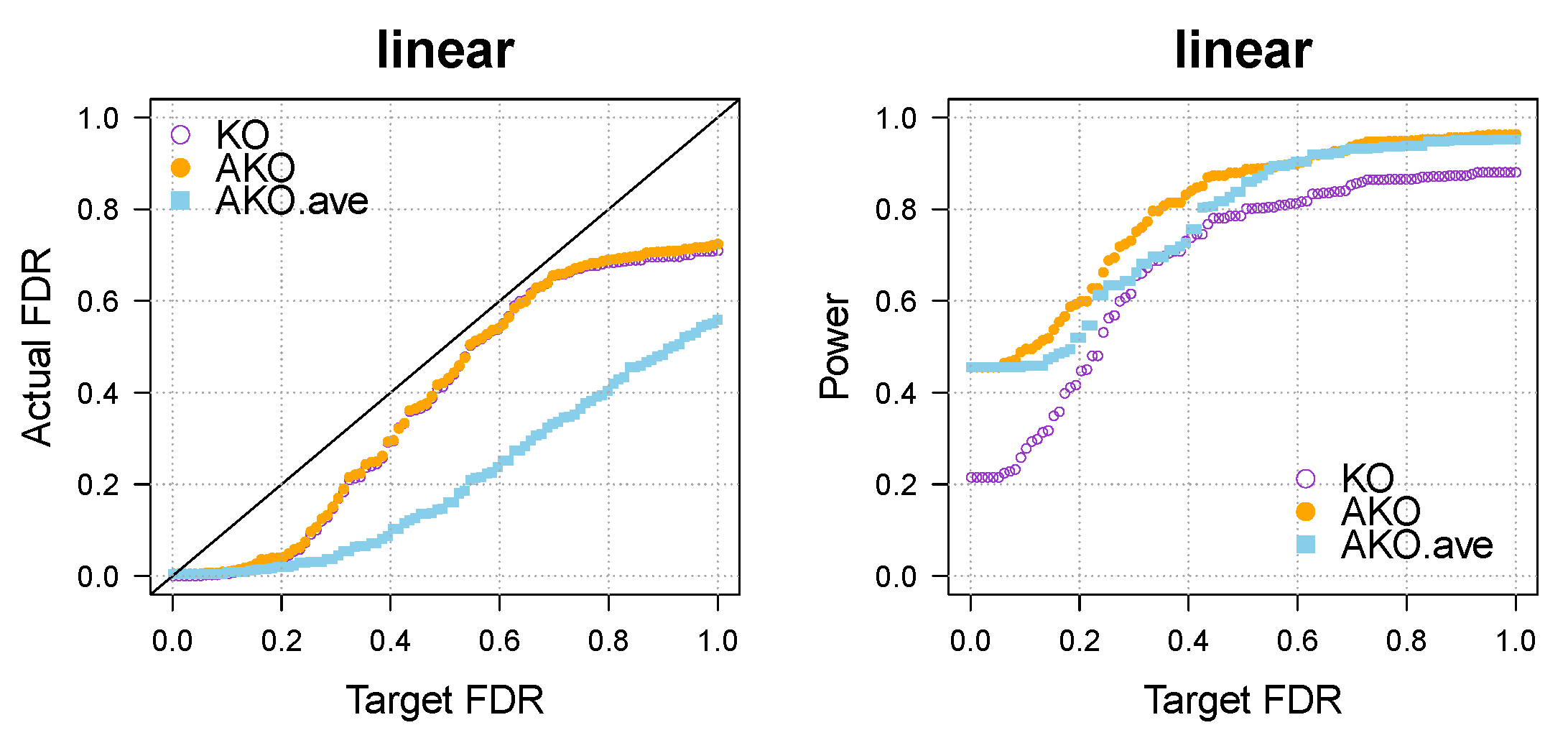
Appendix A.1. Further Simulations for Comparison to Multiple Knockoffs (MKO)

Appendix A.2. Choice of q1,…,qk
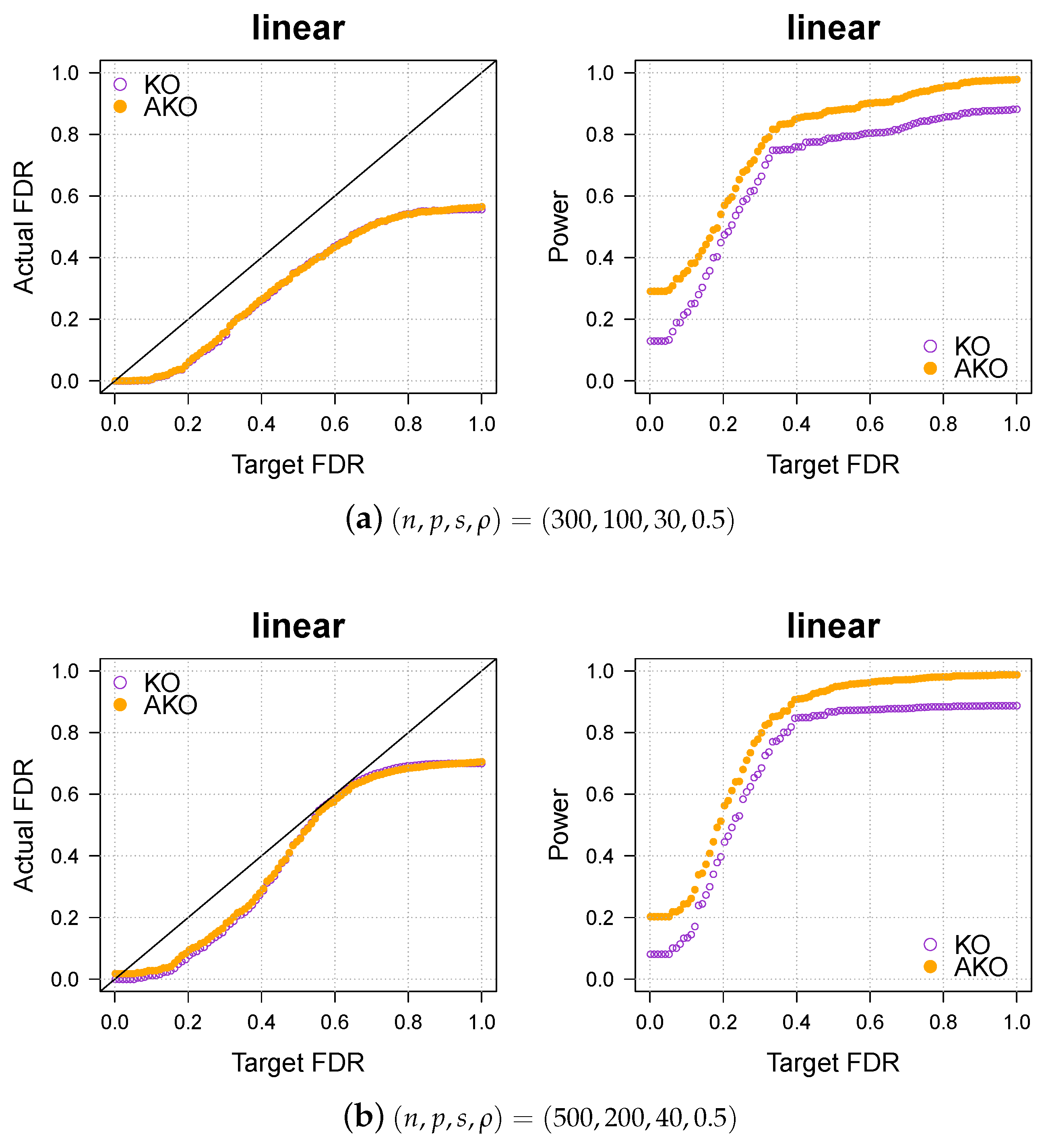
Appendix A.3. Various Settings for the Simulation Part
Appendix A.4. Better Than other Competitors (under the AGP Data)
| (i) ALL | |||
|---|---|---|---|
| BH | TreeFDR | KO | AKO |
| Actinobacteria | Actinobacteria | ||
| Bacteroidetes | |||
| Cyanobacteria | Cyanobacteria | Cyanobacteria | |
| Proteobacteria | Proteobacteria | ||
| Spirochaetes | |||
| Synergistetes | Synergistetes | ||
| Tenericutes | Tenericutes | ||
| Verrucomicrobia | Verrucomicrobia | Verrucomicrobia | |
Appendix B. Additional Results on the Genera Rank
| Phylum | KO | AKO |
|---|---|---|
| Actinobacteria | Collinsella | Collinsella |
| Firmicutes | Lachnospira | |
| Acidaminococcus | ||
| Catenibacterium | ||
| Tenericutes | RF39 | RF39 |
| Phylum | KO | AKO |
|---|---|---|
| Actinobacteria | Actinomyces | Actinomyces |
| Collinsella | Collinsella | |
| Cyanobacteria | YS2 | YS2 |
| Firmicutes | Bacillus | Bacillus |
| Lactococcus | ||
| Lachnospira | Lachnospira | |
| Ruminococcus | Ruminococcus | |
| Acidaminococcus | Acidaminococcus | |
| Megasphaera | Megasphaera | |
| Mogibacteriaceae | ||
| Erysipelotrichaceae | ||
| Catenibacterium | Catenibacterium | |
| Proteobacteria | RF32 | RF32 |
| Haemophilus | ||
| Tenericutes | RF39 | RF39 |
| ML615J-28 |
| Phylum | KO | AKO |
|---|---|---|
| Actinobacteria | Eggerthella | Eggerthella |
| Cyanobacteria | YS2 | YS2 |
| Streptophyta | Streptophyta | |
| Firmicutes | Bacillus | |
| Clostridium | Clostridium | |
| Lachnospira | Lachnospira | |
| Acidaminococcus | Acidaminococcus | |
| 1-68 | ||
| Erysipelotrichaceae | Erysipelotrichaceae | |
| Catenibacterium | ||
| Proteobacteria | Haemophilus | Haemophilus |
| Phylum | KO | AKO |
|---|---|---|
| Actinobacteria | Actinomyces | Actinomyces |
| Collinsella | Collinsella | |
| Cyanobacteria | YS2 | YS2 |
| Firmicutes | Bacillus | Bacillus |
| Lactococcus | ||
| Lachnospira | Lachnospira | |
| Ruminococcus | Ruminococcus | |
| Acidaminococcus | Acidaminococcus | |
| Megasphaera | Megasphaera | |
| Mogibacteriaceae | ||
| SHA-98 | ||
| Erysipelotrichaceae | ||
| Catenibacterium | Catenibacterium | |
| Proteobacteria | RF32 | RF32 |
| Haemophilus | ||
| Tenericutes | RF39 | RF39 |
| ML615J-28 |
| Phylum | KO | AKO |
|---|---|---|
| Actinobacteria | Eggerthella | Eggerthella |
| Cyanobacteria | YS2 | YS2 |
| Streptophyta | Streptophyta | |
| Firmicutes | Bacillus | Bacillus |
| Lactobacillus | ||
| Clostridium | Clostridium | |
| Lachnospira | Lachnospira | |
| Veillonellaceaes | ||
| Acidaminococcus | Acidaminococcus | |
| 1-68 | 1-68 | |
| Erysipelotrichaceae | Erysipelotrichaceae | |
| Catenibacterium | Catenibacterium | |
| Eubacterium | Eubacterium | |
| Proteobacteria | RF32 | |
| Haemophilus | Haemophilus |
| Phylum | KO | AKO |
|---|---|---|
| Actinobacteria | Actinomyces | Actinomyces |
| Collinsella | Collinsella | |
| Eggerthella | Eggerthella | |
| Cyanobacteria | YS2 | YS2 |
| Firmicutes | Bacillus | Bacillus |
| Lachnospira | Lachnospira | |
| Ruminococcus | Ruminococcus | |
| Acidaminococcus | Acidaminococcus | |
| Megasphaera | Megasphaera | |
| Erysipelotrichaceae | Erysipelotrichaceae | |
| Catenibacterium | Catenibacterium | |
| Proteobacteria | RF32 | RF32 |
| Haemophilus | Haemophilus | |
| Tenericutes | RF39 |
References
- Evans, J.M.; Morris, L.S.; Marchesi, J.R. The gut microbiome: The role of a virtual organ in the endocrinology of the host. J. Endocrinol. 2013, 218, R37–R47. [Google Scholar] [CrossRef]
- Huttenhower, C.; Gevers, D.; Knight, R.; Abubucker, S.; Badger, J.H.; Chinwalla, A.T.; Creasy, H.H.; Earl, A.M.; FitzGerald, M.G.; Fulton, R.S.; et al. The Human Microbiome Project Consortium: Structure, function and diversity of the healthy human microbiome. Nature 2012, 486, 207–214. [Google Scholar]
- Koliada, A.; Syzenko, G.; Moseiko, V.; Budovska, L.; Puchkov, K.; Perederiy, V.; Gavalko, Y.; Dorofeyev, A.; Romanenko, M.; Tkach, S. Association between body mass index and Firmicutes/Bacteroidetes ratio in an adult Ukrainian population. BMC Microbiol. 2017, 17, 120. [Google Scholar] [CrossRef]
- Ley, R.E.; Turnbaugh, P.J.; Klein, S.; Gordon, J.I. Microbial ecology: Human gut microbes associated with obesity. Nature 2006, 444, 1022. [Google Scholar] [CrossRef] [PubMed]
- Knight Lab. American Gut Project. Available online: http://americangut.org (accessed on 11 June 2019).
- Ng, A.Y. Feature selection, L 1 vs. L 2 regularization, and rotational invariance. In Proceedings of the 21st International Conference on Machine Learning, Banff, AL, Canada, 4–8 July 2004; p. 78. [Google Scholar]
- Barber, R.F.; Candès, E.J. Controlling the false discovery rate via knockoffs. Ann. Stat. 2015, 43, 2055–2085. [Google Scholar] [CrossRef]
- Barber, R.F.; Candès, E.J.; Samworth, R.J. Robust inference with knockoffs. arXiv 2018, arXiv:1801.03896. [Google Scholar] [CrossRef]
- Candès, E.J.; Fan, Y.; Janson, L.; Lv, J. Panning for gold: ‘Model-X’knockoffs for high dimensional controlled variable selection. J. R. Stat. Soc. Ser. (Stat. Methodol.) 2018, 80, 551–577. [Google Scholar] [CrossRef]
- Romano, Y.; Sesia, M.; Candès, E.J. Deep Knockoffs. J. Am. Stat. Assoc. 2019, 115, 1861–1872. [Google Scholar] [CrossRef]
- Jordon, J.; Yoon, J.; van der Schaar, M. KnockoffGAN: Generating Knockoffs for Feature Selection using Generative Adversarial Networks. In Proceedings of the International Conference on Learning Representations, New Orleans, LA, USA, 9 May 2019. [Google Scholar]
- Holden, L.; Hellton, K.H. Multiple Model-Free Knockoffs. arXiv 2018, arXiv:1812.04928. [Google Scholar]
- Gimenez, J.R.; Zou, J. Improving the stability of the knockoff procedure: Multiple simultaneous knockoffs and entropy maximization. In Proceedings of the 22nd International Conference on Artificial Intelligence and Statistics, Naha, Okinawa, Japan, 19 April 2019; pp. 2184–2192. [Google Scholar]
- Lu, J.; Shi, P.; Li, H. Generalized linear models with linear constraints for microbiome compositional data. Biometrics 2019, 75, 235–244. [Google Scholar] [CrossRef]
- Aitchison, J. The statistical analysis of compositional data. J. R. Stat. Soc. Ser. (Methodol.) 1982, 44, 139–160. [Google Scholar] [CrossRef]
- Naqvi, A.; Rangwala, H.; Keshavarzian, A.; Gillevet, P. Network-based modeling of the human gut microbiome. Chem. Biodivers. 2010, 7, 1040–1050. [Google Scholar] [CrossRef]
- Aitchison, J. The Statistical Analysis of Compositional Data; Blackburn Press: Caldwell, NJ, USA, 2003. [Google Scholar]
- Kurtz, Z.D.; Müller, C.L.; Miraldi, E.R.; Littman, D.R.; Blaser, M.J.; Bonneau, R.A. Sparse and Compositionally Robust Inference of Microbial Ecological Networks. PLoS Comput. Biol. 2015, 11, 1–25. [Google Scholar] [CrossRef] [PubMed]
- Klose, S.; Lederer, J. A Pipeline for Variable Selection and False Discovery Rate Control With an Application in Labor Economics. arXiv 2020, arXiv:2006.12296. [Google Scholar]
- Escobar, J.S.; Klotz, B.; Valdes, B.E.; Agudelo, G.M. The gut microbiota of Colombians differs from that of Americans, Europeans and Asians. BMC Microbiol. 2014, 14, 311. [Google Scholar] [CrossRef] [PubMed]
- Gérard, P. Gut microbiota and obesity. Cell. Mol. Life Sci. 2016, 73, 147–162. [Google Scholar] [CrossRef]
- Turnbaugh, P.J.; Gordon, J.I. The core gut microbiome, energy balance and obesity. J. Physiol. 2009, 587, 4153–4158. [Google Scholar] [CrossRef] [PubMed]
- Bai, J.; Hu, Y.; Bruner, D.W. Composition of gut microbiota and its association with body mass index and lifestyle factors in a cohort of 7-18 years old children from the American Gut Project. Pediatr. Obes. 2019, 14, e12480. [Google Scholar] [CrossRef] [PubMed]
- Clarke, S.F.; Murphy, E.F.; Nilaweera, K.; Ross, P.R.; Shanahan, F.; O’Toole, P.W.; Cotter, P.D. The gut microbiota and its relationship to diet and obesity. Gut Microbes 2012, 3, 186–202. [Google Scholar] [CrossRef] [PubMed]
- Depommier, C.; Everard, A.; Druart, C.; Plovier, H.; Van Hul, M.; Vieira-Silva, S.; Falony, G.; Raes, J.; Maiter, D.; Delzenne, N.M.; et al. Supplementation with Akkermansia muciniphila in overweight and obese human volunteers: A proof-of-concept exploratory study. Nat. Med. 2019, 25, 1096–1103. [Google Scholar] [CrossRef] [PubMed]
- Gao, X.; Zhang, M.; Xue, J.; Huang, J.; Zhuang, R.; Zhou, X.; Zhang, H.; Fu, Q.; Hao, Y. Body Mass Index Differences in the Gut Microbiota Are Gender Specific. Front. Microbiol. 2018, 9, 1250. [Google Scholar] [CrossRef] [PubMed]
- Benjamini, Y.; Hochberg, Y. Controlling the false discovery rate: A practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B 1995, 289–300. [Google Scholar] [CrossRef]
- Xiao, J.; Cao, H.; Chen, J. False discovery rate control incorporating phylogenetic tree increases detection power in microbiome-wide multiple testing. Bioinformatics 2017, 33, 2873–2881. [Google Scholar] [CrossRef] [PubMed]
- Srinivasan, A.; Xue, L.; Zhan, X. Compositional knockoff filter for high-dimensional regression analysis of microbiome data. Biometrics 2020. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, T.B.; Chevalier, J.A.; Thirion, B.; Arlot, S. Aggregation of multiple knockoffs. In Proceedings of the 37th International Conference on Machine Learning, Virtual Conference, Online. 18 July 2020. [Google Scholar]
| (i) all | (ii) uw + ob | ||
| KO | AKO | KO | AKO |
| Actinobacteria | Actinobacteria | Actinobacteria | Actinobacteria |
| Bacteroidetes | |||
| Cyanobacteria | Cyanobacteria | Cyanobacteria | |
| Firmicutes | |||
| Proteobacteria | Proteobacteria | ||
| Spirochaetes | |||
| Synergistetes | Synergistetes | Synergistetes | |
| Tenericutes | Tenericutes | Tenericutes | Tenericutes |
| Verrucomicrobia | |||
| (iii) nor + ob | (iv) ow + ob | ||
| KO | AKO | KO | AKO |
| Actinobacteria | Actinobacteria | Actinobacteria | |
| Bacteroidetes | Bacteroidetes | ||
| Cyanobacteria | Cyanobacteria | Cyanobacteria | Cyanobacteria |
| Firmicutes | |||
| Lentisphaerae | |||
| Proteobacteria | Proteobacteria | Proteobacteria | |
| Spirochaetes | Spirochaetes | ||
| Synergistetes | Synergistetes | Synergistetes | |
| TM7 | |||
| Tenericutes | Tenericutes | Tenericutes | Tenericutes |
| Verrucomicrobia | |||
| Thermi | |||
| (v) uw + nor + ob | (vi) uw + ow + ob | ||
| KO | AKO | KO | AKO |
| Actinobacteria | Actinobacteria | Actinobacteria | |
| Bacteroidetes | Bacteroidetes | ||
| Cyanobacteria | Cyanobacteria | Cyanobacteria | Cyanobacteria |
| Firmicutes | |||
| Lentisphaerae | |||
| Proteobacteria | Proteobacteria | Proteobacteria | |
| Spirochaetes | Spirochaetes | ||
| Synergistetes | Synergistetes | Synergistetes | |
| TM7 | |||
| Tenericutes | Tenericutes | Tenericutes | Tenericutes |
| (vii) nor+ow+ob | |||
| KO | AKO | ||
| Actinobacteria | |||
| Bacteroidetes | |||
| Cyanobacteria | Cyanobacteria | ||
| Proteobacteria | Proteobacteria | ||
| Spirochaetes | |||
| Synergistetes | Synergistetes | ||
| Tenericutes | Tenericutes | ||
| Verrucomicrobia | |||
| Phylum | KO | AKO |
|---|---|---|
| Actinobacteria | Actinomyces | Actinomyces |
| Collinsella | Collinsella | |
| Eggerthella | Eggerthella | |
| Cyanobacteria | YS2 | YS2 |
| Streptophyta | ||
| Firmicutes | Bacillus | Bacillus |
| Lactobacillus | ||
| Lactococcus | Lactococcus | |
| Clostridium | ||
| Lachnospira | Lachnospira | |
| Ruminococcus | Ruminococcus | |
| Peptostreptococcaceae | ||
| Acidaminococcus | Acidaminococcus | |
| Megasphaera | Megasphaera | |
| Mogibacteriaceae | ||
| Erysipelotrichaceae | Erysipelotrichaceae | |
| Catenibacterium | Catenibacterium | |
| Proteobacteria | RF32 | RF32 |
| Haemophilus | Haemophilus | |
| Tenericutes | RF39 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Xie, F.; Lederer, J. Aggregating Knockoffs for False Discovery Rate Control with an Application to Gut Microbiome Data. Entropy 2021, 23, 230. https://doi.org/10.3390/e23020230
Xie F, Lederer J. Aggregating Knockoffs for False Discovery Rate Control with an Application to Gut Microbiome Data. Entropy. 2021; 23(2):230. https://doi.org/10.3390/e23020230
Chicago/Turabian StyleXie, Fang, and Johannes Lederer. 2021. "Aggregating Knockoffs for False Discovery Rate Control with an Application to Gut Microbiome Data" Entropy 23, no. 2: 230. https://doi.org/10.3390/e23020230
APA StyleXie, F., & Lederer, J. (2021). Aggregating Knockoffs for False Discovery Rate Control with an Application to Gut Microbiome Data. Entropy, 23(2), 230. https://doi.org/10.3390/e23020230






