A Systems Approach to Study Immuno- and Neuro-Modulatory Properties of Antiviral Agents
Abstract
1. Introduction
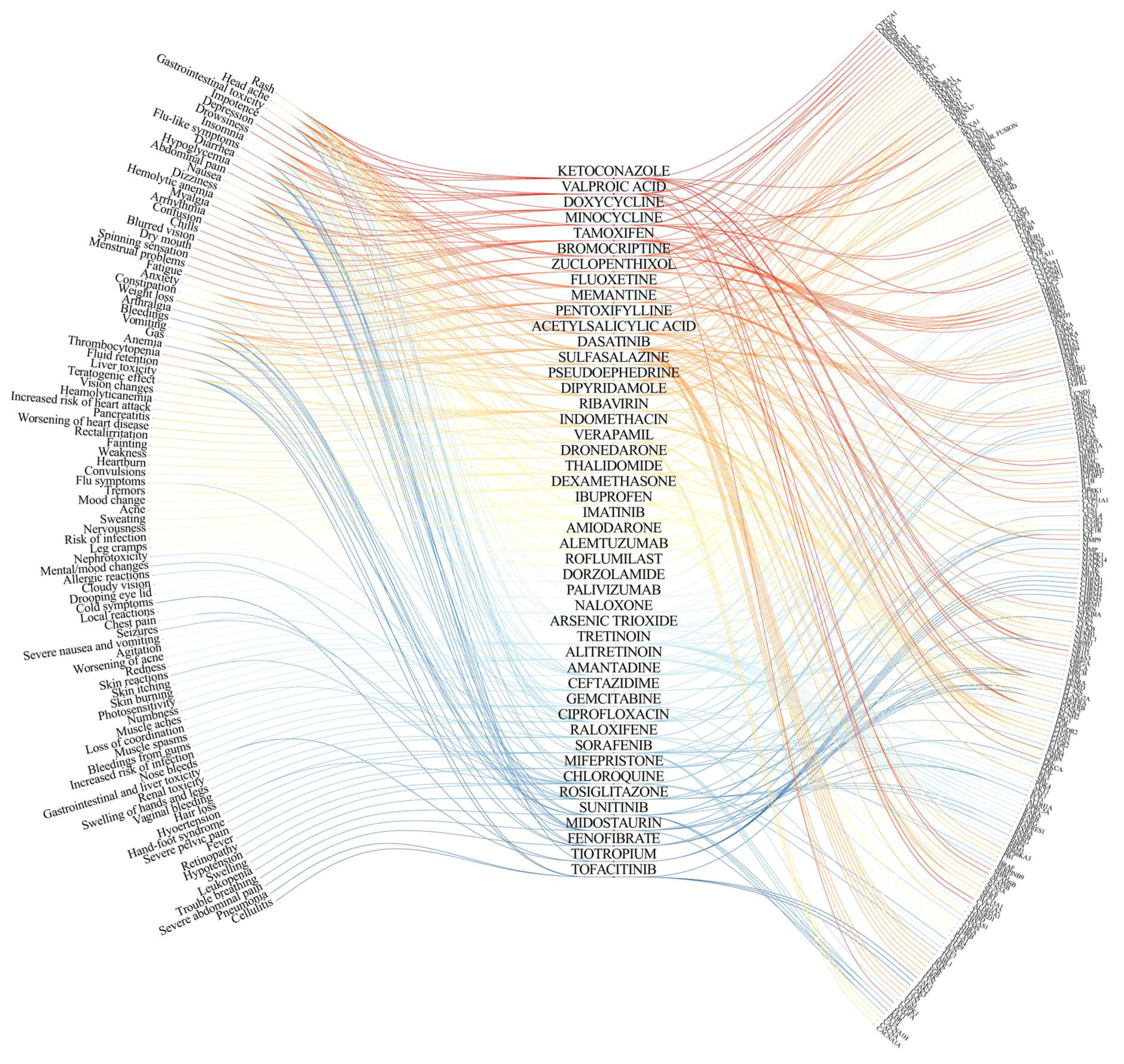
2. On- and Off-Target Side Effects of Antiviral Agents
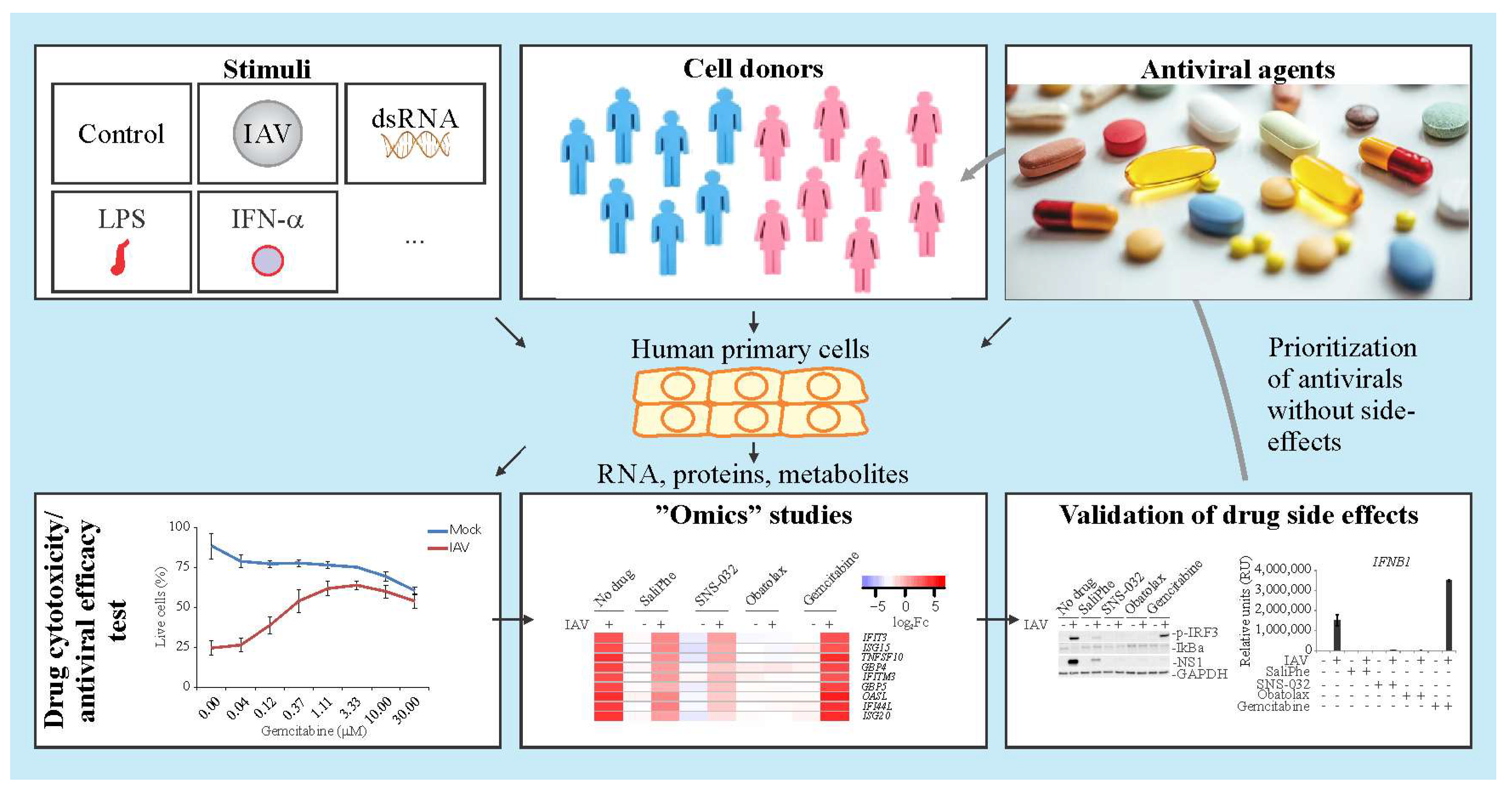
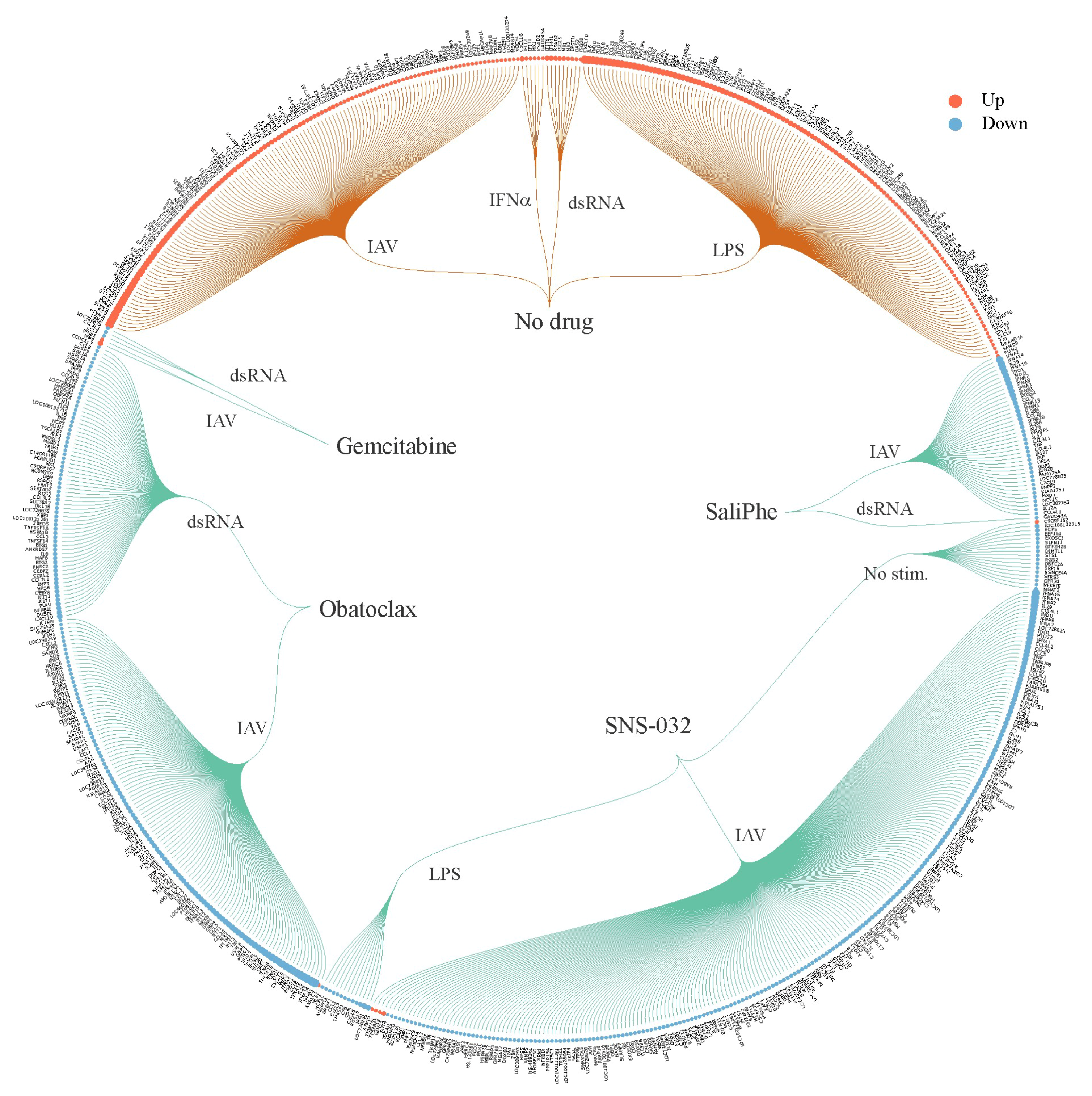
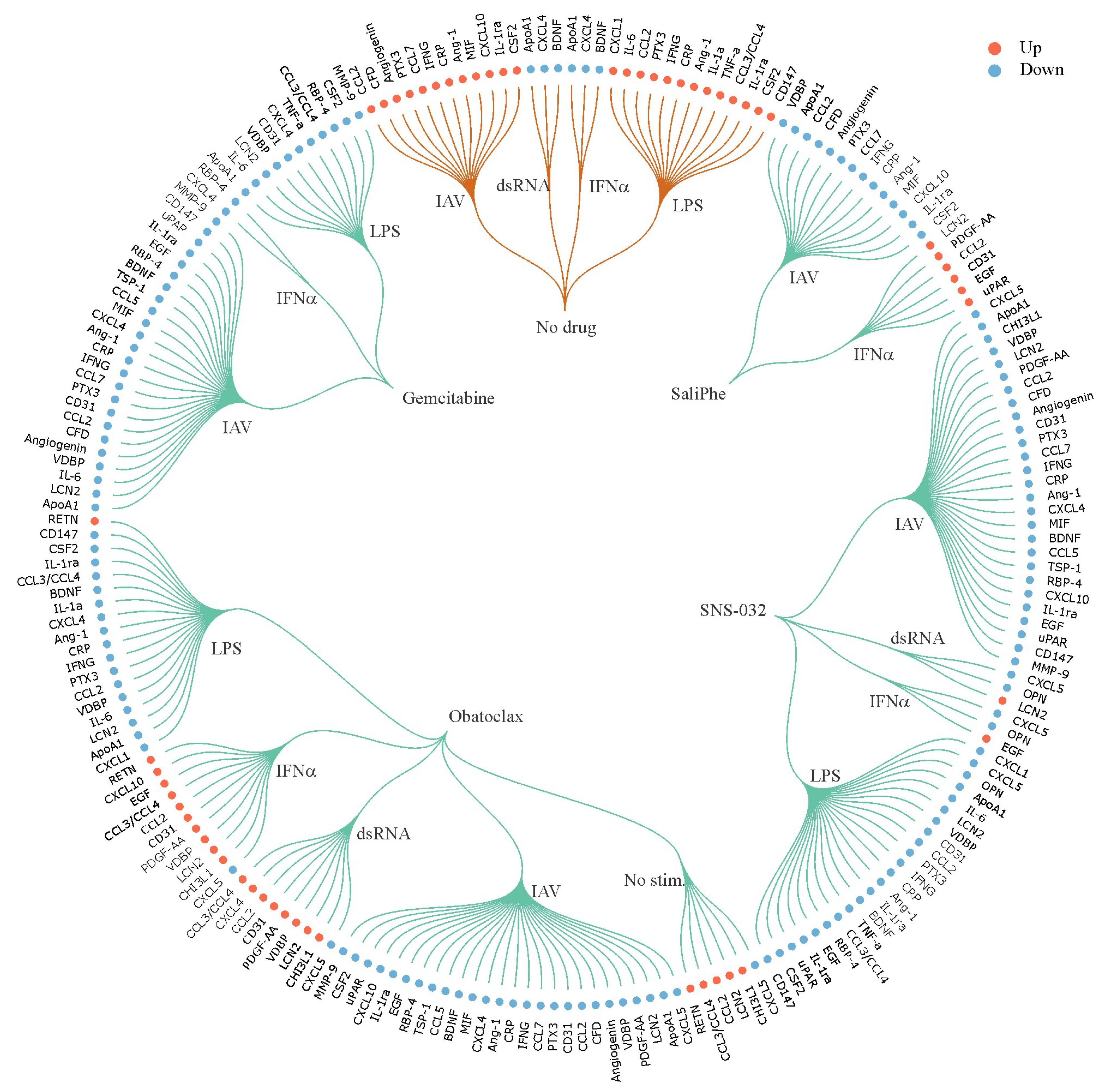
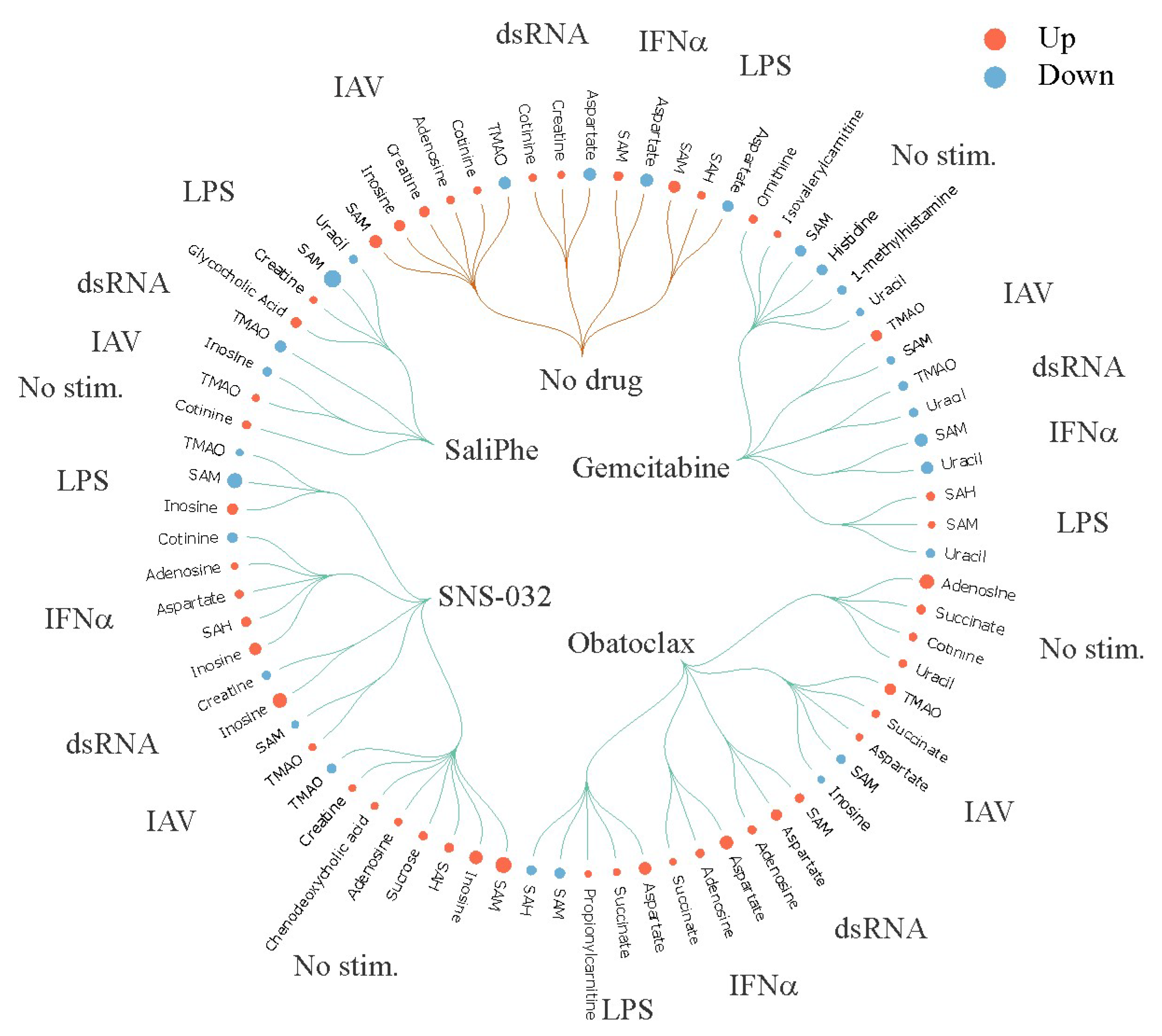
3. Systems Biology Approach to Study Immuno- and Neuro-Modulatory Properties of Antiviral Agents
4. Advantages and Disadvantages of the Approach
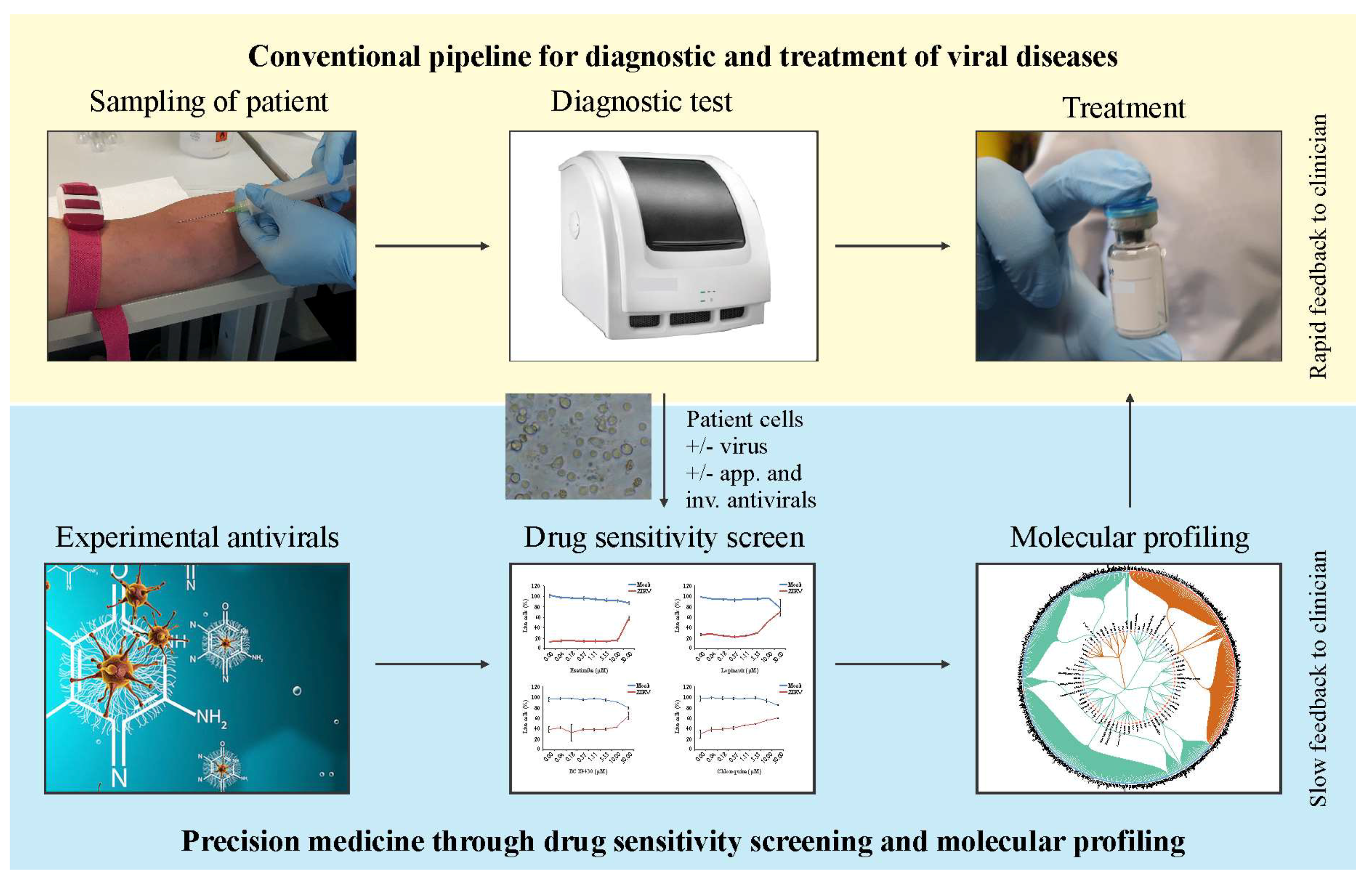
5. Future Perspectives: Personalized Treatment of Viral Diseases
6. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
References
- Ianevski, A.; Zusinaite, E.; Kuivanen, S.; Strand, M.; Lysvand, H.; Teppor, M.; Kakkola, L.; Paavilainen, H.; Laajala, M.; Kallio-Kokko, H.; et al. Novel activities of safe-in-human broad-spectrum antiviral agents. Antivir. Res. 2018, 154, 174–182. [Google Scholar] [CrossRef] [PubMed]
- De Clercq, E.; Li, G. Approved Antiviral Drugs over the Past 50 Years. Clin. Microbiol. Rev. 2016, 29, 695–747. [Google Scholar] [CrossRef] [PubMed]
- Smith, L.E.; D’Antoni, D.; Jain, V.; Pearce, J.M.; Weinman, J.; Rubin, G.J. A systematic review of factors affecting intended and actual adherence with antiviral medication as treatment or prophylaxis in seasonal and pandemic flu. Influ. Other Respir. Viruses 2016, 10, 462–478. [Google Scholar] [CrossRef] [PubMed]
- Esposito, I.; Labarga, P.; Barreiro, P.; Fernandez-Montero, J.V.; de Mendoza, C.; Benitez-Gutierrez, L.; Pena, J.M.; Soriano, V. Dual antiviral therapy for HIV and hepatitis C—Drug interactions and side effects. Expert Opin. Drug Saf. 2015, 14, 1421–1434. [Google Scholar] [CrossRef] [PubMed]
- Reust, C.E. Common adverse effects of antiretroviral therapy for HIV disease. Am. Fam. Phys. 2011, 83, 1443–1451. [Google Scholar]
- Esposito, I.; Benitez-Gutierrez, L.; Trevino, A.; Arias, A.; Citores, M.J.; Requena, S.; Soriano, V.; Cuervas-Mons, V.; de Mendoza, C. Impact of ITPA gene polymorphisms on the risk of ribavirin-induced haemolytic anaemia using interferon-free antivirals for chronic hepatitis C. Antivir. Ther. 2017, 22, 571–575. [Google Scholar] [CrossRef] [PubMed]
- Guaraldi, G.; Stentarelli, C.; Zona, S.; Santoro, A. HIV-associated lipodystrophy: Impact of antiretroviral therapy. Drugs 2013, 73, 1431–1450. [Google Scholar] [CrossRef] [PubMed]
- Keiser, M.J.; Setola, V.; Irwin, J.J.; Laggner, C.; Abbas, A.I.; Hufeisen, S.J.; Jensen, N.H.; Kuijer, M.B.; Matos, R.C.; Tran, T.B.; et al. Predicting new molecular targets for known drugs. Nature 2009, 462, 175–181. [Google Scholar] [CrossRef] [PubMed]
- Williamson, J.C.; Pegram, P.S. Neuraminidase inhibitors in patients with underlying airways disease. Am. J. Respir. Med. 2002, 1, 85–90. [Google Scholar] [CrossRef] [PubMed]
- Onakpoya, I.J.; Heneghan, C.J.; Aronson, J.K. Post-marketing withdrawal of 462 medicinal products because of adverse drug reactions: A systematic review of the world literature. BMC Med. 2016, 14, 10. [Google Scholar] [CrossRef] [PubMed]
- Siramshetty, V.B.; Nickel, J.; Omieczynski, C.; Gohlke, B.O.; Drwal, M.N.; Preissner, R. WITHDRAWN—A resource for withdrawn and discontinued drugs. Nucleic Acids Res. 2016, 44, D1080–D1086. [Google Scholar] [CrossRef] [PubMed]
- Belanov, S.S.; Bychkov, D.; Benner, C.; Ripatti, S.; Ojala, T.; Kankainen, M.; Kai Lee, H.; Wei-Tze Tang, J.; Kainov, D.E. Genome-Wide Analysis of Evolutionary Markers of Human Influenza A(H1N1)pdm09 and A(H3N2) Viruses May Guide Selection of Vaccine Strain Candidates. Genome Biol. Evol. 2015, 7, 3472–3483. [Google Scholar] [CrossRef] [PubMed]
- Kaefer, V.; Semedo, J.G.; Silva Kahl, V.F.; Von Borowsky, R.G.; Gianesini, J.; Ledur Kist, T.B.; Pereira, P.; Picada, J.N. DNA damage in brain cells and behavioral deficits in mice after treatment with high doses of amantadine. J. Appl. Toxicol. 2010, 30, 745–753. [Google Scholar] [CrossRef] [PubMed]
- Benschop, K.S.; Wildenbeest, J.G.; Koen, G.; Minnaar, R.P.; van Hemert, F.J.; Westerhuis, B.M.; Pajkrt, D.; van den Broek, P.J.; Vossen, A.C.; Wolthers, K.C. Genetic and antigenic structural characterization for resistance of echovirus 11 to pleconaril in an immunocompromised patient. J. Gen. Virol. 2015, 96, 571–579. [Google Scholar] [CrossRef] [PubMed]
- Mercorelli, B.; Palu, G.; Loregian, A. Drug Repurposing for Viral Infectious Diseases: How Far Are We? Trends Microbiol. 2018. [Google Scholar] [CrossRef] [PubMed]
- Berger, S.I.; Iyengar, R. Role of systems pharmacology in understanding drug adverse events. Wiley Interdiscip. Rev. Syst. Biol. Med. 2011, 3, 129–135. [Google Scholar] [CrossRef] [PubMed]
- Gallay, P.A.; Lin, K. Profile of alisporivir and its potential in the treatment of hepatitis C. Drug Des. Dev. Ther. 2013, 7, 105–115. [Google Scholar] [CrossRef] [PubMed]
- Sharma, V.D.; Lyons, K.E.; Pahwa, R. Amantadine extended-release capsules for levodopa-induced dyskinesia in patients with Parkinson’s disease. Ther. Clin. Risk Manag. 2018, 14, 665–673. [Google Scholar] [CrossRef] [PubMed]
- Schor, S.; Einav, S. Repurposing of Kinase Inhibitors as Broad-Spectrum Antiviral Drugs. DNA Cell Biol. 2018, 37, 63–69. [Google Scholar] [CrossRef] [PubMed]
- Soderholm, S.; Anastasina, M.; Islam, M.M.; Tynell, J.; Poranen, M.M.; Bamford, D.H.; Stenman, J.; Julkunen, I.; Sauliene, I.; De Brabander, J.K.; et al. Immuno-modulating properties of saliphenylhalamide, SNS-032, obatoclax, and gemcitabine. Antivir. Res. 2016, 126, 69–80. [Google Scholar] [CrossRef] [PubMed]
- Soderholm, S.; Kainov, D.E.; Ohman, T.; Denisova, O.V.; Schepens, B.; Kulesskiy, E.; Imanishi, S.Y.; Corthals, G.; Hintsanen, P.; Aittokallio, T.; et al. Phosphoproteomics to Characterize Host Response During Influenza A Virus Infection of Human Macrophages. Mol. Cell. Proteom. 2016, 15, 3203–3219. [Google Scholar] [CrossRef] [PubMed]
- Kuivanen, S.; Bespalov, M.M.; Nandania, J.; Ianevski, A.; Velagapudi, V.; De Brabander, J.K.; Kainov, D.E.; Vapalahti, O. Obatoclax, saliphenylhalamide and gemcitabine inhibit Zika virus infection in vitro and differentially affect cellular signaling, transcription and metabolism. Antivir. Res. 2017, 139, 117–128. [Google Scholar] [CrossRef] [PubMed]
- Varghese, F.S.; Rausalu, K.; Hakanen, M.; Saul, S.; Kummerer, B.M.; Susi, P.; Merits, A.; Ahola, T. Obatoclax Inhibits Alphavirus Membrane Fusion by Neutralizing the Acidic Environment of Endocytic Compartments. Antimicrob. Agents Chemother. 2017, 61. [Google Scholar] [CrossRef] [PubMed]
- Adcock, R.S.; Chu, Y.K.; Golden, J.E.; Chung, D.H. Evaluation of anti-Zika virus activities of broad-spectrum antivirals and NIH clinical collection compounds using a cell-based, high-throughput screen assay. Antivir. Res. 2017, 138, 47–56. [Google Scholar] [CrossRef] [PubMed]
- Muller, K.H.; Spoden, G.A.; Scheffer, K.D.; Brunnhofer, R.; De Brabander, J.K.; Maier, M.E.; Florin, L.; Muller, C.P. Inhibition by cellular vacuolar ATPase impairs human papillomavirus uncoating and infection. Antimicrob. Agents Chemother. 2014, 58, 2905–2911. [Google Scholar] [CrossRef] [PubMed]
- Denisova, O.V.; Kakkola, L.; Feng, L.; Stenman, J.; Nagaraj, A.; Lampe, J.; Yadav, B.; Aittokallio, T.; Kaukinen, P.; Ahola, T.; et al. Obatoclax, saliphenylhalamide, and gemcitabine inhibit influenza a virus infection. J. Biol. Chem. 2012, 287, 35324–35332. [Google Scholar] [CrossRef] [PubMed]
- Morales, F.; Giordano, A. Overview of CDK9 as a target in cancer research. Cell Cycl. 2016, 15, 519–527. [Google Scholar] [CrossRef] [PubMed]
- Lebreton, S.; Jaunbergs, J.; Roth, M.G.; Ferguson, D.A.; De Brabander, J.K. Evaluating the potential of vacuolar ATPase inhibitors as anticancer agents and multigram synthesis of the potent salicylihalamide analog saliphenylhalamide. Bioorg. Med. Chem. Lett. 2008, 18, 5879–5883. [Google Scholar] [CrossRef] [PubMed]
- Lee, M.N.; Ye, C.; Villani, A.C.; Raj, T.; Li, W.; Eisenhaure, T.M.; Imboywa, S.H.; Chipendo, P.I.; Ran, F.A.; Slowikowski, K.; et al. Common genetic variants modulate pathogen-sensing responses in human dendritic cells. Science 2014, 343, 1246980. [Google Scholar] [CrossRef] [PubMed]
- Baba, T.; Mukaida, N. Role of macrophage inflammatory protein (MIP)-1alpha/CCL3 in leukemogenesis. Mol. Cell. Oncol. 2014, 1, e29899. [Google Scholar] [CrossRef] [PubMed]
- Robertson, M.J. Role of chemokines in the biology of natural killer cells. J. Leukoc. Biol. 2002, 71, 173–183. [Google Scholar] [PubMed]
- Koper, O.M.; Kaminska, J.; Sawicki, K.; Kemona, H. CXCL9, CXCL10, CXCL11, and their receptor (CXCR3) in neuroinflammation and neurodegeneration. Adv. Clin. Exp. Med. 2018, 27, 849–856. [Google Scholar] [CrossRef] [PubMed]
- Ferrari, S.M.; Ruffilli, I.; Colaci, M.; Antonelli, A.; Ferri, C.; Fallahi, P. CXCL10 in psoriasis. Adv. Med. Sci. 2015, 60, 349–354. [Google Scholar] [CrossRef] [PubMed]
- Antonelli, A.; Ferrari, S.M.; Giuggioli, D.; Ferrannini, E.; Ferri, C.; Fallahi, P. Chemokine (C-X-C motif) ligand (CXCL)10 in autoimmune diseases. Autoimmun. Rev. 2014, 13, 272–280. [Google Scholar] [CrossRef] [PubMed]
- Gaelings, L.; Soderholm, S.; Bugai, A.; Fu, Y.; Nandania, J.; Schepens, B.; Lorey, M.B.; Tynell, J.; Vande Ginste, L.; Le Goffic, R.; et al. Regulation of kynurenine biosynthesis during influenza virus infection. FEBS J. 2017, 284, 222–236. [Google Scholar] [CrossRef] [PubMed]
- Duan, Z.; Duan, Y.; Lei, H.; Hu, N.; Shi, J.; Shen, D.; Wang, X.; Hu, Y. Attenuation of antigenic immunogenicity by kynurenine, a novel suppressive adjuvant. Hum. Vaccines Immunother. 2014, 10, 1295–1305. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.C.; Holtze, M.; Powell, S.B.; Terrando, N.; Larsson, M.K.; Persson, A.; Olsson, S.K.; Orhan, F.; Kegel, M.; Asp, L.; et al. Behavioral disturbances in adult mice following neonatal virus infection or kynurenine treatment--role of brain kynurenic acid. Brain Behav. Immun. 2014, 36, 80–89. [Google Scholar] [CrossRef] [PubMed]
- Lucido, M.J.; Orlando, B.J.; Vecchio, A.J.; Malkowski, M.G. Crystal Structure of Aspirin-Acetylated Human Cyclooxygenase-2: Insight into the Formation of Products with Reversed Stereochemistry. Biochemistry 2016, 55, 1226–1238. [Google Scholar] [CrossRef] [PubMed]
- Antonioli, L.; Csoka, B.; Fornai, M.; Colucci, R.; Kokai, E.; Blandizzi, C.; Hasko, G. Adenosine and inflammation: What’s new on the horizon? Drug Discov. Today. 2014, 19, 1051–1068. [Google Scholar] [CrossRef] [PubMed]
- Benarroch, E.E. Adenosine and its receptors: Multiple modulatory functions and potential therapeutic targets for neurologic disease. Neurology 2008, 70, 231–236. [Google Scholar] [CrossRef] [PubMed]
- Tang, W.H.; Wang, Z.; Levison, B.S.; Koeth, R.A.; Britt, E.B.; Fu, X.; Wu, Y.; Hazen, S.L. Intestinal microbial metabolism of phosphatidylcholine and cardiovascular risk. N. Engl. J. Med. 2013, 368, 1575–1584. [Google Scholar] [CrossRef] [PubMed]
- Fu, Y.; Gaelings, L.; Soderholm, S.; Belanov, S.; Nandania, J.; Nyman, T.A.; Matikainen, S.; Anders, S.; Velagapudi, V.; Kainov, D.E. JNJ872 inhibits influenza A virus replication without altering cellular antiviral responses. Antivir. Res. 2016, 133, 23–31. [Google Scholar] [CrossRef] [PubMed]
- Denisova, O.V.; Soderholm, S.; Virtanen, S.; Von Schantz, C.; Bychkov, D.; Vashchinkina, E.; Desloovere, J.; Tynell, J.; Ikonen, N.; Theisen, L.L.; et al. Akt inhibitor MK2206 prevents influenza pH1N1 virus infection in vitro. Antimicrob. Agents Chemother. 2014, 58, 3689–3696. [Google Scholar] [CrossRef] [PubMed]
- Bulanova, D.; Ianevski, A.; Bugai, A.; Akimov, Y.; Kuivanen, S.; Paavilainen, H.; Kakkola, L.; Nandania, J.; Turunen, L.; Ohman, T.; et al. Antiviral Properties of Chemical Inhibitors of Cellular Anti-Apoptotic Bcl-2 Proteins. Viruses 2017, 9, 271. [Google Scholar] [CrossRef] [PubMed]
- Shim, J.M.; Kim, J.; Tenson, T.; Min, J.Y.; Kainov, D.E. Influenza Virus Infection, Interferon Response, Viral Counter-Response, and Apoptosis. Viruses 2017, 9, 223. [Google Scholar] [CrossRef] [PubMed]
- Kakkola, L.; Denisova, O.V.; Tynell, J.; Viiliainen, J.; Ysenbaert, T.; Matos, R.C.; Nagaraj, A.; Ohman, T.; Kuivanen, S.; Paavilainen, H.; et al. Anticancer compound ABT-263 accelerates apoptosis in virus-infected cells and imbalances cytokine production and lowers survival rates of infected mice. Cell Death Dis. 2013, 4, e742. [Google Scholar] [CrossRef] [PubMed]
- Jurado, K.A.; Iwasaki, A. Zika virus targets blood monocytes. Nat. Microbiol. 2017, 2, 1460–1461. [Google Scholar] [CrossRef] [PubMed]
- Marino-Merlo, F.; Papaianni, E.; Medici, M.A.; Macchi, B.; Grelli, S.; Mosca, C.; Borner, C.; Mastino, A. HSV-1-induced activation of NF-kappaB protects U937 monocytic cells against both virus replication and apoptosis. Cell Death Dis. 2016, 7, e2354. [Google Scholar] [CrossRef] [PubMed]
- Kumar, A.; Abbas, W.; Herbein, G. HIV-1 latency in monocytes/macrophages. Viruses 2014, 6, 1837–1860. [Google Scholar] [CrossRef] [PubMed]
- Her, Z.; Malleret, B.; Chan, M.; Ong, E.K.; Wong, S.C.; Kwek, D.J.; Tolou, H.; Lin, R.T.; Tambyah, P.A.; Renia, L.; et al. Active infection of human blood monocytes by Chikungunya virus triggers an innate immune response. J. Immunol. 2010, 184, 5903–5913. [Google Scholar] [CrossRef] [PubMed]
- Wishart, D.S.; Feunang, Y.D.; Guo, A.C.; Lo, E.J.; Marcu, A.; Grant, J.R.; Sajed, T.; Johnson, D.; Li, C.; Sayeeda, Z.; et al. DrugBank 5.0: A major update to the DrugBank database for 2018. Nucleic Acids Res. 2018, 46, D1074–D1082. [Google Scholar]
- Beger, R.D.; Dunn, W.; Schmidt, M.A.; Gross, S.S.; Kirwan, J.A.; Cascante, M.; Brennan, L.; Wishart, D.S.; Oresic, M.; Hankemeier, T.; et al. Metabolomics enables precision medicine: “A White Paper, Community Perspective”. Metabolomics 2016, 12, 149. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Wang, J.; Ning, C.; Zheng, X.; Fu, J.; Wang, A.; Zhang, Q.; Liu, J.F. Genome-wide DNA methylation and transcriptome analyses reveal genes involved in immune responses of pig peripheral blood mononuclear cells to poly I:C. Sci. Rep. 2017, 7, 9709. [Google Scholar] [CrossRef] [PubMed]
- Yan, H.; Tian, S.; Slager, S.L.; Sun, Z.; Ordog, T. Genome-Wide Epigenetic Studies in Human Disease: A Primer on-Omic Technologies. Am. J. Epidemiol. 2016, 183, 96–109. [Google Scholar] [CrossRef] [PubMed]
- Yip, V.L.; Alfirevic, A.; Pirmohamed, M. Genetics of immune-mediated adverse drug reactions: A comprehensive and clinical review. Clin. Rev. Allergy Immunol. 2015, 48, 165–175. [Google Scholar] [CrossRef] [PubMed]
- Fairfax, B.P.; Humburg, P.; Makino, S.; Naranbhai, V.; Wong, D.; Lau, E.; Jostins, L.; Plant, K.; Andrews, R.; McGee, C.; et al. Innate immune activity conditions the effect of regulatory variants upon monocyte gene expression. Science 2014, 343, 1246949. [Google Scholar] [CrossRef] [PubMed]
- Rotival, M.; Zeller, T.; Wild, P.S.; Maouche, S.; Szymczak, S.; Schillert, A.; Castagne, R.; Deiseroth, A.; Proust, C.; Brocheton, J.; et al. Integrating genome-wide genetic variations and monocyte expression data reveals trans-regulated gene modules in humans. PLoS Genet. 2011, 7, e1002367. [Google Scholar] [CrossRef] [PubMed]
- Ginsburg, G.S.; McCarthy, J.J. Personalized medicine: Revolutionizing drug discovery and patient care. Trends Biotechnol. 2001, 19, 491–496. [Google Scholar] [CrossRef]
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zusinaite, E.; Ianevski, A.; Niukkanen, D.; Poranen, M.M.; Bjørås, M.; Afset, J.E.; Tenson, T.; Velagapudi, V.; Merits, A.; Kainov, D.E. A Systems Approach to Study Immuno- and Neuro-Modulatory Properties of Antiviral Agents. Viruses 2018, 10, 423. https://doi.org/10.3390/v10080423
Zusinaite E, Ianevski A, Niukkanen D, Poranen MM, Bjørås M, Afset JE, Tenson T, Velagapudi V, Merits A, Kainov DE. A Systems Approach to Study Immuno- and Neuro-Modulatory Properties of Antiviral Agents. Viruses. 2018; 10(8):423. https://doi.org/10.3390/v10080423
Chicago/Turabian StyleZusinaite, Eva, Aleksandr Ianevski, Diana Niukkanen, Minna M. Poranen, Magnar Bjørås, Jan Egil Afset, Tanel Tenson, Vidya Velagapudi, Andres Merits, and Denis E. Kainov. 2018. "A Systems Approach to Study Immuno- and Neuro-Modulatory Properties of Antiviral Agents" Viruses 10, no. 8: 423. https://doi.org/10.3390/v10080423
APA StyleZusinaite, E., Ianevski, A., Niukkanen, D., Poranen, M. M., Bjørås, M., Afset, J. E., Tenson, T., Velagapudi, V., Merits, A., & Kainov, D. E. (2018). A Systems Approach to Study Immuno- and Neuro-Modulatory Properties of Antiviral Agents. Viruses, 10(8), 423. https://doi.org/10.3390/v10080423





