Abstract
Cancer cachexia is a multifactorial metabolic syndrome that profoundly impairs treatment tolerance and prognosis; however, how tumor-intrinsic transcriptional programs contribute to cachexia induction and shape responses to chemotherapeutic stress remains poorly understood. In this study, we analyzed a paired human duodenal neuroendocrine carcinoma cell-line model consisting of the non-cachexia-inducing parental line TCC-NECT-2 and its cachexia-inducing derivative AkuNEC. Bulk RNA sequencing was performed under baseline conditions and after doxorubicin treatment (10 μM, 24 h), and transcriptomic differences were assessed using log2 fold-change–based analyses to characterize baseline reprogramming, drug-induced responses, and differential stress adaptation. Despite comparable morphology and similar levels of doxorubicin-induced acute cytotoxicity, AkuNEC cells exhibited extensive baseline transcriptional reprogramming relative to TCC-NECT-2, including coordinated upregulation of inflammatory, secretory, and metabolic regulators previously implicated in cancer cachexia, together with suppression of structural and homeostatic programs. Following doxorubicin exposure, AkuNEC cells showed a distinct transcriptional response characterized by selective reorganization of proliferation-, metabolism-, and stress-related pathways, indicating enhanced transcriptional plasticity rather than uniform stress suppression. Differential response analyses further revealed preferential induction of genes involved in cell cycle control, DNA replication, and metabolic adaptation in AkuNEC under chemotherapeutic stress. These findings indicate that cachexia-inducing capacity is embedded within tumor-intrinsic transcriptional states and is amplified by stress-induced plasticity, supporting a network-level model of cancer cachexia that links systemic host effects with tumor adaptation to therapy. The TCC-NECT-2/AkuNEC model provides a tractable framework for dissecting these tumor-intrinsic mechanisms and their relevance to cachexia-associated cancer biology.
1. Introduction
Cancer cachexia is a progressive metabolic syndrome characterized by involuntary weight loss, skeletal muscle wasting, and reduced appetite, and these symptoms cannot be fully reversed by nutritional support [1,2]. Cachexia has a substantial impact on physical function, tolerance to anticancer treatment, and overall survival, yet effective therapeutic options remain limited [2]. Because current diagnostic criteria rely mainly on visible and late-stage manifestations, many patients are diagnosed only after systemic homeostasis has already been markedly disrupted [1]. Recent multi-organ studies suggest that tumor–host metabolic and inflammatory interactions begin long before overt wasting becomes apparent [3,4], indicating that cachexia represents a gradual and dynamic pathological process. However, the molecular mechanisms that drive these early changes are still incompletely understood.
There is increasing evidence that cachexia may also influence the responsiveness of tumors to chemotherapy. A recent meta-analysis reported that cachexia is associated with reduced treatment efficacy and poorer survival across several malignancies [5]. Altered body composition, including decreased muscle mass and disrupted lipid metabolism, can affect drug pharmacokinetics, increase toxicity, and lead to dose reductions or early discontinuation [6]. Experimental studies further show that tumor-derived factors such as interleukin-6 (IL-6), PTH-related protein (PTHrP), myostatin/activin, and ATGL-mediated lipolysis can promote adipose tissue browning, muscle catabolism, and systemic metabolic disturbance [7,8,9,10,11]. These findings support the hypothesis that tumors capable of inducing cachexia may also possess distinct drug-response programs. Yet, tumor-intrinsic mechanisms that connect cachexia-inducing ability with altered transcriptional responses to therapeutic stress remain largely unexplored.
To address this gap, suitable experimental models are essential [12]. Human cancer cell lines that reproducibly induce cachexia in vivo are rare, and commonly used xenograft models often show variable or late-onset wasting [12]. The human cell-line pair TCC-NECT-2 [13] and AkuNEC provides a unique opportunity to investigate tumor-intrinsic differences underlying cachexia induction [13,14]. TCC-NECT-2, derived from a duodenal neuroendocrine carcinoma, does not induce cachexia when transplanted into mice, whereas its in vivo–passaged derivative AkuNEC reproducibly causes systemic cachexia. Because these two lines share a common origin, they enable controlled comparison of cachexia-inducing and non-inducing tumor states without confounding genomic background. Such paired models remain uncommon but offer a tractable system for mechanistic studies.
Although interest in tumor–host metabolic crosstalk has grown, relatively few studies have examined tumor-intrinsic transcriptional programs linked to cachexia. Most previous research has focused on host tissues, including skeletal muscle, adipose tissue, and liver [10,11], whereas far less is known about gene-expression programs within cachexia-inducing tumors themselves, either at baseline or under therapeutic stress. This remains an important gap in the field. In this study, we performed RNA sequencing of TCC-NECT-2 and AkuNEC cells before and after exposure to doxorubicin to identify (i) baseline transcriptional differences associated with cachexia-inducing capacity and (ii) differential chemotherapy-induced responses. Through this paired-cell-line approach, we aim to define tumor-intrinsic transcriptional signatures that may contribute to cachexia induction and potentially influence chemotherapeutic sensitivity. Based on prior experimental and clinical studies, a literature-curated reference set of tumor-derived genes previously implicated in cancer cachexia was compiled and used to guide downstream transcriptomic analyses (Supplementary Table S1).
2. Materials and Methods
2.1. Cell Lines
Two human cell lines derived from a duodenal neuroendocrine carcinoma were examined: the parental line TCC-NECT-2 [13] and its cachexia-inducing subline AkuNEC [14]. The procedures for establishing these cell lines and their initial characterization have been described previously. Cells were routinely maintained in RPMI-1640 medium (Gibco, Thermo Fisher Scientific, Waltham, MA, USA) supplemented with 10% heat-inactivated fetal bovine serum (FBS), 100 IU/mL penicillin, and 100 µg/mL streptomycin. All cultures were kept at 37 °C in a humidified incubator with 5% CO2. For 2D culture, cell suspensions were seeded onto 10 cm tissue culture dishes (Nunc Easydish; Thermo Fisher Scientific Inc.) and maintained at 37 °C in a humidified incubator with 5% CO2. For 3D culture, the cells were kept in round-bottom 96-well ultra-low attachment plates (Thermo Scientific™ 96 Well Round (U) Bottom Plate, TC Surface; Thermo Fisher Scientific Inc., Waltham, MA, USA). The cells were seeded at 5000 cells per well in the ultra-low attachment 96-well plates. 3D culture in ultra-low attachment plates was selected for all drug treatments and transcriptomic assays to provide a model of tumor architecture.
2.2. Treatments with Doxorubicin
The cells were seeded at 5000 cells per well in the ultra-low attachment 96-well plates. After 24 h of incubation to allow for spheroid stabilization, doxorubicin (doxorubicin hydrochloride, Selleck Chemicals LLC, Houston, TX, USA) was added to each well at a final concentration of 10 µM and the cells were exposed for an additional 24 h. Doxorubicin was selected as a prototypical genotoxic stressor to induce acute DNA damage and capture the resulting transcriptional plasticity, rather than to simulate a specific clinical regimen for neuroendocrine carcinoma. Cell viability following drug treatment was assessed using the Cell Counting Kit-8 (Dojindo Laboratories, Kumamoto, Japan). After adding the reagent to each well, plates were incubated at 37 °C for the recommended period, and absorbance at 450 nm was measured using a Multiskan SkyHigh microplate reader (Thermo Fisher Scientific, Waltham, MA, USA). As a vehicle control, to represent 100% cell viability, the same final concentration of DMSO used to prepare doxorubicin (10 µM) was added to the culture medium. We confirmed in advance that this concentration of DMSO did not affect cell proliferation. The absorbance of the vehicle DMSO control was defined as 100%, and relative cell viability (%) was calculated for the doxorubicin-treated samples. All experiments were performed in duplicate, and mean values were reported. All experiments were performed in duplicate (n = 2 biological replicates, each with technical duplicates).
2.3. Scanning Electron Microscopy (SEM)
Scanning electron microscopy was carried out following a previously described procedure [15], with several minor adjustments to suit our experimental conditions. The spheroids were fixed at 4 °C overnight in 2.5% glutaraldehyde prepared in 0.1 M phosphate buffer (pH 7.4). After fixation, the spheroids were gently rinsed with phosphate-buffered saline (PBS) and subsequently post-fixed in 1% osmium tetroxide (OsO4) for 30 min at room temperature. The samples were then dehydrated through a graded ethanol series, transferred to 100% ethanol, and allowed to dry in air. Once fully dried, the spheroids were coated with a thin platinum film using an MSP-1S sputter coater (Shinku Device, Ibaraki, Japan). SEM imaging was performed using a Phenom Pro desktop scanning electron microscope (Thermo Fisher Scientific, Waltham, MA, USA), and representative images were obtained to document the surface morphology of the spheroids.
2.4. Transcriptomic Study
2.4.1. RNA Extraction and Sample Preparation
The cells were washed twice with phosphate-buffered saline (PBS) directly in the plates. Cells were then resuspended in PBS per well, followed by the QIAzol Lysis Reagent (QIAGEN K.K., Tokyo, Japan) for cell lysis. Cell lysates were collected and shipped to Zeon BioSolutions Inc. (Hyogo, Japan), where total RNA extraction and purification were performed using TRIzol reagent according to the manufacturer’s protocol.
2.4.2. RNA Quality Assessment
RNA purity was assessed by spectrophotometric measurement of the A260/280 ratio, and RNA integrity was evaluated using an Agilent Bioanalyzer or TapeStation system (Agilent Technologies, Santa Clara, CA, USA). Only RNA samples with an RNA integrity number (RIN) greater than 8.0 were used for downstream analyses. RNA concentration and RNA integrity were applied as the primary quality-control criteria for RNA sequencing. For library preparation, 20 ng of total RNA per sample was used as starting material.
2.4.3. RNA-Seq Library Preparation and Sequencing
RNA-seq libraries were prepared using the Quartz-Seq2 protocol. Neither ribosomal RNA depletion nor poly(A) selection was performed. Libraries were multiplexed and sequenced on an Illumina NextSeq platform. Quartz-Seq2 is designed for low-input transcriptome profiling and has been shown to provide robust relative expression quantification at modest sequencing depth. Quality control of raw sequencing data was performed using FastQC to evaluate read quality, base composition, and potential adapter contamination prior to downstream bioinformatic analyses.
2.5. Statistical and Bioinformatic Analyses
Gene expression analyses were primarily based on effect-size–driven comparisons in this exploratory pilot study. Due to the small sample size (n = 2 biological replicates per condition), formal statistical significance testing or multiple-testing correction was not applied to the RNA-seq data. RNA-seq expression values were log2-transformed prior to downstream analyses. For pairwise comparisons between experimental conditions, log2 fold-change (log2FC) values were calculated using the mean expression of two biological replicates per condition. The log2-normalized gene expression matrix for all genes and all experimental conditions is provided in Supplementary Table S2.
Baseline transcriptional differences between TCC-NECT-2 and AkuNEC were evaluated under no-treatment conditions. Doxorubicin-induced transcriptional responses were assessed by comparing doxorubicin-treated samples with DMSO-treated controls within each cell line. Differential drug response (Δresponse) was defined as the difference in log2FC between AkuNEC and TCC-NECT-2, calculated as (AkuNEC_Dox − AkuNEC_DMSO) − (TCC-NECT-2_Dox−TCC-NECT-2_DMSO).
Because of the limited number of biological replicates (n = 2 per condition), no formal statistical significance testing or multiple-testing correction was applied. Instead, transcriptomic differences were interpreted based on the magnitude and consistency of log2FC values across comparisons. This approach was chosen to identify robust and reproducible transcriptional patterns while avoiding overinterpretation of statistical significance.
Pathway enrichment analyses were performed using the Enrichr platform implemented via the gseapy Python package. Gene ontology (GO) biological process databases were used to assess functional enrichment among genes upregulated and downregulated in AkuNEC relative to TCC-NECT-2. Enrichment results were ranked by combined score, and representative pathways were selected for visualization. Enrichment results were ranked by “Combined Score”, which integrates the p-value from Fisher’s exact test with the z-score of the deviation from the expected rank, providing a prioritized list of biological processes even in limited datasets. All enrichment analyses were conducted without statistical filtering to reflect effect-size–driven differences.
All computational analyses and data visualizations were performed using Python 3.14 with the pandas, numpy, matplotlib, seaborn, and gseapy libraries. Figures were generated using log2-transformed expression values and exported at publication-quality resolution.
3. Results
3.1. Morphological Features and Doxorubicin Sensitivity of TCC-NECT-2 and AkuNEC
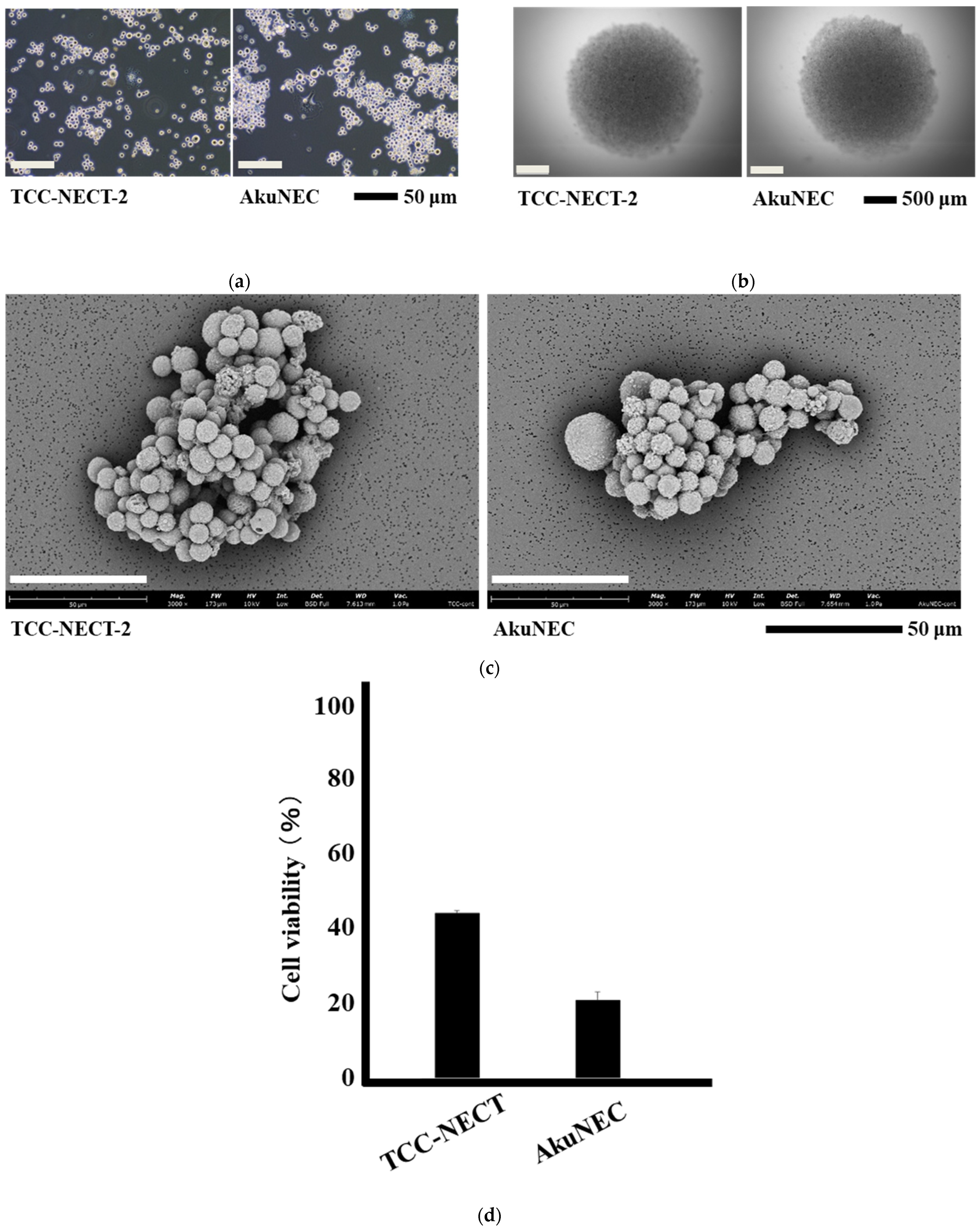
To establish a phenotypic baseline for subsequent transcriptomic analyses, we first compared the morphological characteristics and doxorubicin sensitivity of the non–cachexia-inducing parental cell line TCC-NECT-2 and its cachexia-inducing derivative AkuNEC. Under standard culture conditions, phase-contrast microscopy revealed no overt morphological differences between the two cell lines (Figure 1a,b). Both TCC-NECT-2 and AkuNEC exhibited similar cell shape and growth patterns at the gross morphological level.
Figure 1.
Morphological features and doxorubicin sensitivity of TCC-NECT-2 and AkuNEC cells. (a,b) Phase-contrast microscopy images of TCC-NECT-2 and AkuNEC cells, showing no overt morphological differences under standard culture conditions. (c) Scanning electron microscopy images illustrating subtle differences in cell surface architecture between the two cell lines. Scale bars, 50 μm. (d) Cell viability assessed using the Cell Counting Kit-8 (CCK-8) assay after 24 h exposure to doxorubicin (10 μM). Absorbance values were normalized to DMSO-treated controls, which were set to 100%. TCC-NECT-2 cells retained approximately 41% viability, whereas AkuNEC cells exhibited significantly lower viability (~19%), indicating enhanced sensitivity to doxorubicin in the cachexia-inducing AkuNEC line. Data are presented as mean ± SD. Statistical significance was evaluated using a two-tailed Student’s t-test.
We next examined cell surface architecture using scanning electron microscopy (SEM). Although overall cellular morphology was largely comparable, SEM imaging suggested subtle differences in surface features between TCC-NECT-2 and AkuNEC cells (Figure 1c). These differences were qualitative in nature and were not subjected to quantitative morphometric analysis, but they raised the possibility that cachexia-inducing capacity may be associated with modest alterations in cell surface organization rather than dramatic changes in cell shape. These differences were qualitative in nature and were not subjected to quantitative morphometric analysis, but they raised the possibility that cachexia-inducing capacity may be associated with modest alterations in cell surface organization.
To assess whether cachexia-inducing capacity is accompanied by altered sensitivity to chemotherapeutic stress at the phenotypic level, we evaluated acute cytotoxicity following doxorubicin treatment. Under the tested conditions, doxorubicin treatment resulted in a significantly lower viability in AkuNEC cells compared with TCC-NECT-2 (Figure 1d). Cell viability was reduced to approximately 19% in AkuNEC, whereas TCC-NECT-2 retained ~41% viability relative to DMSO-treated controls. Notably, despite this quantitative difference in drug sensitivity, no overt morphological differences were observed between the two cell lines under standard culture conditions (Figure 1a–c), suggesting that enhanced chemosensitivity in AkuNEC is not accompanied by gross phenotypic alterations.
Together, these observations demonstrate that TCC-NECT-2 and AkuNEC cells are largely indistinguishable at the level of gross morphology and short-term acute cytotoxicity by doxorubicin. These findings suggest that cachexia-inducing capacity is not readily captured by conventional phenotypic assays, thereby motivating a deeper investigation into tumor-intrinsic transcriptional programs and stress-responsive molecular differences addressed in subsequent analyses.
3.2. Literature-Based Identification of Cachexia-Associated Tumor Factors
To identify tumor-derived factors potentially involved in cachexia induction, we first conducted a systematic literature-based survey of genes previously reported to drive cancer cachexia. Using PubMed-indexed studies, we compiled a curated list of tumor-secreted molecules that have been experimentally shown to promote key features of cachexia, including muscle wasting, adipose tissue mobilization, systemic inflammation, and anorexia (Supplementary Table S1 for detailed descriptions and a tabulated summary). This approach ensured that subsequent transcriptomic analyses were anchored in established biological mechanisms rather than relying solely on unbiased differential expression.
Based on this literature survey, the identified cachexia-associated factors clustered into three major functional categories: (i) pro-inflammatory cytokines, including IL6, TNF, and IL1B, which mediate systemic inflammation and muscle proteolysis; (ii) TGF-β superfamily ligands, such as GDF15, MSTN, and INHBA (encoding activin A), which suppress muscle anabolism and activate catabolic pathways; and (iii) metabolic and lipid-mobilizing factors, notably AZGP1 (ZAG) and LIF, which enhance lipolysis, alter energy expenditure, and promote anorexia in several tumor models. Extensive experimental work has validated the cachexia-inducing activity of these mediators: IL-6 neutralization attenuates wasting in the C26 carcinoma model; TNF-α was identified as the macrophage-derived cachexia factor “cachectin”; GDF15 drives tumor-induced anorexia via its receptor GFRAL; myostatin and activin A potently inhibit muscle growth; AZGP1 functions as a lipid-mobilizing factor; and tumor-derived LIF induces severe cachexia in melanoma-bearing mice. Together, these findings establish a mechanistic framework for interpreting tumor-intrinsic transcriptional changes in cachexia-inducing cancer cells.
Having defined this literature-based reference framework, we next investigated whether acquisition of cachexia-inducing capacity in AkuNEC cells is accompanied by transcriptional upregulation of these known tumor-derived factors. To this end, we analyzed RNA-seq data obtained from the non–cachexia-inducing parental line TCC-NECT-2 and its cachexia-inducing derivative AkuNEC, both at baseline and following exposure to doxorubicin. The list of candidate genes extracted from the literature was used as a predefined reference panel for interpretation of the transcriptomic data (Supplementary Table S1).
3.3. Baseline Transcriptional Differences Between TCC-NECT-2 and AkuNEC
We first compared baseline gene expression profiles between TCC-NECT-2 and AkuNEC under no-treatment conditions to identify tumor-intrinsic transcriptional differences associated with cachexia-inducing capacity. A complete list of baselines, differentially expressed genes and corresponding log2FC values is provided in Supplementary Table S3, which served as the basis for ranked and pathway analyses. Differential expression was evaluated using log2 fold-change (log2FC) values calculated from the mean expression of two biological replicates per condition.
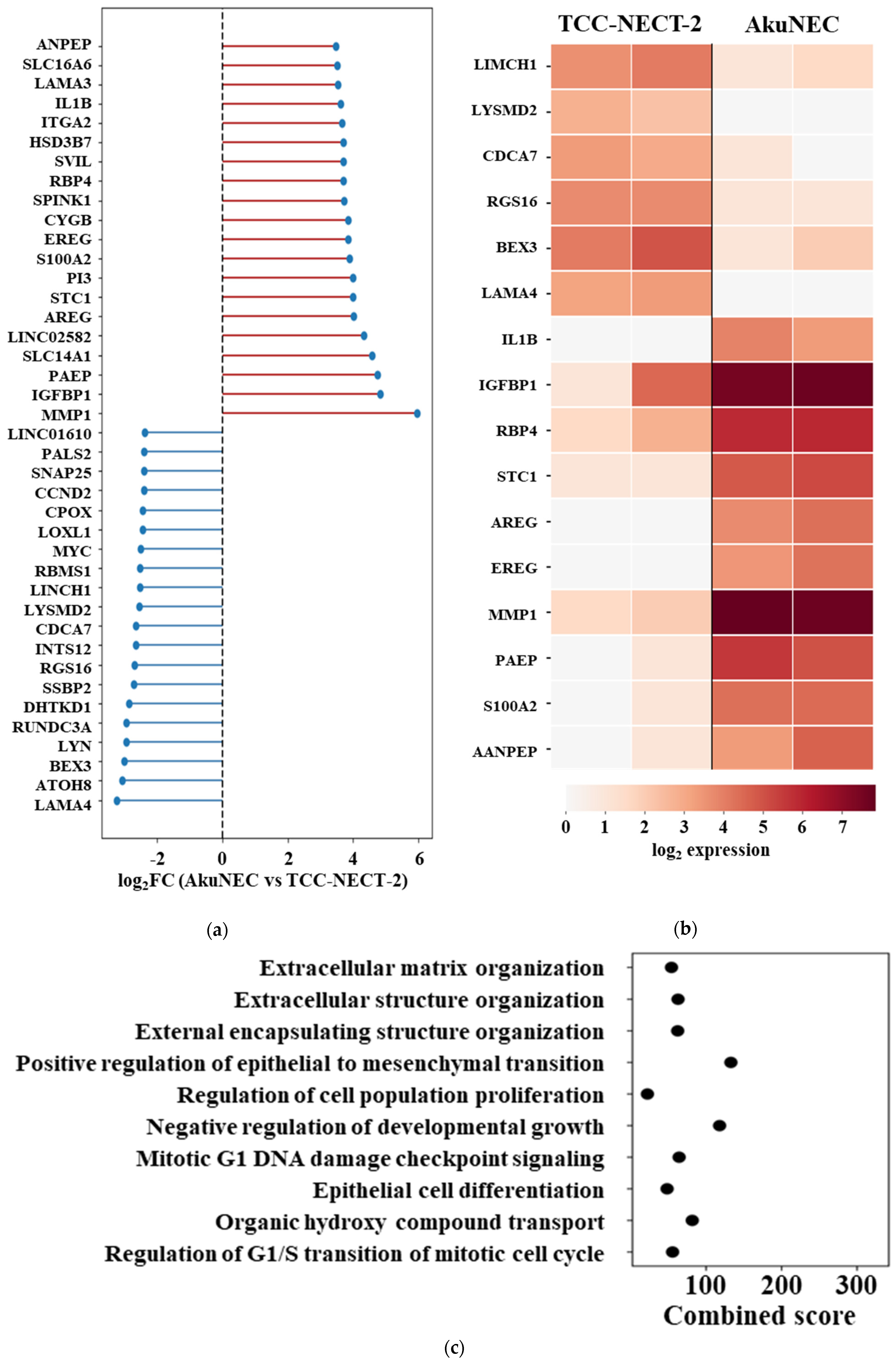
A bidirectional ranked plot demonstrated extensive baseline transcriptional reprogramming in AkuNEC cells relative to TCC-NECT-2, with both upregulated and downregulated gene sets clearly evident (Figure 2a). Genes exhibiting increased expression in AkuNEC included multiple inflammatory, secretory, and metabolic regulators, whereas genes reduced in AkuNEC were enriched for factors associated with cellular structure, differentiation, and homeostatic regulation.
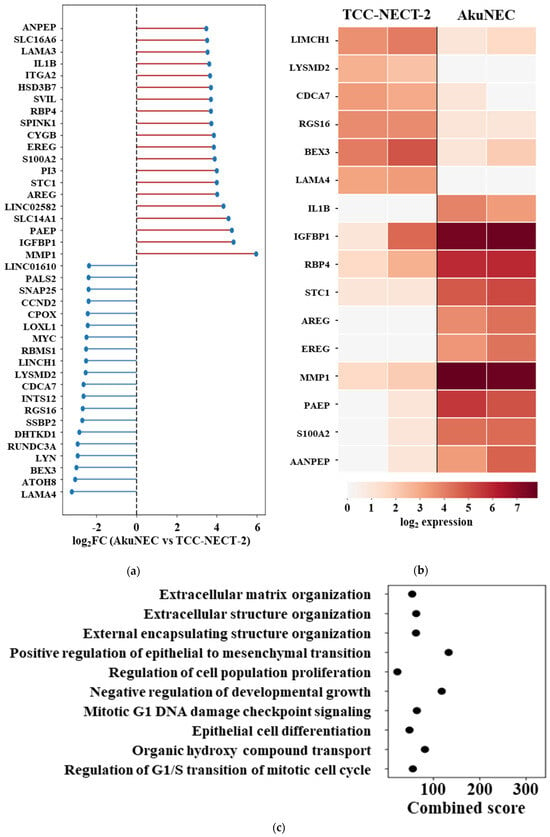
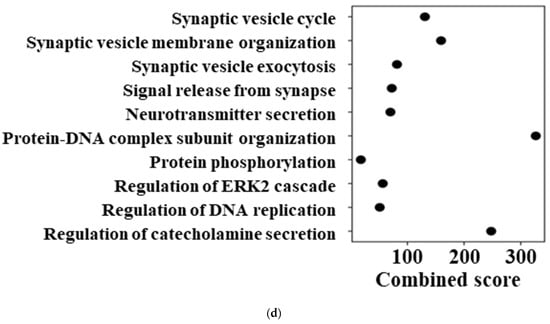
Figure 2.
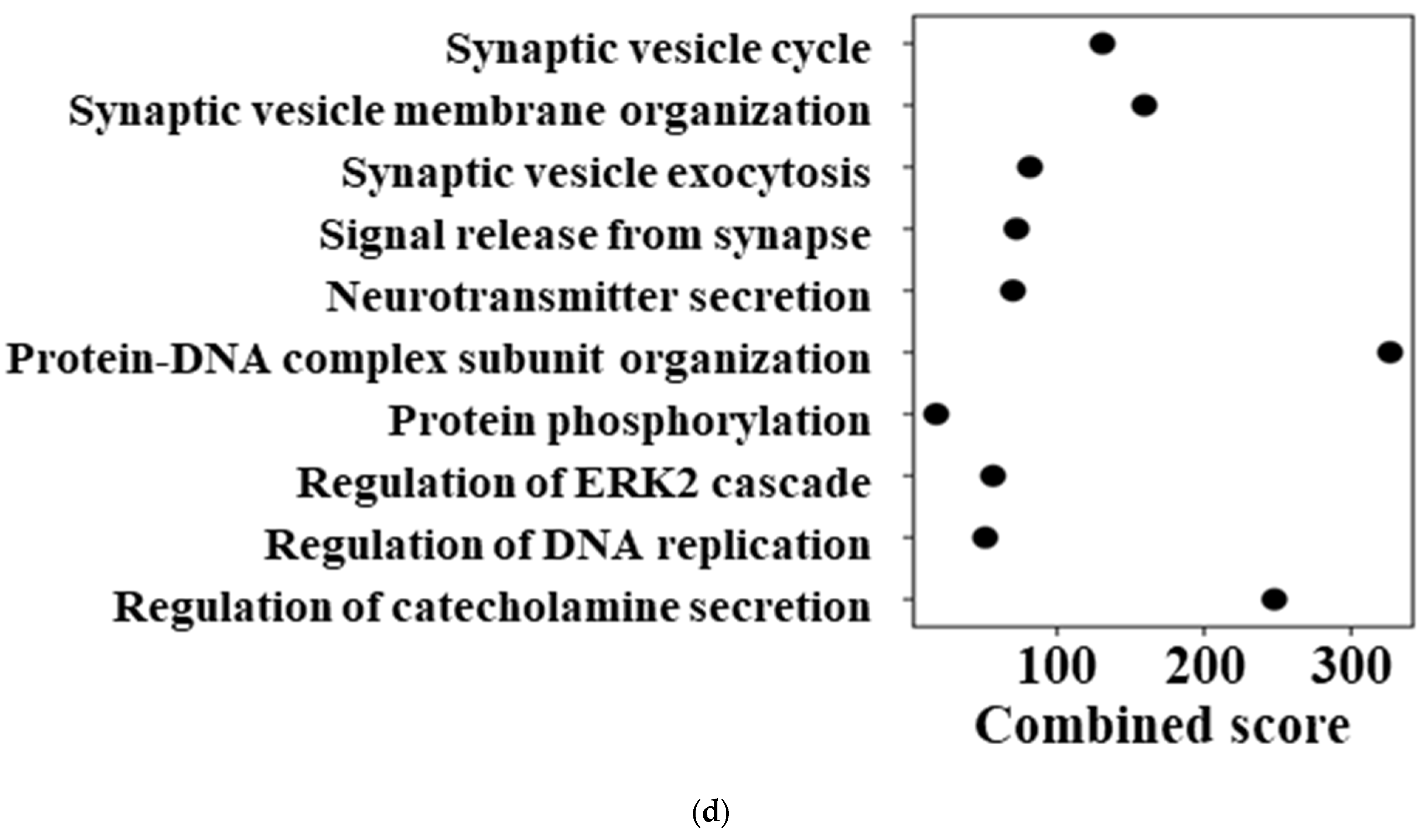
Baseline transcriptional differences between AkuNEC and TCC-NECT-2. (a) Ranked plot showing log2 fold changes (log2FC) of gene expression between AkuNEC and TCC-NECT-2 under no-treatment conditions. Genes are ordered according to log2FC, with positive values indicating higher expression in AkuNEC and negative values indicating lower expression relative to TCC-NECT-2, illustrating bidirectional baseline transcriptional reprogramming associated with cachexia-inducing capacity. Log2FC values were calculated from the mean expression of two biological replicates per condition (n = 2). (b) Heatmap showing expression patterns of 16 representative genes selected to illustrate bidirectional baseline transcriptional differences. Gene selection was based on a combination of literature-curated cachexia-associated tumor-derived factors (Supplementary Table S1) and data-driven ranking of baseline log2FC. Six genes downregulated in AkuNEC relative to TCC-NECT-2 are shown in the upper panel, while ten upregulated genes are shown in the lower panel. Expression values are log2-transformed, and each column represents an individual biological replicate (n = 2 per condition). The vertical line separates TCC-NECT-2 and AkuNEC samples. (c,d) Gene ontology (GO) biological process enrichment analysis performed separately for genes upregulated (c) and downregulated (d) in AkuNEC relative to TCC-NECT-2 under no-treatment conditions. Enrichment results are shown as dot plots ranked by combined score. Upregulated genes were primarily associated with inflammatory responses, secretory pathways, and extracellular matrix remodeling, whereas downregulated genes were enriched for processes related to cellular structure, differentiation, and homeostatic regulation. Gene sets were derived from baseline log2FC values and analyzed without statistical filtering to reflect effect-size–driven transcriptional differences (n = 2).
To visualize representative features of this bidirectional reprogramming, we constructed a heatmap of 16 selected genes integrating literature-curated cachexia-associated factors with data-driven baseline expression differences (Figure 2b). The log2-normalized expression values of these 16 genes are shown in Supplementary Table S7. Six genes downregulated in AkuNEC (LIMCH1, LYSMD2, CDCA7, RGS16, BEX3, and LAMA4) were displayed in the upper portion of the heatmap, while ten upregulated genes (IL1B, IGFBP1, RBP4, STC1, AREG, EREG, MMP1, PAEP, S100A2, and ANPEP) were shown in the lower portion. This analysis revealed a coordinated loss of baseline cellular and structural programs alongside the acquisition of inflammatory and secretory transcriptional states in AkuNEC cells.
We next examined the functional implications of these baseline transcriptional differences using pathway enrichment analysis. Gene ontology (GO) biological process enrichment was performed separately for genes upregulated and downregulated in AkuNEC based on baseline log2FC rankings. Gene lists used for enrichment analyses are provided in Supplementary Table S5 (upregulated) and Table S6 (downregulated). Pathways enriched among upregulated genes (Figure 2c) were predominantly associated with extracellular matrix organization, epithelial–mesenchymal transition, regulation of cell population proliferation, inflammatory signaling, and stress-related processes. In contrast, pathways enriched among downregulated genes (Figure 2d) were associated with synaptic vesicle organization, neurotransmitter secretion, protein–DNA complex organization, regulation of ERK1/2 signaling, and DNA replication-related processes. The same combined score scale was applied to both panels to allow direct comparison of functional enrichment patterns.
Collectively, these results indicate that acquisition of cachexia-inducing capacity in AkuNEC cells is accompanied by extensive baseline transcriptional reprogramming, characterized by upregulation of inflammatory, extracellular, and secretory gene programs and concomitant suppression of cellular structural and homeostatic pathways. This baseline state provides a molecular context for subsequent analyses of chemotherapeutic stress responses.
3.4. Doxorubicin-Induced Transcriptional Responses in TCC-NECT-2 and AkuNEC
Having established that AkuNEC cells exhibit extensive baseline transcriptional reprogramming associated with cachexia-inducing capacity (Figure 2), we next examined how these intrinsic differences influence transcriptional responses to chemotherapeutic stress. To this end, we analyzed gene expression changes following doxorubicin treatment in TCC-NECT-2 and AkuNEC cells and compared both the individual drug responses and their differential patterns.
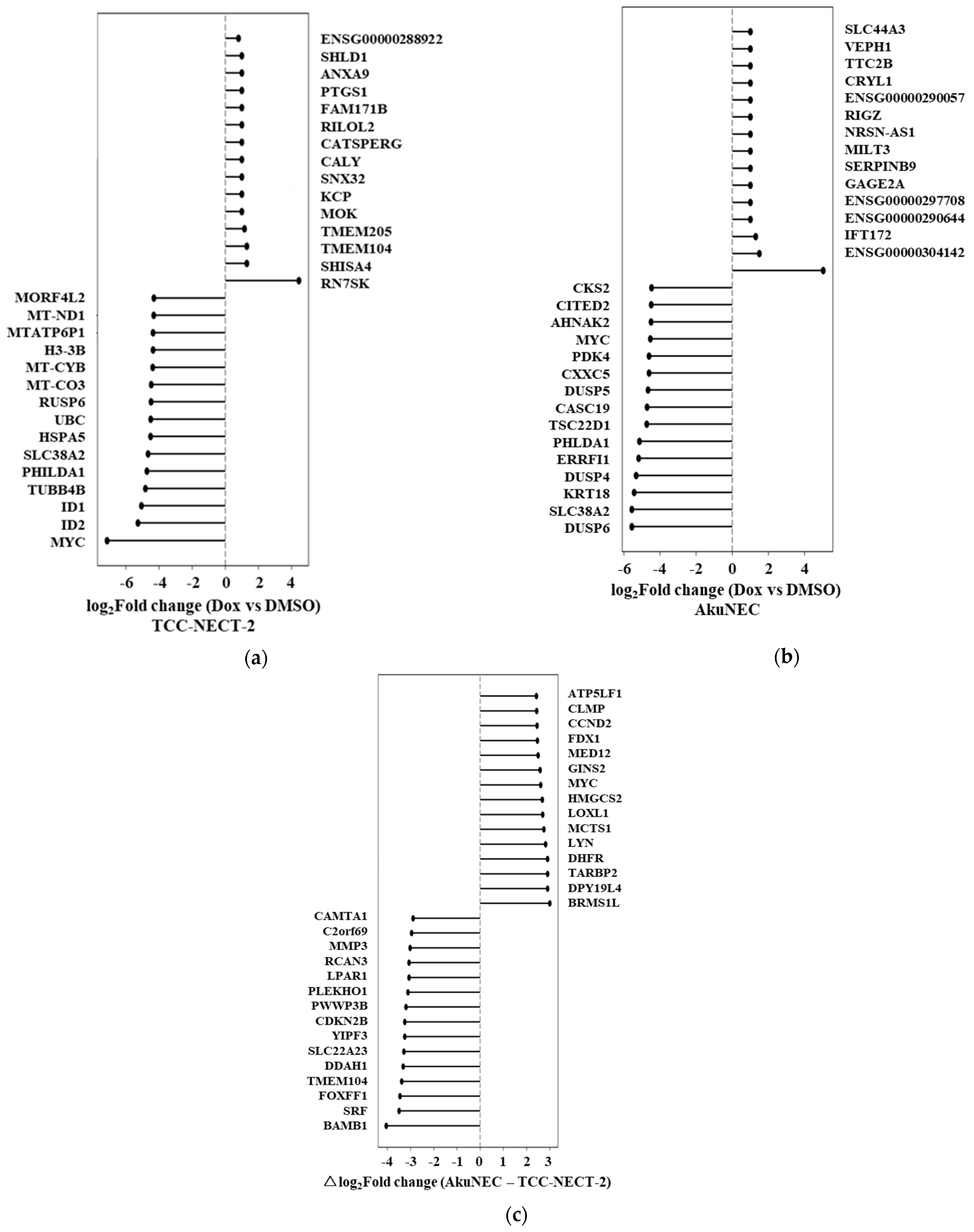
In TCC-NECT-2 cells, doxorubicin exposure induced a transcriptional response characterized primarily by activation of general stress- and transcription-associated pathways (Figure 3a). Prominent changes included strong induction of RN7SK, a regulator of transcriptional elongation, along with alterations in genes associated with membrane trafficking, stress signaling, and prostaglandin metabolism, such as PTGS1. Concurrently, several genes related to proliferation and mitochondrial function, including MYC and mitochondrial-encoded transcripts, were markedly downregulated. Overall, this response pattern is consistent with a canonical DNA damage–associated stress response accompanied by suppression of proliferative programs. The full list of doxorubicin-responsive genes in TCC-NECT-2 cells is provided in Supplementary Table S7.
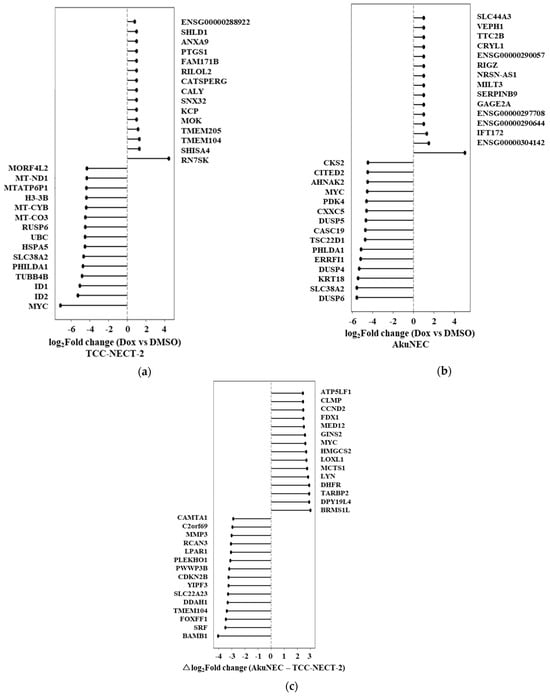

Figure 3.
Doxorubicin-induced transcriptional responses in TCC-NECT-2 and AkuNEC. (a) Bidirectional ranked plot showing log2 fold changes (log2FC) of gene expression in TCC-NECT-2 cells following doxorubicin treatment relative to DMSO-treated controls. Genes are ordered according to log2FC, with positive values indicating induction and negative values indicating suppression upon drug exposure. Log2FC values were calculated from the mean expression of two biological replicates per condition (n = 2). Genes displayed represent the top and bottom differentially expressed genes and are listed in Supplementary Table S10. (b) Bidirectional ranked plot showing doxorubicin-induced transcriptional changes in AkuNEC cells, calculated in the same manner as in (a). Compared with TCC-NECT-2, AkuNEC cells exhibited a distinct pattern of drug-responsive gene expression, reflecting qualitative differences in stress-adaptive transcriptional programs. Genes displayed represent the top- and bottom-ranked genes and are listed in Supplementary Table S11. (c) Bidirectional ranked plot of the differential doxorubicin response (Δresponse) between AkuNEC and TCC-NECT-2, defined as: Δlog2FC = (AkuNEC_Dox − AkuNEC_DMSO) − (TCC-NECT-2_Dox − TCC-NECT-2_DMSO). Genes are ranked according to Δlog2FC, with positive values indicating stronger induction in AkuNEC and negative values indicating attenuated or reduced responses relative to TCC-NECT-2. Genes displayed represent the top- and bottom-ranked genes and are listed in Supplementary Table S12.
In contrast, AkuNEC cells displayed a distinct transcriptional response to doxorubicin treatment (Figure 3b). Although induction of RN7SK was also observed, the AkuNEC response was notable for the additional upregulation of genes implicated in immune-like stress signaling, apoptosis resistance, and adaptive survival pathways, including SERPINB9, DRAM1, IFIT2, and multiple dual-specificity phosphatases. At the same time, downregulated genes in AkuNEC included regulators of transcriptional control and cellular metabolism, suggesting a coordinated reprogramming rather than uniform suppression of gene expression. These features indicate that doxorubicin elicits a more complex and adaptive transcriptional response in AkuNEC cells than in the non–cachexia-inducing parental line. The full list of doxorubicin-responsive genes in AkuNEC is provided in Supplementary Table S8.
To directly compare drug-induced responses between the two cell lines, we calculated the differential doxorubicin response (Δresponse), defined as the difference in log2 fold-change between AkuNEC and TCC-NECT-2 (Figure 3c). This analysis revealed a set of genes that were preferentially induced in AkuNEC under doxorubicin stress, including CCND2, GINS2, MYC, HMGCS2, FDX1, DHFR, and BRMS1L. These genes are involved in cell cycle regulation, DNA replication, metabolic reprogramming, and mitochondrial function, suggesting enhanced transcriptional plasticity in AkuNEC cells. Conversely, genes exhibiting relatively attenuated responses in AkuNEC included BAMBI, CDKN2B, FOXF1, SRF, and other regulators associated with differentiation and growth suppression. Genes ranked by Δresponse are provided in Supplementary Table S9.
Taken together, these results demonstrate that the cachexia-inducing cell line AkuNEC not only exhibits a distinct baseline transcriptional state (Figure 2) but also responds to chemotherapeutic stress through a qualitatively different transcriptional program. Rather than displaying a uniformly heightened sensitivity or resistance, AkuNEC cells undergo selective reorganization of proliferation-, metabolism-, and stress-related pathways in response to doxorubicin. This dynamic response contrasts with the more stereotypical stress-induced suppression observed in TCC-NECT-2 cells and suggests that cachexia-associated tumor-intrinsic programs may shape how cancer cells adapt to chemotherapy. Genes displayed in Figure 3a–c are listed in Supplementary Tables S10–S12.
4. Discussion
4.1. Phenotypic Similarity Masks Tumor-Intrinsic Reprogramming Associated with Cachexia-Inducing Capacity
In this study, we found that the cachexia-inducing cell line AkuNEC and its non–cachexia-inducing parental counterpart TCC-NECT-2 were largely indistinguishable at the level of gross cellular morphology (Figure 1). The absence of overt morphological differences is consistent with the concept that cachexia is primarily driven by tumor–host systemic interactions and endocrine-like signaling rather than by easily detectable changes in cancer cell shape under routine culture conditions [1,2,3,4]. In addition, the limited ability of conventional phenotypic assays to capture clinically relevant tumor–host pathophysiology has been discussed in broader contexts of translational cancer research and preclinical modeling [16]. From a clinical perspective, the constitutive upregulation of inflammatory and metabolic factors in AkuNEC cells suggests that these signatures could serve as candidate prognostic biomarkers to identify patients at higher risk for therapy intolerance and early-onset wasting.
Although scanning electron microscopy suggested subtle differences in surface features, these changes were qualitative and modest in scale. Such subtle alterations may reflect differences in cell–cell or cell–matrix interactions, which can influence secretion, extracellular matrix remodeling, and host signaling [17,18,19]. Importantly, the lack of striking phenotypic divergence supports the interpretation that cachexia-inducing capacity may arise from tumor-intrinsic transcriptional reprogramming that is not necessarily mirrored by gross morphology [1,2,3,4].
Although AkuNEC cells displayed increased sensitivity to doxorubicin in short-term viability assays, this difference was not accompanied by overt morphological changes, underscoring that quantitative drug sensitivity and qualitative transcriptional reprogramming represent partially distinct aspects of the cachexia-associated tumor phenotype. In this context, transcriptomic profiling is useful for capturing stress-induced pathway reorganization that may not translate into large differences in short-term viability assays [20,21].
A central finding of this study is that multiple factors previously implicated in cancer cachexia were constitutively upregulated in AkuNEC cells at baseline (Figure 2). The literature-curated cachexia-associated reference gene set used in this analysis is provided in Supplementary Table S1. Inflammatory and secretory mediators such as IL1B, together with metabolic/endocrine-related regulators including IGFBP1, RBP4, STC1, AREG, and EREG, were increased in AkuNEC relative to TCC-NECT-2. These observations are consistent with a large body of literature supporting tumor-derived inflammatory cytokines and secreted factors as drivers or correlates of systemic metabolic dysregulation, anorexia, and tissue wasting [1,2,3,4,22,23,24,25,26,27]. Classical experimental work established causal links between cytokines and wasting phenotypes (e.g., IL-6 in the C26 model; TNF/cachectin; IL-1–associated anorexia pathways), providing a mechanistic basis for interpreting inflammatory gene programs within cachexia-inducing tumor contexts [22,23,24,25,26,27].
Importantly, cachexia-associated genes were not upregulated in isolation. We observed coordinated induction of additional genes with comparable effect sizes (e.g., MMP1, PAEP, S100A2, ANPEP), many of which are linked to extracellular matrix remodeling, stress responses, and secretory biology. Such coordinated shifts are consistent with the concept that cachexia is rarely attributable to a single “master factor” and instead reflects network-level tumor programs that collectively perturb host metabolism and tissue homeostasis [1,2,3,4,22,23,24,25,26,27,28,29]. In support of this view, multiple mechanistic studies have demonstrated that distinct tumor-derived pathways can converge on muscle proteolysis, adipose browning/lipolysis, and systemic inflammation, including ATGL-mediated lipolysis, PTHrP-driven thermogenic programming, and activin/myostatin signaling via ActRIIB [7,8,9,10,30,31].
At the pathway level, enrichment among genes upregulated in AkuNEC highlighted inflammatory signaling, extracellular matrix organization, and stress-associated processes (Figure 2c), whereas downregulated programs were enriched for cellular structural and homeostatic regulation (Figure 2d). Gene lists underlying these enrichment analyses are provided in Supplementary Tables S11 and S12. This bidirectional remodeling is compatible with contemporary multi-organ and multi-omics views of cachexia, which emphasize early systemic host abnormalities, broad endocrine and immune changes, and progressive rewiring of metabolism and inflammation across tissues [32,33,34]. Together, our data support a model in which cachexia-inducing capacity is embedded in coordinated tumor-intrinsic transcriptional states that generate a cachexia-promoting milieu through cumulative or synergistic effects of multiple factors rather than isolated mediators.
4.2. Stress-Induced Transcriptional Reprogramming Builds upon a Cachexia-Associated Baseline Program
Baseline transcriptional reprogramming in AkuNEC provides a framework for interpreting the response to chemotherapeutic stress. Chemotherapy can modulate systemic inflammation and metabolic states, and several studies have suggested that stress-induced endocrine and inflammatory pathways may influence appetite, muscle wasting, and immune competence [35,36,37]. In our data, doxorubicin elicited general stress-associated changes in both cell lines, including induction of RN7SK, consistent with transcriptional stress and global regulatory responses [38,39]. However, the overall response pattern differed between TCC-NECT-2 and AkuNEC (Figure 3a,b), suggesting that cachexia-associated baseline programs may predispose tumor cells to qualitatively distinct stress-adaptive transcriptional trajectories. Since doxorubicin is not the first-line chemotherapy for neuroendocrine carcinoma, the observed responses represent a general model of stress-induced plasticity rather than a direct clinical simulation.
AkuNEC cells showed selective induction of genes linked to immune-like stress signaling, autophagy/apoptosis control, and adaptive survival pathways. Such responses align with broader concepts of transcriptional plasticity under stress, in which tumor cells reorganize regulatory networks to sustain viability and metabolic flexibility [40,41,42,43,44]. The Δresponse analysis further identified preferential induction of genes involved in cell cycle regulation, DNA replication, and metabolic remodeling (Figure 3c). Gene lists used for Δresponse-based enrichment analyses are provided in Supplementary Tables S13 and S14. Notably, metabolic rewiring and mitochondrial programs have been implicated in cachexia-related host metabolism and systemic immune modulation, suggesting possible links between tumor stress responses and host-pathophysiology–relevant signaling [45,46,47,48]. Genes commonly responsive to doxorubicin are summarized in Supplementary Table S15, whereas AkuNEC-specific responses are summarized in Supplementary Table S16.
Taken together, our findings support a two-tiered framework: (i) baseline cachexia-associated tumor-intrinsic transcriptional programs that may shape tumor–host interactions and systemic wasting, and (ii) stress-induced transcriptional plasticity that reorganizes proliferation- and metabolism-related pathways under chemotherapeutic challenge. This framework reconciles the limited phenotypic divergence observed in standard assays (Figure 1) with pronounced molecular divergence at the transcriptomic level (Figure 2 and Figure 3) and argues that cachexia-associated tumor evolution may affect not only host physiology but also qualitative aspects of tumor stress adaptation [1,2,3,4,5,6,20].
4.3. Limitations and Future Perspectives
Several limitations of the present study should be acknowledged. First, the transcriptomic analyses were performed with a limited number of biological replicates (n = 2 per condition). While this design was sufficient to detect robust and reproducible effect-size–driven transcriptional differences between TCC-NECT-2 and AkuNEC cells, it restricts the use of formal statistical testing and precludes definitive conclusions regarding variability across broader biological contexts. The findings should be viewed as hypothesis-generating, and further validation in larger cohorts is required. Future studies using standard-of-care agents, such as etoposide or platinum-based drugs, will be necessary to confirm the clinical relevance of these sensitivity programs.
Second, this study was conducted exclusively using in vitro cell line models. Although the paired nature of the TCC-NECT-2/AkuNEC system provides a controlled framework for dissecting tumor-intrinsic programs associated with cachexia-inducing capacity, it does not fully recapitulate the complex tumor–host interactions that underlie cancer cachexia in vivo. In particular, interactions with immune cells, stromal components, and systemic metabolic organs are not captured in this experimental setting [12].
Third, while our analyses identified coordinated upregulation of multiple inflammatory, secretory, and metabolic regulators in AkuNEC cells, functional validation of individual factors was beyond the scope of this study. We did not directly assess the secretion, bioactivity, or host-modulating effects of candidate cachexia-associated proteins identified at the transcriptomic level. As such, the relative contribution of individual factors versus their combined or synergistic effects remains to be determined. Moreover, this study lacks protein-level validation of the identified factors, and the validation study at the protein-level using the proteomics modalities and the direct assessment of secreted proteins via ELISA or Western blotting, alongside functional assays such as myotube atrophy models, will be critical to establish the causal contribution and should be performed.
Despite these limitations, the present study provides several important perspectives for future research. The observation that cachexia-inducing capacity is associated with coordinated baseline transcriptional reprogramming and enhanced stress-induced plasticity suggests that cancer cachexia may emerge from network-level tumor-intrinsic programs rather than from single dominant mediators. Future studies integrating transcriptomic data with proteomic profiling of tumor-conditioned media, functional assays using muscle or adipose cell models, and in vivo validation in xenograft or organoid-based systems will be critical to establish causal links between tumor-derived programs and systemic cachexia phenotypes.
Moreover, extending this analytical framework to additional cachexia-inducing and non-inducing tumor models, as well as to patient-derived samples, may help clarify whether the transcriptional architecture identified here represents a generalizable feature of cachexia-associated cancers. Such efforts could ultimately inform strategies aimed at disrupting coordinated tumor-derived programs, rather than targeting individual factors in isolation, as a means of mitigating cancer cachexia. Future investigations utilizing confocal microscopy for intracellular localization and organelle-specific staining will be necessary to provide a comprehensive internal morphological assessment under stress.
5. Conclusions
In summary, this study demonstrates that cachexia-inducing capacity in tumor cells is associated with coordinated tumor-intrinsic transcriptional reprogramming rather than with overt phenotypic differences in morphology, suggesting that qualitative transcriptional divergence may be more pronounced than quantitative differences in bulk cell viability. Using a paired duodenal neuroendocrine carcinoma cell line system, we show that the cachexia-inducing subline AkuNEC exhibits constitutive upregulation of inflammatory, secretory, and metabolic gene programs at baseline, alongside suppression of cellular structural and homeostatic pathways.
Importantly, this baseline transcriptional state shapes the response to chemotherapeutic stress. Upon doxorubicin exposure, AkuNEC cells display a qualitatively distinct transcriptional response characterized by selective reorganization of proliferation-, metabolism-, and stress-related pathways, indicative of enhanced transcriptional plasticity. These findings suggest that cachexia-associated tumor programs and stress-induced adaptive responses are functionally linked.
Together, our results support a model in which cancer cachexia emerges from network-level tumor-intrinsic programs that influence both systemic host interactions and tumor adaptability under therapeutic stress. This work provides a framework for exploring cachexia as a consequence of coordinated transcriptional states rather than isolated molecular factors and highlights the importance of integrating baseline and stress-induced tumor programs in future cachexia research.
Supplementary Materials
The following supporting information can be downloaded at: https://www.mdpi.com/article/10.3390/targets4010009/s1. Supplementary Table S1. Literature-curated tumor-derived factors implicated in cancer cachexia. Supplementary Table S2. Log2-normalized gene expression profiles of TCC-NECT-2 and AkuNEC cells. Supplementary Table S3: Baseline differential gene expression between AkuNEC and TCC-NECT-2 cells. Supplementary Table S4: Log2-normalized expression values of the 16 representative genes shown in Figure 2b. Supplementary Table S5. Gene list used for pathway enrichment analysis of baseline upregulated genes. Supplementary Table S6. Gene list used for pathway enrichment analysis of baseline downregulated genes. Supplementary Table S7. Doxorubicin-induced transcriptional changes in TCC-NECT-2 cells. Supplementary Table S8. Doxorubicin-induced transcriptional changes in AkuNEC cells. Supplementary Table S9. Differential doxorubicin response between AkuNEC and TCC-NECT-2 cells. Supplementary Table S10. Top and bottom differentially expressed genes in TCC-NECT-2 cells (Figure 3a). Supplementary Table S11. Top and bottom differentially expressed genes in AkuNEC cells (Figure 3b). Supplementary Table S12. Top and bottom genes ranked by differential doxorubicin response (Figure 3c). Supplementary Table S13. Gene list used for pathway enrichment analysis of genes with increased Δresponse. Supplementary Table S14. Gene list used for pathway enrichment analysis of genes with decreased Δresponse. Supplementary Table S15. Genes commonly responsive to doxorubicin in TCC-NECT-2 and AkuNEC cells. Supplementary Table S16. Genes showing AkuNEC-specific transcriptional responses to doxorubicin.
Author Contributions
Conceptualization, Y.S.; methodology, Y.Y.; formal analysis, T.K.; investigation, J.O.; resources, K.Y.; data curation, Y.Y.; writing—original draft preparation, T.K. and Y.Y.; writing—review and editing, T.K. and Y.Y.; visualization, T.K.; supervision, Y.Y.; project administration, Y.Y. All authors have read and agreed to the published version of the manuscript.
Funding
This research was supported by internal research funding provided to Yuki Yoshimatsu by the Tochigi Cancer Center in fiscal year 2025.
Institutional Review Board Statement
Ethical review and approval were not required for this study because it involved established human cell lines and did not include human participants or identifiable personal data.
Informed Consent Statement
Not applicable. Ethical approval was not required for this study, as it involved the use of established patient-derived cell lines that are publicly available and did not include identifiable human subjects.
Data Availability Statement
The data are available upon reasonable request.
Acknowledgments
The authors thank all members of the participating laboratories for their valuable discussions. RNA sequencing data were generated by Zeon BioSolutions, Inc. (Hyogo, Japan) as a contracted service provider. The authors also acknowledge the use of ChatGPT (OpenAI, San Francisco, CA, USA) for improving the clarity and readability of the English text. The authors remain fully responsible for the scientific content, interpretation, and conclusions presented in this study.
Conflicts of Interest
The authors declare no conflicts of interest associated with this manuscript.
References
- Fearon, K.; Strasser, F.; Anker, S.D.; Bosaeus, I.; Bruera, E.; Fainsinger, R.L.; Jatoi, A.; Loprinzi, C.; MacDonald, N.; Mantovani, G.; et al. Definition and classification of cancer cachexia: An international consensus. Lancet Oncol. 2011, 12, 489–495. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, H.; Asakawa, A.; Amitani, H.; Nakamura, N.; Inui, A. Cancer cachexia--pathophysiology and management. J. Gastroenterol. 2013, 48, 574–594. [Google Scholar] [CrossRef]
- Schmidt, S.F.; Rohm, M.; Herzig, S.; Berriel Diaz, M. Cancer Cachexia: More Than Skeletal Muscle Wasting. Trends Cancer 2018, 4, 849–860. [Google Scholar] [CrossRef]
- Li, L.; Ling, Z.Q. Mechanisms of cancer cachexia and targeted therapeutic strategies. Biochim. Biophys. Acta Rev. Cancer 2024, 1879, 189208. [Google Scholar] [CrossRef]
- Khalil, M.A.; Jafri, S.A.A.; Zahra Naqvi, M.E.; Jauhar, I.; Kumar, V.; Ali, S.M.S.; Khalil, S.; Raheem, A.; Kumar, R.; Haseeb, A.; et al. Impact of Cachexia on Chemotherapy Efficacy and Survival in Pancreatic Cancer: A Systematic Review and Meta-Analysis. Cancer Control 2024, 31, 10732748241292784. [Google Scholar] [CrossRef]
- Pin, F.; Couch, M.E.; Bonetto, A. Preservation of muscle mass as a strategy to reduce the toxic effects of cancer chemotherapy on body composition. Curr. Opin. Support. Palliat. Care 2018, 12, 420–426. [Google Scholar] [CrossRef]
- Petruzzelli, M.; Schweiger, M.; Schreiber, R.; Campos-Olivas, R.; Tsoli, M.; Allen, J.; Swarbrick, M.; Rose-John, S.; Rincon, M.; Robertson, G.; et al. A switch from white to brown fat increases energy expenditure in cancer-associated cachexia. Cell Metab. 2014, 20, 433–447. [Google Scholar] [CrossRef]
- Kir, S.; White, J.P.; Kleiner, S.; Kazak, L.; Cohen, P.; Baracos, V.E.; Spiegelman, B.M. Tumour-derived PTH-related protein triggers adipose tissue browning and cancer cachexia. Nature 2014, 513, 100–104. [Google Scholar] [CrossRef]
- Kir, S.; Komaba, H.; Garcia, A.P.; Economopoulos, K.P.; Liu, W.; Lanske, B.; Hodin, R.A.; Spiegelman, B.M. PTH/PTHrP Receptor Mediates Cachexia in Models of Kidney Failure and Cancer. Cell Metab. 2016, 23, 315–323. [Google Scholar] [CrossRef] [PubMed]
- Das, S.K.; Eder, S.; Schauer, S.; Diwoky, C.; Temmel, H.; Guertl, B.; Gorkiewicz, G.; Tamilarasan, K.P.; Kumari, P.; Trauner, M.; et al. Adipose triglyceride lipase contributes to cancer-associated cachexia. Science 2011, 333, 233–238. [Google Scholar] [CrossRef] [PubMed]
- Dahlman, I.; Mejhert, N.; Linder, K.; Agustsson, T.; Mutch, D.M.; Kulyte, A.; Isaksson, B.; Permert, J.; Petrovic, N.; Nedergaard, J.; et al. Adipose tissue pathways involved in weight loss of cancer cachexia. Br. J. Cancer 2010, 102, 1541–1548. [Google Scholar] [CrossRef] [PubMed]
- Talbert, E.E.; Cuitiño, M.C.; Ladner, K.J.; Rajasekerea, P.V.; Siebert, M.; Shakya, R.; Leone, G.W.; Ostrowski, M.C.; Paleo, B.; Weisleder, N.; et al. Modeling Human Cancer-induced Cachexia. Cell Rep. 2019, 28, 1612–1622.e4. [Google Scholar] [CrossRef]
- Yanagihara, K.; Kubo, T.; Mihara, K.; Kuwata, T.; Ochiai, A.; Seyama, T.; Yokozaki, H. Establishment of a novel cell line from a rare human duodenal poorly differentiated neuroendocrine carcinoma. Oncotarget 2018, 9, 36503–36514. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Yanagihara, K.; Kubo, T.; Iino, Y.; Mihara, K.; Morimoto, C.; Seyama, T.; Kuwata, T.; Ochiai, A.; Yokozaki, H. Development and characterization of a cancer cachexia model employing a rare human duodenal neuroendocrine carcinoma-originating cell line. Oncotarget 2019, 10, 2435–2450. [Google Scholar] [CrossRef]
- Minami, F.; Sasaki, N.; Shichi, Y.; Gomi, F.; Michishita, M.; Ohkusu-Tsukada, K.; Toyoda, M.; Takahashi, K.; Ishiwata, T. Morphofunctional analysis of human pancreatic cancer cell lines in 2- and 3-dimensional cultures. Sci. Rep. 2021, 11, 6775. [Google Scholar] [CrossRef]
- Hou, X.; Du, C.; Lu, L.; Yuan, S.; Zhan, M.; You, P.; Du, H. Opportunities and challenges of patient-derived models in cancer research: Patient-derived xenografts, patient-derived organoid and patient-derived cells. World J. Surg. Oncol. 2022, 20, 37. [Google Scholar] [CrossRef]
- Lu, P.; Weaver, V.M.; Werb, Z. The extracellular matrix: A dynamic niche in cancer progression. J. Cell Biol. 2012, 196, 395–406. [Google Scholar] [CrossRef]
- Mina-Osorio, P. The moonlighting enzyme CD13: Old and new functions to target. Trends Mol. Med. 2008, 14, 361–371. [Google Scholar] [CrossRef] [PubMed]
- Pickup, M.W.; Mouw, J.K.; Weaver, V.M. The extracellular matrix modulates the hallmarks of cancer. EMBO Rep. 2014, 15, 1243–1253. [Google Scholar] [CrossRef]
- Kim, J.; DeBerardinis, R.J. Mechanisms and Implications of Metabolic Heterogeneity in Cancer. Cell Metab. 2019, 30, 434–446. [Google Scholar] [CrossRef]
- Hata, A.N.; Engelman, J.A.; Faber, A.C. The BCL2 Family: Key Mediators of the Apoptotic Response to Targeted Anticancer Therapeutics. Cancer Discov. 2015, 5, 475–487. [Google Scholar] [CrossRef] [PubMed]
- Strassmann, G.; Fong, M.; Kenney, J.S.; Jacob, C.O. Evidence for the involvement of interleukin 6 in experimental cancer cachexia. J. Clin. Investig. 1992, 89, 1681–1684. [Google Scholar] [CrossRef]
- Beutler, B.; Greenwald, D.; Hulmes, J.D.; Chang, M.; Pan, Y.C.; Mathison, J.; Ulevitch, R.; Cerami, A. Identity of tumour necrosis factor and the macrophage-secreted factor cachectin. Nature 1985, 316, 552–554. [Google Scholar] [CrossRef]
- Uehara, A.; Sekiya, C.; Takasugi, Y.; Namiki, M.; Arimura, A. Anorexia induced by interleukin 1: Involvement of corticotropin-releasing factor. Am. J. Physiol. 1989, 257, R613–R617. [Google Scholar] [CrossRef]
- Roeland, E.J.; Bohlke, K.; Baracos, V.E.; Bruera, E.; Del Fabbro, E.; Dixon, S.; Fallon, M.; Herrstedt, J.; Lau, H.; Platek, M.; et al. Management of Cancer Cachexia: ASCO Guideline. J. Clin. Oncol. 2020, 38, 2438–2453. [Google Scholar] [CrossRef]
- Argilés, J.M.; Busquets, S.; Stemmler, B.; López-Soriano, F.J. Cancer cachexia: Understanding the molecular basis. Nat. Rev. Cancer 2014, 14, 754–762. [Google Scholar] [CrossRef]
- Baracos, V.E.; Martin, L.; Korc, M.; Guttridge, D.C.; Fearon, K.C.H. Cancer-associated cachexia. Nat. Rev. Dis. Primers 2018, 4, 17105. [Google Scholar] [CrossRef] [PubMed]
- Fearon, K.C.; Glass, D.J.; Guttridge, D.C. Cancer cachexia: Mediators, signaling, and metabolic pathways. Cell Metab. 2012, 16, 153–166. [Google Scholar] [CrossRef]
- Nakamura, Y.; Saldajeno, D.P.; Kawaguchi, K.; Kawaoka, S. Progressive, multi-organ, and multi-layered nature of cancer cachexia. Cancer Sci. 2024, 115, 715–722. [Google Scholar] [CrossRef]
- Zhou, X.; Wang, J.L.; Lu, J.; Song, Y.; Kwak, K.S.; Jiao, Q.; Rosenfeld, R.; Chen, Q.; Boone, T.; Simonet, W.S.; et al. Reversal of cancer cachexia and muscle wasting by ActRIIB antagonism leads to prolonged survival. Cell 2010, 142, 531–543. [Google Scholar] [CrossRef] [PubMed]
- Zimmers, T.A.; Davies, M.V.; Koniaris, L.G.; Haynes, P.; Esquela, A.F.; Tomkinson, K.N.; McPherron, A.C.; Wolfman, N.M.; Lee, S.J. Induction of cachexia in mice by systemically administered myostatin. Science 2002, 296, 1486–1488. [Google Scholar] [CrossRef]
- Flint, T.R.; Janowitz, T.; Connell, C.M.; Roberts, E.W.; Denton, A.E.; Coll, A.P.; Jodrell, D.I.; Fearon, D.T. Tumor-Induced IL-6 Reprograms Host Metabolism to Suppress Anti-tumor Immunity. Cell Metab. 2016, 24, 672–684. [Google Scholar] [CrossRef] [PubMed]
- Hiam-Galvez, K.J.; Allen, B.M.; Spitzer, M.H. Systemic immunity in cancer. Nat. Rev. Cancer 2021, 21, 345–359. [Google Scholar] [CrossRef] [PubMed]
- Skipworth, R.J.; Stewart, G.D.; Dejong, C.H.; Preston, T.; Fearon, K.C. Pathophysiology of cancer cachexia: Much more than host-tumour interaction? Clin. Nutr. 2007, 26, 667–676. [Google Scholar] [CrossRef]
- Lerner, L.; Hayes, T.G.; Tao, N.; Krieger, B.; Feng, B.; Wu, Z.; Nicoletti, R.; Chiu, M.I.; Gyuris, J.; Garcia, J.M. Plasma growth differentiation factor 15 is associated with weight loss and mortality in cancer patients. J. Cachexia Sarcopenia Muscle 2015, 6, 317–324. [Google Scholar] [CrossRef]
- Cheng, L.Y.; Desai, J.; Neeson, P.J. Targeting GDF-15 in cancer cachexia and immunity. Nat. Med. 2025, 31, 1069–1070. [Google Scholar] [CrossRef]
- Patel, H.J.; Patel, B.M. TNF-α and cancer cachexia: Molecular insights and clinical implications. Life Sci. 2017, 170, 56–63. [Google Scholar] [CrossRef]
- Nguyen, V.T.; Kiss, T.; Michels, A.A.; Bensaude, O. 7SK small nuclear RNA binds to and inhibits the activity of CDK9/cyclin T complexes. Nature 2001, 414, 322–325. [Google Scholar] [CrossRef]
- Yang, F.; Kemp, C.J.; Henikoff, S. Doxorubicin enhances nucleosome turnover around promoters. Curr. Biol. 2013, 23, 782–787. [Google Scholar] [CrossRef] [PubMed]
- Amaravadi, R.; Kimmelman, A.C.; White, E. Recent insights into the function of autophagy in cancer. Genes Dev. 2016, 30, 1913–1930. [Google Scholar] [CrossRef]
- Ibáñez-Molero, S.; van Vliet, A.; Pozniak, J.; Hummelink, K.; Terry, A.M.; Monkhorst, K.; Sanders, J.; Hofland, I.; Landeloos, E.; Van Herck, Y.; et al. SERPINB9 is commonly amplified and high expression in cancer cells correlates with poor immune checkpoint blockade response. Oncoimmunology 2022, 11, 2139074. [Google Scholar] [CrossRef]
- Budanov, A.V.; Karin, M. p53 target genes sestrin1 and sestrin2 connect genotoxic stress and mTOR signaling. Cell 2008, 134, 451–460. [Google Scholar] [CrossRef]
- Tisdale, M.J. Cancer cachexia: Metabolic alterations and clinical manifestations. Nutrition 1997, 13, 1–7. [Google Scholar] [CrossRef]
- Goldman, A.; Majumder, B.; Dhawan, A.; Ravi, S.; Goldman, D.; Kohandel, M.; Majumder, P.K.; Sengupta, S. Temporally sequenced anticancer drugs overcome adaptive resistance by targeting a vulnerable chemotherapy-induced phenotypic transition. Nat. Commun. 2015, 6, 6139. [Google Scholar] [CrossRef]
- Puchalska, P.; Crawford, P.A. Multi-dimensional Roles of Ketone Bodies in Fuel Metabolism, Signaling, and Therapeutics. Cell Metab. 2017, 25, 262–284. [Google Scholar] [CrossRef] [PubMed]
- Weinberg, S.E.; Chandel, N.S. Targeting mitochondria metabolism for cancer therapy. Nat. Chem. Biol. 2015, 11, 9–15. [Google Scholar] [CrossRef] [PubMed]
- Tsvetkov, P.; Coy, S.; Petrova, B.; Dreishpoon, M.; Verma, A.; Abdusamad, M.; Rossen, J.; Joesch-Cohen, L.; Humeidi, R.; Spangler, R.D.; et al. Copper induces cell death by targeting lipoylated TCA cycle proteins. Science 2022, 375, 1254–1261. [Google Scholar] [CrossRef] [PubMed]
- Shaffer, S.M.; Dunagin, M.C.; Torborg, S.R.; Torre, E.A.; Emert, B.; Krepler, C.; Beqiri, M.; Sproesser, K.; Brafford, P.A.; Xiao, M.; et al. Rare cell variability and drug-induced reprogramming as a mode of cancer drug resistance. Nature 2017, 546, 431–435. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.