Abstract
Bacteria play an important role in addressing challenges in rice production by promoting plant growth and enhancing stress tolerance through multiple mechanisms. Different types of soil bacteria affect rice growth by improving nutrient absorption, managing stress, and enhancing root structure. The relationship between rice plants and bacteria is intricate, as these bacteria can help reduce problems like salt stress, heavy metal toxicity, and infections. This review summarises studies published up to 2025 on how bacteria influence rice roots, including aspects like root length, density, biomass, and volume. Bibliometric analysis shows an increase of over 900% in research interest after 2020, with most studies conducted under controlled conditions and limited field validation. In addition to identifying key bacterial groups such as Bacillus, Pseudomonas, Burkholderia, and Azospirillum, this review identifies research gaps related to context dependency, strain specificity, and scalability. We have also emphasised the need for multi-strain inoculation strategies, field-scale experiments, and integration of microbial selection with rice breeding. The synthesis has highlighted that bacterial strains do not simply stimulate root growth but actively reprogram rice root architecture, modulating elongation, branching, density, and surface area as a response to environmental constraints. These effects are mediated by interconnected mechanisms that include phytohormone production, nutrient solubilisation, deaminase activity, stress-related gene regulation, and microbiome-driven feedback involving root exudation. Overall, viewing bacteria as regulators of root developmental dynamics rather than simple biofertilisers provides new insights for designing climate-adapted and sustainable rice production systems.
1. Introduction
Rice (Oryza sativa L.) is a key food source for nearly half of the world’s population, particularly in Asia, Latin America, and Africa, with Asia accounting for over 90% of global rice production [1,2]. Given its nutritional value, including essential amino acids, vitamins, minerals, fibres, and carbohydrates, most rice is cultivated for human consumption, with little production allocated to other uses [3,4]. To meet the needs of a growing population, global cereal production, including rice, needs to rise to around three billion tons by 2050 [5].
To secure rice yields and food security, farmers have traditionally relied on chemical fertilisers to supply essential nutrients such as nitrogen, phosphorus, and potassium [6]. However, excessive use of synthetic fertilisers has led to soil degradation, reduced fertility, and adverse environmental impacts [7]. This highlights the urgent need for more sustainable farming practices. Approaches such as organic and inorganic amendments (e.g., cow manure, nitrogen, and potassium), improved breeding programmes, and enhanced crop and water management practices have been explored to boost rice productivity [8,9,10,11,12].
In recent years, there has been a growing interest in using beneficial bacteria to support rice growth and yield [13,14]. These plant growth-promoting rhizobacteria (PGPR) can enhance nutrient uptake, modulate stress responses, and influence root architecture, offering a more environmentally friendly alternative to conventional approaches. More broadly, over the past decade, plant–microbiome research has increasingly framed microbes not only as biofertilisers but also as regulators of plant development and root system plasticity. However, this conceptual perspective has mostly been developed at general or cross-crop levels, with limited system-focused synthesis for rice. Rather than proposing a wholly new conceptual paradigm, this review provides a rice-specific integrative synthesis that reorganises dispersed experimental evidence on bacterial effects on root traits into a coherent interpretative framework centred on root architecture modulation and functional plant responses. In this sense, the contribution is primarily synthetic and system-focused, while offering a refined interpretative lens for rice-based systems. Accordingly, the novelty of this review lies not in introducing the idea that microbes can regulate plant root traits per se, but in systematically compiling and critically interpreting how distinct bacterial strains and consortia influence rice root development and associated functional outcomes across experimental contexts. Nonetheless, despite extensive research on rice–microbe interactions, no comprehensive overview has specifically examined how different bacterial strains influence root traits and overall plant health.
This review addresses the following key research questions: (i) What are the predominant bacteria that influence rice root traits and plant growth? (ii) Through which biological mechanisms do these bacteria impact nutrient uptake, stress tolerance, and growth? (iii) What are the gaps and contradictions in current knowledge that require further investigation? To address these questions, this review brings together and looks at peer-reviewed studies on how bacteria affect rice, focusing on root-level effects. It also discusses new trends, points out limitations in current research, and suggests future directions to help both basic understanding and practical applications.
2. Data Collection and Processing
This systematic review was carried out according to the PRISMA 2020 guidelines [15] to ensure transparency and reproducibility in the article selection and analysis. The related steps were the following (Table A1):
- (1)
- Search strategy: in November 2025, a comprehensive search was performed on three bibliometric databases: Scopus®, Web of Science®, and Google Scholar®. The following keywords and their combinations were used to select relevant literature: “rice root”, “bacteria”, “root morphology”, “bacterial strains”, and “rhizosphere”. No restrictions were applied regarding publication years to include all potentially relevant studies up to date;
- (2)
- Inclusion criteria: (a) Articles in English; (b) Original research articles about the interaction between bacteria and rice root systems, including laboratory, greenhouse, or field studies; (c) Studies investigating the impact of bacterial strains on rice root morphology or rhizosphere characteristics; (d) studies dated 1995 to present;
- (3)
- Exclusion criteria: (a) Duplicate articles in the aforementioned databases; (b) Non-English publications; (c) Reviews, meta-analyses, editorials, conference abstracts, book chapters or articles not directly related to the topics;
- (4)
- Screening process: An initial screening was carried out by reviewing titles and abstracts for relevance. Articles that did not meet the inclusion criteria were excluded. Full-text versions of potentially eligible articles were then downloaded and evaluated for eligibility according to the criteria reported above;
- (5)
- Data extraction and quality assessment: For the selected studies, data were extracted according to the following parameters: publication year, country of study, publishing journal, number of citations, and bacterial strains investigated. These data were used for both bibliometric and critical analysis, and discussion of the literature. While formal bias assessment tools were not applied, studies were evaluated for relevance and quality during the analysis; citations were retrieved from the Scopus database, and self-citations were included;
- (6)
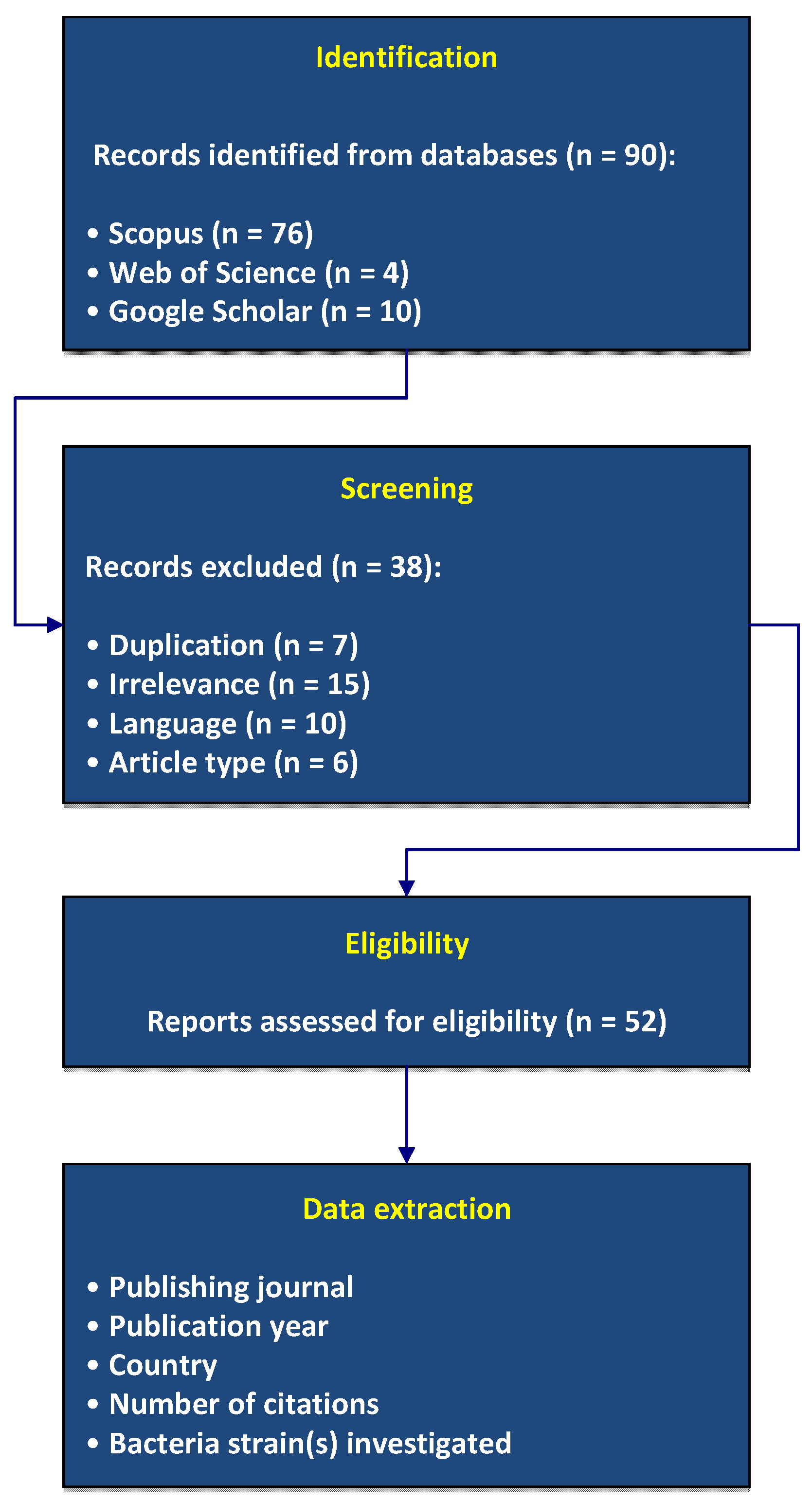
- Study selection summary: The search initially found 90 records. After removing 17 duplicates or non-English articles, 73 articles remained for full screening. Of these, 15 were general papers (without a specific focus on bacteria or rice), 6 were other reviews, while the 52 studies that specifically investigated bacterial impacts on rice plants were finally included.
Figure 1 presents the PRISMA 2020 flowchart detailing the selection process.
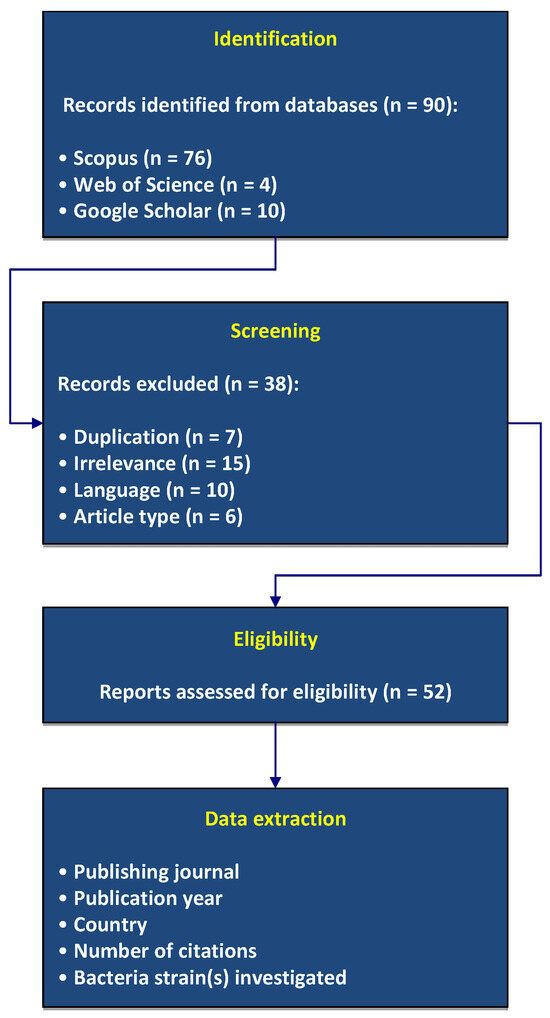
Figure 1.
Flow-chart of this review following the PRISMA 2020 guidelines.
The scope of the bibliometric analysis was descriptive, aimed at contextualising publication trends rather than performing network or citation-structure inference. A formal risk-of-bias assessment was not applied because this review aims to provide a structured, PRISMA-guided synthesis of heterogeneous experimental studies with diverse designs and outcomes, for which standardised bias assessment tools are not readily applicable.
3. Results of the Bibliometric Analysis
The selected papers were published in 42 journals. Ten journals published 38.5% of the total papers, while the remaining papers were included in 32 other journals (Table 1).

Table 1.
Distribution of the analysed papers by publishing journals.
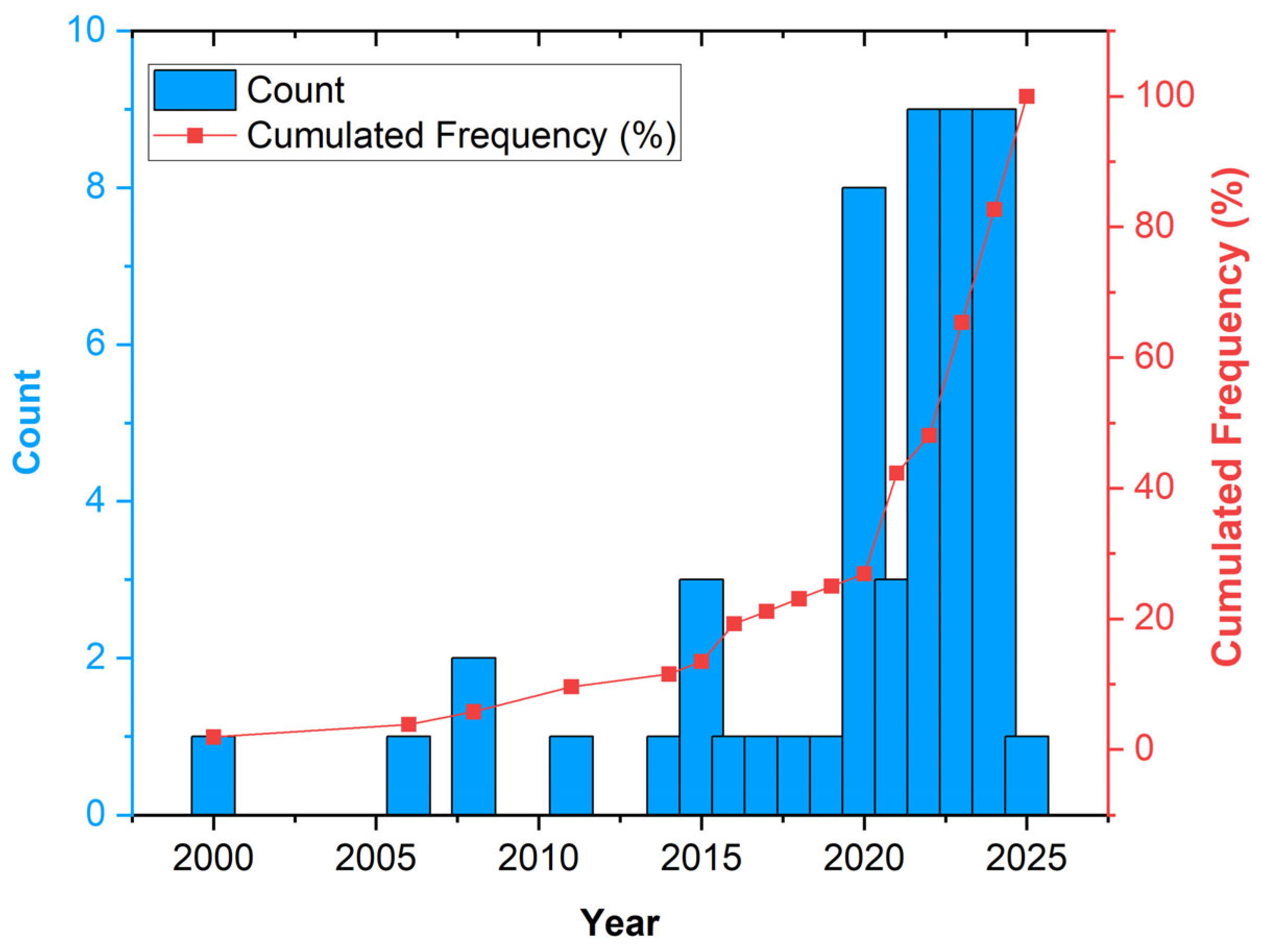
We noted an increasing interest in this topic in recent years. The highest number of papers (27, more than 50% of the total) was published between 2022 and 2024, and 8 (15%) papers were published in 2020 (Figure 2).
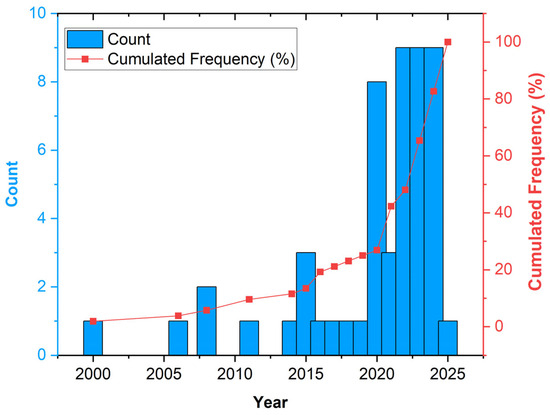
Figure 2.
Distribution of the analysed papers by publication year.
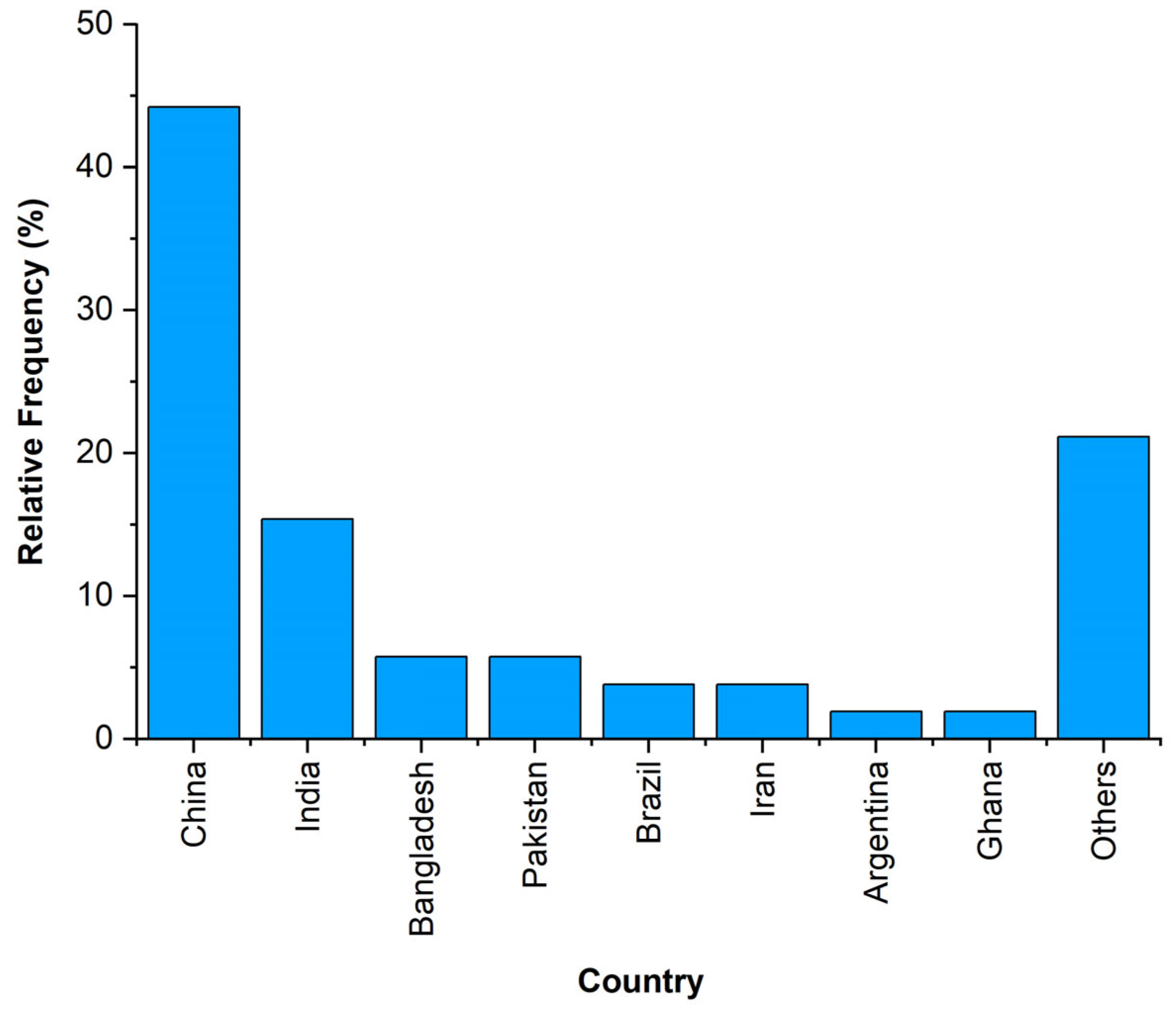
The selected papers contain case studies carried out in China (23 papers, 44% of the total) and India (8, 15%), both accounting for more than 50% of the total number (Figure 3).
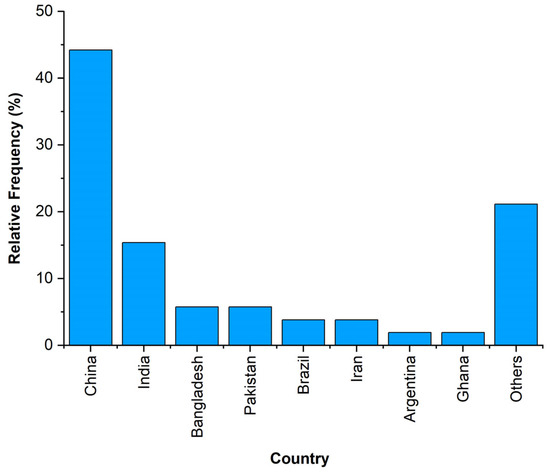
Figure 3.
Distribution of the analysed case studies by countries.
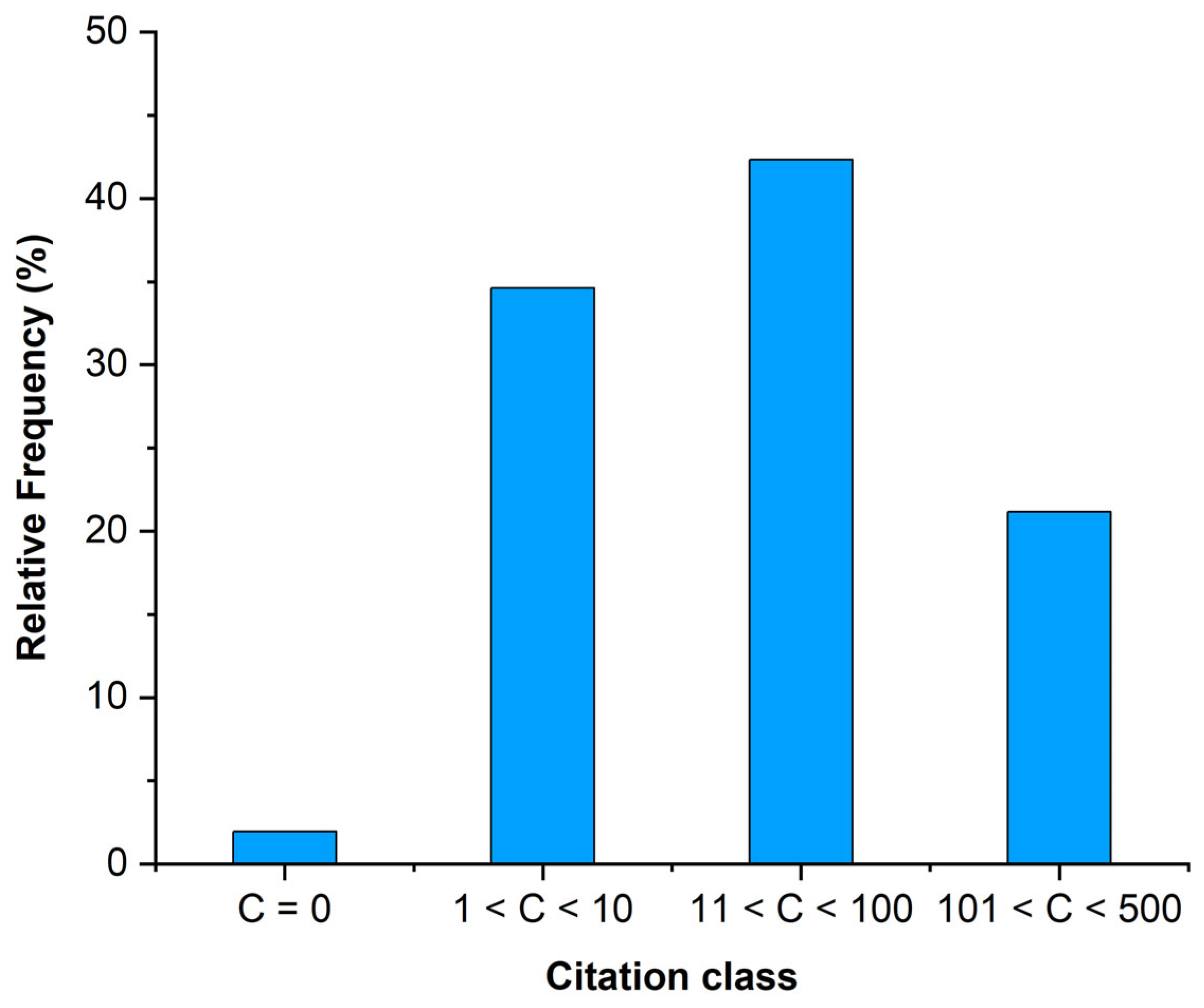
The analysed papers totalled 3464 citations (update 25 November 2025). Thirty-three (63% of the total) papers achieved over 10 citations, while 11 papers (21%) were in the class of 101 to 500 citations (Figure 4). Two papers [16,17] had more than 300 citations.
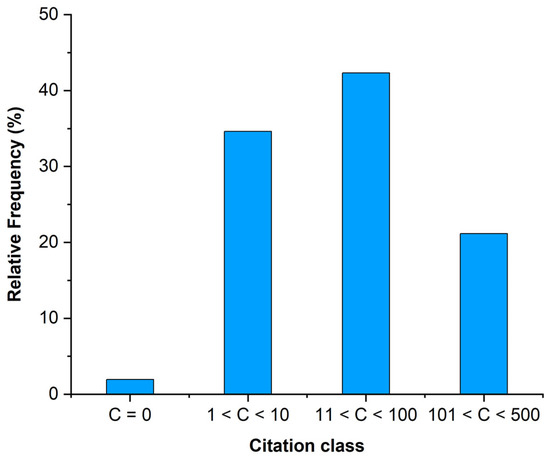
Figure 4.
Citation class of the analysed papers based on Google Scholar.
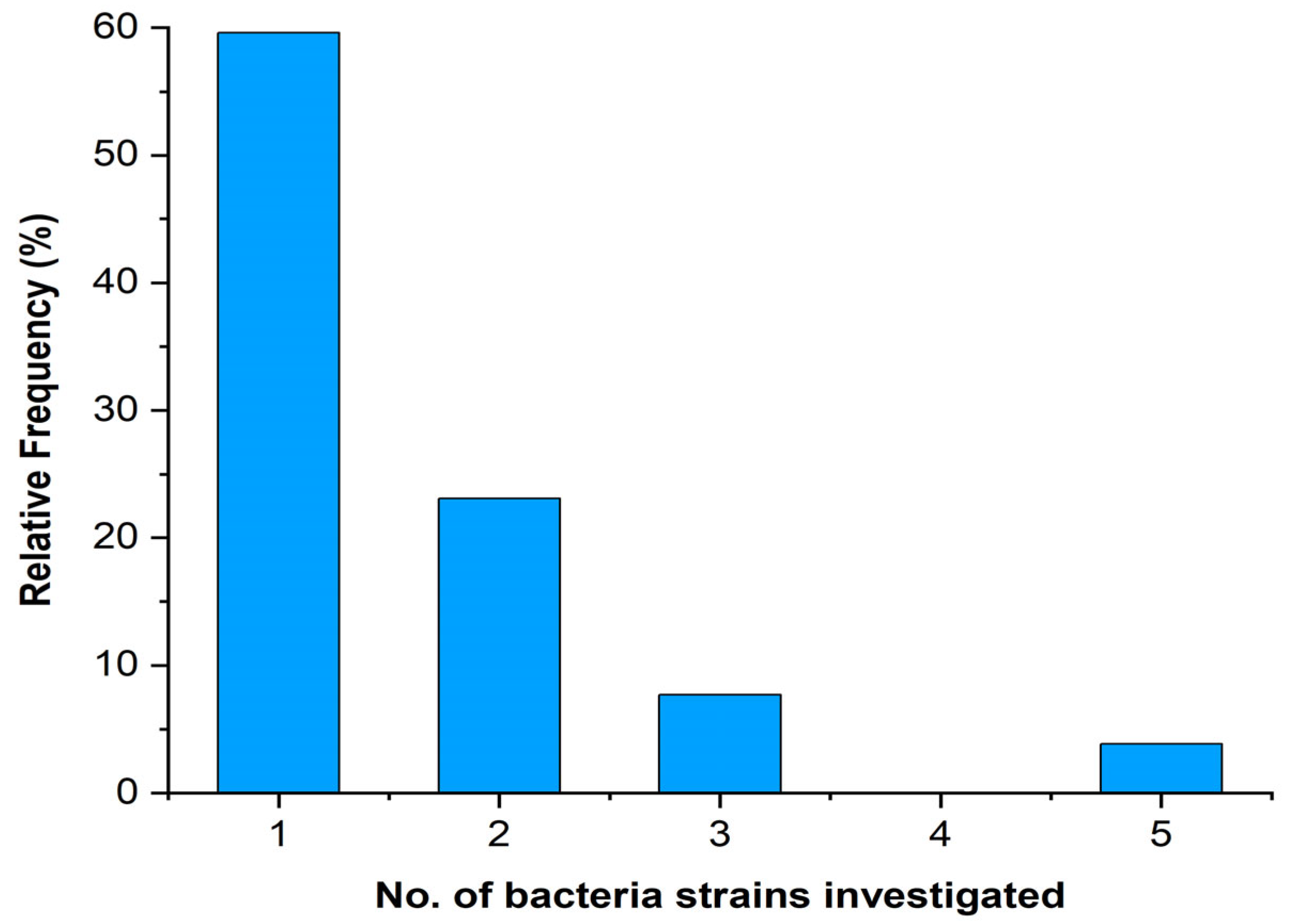
Three studies investigated more than five bacterial strains, while most papers (31, 60% of the total) studied only one bacterium (Figure 5).
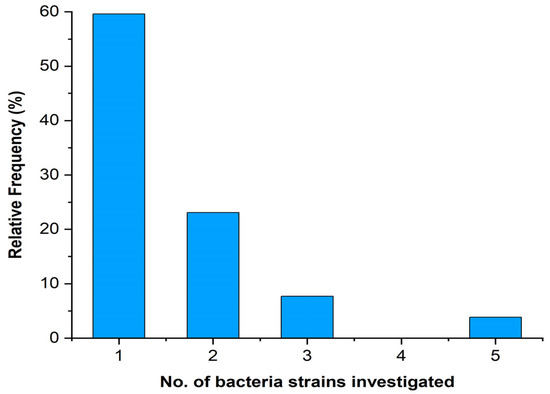
Figure 5.
Number of bacterial strains investigated in the analysed papers.
4. Discussion
Table 2 reports the beneficial effects of bacteria on rice from the reviewed studies. The analysed investigations have focused on bacterial impacts on disease resistance, stress tolerance, nutrient absorption, and rice growth and yield, often with synergistic effects. The evidence summarised in Table 2 is discussed below by distinguishing primary mechanisms of bacterial action, resulting plant phenotypic responses, and broader ecological and agronomic implications.
4.1. Bacteria Effects on Rice Plants
A wide range of microorganisms, particularly bacteria, play a key role in rice growth [18,19,20,21,22]. These bacteria help rice by improving how plants absorb nutrients and supporting their growth in the paddies [18,21]. For instance, Burkholderia, Pseudomonas, Bacillus, and Azospirillum were found to noticeably improve rice growth and yield [23,24,25,26]. An increasing number of studies show how these bacteria interact with rice roots to form beneficial networks that are vital for plant growth [27,28,29,30].
4.1.1. Bacterial Effects on Abiotic Stress Tolerance
Several bacterial strains can enhance rice tolerance to adverse environmental conditions. For example, Sun et al. [31] reported that exopolysaccharides of Pantoea alhagi NX-11 increased drought resistance by enhancing fresh weight, chlorophyll content, proline, soluble sugars, and relative water content of rice seedlings. Halophytic Bacillus endophytes increased rice tolerance to salinity by producing phytohormones [24]. Burkholderia and Azospirillum strains reduce oxidative damage induced by NaCl stress and regulate stress-responsive genes, promoting rice growth under saline conditions [32,33]. Additionally, Bacillus amyloliquefaciens SN13 enhances tolerance to multiple abiotic stresses, including drought and extreme temperatures [34,35].
4.1.2. Bacterial Effects on Nutrient Absorption
Bacteria play a significant role in improving nutrient uptake by rice plants. Pseudomonas species improve root structure and increase nutrient availability, particularly nitrogen, phosphorus, and iron, supporting better growth [36]. Some bacteria even help with the efficiency of phosphorus use [37]. Improvements in aeration of the rhizosphere alter microbial communities, enhancing phosphorus transformation [38]. Combined inoculations with Burkholderia and diazotrophs have increased rice yield by up to 23.6%, in part thanks to improved nitrogen fixation [16]. Moreover, Acinetobacter and Serratia species promote zinc solubilisation, which enhances root colonisation and plant biomass [39]. Some Bacillus strains can immobilise toxic cadmium in soils, reducing metal uptake and promoting plant health [40]. Azospirillum brasilense Az39, the type strain of the novel species Azospirillum argentinense sp. nov., exhibits nitrogenase activity and indole-3-acetic acid production, which are important traits for promoting plant growth [41].
4.1.3. Bacterial Effects on Disease Resistance and Biocontrol
Plant growth-promoting rhizobacteria (PGPR) are essential in protecting rice against pathogens. Burkholderia species exhibits antagonistic activity against pathogenic fungi, and Pseudomonas strain EA105 reduces rice blast lesions [42]. Strains such as Bacillus thuringiensis enhance defence gene expression and systemic resistance [43]. Bacillus atrophaeus FA12 and B. cabrialesii FA26 show broad-spectrum antagonistic activity while promoting rice growth [44]. The synergistic action of bacteria, such as Sphingomonas paucimobilis, further increases disease resistance and stress tolerance [34]. Some endophytic bacteria can reduce the toxicity of heavy metals (e.g., cadmium) [45,46].
4.1.4. Bacterial Effects on Specific Biological Mechanisms
In addition to the reported growth promotion and stress tolerance, bacteria influence specific biological activities in rice plants. For instance, Mycobacterium, Streptomyces, Anaeromyxobacter, and Geotalea species play a key role in silicon-mediated arsenic absorption by roots, contributing to detoxification [47]. Improved oxygen conditions in paddy fields due to aeration increase the content of soil phosphorus in the rhizosphere and regulate the abundance and composition of P-solubilising microorganisms, therefore promoting its transformation to plant-available phosphorus [38]. Azospirillum inoculation biologically increases production of plant biomass and develops the root system (particularly, length and density as well as the number of lateral roots and root hairs), and, finally, the rice yield [48]. Production of 1-aminocyclopropane-1-carboxylate (ACC) deaminase by some strains reduces ethylene levels in plants, supporting stress tolerance and plant development [49]. Moreover, rhizobacteria modulate gene expression related to nutrient uptake and stress responses [50,51,52,53]. Rhizobial bacteria enhance the phytoremediation of heavy metals by legumes, highlighting their potential to improve metal uptake and detoxification of contaminated soils [54].
4.2. Importance of Bacterial Communities in Rice Root Growth
4.2.1. Central Role of Root Traits in Rice Productivity
Rice roots are vital for rice plant growth and production, especially under adverse environmental conditions such as drought and heat stress, since roots form the main interface between the plant and its surrounding environment [55]. Among the analysed characteristics, root length was the most frequently studied trait. However, this predominance likely reflects methodological convenience rather than exclusive biological relevance. Root system architecture is inherently multidimensional and includes branching topology, lateral root density, root angle, diameter, surface area, turnover, and plasticity dynamics, many of which are more functionally informative but less frequently quantified. This emphasises the central role of root length in enhancing nutrient uptake and overall plant productivity. Length is a key aspect of root morphology, since this trait is particularly important for the efficient absorption of nutrients (especially phosphorus) [56]. Nevertheless, reliance on root length alone may oversimplify bacterial effects and potentially overestimate functional improvement when other architectural traits are not concurrently measured. Under stress conditions, rice, like other crops, can adapt to adverse environments by modifying root architecture, primarily through elongation, to improve nutrient absorption [57]. Future studies should therefore adopt multidimensional root phenotyping approaches rather than single-metric indicators.
4.2.2. Bacterial Promotion of Root Growth and Morphology
Bacterial communities significantly promote rice yield by enhancing water and nutrient uptake, which, in turn, supports the development of root traits, such as length, density, and volume [58]. The interaction between bacteria and rice roots has gained increasing attention from researchers in recent years. Several studies have stated the beneficial effects of many bacteria on root development [59,60,61]. For instance, Islam [62] found that inoculation with strains of Paraburkholderia and Delftia significantly increased the root length of rice seedlings in the short term (very few days). These strains promote root elongation and improve seed germination and yield, indicating their potential as eco-friendly alternatives to synthetic fertilisers. A direct correlation between rice growth duration and root characteristics (i.e., length and growth rate) was also reported [63]. The findings of the latter study show that extended growth periods—an important driver of rice yield [64,65]—are associated with greater root elongation.
4.2.3. Bacterial Mitigation of Heavy Metal Toxicity and Stress in Roots
In addition to promoting the development of root morphology, some beneficial bacteria are also able to mitigate the toxicity of heavy metals in rice roots. For example, silica-impregnated biochar composites enhance microbial activity in the rhizosphere of rice plants, favouring the immobilisation of arsenic and reducing its uptake [66]. Several studies, reported in Table 2, show that inoculation with various bacterial strains enhances root traits (e.g., length, density, volume, biomass), and reduces the accumulation of toxic metals. For instance, Yanni [67] and Beneduzi [68] reported that endophytic strains, such as Rhizobium leguminosarum bv. trifolii and Bacillus sp. (SVPR30 isolate), significantly promote root growth. Similarly, length, volume, and biomass of roots noticeably increase after treatments with plant growth–promoting bacteria [69]. Moreover, bacterial inoculation can be effectively used to enhance the root length of rice [61,70]. Bacterial strains Herbaspirillum sp. R3 and Bacillus cereus T4 improve root morphology and support cadmium uptake, which clearly shows their dual role in growth promotion and bioremediation [71]. Finally, the Piriformospora indica functional microbe is able to increase root length and enhance cold tolerance in rice [72].
4.2.4. Microbiome-Induced Effects on Root Architecture
Some studies have also focused on microbiome-induced effects on rice root architecture beyond growth promotion. In this regard, most papers report bacterial effects on rice roots as increases in length or biomass. This reporting pattern reinforces the measurement bias noted above and may obscure more complex architectural effects. However, the reviewed literature indicates that bacteria actively reprogram root architectural strategies rather than merely stimulating growth [48,68]. Several bacterial strains modulate density, lateral initiation, and surface expansion of roots, leading to functionally optimised systems under different soil constraints [73]. Auxin-producing and ACC deaminase-producing bacteria shift priorities for development by altering ethylene signalling pathways, thus influencing root branching patterns instead of uniform elongation [34,49]. Under nutrient limitation, bacteria-induced elongation enhances soil penetration, while, under salinity or heavy metal stress, altered root density and surface traits limit toxic ion uptake [40,53]. It is worth noting that these architectural changes are strongly context-dependent and vary across soil chemistry, flooding regime, and rice genotype [16,24]. Accordingly, bacterial effects should be interpreted as regulatory and plastic responses rather than additive growth increments. This variability further indicates that bacterial effects on roots are regulatory rather than additive. Rice roots thus act as adaptive systems whose developmental programmes are dynamically shaped by microbial cues [27].

Table 2.
The bacterial strains’ impacts on the growth and yield of rice and their contributions to improved rice root characteristics.
Table 2.
The bacterial strains’ impacts on the growth and yield of rice and their contributions to improved rice root characteristics.
| Reference | Bacterial Strains | Mechanisms | Experimental Context | Inoculation Type |
|---|---|---|---|---|
| [16] | Burkholderia vietnamensis | Efficient colonisation of rice roots and N2 fixation Root surface area and root dry weight increased by ~20–30% following bacterial inoculation | Greenhouse | Single strain |
| [17] | Burkholderia | Prevention of infections in young plants | – | – |
| [74] | Pseudomonas EA105 | Inhibition of growth and appressoria formation; reduced blast lesions | Laboratory | Single strain |
| [49] | Pseudomonas stutzeri A1501 | Shoot and root biomass increased by ~20–35% under salt and heavy-metal stress; ACC deaminase activity significantly reduced stress-induced ethylene levels (quantified enzymatic activity reported). | Laboratory | Single strain |
| [39] | Acinetobacter sp., Serratia sp. | Plant height, root length, and total biomass increased by ~15–30%; chlorophyll content increased by ~18–23% compared with non-inoculated controls; Zn solubilisation Increased root volume and length | Greenhouse | Consortium |
| [54] | Bradyrhizobium sp. 750 | Increased growth under heavy metal stress | Greenhouse | Single strain |
| [34] | Bacillus amyloliquefaciens SN13 | Shoot biomass increased by ~22–28%; relative water content increased by ~18–22%; proline content increased by ~30–40% under salinity stress. | Laboratory | Single strain |
| [75] | B. licheniformis A21, B. haynesii EN43, Paralicheniformis EN107 | Grain yield increased by ~18–30%; shoot and root biomass increased by ~20–25%; Superoxide Dismutase (SOD), Catalase (CAT), and Guaiacol peroxidase (GPOX) activities decreased by ~15–30% under drought stress | Field | Consortium |
| [48] | Azospirillum amazonense | Number of panicles increased by ~20–28%; grain dry matter accumulation increased by ~18–24%; nitrogen uptake increased by ~15–22% Increased root length, lateral roots, hairs, and volume | Field | Single strain |
| [40] | Bacillus strains NMTD17, GBSW22, LLTC96 | Cadmium concentration in rice roots reduced by ~30–50%; shoot and root biomass increased by ~20–30%, depending on strain Reduced cadmium concentration in roots | Greenhouse | Consortium |
| [41] | Azospirillum amazonense | Nitrogenase activity and IAA production | Laboratory | Single strain |
| [43] | Bacillus thuringiensis GBAC46, NMTD81 | Nematode control via crystal proteins Increased root length | Laboratory | Consortium |
| [44] | Bacillus atrophaeus FA12, B. cabrialesii FA26, velezensis FZB42 | Antagonistic activity and growth promotion Increased root length | Laboratory + Greenhouse | Consortium |
| Bacillus | ||||
| [76] | Rhodopseudomonas palustris + Bacillus subtilis | Grain yield increased by ~10–18%; relative abundance of beneficial rare taxa increased by >25%, with significant enrichment of membrane-transport and signalling pathways | Greenhouse | Consortium |
| [35] | Bacillus spp. (FZB42, NMTD17, LLTC93, GBSC56, SYST2, IHBT-705 | Heat tolerance and abiotic stress resilience Increased root length | Greenhouse | Consortium |
| [77] | Sphingomonas paucimobilis | Tolerance to salt, drought, and cadmium stress | – | Single strain |
| [37] | Pseudomonas | Phosphorus uptake and use efficiency increased by ~25–40%, associated with increased recruitment of Pseudomonas spp. | Field | – |
| [47] | Mycobacterium, Streptomyces, Geotalea, Anaeromyxobacter | Si-regulated arsenic absorption in roots | Field | Consortium |
| [38] | Multiple phyla (e.g., Actinobacteria, Proteobacteria) | P transformation and enhanced soil P availability | Field | Consortium |
| [50] | Chryseobacterium sp., Ochrobactrum sp. | Root length increased by ~18–25%; root dry matter increased by ~15–20% | Greenhouse | Consortium |
| [51] | Serratia nematodiphila | Root surface area and root biomass increased by ~15–25%, depending on the microbial consortium | Greenhouse | Single strain |
| [52] | B. altitudinis FD48, B. methylotrophicus RABA6 | Root–shoot ratio increased by ~20–25% under moisture stress conditions | Greenhouse | Consortium |
| [53] | Bacillus marisflavi, Pantoea stewartii A | Reduced sodium content in roots | Greenhouse | Consortium |
| [61] | Enterobacteraero genes | Increased root length | Greenhouse | Single strain |
| [66] | Bacillus and Geobacter | Arsenic concentration in rice roots reduced by ~35–55% due to microbe-mediated immobilisation | Field | Consortium |
| [67] | Rhizobium leguminosarum bv. trifolii | Root dry weight and length increased by ~20–25% compared with uninoculated plants | Greenhouse | Single strain |
| [68] | Bacillus sp., Paenibacillus sp. | Root length increased by ~15–22% in inoculated rice plants | Greenhouse | Consortium |
| [69] | Rhizobium BU Ls 28 | Root length increased by ~25–35%; root volume increased by ~30–38%; dry biomass increased by ~20–25% | Greenhouse | Single strain |
| [70] | Bacillus spp. GBSC56, SYST2, FZB42 | Increased root length | Greenhouse | Consortium |
| [71] | Burkholderia and Acidovorax | Reduced cadmium concentration in roots | Field | Consortium |
| [72] | Piriformospora indica | Root length increased by ~20–30% under cold stress following inoculation with Piriformospora indica | Greenhouse | Single strain |
| [73] | ACC deaminase-producing bacteria | Root length increased by ~25–35%; fresh and dry biomass increased by ~20–28% under salt stress | Laboratory | Consortium |
4.2.5. Root Exudation and Self-Reinforcing Rhizosphere Feedbacks
Recognising these microbiome-induced effects highlights bacteria as developmental regulators, with important implications for field-scale predictability. To avoid overlap with previously described mechanisms, we focus here specifically on rhizosphere feedback dynamics rather than general growth promotion. From the reviewed studies, it emerges that bacterial shaping of root exudation is a driver of self-reinforcing functionality of the rhizosphere. Although rarely measured directly, root exudation emerges as a central mechanism that underlies bacterial effects on rice roots across multiple studies [30,37]. Bacteria influence plant carbon stock and metabolic pathways, resulting in qualitative changes in the composition of root exudates, such as organic acids, sugars, and amino acids [51]. These altered exudates selectively recruit beneficial microbial taxa, which reinforce functional communities in nutrient solubilisation and stress mitigation [62,76]. This feedback loop helps explain variability in inoculant performance across environments and supports a systems-level interpretation of bacteria–root interactions. In flooded rice systems, such selective recruitment stabilises phosphorus-solubilising and nitrogen-fixing populations that directly enhance root nutrient acquisition [37,38]. Organic acid exudation promoted by bacteria increases the availability of phosphorus and micronutrients, while changes in sugar profiles regulate microbial competition close to the root surface [36]. This feedback helps explain why some bacterial inoculants maintain long-term effectiveness, while others fail under field conditions [69,75]. Soil heterogeneity and redox conditions further modulate these interactions by driving diffusion and microbial turnover [78]. Thus, bacteria reshape not only roots but also the biochemical dialogue governing rhizosphere stability.
4.2.6. Bacterial Mediation of Root–Soil Interfaces in Flooded Systems
Bacteria also mediate root–soil interfaces under redox and metal stress in paddy ecosystems, as shown by several studies. Flooded rice cultivation exposes roots to fluctuation in redox conditions that heavily influence nutrient availability and metal mobility [38,47]. The reviewed literature demonstrates that bacteria act as critical mediators at the root–soil interface by buffering chemical instabilities associated with these environments [47,66]. Some bacterial taxa promote iron plaque formation and biofilm-like structures on root surfaces, modifying ion exchange processes and limiting arsenic and cadmium uptake [66]. At the same time, silicon-regulated microbial communities indirectly control metalloid transport into root tissues [47]. These processes are closely linked to root morphology, as thicker roots and modified epidermal layers exhibit distinct microbial colonisation patterns [51]. Anaerobic-tolerant bacteria sustain nutrient cycling under prolonged flooding, maintaining nitrogen and phosphorus availability when oxygen diffusion is restricted [14,38]. This microbial mediation allows rice roots to sustain metabolic activity despite extreme chemical fluctuations. The co-adaptive nature of these interactions explains the heightened importance of bacteria in paddy soils compared with upland systems.
4.2.7. From Single-Strain Inoculation to Predictive Root–Microbiome Techniques
A critical synthesis of the reviewed studies reveals inherent limitations of single-strain bacterial inoculation strategies [24,75]. Although many strains perform well under controlled conditions, their effectiveness is inconsistent across soils, climates, and management practices [16,17]. This inconsistency highlights the need for precision microbial design targeting specific root functional traits rather than general growth promotion [21]. To operationalise context dependency, we propose a simple selection framework linking environmental and host factors to microbial functional traits and experimental design choices: (i) under nutrient limitation, prioritise strains with documented solubilisation and mobilisation traits and measure multidimensional root foraging architecture; (ii) under salinity or metal stress, prioritise ACC-deaminase and osmoprotectant-associated strains and quantify branching density and surface traits rather than length alone; (iii) under flooded or redox-variable systems, prioritise anaerobic-tolerant and biofilm-forming taxa and measure root–soil interface traits; (iv) across genotypes, explicitly test host × microbe interactions using factorial designs. Such a framework translates descriptive context dependency into testable and comparable experimental strategies. Genomic and transcriptomic analyses have identified bacterial genes involved in hormone modulation, stress signalling, and nutrient mobilisation that directly influence root developmental pathways [32]. Combining strains with complementary functions may enable targeted manipulation of root depth, density, and longevity [62]. In addition, inoculation timing is increasingly recognised as critical, as bacterial influence on roots varies across growth stages [48]. Integrating microbial selection with rice breeding programmes could align host genetic potential with microbial functionality [55]. This shift supports a transition from descriptive bio-inoculation studies toward predictive, design-oriented root–microbiome approaches, moving from empirical biofertilisation toward predictive root–microbiome techniques.
4.3. Replies to the Research Questions
Based on the reviewed articles and thematic analysis, the following responses address the research questions formulated in the introduction. First, as a reply to the first question (‘What are the predominant bacteria that influence rice root traits and plant growth?’), the study has shown that several bacterial strains emerge as key drivers of rice growth, health, and resilience. The most frequently reported bacterial genera with significant positive effects on rice roots and overall plant performance include Pseudomonas, Bacillus, Burkholderia, and Azospirillum [17,23,24,25,26,36]. These genera are well studied for their roles in promoting growth, enhancing nutrient availability, and improving stress tolerance. For example, Pseudomonas enhances root architecture and nutrient uptake [36]. Burkholderia and Azospirillum have demonstrated the capacity to alleviate salinity stress and improve yield [24,32]. Additional genera, such as Acinetobacter and Serratia [39], Streptomyces [47], Sphingomonas [77], and Mycobacterium [47], have also been identified as beneficial in more specialised functions, such as solubilisation and biocontrol of micronutrients.
While phytohormone-related mechanisms (notably indole-3-acetic acid production and ACC deaminase activity) are frequently reported, their effects appear highly context-dependent. We argue that root architectural modulation primarily emerges from the convergence of multiple bacterial traits rather than from single mechanisms acting in isolation. In particular, strains combining hormone modulation, nutrient solubilisation, and stress-alleviation pathways consistently promote the formation of lateral roots and their surface expansion. Accordingly, bacterial efficacy should be interpreted as a functional syndrome rather than as the sum of individual traits.
In reply to the second question (‘Through which biological mechanisms do these bacteria impact nutrient uptake, stress tolerance, and growth?’), the reviewed literature has indicated that bacterial strains enhance rice growth by several interconnected mechanisms. The key pathways involved are: (i) Nutrient solubilisation and mobilisation (several strains solubilise phosphorus [37] and zinc [39], and fix atmospheric nitrogen [16], improving nutrient availability and uptake); (ii) Phytohormone production (several strains, such as halophytic Bacillus spp., produce phytohormones (e.g., auxins), which promote elongation and adaptation of roots to abiotic stress [24,35]; (iii) Induced systemic resistance and gene modulation (strains, such as Bacillus thuringiensis, stimulate defence gene expression, reducing pathogen load [43,44]; (iv) Stress mitigation via osmoregulation and antioxidant activity (Pantoea strains produce exopolysaccharides, which increase levels of proline and soluble sugars, thus enhancing drought tolerance [31], while Burkholderia and Azospirillum reduce oxidative damage in environments subjected to salinity stress [32,33]; (v) Rhizosphere drivers and microbial recruitment (bacteria can influence root exudation activities and modify microbial communities, thereby indirectly enhancing nutrient cycling and root health [37,38]. From a synthetic perspective, bacterial effects on plant growth should be viewed as emergent properties of dynamic plant-microbe-soil feedback rather than as additive trait effects (e.g., N fixation + IAA production), but rather as emerging properties of dynamic plant-microbe-soil feedback. The same bacterial strain may activate different mechanisms depending on soil texture, nutrient status, plant developmental stage, and stress intensity. Consequently, future research should explore integrative experimental designs that combine molecular, physiological, and soil-scale measurements, rather than focusing on single traits in isolation.
4.4. Limitations of Current Knowledge
Several critical gaps and contradictions in current knowledge require further investigation, and this aspect is the core of the third research question (‘Through which biological mechanisms do these bacteria impact nutrient uptake, stress tolerance, and growth?’). To distinguish evidence limitations from mechanistic interpretation, we group the main constraints into experimental, mechanistic, and scaling gaps. Although the body of literature is extensive, several issues remain unresolved. The comparative evaluations of bacterial strains under comparable experimental conditions are few, and the generalisation of their effectiveness across environments is difficult [24,75]. Contradictory findings (e.g., variable performance of the same genus in different soil types or climates) suggest a need for more investigations in specific contexts [17,23]. Moreover, this review has highlighted that most studies about the beneficial effects of bacteria on rice have been carried out in the laboratory (e.g., [52,69,75,76]). Field studies remain limited compared with laboratory experiments conducted under controlled conditions, reducing confidence in the general results [16]. This imbalance also contributes to trait-selection bias, including the dominance of easily measurable variables such as root length. However, these experiments may not accurately reflect the complex and dynamic conditions of the field experiments and therefore limit the generalisability of results. This challenge is particularly pronounced in studies addressing plant–soil–microorganism interactions. Particularly, transferring information in the field of the relationship between soil microorganisms and plants shows one of the important challenges facing environmental scientists. Finally, molecular mechanisms (e.g., hormone signalling and gene regulation) are often cited, but not always thoroughly investigated or validated; this gap indicates the need for more omics-based research to clarify mode-of-action pathways [49,50]. Long-term ecological impacts and consistency with existing agricultural practices also have not been sufficiently explored.
From a critical standpoint, we think that progress in this field requires a shift from descriptive reporting toward hypothesis-driven and mechanism-testing frameworks. Key priorities include: (i) standardisation of experimental protocols to improve comparability among studies; (ii) explicit testing of causal pathways using omics and mutant-based approaches; and (iii) greater integration of ecological theory to understand stability, resilience, and context dependency of bacterial effects.
Recent omics-based studies have begun to unravel the molecular and community-level processes that underlie bacterial impacts on rice roots. For example, integrated 16S rRNA sequencing and metabolomic profiling have shown that drought and genotype jointly shape rhizosphere bacterial communities and root exudate compositions in rice, linking microbial shifts to specific metabolite profiles [79]. Transcriptomic analyses have further demonstrated that the presence of root microbiomes can induce major changes in rice root gene expression, including downregulation of immune-related receptor kinase families [80]. Metagenomic investigations reveal how stress-induced root exudates modulate rhizosphere microbiome composition and functional gene expression related to stress tolerance and metal resistance [81]. Moreover, comparative metagenomics across rice varieties under salinity and alkalinity stress indicates environment-dependent functional differentiation in rhizosphere microbial communities [82]. These omics insights underscore the complexity of plant–microbe regulatory networks and illustrate the value of multi-omics integration for identifying microbial genes, plant pathways, and metabolite signals involved in root–microbiome interactions [83].
4.5. Directions for Future Research
Upscaling soil microbiology research from the laboratory to ecosystems represents one of the grand challenges facing environmental scientists and is essential for informing land managers towards resource-efficient ecosystems. Scaling information from laboratory to field can be explored through experimental, theoretical, and modelling studies in future [84]. One important concern is the spatial upscale of plant–microbe interactions. Microbial functions (e.g., enzyme production and substrate metabolism) that influence soil properties [85] and root characteristics [86] strongly depend on the heterogeneity of soil [78]. Consequently, field studies are essential to consider this spatial variability and to validate laboratory results in more complex and real conditions. Therefore, scaling microbial processes from controlled environments to open fields or ecosystem scales remains a key challenge for future research [87].
Furthermore, the analysed literature often neglects the influence of variations in soil properties on bacterial behaviour and effectiveness. For instance, soil pH and salinity are acknowledged as dominant factors that indirectly influence rice growth, since they affect microbial diversity and functionality [88]. High pH and salinity can reduce microbial biomass, slow down or even block extracellular enzymatic activities, and reduce microbial diversity. Moreover, intensive tillage can disrupt bacterial communities in soil by constraining nitrification processes, which limits the potential yield of rice crops [89]. The impacts of climate change on soil microbial communities are another underexplored but critical issue. Changes in temperature and precipitation can lead to significant alterations in the physico-chemical and biological properties of soils [90]. Soil water content is extremely sensitive to climate change, and this sensitivity plays a crucial role in regulating microbial activity and plant–soil interactions [91,92,93]. Temporal and spatial fluctuations in soil moisture in response to climate change can influence root development and nutrient availability, in turn affecting plant growth and the structure of microbial communities [94,95]. Increasing temperatures, particularly in the topsoil (10–30 cm), may further reduce microbial diversity and complexity, impact nutrient cycling, and worsen rice productivity [95,96].
To fully recognise the potential of bacterial inoculants in rice cultivation, integrated field investigations are needed under different environmental conditions with particular attention to variations in climate, soil, and management practices. These types of studies are essential to verify whether those bacteria that appear beneficial in laboratory conditions can similarly help to improve rice yield in croplands.
Large-scale field experiments across contrasting soil types and climatic zones are essential to validate bacterial impacts on rice root systems under realistic conditions [29]. Moreover, multi-strain and community-based inoculation strategies should be prioritised to reflect natural rhizosphere complexity and enhance functional stability [21,62]. Advances in genomics, transcriptomics, and phenomics offer new opportunities to identify rice genotypes with heightened responsiveness to beneficial microbes and to disentangle plant–microbe signalling pathways [55].
Integrating these approaches can enable the development of microbiome-informed root ideotypes, where optimal root architectures are co-defined by plant genetics and microbial recruitment capacity [27,30]. Such ideotypes would be particularly valuable under climate change scenarios characterised by water scarcity, salinity intrusion, and metal contamination [32,53]. Ultimately, aligning microbial ecology with rice breeding and agronomic management represents a paradigm shift from empirical biofertilisation toward predictive, climate-smart rice production systems.
5. Conclusions
Rice, as an important food for half the population on a global scale, faces important growth and yield challenges under environmental stress (e.g., drought, heavy metal contamination, and climate change). This review has highlighted the essential role of bacterial strains (particularly Bacillus species) in supporting rice tolerance to these stresses. These bacteria substantially contribute to rice plant resilience and productivity by promoting mechanisms, such as improving biomass, water retention, antioxidant activity and heavy metal detoxification.
From this review, root length appears to be a key rice plant trait influenced by bacterial communities, since this trait is closely associated with nutrient uptake, and plant growth and production. However, to date, most studies have been conducted under laboratory conditions or as small-scale experiments, often focusing on single case studies (e.g., one bacterial strain). This limits the generalisation of the study findings, considering the influence of variability in soil properties, cultivation practices, and climate.
Recent evidence further indicates that bacterial strains do not simply enhance root growth but actively regulate rice root system architecture by modulating elongation, branching patterns, and root surface traits in response to environmental constraints. These regulatory effects highlight that bacterial influences on roots are context-dependent rather than additive, varying across soil conditions, flooding regimes, and rice genotypes. Future research should prioritise large-scale and field-based investigations that must integrate the aforementioned variables to better understand how bacteria interact with root architecture and the yield of rice plants.
Finally, this review underscores the potential of bacterial inoculation as an effective strategy to improve rice root systems and yields, paving the way for more resilient and sustainable rice cultivation. The integration of microbiological studies with agronomic management is promising for optimisation of the use of resources (water, fertilisers, tillage) and improvement of yield quality. Advances in biological techniques studying the action of microbial inoculants may provide sustainable tools to control climate stress and enhance rice yields. To meet these challenges, collaboration among researchers and farmers is essential to develop and experiment with quick and beneficial bacterial-based solutions in the field.
Viewing bacteria as co-regulators of root development rather than simple biofertilisers provides a conceptual framework for designing more predictable and climate-smart rice production systems.
Author Contributions
Conceptualisation, M.P., D.A.Z. and M.E.L.-B.; methodology, M.P., D.A.Z. and M.E.L.-B.; validation, M.P., D.A.Z. and M.E.L.-B.; formal analysis, M.P., D.A.Z. and M.E.L.-B.; investigation, M.P.; resources, D.A.Z.; data curation, M.P.; writing—original draft preparation, M.P.; writing—review and editing, M.P., D.A.Z. and M.E.L.-B.; supervision, D.A.Z. and M.E.L.-B.; project administration, D.A.Z.; funding acquisition, D.A.Z. All authors have read and agreed to the published version of the manuscript.
Funding
This research received no specific grant from any funding agency.
Data Availability Statement
The original contributions presented in the study are included in the article, further inquiries can be directed to the corresponding author.
Conflicts of Interest
The authors declare no conflicts of interest.
Appendix A

Table A1.
Complete search strategies of this review following the PRISMA 2020 guidelines.
Table A1.
Complete search strategies of this review following the PRISMA 2020 guidelines.
| Filter | Values |
|---|---|
| Keywords | Rice root, Bacteria, Root morphology, Bacterial strains, Rhizosphere |
| Boolean combinations of keywords | rice root and Bacteria Rice root and Rhizosphere and Bacterial Strains Bacterial Strains and Rice root or Root Morphology Bacteria and Rhizosphere |
| Document type | Article |
| Language | English |
| Time range | 1995–2025 |
| Subject area | Agricultural and Biological Sciences, Environmental Sciences |
| Subject category | Agricultural and Biological Sciences (miscellaneous), Plant Science Ecology, Environmental Science (miscellaneous) |
References
- FAOSTAT. Food and Agriculture Organization of the United Nations. Available online: www.fao.org/faostat/en/#data/QCL (accessed on 20 September 2025).
- Mohidem, N.A.; Hashim, N.; Shamsudin, R.; Che Man, H. Rice for food security: Revisiting its production, diversity, rice milling process and nutrient content. Agriculture 2022, 12, 741. [Google Scholar] [CrossRef]
- Zafar, S.; Xu, J. Recent advances to enhance nutritional quality of rice. Rice Sci. 2023, 30, 523–536. [Google Scholar] [CrossRef]
- Sen, S.; Chakraborty, R.; Kalita, P. Rice—Not just a staple food: A comprehensive review on its phytochemicals and therapeutic potential. Trends Food Sci. Technol. 2020, 97, 265–285. [Google Scholar] [CrossRef]
- Food and Agriculture Organization of the United Nations (FAO). The State of Food and Agriculture; United Nations: Rome, Italy, 2009. [Google Scholar]
- Devi, N.T.; Devi, A.S.; Singh, K.R. Impact of chemical fertilizer and composts application on growth and yield of rice in Northeast India. Sci. Rep. 2025, 15, 8575. [Google Scholar] [CrossRef]
- Tadesse, K.A.; Lu, Z.; Shen, Z.; Daba, N.A.; Li, J.; Alam, M.D.A.; Liu, L.; Gilbert, N.; Legesse, T.G.; Huimin, Z. Impacts of long-term chemical nitrogen fertilization on soil quality, crop yield, and greenhouse gas emissions: With insights into post-lime application responses. Sci. Total Environ. 2024, 944, 173827. [Google Scholar] [CrossRef]
- Chen, G.; Duan, Q.; Wu, C.; He, X.; Hu, M.; Li, C.; Ouyang, Y.; Peng, L.; Yang, H.; Zhang, Q.; et al. Optimizing rice yield, quality and nutrient use efficiency through combined application of nitrogen and potassium. Front. Plant Sci. 2024, 15, 1335744. [Google Scholar] [CrossRef]
- Johnson, J.M.; Becker, M.; Dossou-Yovo, E.R.; Saito, K. Improving rice yield and water productivity in dry climatic zones of West Africa: Season-specific strategies. Field Crops Res. 2024, 316, 109519. [Google Scholar] [CrossRef]
- Khan, M.; Dar, Z.; Dar, S. Breeding strategies for improving rice yield—A review. Agric. Sci. 2015, 6, 467–478. [Google Scholar] [CrossRef]
- Zhou, J.; Tang, S.; Pan, W.; Xu, M.; Liu, X.; Ni, L.; Mao, X.; Sun, T.; Fu, H.; Han, K.; et al. Long-term application of controlled-release fertilizer enhances rice production and soil quality under non-flooded plastic film mulching cultivation conditions. Agric. Ecosyst. Environ. 2023, 358, 108720. [Google Scholar] [CrossRef]
- Zou, Y.; Zhang, Y.; Cui, J.; Gao, J.; Guo, L.; Zhang, Q. Nitrogen fertilization application strategies improve yield of the rice cultivars with different yield types by regulating phytohormones. Sci. Rep. 2023, 13, 21803. [Google Scholar] [CrossRef]
- Li, P.; Tian, Y.; Yang, K.; Tian, M.; Zhu, Y.; Chen, X.; Hu, R.; Qin, T.; Liu, Y.; Peng, S.; et al. Mechanism of microbial action of the inoculated nitrogen-fixing bacterium for growth promotion and yield enhancement in rice (Oryza sativa L.). Adv. Biotechnol. 2024, 2, 32. [Google Scholar] [CrossRef] [PubMed]
- Zhong, Y.; Hu, J.; Xia, Q.; Zhang, S.; Li, X.; Pan, X.; Zhao, R.; Wang, R.; Yan, W.; Shangguan, Z.; et al. Soil microbial mechanisms promoting ultrahigh rice yield. Soil Biol. Biochem. 2020, 143, 107741. [Google Scholar] [CrossRef]
- Page, M.J.; McKenzie, J.E.; Bossuyt, P.M.; Boutron, I.; Hoffmann, T.C.; Mulrow, C.D.; Shamseer, L.; Tetzlaff, J.M.; Akl, E.A.; Brennan, S.E.; et al. The PRISMA 2020 statement: An updated guideline for reporting systematic reviews. BMJ 2021, 372, n71. [Google Scholar] [CrossRef] [PubMed]
- Govindarajan, M.; Balandreau, J.; Kwon, S.W.; Weon, H.Y.; Lakshminarasimhan, C. Effects of the inoculation of Burkholderia vietnamensis and related endophytic diazotrophic bacteria on grain yield of rice. Microb. Ecol. 2008, 55, 21–37. [Google Scholar] [CrossRef]
- Trân Van, V.; Berge, O.; Ngô Kê, S.; Balandreau, J.; Heulin, T. Repeated beneficial effects of rice inoculation with a strain of Burkholderia vietnamiensis on early and late yield components in low fertility sulphate acid soils of Vietnam. Plant Soil 2000, 218, 273–284. [Google Scholar] [CrossRef]
- Doni, F.; Suhaimi, N.S.M.; Mispan, M.S.; Fathurrahman, F.; Marzuki, B.M.; Kusmoro, J.; Uphoff, N. Microbial contributions for rice production: From conventional crop management to the use of ‘omics’ technologies. Int. J. Mol. Sci. 2022, 23, 737. [Google Scholar] [CrossRef]
- Gao, Y.; Yu, X.Z.; Wu, S.C.; Cheung, K.C.; Tam, N.F.Y.; Qian, P.Y. Interactions of rice (Oryza sativa L.) and PAH-degrading bacteria (Acinetobacter sp.) on enhanced dissipation of spiked phenanthrene and pyrene in waterlogged soil. Sci. Total Environ. 2006, 372, 1–11. [Google Scholar] [CrossRef]
- Liu, A.; Yang, X.; Guo, Q.; Li, B.; Zheng, Y.; Shi, Y.; Zhu, L. Microbial communities and flavor compounds during the fermentation of traditional Hong Qu glutinous rice wine. Foods 2022, 11, 1097. [Google Scholar] [CrossRef]
- Misu, I.J.; Kayess, M.O.; Siddiqui, M.N.; Gupta, D.R.; Islam, M.N.; Islam, T. Microbiome engineering for sustainable rice production: Strategies for biofertilization, stress tolerance, and climate resilience. Microorganisms 2025, 13, 233. [Google Scholar] [CrossRef]
- Ríos-Ruiz, W.F.; Jave-Concepción, H.G.; Torres-Chávez, E.E.; Rios-Reategui, F.; Padilla-Santa-Cruz, E.; Guevara-Pinedo, N.E. Plant-growth-promoting microorganisms: Their impact on crop quality and yield, with a focus on rice. Int. J. Plant Biol. 2025, 16, 9. [Google Scholar] [CrossRef]
- Arora, N.K.; Mishra, J.; Singh, P.; Fatima, T. Salt-tolerant plant growth-promoting Pseudomonas atacamensis KSS-6 in combination with organic manure enhances rice yield, improves nutrient content and soil properties under salinity stress. J. Basic Microbiol. 2024, 64, 2300767. [Google Scholar] [CrossRef]
- Khan, M.A.; Asaf, S.; Khan, A.L.; Adhikari, A.; Jan, R.; Ali, S.; Imran, M.; Kim, K.-M.; Lee, I.-J. Plant growth-promoting endophytic bacteria augment growth and salinity tolerance in rice plants. Plant Biol. 2020, 22, 850–862. [Google Scholar] [CrossRef] [PubMed]
- Sarkodee-Addo, E.; Tokiwa, C.; Bonney, P.; Aboagye, D.A.; Yeboah, A.; Abebrese, S.O.; Bam, R.; Nartey, E.K.; Okazaki, S.; Yasuda, M. Biofertilizer activity of Azospirillum sp. B510 on the rice productivity in Ghana. Microorganisms 2021, 9, 2000. [Google Scholar] [CrossRef] [PubMed]
- Shakeel, M.; Rais, A.; Hassan, M.N.; Hafeez, F.Y. Root associated Bacillus sp. improves growth, yield and zinc translocation for Basmati rice (Oryza sativa) varieties. Front. Microbiol. 2015, 6, 1286. [Google Scholar] [CrossRef] [PubMed]
- Domingo, C.; San Segundo, B. Rice thematic special issue: Beneficial plant-microbe interactions in rice. Rice 2023, 16, 50. [Google Scholar] [CrossRef]
- Xu, D.; Yu, X.; Chen, J.; Liu, H.; Zheng, Y.; Qu, H.; Bao, Y. Microbial assemblages associated with the soil-root continuum of an endangered plant, Helianthemum songaricum Schrenk. Microbiol. Spectr. 2023, 11, e0338922. [Google Scholar] [CrossRef]
- Zhao, J.; Yu, X.; Zhang, C.; Hou, L.; Wu, N.; Zhang, W.; Wang, Y.; Yao, B.; Delaplace, P.; Tian, J. Harnessing microbial interactions with rice: Strategies for abiotic stress alleviation in the face of environmental challenges and climate change. Sci. Total Environ. 2024, 912, 168847. [Google Scholar] [CrossRef]
- Zhou, Y.; Wei, Y.; Zhao, Z.; Li, J.; Li, H.; Yang, P.; Tian, S.; Ryder, M.; Toh, R.; Yang, H.; et al. Microbial communities along the soil-root continuum are determined by root anatomical boundaries, soil properties, and root exudation. Soil Biol. Biochem. 2022, 171, 108721. [Google Scholar] [CrossRef]
- Sun, L.; Yang, Y.; Wang, R.; Li, S.; Qiu, Y.; Lei, P.; Gao, J.; Xu, H.; Zhang, F.; Lv, Y. Effects of exopolysaccharide derived from Pantoe aalhagi NX-11 on drought resistance of rice and its efficient fermentation preparation. Int. J. Biol. Macromol. 2020, 162, 946–955. [Google Scholar] [CrossRef]
- Degon, Z.; Dixon, S.; Rahmatallah, Y.; Galloway, M.; Gulutzo, S.; Price, H.; Cook, J.; Glazko, G.; Mukherjee, A. Azospirillum brasilense improves rice growth under salt stress by regulating the expression of key genes involved in salt stress response, abscisic acid signaling, and nutrient transport, among others. Front. Agron. 2023, 5, 1216503. [Google Scholar] [CrossRef]
- Maqsood, A.; Shahid, M.; Hussain, S.; Mahmood, F.; Azeem, F.; Tahir, M.; Ahmed, T.; Noman, M.; Manzoor, I.; Basit, F. Root colonizing Burkholderia sp. AQ12 enhanced rice growth and upregulated tillering-responsive genes in rice. Appl. Soil Ecol. 2021, 157, 103769. [Google Scholar] [CrossRef]
- Chauhan, P.S.; Lata, C.; Tiwari, S.; Chauhan, A.S.; Mishra, S.K.; Agrawal, L.; Nautiyal, C.S. Transcriptional alterations reveal Bacillus amyloliquefaciens-rice cooperation under salt stress. Sci. Rep. 2019, 9, 11912. [Google Scholar] [CrossRef]
- Ali, N.; Swarnkar, M.K.; Veer, R.; Kaushal, P.; Pati, A.M. Temperature-induced modulation of stress-tolerant PGP genes bioprospected from Bacillus sp. IHBT-705 associated with saffron (Crocus sativus) rhizosphere: A natural-treasure trove of microbial biostimulants. Front. Plant Sci. 2023, 14, 1141538. [Google Scholar] [CrossRef]
- Sultana, R.; Jashim, A.I.I.; Islam, S.M.N.; Rahman, M.D.H.; Haque, M.M. Bacterial endophyte Pseudomonas mosselii PR5 improves growth, nutrient accumulation, and yield of rice (Oryza sativa L.) through various application methods. BMC Plant Biol. 2024, 24, 1030. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Xu, W.; Zhang, Q.; Liao, W.; Li, L.; Chen, S.; Yang, J.; Wang, Z.; Xu, F. OsPHR2-mediated recruitment of Pseudomonadaceae enhances rice phosphorus uptake. Plant Commun. 2024, 5, 100930. [Google Scholar] [CrossRef] [PubMed]
- Xiao, D.; Tang, X.; Chen, S.; Chu, G.; Liu, Y.; Wang, D. Aeration treatment promotes transformation of soil phosphorus fractions to plant-available phosphorus by modulating rice rhizosphere microbiota. Soil Tillage Res. 2025, 245, 106318. [Google Scholar] [CrossRef]
- Othman, N.M.I.; Othman, R.; Saud, H.M.; Wahab, P.E.M. Effects of root colonization by zinc-solubilizing bacteria on rice plant (Oryza sativa MR219) growth. Agric. Nat. Resour. 2017, 51, 532–537. [Google Scholar] [CrossRef]
- Ali, Q.; Ayaz, M.; Yu, C.; Wang, Y.; Gu, Q.; Wu, H.; Gao, X. Cadmium tolerant microbial strains possess different mechanisms for cadmium biosorption and immobilization in rice seedlings. Chemosphere 2022, 303, 135206. [Google Scholar] [CrossRef]
- dos Santos Ferreira, N.; Coniglio, A.; Puente, M.; Sant’Anna, F.H.; Maroniche, G.; García, J.; Molina, R.; Nievas, S.; Volpiano, C.G.; Ambrosini, A.; et al. Genome-based reclassification of Azospirillum brasilense Az39 as the type strain of Azospirillum argentinense sp. nov. Int. J. Syst. Evol. Microbiol. 2022, 72, 005475. [Google Scholar] [CrossRef]
- Loaces, I.; Ferrando, L.; Fernández Scavino, A. Dynamics, diversity and function of endophytic siderophore-producing bacteria in rice. Microb. Ecol. 2011, 61, 606–618. [Google Scholar] [CrossRef]
- Liang, Z.; Ali, Q.; Wang, Y.; Mu, G.; Kan, X.; Ren, Y.; Manghwar, H.; Gu, Q.; Wu, H.; Gao, X. Toxicity of Bacillus thuringiensis strains derived from the novel crystal protein Cry31Aa with high nematicidal activity against rice parasitic nematode Aphelenchoides besseyi. Int. J. Mol. Sci. 2022, 23, 8189. [Google Scholar] [CrossRef]
- Rajer, F.U.; Samma, M.K.; Ali, Q.; Rajar, W.A.; Wu, H.; Raza, W.; Xie, Y.; Tahir, H.A.S.; Gao, X. Bacillus spp.-mediated growth promotion of rice seedlings and suppression of bacterial blight disease under greenhouse conditions. Pathogens 2022, 11, 1251. [Google Scholar] [CrossRef]
- Thanwisai, L.; Kim Tran, H.T.; Siripornadulsil, W.; Siripornadulsil, S.A. Cadmium-tolerant endophytic bacterium reduces oxidative stress and Cd uptake in KDML105 rice seedlings by inducing glutathione reductase-related activity and increasing the proline content. Plant Physiol. Biochem. 2022, 192, 72–86. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Zhou, Q.; Gao, C.; Lu, Y.; Sheng, Y.; Xiao, M.; Yun, Y.; Selvaraj, J.N.; Zhang, X.; Li, Y.; et al. Endophytic bacteria for Cd remediation in rice: Unraveling the Cd tolerance mechanisms of Cupriavidus metallidurans CML2. J. Hazard. Mater. 2024, 469, 133846. [Google Scholar] [CrossRef] [PubMed]
- Gao, Z.; Jiang, Y.; Li, W.; Chen, H.; Ye, M.; Liang, Y. Evidence for the role of microbes in the silicon-regulated arsenic concentrations of rice roots in the soil environment. Sci. Total Environ. 2024, 908, 168321. [Google Scholar] [CrossRef] [PubMed]
- Namdeo, V.; Mitra, N.; Jakhar, S.; Sahu, R. Increase in growth, productivity, nutritional status of rice (Oryza sativa L.) and enrichment in soil fertility applied with Azospirillum and nitrogen level. Int. J. Curr. Microbiol. App. Sci. 2020, 9, 242–248. [Google Scholar] [CrossRef]
- Han, Y.; Wang, R.; Yang, Z.; Zhan, Y.; Ma, Y.; Ping, S.; Zhang, L.; Lin, M.; Yan, Y. 1-Aminocyclopropane-1-carboxylate deaminase from Pseudomonas stutzeri A1501 facilitates the growth of rice in the presence of salt or heavy metals. J. Microbiol. Biotechnol. 2015, 25, 1119–1128. [Google Scholar] [CrossRef]
- de Souza, R.; Meyer, J.; Schoenfeld, R.; da Costa, P.B.; Passaglia, L.M.P. Characterization of plant growth-promoting bacteria associated with rice cropped in iron-stressed soils. Ann. Microbiol. 2015, 65, 951–964. [Google Scholar] [CrossRef]
- Hong, Y.; Huang, H. The role of soil microbiota in rice cultivation and its implications for agricultural sustainability. Mol. Soil Biol. 2024, 15, 87–98. [Google Scholar] [CrossRef]
- Narayanasamy, S.; Thangappan, S.; Uthandi, S. Plant growth-promoting Bacillus sp. cahoots moisture stress alleviation in rice genotypes by triggering antioxidant defense system. Microbiol. Res. 2020, 239, 126518. [Google Scholar] [CrossRef]
- Yang, X.; Yuan, R.; Yang, S.; Dai, Z.; Di, N.; Yang, H.; He, Z.; Wei, M. A salt-tolerant growth-promoting phyllosphere microbial combination from mangrove plants and its mechanism for promoting salt tolerance in rice. Microbiome 2024, 12, 270. [Google Scholar] [CrossRef] [PubMed]
- Fagorzi, C.; Checcucci, A.; DiCenzo, G.C.; Debiec-Andrzejewska, K.; Dziewit, L.; Pini, F.; Mengoni, A. Harnessing rhizobia to improve heavy-metal phytoremediation by legumes. Genes 2018, 9, 542. [Google Scholar] [CrossRef] [PubMed]
- Klinsawang, S.; Aesomnuk, W.; Mangkalasane, P.; Ruanjaichon, V.; Siangliw, J.L.; Pandey, B.K.; Bennett, M.J.; Wanchana, S.; Arikit, S. Genome-wide association study identifies key F-box genes linked to ethylene responsiveness and root growth in rice (Oryza sativa L.). Front. Plant Sci. 2024, 15, 1501533. [Google Scholar] [CrossRef] [PubMed]
- Reichert, T.; Rammig, A.; Fuchslueger, L.; Lugli, L.F.; Quesada, C.A.; Fleischer, K. Plant phosphorus-use and -acquisition strategies in Amazonia. New Phytol. 2022, 234, 1126–1143. [Google Scholar] [CrossRef]
- Hetrick, B.A.D. Mycorrhizas and root architecture. Experientia 1991, 47, 355–361. [Google Scholar] [CrossRef]
- Huergo, L.F.; Monterio, R.A.; Bonatto, A.C.; Rigo, L.U.; Steffens, M.B.R.; Cruz, L.M.; Chubatsu, L.S.; Souza, E.M.; Pedrosa, F.O. Regulation of nitrogen fixation in Azospirillum brasilense. In Azospirillum sp.: Cell Physiology, Plant Interactions and Agronomic Research in Argentina; Cassan, F.D., Garcia de Salmone, I., Eds.; Asociación Argentina de Microbiología: Buenos Aires, Argentina, 2008; pp. 17–36. [Google Scholar]
- Das, J.; Sultana, S.; Rangappa, K.; Kalita, M.C.; Thakuria, D. Endophyte bacteria alter physiological traits and promote growth of rice (Oryza sativa L.) in aluminium toxic and phosphorus deficient acid Inceptisols. J. Pure Appl. Microbiol. 2020, 14, 627–639. [Google Scholar] [CrossRef]
- Li, S.; Yan, Q.; Wang, J.; Jiang, H.; Li, Z.; Peng, Q. Root endophyte shift and key genera discovery in rice under barnyardgrass stress. Rice Sci. 2023, 30, 160–170. [Google Scholar] [CrossRef]
- Zhang, Y.; Du, H.; Xu, F.; Ding, Y.; Gui, Y.; Zhang, J.; Xu, W. Root-bacteria associations boost rhizosheath formation in moderately dry soil through ethylene responses. Plant Physiol. 2020, 183, 780–792. [Google Scholar] [CrossRef]
- Islam, T.; Fatema, N.; Hoque, M.N.; Gupta, D.R.; Mahmud, N.U.; Sakif, T.I.; Sharpe, A.G. Improvement of growth, yield and associated bacteriome of rice by the application of probiotic Paraburkholderia and Delftia. Front. Microbiol. 2023, 14, 1212505. [Google Scholar] [CrossRef]
- Rosecrance, R.C.; Weinbaum, S.A.; Brown, P.H. Assessment of nitrogen, phosphorus, and potassium uptake capacity and root growth in mature alternate-bearing pistachio (Pistacia vera) trees. Tree Physiol. 1996, 16, 949–956. [Google Scholar] [CrossRef]
- Chen, J.; Cao, F.; Liu, Y.; Tao, Z.; Lei, T.; Abou-Elwafa, S.F.; Huang, M. Comparison of short-duration and long-duration rice cultivars cultivated in various planting densities. Agronomy 2022, 12, 1739. [Google Scholar] [CrossRef]
- Lang, Y.; Dou, Y.; Wang, M.; Zhang, Z.; Zhu, Q. Effects of growth duration on grain yield and quality in rice. Acta Agron. Sin. 2012, 38, 528–534. [Google Scholar] [CrossRef]
- Herath, I.; Zhao, F.J.; Bundschuh, J.; Wang, P.; Wang, J.; Ok, Y.S.; Palansooriya, K.N.; Vithanage, M. Microbe mediated immobilization of arsenic in the rice rhizosphere after incorporation of silica impregnated biochar composites. J. Hazard. Mater. 2020, 398, 123096. [Google Scholar] [CrossRef] [PubMed]
- Yanni, Y.G.; Rizk, R.Y.; Corich, V.; Squartini, A.; Ninke, K.; Philip-Hollingsworth, S. Natural endophytic association between Rhizobium leguminosarum bv. trifolii and rice roots and assessment of its potential to promote rice growth. Plant Soil 1997, 194, 99–114. [Google Scholar] [CrossRef]
- Beneduzi, A.; Peres, D.; Vargas, L.K.; Bodanese-Zanettini, M.H.; Passaglia, L.M.P. Evaluation of genetic diversity and plant growth promoting activities of nitrogen-fixing Bacilli isolated from rice fields in South Brazil. Appl. Soil Ecol. 2008, 39, 311–320. [Google Scholar] [CrossRef]
- Khanam, M.; Solaiman, A.R.M.; Rahman, G.K.M.M.; Haque, M.M.; Alam, M.S. Effect of plant growth promoting bacteria on growth and nutrient content of rice. Asian J. Adv. Agric. Res. 2022, 19, 20–29. [Google Scholar] [CrossRef]
- Ali, Q.; Yu, C.; Wang, Y.; Sheng, T.; Zhao, X. High killing rate of nematode and promotion of rice growth by synthetic volatiles from Bacillus strains due to enhanced oxidative stress response. Physiol. Plant. 2023, 175, e13868. [Google Scholar] [CrossRef]
- Li, P.; Xiong, Z.; Tian, Y.; Zheng, Z.; Liu, Z.; Hu, R.; Wang, Q.; Ao, H.; Yi, Z.; Li, J. Community-based mechanisms underlying the root cadmium uptake regulated by Cd-tolerant strains in rice (Oryza sativa L.). Front. Plant Sci. 2023, 14, 1196130. [Google Scholar] [CrossRef]
- Shi, F.; Zhu, S.; Li, H.; Zhang, B.; Liu, J.; Song, F. Further enhancement of cold tolerance in rice seedlings by Piriformospora indica collaborating with plant growth-promoting bacteria: Evidence from the antioxidant defense, osmoregulation, photosynthesis, and related genes. Plant Stress 2024, 14, 100656. [Google Scholar] [CrossRef]
- Bal, H.B.; Nayak, L.; Das, S.; Adhya, T.K. Isolation of ACC deaminase producing PGPR from rice rhizosphere and evaluating their plant growth promoting activity under salt stress. Plant Soil 2013, 366, 93–105. [Google Scholar] [CrossRef]
- Spence, C.; Alff, E.; Johnson, C.; Ramos, C.; Donofrio, N.; Sundaresan, V.; Bais, H. Natural rice rhizospheric microbes suppress rice blast infections. BMC Plant Biol. 2014, 14, 130. [Google Scholar] [CrossRef] [PubMed]
- Joshi, B.; Chaudhary, A.; Singh, H.; Kumar, P.A. Prospective evaluation of individual and consortia plant growth promoting rhizobacteria for drought stress amelioration in rice (Oryza sativa L.). Plant Soil 2020, 457, 225–240. [Google Scholar] [CrossRef]
- Xiao, X.; Zhu, Y.; Gao, C.; Zhang, Y.; Gao, Y.; Zhao, Y. Microbial inoculations improved rice yields by altering the presence of soil rare bacteria. Microbiol. Res. 2022, 254, 126910. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Wu, H.; Pu, Q.; Zhang, C.; Chen, Y.; Lin, Z.; Hu, X.; Li, O. Complete genome of Sphingomonas paucimobilis ZJSH1, an endophytic bacterium from Dendrobium officinale with stress resistance and growth promotion potential. Arch. Microbiol. 2023, 205, 132. [Google Scholar] [CrossRef]
- Šnajdr, J.; Valášková, V.; Merhautová, V.; Herinková, J.; Cajthaml, T.; Baldrian, P. Spatial variability of enzyme activities and microbial biomass in the upper layers of Quercus petraea forest soil. Soil Biol. Biochem. 2008, 40, 2068–2075. [Google Scholar] [CrossRef]
- Chen, J.; Zhang, Y.; Kuzyakov, Y.; Wang, D.; Olesen, J.E. Challenges in upscaling laboratory studies to ecosystems in soil microbiology research. Glob. Change Biol. 2023, 29, 569–574. [Google Scholar] [CrossRef]
- Li, G.; Wang, K.; Qin, Q.; Li, Q.; Mo, F.; Nangia, V.; Liu, Y. Integrated microbiome and metabolomic analysis reveal responses of rhizosphere bacterial communities and root exudate composition to drought and genotype in rice (Oryza sativa L.). Rice 2023, 16, 19. [Google Scholar] [CrossRef]
- Santos-Medellín, C.; Edwards, J.; Nguyen, B.; Sundaresan, V. Acquisition of a complex root microbiome reshapes the transcriptomes of rice plants. New Phytol. 2022, 235, 2008–2021. [Google Scholar] [CrossRef]
- Zhu, S.; Zhao, W.; Sun, S.; Yang, X.; Mao, H.; Sheng, L.; Chen, Z. Community metagenomics reveals the processes of cadmium resistance regulated by microbial functions in soils with Oryza sativa root exudate input. Sci. Total Environ. 2024, 949, 175015. [Google Scholar] [CrossRef]
- Zhong, Y.; Chi, H.; Wu, T.; Fan, W.; Su, H.; Li, R.; Jiang, W.; Du, X.; Ma, Z. Diversity of rhizosphere microbial communities in different rice varieties and their diverse adaptive responses to saline and alkaline stress. Front. Microbiol. 2025, 16, 1537846. [Google Scholar] [CrossRef]
- Jain, A.; Sarsaiya, S.; Singh, R.; Gong, Q.; Wu, Q.; Shi, J. Omics approaches in understanding the benefits of plant-microbe interactions. Front. Microbiol. 2024, 15, 1391059. [Google Scholar] [CrossRef] [PubMed]
- Parhizkar, M.; Lucas-Borja, M.E.; Zema, D.A. Effects of inoculation with Bacillus arenosi on physical and biochemical properties of soil, surface runoff and erosion in degraded forestlands of Northern Iran. Ecol. Eng. 2024, 209, 107413. [Google Scholar] [CrossRef]
- Ghasemzadeh, Z.; Parhizkar, M.; Zomorodian, M.; Shamsi, R.; Mirmohammadmeygooni, S.; Shabanpour, M. The role of extracellular polysaccharide produced by Bradyrhizobium strain in root growth, improvement of soil aggregate stability and reduction of soil detachment capacity. Rhizosphere 2023, 27, 100771. [Google Scholar] [CrossRef]
- Classen, A.T.; Sundqvist, M.K.; Henning, J.A.; Newman, G.S.; Moore, J.A.M.; Cregger, M.A.; Moorhead, L.C.; Patterson, C.M. Direct and indirect effects of climate change on soil microbial and soil microbial-plant interactions: What lies ahead? Ecosphere 2015, 6, 130. [Google Scholar] [CrossRef]
- Li, Y.; Hu, Y.; Yan, C.; Xiong, J.; Qiu, Q. pH and salinity are the dominant limiting factors for the application of mariculture sludge to paddy soil. Appl. Soil Ecol. 2022, 175, 104463. [Google Scholar] [CrossRef]
- Liu, Y.; Zhang, J.; Wang, Z.; Ke, W.; Wang, L.; Peng, T.; Zhao, Q. Restriction of soil bacteria promoting high yield of super hybrid rice in the Huaihe Valley in central China by conventional ploughing intensity. Soil Tillage Res. 2021, 214, 105169. [Google Scholar] [CrossRef]
- Brevik, E.C. The potential impact of climate change on soil properties and processes and corresponding influence on food security. Agriculture 2013, 3, 398–417. [Google Scholar] [CrossRef]
- Banerjee, S.; Helgason, B.; Wang, L.; Winsley, T.; Ferrari, B.C.; Siciliano, S.D. Legacy effects of soil moisture on microbial community structure and N2O emissions. Soil Biol. Biochem. 2016, 95, 40–50. [Google Scholar] [CrossRef]
- Borowik, A.; Wyszkowska, J. Soil moisture as a factor affecting the microbiological and biochemical activity of soil. Plant Soil Environ. 2016, 62, 250–255. [Google Scholar] [CrossRef]
- Chen, Y.; Ma, S.; Jiang, H.; Hu, Y.; Lu, X. Influences of litter diversity and soil moisture on soil microbial communities in decomposing mixed litter of alpine steppe species. Geoderma 2020, 377, 114577. [Google Scholar] [CrossRef]
- Chen, X.; Chen, Y.; Zhang, W.; Zhang, W.; Wang, H.; Zhou, Q. Response characteristics of root to moisture change at seedling stage of Kengyilia hirsuta. Front. Plant Sci. 2023, 13, 1052791. [Google Scholar] [CrossRef]
- Hao, Y.; Mao, J.; Bachmann, C.M.; Chen, B.; Zhang, Q.; Gu, J.; He, J.Z. Soil moisture controls over carbon sequestration and greenhouse gas emissions: A review. npj Clim. Atmos. Sci. 2025, 8, 16. [Google Scholar] [CrossRef]
- Fu, F.; Li, J.; Li, Y.; Chen, W.; Ding, H.; Xiao, S. Simulating the effect of climate change on soil microbial community in an Abies georgei var. smithii forest. Front. Microbiol. 2023, 14, 1189859. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.