Abstract
Plant diseases pose significant threats globally due to the high economic losses and effects on food security. Traditional disease identification methods usually have limitations regarding their accuracy and efficiency. This study discusses six advanced deep learning models: VGG19, DenseNet201, Xception, InceptionResNetV2, MobileNetV2, and EfficientNetV2B3. A dataset is used that is rich in diversity and contains high-quality images of diseased sections or parts of plants. These deep models are discussed and compared for studying their efficiencies in recognizing plant diseases accurately. EfficientNetV2B3 and Xception outperformed the rest of the models due to the ability of the model to capture major features from the image of the infected region. MobileNetV2 was also useful which provided a good trade-off between accuracy and computational efficiency. The study further applied transfer learning and image augmentation in boosting model performance and addressing the issue of class imbalance in the dataset. Results showed that the proposed approach proved much more reliable and efficient compared to conventional approaches to plant disease detection. Future efforts will be geared towards early detection of diseases to further assist farmers and researchers in order to upgrade the practices related to crop management. Additional data will be integrated, including hyperspectral images and environmental factors, for developing a robust and efficient system for plant disease detection. These models will be deployed in intelligent farming systems.
1. Introduction
Plant diseases present a serious threat to agriculture in every part of the world. The diseases reduce yields and threaten economic stability and food security. Traditional methods of disease detection relied on inspecting infected crops, followed by confirmatory tests in the laboratory. These methods are slow and require expert skill, thus making large-scale applications difficult. Due to the disadvantages of conventional approaches, advanced technologies like deep learning are being developed, which can identify diseases more quickly and accurately [1]. Deep learning in general, and CNNs particularly, has shown significant improvement in identifying and diagnosing plant diseases from images of infected plant parts—leaves, stems, roots, and fruits [2].
Compared with traditional methods, deep-learning-based models are easier to use and require less domain expertise. They leverage large image datasets for rapid disease detection, enabling farmers to take remedial actions before serious crop damage occurs (as discussed in the Literature Review). Deep learning models extract the most indicative features of infected plants, balancing accuracy and computational efficiency for reliable diagnoses. Training on diverse datasets improves early disease detection, allowing timely intervention for effective crop management [3].
Training on large datasets enhances the performance of these systems to detect diseases early, thus allowing timely intervention by farmers and researchers for effective control. In the future, continued improvement will certainly push these models toward greater precision and wider availability. When integrated into smart farming, they can significantly lift yields by cutting disease outbreaks early. All in all, these technologies cultivate more productive means of farming. Additionally, models with real-time monitoring and automated decision-making capabilities will enable even higher agricultural efficiency. Agricultural experts take advantage of transfer learning and augmenting data to optimize learning. Transfer learning uses a pre-trained model that has already acquired key features from big datasets, which improves model performance in the recognition of diverse plant images. Mainly, early disease identification remains an important area of interest for further research in the detection of plant diseases. Researchers also employ advanced techniques, such as hyperspectral imaging and environmental data, to enhance disease detection. Environmental factors like temperature, humidity, and soil conditions influence the development of diseases, while hyperspectral images provide comprehensive information about plant diseases. With these additional features, the models become more efficient and their results highly accurate. Deep learning models in mobile applications are making farming easier. These models also lead to enhanced efficiency and accuracy in farm operations when integrated with smart farming methods. Generally, farmers can identify diseases in plants more rapidly using smartphone apps, reducing their dependence on expert consultations and expensive laboratory tests. Smart farming systems support the detection of plant diseases and help farmers make better decisions for improved health and productivity of crops. Deep learning remains among the most effective approaches to the identification of plant diseases, which offers fast, precise, and consistent solutions.
Finally, all these contributions support healthy crop management and ensure enhanced food security. Despite these advancements, challenges persist. Many existing studies focus on a single model or specific disease types, without systematically comparing the performance of multiple deep learning architectures on the same dataset. They often do not fully address issues such as class imbalance, model generalization under real field conditions, and practical applicability for real-world deployment. This study aims to fill some of these gaps by evaluating six deep learning models namely VGG19, DenseNet201, Xception, InceptionResNetV2, MobileNetV2, and EfficientNetV2B3 using a self-collected paddy disease dataset. By incorporating transfer learning and data augmentation, the study seeks to improve both the accuracy and stability of disease recognition and to provide guidance for selecting the most suitable model for intelligent agricultural disease identification systems.
The main contributions of this study are as follows:
- A systematic comparison of six widely used deep learning architectures for paddy disease classification under identical experimental conditions.
- An extensive evaluation of transfer learning and fine-tuning strategies to address class imbalance and improve model generalization.
- A detailed analysis of model performance before and after fine-tuning using consistent evaluation metrics.
- Practical insights into model selection for intelligent agricultural disease identification systems, considering both accuracy and computational efficiency.
2. Literature Survey
Plant diseases cause a significant reduction in crop production and quality, thus threatening food security and economic sustainability. The early detection of plant diseases in plant tissues inhibits the spread of the disease [4]. Various research studies have been conducted to explore the use of deep learning models for the task. Early work has been done to create disease image datasets for rice, wheat, and maize plants and to analyze the performance of different CNN models, which have shown promising results in classification tasks [5]. Improvements in preprocessing techniques and the combination of CNN models improve classification results, allowing farmers to take preventive actions [6,7].
Deep transfer learning models have been widely used for enhancing generalization capabilities on various datasets [8]. Variants of DenseNet have shown promising results in classification accuracy for rice disease diagnosis [3], while Bayesian learning methods implemented in CNN architectures have ensured effective early diagnosis [9]. Certain efforts for paddy disease classification were removed owing to concerns regarding dataset quality and incorrect labeling, thus emphasizing difficulties in data acquisition and labeling [10]. Systematic reviews on rice leaf disease diagnosis reveal the evolving landscape of transfer learning, ensemble learning, and hybrid deep learning models, while emphasizing drawbacks such as class imbalance and inconsistent metrics [11].
Hybrid models that combine CNNs with simulated thermal images have improved disease diagnosis in varying field conditions [12]. Traditional machine learning algorithms, such as SVM and decision tree classifiers, have also been used for rice leaf disease recognition but tend to be less effective compared to deep learning methods, especially when working with small and imbalanced datasets [13,14]. Similarly, research on fruit and vegetable disease identification suggests the limitations of data availability and efficiency, hence the need for deep learning [15].
Current research focuses on multi-prediction and generalization of models. Transfer learning methods have shown high accuracy for particular diseases like blast and bacterial blight in paddy crops [16,17,18]. VGG16 with transfer learning has proved to be useful for smaller datasets, providing real-world applications [19]. Multi-scale feature extraction and combination techniques, such as Xception for wheat disease classification, improve results further [20]. One-shot learning techniques have been proposed for small datasets, providing better early disease diagnosis in precision agriculture [21].
Real-time and deployment-driven research focuses on applicability. CNN-based monitoring systems facilitate real-time identification of leaf diseases with promising accuracy [22]. ResNet and other models have been used on various crops such as rice, tomatoes, and potatoes, with high accuracy while emphasizing the need for varied datasets and robustness to variations in the field [23,24]. Various reviews highlight the significance of standardized testing, larger datasets, and comparisons among models for deployment [25]. Recent developments include the integration of vision transformers and hybrid deep learning models for improved accuracy [26,27,28]. Other research works have benchmarked machine learning and deep learning models for plant disease classification, emphasizing parameter optimization and data set development for sustainable agriculture practices [29,30,31].
In general, the literature shows that there has been great progress in the use of deep learning for plant disease detection, but there are still some limitations:
- Class imbalance: There is a tendency for many datasets to have a low number of instances of the minority classes of diseases, which could affect model training and bias results.
- Limited generalization: The models developed on controlled data may not perform well in varying field conditions [32].
- Incomplete reporting of metrics: Some studies only report accuracy, which hinders reproducibility and comparison [33].
- Applicability: There are very few studies that show the applicability of their work in real-life agricultural settings [34].
The existence of these gaps makes the current study valid and necessary, as it compares six architectures of deep learning on a self-collected paddy disease dataset, using transfer learning and data augmentation.
3. Dataset Description
The dataset used in this study consists of paddy leaf images representing ten major disease classes. The original dataset exhibits significant class imbalance, which was addressed using data augmentation techniques applied exclusively to the training set as shown in Table 1.

Table 1.
Class-wise distribution of the paddy disease dataset.
The dataset used in this study was assembled specifically for experimental evaluation of paddy disease classification models and consists of images representing ten commonly reported paddy leaf disease categories. The dataset was curated from previously available image collections used in prior experimental studies and manually assigned to disease categories based on observable visual symptoms. As the dataset was prepared prior to this study, detailed field-level acquisition metadata such as exact capture location, imaging device specifications, and environmental conditions are not available. Nevertheless, the dataset exhibits sufficient visual diversity in terms of disease patterns, severity levels, and background variations to support reliable comparative deep learning experiments.
4. Train–Validation Split
The augmented dataset was divided using an 80:20 split, resulting in:
- Training set: 80% of 6524 images
- Validation set: 20% of 6524 images
Data augmentation was applied only to the training set, while the validation set was kept unchanged except for normalization.
5. Class Imbalance Handling
The original dataset shows substantial class imbalance, particularly for diseases such as grain discoloration, sheath blight, sheath rot, and tungro, which contain far fewer samples than other classes. To mitigate this issue and improve model generalization, image augmentation techniques including rotation, shifting, zooming, and horizontal flipping were employed. This strategy ensured a more balanced class distribution and enabled the models to learn robust and discriminative features from limited data.
6. Methodology
The primary objective of this methodology is not to propose a new network architecture, but to provide a systematic and fair comparison of six widely used deep learning models for paddy disease classification under identical experimental conditions. Unlike many prior studies that evaluate a single model or use inconsistent training settings, this work applies the same dataset, augmentation strategy, and evaluation protocol across all models. This enables an objective assessment of how different architectures respond to transfer learning, fine-tuning, and class imbalance, thereby offering practical guidance for selecting suitable models in real-world agricultural disease identification systems.
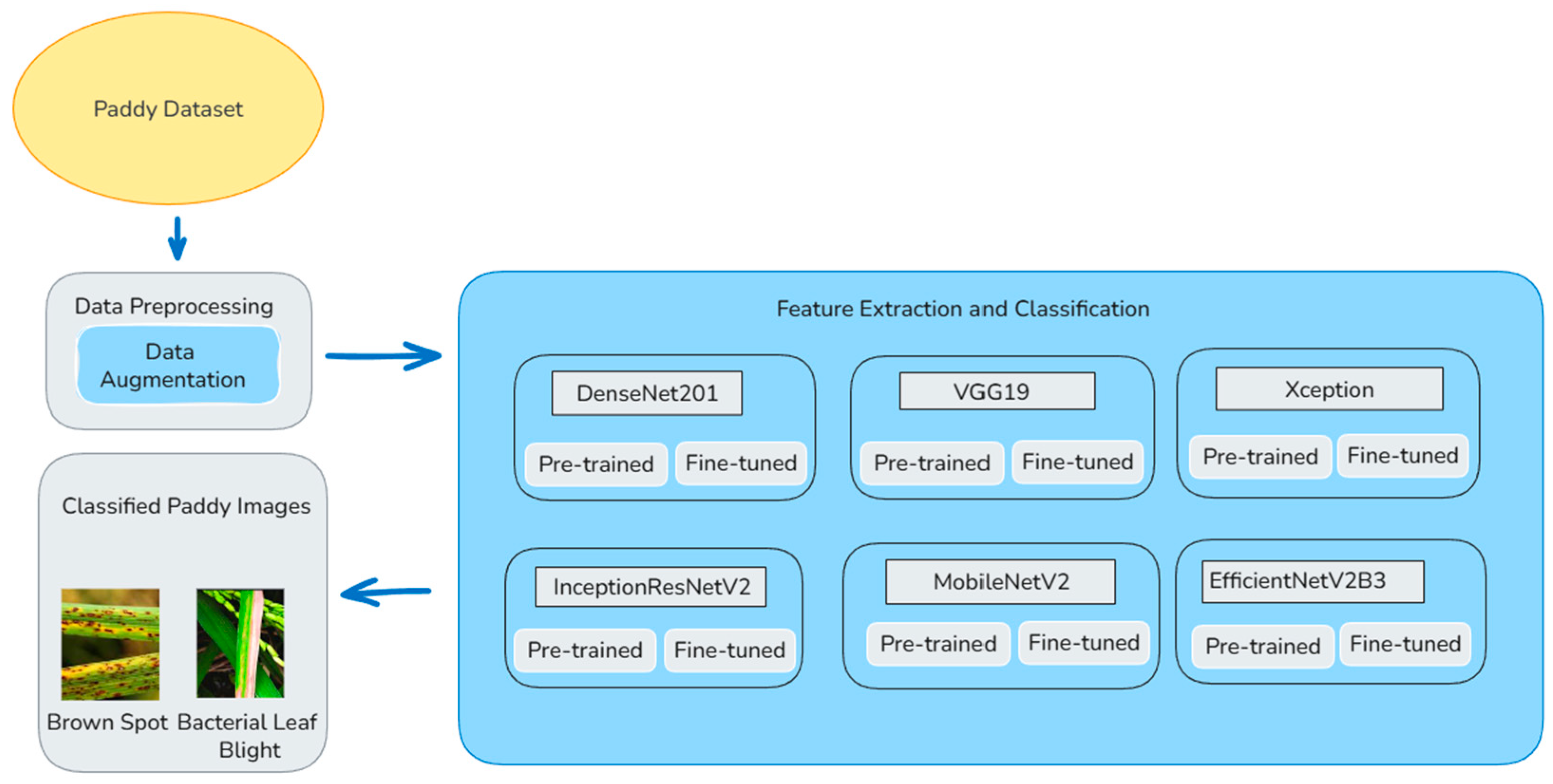
The overall classification of the paddy diseases is displayed in Figure 1. The size of the input images depends on which model is used. The size of the images should be 224 × 224 pixels when using models like DenseNet201, VGG19, and MobileNetV2. The size of the image should be 299 × 299 pixels when using models like Xception and InceptionResNetV2. The size of the images should be 300 × 300 pixels when using a model like EfficientNetV2B3. The model processes 32 images at once during training. The dataset carries 10 classes, adjusted based on the actual number of disease classes. To improve the model and avoid overfitting, data augmentation is done only on the training images using Image Data Generator. The images are rescaled from 0 to 255 to 0–1 for easier learning. The images are randomly rotated, moved sideways and up or down, slightly tilted, zoomed in, and flipped sideways. Also, 80% of the images are used for training and 20% for validation. The validation set is only rescaled to 0–1. This helps to check the performance of the model fairly. 20% of the total data is used for validation using validation split = 0.2. The training set uses data augmentation and shuffles the images to add variety. The validation set keeps the images unchanged. This helps the model to learn well and be tested fairly.
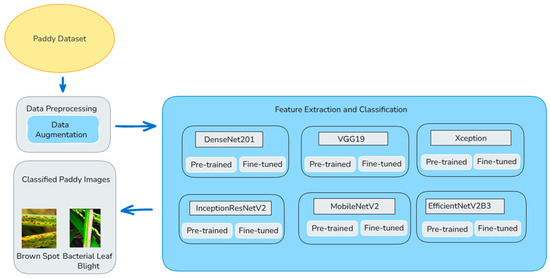
Figure 1.
Paddy disease classification using six deep learning models.
6.1. Hyperparameter Selection and Fine-Tuning Strategy
A batch size of 32 was chosen to strike a balance between the stability of training and computational requirements. The Adam optimizer was used due to its adaptive learning rates and stable convergence properties during image classification tasks. During the feature extraction stage, a learning rate of 0.001 was used to train the classifier layers added to the network. During the fine-tuning stage, the learning rate was set to 0.0001 to prevent sudden changes in the pre-trained weights.
A dropout rate of 0.3 was used to prevent overfitting while retaining sufficient representational power. The number of unfrozen layers in the fine-tuning process was set based on the depth of the architectures to allow higher-level task-specific features to adapt to the characteristics of paddy diseases while retaining the integrity of lower-level generic features as given in the Table 2.

Table 2.
Training and fine-tuning hyperparameters used for all models.
6.2. DenseNet201: Architecture and Working
DenseNet201 is a deep convolutional neural network. This improves feature learning by connecting each layer to every previous layer. This improves feature propagation and smooth gradient flow. Its architecture includes dense blocks, which are groups of convolutional layers where each layer connects to every previous layer, bottleneck layers to reduce the number of parameters, transition layers, which are the pooling layers between dense blocks to down sample feature maps, Global Average Pooling (GAP) that replaces fully connected layers to reduce parameters and overfitting, Softmax Classifier, which is the final layer that outputs class probabilities for classification. Its working before fine-tuning includes the following. The DenseNet201 model, which is trained on ImageNet is loaded by excluding the top classifier. The convolutional base is frozen in order to keep their weights unchanged. This allows it to act as a feature extractor. A new classifier has been added on top. It consists of GAP to reduce spatial dimensions and is followed by dense layers and dropout layers to prevent overfitting. The final layer is a Softmax classifier that detects the probability of each class. The model is compiled using the Adam optimizer and categorical cross-entropy loss. Early stopping is used to prevent overfitting. Its working after fine-tuning includes the following. After initial training, selective layers (after layer 600) are unfrozen. This allows the model to fine-tune deeper layers, keeping the earlier layers frozen in order to retain pre-trained knowledge. The learning rate is reduced for smooth convergence so that drastic updates of weights may be avoided. Fine-tuning improves the model’s discrimination capability for subtle disease patterns in paddy leaves with the added advantage of computational efficiency. This allows the model to adapt better for specific tasks, say, paddy leaf disease detection, with computational efficiency.
6.3. VGG19: Architecture and Working
VGG19 is one of those deep convolutional neural networks that have both a simple and effective architecture. It has a total of 19 layers, consisting of 16 convolutional layers and 3 fully connected layers. It has a very organized and sequential design consisting of convolutional layers followed by max pooling layers. Other key architectural elements include sequential convolutions that apply tiny filters to detect complex features, and max pooling layers that subsample feature maps to lower the calculations. Fully connected layers represent the normal dense layers used in classification. GAP is used instead of a fully connected layer in custom models to reduce overfitting. Softmax classifier outputs class probabilities for final predictions. Its working before fine-tuning (feature extraction phase) includes the following. The VGG19 model which is trained on ImageNet is loaded by excluding the top classifier. The convolutional base is frozen in order to keep their weights unchanged. A new classifier has been added. It consists of GAP that is followed by dense layers and dropout layers to improve generalization. The final Softmax layer detects the probability of each class. The model is compiled using the Adam optimizer with categorical cross-entropy loss. Early stopping is used to monitor validation loss to prevent overfitting. Its working after fine-tuning includes the following. To improve performance, the last layers (after layer 16) are unfrozen. It allows the model to adjust deeper feature representations while keeping earlier layers frozen to retain pre-trained knowledge. A lower learning rate is employed to avoid abrupt updates to the weights and to make the fine-tuning stable. The strategy promotes the capacity of the model to identify distinct features within paddy disease classification, aside from exacting computational procedures to accomplish the task.
6.4. Xception: Architecture and Working
Xception (Extreme Inception) is a deep CNN. This model works by removing the need for regular convolutions within the Inception module and satisfying the requirement by performing depthwise separable convolutions. Key components of the architecture include depthwise separable convolutions that factorize standard convolutions into depthwise, pointwise convolutions that improve efficiency, residual connections that help to stabilize deep network training and improve gradient flow, entry, middle and exit flow, the three sections that are divided by the network to process features hierarchically and GAP which reduces overfitting by averaging feature maps before classification. Its working before fine-tuning. The Xception model, which is trained on ImageNet is loaded by excluding the top classifier. The first 86 layers are frozen in order to keep their weights unchanged. A new classifier has been added. It consists of Global Average Pooling that is followed by two dense layers and dropout layers to prevent overfitting. The final Softmax layer detects the probability of every class. The model is compiled using the Adam optimizer with categorical cross-entropy loss. Early stopping monitors validation loss to prevent overfitting. Its working after fine-tuning includes the following. The last layers are unfrozen after layer 86 to improve performance. It allows the model to fine-tune deeper features while keeping the initial layers frozen to retain general feature representations. A lower learning rate is used to ensure stable weight updates and prevent overfitting. This fine-tuning strategy allows Xception to adapt its learned representations to capture paddy disease patterns while maintaining computational efficiency.
6.5. InceptionResNetV2: Architecture
InceptionResNetV2 is a hybrid deep learning architecture. InceptionResNetV2 inherits the best parts of both architectures. The architecture of InceptionResNetV2 contains Inception modules that apply a set of multiple convolutional filters in parallel for multiscale features, residual connections that add shortcut connections to eliminate vanishing gradi-ents during backpropagation, factorized convolutions that expedite computation by replacing large kernels with small ones, and GAP Layers that reduce spatial dimensions of images for classification. The working prior fine-tuning involves: The InceptionResNetV2 deep learning architecture that was trained on ImageNet is loaded by omitting the top classifier. The convolutional layers are frozen, meaning that during fine-tuning, all the values of the convolutional layers’ weights remain unchanged. There is a newly attached classifier. It consists of GAP that is followed by two dense layers and dropout layers to prevent overfitting. The final Softmax layer detects the probability of each class. The model is compiled using the Adam optimizer with categorical cross-entropy loss. Early stopping is used to halt training if validation loss does not improve. Its working after fine-tuning (unfreezing layers for training) includes the following. To improve performance, the last layers (after layer 682) are unfrozen. It allows the model to fine-tune deeper feature representations while keeping initial layers frozen to preserve pre-trained knowledge. A lower learning rate ensures smooth and stable weight changes without sudden changes. This fine-tuning strategy helps the model to adapt more effectively to the paddy disease classification task while maintaining efficiency.
6.6. MobileNetV2: Architecture and Working
MobileNetV2 is a lightweight convolutional neural network which is made for mobile and embedded applications. Its architecture improves efficiency using some key architectural characteristics. These include depthwise separable convolutions, which reduce computational cost by factorizing standard convolutions into depthwise and pointwise convolutions, inverted residual blocks, which use bottleneck layers to communicate while reducing the number of computations, linear bottlenecks, which resist the loss of information due to non-linearity in deeper layers and GAP which reduces the number of parameters before classification and enhances generalization. Its working before fine-tuning (feature extraction phase), which includes the following. A pre-trained MobileNetV2 model which is trained on ImageNet is loaded by excluding the top classifier. All convolutional layers are frozen in order to keep their weights unchanged. A new classifier has been added. It consists of GAP that is followed by two dense layers and dropout layers to prevent overfitting. The final Softmax layer detects the probability of each class. The model is compiled using the Adam optimizer with categorical cross-entropy loss. Early stopping is used to observe validation loss and prevent overfitting. Its working after fine-tuning (unfreezing layers for training) includes the following. To improve performance, the last layers (after layer 106) are unfrozen. This allows the model to fine-tune deeper feature representations while keeping earlier layers unchanged to maintain pre-trained knowledge. A lower learning rate confirms smooth weight changes without sudden changes. This fine-tuning strategy improves the ability of the model to acquire domain-specific features for paddy disease classification without disturbing computational efficiency.
6.7. EfficientNetV2B3: Architecture and Working
EfficientNetV2B3 is an advanced version of EfficientNet. It is improvised for rapid training and improved accuracy. It supports compound scaling that focuses on the balancing of depth, width and resolution simultaneously, which entails improved efficiencies and also reduces the computations needed. There are also fused MBConv layers that improve the network using fused convolutions rather than the normal depthwise convolutions. Progressive learning is also involved that trains the network using an escalating image size that boosts the learning of the features. Other advancements include the application of Squeeze and Excitation (SE) Blocks that improve feature attention in the channels and also the use of GAP that boosts the feature abstraction and reduces the dimensions of the vector.
A description of the working of the code even prior to the fine-tuning phase is provided below. A pre-trained EfficientNetV2B3 is utilized that is pre-trained on the ImageNet dataset and is excluded from the top classifier. All convolutional layers are frozen in order to keep their weights unchanged. A new classifier has been added. It consists of Global Average Pooling GAP that is followed by two dense layers and dropout layers to prevent overfitting. The final Softmax layer detects the probability of each class. The model is compiled using Adam and categorical cross-entropy loss. Early stopping is used to prevent overfitting. Its working after fine-tuning includes the following. To improve performance, the last layers (after layer 350) are unfrozen. It allows deeper feature adaptation while keeping early layers frozen for general feature extraction. A lower learning rate ensures stable updates. This fine-tuning approach helps the model to refine its learned representations for paddy disease classification and optimizes both speed and accuracy.
7. Results and Discussion
DenseNet201:
Before fine-tuning
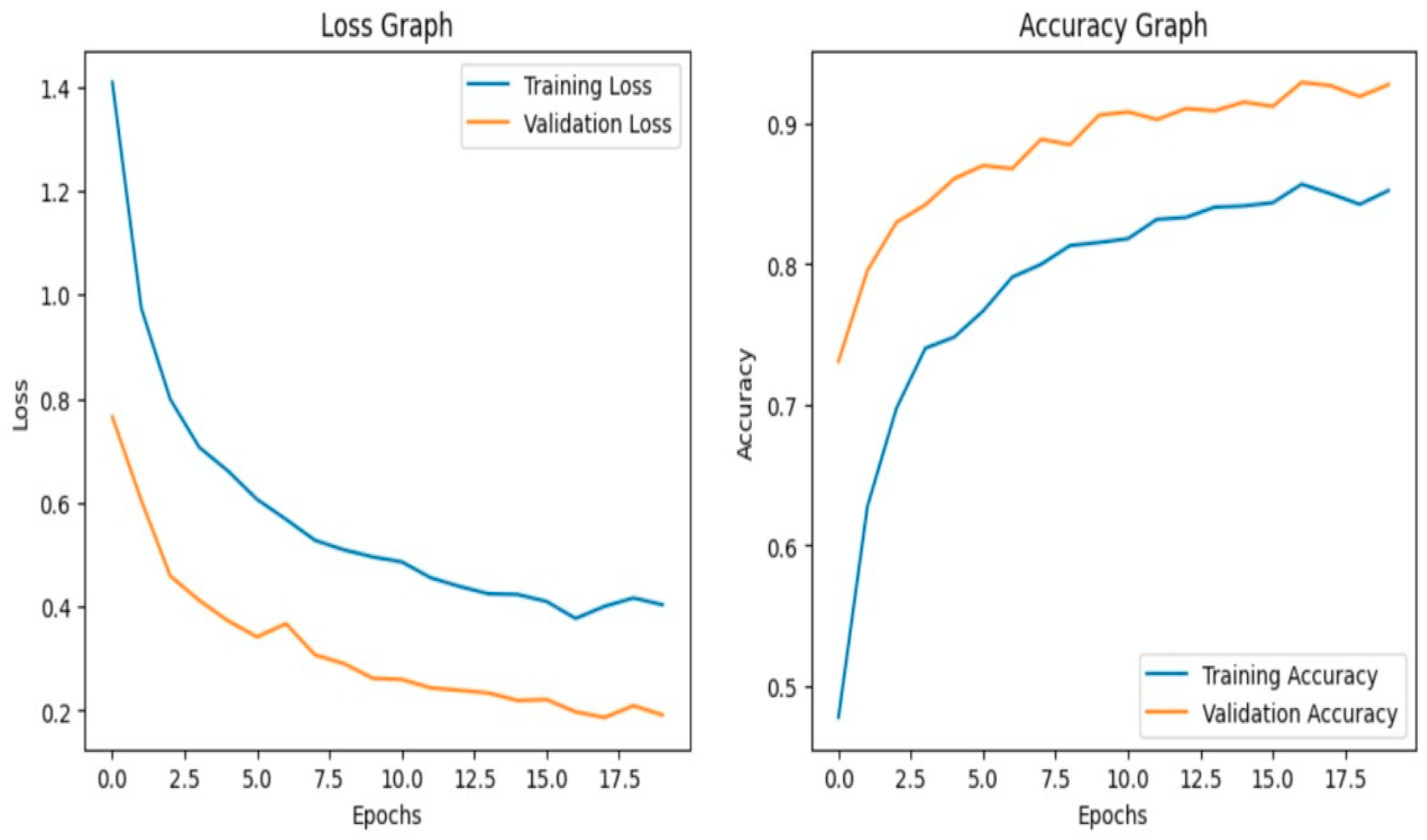
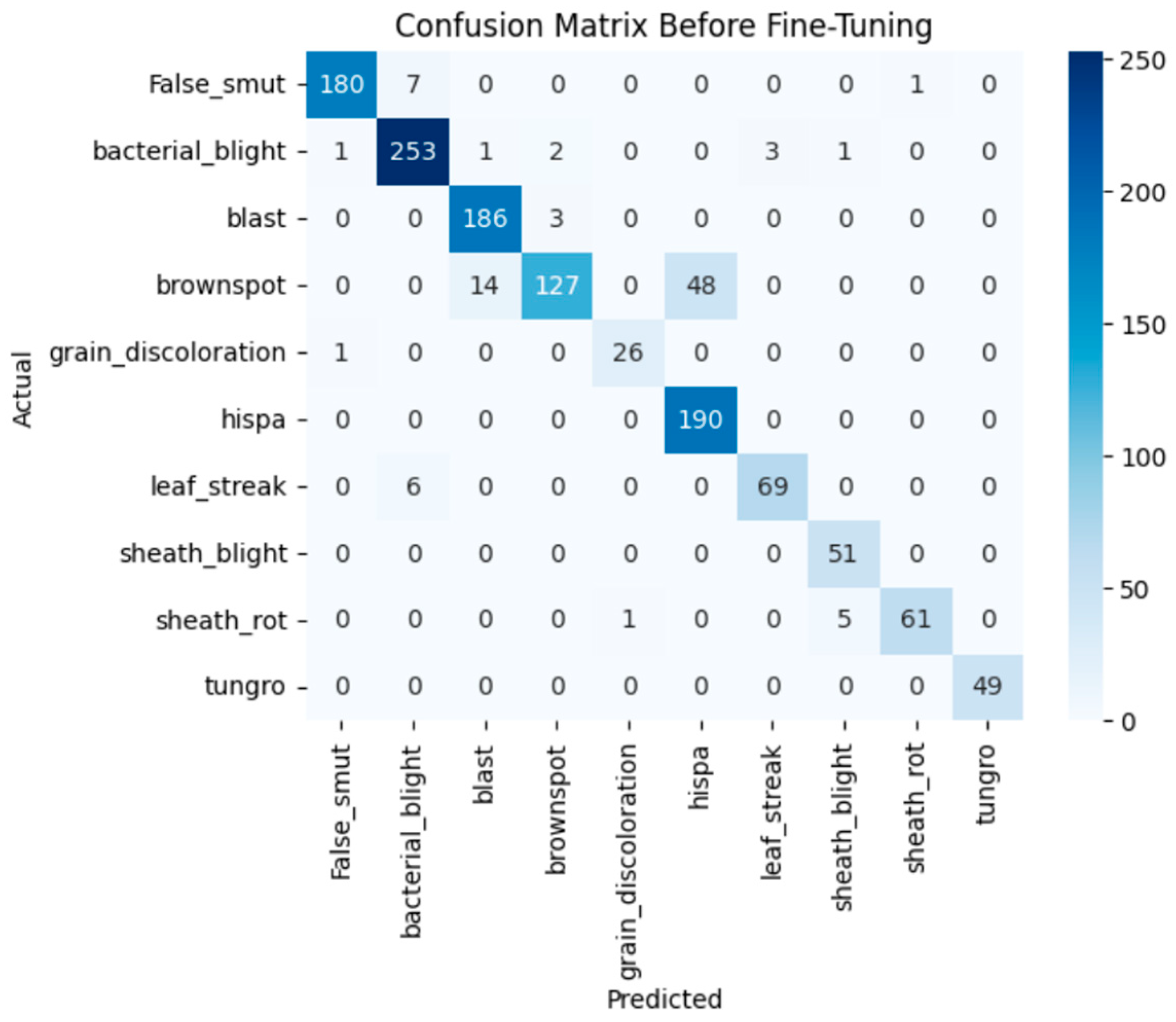
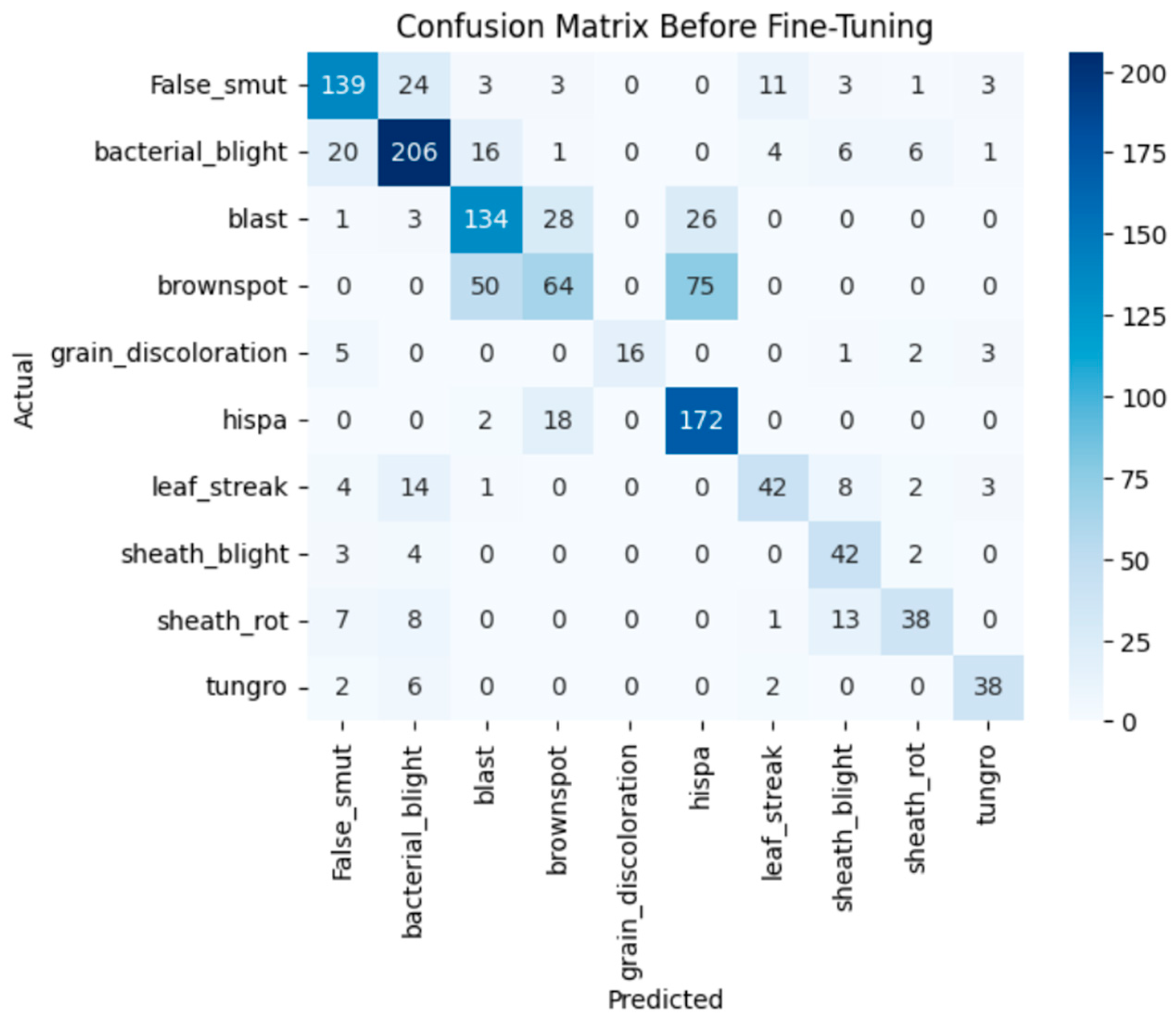
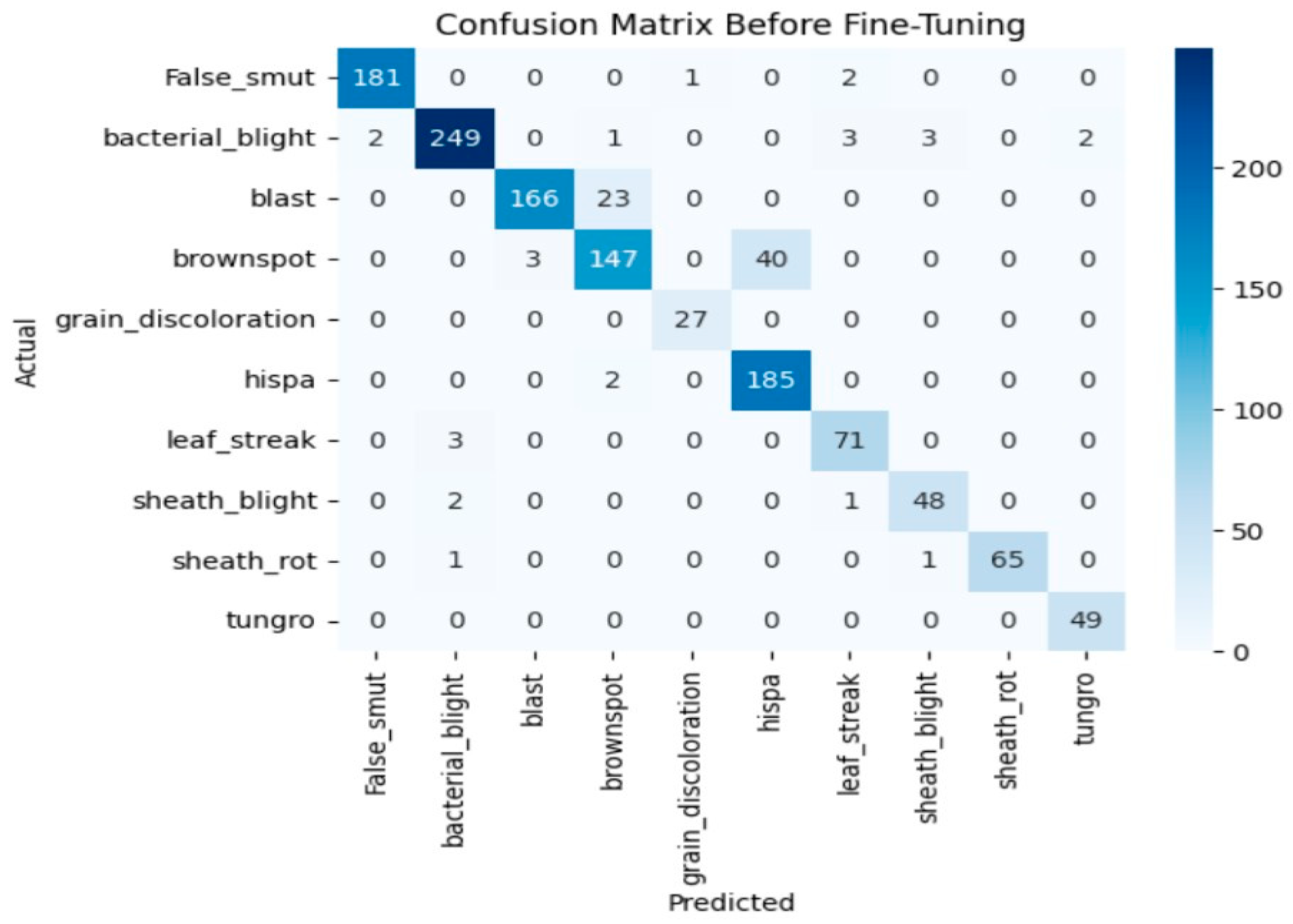
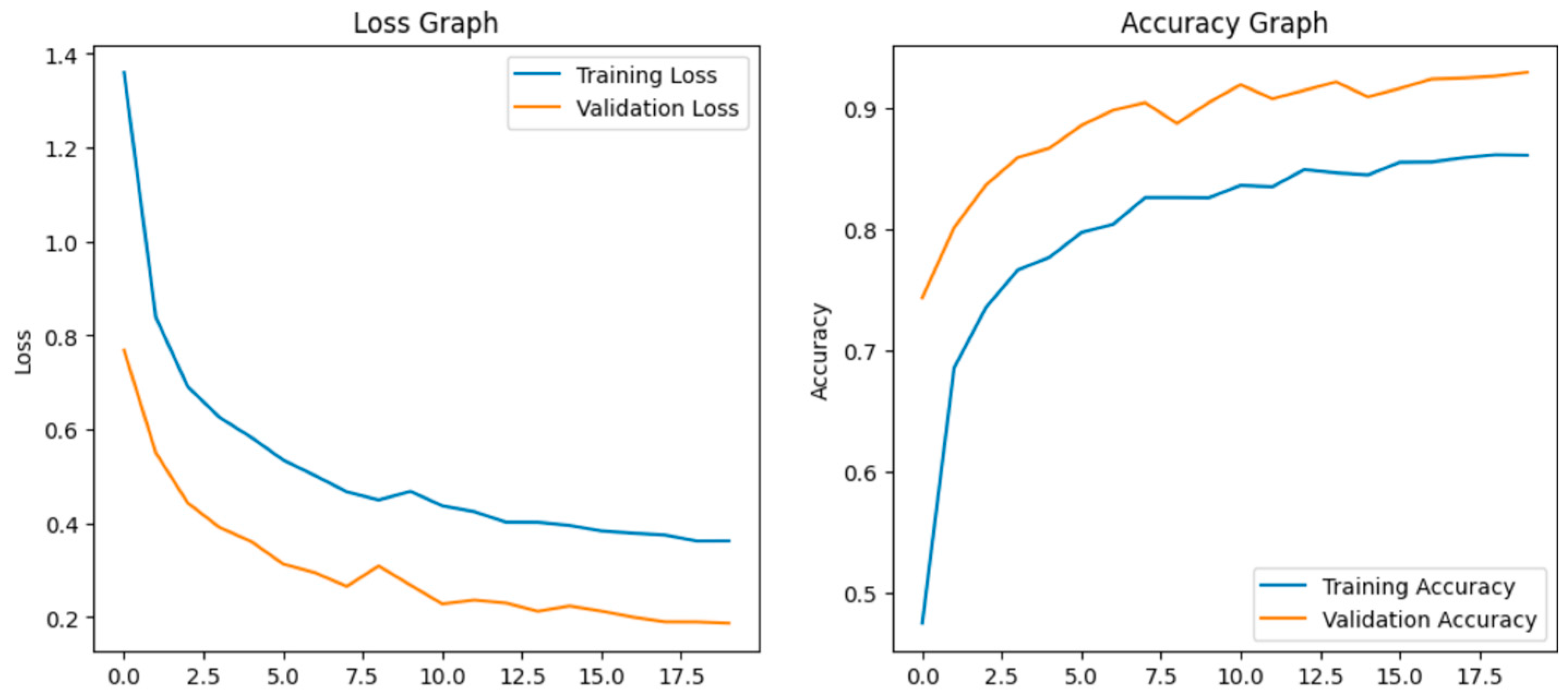
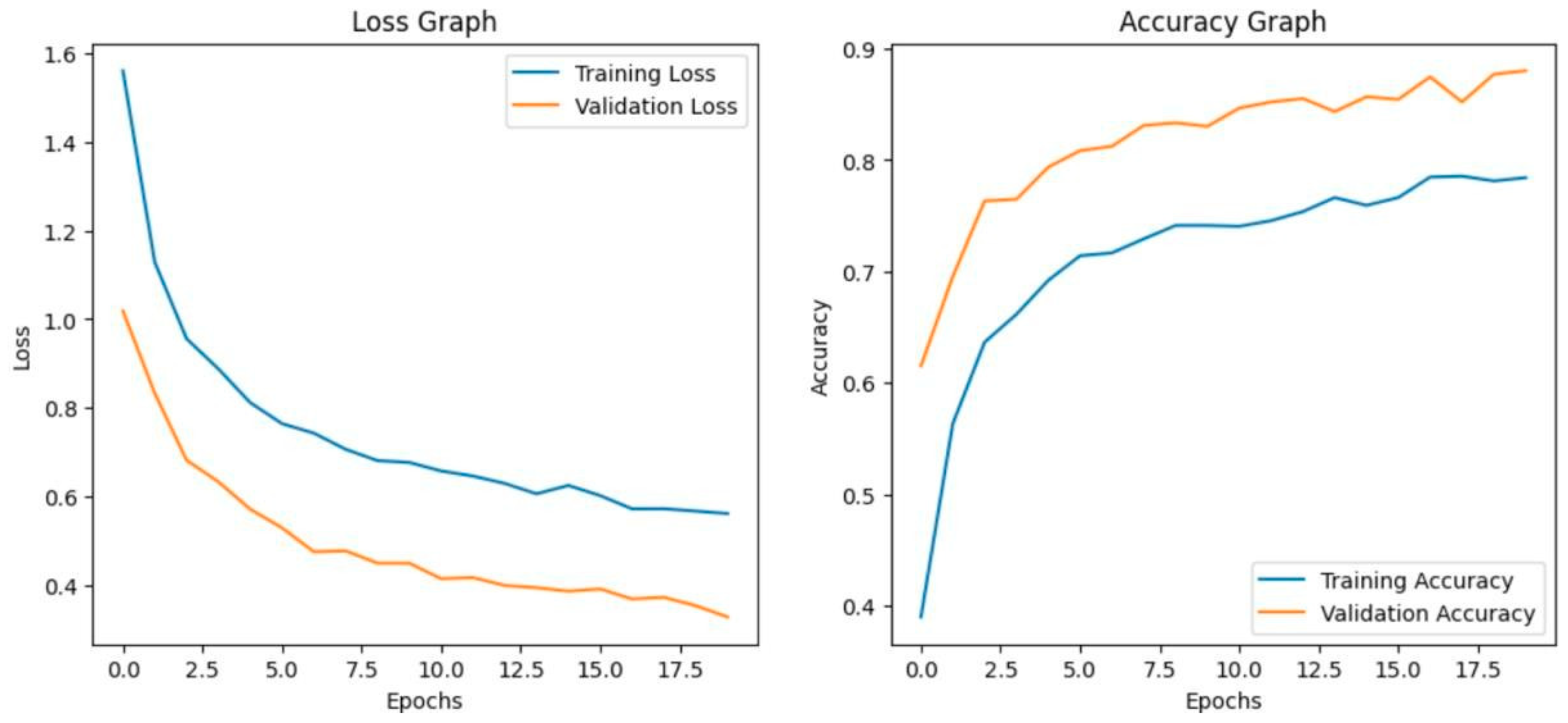
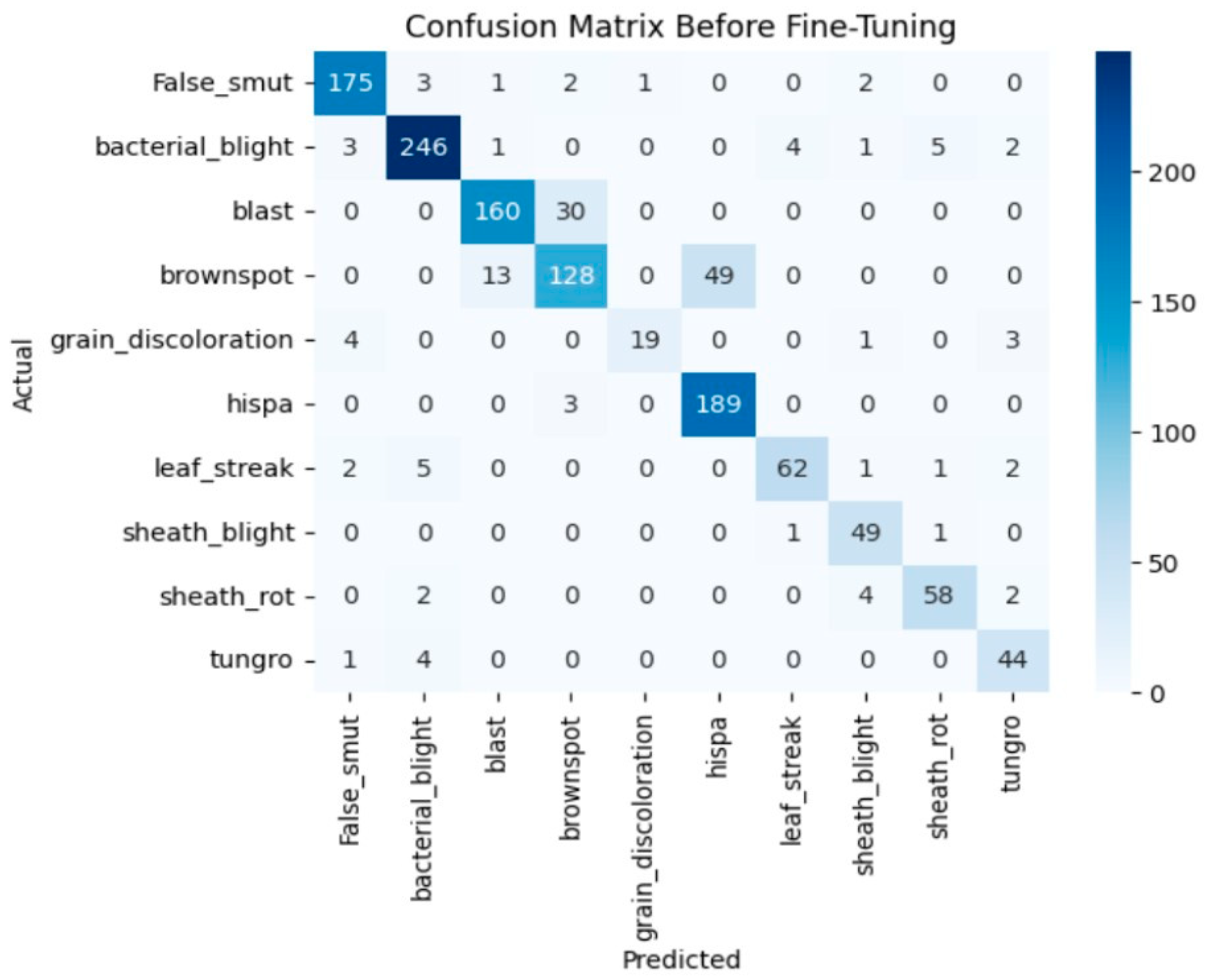
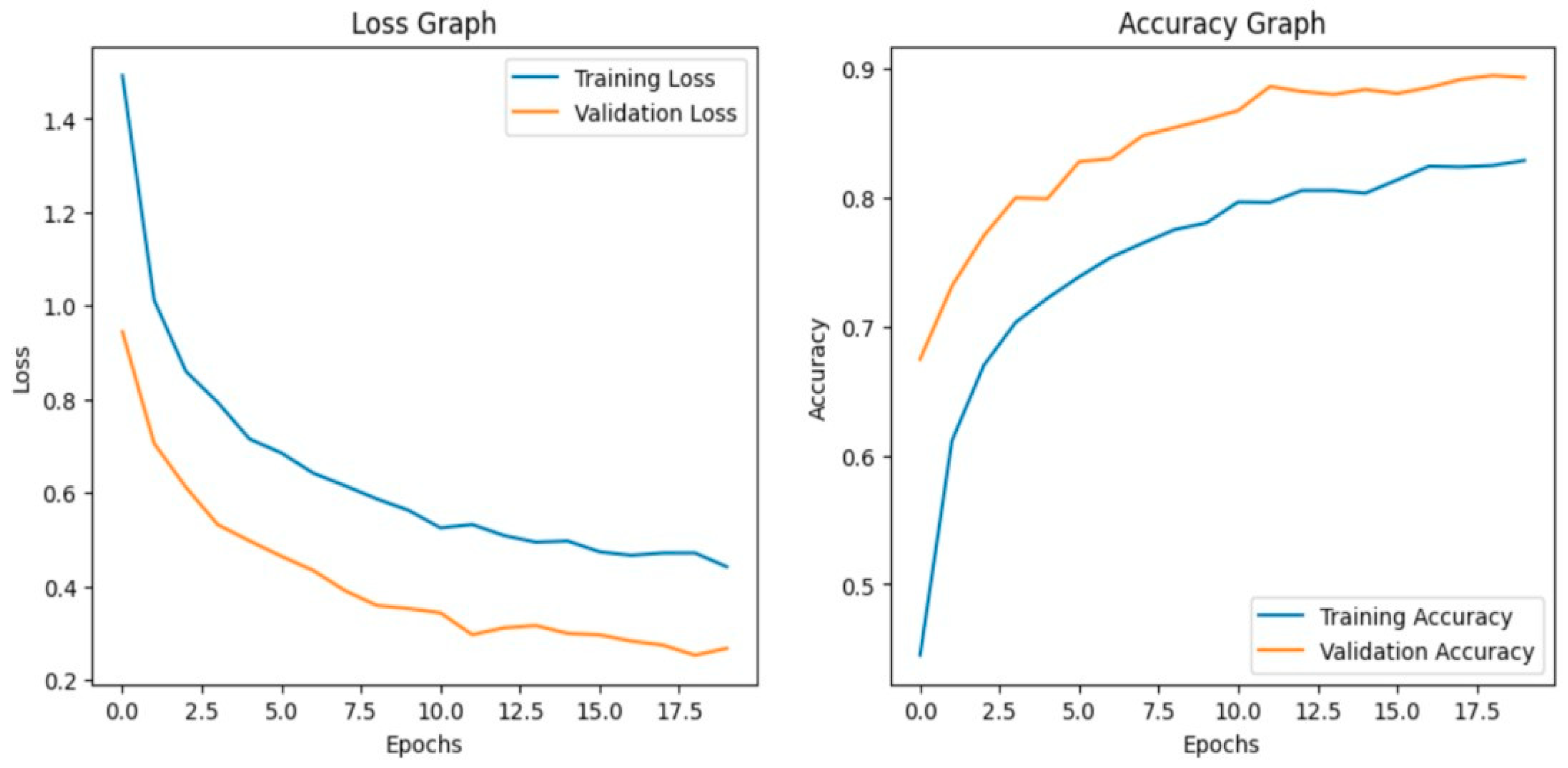
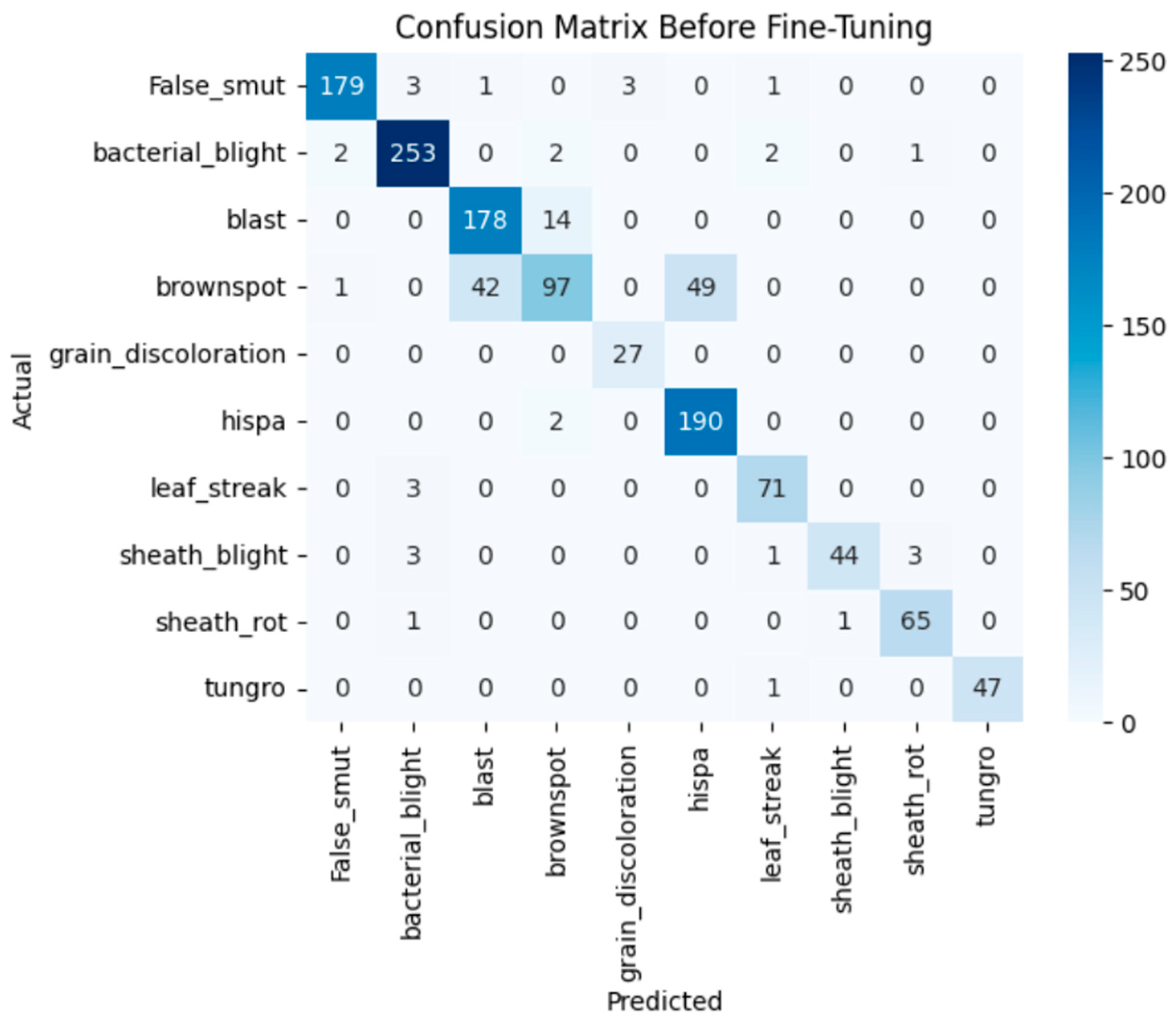
The model achieves 93% accuracy before fine-tuning. The accuracy graph and loss graph before the DenseNet201 fine-tuning are displayed in Figure 2. Training and validation losses decrease slowly and accuracy improves steadily. These changes show effective learning. Most classes are classified accurately but there are some misclassifications in “brown spot” and “hispa”. The confusion matrix confirms the class overlaps. The confusion matrix is shown in Figure 3. Therefore, the model is reliable but could benefit from fine-tuning to improve specific class performance.
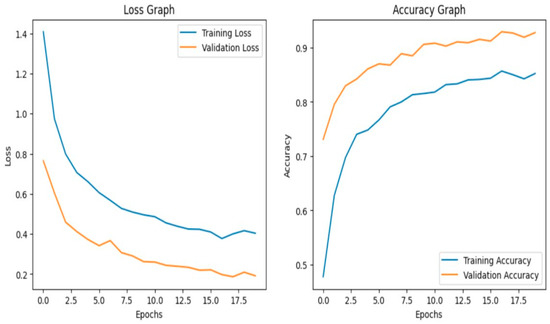
Figure 2.
Loss graph and Accuracy graph of the DenseNet201 before fine-tuning.
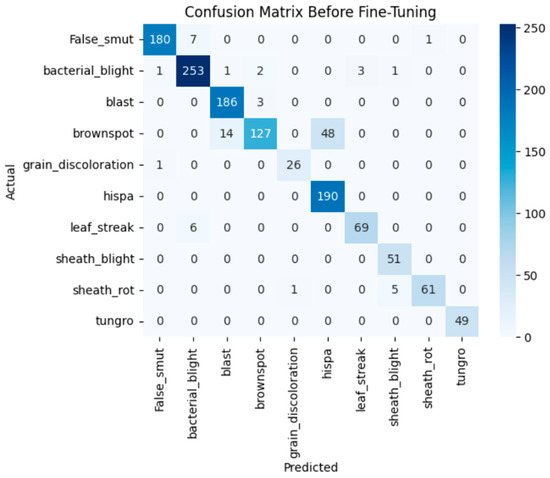
Figure 3.
Confusion matrix of the DenseNet201 before fine-tuning.
After fine-tuning:
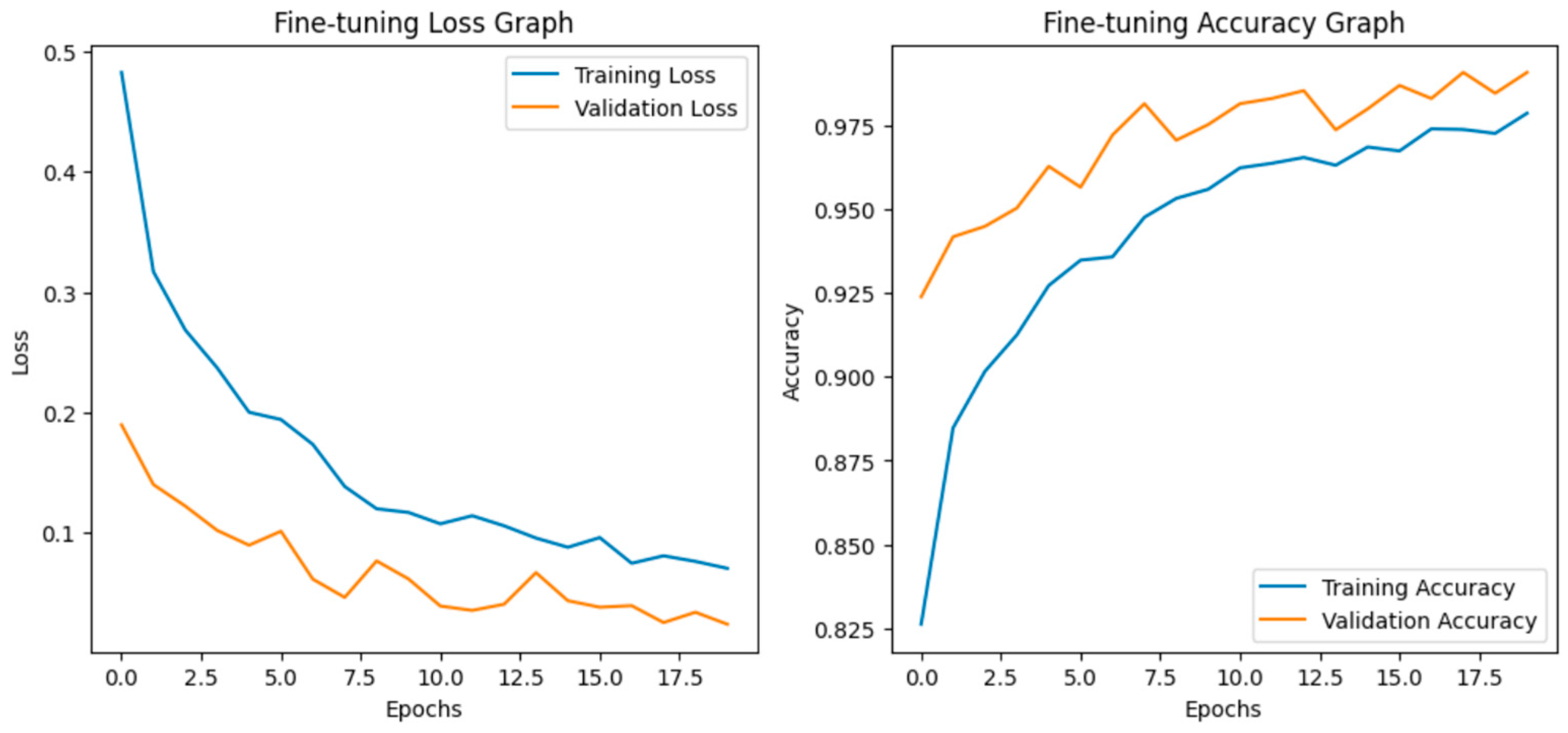
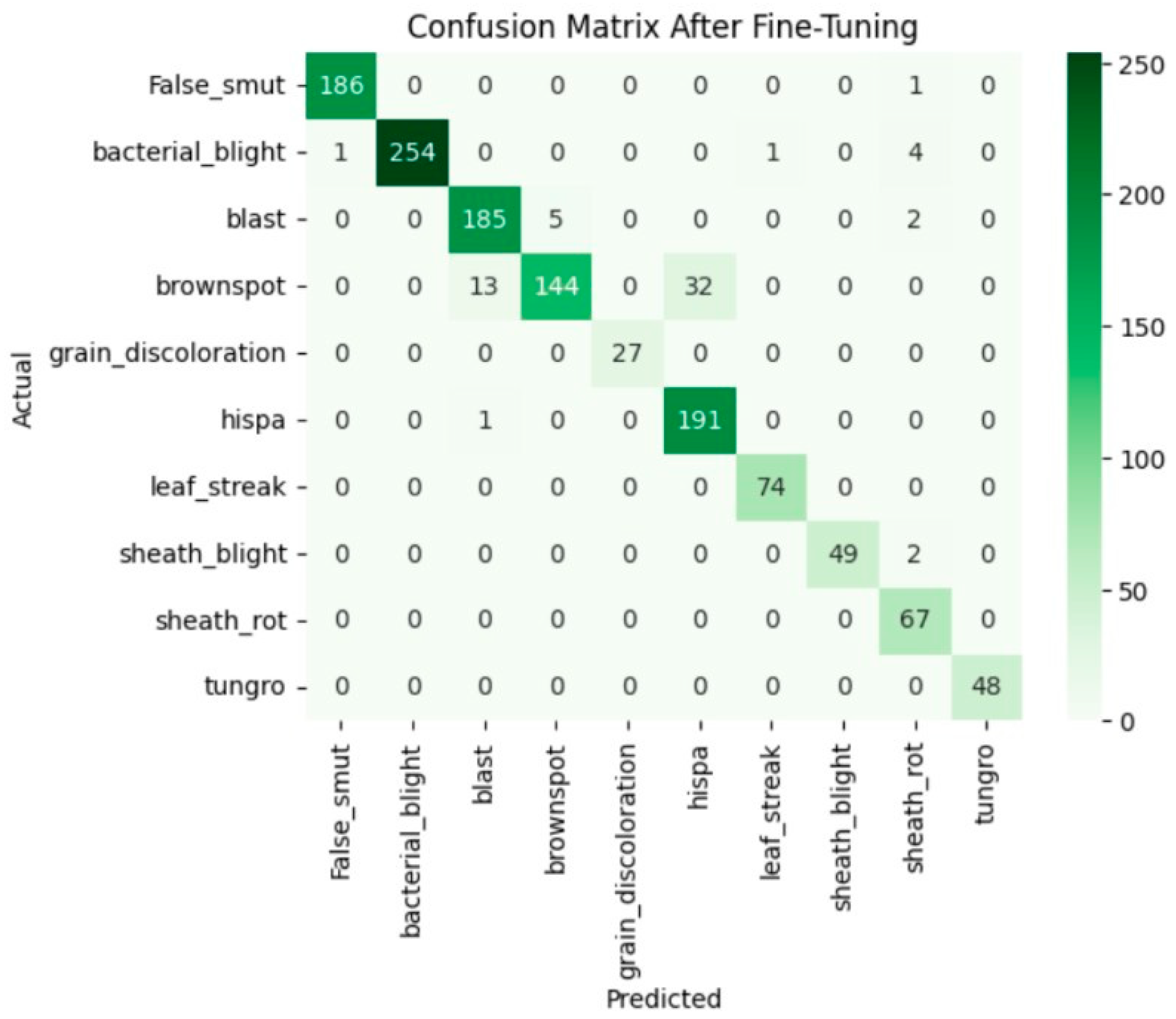
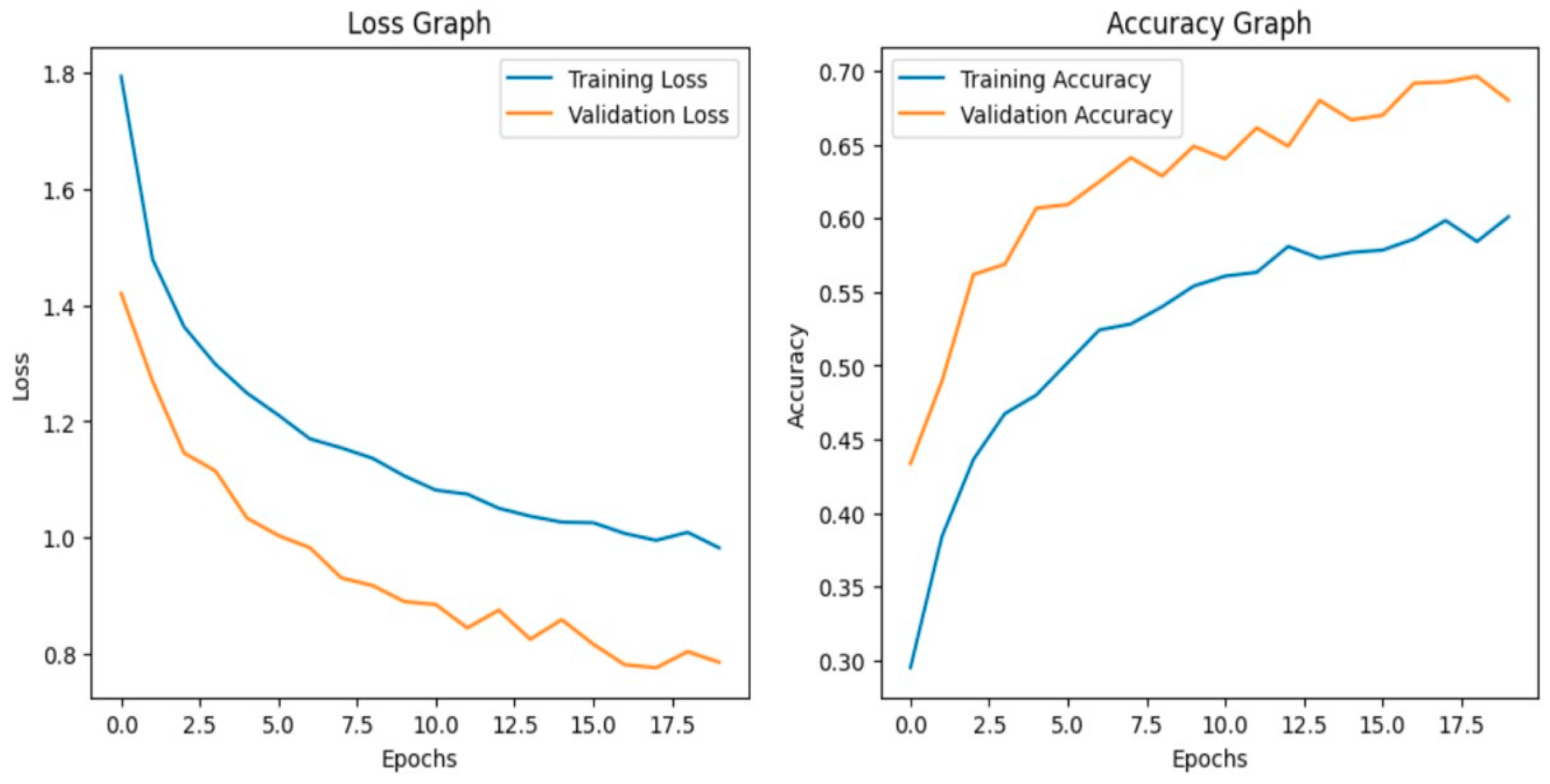
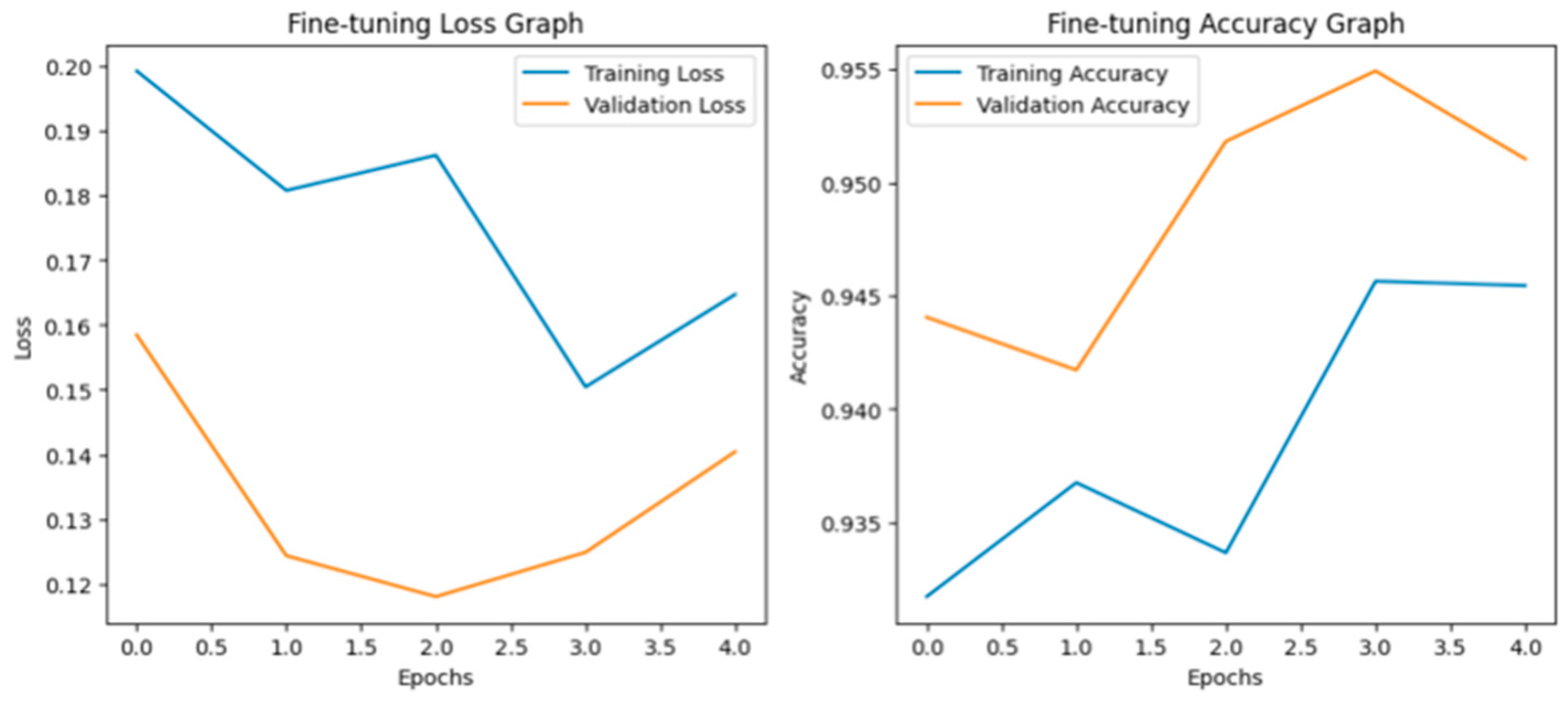
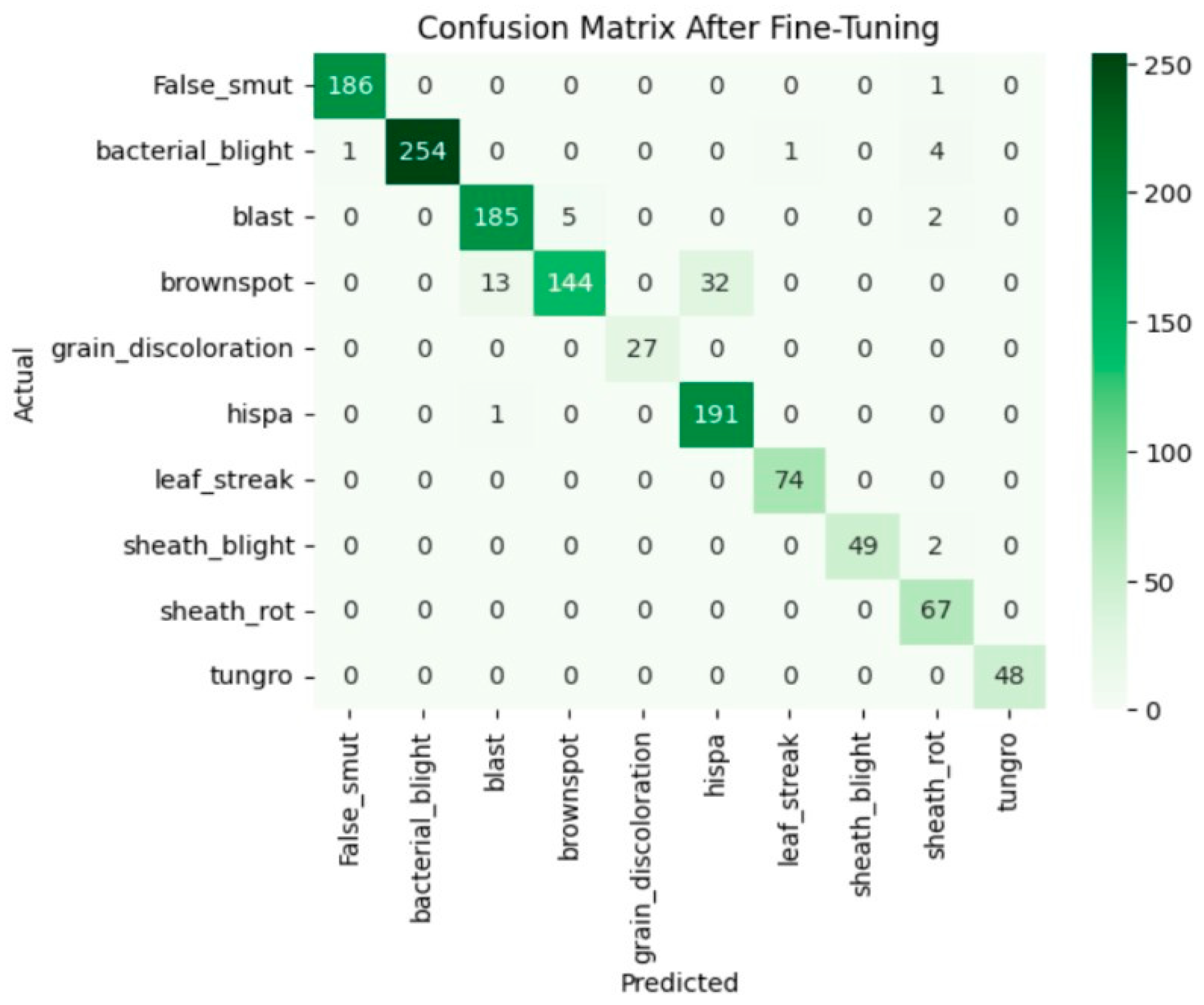
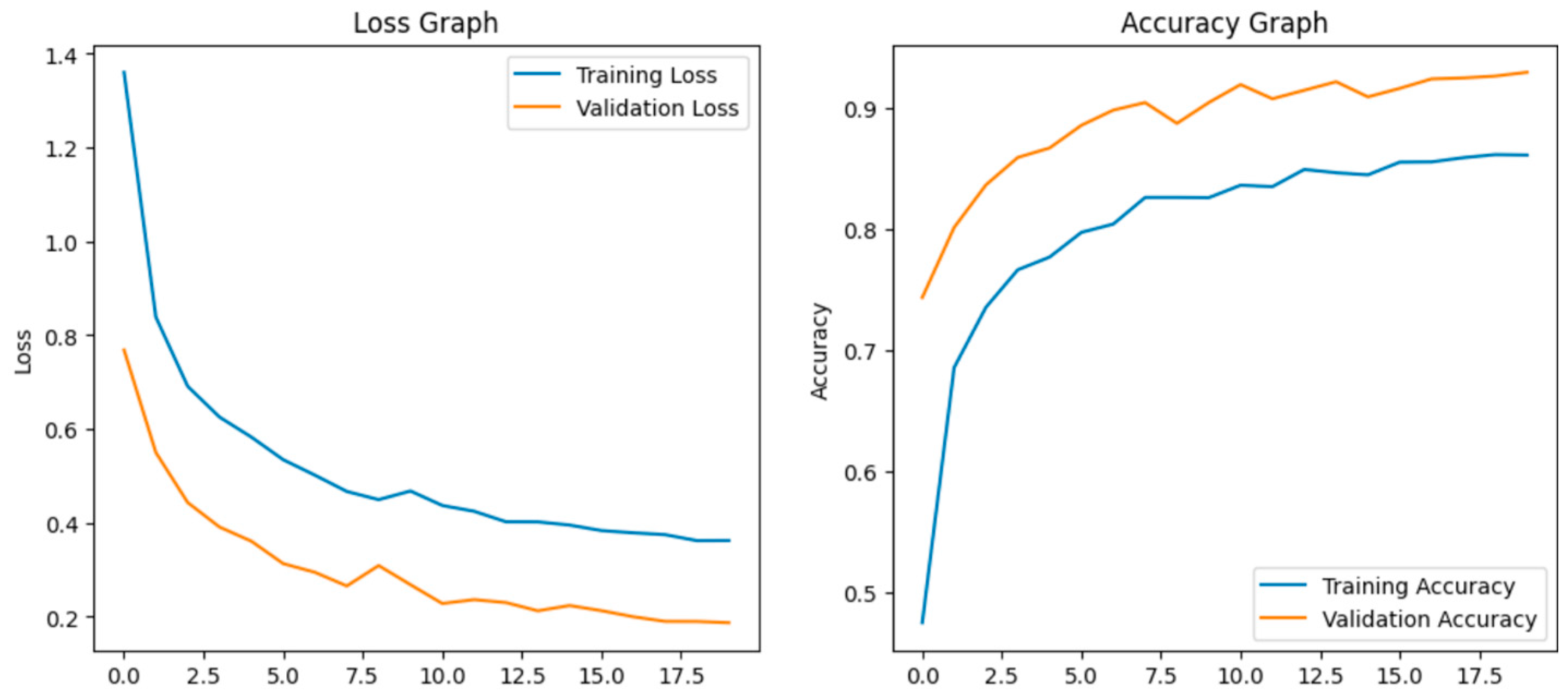
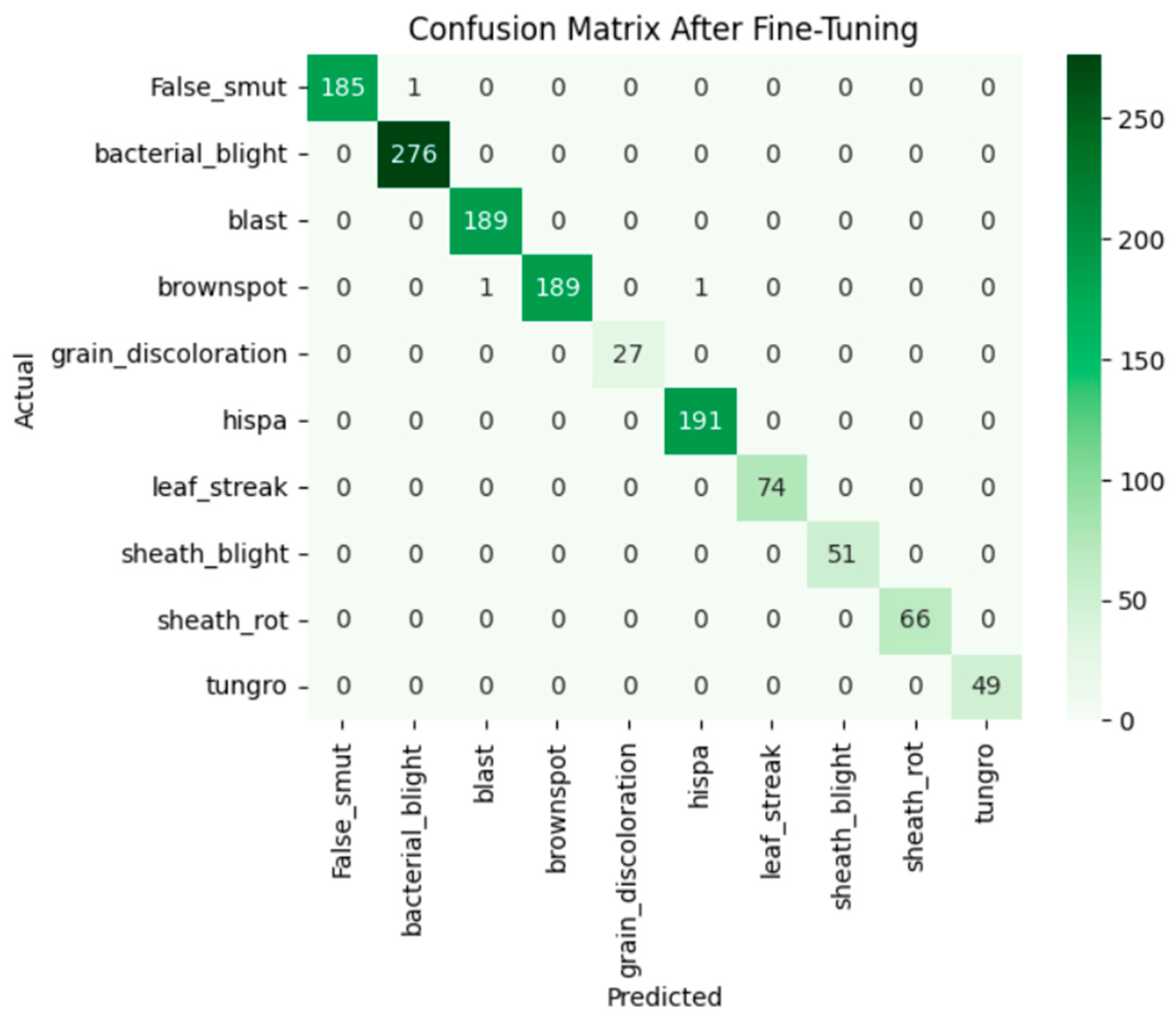
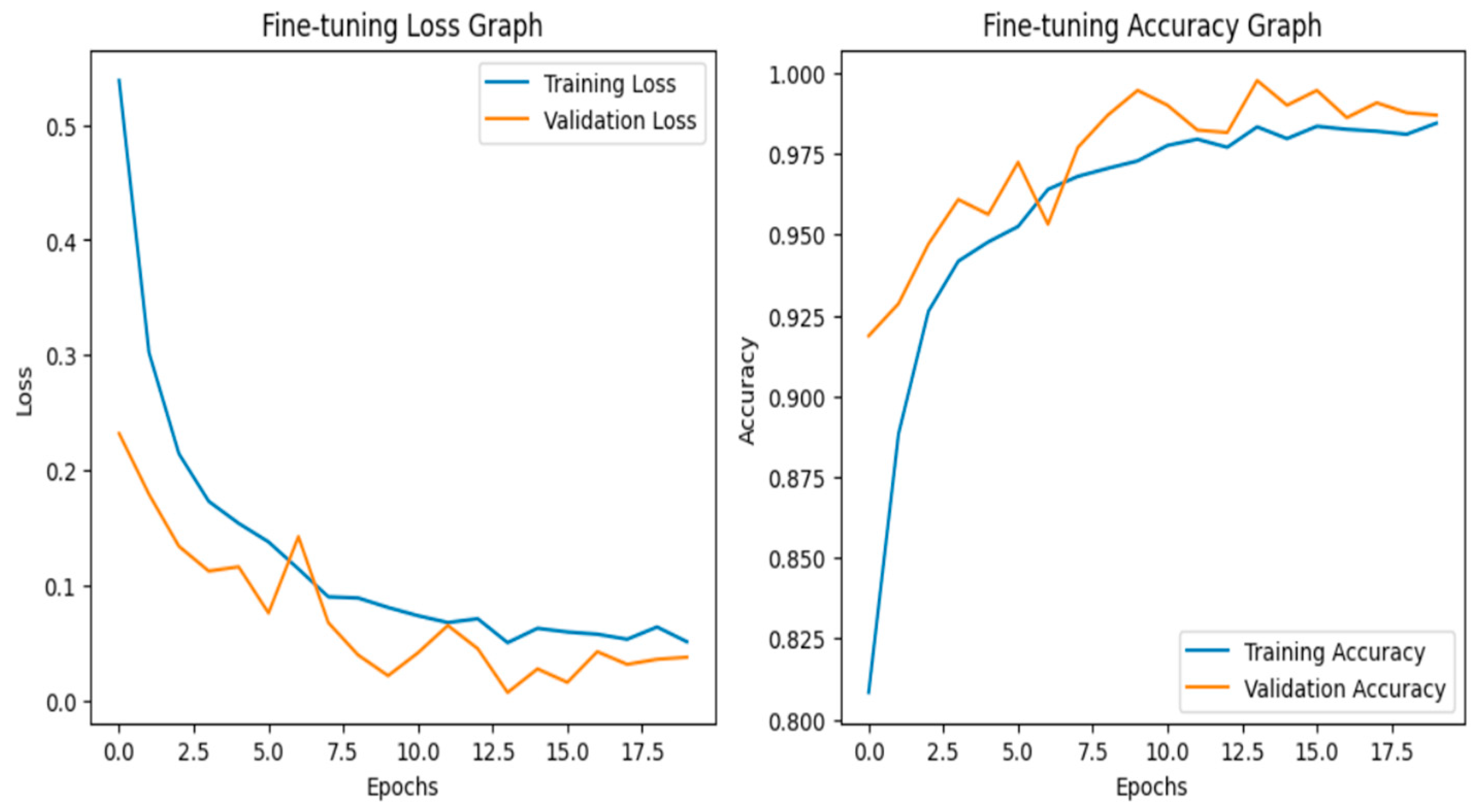
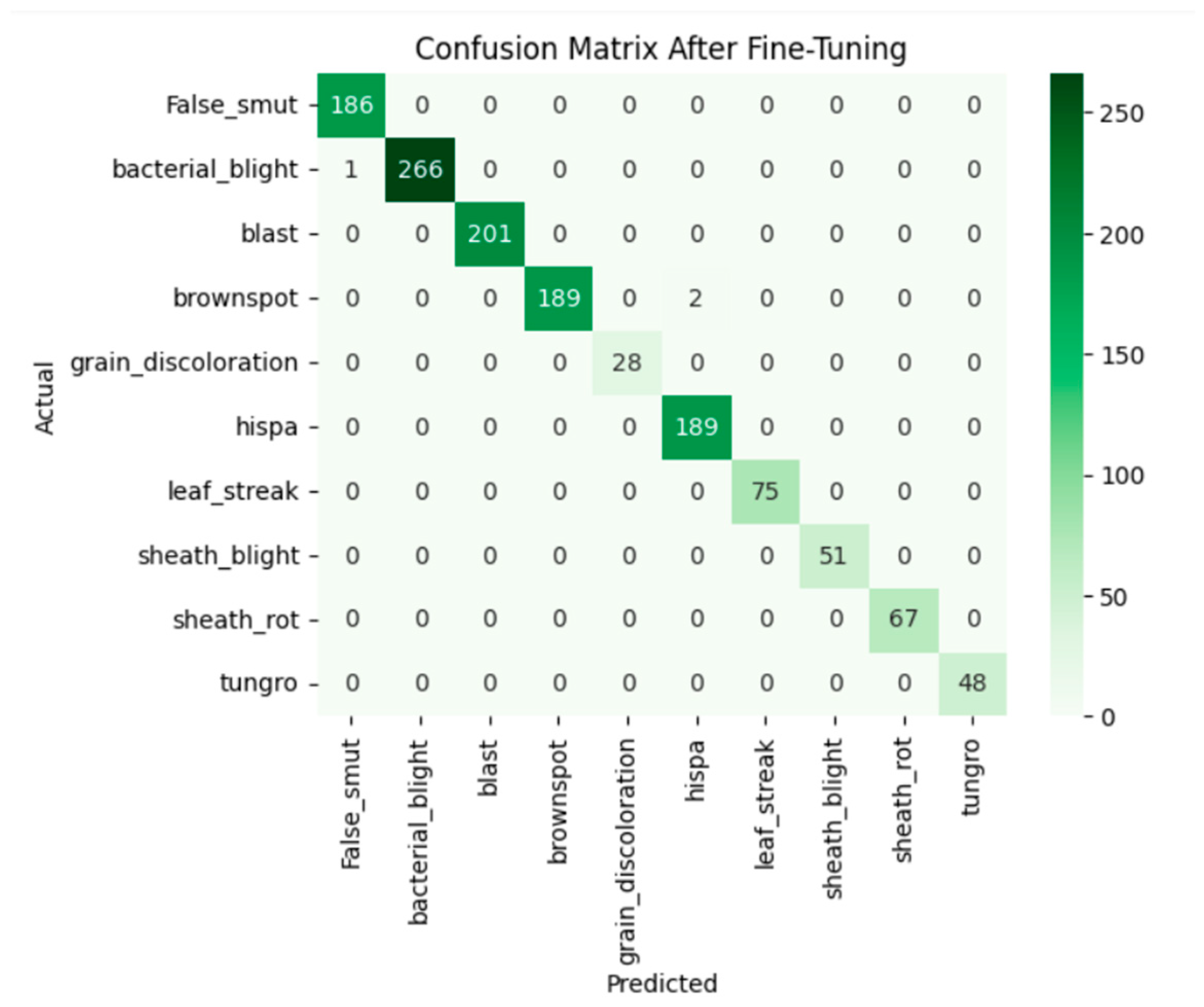
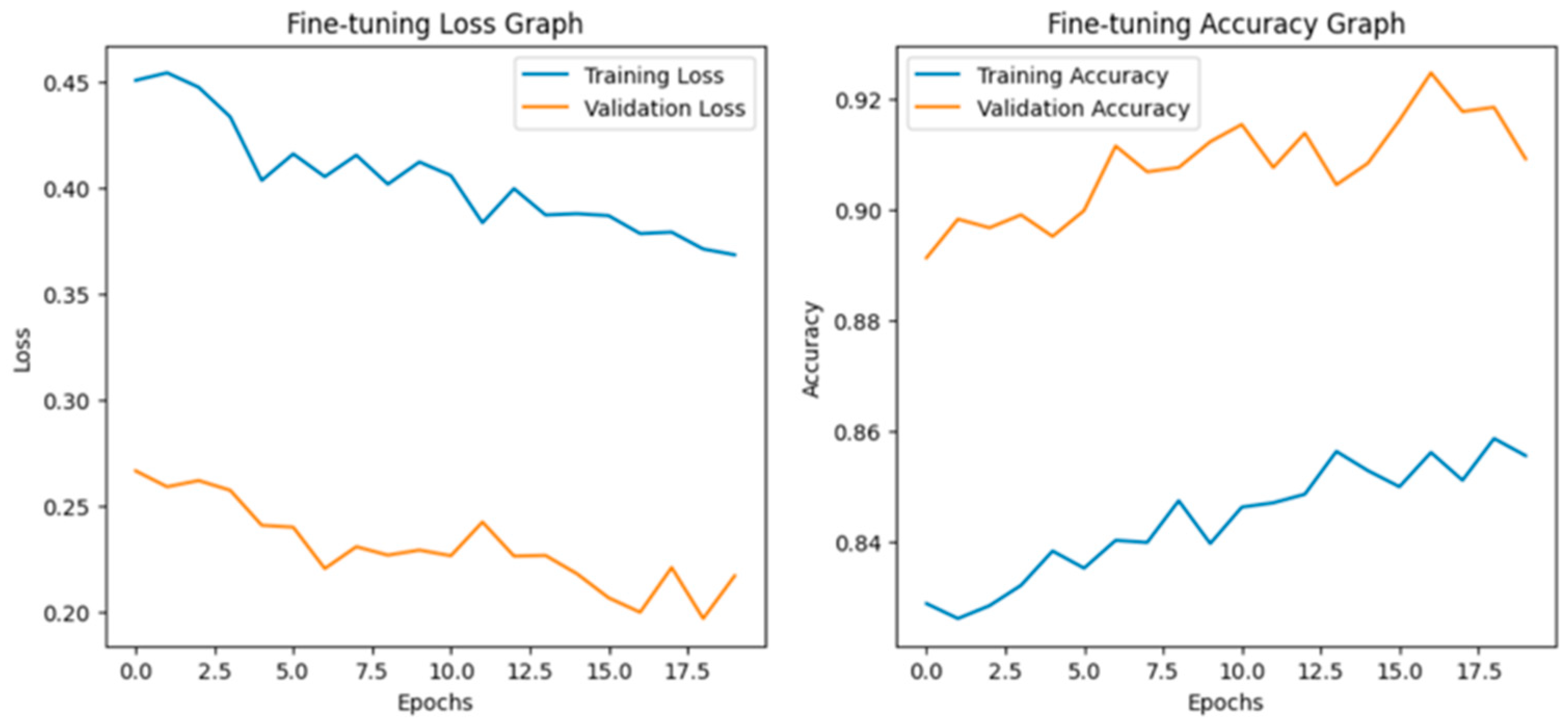
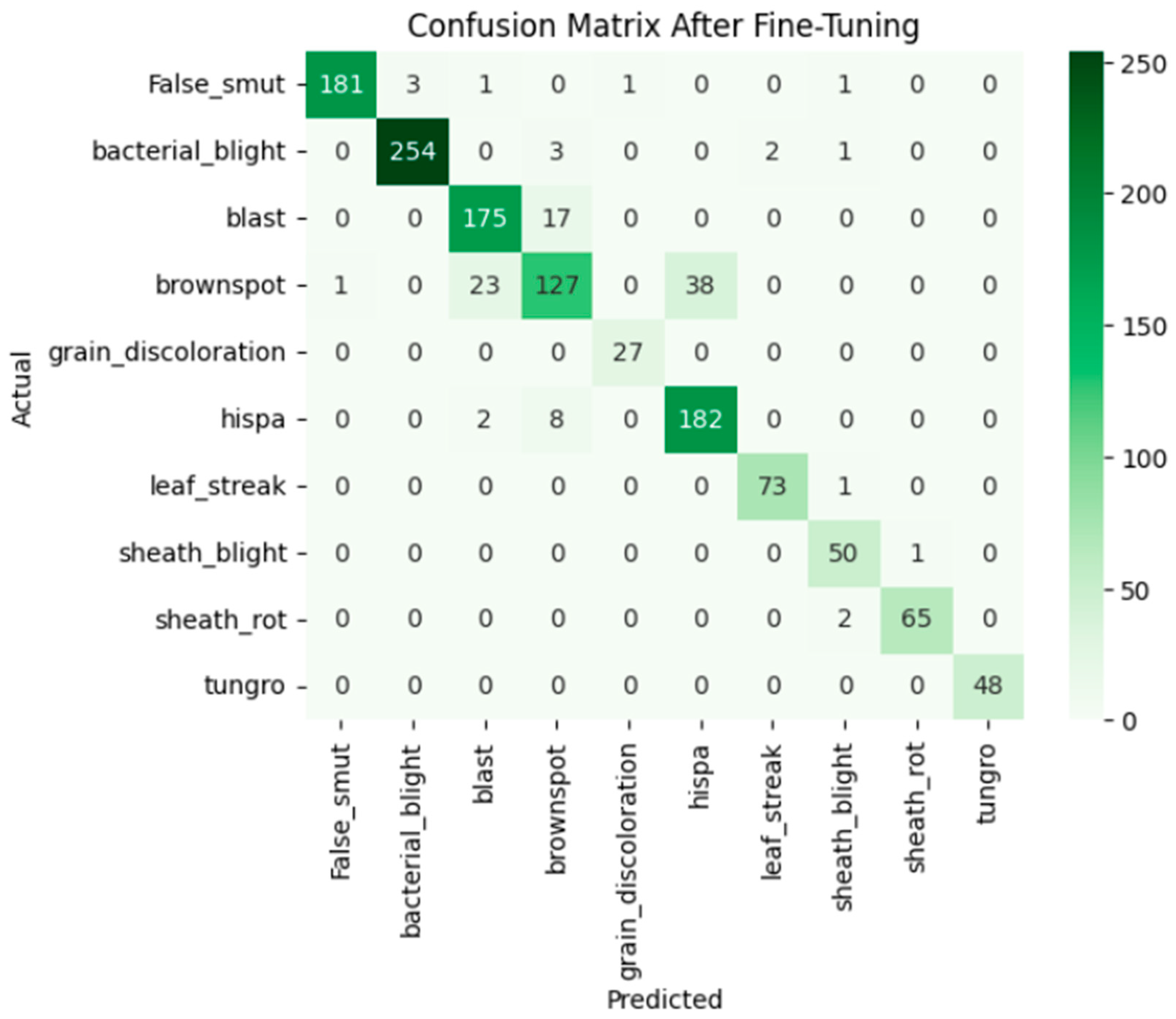
Fine-tuning significantly enhanced the performance of the model. The accuracy graph and loss graph of the DenseNet201 after fine-tuning are displayed in Figure 4. This increases the overall accuracy from 93% to 95%. It led to noticeable improvements in precision, recall, and F1-scores across most classes and indicates better classification reliability. The confusion matrix shown in Figure 5 demonstrates reduced misclassifications indicating that the model has become more effective at distinguishing between different disease categories. Therefore, fine-tuning contributed accurate model.
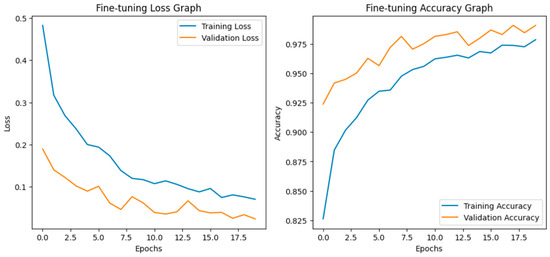
Figure 4.
Loss graph and accuracy graph of the DenseNet201 after fine-tuning.
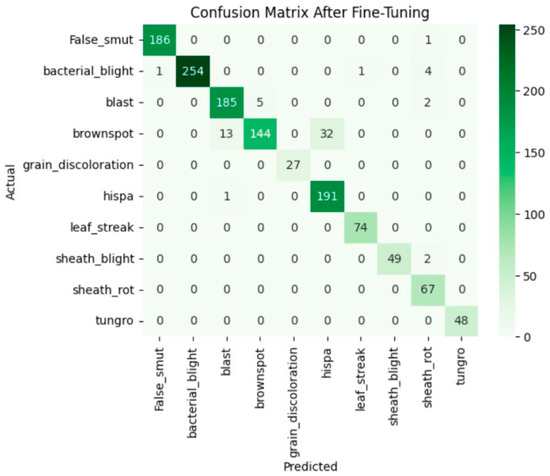
Figure 5.
Confusion matrix of the DenseNet201 after fine-tuning.
VGG19: Architecture and Working:
Before fine-tuning
Before fine-tuning, the VGG19 model shows fair performance in classifying the custom paddy disease dataset. With the deeper convolutional layers frozen, the model leveraged mainly generic ImageNet features and failed to capture the disease-specific texture patterns along with fine-grained visual features of the diseases. It achieved an overall validation accuracy of 69%, with a macro-average F1-score of 0.69. From Figure 6, although both the training and validation loss curves converge gradually, there is significant misclassification between visually similar disease categories. The associated confusion matrix in Figure 7 illustrates confusion between closely related classes such as brown spot, blast, and hispa.
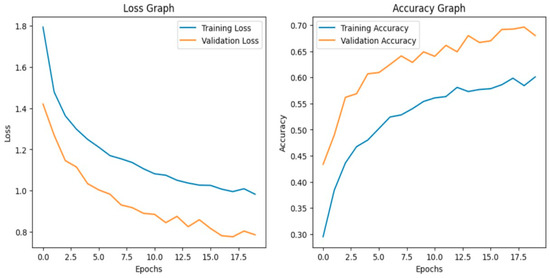
Figure 6.
Loss graph and accuracy graph of the VGG19 before fine-tuning.
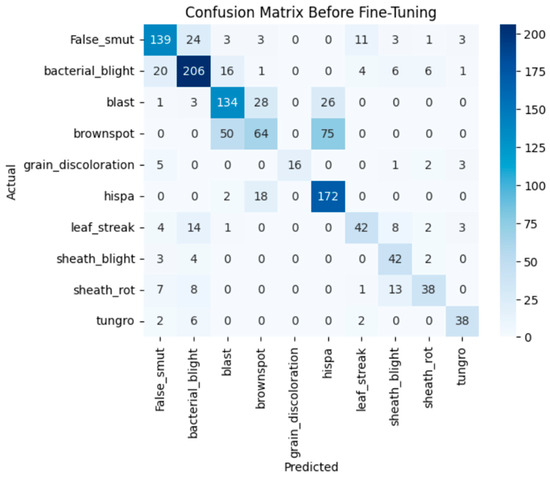
Figure 7.
Confusion matrix of the VGG19 before fine-tuning.
After fine-tuning:
Fine-tuning significantly improved VGG19 classification performance due to the adaptation of higher convolutional layers toward domain-specific features. It obtained an overall validation accuracy of 95%, with a macro-average F1-score of 0.96, showing strong and balanced performance across all the classes of diseases. In Figure 8, the accuracy and loss curves reveal stable convergence and decreased generalization error after fine-tuning. Moreover, the confusion matrix in Figure 9 shows significant reductions in the inter-class confusion of minority and visually similar classes. The obtained results confirm that fine-tuning enables VGG19 to learn effective discriminative patterns from the custom paddy disease dataset.
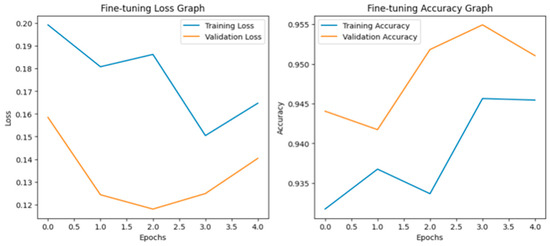
Figure 8.
Loss graph and accuracy graph of the VGG19 after fine-tuning.
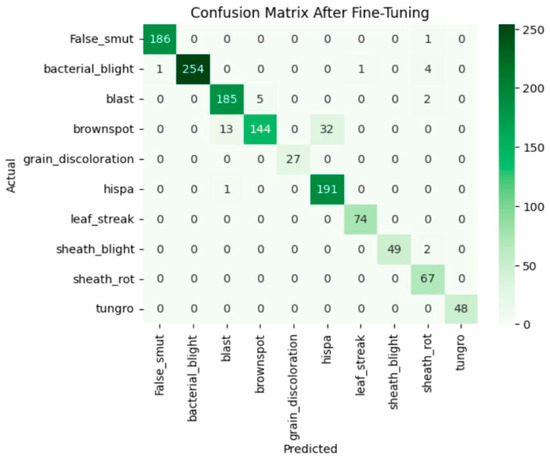
Figure 9.
Confusion matrix of the VGG19 after fine-tuning.
Xception: Architecture and Working
Before fine-tuning:
Before fine-tuning, the Xception model showed moderate performance with training accuracy of 65% and validation accuracy of 80% which indicates overfitting. The accuracy graph and loss graph of the Xception before fine-tuning are displayed in Figure 10. Training loss decreased gradually but validation loss remained high showing limited generalization. Performance across classes varied due to visual similarity as shown in Figure 11.
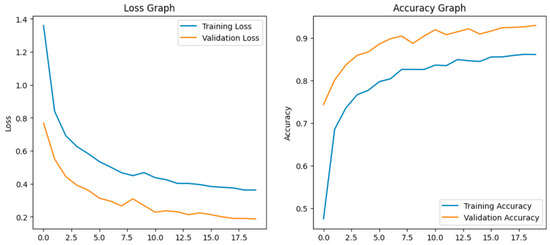
Figure 10.
Loss graph and accuracy graph of the Xception before fine-tuning.
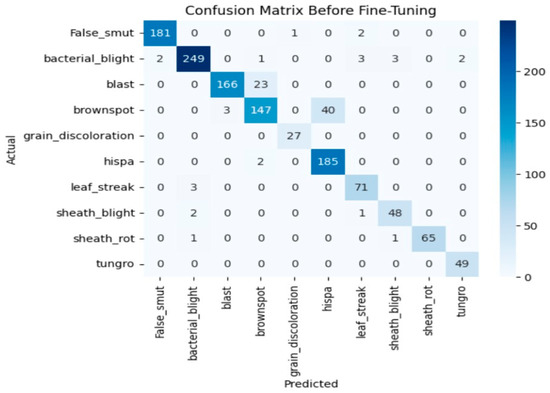
Figure 11.
Confusion matrix of the Xception before fine-tuning.
After fine-tuning:
After fine-tuning, the model adapted well. The training and validation accuracy rapidly increases and loss curves show a consistent decline. The accuracy graph and loss graph of the Xception after fine-tuning are displayed in Figure 12. Precision, recall, and F1-scores were balanced and high across all classes. The confusion matrix exhibits minimal misclassification. The confusion matrix shown in Figure 13 exhibits minimal misclassification, confirming strong model generalization and optimal performance.
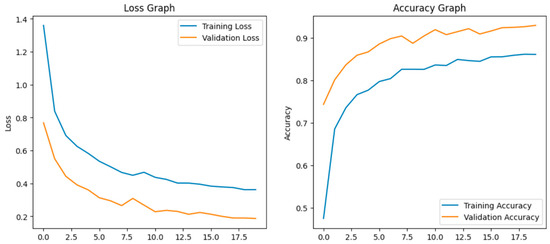
Figure 12.
Loss graph and accuracy graph of the Xception after fine-tuning.
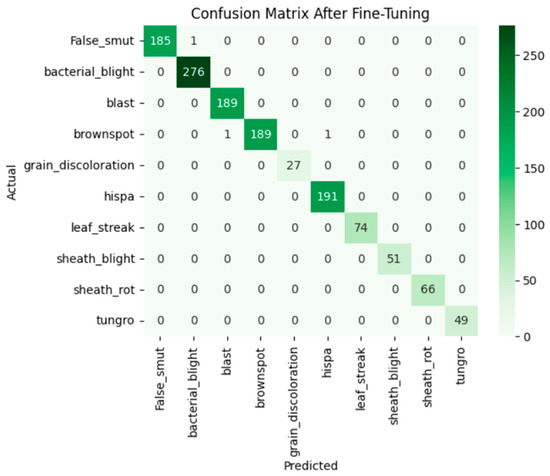
Figure 13.
Confusion matrix of the Xception after fine-tuning.
InceptionResNetV2:
Before fine-tuning:
Before fine-tuning, InceptionResNetV2 showed inconsistent performance. The accuracy graph and loss graph of the InceptionResNetV2 before fine-tuning are displayed in Figure 14. The training accuracy improved slowly and validation accuracy fluctuated around 70–80% which indicates poor generalization. The deeper frozen layers did not contribute effectively to feature learning that leads to higher validation loss and occasional class misclassifications. The confusion matrix is shown in Figure 15.
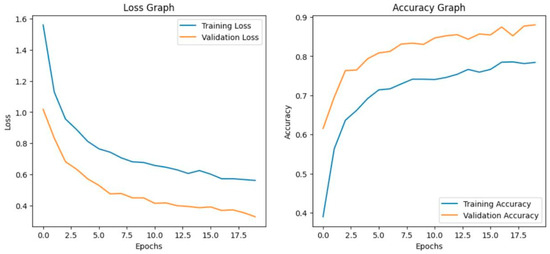
Figure 14.
Loss graph and accuracy graph of the InceptionResNetV2 before tuning.
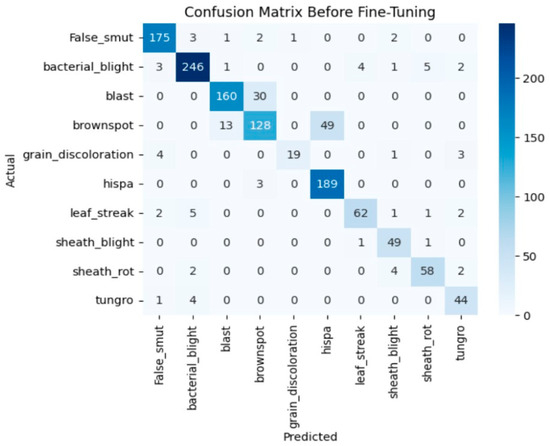
Figure 15.
Confusion matrix of the InceptionResNetV2 before fine-tuning.
After fine-tuning:
After fine-tuning, the model achieves 0.99 accuracy. The accuracy graph and loss graph of the InceptionResNetV2 after fine-tuning are displayed in Figure 16. Both training and validation accuracy curves converged. Loss steadily decreased to low values which shows stable and efficient learning. Precision, recall, and F1-scores became uniformly high across all classes. The confusion matrix exhibits minimal misclassification. The confusion matrix is shown in Figure 17. Thus, this confirms that the model successfully adapted to the dataset.
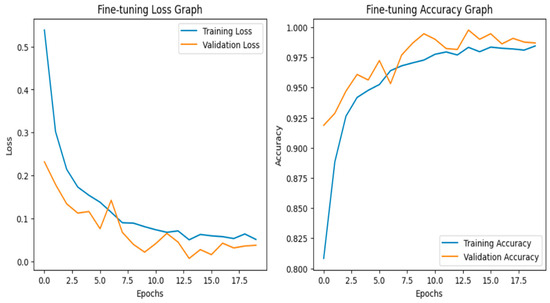
Figure 16.
Loss graph and accuracy graph of the InceptionResNetV2 after fine-tuning.
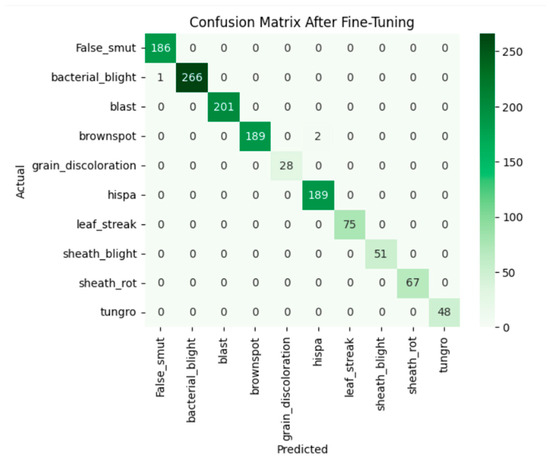
Figure 17.
Confusion matrix of the InceptionResNetV2 after fine-tuning.
MobileNetV2:
Before fine-tuning
Before fine-tuning, MobileNetV2 showed limited performance. The training and validation accuracy indicate underfitting. The accuracy graph and loss graph of the MobileNetV2 before fine-tuning are displayed in Figure 18. The model struggled to generalize due to its lightweight architecture and fewer parameters. Loss graphs showed a slow decline in training loss while validation loss fluctuated. This reflects difficulty in capturing complex disease patterns. Precision, recall, and F1-scores varied across classes. The confusion matrix exhibits frequent misclassifications. The confusion matrix is shown in Figure 19.
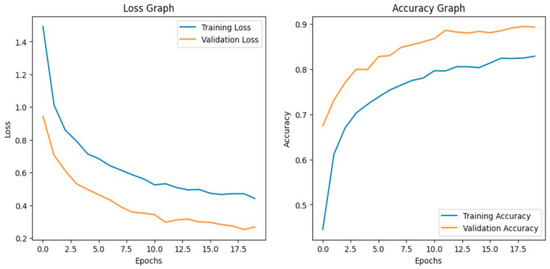
Figure 18.
Loss graph and accuracy graph of the MobileNetV2 before fine-tuning.
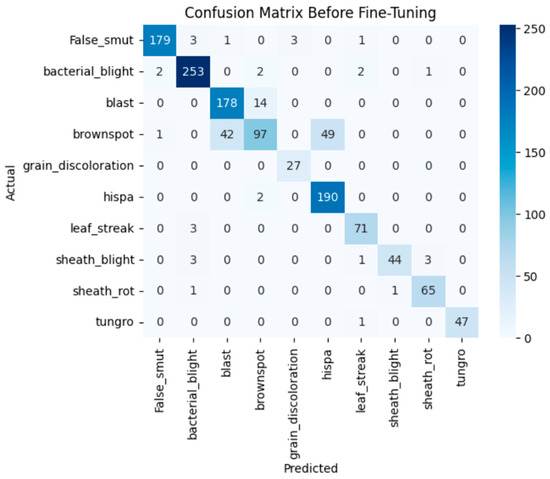
Figure 19.
Confusion matrix of the MobileNetV2 before fine-tuning.
After fine-tuning:
After fine-tuning, both training and validation accuracy increased steadily and the losses decreased consistently. The accuracy graph and loss graph of the MobileNetV2 after fine-tuning are displayed in Figure 20. Precision, recall, and F1-scores also improved across all classes. The confusion matrix became more diagonal, indicating accurate and stable classification performance. The confusion matrix after fine-tuning is shown in Figure 21.
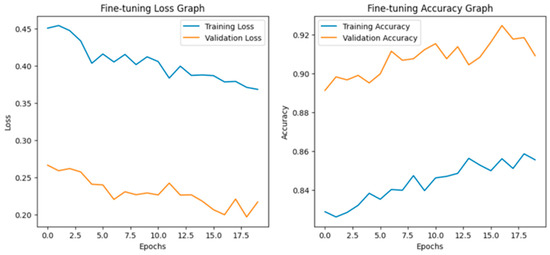
Figure 20.
Loss graph and accuracy graph of the MobileNetV2 after fine-tuning.
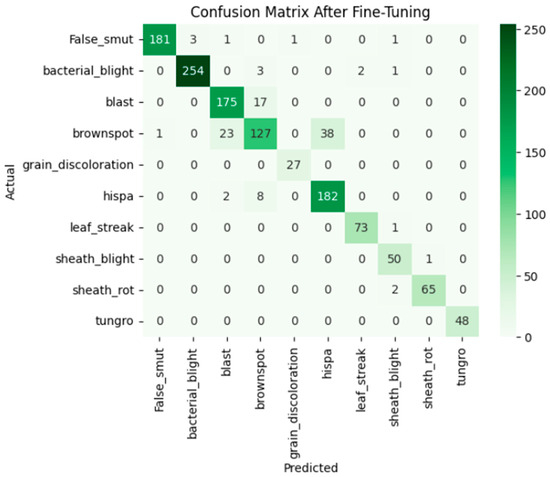
Figure 21.
Confusion matrix of the MobileNetV2 after fine-tuning.
The evaluation of six deep learning models on a plant disease dataset displays a clear development in performance after fine-tuning. The overall performance of the six deep learning models is displayed in Table 3. DenseNet201 was already performing well with 0.93 accuracy and fine-tuning gave it a small increase to 0.95. VGG19 and Efficient NetV2B3 showed moderate improvement, moving from about 0.85 to 0.90 and 0.98, respectively. The biggest improvements were seen in Xception and InceptionResNetV2. Their accuracy went from 0.80 and 0.75 to 0.99. This shows that these models responded very well to fine-tuning methods like adjusting the learning rate, unfreezing layers, or using data augmentation. MobileNetV2 improved a lot, going from about 0.70 to 0.96. This shows that even smaller models can achieve high accuracy with proper fine-tuning. In general, these findings show that fine-tuning is very important to get the best performance from deep learning models.

Table 3.
Overall performance of deep learning models.
8. Conclusions
In this research work, six deep learning models were used, including DenseNet201, VGG19, Xception, InceptionResNetV2, MobileNetV2, and EfficientNetV2B3, were tested with high-quality datasets. This work shows the effective and reliable solution of deep learning models in plant disease detection compared to traditional methods of manual inspection. Each model’s strengths and limitations are briefed in the paper. Among them, EfficientNetV2B3 and DenseNet201 yielded the best performance due to their strong feature extraction ability and balanced computational efficiency. The Xception and InceptionResNetV2 reached the highest value of accuracy with a balanced performance and VGG19 achieved moderate performance but was slower and less efficient than newer architectures, making it less suitable for large-scale deployment. MobileNetV2 demonstrated its high suitability for practical usage by taking into consideration its lightweight architecture. From the improved validation accuracy across the models, it became clear that fine-tuning played a significant role in improving the accuracy and stability of these models. Also, data augmentation and transfer learning significantly tackled data imbalance and improved the generalization of the model, indicating that dataset diversity especially in classes with fewer or visually similar images. This ensures that the models learnt robust features despite limited or uneven data. Thus, the work confirms that deep-learning-based approaches can enable fast and scalable disease detection. This reduces crop losses and strengthens food security, thus allowing farmers and researchers to act efficiently. Further advancements should focus on building lightweight models optimized for devices and smartphones that make the detection of diseases more accessible to farmers. Also, the integration of multi-modal data, such as thermal imaging and environmental parameters, in smart farming systems allows the improvement of early disease prediction for sustainable agriculture.
During the preparation of this work, the authors used Grammarly and Quillbot to correct grammatical errors.
Author Contributions
Conceptualization, K.S.S. and J.N.; Methodology, B.P.S., K.S.S. and J.N.; Validation, K.S.S.; Formal analysis, K.S.S.; Resources, J.N.; Data curation, K.S.S.; Writing—original draft, V.K.S. and R.K.J.; Writing—review & editing, B.P.S., K.S.S. and R.K.J.; Visualization, K.S.S.; Supervision, B.P.S. and K.S.S.; Project administration, B.P.S. and K.S.S. All authors have read and agreed to the published version of the manuscript.
Funding
This research received no external funding.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
The data presented in this study are available on request from the corresponding author due to future research work.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Prakash, M.; Neelakandan, S.; Tamilselvi, M.; Velmurugan, S.; Priya, S.B.; Martinson, E.O. Deep Learning-Based Wildfire Image Detection and Classification Systems for Controlling Biomass. Int. J. Intell. Syst. 2023, 2023, 7939516. [Google Scholar] [CrossRef]
- Indukuri, G.K.; Yarasuri, V.K.; Nair, A.K. Paddy Disease Classifier using Deep Learning Techniques. In Proceedings of the 2021 5th International Conference on Trends in Electronics and Informatics (ICOEI), Tirunelveli, India, 3–5 June 2021; IEEE: Washington, DC, USA, 2021. [Google Scholar]
- Narmadha, R.P.; Sengottaiyan, N.; Kavitha, R.J. Deep transfer learning based rice plant disease detection model. Intell. Autom. Soft Comput. 2022, 31, 1257–1271. [Google Scholar] [CrossRef]
- Joseph, D.S.; Pawar, P.M.; Chakradeo, K. Real-time plant disease dataset development and detection of plant disease using deep learning. IEEE Access 2024, 12, 1631016333. [Google Scholar] [CrossRef]
- Demilie, W.B. Plant disease detection and classification techniques: A comparative study of the performances. J. Big Data 2024, 11, 5. [Google Scholar] [CrossRef]
- Bhaskar, N.; Tupe-Waghmare, P.; Shetty, P.; Shetty, S.S.; Rai, T. A Deep Learning Hybrid Approach for Automated Leaf Disease Identification in Paddy Crops. In Proceedings of the 2024 International Conference on Distributed Computing and Optimization Techniques (ICDCOT), Bengaluru, India, 15–16 March 2024; IEEE: Washington, DC, USA, 2024; pp. 1–5. [Google Scholar]
- Sangar, G.; Rajasekar, V. Crop Disease Recognition and Classification: A Deep Dive into Machine Learning Techniques-A Survey. In Proceedings of the International Research Conference on Computing Technologies for Sustainable Development, Chennai, India, 9–10 May 2024; Springer Nature: Cham, Switzerland, 2024; pp. 38–57. [Google Scholar]
- Liu, N.; Zhao, Q.; Williams, R.; Duan, S.B.; Sun, Y.; Barrett, B. Ensemble modelling based on transfer learning for enhancing crop mapping through synergistic integration of InSAR coherence and multispectral satellite data. Comput. Electron. Agric. 2026, 242, 111332. [Google Scholar] [CrossRef]
- Sachdeva, G.; Singh, P.; Kaur, P. Plant leaf disease classification using deep Convolutional neural network with Bayesian learning. Mater. Today Proc. 2021, 45, 5584–5590. [Google Scholar] [CrossRef]
- Dubey, R.K.; Choubey, D.K. An efficient adaptive feature selection with deep learning model-based paddy plant leaf disease classification. Multimed. Tools Appl. 2024, 83, 22639–22661. [Google Scholar] [CrossRef]
- Simhadri, C.G.; Kondaveeti, H.K.; Vatsavayi, V.K.; Mitra, A.; Ananthachari, P. Deep learning for rice leaf disease detection: A systematic literature review on emerging trends, methodologies and techniques. Inf. Process. Agric. 2024, 12, 151–168. [Google Scholar] [CrossRef]
- Padhi, J.; Mishra, K.; Ratha, A.K.; Behera, S.K.; Sethy, P.K.; Nanthaamornphong, A. Enhancing Paddy Leaf Disease Diagnosis-a Hybrid CNN Model using Simulated Thermal Imaging. Smart Agric. Technol. 2025, 10, 100814. [Google Scholar] [CrossRef]
- Kulkarni, P.; Shastri, S. Rice leaf diseases detection using machine learning. J. Sci. Res. Technol. 2024, 2, 17–22. [Google Scholar] [CrossRef]
- Bollimuntha, K.S.; Priya, S.B. Enhancing agricultural supply chains with Industry 6.0 technologies for sustainable efficiency: Blockchain, AI and IoT for smart agriculture. In Industry 6.0 for Sustainable Supply Chains in Agriculture, Healthcare, and Asset Management; IGI Global Scientific Publishing: Hershey, PA, USA, 2026. [Google Scholar]
- Gupta, S.; Tripathi, A.K. Fruit and vegetable disease detection and classification: Recent trends, challenges, and future opportunities. Eng. Appl. Artif. Intell. 2024, 133, 108260. [Google Scholar] [CrossRef]
- Singh, A.; Kaur, J.; Singh, K.; Singh, M.L. Deep transfer learning-based automated detection of blast disease in paddy crop. Signal Image Video Process. 2024, 18, 569–577. [Google Scholar] [CrossRef]
- Shrivastava, V.K.; Pradhan, M.K.; Minz, S.; Thakur, M.P. Rice plant disease classification using transfer learning of deep convolution neural network. Int. Arch. Photogramm. Remote Sens. Spat. Inf. Sci. 2019, 42, 631635. [Google Scholar] [CrossRef]
- Kishore, G.; Kumar, K.P.; Vamsikrishna, C.D.; Vignesh, E.D.; Reddy, P.R. Paddy Leaf Disease Detection using Deep Learning Methods. In Proceedings of the 2022 Third International Conference on Intelligent Computing Instrumentation and Control Technologies (ICICICT), Kannur, India, 11–12 August 2022; IEEE: Washington, DC, USA, 2022. [Google Scholar]
- Elakya, R.; Manoranjitham, T. Transfer learning by VGG-16 with convolutional neural network for paddy leaf disease classification. Int. J. Image Data Fusion 2024, 15, 461–484. [Google Scholar] [CrossRef]
- Shafik, W.; Tufail, A.; De Silva Liyanage, C.; Apong, R.A.A.H.M. Using transfer learning-based plant disease classification and detection for sustainable agriculture. BMC Plant Biol. 2024, 24, 136. [Google Scholar] [CrossRef]
- Saad, M.H.; Salman, A.E. A plant disease classification using one-shot learning technique with field images. Multimed. Tools Appl. 2024, 83, 58935–58960. [Google Scholar] [CrossRef]
- Rahman, K.N.; Banik, S.C.; Islam, R.; Al Fahim, A. A real time monitoring system for accurate plant leaves disease detection using deep learning. Crop Des. 2025, 4, 100092. [Google Scholar] [CrossRef]
- Sharma, R.; Singh, A.; Jhanjhi, N.Z.; Masud, M.; Jaha, E.S.; Verma, S. Plant Disease Diagnosis and Image Classification Using Deep Learning. Comput. Mater. Contin. 2022, 71, 2125–2140. [Google Scholar] [CrossRef]
- Kalaivani, S.; Tharini, C.; Viswa, T.S.; Sara, K.F.; Abinaya, S.T. ResNet-based classification for leaf disease detection. J. Inst. Eng. 2025, 106, 1–14. [Google Scholar] [CrossRef]
- Yao, J.; Tran, S.N.; Garg, S.; Sawyer, S. Deep learning for plant identification and disease classification from leaf images: Multi-prediction approaches. ACM Comput. Surv. 2024, 56, 1–37. [Google Scholar] [CrossRef]
- Borhani, Y.; Khoramdel, J.; Najafi, E. A deep learning based approach for automated plant disease classification using vision transformer. Sci. Rep. 2022, 12, 11554. [Google Scholar] [CrossRef]
- Ahmad, W.; Azhar, E.; Anwar, M.; Ahmed, S.; Noor, T. Machine learning approaches for detecting vine diseases: A comparative analysis. J. Inform. Web Eng. 2025, 4, 99–110. [Google Scholar] [CrossRef]
- Jansi, K.R.; Amutha, A.L.; Shankar, A.B.; Adesh, J.P.; Kant, K. Plant Disease Classification Using Deep Learning for Agricultural Applications. In Harnessing AI in Geospatial Technology for Environmental Monitoring and Management; IGI Global Scientific Publishing: Hershey, PA, USA, 2025; pp. 213–238. [Google Scholar]
- Appalanaidu, M.V.; Kumaravelan, G. Classification of Plant Disease using Machine Learning Algorithms. In Proceedings of the 2024 Sixth International Conference on Computational Intelligence and Communication Technologies (CCICT), Bhubaneswar, India, 19–21 December 2024; IEEE: Washington, DC, USA, 2024; pp. 1–7. [Google Scholar]
- Ahmed, K.; Shahidi, T.R.; Alam, S.M.I.; Momen, S. Rice leaf disease detection using machine learning techniques. In Proceedings of the 2019 International Conference on Sustainable Technologies for Industry 4.0 (STI), Dhaka, Bangladesh, 24–25 December 2019; IEEE: Washington, DC, USA, 2019; pp. 1–5. [Google Scholar]
- Kaur, A.; Guleria, K.; Trivedi, N.K. A deep learning-based model for biotic rice leaf disease detection. Multimed. Tools Appl. 2024, 83, 83583–83609. [Google Scholar] [CrossRef]
- Khalid, M.M.; Karan, O. Deep learning for plant disease detection. Int. J. Math. Stat. Comput. Sci. 2024, 2, 75–84. [Google Scholar]
- Bagga, M.; Goyal, S. Image-based detection and classification of plant diseases using deep learning: State-of-the-art review. Urban Agric. Reg. Food Syst. 2024, 9, e20053. [Google Scholar] [CrossRef]
- Mukherjee, R.; Ghosh, A.; Chakraborty, C.; De, J.N.; Mishra, D.P. Rice leaf disease identification and classification using machine learning techniques: A comprehensive review. Eng. Appl. Artif. Intell. 2025, 139, 109639. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.