Abstract
Accurate identification of Alzheimer’s disease (AD)-related cellular characteristics from microscopy images is essential for understanding neurodegenerative mechanisms at the cellular level. While most computational approaches focus on macroscopic neuroimaging modalities, cell type classification from microscopy remains relatively underexplored. In this study, we propose a hybrid vision transformer–convolutional neural network (ViT–CNN) framework that integrates DeiT-Small and EfficientNet-B7 to classify three AD-related cell types—astrocytes, cortical neurons, and SH-SY5Y neuroblastoma cells—from phase-contrast microscopy images. We perform a comparative evaluation against conventional CNN architectures (DenseNet, ResNet, InceptionNet, and MobileNet) and prompt-based multimodal vision–language models (GPT-5, GPT-4o, and Gemini 2.5-Flash) using zero-shot, few-shot, and chain-of-thought prompting. Experiments conducted with stratified fivefold cross-validation show that the proposed hybrid model achieves a test accuracy of 61.03% and a macro F1 score of 61.85, outperforming standalone CNN baselines and prompt-only LLM approaches under data-limited conditions. These results suggest that combining convolutional inductive biases with transformer-based global context modeling can improve generalization for cellular microscopy classification. While constrained by dataset size and scope, this work serves as a proof of concept and highlights promising directions for future research in domain-specific pretraining, multimodal data integration, and explainable AI for AD-related cellular analysis.
1. Introduction
Alzheimer’s disease (AD) is a neurodegenerative disorder caused by the accumulation of misfolded amyloid-beta protein, which severely impacts the brain and central nervous system, leading to the progressive loss of memory and the ability to function independently. Despite numerous hypotheses regarding the origins and causes of AD, current medical treatments are unable to cure most of these conditions [1,2]. Over the past three decades, the pathology of Alzheimer’s disease (AD) has been extensively studied through the key pathological features of AD, including the accumulation of proteins, the loss of neurons and synapses, and changes associated with reactive processes [3].
The prevalence of AD continues to grow, with approximately 6 million individuals affected in the United States alone. This number is expected to rise to 12.7 million by 2050, driven by an aging population and the absence of major breakthroughs in prevention or treatment [4]. The ability to delineate specific brain cell types in AD pathology is critical for understanding the cellular mechanisms that contribute to disease progression and developing targeted therapeutic strategies [5]. However, studying the interactions and adaptability of different brain cell types in living organisms remains a significant challenge, particularly when investigating large-scale structural brain changes linked to physiological and pathological states [6,7].
Artificial intelligence (AI) has emerged as a transformative tool in biomedical imaging, providing advanced analytical frameworks to support early diagnosis and mechanistic understanding of diseases. While AI-based models have achieved remarkable success in detecting AD stages from magnetic resonance imaging (MRI) data—such as the convolutional neural network (CNN)-based approach by Alshammari et al., which achieved 97% accuracy across four AD stages [8]—the classification of AD-associated cell types remains an underexplored but critical frontier. Accurate identification of cellular phenotypes can reveal insights into disease progression at the microscopic level and support the discovery of novel drug targets.
The classification of key cell types involved in AD, such as astrocytes, cortical neurons, and SH-SY5Y neuroblastoma cells, using computer vision and AI techniques offers great potential for advancing therapeutic research [9,10]. Several state-of-the-art deep learning architectures have shown promise in AD-related image analysis. For instance, DenseNet [11], which employs dense connectivity to promote feature reuse, achieved an 86% classification accuracy in AD MRI categorization [12]. Similarly, ResNet’s residual learning framework effectively mitigates vanishing gradient problems, with pretrained ResNet-50 models achieving up to 98.99% accuracy in AD MRI classification [13,14]. MobileNet and InceptionNet architectures have also been leveraged for efficient AD diagnosis. MobileNet’s depthwise separable convolutions optimize computational costs while maintaining accuracy, yielding 94% accuracy in AD classification [15,16], whereas InceptionNet’s multi-scale design enhances feature extraction and performance with minimal resource overhead [17,18].
Furthermore, EfficientNet models overcome conventional CNN limitations by uniformly scaling the network depth, width, and resolution, thereby achieving superior performance with reduced manual tuning. Sethi et al. [19] demonstrated that by employing a composite scaling coefficient to jointly adjust these parameters, the EfficientNet transfer learning (TL) model effectively mitigates architectural imbalance and enhances representational efficiency. The model is characterized by two guiding principles: first, an optimized baseline architecture that accelerates convergence during training, and second, a compound scaling strategy that maintains accuracy while deepening the network. Its design relies on 2D depthwise separable convolutions composed of multiple inverted mobile bottleneck blocks, which contribute to computational efficiency and improved feature extraction. One of EfficientNet’s major advantages over conventional CNNs lies in this systematic scaling approach, which employs predefined scaling factors to balance the model capacity and computational costs. Consequently, the EfficientNet TL strategy has been successfully applied for Alzheimer’s disease (AD) diagnosis, achieving 91.36% accuracy and an AUC of 83% on the ADNI dataset [19]. Similarly, Zheng et al. [20] proposed a modified 3D EfficientNet architecture that addresses the limitations of conventional scaling methods by introducing a max pooling layer after the initial convolution, ReLU activation outside the main building block, and replacing the global average pooling layer with three fully connected layers. These enhancements led to substantial performance gains, achieving 83.33% classification accuracy and 88.75% AUC, an improvement of more than 10% over the original EfficientNet model.
Despite these advancements, EfficientNet-based architectures remain limited by their reliance on large-scale pretraining datasets such as ImageNet, which may introduce domain bias when applied to biomedical images. Future research should focus on domain-specific pretraining and adaptive scaling strategies tailored to medical and cellular imaging data.
Astrocytes, cortical neurons, and SH-SY5Y cells each play critical roles in AD pathology. Astrocytes regulate key neuropathological processes, cortical neurons experience significant atrophy, and SH-SY5Y cells undergo structural and molecular alterations linked to AD progression [21,22,23]. Leveraging AI-driven deep learning frameworks to classify these cell types from microscopic images can provide valuable insights into disease mechanisms and inform drug development strategies [5,24,25].
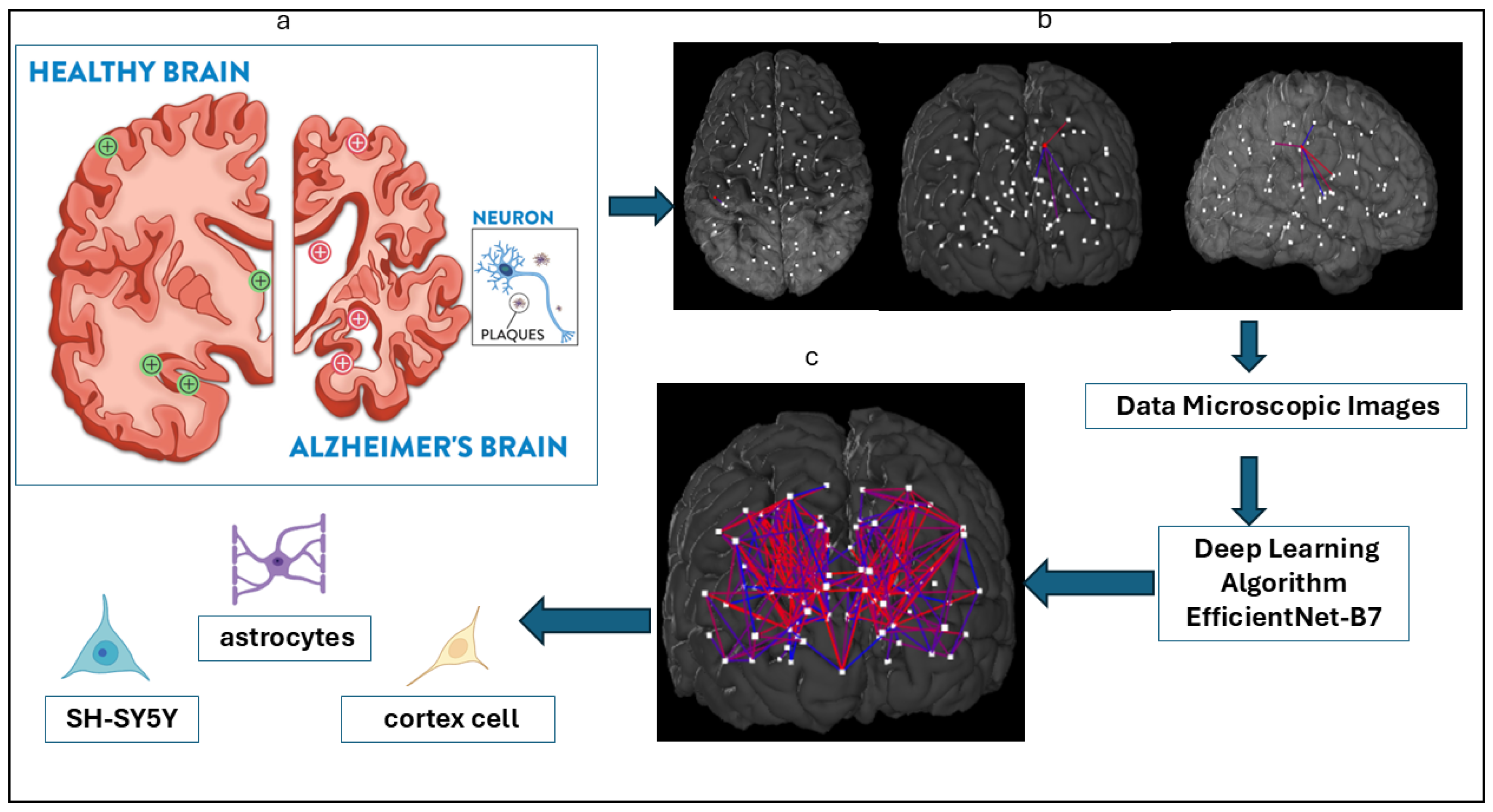
While accurate cell classification is an essential step in addressing fundamental biological questions in AD research, challenges such as low image contrast, high cell density, and limited annotated data often hinder conventional analysis [7,26]. To address this gap, the present study focuses on three key AD-related cell types—astrocytes, cortical cells, and SH-SY5Y cells—using a hybrid transformer–CNN architecture built upon EfficientNet-B7. This model integrates convolutional and attention-based mechanisms to enhance feature extraction and improve generalization in multi-class cellular image classification. Figure 1 illustrates the overall schematic of our approach.
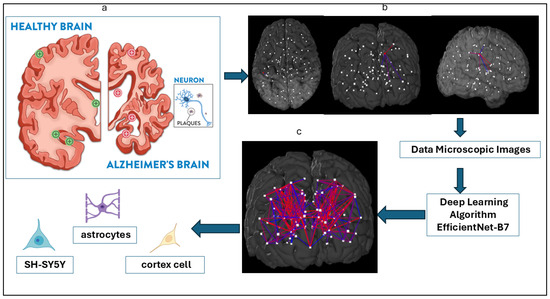
Figure 1.
Study schematics of Alzheimer’s disease cell type detection. (a) Sketch of the brain and Alzheimer’s disease by the Fisher Center for Alzheimer’s Research Foundation. (b) Brain parcellation in sagittal, coronal, and axial views drawn from Budapest Reference Connectome version 3.0. (c) Classification of three key cell types—astrocytes, cortex cells, and SH-SY5Y cells—associated with AD using an AI-driven EfficientNet-B7 architecture applied to cell type-specific microscopic images.
This study presents an advanced CNN-based hybrid architecture for classifying AD-associated cell types using transfer learning and attention-based enhancement. By improving the precision of cell type identification, our approach contributes to a deeper understanding of AD pathology and provides a foundation for developing targeted therapeutic interventions.
2. Related Work
2.1. CNN-Based Approaches
In this section, we review several CNN-based methods that have been used for image classification in the past and how they inform the proposed approach.
One highly relevant CNN-based model is DenseNet [11]. DenseNet has a distinct connectivity scheme because it adds direct links between any layer and successive layers. This allows the feature maps of all previous layers to be sent to the lth layer. This scheme facilitates the extraction and sharing of feature maps throughout the network, which improves accuracy and speeds up inference times. Consequently, it is defined as follows: , where denotes the combination of feature maps created in layers . The developers called this network architecture a “Dense Convolutional Network”, or DenseNet, because of its dense interconnectedness. The several inputs of in equation can be combined into a single tensor for simplicity of implementation [11].
Another relevant CNN-based model is ResNet [13], which considers as an underlying mapping that can be fitted by a few stacked layers, though not necessarily the entire network, with x representing the inputs to these layers, as in DenseNet. It is analogous to hypothesizing that numerous nonlinear layers can asymptotically approximate the residual functions; that is, (assuming that the input and output are of the same dimensions). Therefore, the layers explicitly approximate the residual function instead of assuming the stacked layers to approach . In this framework, the initial function is rather than . Both representations can provide an asymptotic approximation of the required functions; however, their respective learning curves may differ. From this perspective, it is possible to apply residual learning to every stacked level. Formally, they defined a building block as follows: [13]. In this case, the vectors x and y serve both as input and output for the layers being considered. The function serves as a representation of the residual mapping that must be learned. Then, using a skip connection and element-wise addition, the operation is carried out. Note that in the equation , the dimensions of x and F must be equivalent. In the case that they are not, a linear projection using the skip connections to fit the dimensions () [13] is a reasonable solution.
The MobileNet model is built upon depthwise separable convolutions, which are a kind of factorized convolution. This approach decomposes a standard convolution into two separate convolutions: a depth-wise and a point-wise convolution. In MobileNets, each input channel is assigned a single filter as a result of the use of depthwise convolution. The outputs of the depthwise convolution are combined with an additional 11 convolutions after the point-wise convolution. The inputs are filtered and combined using a typical convolution algorithm, which results in a new set of outputs. The factorization leads to a significant reduction in both the computational cost and model size. To apply a single filter to each input channel, the authors applied depth-wise convolutions (input depth), and the depth-wise layer’s output was then combined linearly using point-wise convolution. In both layers of MobileNets, batch normalization and ReLU nonlinearities are used [15].
Tan et al. [27] proposed EfficientNet, which is a new compound scaling method using a compound coefficient. The compound scaling of the network’s breadth, depth, and resolution are as follows:
Recent studies have demonstrated the effectiveness of ML techniques in diagnosing Alzheimer’s disease (AD) using neuroimaging data such as magnetic resonance imaging (MRI) and positron emission tomography (PET). Fathi et al.[28] reviewed 74 studies on eight different databases, resulting in 736 studies on deep learning for early Alzheimer’s disease diagnosis. They found that CNN-based models outperform other deep learning models but lack explainability due to their black box nature. Moreover, most of the models focus on discriminative features for AD or MCI classification, improving performance but limiting interpretability. Additionally, they found there were 13 studies reporting this classification group for eight different databases, with the accuracy being in the range of 64.04–99.7% and the mean value being 87.43% [28].
Jain et al. [29] achieved 95.3% accuracy using a 2D-CNN on sMRI images from the ADNI dataset. Ding et al. [30] employed an Inception v3 CNN on PET scans, achieving 82% specificity and 100% sensitivity. Chitradevi et al. [31] utilized an AlexNet CNN on MRI images, reporting 95% accuracy, 94% specificity, and 95% sensitivity. Nawaz et al. [32] achieved the highest accuracy of 99.21% using SVMs on MRI data from the OASIS dataset. Kundaram et al. [33] reported 98.57% accuracy using a CNN on MRI images from ADNI. These studies highlight the potential of various deep learning architectures and traditional machine learning methods in AD diagnosis across different neuroimaging modalities and datasets. However, these studies were unable to validate their analyses with cell type-specific data, despite the fact that Alzheimer’s disease (AD) is characterized by neuronal death and tissue loss in the brain.
Several studies have addressed the challenge of data limitation in Alzheimer’s disease (AD) research using various approaches. Vu et al. [34] combined MRI and PET modalities from the ADNI dataset to increase data volume, albeit at the cost of increased computational complexity. Jain et al. [29] employed transfer learning to mitigate the scarcity of sMRI data, reducing computational costs. Salehi et al. [35] merged two ADNI MRI datasets to expand their sample size, though this approach may introduce inconsistencies in image formats and sizes. Basaia et al. [36] combined ADNI and Milan datasets during the testing phase, achieving higher accuracy in various classification tasks, except for AD vs. HC classification. Additionally, Khvostikov et al. utilized both structural MRI (sMRI) and diffusion tensor imaging (DTI) modalities from the ADNI dataset for Alzheimer’s disease classification, demonstrating that while the addition of DTI data can provide complementary information, it may also introduce noise in smaller datasets, highlighting the trade-offs in multi-modal approaches for neuroimaging analysis [37]. However, this method faced challenges in feature interpretability. These strategies demonstrate the trade-offs between data augmentation, computational complexity, and model performance in overcoming data limitations in AD research.
This review of the major CNN-based models—DenseNet, ResNet, MobileNet, and EfficientNet—highlights their unique architectural innovations and their application to AD diagnosis. Recent SOTA approaches have used multimodal data, hybrid architectures, and explainable AI to improve diagnostic performance. However, most studies lack validation on cell-type-specific data, despite AD’s cellular pathology involving astrocytes, cortical neurons, and SH-SY5Y cells. This limitation underscores the need for models tailored to cellular-level analysis. Our study addresses this gap by extending these architectures to classify AD-related cell types, bridging the divide between computational and cellular neuroscience.
2.2. Prompt-Based LLM
Recent research has extended instruction tuning and reasoning prompting from pure language models to vision–language and multimodal systems. For example, InstructBLIP [38] reformulates 26 public vision–language tasks into an instruction format and introduces an instruction-aware query transformer, achieving strong zero-shot performance across held-out datasets. Similarly, LLaVA uses machine-generated image-instruction pairs to train a vision encoder plus LLM that shows effective image-language chat and reasoning capabilities [39]. In the biomedical domain, MedCLIP [40] applies contrastive learning on unpaired medical image and text data and demonstrates zero-shot prediction gains on medical imaging tasks. These works show that while multimodal LLMs offer strong global reasoning and conversational abilities, they often require domain-specific adaptation (especially for fine-grained or domain-specialized tasks) and may still lag supervised models in highly specialized visual discrimination in Table 1. In this study, we therefore include prompt-based evaluations (zero shot, few shot, and chain of thought) of multimodal LLMs alongside our proposed hybrid ViT–CNN architecture to quantify this trade-off in the Alzheimer’s disease cell type classification setting.

Table 1.
Recent state-of-the-art (SOTA) studies in Alzheimer’s disease classification.
3. Methodology
This study investigates Alzheimer’s cell classification from microscopy imagery through three complementary methodological paradigms: (1) deep learning-based visual modeling, (2) multi-modal large language model (LLM) reasoning via prompt-based evaluation, and (3) a proposed hybrid transformer–CNN model that integrates global semantic and fine-grained morphological information. In the following, a detailed description of the data is provided, followed by a summary of the baseline and proposed models that were evaluated on that data.
3.1. Data Preparation
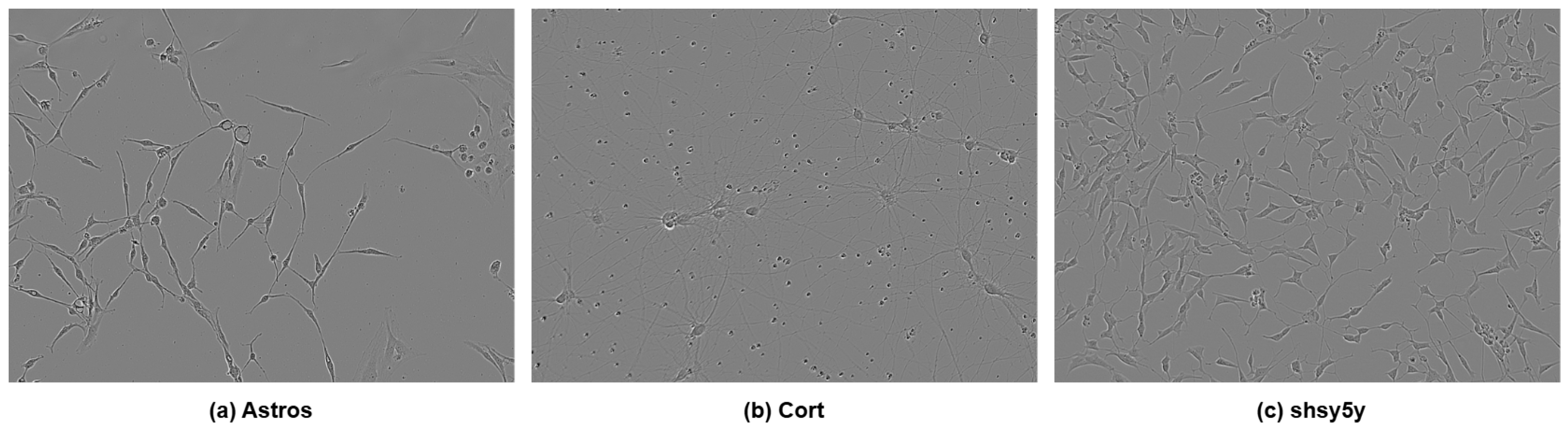
The dataset consisted of 606 label-free phase-contrast microscopy images, including 320 cortical images, 150 SH-SY5Y images, and 130 astrocyte images. These were obtained from the Incucyte Live-Cell Analysis System [7]. The dataset exhibited substantial class imbalance, which poses challenges for supervised learning under limited sample regimes. In Figure 2, three types of cells are shown.

Figure 2.
Three types of cells: (a) astrocyte, (b) cortical, and (c) SH-SY5Y.
To address this imbalance, we employed the synthetic minority oversampling technique (SMOTE) [47] under a strictly controlled experimental protocol. Importantly, the SMOTE was applied exclusively to the training data within each cross-validation fold and never to the validation or test sets, thereby eliminating any possibility of data leakage during evaluation. Furthermore, the SMOTE was applied in feature space rather than pixel space. Specifically, deep feature embeddings were first extracted from pretrained backbone networks, and the SMOTE was used to interpolate between minority-class feature vectors. The resulting synthetic feature representations were then used only during model training. This approach avoids generating unrealistic pixel-level images while enabling controlled rebalancing of class distributions in representation space. After applying the SMOTE within the training folds, the effective training set size increased to 960 samples, while the validation and test sets remained composed solely of original, unmodified images. All reported test results therefore reflect performance on real microscopy data only.
Stratified fivefold cross-validation was employed to ensure consistent class proportions across splits, and identical preprocessing pipelines were applied across all folds. The training set facilitated the acquisition of knowledge by the model, the validation set aided in fine-tuning the model, and the test set offered an impartial assessment of the model’s performance.
Data Augmentation
Robust data augmentation was applied to enhance generalization and reduce overfitting. The augmentations include RandomResizedCrop, ColorJitter, GaussianBlur, and MixUp/CutMix regularization. These transformations simulate realistic variations in microscopy conditions while maintaining biological integrity.
3.2. Deep Learning Baseline Models
To establish strong and reliable baselines, we employed several widely adopted convolutional neural network (CNN) architectures pretrained on ImageNet, including ResNet-50, VGG16, DenseNet121, and EfficientNet-B7. These models were selected due to their proven effectiveness in medical image analysis and transfer learning scenarios.
All baseline networks followed a unified transfer learning framework. For each architecture, the original ImageNet classification head was removed, and the network was initialized with pretrained weights. The input images were processed using the model-specific preprocessing function to ensure consistency with the feature distribution learned during ImageNet pretraining. The pretrained backbone served as a feature extractor, generating high-level convolutional representations of the microscopy images. During fine-tuning, only the last portion of the backbone layers was updated, while earlier layers remained frozen. To maintain stable feature statistics under domain adaptation, all batch normalization layers within the backbone were kept frozen.
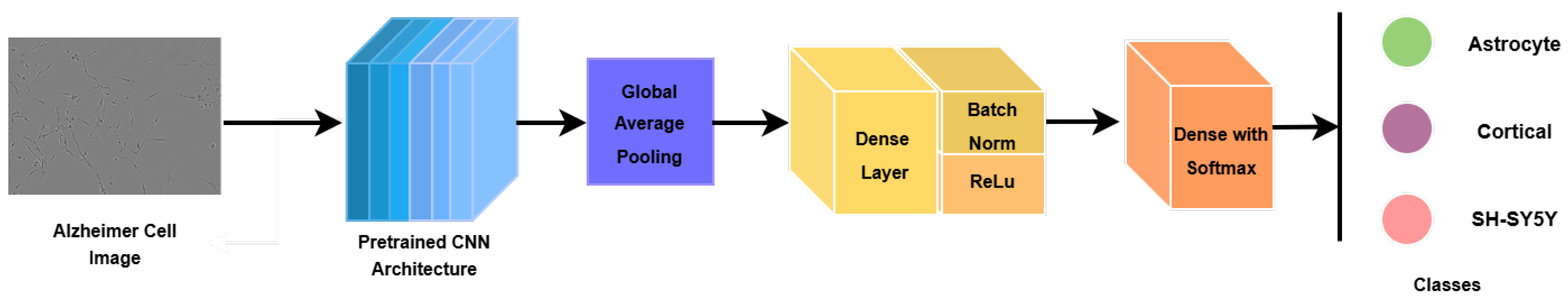
To adapt the pretrained backbones to the Alzheimer’s microscopy classification task, a lightweight task-specific classification head was appended. The extracted feature maps were first aggregated using global average pooling (GAP) to obtain a compact feature representation. This representation was passed through a fully connected layer, followed by batch normalization, ReLU activation, and dropout for regularization. A final fully connected layer with three output neurons and Softmax activation produced class probabilities for the three diagnostic categories.
Figure 3 illustrates this transfer learning architecture.
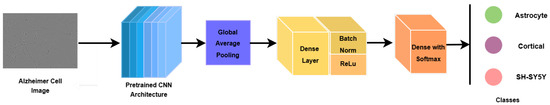
Figure 3.
Deep learning model architectural pipeline.
3.2.1. Optimization and Training Set-Up for Deep Learning Models
- Optimizer: AdamW, known for stable convergence and adaptive weight decay.
- Loss Function: CrossEntropyLoss with class weighting to counter data imbalance.
- Batch Size: 32.
- Epochs: 50–100 (based on convergence criteria).
3.2.2. Evaluation Objective for Deep Learning Models
These baseline CNN models act as a reference framework for evaluating both prompt-based and hybrid transformer–CNN models. The performance of each model was measured in terms of accuracy, macro F1 score, and class-wise confusion matrices.
3.3. Multi-Modal LLM Evaluation via Prompting
To examine the reasoning and interpretative capabilities of large-scale multimodal vision–language models on cellular microscopy images, we evaluated Gemini 2.5 Flash and GPT-5, both of which support joint visual and textual inputs. These models were assessed without gradient-based fine-tuning, relying exclusively on prompt engineering to perform the classification task. Consequently, the LLM results are intended to serve as prompt-only baselines rather than fully optimized vision classifiers.
All multimodal LLM experiments were conducted for cell type classification and not disease stage prediction. Each model was instructed to categorize microscopy images into one of three cell types: astrocyte, cortical neuron, or SH-SY5Y neuroblastoma cells. This formulation ensured consistency with the supervised deep learning and hybrid models evaluated in this study.
3.3.1. LLM Prompting Strategies
We employed three complementary prompting paradigms to analyze how contextual guidance and explicit reasoning affect multimodal LLM performance. The exact prompt templates used in all experiments are reported below for reproducibility:
- Zero-Shot Prompting: The model is provided with a task instruction without labeled examples, evaluating its intrinsic visual reasoning capability.

- Few-Shot Prompting: The prompt includes a small number (3–5) of labeled examples to provide contextual guidance via in-context learning.
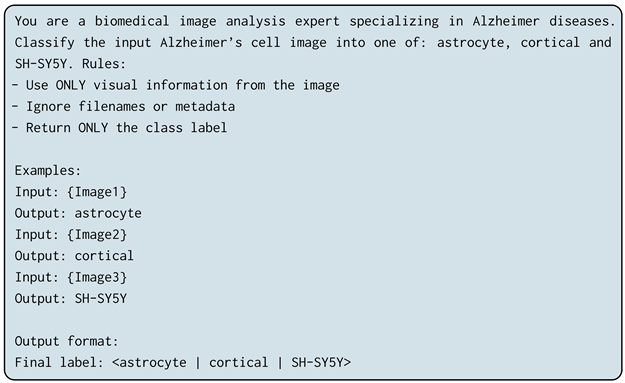
- Chain-of-Thought (CoT) Prompting: The model is explicitly instructed to reason through intermediate steps prior to classification, encouraging structured visual analysis similar to human expert reasoning.

3.3.2. Evaluation Procedure for LLM Models
Each multimodal LLM processes the same microscopy images under all three prompting configurations. Model outputs are parsed and normalized to one of the three predefined cell type labels. Quantitative performance was evaluated according to accuracy, precision, recall, and macro F1 score, enabling direct comparison with supervised CNN and hybrid CNN–transformer models.
While multimodal LLMs demonstrate promising interpretability through natural language explanations and reasoning traces, their quantitative performance remained limited in this setting. This limitation is attributed to the absence of domain-specific visual fine-tuning and the inherent difficulty of transferring general-purpose vision–language representations to specialized microscopy imagery. Accordingly, the LLM results are interpreted as exploratory benchmarks that contextualize the performance of learned visual representations under data-limited conditions.
3.4. Proposed Hybrid Transformer–CNN Model
To overcome the limitations of both standalone CNNs (local feature bias) and prompt-based LLM reasoning (lack of domain adaptation), we introduce a fine-tuned HybridDeiT_EfficientNet_B7 architecture. This model synergistically combines the global semantic awareness of a distilled transformer (DeiT-Small) with the high-resolution morphological precision of a deep CNN (EfficientNet-B7).
3.4.1. Rationale for Hybridization
The hybrid architecture explicitly leverages complementary inductive biases of CNNs and transformers. While the transformers capture global semantic relationships and long-range dependencies, which are critical for identifying distributed pathological cues, the CNNs capture the localized spatial textures and morphological granularity essential for fine cellular differentiation. This combination ensures that both global and local cues are utilized during training, leading to more robust classification in complex biomedical imagery.
3.4.2. Architectural Design of the Proposed Model
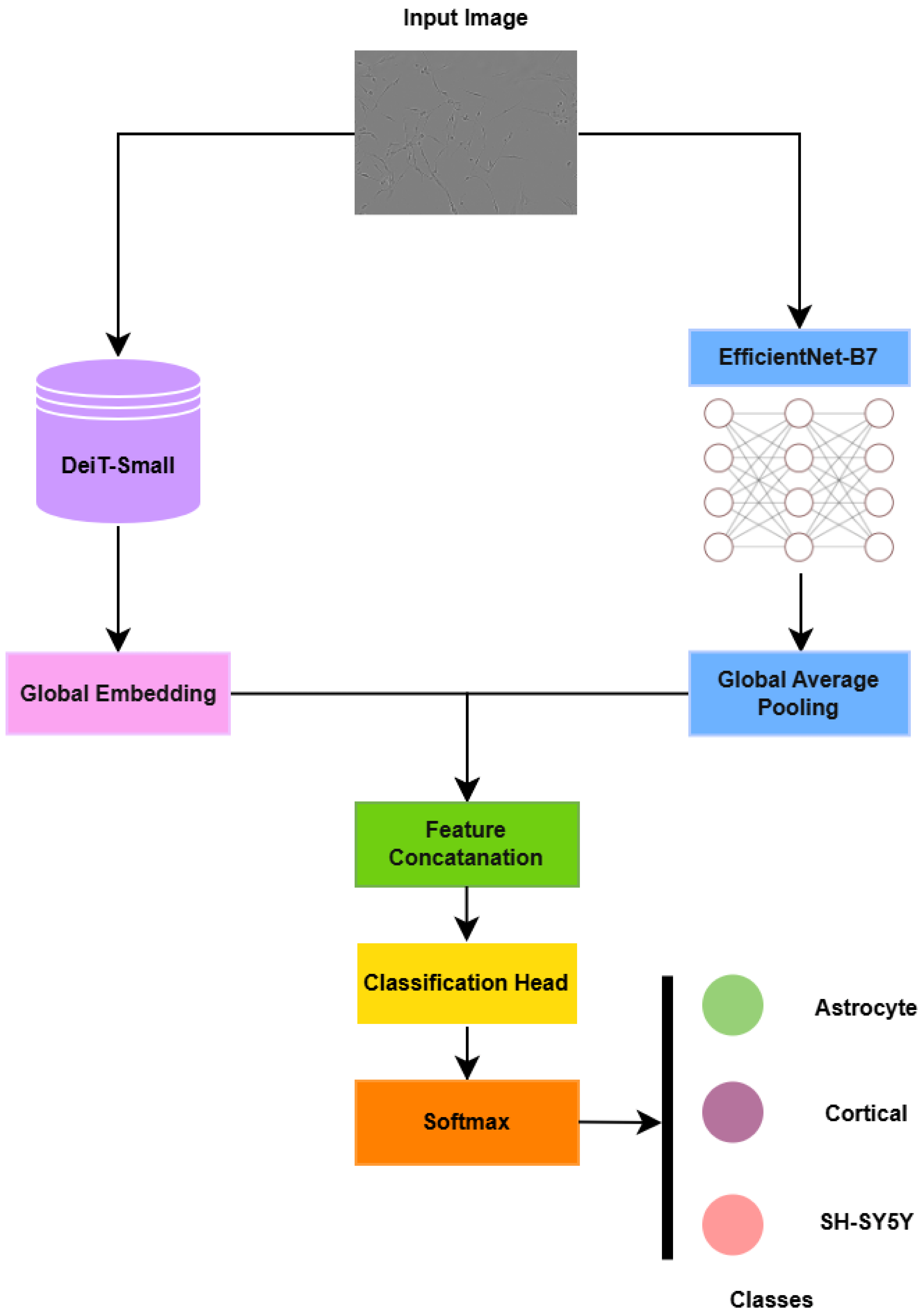
The proposed hybrid architecture integrates two pretrained visual backbones that operate in parallel to jointly capture global contextual semantics and fine-grained cellular morphology.
The first branch employs DeiT-Small, a distilled vision transformer, to encode global semantic relationships through self-attention. Given an input microscopy image , the transformer produces a compact global embedding derived from the class token of the final encoder layer:
In parallel, the second branch utilizes EfficientNet-B7 to extract hierarchical convolutional features that encode a localized texture and morphological structure. The convolutional feature maps
are aggregated using global average pooling (GAP) to obtain a compact visual descriptor:
Feature fusion is performed at the embedding level through concatenation of the transformer and CNN representations:
where denotes vector concatenation.
The fused representation is subsequently mapped into a shared latent space using a learnable linear projection implemented as a fully connected layer:
where and are trainable parameters that project the concatenated features into a 2048-dimensional embedding space.
The projected representation is then passed to a classification head consisting of a fully connected layer followed by Softmax activation, producing probabilities over the three Alzheimer’s-related cell types (astrocyte, cortical, and SH-SY5Y).
The overall architectural pipeline is illustrated in Figure 4.
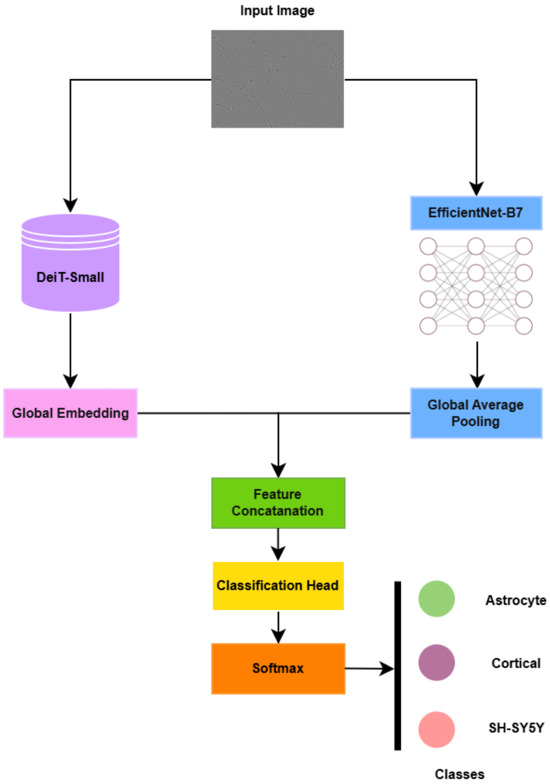
Figure 4.
Proposed model’s architectural pipeline.
3.4.3. Training Configuration for Proposed Model
The training configuration we used for the proposed model is as follows:
- Optimizer: AdamW.
- Scheduler: Cosine annealing.
- Loss Function: Weighted CrossEntropyLoss to address class imbalance.
- Batch Size: 16.
- Epochs: 80.
- Regularization: Stochastic depth and label smoothing.
- Data Augmentation: Augmentations identical to the baselines are used—Random ResizedCrop, ColorJitter, GaussianBlur, and MixUp/CutMix—to maintain consistent preprocessing while increasing sample diversity.
- Input Resolution: DeiT path (224 × 224) and EfficientNet path (448 × 448–600 × 600). These images are downsampled separately for each pipeline to preserve the optimal receptive field for both networks.
3.5. Comparitive Evaluation Protocol
All models, comprising the deep learning baselines, multi-modal LLMs, and the proposed hybrid architecture, were evaluated under a consistent experimental protocol to ensure fairness and comparability. The evaluation metrics included the overall accuracy, macro F1 score, and per-class precision–recall curves, providing a comprehensive assessment of both global and class-specific performance. The dataset was partitioned into 70% for training, 15% for validation, and 15% for testing, maintaining stratified distribution across classes. To ensure reproducibility and statistical reliability, each experiment was repeated with three distinct random seeds, and the final reported results represent the averaged performance across these runs.
The methodological framework systematically contrasts three distinct learning paradigms—supervised CNNs, reasoning-based multi-modal LLMs, and a fine-tuned hybrid transformer–CNN model. Empirically, the HybridDeiT_EfficientNet_B7 achieved superior classification performance, demonstrating that explicit multi-modal feature fusion remains more effective than prompt-based inference in specialized biomedical image understanding tasks.
4. Results
4.1. Experimental Set-Up and Evaluation Metrics
All experiments were conducted using stratified fivefold cross-validation to ensure robust evaluation across different data splits. Class balancing via the SMOTE was applied exclusively within the training portion of each fold, while the validation and test sets consisted solely of original, unmodified microscopy images. This protocol ensured that no synthetic samples appeared in the evaluation data and eliminated any possibility of data leakage.
Following SMOTE-based rebalancing of the training folds, the effective training set size increased to 960 samples, while evaluation metrics were computed only on real images. Model performance was assessed using the accuracy, precision, recall, and macro F1 score. Particular emphasis was placed on the F1 score due to its balanced consideration of false positives and false negatives, which is especially important in medical imaging tasks, where misclassification can lead to misleading biological interpretations [48]. The experiments were conducted on an NVIDIA T4 GPU using PyTorch 2.9.0, CUDA 13.1, and Python 3.9.
4.2. Comparative Analysis of Methodological Approaches
4.2.1. Prompt-Based Large Language Model Performance
Table 2 summarizes the performance of prompt-based multimodal large language models (LLMs) under zero-shot, few-shot, and chain-of-thought prompting strategies for Alzheimer’s-related cell type classification. These models were evaluated without task-specific fine-tuning and should therefore be interpreted as prompt-only baselines rather than optimized vision classifiers.

Table 2.
Performance comparison of prompt-based LLM methods.
Across all prompting strategies, performance exhibited a clear hierarchy consistent with the model scale and architectural sophistication. GPT-5 achieved the highest accuracy range (50.94–58.82%), outperforming GPT-4o (41.98–45.32%) and Gemini 2.5 Flash (30.37–35.07%). Chain-of-thought prompting consistently improved performance for Gemini and GPT-5, suggesting that structured reasoning benefits complex visual categorization tasks. However, few-shot prompting yielded mixed results for GPT-4o, indicating sensitivity to prompt formulation.
The precision–recall trade-offs varied across the models; GPT-4o zero shot achieved relatively high precision (53.43%) at the expense of recall (44.37%), whereas GPT-5 with chain-of-thought prompting maintained a more balanced precision–recall profile (56.58% and 57.86%, respectively). Overall, while the prompt-based LLMs demonstrated encouraging zero- and few-shot reasoning capabilities, their performance remained constrained in this domain due to the absence of domain-specific visual adaptation.
4.2.2. Fine-Tuned Deep Learning Model Performance
Table 3 presents the performance of fine-tuned deep learning models trained using ImageNet-pretrained weights. Across all CNN-only architectures, a consistent pattern emerged: validation accuracies remained moderate (approximately 64–66%), while test performance dropped sharply to near chance levels (accuracy of 32–35% and F1 score of about 31–35%). This discrepancy resulted in large generalization gaps exceeding 30% for most CNN architectures.

Table 3.
Performance comparison of fine-tuned models.
These results indicate that despite transfer learning, the standalone CNN models struggled to generalize effectively under extreme data scarcity and domain shifts from natural images to microscopy imagery. The consistency of this behavior across diverse architectures—including MobileNet variants, ResNet, DenseNet, InceptionNet, and EfficientNet-B7—suggests that the observed limitation is dataset-driven rather than architecture-specific.
In contrast, the transformer-based and hybrid models demonstrated substantially improved generalization. The DeiT-Small model alone achieved a test accuracy of 57.39% and F1 score of about 56.75% with a reduced generalization gap (15.72%), indicating that global self-attention mechanisms offer increased robustness under limited data conditions. The proposed hybrid DeiT-Small + EfficientNet-B7 architecture further improved performance, achieving the highest test accuracy (61.03%) and macro F1 score (61.85%). Compared with the CNN-only baselines, this represents a substantial improvement in precision–recall balance and overall robustness.
4.3. Architectural Analysis and Performance Trends
4.3.1. Hybrid Architecture Advantages
The improved performance of the hybrid DeiT-Small + EfficientNet-B7 model can be attributed to complementary feature integration rather than reliance on a single architectural component. EfficientNet-B7 captures localized morphological and textural features relevant to cellular structure, while the DeiT-Small transformer models global spatial relationships and long-range dependencies within microscopy images.
The reduced generalization gap (17.48%) relative to CNN-only models (>31%) suggests that architectural diversity provides implicit regularization under SMOTE-balanced training. Attention mechanisms may help suppress spurious correlations amplified by oversampling, while convolutional features preserve a biologically meaningful local structure. Together, these components yield more stable representations under class imbalance.
4.3.2. Cross-Methodological Performance Analysis
A comparison across all evaluated paradigms revealed distinct performance characteristics:
- The prompt-based LLMs (30.37–58.82% accuracy) demonstrated variable performance, depending on the model scale and prompting strategy, but remained limited by the lack of domain-specific visual fine-tuning.
- The CNN-only models (32.01–35.12% test accuracy) exhibited near-random performance under data scarcity, highlighting the limitations of conventional transfer learning for microscopy imagery.
- The transformer and hybrid models (57.39–61.03% test accuracy) achieved improved generalization, with the hybrid CNN–transformer architecture providing the most consistent precision–recall balance.
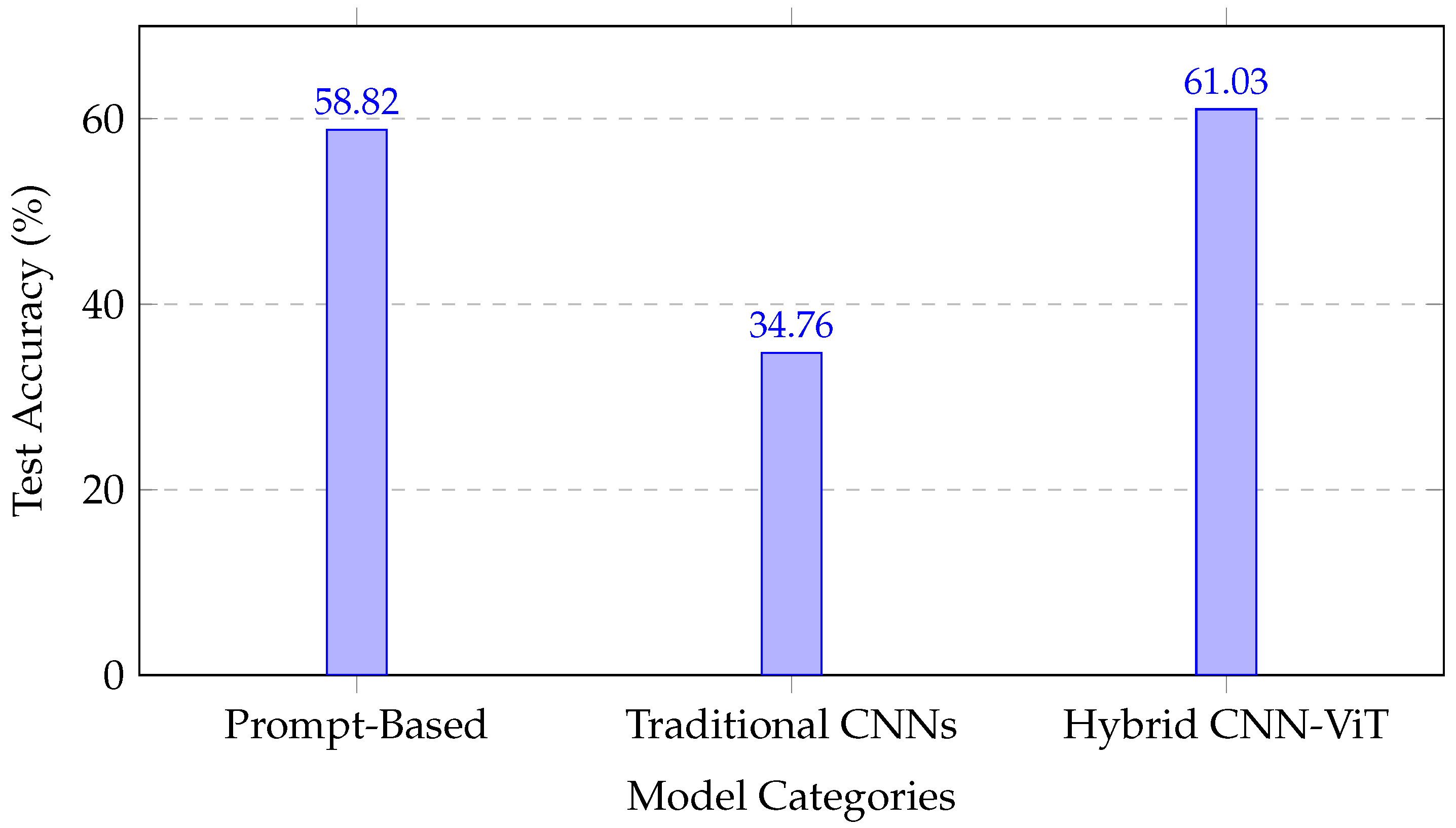
Figure 5 illustrates the comparative performance across methodological categories, emphasizing the robustness gains achieved by integrating convolutional and attention-based representations.
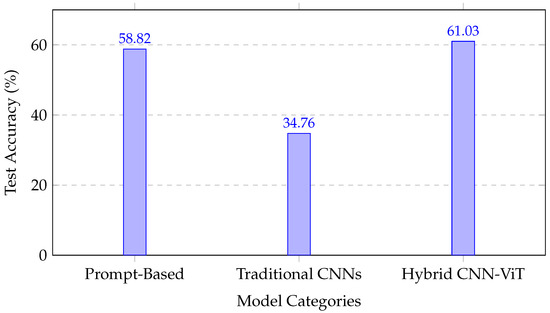
Figure 5.
Performance comparison across methodological approaches, showing the superior performance of the hybrid CNN–transformer architecture.
4.3.3. Statistical Significance and Clinical Relevance
The increase in the macro F1 score from approximately 33% for the CNN-only models to 61.85% for the hybrid architecture represents a substantial improvement in classification reliability under controlled SMOTE-balanced training. While these results do not yet meet the thresholds required for clinical deployment, they demonstrate meaningful gains in automated cell type discrimination relevant to exploratory Alzheimer’s disease research.
Importantly, all evaluation metrics were computed on original microscopy images, ensuring that the observed improvements reflect enhanced generalization rather than memorization of synthetic samples. Table 4 provides a consolidated summary of performance across all evaluated approaches.

Table 4.
Summary of key performance metrics across all approaches. Bold indicates our model.
The results position hybrid CNN–transformer architectures as a promising direction for cell-level Alzheimer’s disease image analysis under data-limited conditions, while underscoring the continued need for larger datasets and domain-specific pretraining to enable clinical translation.
5. Discussion and Limitations
This study presented a comparative evaluation of conventional convolutional neural networks (CNNs), prompt-based multimodal large language models (LLMs), and a hybrid CNN–transformer architecture for Alzheimer’s disease (AD) cell type classification using phase-contrast microscopy images. Among the evaluated approaches, the hybrid DeiT-Small + EfficientNet-B7 model achieved the strongest performance, attaining a test accuracy of 61.03% and a macro F1 score of 61.85 (Table 4). This constitutes a substantial improvement over standalone CNN baselines, whose F1 scores remained near chance levels (32.01–34.76%). These results indicate that coupling convolutional inductive biases with attention-based global reasoning yields more discriminative representations for fine-grained cellular image analysis, consistent with recent advances in hybrid vision architectures [49].
The observed performance gains can be attributed to complementary feature learning mechanisms. Convolutional layers effectively capture localized morphological characteristics such as texture, cell boundaries, and neuritic structures, whereas transformer-based self-attention enables modeling of long-range spatial dependencies and global contextual organization. This synergy is particularly well suited to microscopy imagery, where accurate classification depends on both cellular-scale morphology and broader spatial arrangements. Similar hybrid designs have demonstrated improved robustness and generalization across biomedical imaging tasks, further supporting the suitability of this architectural paradigm for cellular-level AD analysis [49,50].
In contrast, transfer learning from natural image datasets (e.g., ImageNet) proved inadequate for this task. The CNN-only baselines exhibited pronounced validation–test performance gaps exceeding 31%, indicating limited generalization under conditions of data scarcity and substantial domain shifts. This finding aligns with prior work demonstrating that ImageNet-pretrained representations often encode inductive biases poorly aligned with biomedical and microscopy image statistics [51,52]. Collectively, these results underscore the importance of domain-aware pretraining strategies, as emphasized in recent foundation-model research for pathology and neuroscience imaging [53,54].
The prompt-based multimodal LLMs achieved moderate performance (30.37–58.82% accuracy), with outcomes strongly influenced by the model scale and prompting strategy. While these models offer interpretability advantages through natural language reasoning, their quantitative performance remains limited in the absence of domain-specific visual adaptation. The difficulty of capturing subtle cellular morphology via text-guided inference alone likely contributes to this gap. Nevertheless, the observed performance ordering (GPT-5 > GPT-4o > Gemini 2.5 Flash) suggests that increasing the model capacity and improving multimodal alignment may enhance applicability to specialized biomedical tasks, consistent with broader trends in vision–language modeling [38].
Despite the relative advantages of the hybrid architecture, several limitations warrant consideration. First, the absolute classification accuracy (61.03%) remains substantially below thresholds typically required for clinical deployment, which often exceed 90% [43,55]. Second, although the synthetic minority over-sampling technique (SMOTE) was applied in a strictly controlled manner—restricted to feature space and training folds—the effective dataset size (960 samples) remains limited for training high-capacity transformer-based models. The persistent 17.48% validation–test gap suggests residual overfitting, reflecting constraints imposed by the limited sample diversity and partial reliance on synthetic balancing.
Additionally, this study was restricted to a single imaging modality and three cell types: astrocytes, cortical neurons, and SH-SY5Y neuroblastoma cells. While these classes are biologically relevant to AD pathology, the narrow scope limits the generalizability of the conclusions. Furthermore, the evaluation of multimodal LLMs was intentionally confined to prompt-based inference without weight adaptation to ensure fairness and reproducibility; however, this design choice precludes assessment of their full potential under domain-specific fine-tuning.
Overall, the findings indicate that effective AD cell type classification requires architectures explicitly tailored to biomedical imagery rather than direct repurposing of natural image models. Hybrid CNN–fransformer frameworks have emerged as a promising direction, as they integrate fine-grained cellular morphology with global spatial context, both of which are critical for modeling neurodegenerative disease mechanisms [46,56]. Future work should prioritize the expansion of microscopy datasets through multi-center collaborations, the development of microscopy-specific pretraining and augmentation strategies, and the integration of complementary modalities such as gene expression and protein biomarkers. Incorporating explainable AI (XAI) techniques will also be essential to improve interpretability and foster clinical trust in automated cellular analysis systems [57,58].
In summary, despite constraints related to dataset size and scope, this study provides a methodologically rigorous proof of concept demonstrating the advantages of hybrid CNN–transformer architectures for AD-related cell type classification. The results establish a foundation for future biologically informed and clinically relevant computational models while clearly delineating the challenges that must be addressed to advance toward real-world deployment.
6. Conclusions
This study presented a hybrid vision transformer–CNN framework for Alzheimer’s disease (AD) cell type classification from phase-contrast microscopy images. By integrating the global contextual modeling of a DeiT-Small transformer with the fine-grained morphological feature extraction of EfficientNet-B7, the proposed architecture demonstrated improved generalization compared with conventional CNN and standalone transformer baselines under data-limited conditions. The model achieved a test accuracy of 61.03% and a macro F1 score of 61.85, highlighting the advantages of combining convolutional and attention-based representations for cellular image analysis.
The results indicate that direct transfer learning from natural image datasets is insufficient for microscopy-based AD cell classification, whereas hybrid architectures better capture the complementary local and global visual cues required for this task. Prompt-based multimodal large language models, evaluated as zero- and few-shot baselines, provide interpretability benefits but remain limited in quantitative performance without domain-specific visual adaptation.
Despite these promising findings, the study was constrained by a modest dataset size, class imbalance, and a single imaging modality, which collectively limit clinical applicability. Accordingly, the presented results should be interpreted as a proof of concept rather than a deployable diagnostic system.
Future work will focus on expanding microscopy datasets through multi-center collaborations, exploring domain-specific pretraining strategies, incorporating additional biological modalities, and integrating explainable AI (XAI) techniques to improve interpretability and reliability. These directions are essential for advancing toward biologically grounded and clinically meaningful computational tools for Alzheimer’s disease research.
Author Contributions
Conceptualization, M.E.H.; Methodology, M.E.H. and M.T.H.F.; Formal analysis, M.E.H. and M.T.H.F.; Investigation, M.E.H. and M.T.H.F.; Software, M.E.H. and M.T.H.F.; Visualization, M.E.H., M.T.H.F. and O.S.; Writing—Original Draft, M.E.H.; Writing—Review & Editing, M.E.H., M.T.H.F., O.S. and A.W.; Supervision, A.W. All authors have read and agreed to the published version of the manuscript.
Funding
This research received no external funding.
Institutional Review Board Statement
Ethical review and approval were waived for this study due to the data used were obtained from the public databases.
Informed Consent Statement
Patient consent was waived due to the data used were obtained from the public databases.
Data Availability Statement
The data that support the findings of this study are available upon reasonable request.
Conflicts of Interest
The authors declare no conflicts of interest.
References
- Goedert, M.; Spillantini, M.G. A century of alzheimer’s disease. Science 2006, 314, 777–781. [Google Scholar] [CrossRef]
- Miller, E.M.; Chan, T.C.D.; Montes-Matamoros, C.; Sharif, O.; Pujo-Menjouet, L.; Lindstrom, M.R. Oscillations in neuronal activity: A neuron-centered spatiotemporal model of the unfolded protein response in prion diseases. Bull. Math. Biol. 2024, 86, 82. [Google Scholar] [CrossRef]
- Duyckaerts, C.; Delatour, B.; Potier, M.-C. Classification and basic pathology of alzheimer disease. Acta Neuropathol. 2009, 118, 5–36. [Google Scholar] [CrossRef]
- Alzheimer’s Disease Facts and Figures. Available online: https://www.alz.org/alzheimers-dementia/facts-figures#:~:text=more%20than%206%20million%20americans%20of%20all%20ages%20have%20alzheimer’s,older%20(10.7%25)%20has%20alzheimer’s (accessed on 1 December 2022).
- Garwood, C.; Ratcliffe, L.; Simpson, J.; Heath, P.; Ince, P.; Wharton, S. astrocytes in alzheimer’s disease and other age-associated dementias: A supporting player with a central role. Neuropathol. Appl. Neurobiol. 2017, 43, 281–298. [Google Scholar] [CrossRef]
- Gupta, A.D.; Asan, L.; John, J.; Beretta, C.; Kuner, T.; Knabbe, J. Accurate classification of major brain cell types using in vivo imaging and neural network processing. PLoS Biol. 2023, 21, e3002357. [Google Scholar]
- Edlund, C.; Jackson, T.R.; Khalid, N.; Bevan, N.; Dale, T.; Dengel, A.; Ahmed, S.; Trygg, J.; Sjögren, R. Livecell—A large-scale dataset for label-free live cell segmentation. Nat. Methods 2021, 18, 1038–1045. [Google Scholar] [CrossRef]
- Alshammari, M.; Mezher, M. A modified convolutional neural networks for mri-based images for detection and stage classification of alzheimer disease. In Proceedings of the 2021 National Computing Colleges Conference (NCCC), Taif, Saudi Arabia, 27–28 March 2021; IEEE: Piscataway, NJ, USA, 2021; pp. 1–7. [Google Scholar]
- El-Sappagh, S.; Alonso-Moral, J.M.; Abuhmed, T.; Ali, F.; Bugarín-Diz, A. Trustworthy artificial intelligence in alzheimer’s disease: State of the art, opportunities, and challenges. Artif. Intell. Rev. 2023, 56, 11149–11296. [Google Scholar] [CrossRef]
- Wäldchen, J.; Mäder, P. Machine learning for image based species identification. Methods Ecol. Evol. 2018, 9, 2216–2225. [Google Scholar] [CrossRef]
- Huang, G.; Liu, Z.; Maaten, L.V.D.; Weinberger, K.Q. Densely connected convolutional networks. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Honolulu, HI, USA, 21–26 July 2017; pp. 4700–4708. [Google Scholar]
- Solano-Rojas, B.; Villalón-Fonseca, R. A low-cost three-dimensional densenet neural network for alzheimer’s disease early discovery. Sensors 2021, 21, 1302. [Google Scholar] [CrossRef]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep residual learning for image recognition. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Las Vegas, NV, USA, 27–30 June 2016; pp. 770–778. [Google Scholar]
- Fulton, L.V.; Dolezel, D.; Harrop, J.; Yan, Y.; Fulton, C.P. Classification of alzheimer’s disease with and without imagery using gradient boosted machines and resnet-50. Brain Sci. 2019, 9, 212. [Google Scholar] [CrossRef]
- Howard, A.G.; Zhu, M.; Chen, B.; Kalenichenko, D.; Wang, W.; Weyand, T.; Andreetto, M.; Adam, H. Mobilenets: Efficient convolutional neural networks for mobile vision applications. arXiv 2017, arXiv:1704.04861. [Google Scholar] [CrossRef]
- Lu, X.; Wu, H.; Zeng, Y. Classification of alzheimer’s disease in mobilenet. In Journal of Physics: Conference Series; IOP Publishing: Bristol, UK, 2019; Volume 1345, p. 042012. [Google Scholar]
- Szegedy, C.; Liu, W.; Jia, Y.; Sermanet, P.; Reed, S.; Anguelov, D.; Erhan, D.; Vanhoucke, V.; Rabinovich, A. Going deeper with convolutions. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Boston, MA, USA, 7–12 June 2015; pp. 1–9. [Google Scholar]
- Cui, Z.; Gao, Z.; Leng, J.; Zhang, T.; Quan, P.; Zhao, W. Alzheimer’s disease diagnosis using enhanced inception network based on brain magnetic resonance image. In Proceedings of the 2019 IEEE International Conference on Bioinformatics and Biomedicine (BIBM), San Diego, CA, USA, 18–21 November 2019; IEEE: Piscataway, NJ, USA, 2019; pp. 2324–2330. [Google Scholar]
- Sethi, M.; Ahuja, S.; Singh, S.; Snehi, J.; Chawla, M. An intelligent framework for alzheimer’s disease classification using efficientnet transfer learning model. In Proceedings of the 2022 International Conference on Emerging Smart Computing and Informatics (ESCI), Pune, India, 9–11 March 2022; IEEE: Piscataway, NJ, USA, 2022; pp. 1–4. [Google Scholar]
- Zheng, B.; Gao, A.; Huang, X.; Li, Y.; Liang, D.; Long, X. A modified 3d efficientnet for the classification of alzheimer’s disease using structural magnetic resonance images. IET Image Process 2023, 17, 77–87. [Google Scholar] [CrossRef]
- Verkhratsky, A.; Olabarria, M.; Noristani, H.N.; Yeh, C.-Y.; Rodriguez, J.J. Astrocytes in alzheimer’s disease. Neurotherapeutics 2010, 7, 399–412. [Google Scholar] [CrossRef]
- Gómez-Isla, T.; Price, J.L.; McKeel, D.W., Jr.; Morris, J.C.; Growdon, J.H.; Hyman, B.T. Profound loss of layer ii entorhinal cortex neurons occurs in very mild alzheimer’s disease. J. Neurosci. 1996, 16, 4491–4500. [Google Scholar] [CrossRef]
- Riegerova, P.; Brejcha, J.; Bezděková, D.; Chum, T.; Mašínová, E.; Čermáková, N.; Ovsepian, S.V.; Cebecauer, M.; Štefl, M. Expression and localization of aβpp in sh-sy5y cells depends on differentiation state. J. Alzheimer’s Dis. 2021, 82, 485–491. [Google Scholar] [CrossRef]
- Querbes, O.; Aubry, F.; Pariente, J.; Lotterie, J.-A.; Démonet, J.-F.; Duret, V.; Puel, M.; Berry, I.; Fort, J.-C.; Celsis, P.; et al. Early diagnosis of alzheimer’s disease using cortical thickness: Impact of cognitive reserve. Brain 2009, 132, 2036–2047. [Google Scholar] [CrossRef]
- Yin, Q.-Q.; Dong, C.-F.; Dong, S.-Q.; Dong, X.-L.; Hong, Y.; Hou, X.-Y.; Luo, D.-Z.; Pei, J.-J.; Liu, X.-P. Ages induce cell death via oxidative and endoplasmic reticulum stresses in both human sh-sy5y neuroblastoma cells and rat cortical neurons. Cell. Mol. Neurobiol. 2012, 32, 1299–1309. [Google Scholar] [CrossRef]
- Zhang, C.; Lu, Y. Study on artificial intelligence: The state of the art and future prospects. J. Ind. Inf. Integr. 2021, 23, 100224. [Google Scholar] [CrossRef]
- Tan, M.; Le, Q. Efficientnet: Rethinking model scaling for convolutional neural networks. In Proceedings of the International Conference on Machine Learning, Long Beach, CA, USA, 9–15 June 2019; PMLR: New York, NY, USA, 2019; pp. 6105–6114. [Google Scholar]
- Fathi, S.; Ahmadi, M.; Dehnad, A. Early diagnosis of alzheimer’s disease based on deep learning: A systematic review. Comput. Biol. Med. 2022, 146, 105634. [Google Scholar] [CrossRef]
- Jain, R.; Jain, N.; Aggarwal, A.; Hemanth, D.J. Convolutional neural network based alzheimer’s disease classification from magnetic resonance brain images. Cogn. Syst. Res. 2019, 57, 147–159. [Google Scholar] [CrossRef]
- Ding, Y.; Sohn, J.H.; Kawczynski, M.G.; Trivedi, H.; Harnish, R.; Jenkins, N.W.; Lituiev, D.; Copeland, T.P.; Aboian, M.S.; Aparici, C.M.; et al. A deep learning model to predict a diagnosis of alzheimer disease by using 18f-fdg pet of the brain. Radiology 2019, 290, 456–464. [Google Scholar] [CrossRef]
- Chitradevi, D.; Prabha, S. Analysis of brain sub regions using optimization techniques and deep learning method in alzheimer disease. Appl. Soft Comput. 2020, 86, 105857. [Google Scholar] [CrossRef]
- Nawaz, H.; Maqsood, M.; Afzal, S.; Aadil, F.; Mehmood, I.; Rho, S. A deep feature-based real-time system for alzheimer disease stage detection. Multimed. Tools Appl. 2021, 80, 35789–35807. [Google Scholar] [CrossRef]
- Kundaram, S.S.; Pathak, K.C. Deep learning-based alzheimer disease detection. In Proceedings of the Fourth International Conference on Microelectronics, Computing and Communication Systems: MCCS 2019, Coimbatore, India, 17–19 July 2019; Springer: Berlin/Heidelberg, Germany, 2021; pp. 587–597. [Google Scholar]
- Vu, T.D.; Yang, H.-J.; Nguyen, V.Q.; Oh, A.-R.; Kim, M.-S. Multimodal learning using convolution neural network and sparse autoencoder. In Proceedings of the 2017 IEEE International Conference on Big Data and Smart Computing (BigComp), Jeju, Republic of Korea, 13–16 February 2017; IEEE: Piscataway, NJ, USA, 2017; pp. 309–312. [Google Scholar]
- Salehi, A.W.; Baglat, P.; Sharma, B.B.; Gupta, G.; Upadhya, A. A cnn model: Earlier diagnosis and classification of alzheimer disease using mri. In Proceedings of the 2020 International Conference on Smart Electronics and Communication (ICOSEC), Trichy, India, 10–12 September 2020; IEEE: Piscataway, NJ, USA, 2020; pp. 156–161. [Google Scholar]
- Basaia, S.; Agosta, F.; Wagner, L.; Canu, E.; Magnani, G.; Santangelo, R.; Filippi, M.; the Alzheimer’s Disease Neuroimaging Initiative. Automated classification of alzheimer’s disease and mild cognitive impairment using a single mri and deep neural networks. Neuroimage Clin. 2019, 21, 101645. [Google Scholar] [CrossRef] [PubMed]
- Khvostikov, A.; Aderghal, K.; Benois-Pineau, J.; Krylov, A.; Catheline, G. 3d cnn-based classification using smri and md-dti images for alzheimer disease studies. arXiv 2018, arXiv:1801.05968. [Google Scholar]
- Dai, W.; Li, J.; Li, D.; Tiong, A.; Zhao, J.; Wang, W.; Li, B.; Fung, P.N.; Hoi, S. Instructblip: Towards general-purpose vision-language models with instruction tuning. Adv. Neural Inf. Process. Syst. 2023, 36, 49250–49267. [Google Scholar]
- Liu, H.; Li, C.; Wu, Q.; Lee, Y.J. Visual instruction tuning. Adv. Neural Inf. Process. Syst. 2023, 36, 34892–34916. [Google Scholar]
- Wang, Z.; Wu, Z.; Agarwal, D.; Sun, J. Medclip: Contrastive learning from unpaired medical images and text. In Proceedings of the 2022 Conference on Empirical Methods in Natural Language Processing (EMNLP), Abu Dhabi, United Arab Emirates, 7–11 December 2022; pp. 3876–3887. [Google Scholar]
- Ramya, J.; Maheswari, B.U.; Rajakumar, M.; Sonia, R. Alzheimer’s disease segmentation and classification on mri brain images using enhanced expectation maximization adaptive histogram (eem-ah) and machine learning. Inf. Technol. Control 2022, 51, 786–800. [Google Scholar] [CrossRef]
- Rangaraju, B.; Chinnadurai, T.; Natarajan, S.; Raja, V. Dual attention aware octave convolution network for early-stage alzheimer’s disease detection. Inf. Technol. Control 2024, 53, 302–316. [Google Scholar] [CrossRef]
- Odusami, M.; Maskeliūnas, R.; Damaševičius, R. Pixel-level fusion approach with vision transformer for early detection of alzheimer’s disease. Electronics 2023, 12, 1218. [Google Scholar] [CrossRef]
- Odusami, M.; Maskeliūnas, R.; Damaševičius, R.; Misra, S. Explainable deep-learning-based diagnosis of alzheimer’s disease using multimodal input fusion of pet and mri images. J. Med. Biol. Eng. 2023, 43, 291–302. [Google Scholar] [CrossRef]
- Odusami, M.; Maskeliūnas, R.; Damaševičius, R. Pareto optimized adaptive learning with transposed convolution for image fusion alzheimer’s disease classification. Brain Sci. 2023, 13, 1045. [Google Scholar] [CrossRef]
- Odusami, M.; Maskeliūnas, R.; Damaševičius, R. Optimized convolutional fusion for multimodal neuroimaging in alzheimer’s disease diagnosis: Enhancing data integration and feature extraction. J. Pers. Med. 2023, 13, 1496. [Google Scholar] [CrossRef]
- Chawla, N.V.; Bowyer, K.W.; Hall, L.O.; Kegelmeyer, W.P. Smote: Synthetic minority over-sampling technique. J. Artif. Intell. Res. 2002, 16, 321–357. [Google Scholar] [CrossRef]
- Hicks, S.A.; Strümke, I.; Thambawita, V.; Hammou, M.; Riegler, M.A.; Halvorsen, P.; Parasa, S. On evaluation metrics for medical applications of artificial intelligence. Sci. Rep. 2022, 12, 5979. [Google Scholar] [CrossRef]
- Borji, A.; Kronreif, G.; Angermayr, B.; Hatamikia, S. Advanced hybrid deep learning model for enhanced evaluation of osteosarcoma histopathology images. Front. Med. 2025, 12, 1555907. [Google Scholar] [CrossRef] [PubMed]
- He, R.; Wang, Y.; Li, J.; Wang, H. Hybrid cnn-transformer architecture for medical image segmentation: A comprehensive review. Biomed. Signal Process. Control 2023, 84, 104796. [Google Scholar]
- Raghu, M.; Zhang, C.; Kleinberg, J.; Bengio, S. Transfusion: Understanding transfer learning for medical imaging. In Proceedings of the 33rd Conference on Neural Information Processing Systems (NeurIPS 2019), Vancouver, BC, Canada, 8–14 December 2019; Volume 32. [Google Scholar]
- Azizi, S.; Mustafa, B.; Ryan, F.; Beaver, Z.; Freyberg, J.; Deaton, J.; Loh, A.; Karthikesalingam, A.; Kornblith, S.; Chen, T.; et al. Big self-supervised models advance medical image classification. In Proceedings of the IEEE/CVF International Conference on Computer Vision, Montreal, BC, Canada, 11–17 October 2021; pp. 3478–3488. [Google Scholar]
- Tang, R.; Chen, F.; Lu, Z.; Yan, Z.; Chen, M.; Zhang, S. Foundation models for generalist medical artificial intelligence. Nature 2023, 624, 540–548. [Google Scholar]
- Jha, D.; Smedsrud, P.H.; Riegler, M.A.; Halvorsen, P.; Johansen, D.; de Lange, T.; Johansen, H.D. Contrastive learning for label-efficient pathological image analysis. Med. Image Anal. 2022, 81, 102559. [Google Scholar]
- Takahashi, S.; Sakaguchi, Y.; Kouno, N.; Takasawa, K.; Ishizu, K.; Akagi, Y.; Aoyama, R.; Teraya, N.; Bolatkan, A.; Shinkai, N.; et al. Comparison of vision transformers and convolutional neural networks in medical image analysis: A systematic review. J. Med. Syst. 2024, 48, 84. [Google Scholar] [CrossRef] [PubMed]
- Gupta, A.; Kumar, A.; Singh, R. Transformers in medical imaging: A survey on vision transformer applications in medical image analysis. Comput. Biol. Med. 2023, 152, 106376. [Google Scholar]
- Singh, A.; Sengupta, S.; Lakshminarayanan, V. Explainable deep learning models in medical image analysis. J. Imaging 2020, 6, 52. [Google Scholar] [CrossRef] [PubMed]
- Samek, W.; Montavon, G.; Lapuschkin, S.; Anders, C.J.; Müller, K.-R. Explaining deep neural networks and beyond: A review of methods and applications. Proc. IEEE 2021, 109, 247–278. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.