Machine Learning-Driven QSAR Modeling for Predicting Short-Term Exposure Limits of Hydrocarbons and Their Derivatives
Abstract
1. Introduction
2. Principal Theoretical Approaches
2.1. Support Vector Machine (SVM)
2.2. Back-Propagation Artificial Neural Network (BP-ANN)
- (1)
- Forward Propagation
- (2)
- Back-propagation
2.3. Extreme Gradient Boosting (XGBoost)
3. Model Construction Preliminaries
3.1. Determination of the Sample Set
3.2. Determination of Characteristic Molecular Descriptors
3.3. Performance Evaluation Metrics of the Model
4. Results and Discussion
4.1. Model Construction
4.1.1. MLR Model
4.1.2. SVM Model
4.1.3. BP-ANN Model
4.1.4. XGBoost Model
4.2. Model Evaluation and Validation
4.2.1. Statistical Quality and Validation Analysis
4.2.2. Comparative Error Analysis
4.2.3. Applicability Domain Evaluation
4.2.4. Interpretability Analysis of the Optimal XGBoost Model Using SHAP
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Kamal, A.; Malik, R.N.; Fatima, N.; Rashid, A. Chemical exposure in occupational settings and related health risks: A neglected area of research in Pakistan. Environ. Toxicol. Pharmacol. 2012, 34, 46–58. [Google Scholar] [CrossRef]
- Barros, B.; Oliveira, M.; Morais, S. Firefighters’ occupational exposure: Contribution from biomarkers of effect to assess health risks. Environ. Int. 2021, 156, 106704. [Google Scholar] [CrossRef]
- Pyatt, J.R.; Gilmore, I.; Mullins, P.A. Abnormal liver function tests following inadvertent inhalation of volatile hydrocarbons. Postgrad. Med. J. 1998, 74, 747–748. [Google Scholar] [CrossRef] [PubMed]
- Boffetta, P.; Jourenkova, N.; Gustavsson, P. Cancer risk from occupational and environmental exposure to polycyclic aromatic hydrocarbons. Cancer Causes Control 1997, 8, 444–472. [Google Scholar] [CrossRef] [PubMed]
- Yu, Y.; Li, Q.; Wang, H.; Wang, B.; Wang, X.; Ren, A.; Tao, S. Risk of human exposure to polycyclic aromatic hydrocarbons: A case study in Beijing, China. Environ. Pollut. 2015, 205, 70–77. [Google Scholar] [CrossRef] [PubMed]
- U.S. Department of Labor, Occupational Safety and Health Administration (OSHA). Air Contaminants. Code of Federal Regulations. 2023. Available online: https://www.ecfr.gov/current/title-29/subtitle-B/chapter-XVII/part-1910/subpart-Z/section-1910.1000 (accessed on 10 October 2025).
- Song, X. Study on Environmental Control After Chemical Leakage During Storage and Transportation. Ph.D. Thesis, Shanghai Ocean University, Shanghai, China, 2019. [Google Scholar] [CrossRef]
- Zheng, S.; Hu, C.; Huang, S.; Xiao, X.; Luo, L. Occupational Exposure Limits of Air Toxic Substances in the GESTIS Substance Database: Current Status. Chin. J. Ind. Hyg. Occup. Dis. 2024, 42, 417–425. [Google Scholar] [CrossRef]
- Ismail, A.U.; Ibrahim, S.A.; Gambo, M.D.; Muhammad, R.F.; Badamasi, M.M.; Sulaiman, I. Impact of differential occupational LPG exposure on cardiopulmonary indices, liver function, and oxidative stress in Northwestern city of Nigeria. Sci. Total Environ. 2023, 862, 160881. [Google Scholar] [CrossRef] [PubMed]
- Wu, S.; Zhang, Y.; Ma, Y.; Zhang, M.; Huang, D. Carcinogenic Risk Assessment of Occupational Exposure to Low-Level Benzene in a Large-Scale Aromatic Plant. Chin. J. Occup. Med. 2015, 42, 205–207. [Google Scholar] [CrossRef]
- Jiang, X.; Wang, R.; Chang, T.; Zhang, Y.; Zheng, K.; Wan, R.; Wang, X. Effect of short-term air pollution exposure on migraine: A protocol for systematic review and meta-analysis on human observational studies. Environ. Int. 2023, 174, 107892. [Google Scholar] [CrossRef]
- Nunes, L.J.R.; Curado, A. Long-term vs. short-term measurements in indoor Rn concentration monitoring: Establishing a procedure for assessing exposure potential (RnEP). Results Eng. 2023, 17, 100966. [Google Scholar] [CrossRef]
- Zhou, Y.; Xu, C.; Zhang, Y.; Zhao, M.; Hu, Y.; Jiang, Y.; Li, D.; Wu, N.; Wu, L.; Li, C.; et al. Association between short-term nitrogen dioxide exposure and outpatient visits for anxiety: A time-series study in Xi’an, China. Atmos. Environ. 2022, 279, 119122. [Google Scholar] [CrossRef]
- Li, S.-S.; Fang, S.-M.; Chen, J.; Zhang, Z.; Yu, Q.-Y. Effects of short-term exposure to volatile pesticide dichlorvos on the olfactory systems in Spodoptera litura: Calcium homeostasis, synaptic plasticity and apoptosis. Sci. Total Environ. 2023, 864, 161050. [Google Scholar] [CrossRef]
- Liu, Y.; Xu, J.; Shi, J.; Zhang, Y.; Ma, Y.; Zhang, Q.; Su, Z.; Zhang, Y.; Hong, S.; Hu, G.; et al. Effects of short-term high-concentration exposure to PM(2.5) on pulmonary tissue damage and repair ability as well as innate immune events. Environ. Pollut. 2023, 319, 121055. [Google Scholar] [CrossRef]
- Dong, Z.; Li, X.; Chen, Y.; Zhang, N.; Wang, Z.; Liang, Y.-Q.; Guo, Y. Short-term exposure to norethisterone affected swimming behavior and antioxidant enzyme activity of medaka larvae, and led to masculinization in the adult population. Chemosphere 2023, 310, 136844. [Google Scholar] [CrossRef]
- Balasch, J.C.; Brandts, I.; Barria, C.; Martins, M.A.; Tvarijonaviciute, A.; Tort, L.; Oliveira, M.; Teles, M. Short-term exposure to polymethylmethacrylate nanoplastics alters muscle antioxidant response, development and growth in Sparus aurata. Mar. Pollut. Bull. 2021, 172, 112918. [Google Scholar] [CrossRef]
- Pengelly, I.; O’Shea, H.; Smith, G.; Coggins, M.A. Measurement of Diacetyl and 2,3-Pentanedione in the Coffee Industry Using Thermal Desorption Tubes and Gas Chromatography-Mass Spectrometry. Ann. Work Expo. Health 2019, 63, 415–425. [Google Scholar] [CrossRef]
- North, C.M.; Rooseboom, M.; Kocabas, N.A.; Synhaeve, N.; Radcliffe, R.J.; Segal, L. Application of physiologically-based pharmacokinetic modeled toluene blood concentration in the assessment of short term exposure limits. Regul. Toxicol. Pharmacol. 2023, 140, 105380. [Google Scholar] [CrossRef] [PubMed]
- Russell, A.J.; Vincent, M.; Buerger, A.N.; Dotson, S.; Lotter, J.; Maier, A. Establishing short-term occupational exposure limits (STELs) for sensory irritants using predictive and in silico respiratory rate depression (RD50) models. Inhal. Toxicol. 2024, 36, 13–25. [Google Scholar] [CrossRef] [PubMed]
- Du, K.-L.; Jiang, B.; Lu, J.; Hua, J.; Swamy, M.N.S. Exploring Kernel Machines and Support Vector Machines: Principles, Techniques, and Future Directions. Mathematics 2024, 12, 3935. [Google Scholar] [CrossRef]
- Jung, T.; Kim, J. A new support vector machine for categorical features. Expert Syst. Appl. 2023, 229, 120449. [Google Scholar] [CrossRef]
- Baldomero-Naranjo, M.; Martínez-Merino, L.I.; Rodríguez-Chía, A.M. A robust SVM-based approach with feature selection and outliers detection for classification problems. Expert Syst. Appl. 2021, 178, 115017. [Google Scholar] [CrossRef]
- Wu, F.; Zhang, X.; Fang, Z.; Yu, X. Support Vector Machine-Based Global Classification Model of the Toxicity of Organic Compounds to Vibrio fischeri. Molecules 2023, 28, 2703. [Google Scholar] [CrossRef]
- Yao, X.J.; Panaye, A.; Doucet, J.P.; Zhang, R.S.; Chen, H.F.; Liu, M.C.; Hu, Z.D.; Fan, B.T. Comparative Study of QSAR/QSPR Correlations Using Support Vector Machines, Radial Basis Function Neural Networks, and Multiple Linear Regression. J. Chem. Inf. Comput. Sci. 2004, 44, 1257–1266. [Google Scholar] [CrossRef]
- Sengupta, A.; Singh, S.K.; Kumar, R. Support Vector Machine-Based Prediction Models for Drug Repurposing and Designing Novel Drugs for Colorectal Cancer. ACS Omega 2024, 9, 18584–18592. [Google Scholar] [CrossRef] [PubMed]
- Ibrahim, S.I.; Ghoneim, S.S.M.; Taha, I.B.M. DGALab: An extensible software implementation for DGA. IET Gener. Transm. Distrib. 2018, 12, 4117–4124. [Google Scholar] [CrossRef]
- Li, Z.; Huang, J.; Wang, J.; Ding, M. Comparative study of meta-heuristic algorithms for reactor fuel reloading optimization based on the developed BP-ANN calculation method. Ann. Nucl. Energy 2022, 165, 108685. [Google Scholar] [CrossRef]
- Wei, Z.; Su, X.; Wang, D.; Feng, Z.; Gao, Q.; Xu, G.; Zu, G. Three-dimensional processing map based on BP-ANN and interface microstructure of Fe/Al laminated sheet. Mater. Chem. Phys. 2023, 297, 127431. [Google Scholar] [CrossRef]
- Guo, Z.; Guo, C.; Sun, L.; Zuo, M.; Chen, Q.; El-Seedi, H.R.; Zou, X. Identification of the apple spoilage causative fungi and prediction of the spoilage degree using electronic nose. J. Food Process Eng. 2021, 44, e13816. [Google Scholar] [CrossRef]
- Chen, L.-S.; Chung, W.-H.; Chen, Y.; Kuo, S.-Y.J.I.A. AMC with a BP-ANN scheme for 5G enhanced mobile broadband. IEEE Access 2020, 13, 124689–124696. [Google Scholar] [CrossRef]
- Bauer, E.; Kohavi, R. An Empirical Comparison of Voting Classification Algorithms: Bagging, Boosting, and Variants. Mach. Learn. 1999, 36, 105–139. [Google Scholar] [CrossRef]
- The National Institute for Occupational Safety and Health (NIOSH). NIOSH Pocket Guide to Chemical Hazards; Centers for Disease Control and Prevention: Atlanta, GA, USA, 2020. Available online: https://www.cdc.gov/niosh/npg/npgdcas.html (accessed on 4 October 2024).
- Frey, B.J.; Dueck, D. Response to Comment on “Clustering by Passing Messages Between Data Points”. Science 2008, 319, 726. [Google Scholar] [CrossRef]
- Gramatica, P.; Pilutti, P.; Papa, E. Approaches for externally validated QSAR modelling of Nitrated Polycyclic Aromatic Hydrocarbon mutagenicity. SAR QSAR Environ. Res 2007, 18, 169–178. [Google Scholar] [CrossRef]
- Yang, C.; Chen, J.; Wang, R.; Zhang, M.; Zhang, C.; Liu, J. Density Prediction Models for Energetic Compounds Merely Using Molecular Topology. J. Chem. Inf. Model. 2021, 61, 2582–2593. [Google Scholar] [CrossRef]
- Tropsha, A.; Gramatica, P.; Gombar, V.K. The Importance of Being Earnest: Validation is the Absolute Essential for Successful Application and Interpretation of QSPR Models. QSAR Comb. Sci. 2003, 22, 69–77. [Google Scholar] [CrossRef]
- Camellini, F.; Crisci, S.; De Magistris, A.; Franchini, G. A line-search based SGD algorithm with Adaptive Importance Sampling. J. Comput. Appl. Math. 2026, 477, 117120. [Google Scholar] [CrossRef]
- Todeschini, R.; Gramatica, P. The Whim Theory: New 3D Molecular Descriptors for Qsar in Environmental Modelling. SAR QSAR Environ. Res. 1997, 7, 89–115. [Google Scholar] [CrossRef]
- Salmina, E.; Haider, N.; Tetko, I. Extended Functional Groups (EFG): An Efficient Set for Chemical Characterization and Structure-Activity Relationship Studies of Chemical Compounds. Molecules 2015, 21, 1. [Google Scholar] [CrossRef] [PubMed]
- Hair, J.F. Multivariate Data Analysis: An Overview. In International Encyclopedia of Statistical Science; Lovric, M., Ed.; Springer: Berlin/Heidelberg, Germany, 2011; pp. 904–907. [Google Scholar]
- Peng, F.; Lu, Y.; Wang, Y.; Yang, L.; Yang, Z.; Li, H. Predicting the formation of disinfection by-products using multiple linear and machine learning regression. J. Environ. Chem. Eng. 2023, 11, 110612. [Google Scholar] [CrossRef]
- Dufera, A.G.; Liu, T.; Xu, J. Regression models of Pearson correlation coefficient. Stat. Theory Relat. Fields 2023, 7, 97–106. [Google Scholar] [CrossRef]
- Phalen, R.F. Inhalation Studies: Foundations and Techniques, 2nd ed.; CRC Press: Boca Raton, FL, USA, 2002. [Google Scholar]
- Dahl, A.R.; Gerde, P. Uptake and metabolism of toxicants in the respiratory tract. Environ. Health Perspect. 1994, 102, 67–70. [Google Scholar] [CrossRef] [PubMed]
- Enoch, S.J.; Ellison, C.M.; Schultz, T.W.; Cronin, M.T.D. A review of the electrophilic reaction chemistry involved in covalent protein binding relevant to toxicity. Crit. Rev. Toxicol. 2011, 41, 783–802. [Google Scholar] [CrossRef]
- Mansouri, K.; Grulke, C.M.; Judson, R.S.; Williams, A.J. OPERA models for predicting physicochemical properties and environmental fate endpoints. J. Cheminformatics 2018, 10, 10. [Google Scholar] [CrossRef] [PubMed]
- Barber, C.; Heghes, C.; Johnston, L. A framework to support the application of the OECD guidance documents on (Q)SAR model validation and prediction assessment for regulatory decisions. Comput. Toxicol. 2024, 30, 100305. [Google Scholar] [CrossRef]
- Chen, T.; Guestrin, C. XGBoost: A Scalable Tree Boosting System. In Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, San Francisco, CA, USA, 13–17 August 2016; Association for Computing Machinery: New York, NY, USA, 2016; pp. 785–794. [Google Scholar] [CrossRef]
- Nahum, O.E.; Yosipof, A.; Senderowitz, H. A Multi-Objective Genetic Algorithm for Outlier Removal. J. Chem. Inf. Model. 2015, 55, 2507–2518. [Google Scholar] [CrossRef] [PubMed]
- Netzeva, T.I.; Worth, A.; Aldenberg, T.; Benigni, R.; Cronin, M.T.; Gramatica, P.; Jaworska, J.S.; Kahn, S.; Klopman, G.; Marchant, C.A.; et al. Current status of methods for defining the applicability domain of (quantitative) structure-activity relationships. The report and recommendations of ECVAM Workshop 52. Altern. Lab. Anim. 2005, 33, 155–173. [Google Scholar] [CrossRef] [PubMed]













| Sample Count | Median | 25% Quantile | 75% Quantile | Min | Max |
|---|---|---|---|---|---|
| 12 | 0.3868 | 0.3646 | 0.4255 | 0.2857 | 0.65 |
| Name | Type | Definition | VIF |
|---|---|---|---|
| Mor28m | 3D-MoRSe | signal 28/weighted by atomic masses | 1.237 |
| E3u | WHIM | 3rd component accessibility directional WHIM index/unweighted | 1.4 |
| G2m | WHIM | 2nd component symmetry directional WHIM index/weighted by atomic masses | 1.211 |
| n = CHR | functional group | number of secondary C (sp2) | 1.047 |
| Test Variable | R2 | RMSE | SD | p-Value | F |
|---|---|---|---|---|---|
| Result | 0.8043 | 0.8707 | 1.213 | <0.001 | 20.381 |
| Inspection Standards | >0.6 | the smaller, the better | the smaller, the better | <0.05 | >Ftheory |
| Characteristic Molecular Descriptors | Regression Coefficient | Standardized Coefficient | Standard Error | t-Value | p-Value |
|---|---|---|---|---|---|
| Constant | 4.338 | 0.497 | 8.735 | <0.001 | |
| E3u | 5.490 | 0.527 | 1.070 | 5.129 | <0.001 |
| Mor28m | 6.880 | 0.460 | 1.444 | 4.766 | <0.001 |
| n = CHR | −2.216 | −0.339 | 0.582 | −3.811 | <0.001 |
| G2m | −2.313 | −0.360 | 0.613 | −3.771 | <0.001 |
| Performance Parameters | Model | |||||||
|---|---|---|---|---|---|---|---|---|
| MLR | SVM | BP-ANN | XGBoost | |||||
| Training Set | Test Set | Training Set | Test Set | Training Set | Test Set | Training Set | Test Set | |
| R2 | 0.8043 | 0.8095 | 0.8089 | 0.8257 | 0.8396 | 0.8824 | 0.9445 | 0.9152 |
| RMSE | 0.8707 | 0.8595 | 0.854 | 0.8314 | 0.7672 | 0.649 | 0.3608 | 0.6815 |
| MAE | 0.6598 | 0.6994 | 0.6356 | 0.6545 | 0.496 | 0.5069 | 0.1302 | 0.4645 |
| 0.7891 | - | 0.7971 | - | 0.8363 | - | 0.9532 | - | |
| - | 0.7785 | - | 0.7928 | - | 0.8737 | - | 0.9291 | |
| CAS No.: | Name: | Experimental Value ln(STEL): | Molecular Structure |
|---|---|---|---|
| Sample with leverage values above the critical threshold (h*) | |||
| 117-81-7 | di(2-ethylhexyl) phthalate | 2.3 |  |
| Samples outside the ±3 standardized residual range | |||
| 75-31-0 | isopropylamine | 3.17 |  |
| 144-62-7 | oxalic acid | 0.69 | 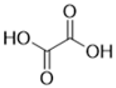 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shi, J.; Wang, C.; Ni, L.; Zhao, W.; Yuan, X. Machine Learning-Driven QSAR Modeling for Predicting Short-Term Exposure Limits of Hydrocarbons and Their Derivatives. Processes 2025, 13, 4025. https://doi.org/10.3390/pr13124025
Shi J, Wang C, Ni L, Zhao W, Yuan X. Machine Learning-Driven QSAR Modeling for Predicting Short-Term Exposure Limits of Hydrocarbons and Their Derivatives. Processes. 2025; 13(12):4025. https://doi.org/10.3390/pr13124025
Chicago/Turabian StyleShi, Jingjie, Cheng Wang, Linli Ni, Wei Zhao, and Xiongjun Yuan. 2025. "Machine Learning-Driven QSAR Modeling for Predicting Short-Term Exposure Limits of Hydrocarbons and Their Derivatives" Processes 13, no. 12: 4025. https://doi.org/10.3390/pr13124025
APA StyleShi, J., Wang, C., Ni, L., Zhao, W., & Yuan, X. (2025). Machine Learning-Driven QSAR Modeling for Predicting Short-Term Exposure Limits of Hydrocarbons and Their Derivatives. Processes, 13(12), 4025. https://doi.org/10.3390/pr13124025






