Efficient Methods for Parameter Estimation of Ordinary and Partial Differential Equation Models of Viral Hepatitis Kinetics
Abstract
1. Introduction
2. Methods
2.1. Development of Mathematical Models
2.1.1. The Standard Biphasic Model
2.1.2. The Multiscale HCV Model
2.2. Data Description
2.3. Solving the Model Equations
2.4. Parameter Estimation
2.4.1. Preliminaries
2.4.2. Optimization by a Constrained Version of Nonlinear Least Squares (Gauss–Newton Method)
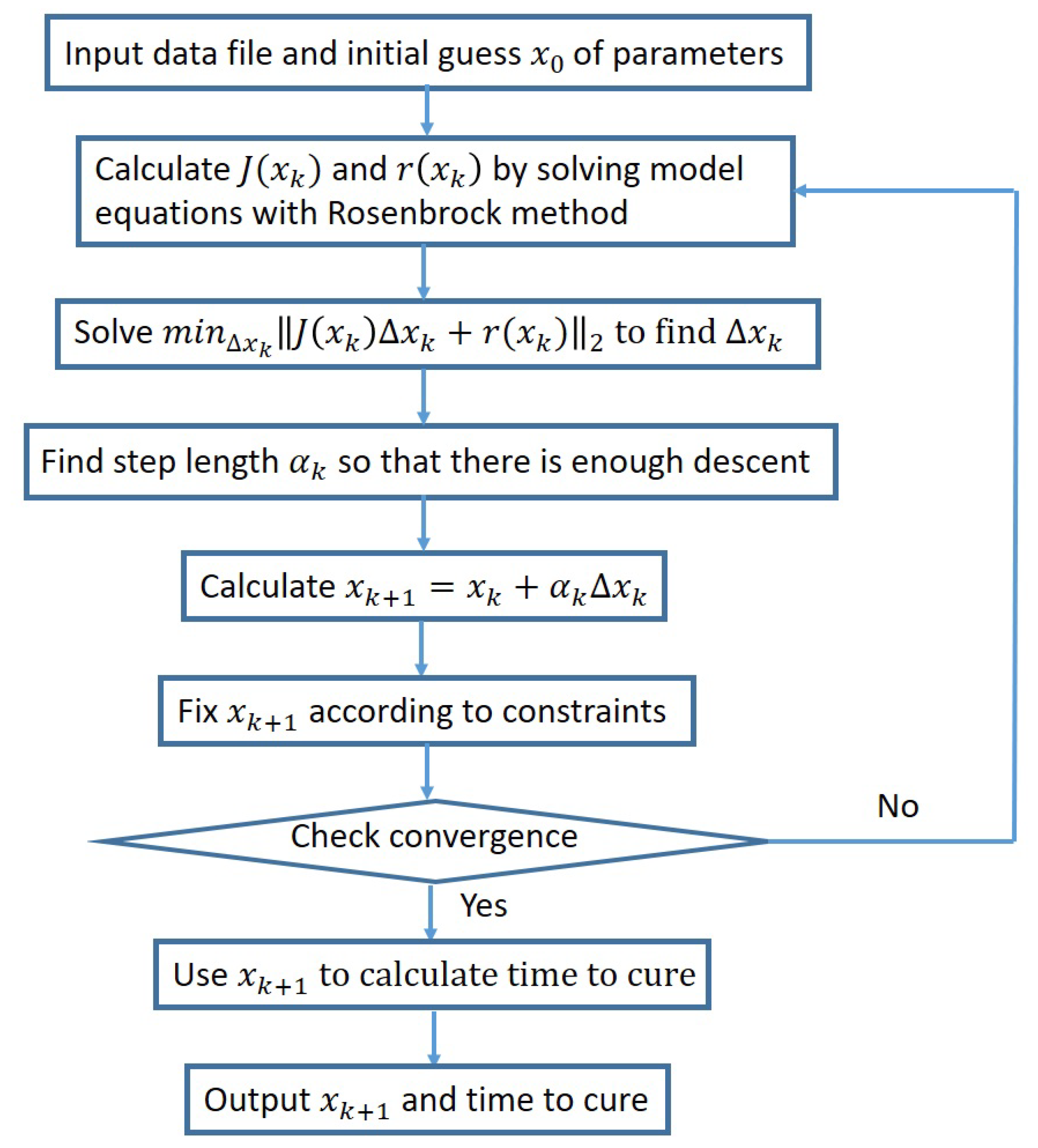
- Start with an initial guess and iterate for
- Solve to compute the correction .
- Choose a step length so that there is enough descent.
- Calculate the new iterate .
- Check for convergence.
2.4.3. Optimization by Derivative-Free Methods (COBYLA Method)
| Algorithm 1: COBYLA method. |
 |
2.5. Method Scope and Other Approaches
2.5.1. Parameters Change When Transforming a PDE Multiscale Model to a System of ODEs
2.5.2. Problematic Issues in Strategies Relying on Canned Methods
3. Results
4. Discussion
5. Conclusions and Future Work
Author Contributions
Funding
Conflicts of Interest
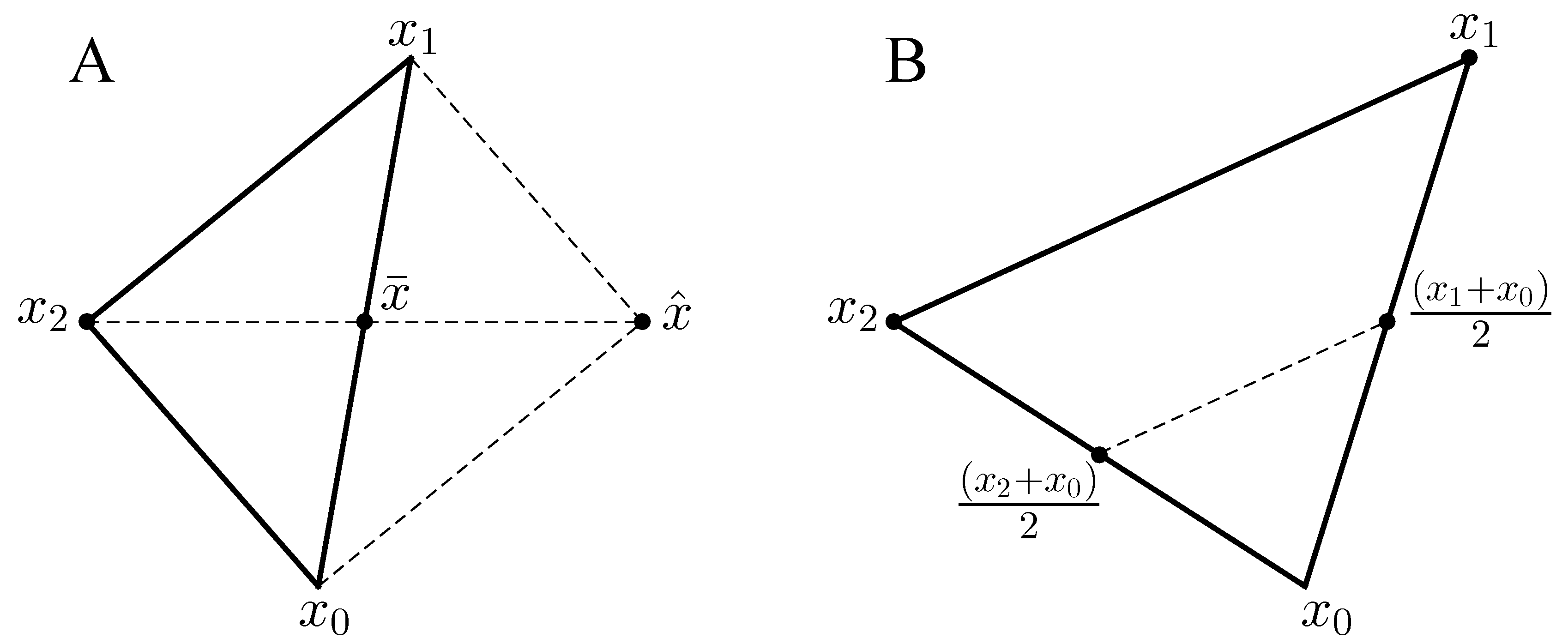
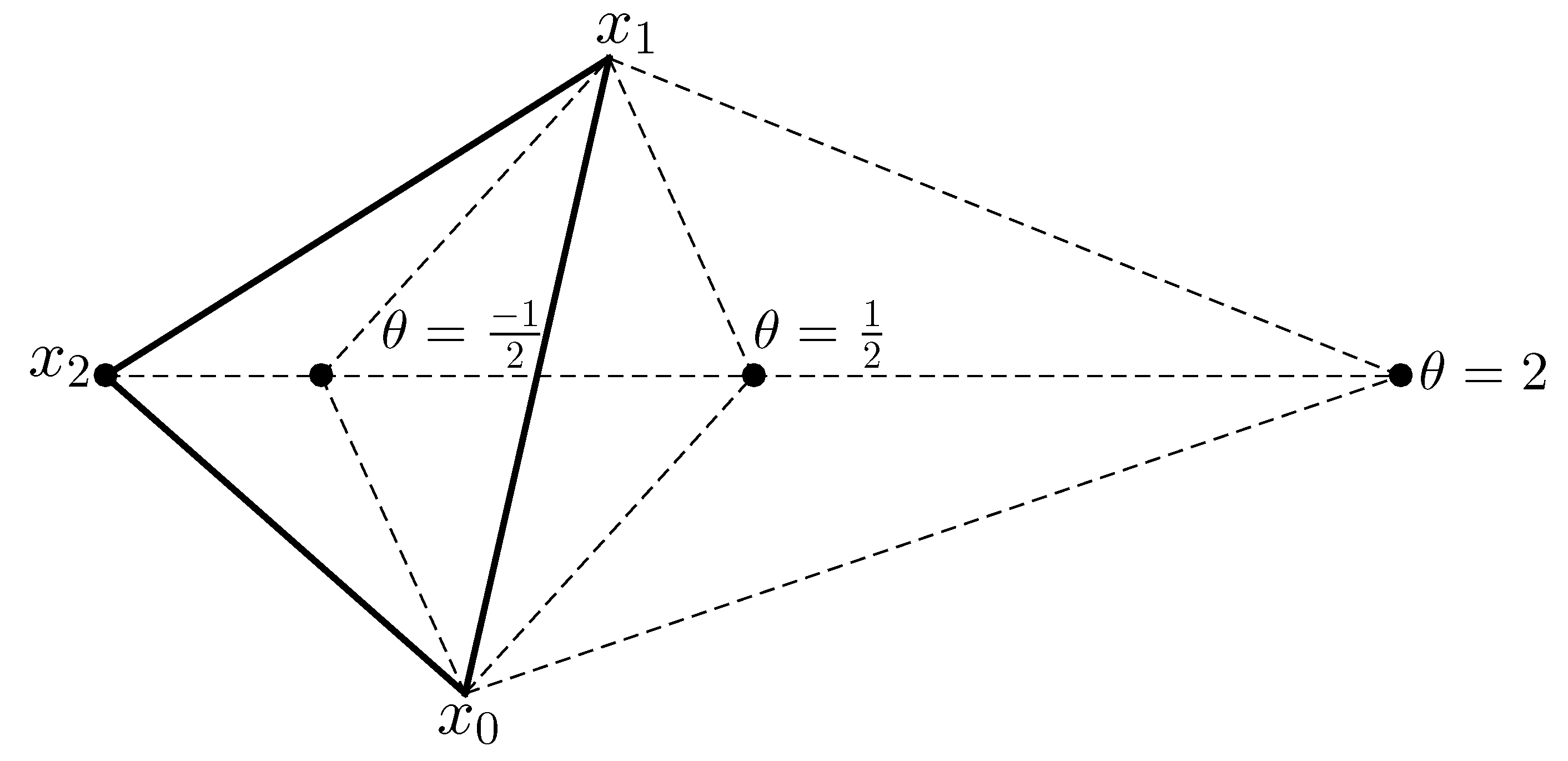
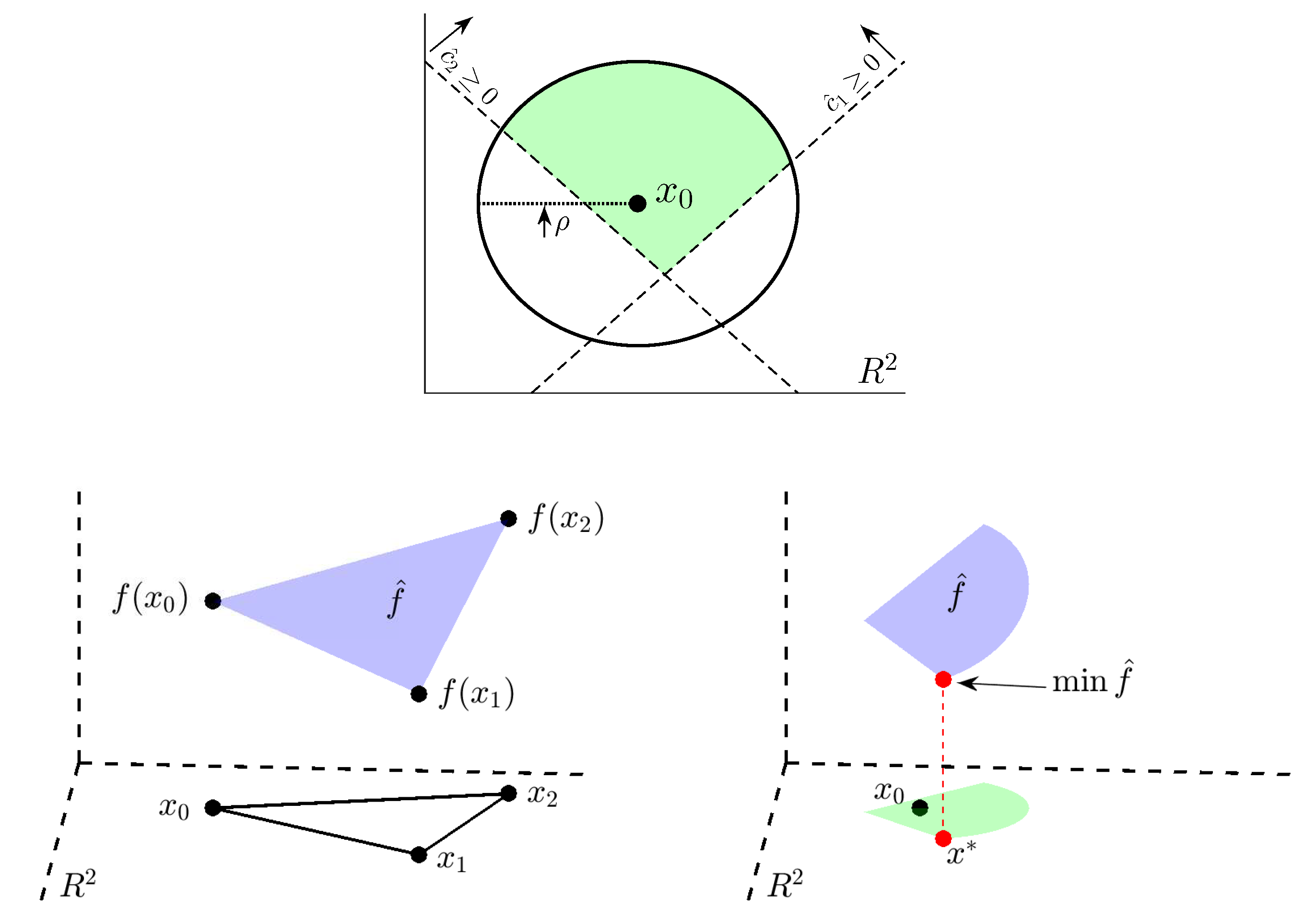
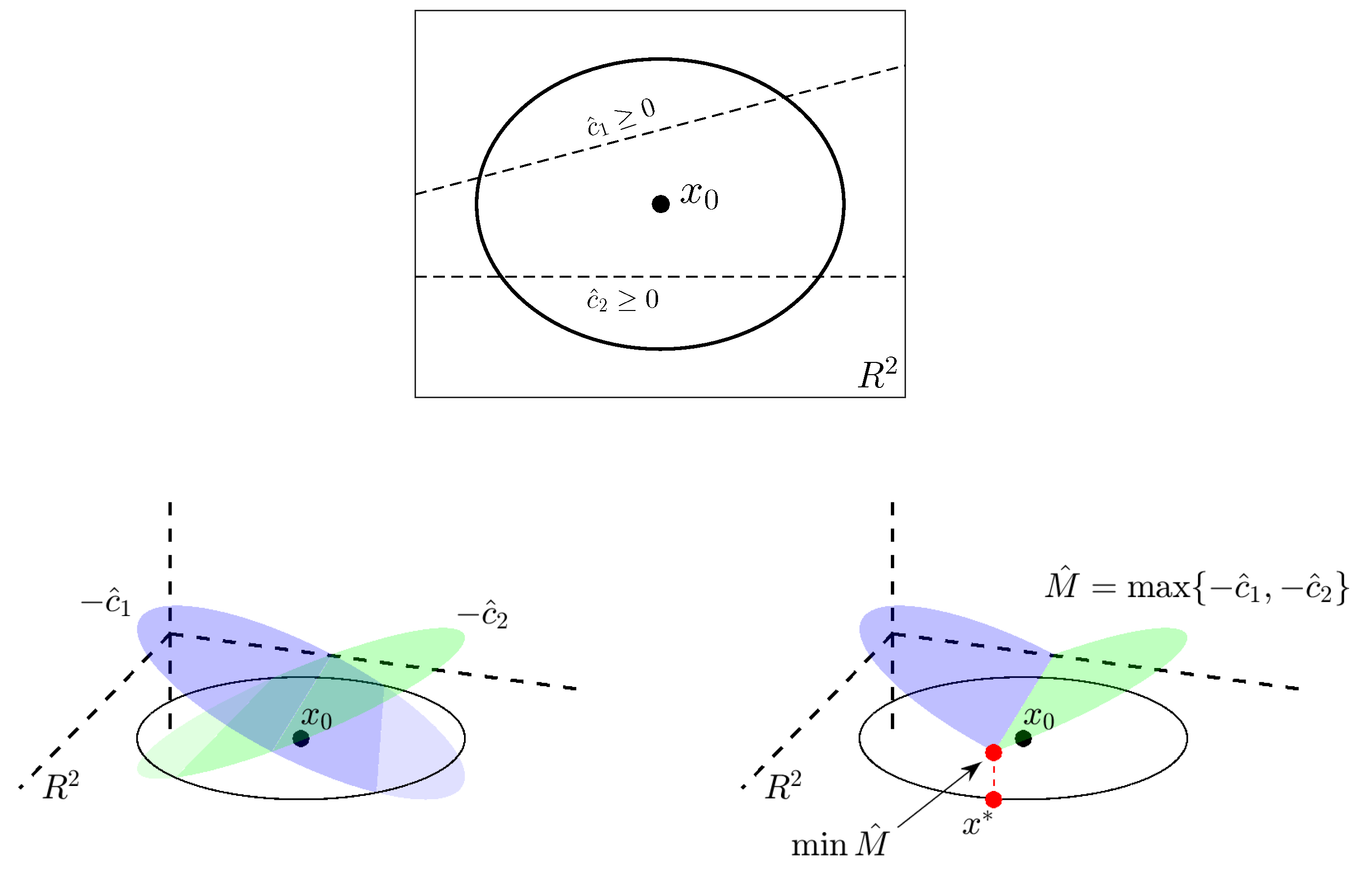
Appendix A. Details of the COBYLA Method

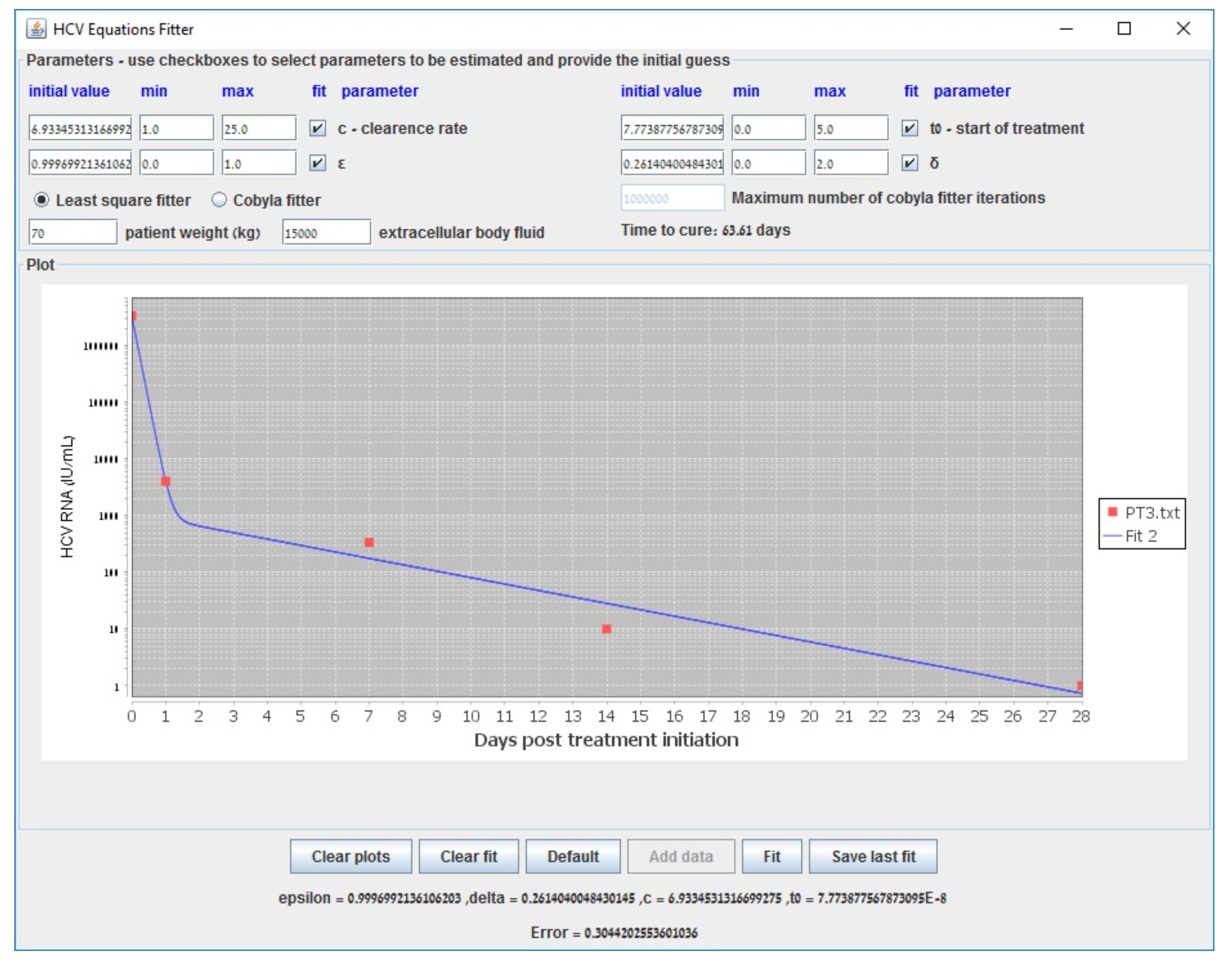
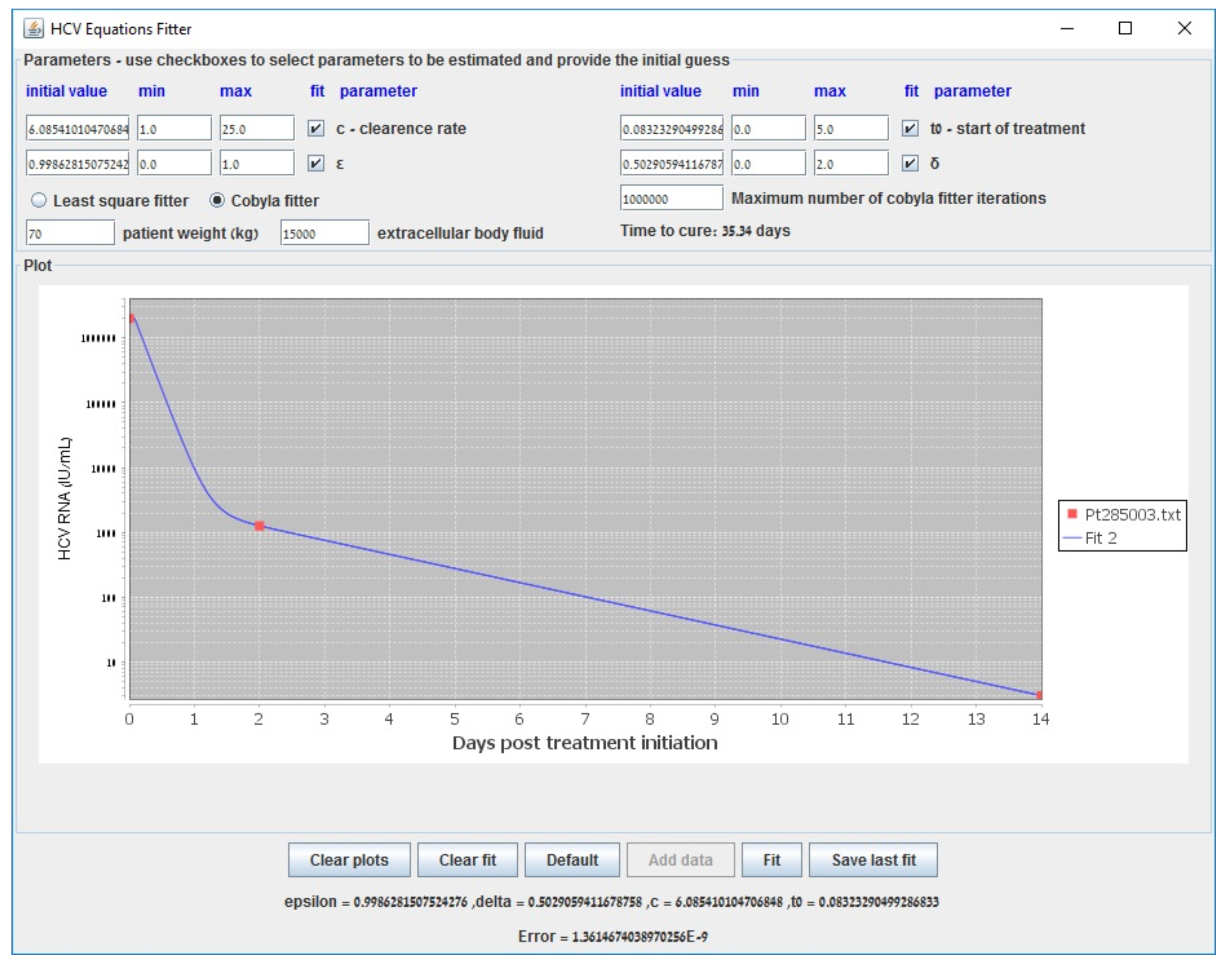
Appendix B. Parameter Estimation in the Biphasic Model
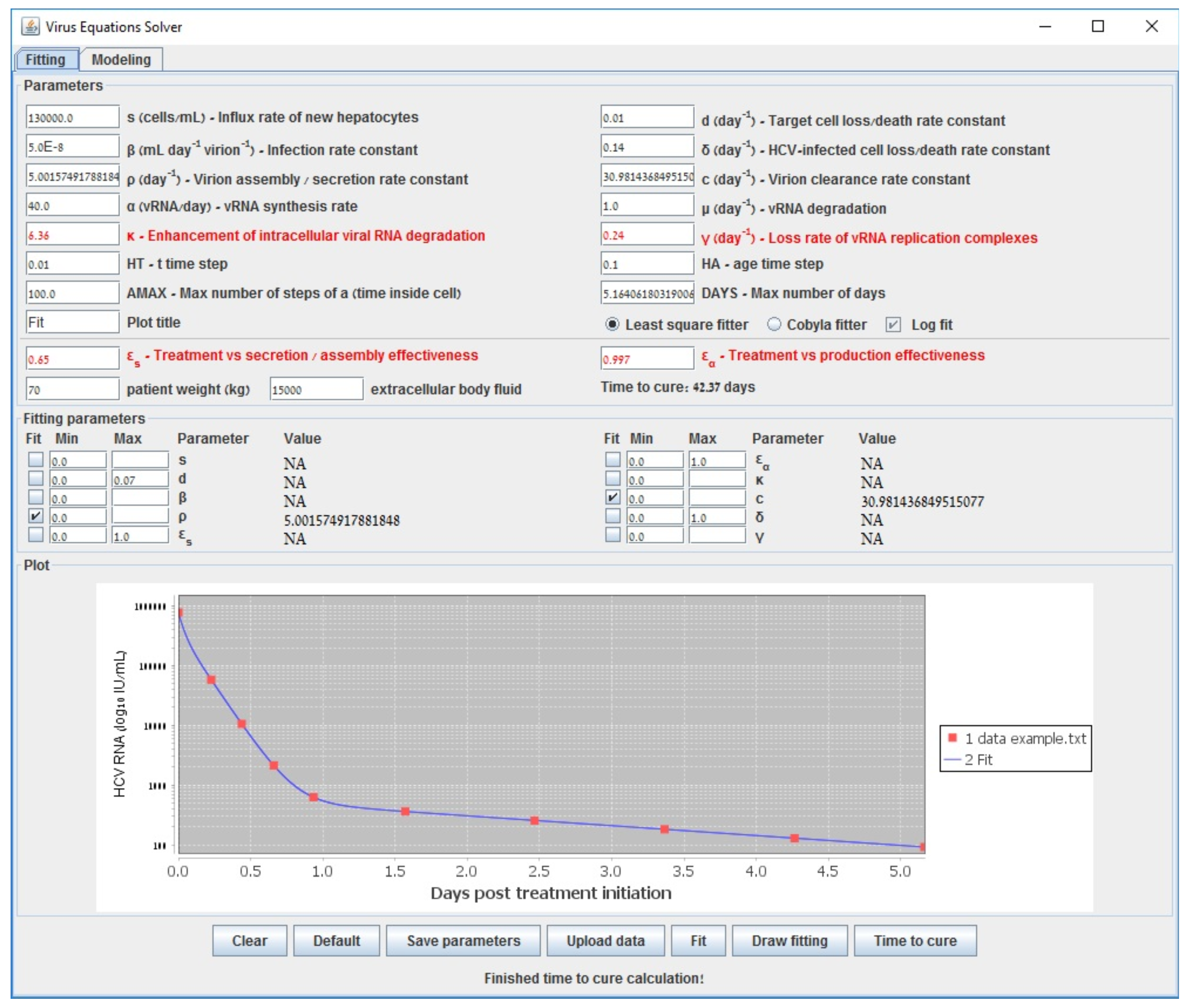
Appendix C. Parameter Estimation in the Multiscale Model

References
- World Health Organization. Global Hepatitis Report 2017: Web Annex A: Estimations of Worldwide Prevalence of Chronic Hepatitis B Virus Infection: A Systematic Review of Data Published between 1965 and 2017; Technical Report; World Health Organization: Geneva, Switzerland, 2018. [Google Scholar]
- Gilman, C.; Heller, T.; Koh, C. Chronic hepatitis delta: A state-of-the-art review and new therapies. World J. Gastroenterol. 2019, 25, 4580. [Google Scholar] [CrossRef] [PubMed]
- Blach, S.; Zeuzem, S.; Manns, M.; Altraif, I.; Duberg, A.S.; Muljono, D.H.; Waked, I.; Alavian, S.M.; Lee, M.H.; Negro, F.; et al. Global prevalence and genotype distribution of hepatitis C virus infection in 2015: A modelling study. Lancet Gastroenterol. Hepatol. 2017, 2, 161–176. [Google Scholar] [CrossRef]
- Stanaway, J.D.; Flaxman, A.D.; Naghavi, M.; Fitzmaurice, C.; Vos, T.; Abubakar, I.; Abu-Raddad, L.J.; Assadi, R.; Bhala, N.; Cowie, B.; et al. The global burden of viral hepatitis from 1990 to 2013: Findings from the Global Burden of Disease Study 2013. Lancet 2016, 388, 1081–1088. [Google Scholar] [CrossRef]
- Foreman, K.J.; Marquez, N.; Dolgert, A.; Fukutaki, K.; Fullman, N.; McGaughey, M.; Pletcher, M.A.; Smith, A.E.; Tang, K.; Yuan, C.W.; et al. Forecasting life expectancy, years of life lost, and all-cause and cause-specific mortality for 250 causes of death: Reference and alternative scenarios for 2016–2040 for 195 countries and territories. Lancet 2018, 392, 2052–2090. [Google Scholar] [CrossRef]
- Ciupe, S.M. Modeling the dynamics of hepatitis B infection, immunity, and drug therapy. Immunol. Rev. 2018, 285, 38–54. [Google Scholar] [CrossRef] [PubMed]
- Means, S.; Ali, M.A.; Ho, H.; Heffernan, J. Mathematical Modeling for Hepatitis B Virus: Would Spatial Effects Play a Role and How to Model It? Front. Physiol. 2020, 11, 146. [Google Scholar] [CrossRef]
- Dahari, H.; Major, M.; Zhang, X.; Mihalik, K.; Rice, C.M.; Perelson, A.S.; Feinstone, S.M.; Neumann, A.U. Mathematical modeling of primary hepatitis C infection: Noncytolytic clearance and early blockage of virion production. Gastroenterology 2005, 128, 1056–1066. [Google Scholar] [CrossRef]
- Dahari, H.; Layden-Almer, J.E.; Kallwitz, E.; Ribeiro, R.M.; Cotler, S.J.; Layden, T.J.; Perelson, A.S. A mathematical model of hepatitis C virus dynamics in patients with high baseline viral loads or advanced liver disease. Gastroenterology 2009, 136, 1402–1409. [Google Scholar] [CrossRef]
- Goyal, A.; Murray, J.M. Dynamics of in vivo hepatitis D virus infection. J. Theor. Biol. 2016, 398, 9–19. [Google Scholar] [CrossRef]
- Goyal, A.; Ribeiro, R.M.; Perelson, A.S. The role of infected cell proliferation in the clearance of acute HBV infection in humans. Viruses 2017, 9, 350. [Google Scholar] [CrossRef]
- Neumann, A.U.; Phillips, S.; Levine, I.; Ijaz, S.; Dahari, H.; Eren, R.; Dagan, S.; Naoumov, N.V. Novel mechanism of antibodies to hepatitis B virus in blocking viral particle release from cells. Hepatology 2010, 52, 875–885. [Google Scholar] [CrossRef] [PubMed]
- Dahari, H.; de Araujo, E.S.A.; Haagmans, B.L.; Layden, T.J.; Cotler, S.J.; Barone, A.A.; Neumann, A.U. Pharmacodynamics of PEG-IFN-α-2a in HIV/HCV co-infected patients: Implications for treatment outcomes. J. Hepatol. 2010, 53, 460–467. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Dahari, H.; Ribeiro, R.M.; Perelson, A.S. Triphasic decline of hepatitis C virus RNA during antiviral therapy. Hepatology 2007, 46, 16–21. [Google Scholar] [CrossRef] [PubMed]
- Dahari, H.; Lo, A.; Ribeiro, R.M.; Perelson, A.S. Modeling hepatitis C virus dynamics: Liver regeneration and critical drug efficacy. J. Theor. Biol. 2007, 247, 371–381. [Google Scholar] [CrossRef] [PubMed]
- Dahari, H.; Shudo, E.; Ribeiro, R.M.; Perelson, A.S. Modeling complex decay profiles of hepatitis B virus during antiviral therapy. Hepatology 2009, 49, 32–38. [Google Scholar] [CrossRef] [PubMed]
- Koh, C.; Dubey, P.; Han, M.A.T.; Walter, P.J.; Garraffo, H.M.; Surana, P.; Southall, N.T.; Borochov, N.; Uprichard, S.L.; Cotler, S.J.; et al. A randomized, proof-of-concept clinical trial on repurposing chlorcyclizine for the treatment of chronic hepatitis C. Antivir. Res. 2019, 163, 149–155. [Google Scholar] [CrossRef]
- Dubey, P.; Koh, C.; Surana, P.; Uprichard, S.L.; Han, M.A.T.; Fryzek, N.; Kapuria, D.; Etzion, O.; Takyar, V.K.; Rotman, Y.; et al. Modeling hepatitis delta virus dynamics during ritonavir boosted lonafarnib treatment-the LOWR HDV-3 study. Hepatology 2017, 66, 21A. [Google Scholar]
- Pawlotsky, J.M.; Dahari, H.; Neumann, A.U.; Hezode, C.; Germanidis, G.; Lonjon, I.; Castera, L.; Dhumeaux, D. Antiviral action of ribavirin in chronic hepatitis C. Gastroenterology 2004, 126, 703–714. [Google Scholar] [CrossRef]
- Neumann, A.U.; Lam, N.P.; Dahari, H.; Davidian, M.; Wiley, T.E.; Mika, B.P.; Perelson, A.S.; Layden, T.J. Differences in viral dynamics between genotypes 1 and 2 of hepatitis C virus. J. Infect. Dis. 2000, 182, 28–35. [Google Scholar] [CrossRef]
- Dahari, H.; Shudo, E.; Ribeiro, R.M.; Perelson, A.S. Mathematical modeling of HCV infection and treatment. Methods Mol. Biol. 2009, 510, 439–453. [Google Scholar]
- DebRoy, S.; Hiraga, N.; Imamura, M.; Hayes, C.N.; Akamatsu, S.; Canini, L.; Perelson, A.S.; Pohl, R.T.; Persiani, S.; Uprichard, S.L.; et al. hepatitis C virus dynamics and cellular gene expression in uPA-SCID chimeric mice with humanized livers during intravenous silibinin monotherapy. J. Viral Hepat. 2016, 23, 708–717. [Google Scholar] [CrossRef] [PubMed]
- Canini, L.; DebRoy, S.; Mariño, Z.; Conway, J.M.; Crespo, G.; Navasa, M.; D’Amato, M.; Ferenci, P.; Cotler, S.J.; Forns, X.; et al. Severity of liver disease affects HCV kinetics in patients treated with intravenous silibinin monotherapy. Antivir. Ther. 2015, 20, 149. [Google Scholar] [CrossRef] [PubMed]
- Goyal, A.; Lurie, Y.; Meissner, E.G.; Major, M.; Sansone, N.; Uprichard, S.L.; Cotler, S.J.; Dahari, H. Modeling HCV cure after an ultra-short duration of therapy with direct acting agents. Antivir. Res. 2017, 144, 281–285. [Google Scholar] [CrossRef] [PubMed]
- Dahari, H.; Shteingart, S.; Gafanovich, I.; Cotler, S.J.; D’Amato, M.; Pohl, R.T.; Weiss, G.; Ashkenazi, Y.J.; Tichler, T.; Goldin, E.; et al. Sustained virological response with intravenous silibinin: Individualized IFN-free therapy via real-time modelling of HCV kinetics. Liver Int. 2015, 35, 289–294. [Google Scholar] [CrossRef]
- Neumann, A.U.; Lam, N.P.; Dahari, H.; Gretch, D.R.; Wiley, T.E.; Layden, T.J.; Perelson, A.S. Hepatitis C viral dynamics in vivo and the antiviral efficacy of interferon-α therapy. Science 1998, 282, 103–107. [Google Scholar] [CrossRef] [PubMed]
- Guedj, J.; Dahari, H.; Pohl, R.T.; Ferenci, P.; Perelson, A.S. Understanding silibinin’s modes of action against HCV using viral kinetic modeling. J. Hepatol. 2012, 56, 1019–1024. [Google Scholar] [CrossRef]
- Lewin, S.R.; Ribeiro, R.M.; Walters, T.; Lau, G.K.; Bowden, S.; Locarnini, S.; Perelson, A.S. Analysis of hepatitis B viral load decline under potent therapy: Complex decay profiles observed. Hepatology 2001, 34, 1012–1020. [Google Scholar] [CrossRef]
- Ribeiro, R.M.; Germanidis, G.; Powers, K.A.; Pellegrin, B.; Nikolaidis, P.; Perelson, A.S.; Pawlotsky, J.M. hepatitis B virus kinetics under antiviral therapy sheds light on differences in hepatitis B e antigen positive and negative infections. J. Infect. Dis. 2010, 202, 1309–1318. [Google Scholar] [CrossRef]
- Nowak, M.A.; Bonhoeffer, S.; Hill, A.M.; Boehme, R.; Thomas, H.C.; McDade, H. Viral dynamics in hepatitis B virus infection. Proc. Natl. Acad. Sci. USA 1996, 93, 4398–4402. [Google Scholar] [CrossRef]
- Tsiang, M.; Rooney, J.F.; Toole, J.J.; Gibbs, C.S. Biphasic clearance kinetics of hepatitis B virus from patients during adefovir dipivoxil therapy. Hepatology 1999, 29, 1863–1869. [Google Scholar] [CrossRef]
- Canini, L.; Koh, C.; Cotler, S.J.; Uprichard, S.L.; Winters, M.A.; Han, M.A.T.; Kleiner, D.E.; Idilman, R.; Yurdaydin, C.; Glenn, J.S.; et al. Pharmacokinetics and pharmacodynamics modeling of lonafarnib in patients with chronic hepatitis delta virus infection. Hepatol. Commun. 2017, 1, 288–292. [Google Scholar] [CrossRef] [PubMed]
- Koh, C.; Canini, L.; Dahari, H.; Zhao, X.; Uprichard, S.L.; Haynes-Williams, V.; Winters, M.A.; Subramanya, G.; Cooper, S.L.; Pinto, P.; et al. Oral prenylation inhibition with lonafarnib in chronic hepatitis D infection: A proof-of-concept randomised, double-blind, placebo-controlled phase 2A trial. Lancet Infect. Dis. 2015, 15, 1167–1174. [Google Scholar] [CrossRef]
- Guedj, J.; Rotman, Y.; Cotler, S.J.; Koh, C.; Schmid, P.; Albrecht, J.; Haynes-Williams, V.; Liang, T.J.; Hoofnagle, J.H.; Heller, T.; et al. Understanding early serum hepatitis D virus and hepatitis B surface antigen kinetics during pegylated interferon-alpha therapy via mathematical modeling. Hepatology 2014, 60, 1902–1910. [Google Scholar] [CrossRef] [PubMed]
- Shekhtman, L.; Cotler, S.J.; Hershkovich, L.; Uprichard, S.L.; Bazinet, M.; Pantea, V.; Cebotarescu, V.; Cojuhari, L.; Jimbei, P.; Krawczyk, A.; et al. Modelling hepatitis D virus RNA and HBsAg dynamics during nucleic acid polymer monotherapy suggest rapid turnover of HBsAg. Sci. Rep. 2020, 10, 1–7. [Google Scholar] [CrossRef]
- Etzion, O.; Dahari, H.; Yardeni, D.; Issachar, A.; Nevo-Shor, A.; Naftaly-Cohen, M.; Uprichard, S.L.; Arbib, O.S.; Munteanu, D.; Braun, M.; et al. Response-Guided Therapy with Direct-Acting Antivirals Shortens Treatment Duration in 50% of HCV Treated Patients. Hepatology 2018, 68, 1469A–1470A. [Google Scholar]
- Dahari, H.; Canini, L.; Graw, F.; Uprichard, S.L.; Araújo, E.S.; Penaranda, G.; Coquet, E.; Chiche, L.; Riso, A.; Renou, C.; et al. HCV kinetic and modeling analyses indicate similar time to cure among sofosbuvir combination regimens with daclatasvir, simeprevir or ledipasvir. J. Hepatol. 2016, 64, 1232–1239. [Google Scholar] [CrossRef]
- Canini, L.; Imamura, M.; Kawakami, Y.; Uprichard, S.L.; Cotler, S.J.; Dahari, H.; Chayama, K. HCV kinetic and modeling analyses project shorter durations to cure under combined therapy with daclatasvir and asunaprevir in chronic HCV-infected patients. PLoS ONE 2017, 12, e0187409. [Google Scholar] [CrossRef]
- Gambato, M.; Canini, L.; Lens, S.; Graw, F.; Perpiñan, E.; Londoño, M.C.; Uprichard, S.L.; Mariño, Z.; Reverter, E.; Bartres, C.; et al. Early HCV viral kinetics under DAAs may optimize duration of therapy in patients with compensated cirrhosis. Liver Int. 2019, 39, 826–834. [Google Scholar] [CrossRef]
- Sandmann, L.; Manns, M.P.; Maasoumy, B. Utility of viral kinetics in HCV therapy—It is not over until it is over? Liver Int. 2019, 39, 815–817. [Google Scholar] [CrossRef]
- Deng, B.; Lou, S.; Dubey, P.; Etzion, O.; Chayam, K.; Uprichard, S.; Sulkowski, M.; Cotler, S.; Dahari, H. Modeling time to cure after short-duration treatment for chronic HCV with daclatasvir, asunaprevir, beclabuvir and sofosbuvir: The FOURward study. J. Viral Hepat. 2018, 25, 58. [Google Scholar]
- Dahari, H.; Sainz, B.; Perelson, A.S.; Uprichard, S.L. Modeling subgenomic hepatitis C virus RNA kinetics during treatment with alpha interferon. J. Virol. 2009, 83, 6383–6390. [Google Scholar] [CrossRef] [PubMed]
- Dahari, H.; Ribeiro, R.M.; Rice, C.M.; Perelson, A.S. Mathematical modeling of subgenomic hepatitis C virus replication in Huh-7 cells. J. Virol. 2007, 81, 750–760. [Google Scholar] [CrossRef] [PubMed]
- Murray, J.M.; Goyal, A. In silico single cell dynamics of hepatitis B virus infection and clearance. J. Theor. Biol. 2015, 366, 91–102. [Google Scholar] [CrossRef] [PubMed]
- Murray, J.M.; Wieland, S.F.; Purcell, R.H.; Chisari, F.V. Dynamics of hepatitis B virus clearance in chimpanzees. Proc. Natl. Acad. Sci. USA 2005, 102, 17780–17785. [Google Scholar] [CrossRef]
- Packer, A.; Forde, J.; Hews, S.; Kuang, Y. Mathematical models of the interrelated dynamics of hepatitis D and B. Math. Biosci. 2014, 247, 38–46. [Google Scholar] [CrossRef]
- Guedj, J.; Dahari, H.; Rong, L.; Sansone, N.D.; Nettles, R.E.; Cotler, S.J.; Layden, T.J.; Uprichard, S.L.; Perelson, A.S. Modeling shows that the NS5A inhibitor daclatasvir has two modes of action and yields a shorter estimate of the hepatitis C virus half-life. Proc. Natl. Acad. Sci. USA 2013, 110, 3991–3996. [Google Scholar] [CrossRef]
- Rong, L.; Guedj, J.; Dahari, H.; Coffield, D.J.J.; Levi, M.; Smith, P.; Perelson, A.S. Analysis of hepatitis C virus decline during treatment with the protease inhibitor danoprevir using a multiscale model. PLoS Comput. Biol. 2013, 9, e1002959. [Google Scholar] [CrossRef]
- Rong, L.; Perelson, A.S. Mathematical analysis of multiscale models for hepatitis C virus dynamics under therapy with direct-acting antiviral agents. Math. Biosci. 2013, 245, 22–30. [Google Scholar] [CrossRef]
- Quintela, B.M.; Conway, J.M.; Hyman, J.M.; Guedj, J.; dos Santos, R.W.; Lobosco, M.; Perelson, A.S. A New Age-Structured Multiscale Model of the Hepatitis C Virus Life-Cycle During Infection and Therapy with Direct-Acting Antiviral Agents. Front. Microbiol. 2018, 9, 601. [Google Scholar] [CrossRef]
- Guedj, J.; Neumann, A.U. Understanding hepatitis C viral dynamics with direct-acting antiviral agents due to the interplay between intracellular replication and cellular infection dynamics. J. Theor. Biol. 2010, 267, 330–340. [Google Scholar] [CrossRef]
- Weickert, J.; ter Haar Romeny, B.; Viergever, M. Efficient and reliable schemes for nonlinear diffusion filtering. IEEE Trans. Imag. Proc. 1998, 7, 398–410. [Google Scholar] [CrossRef]
- Barash, D.; Israeli, M.; Kimmel, R. An Accurate Operator Splitting Scheme for Nonlinear Diffusion Filtering. In Proceedings of the 3rd International Conference on ScaleSpace and Morphology, Vancouver, BC, Canada, 7–8 July 2001; pp. 281–289. [Google Scholar]
- Barash, D. Nonlinear Diffusion Filtering on Extended Neighborhood. Appl. Num. Math. 2005, 52, 1–11. [Google Scholar] [CrossRef]
- Reinharz, V.; Churkin, A.; Dahari, H.; Barash, D. A Robust and Efficient Numerical Method for RNA-mediated Viral Dynamics. Front. Appl. Math. Stat. 2017, 3, 20. [Google Scholar] [CrossRef] [PubMed]
- Reinharz, V.; Dahari, H.; Barash, D. Numerical schemes for solving and optimizing multiscale models with age of hepatitis C virus dynamics. Math. Biosci. 2018, 300, 1–13. [Google Scholar] [CrossRef]
- Reinharz, V.; Churkin, A.; Lewkiewicz, S.; Dahari, H.; Barash, D. A Parameter Estimation Method for Multiscale Models of hepatitis C Virus Dynamics. Bull. Math. Biol. 2019, 81, 3675–3721. [Google Scholar] [CrossRef] [PubMed]
- Levenberg, K. A method for the solution of certain nonlinear problems in least squares. Q. Appl. Math. 1944, 2, 164–168. [Google Scholar] [CrossRef]
- Marquardt, D.W. An algorithm for least-squares estimation of nonlinear parameters. J. Soc. Ind. Appl. Math. 1963, 11, 431–441. [Google Scholar] [CrossRef]
- Kitagawa, K.; Nakaoka, S.; Asai, Y.; Watashi, K.; Iwami, S. A PDE Multiscale Model of hepatitis C virus infection can be transformed to a system of ODEs. J. Theor. Biol. 2018, 448, 80–85. [Google Scholar] [CrossRef]
- Powell, M.J.D. A View of Algorithms for Optimization Without Derivatives. Math. Today 2007, 43, 170–174. [Google Scholar]
- Rohatgi, A. WebPlotDigitizer: Web Based Tool to Extract Data from Plots, Images, and Maps. V 4.1. 2018. Available online: https://automeris.io/WebPlotDigitizer (accessed on 22 August 2020).
- Quarteroni, A.; Valli, A. Springer Series in Computational Mathematics, 1994; Springer: Berlin/Heidelberg, Germany, 1994. [Google Scholar]
- Knabner, P.; Angermann, L. Numerical Methods for Elliptic and Parabolic Partial Differential Equations: An Applications-Oriented Introduction; Springer: Berlin/Heidelberg, Germany, 2004. [Google Scholar]
- Powell, M.J.D. A Direct Search Optimization Method That Models Objective and Constraint Functions by Linear Interpolation. In Advances in Optimization and Numerical Analysis. Mathematics and its Applications; Gomez, S., Hennart, J., Eds.; Springer: Dordrecht, The Netherlands, 1994; Volume 275. [Google Scholar]
- Bastian, P.; Heimann, F.; Marnach, S. Generic implementation of finite element methods in the distributed and unified numerics environment (DUNE). Kybernetika 2010, 46, 294–315. [Google Scholar]
- Flemisch, B.; Darcis, M.; Erbertseder, K.; Faigle, B.; Lauser, A.; Mosthaf, K.; Müthing, S.; Nuske, P.; Tatomir, A.; Wolff, M.; et al. DuMux: DUNE for multi-{phase, component, scale, physics,…} flow and transport in porous media. Adv. Water Resour. 2011, 34, 1102–1112. [Google Scholar] [CrossRef]
- Vogel, A.; Reiter, S.; Rupp, M.; Nägel, A.; Wittum, G. UG 4: A novel flexible software system for simulating PDE based models on high performance computers. Comput. Vis. Sci. 2013, 16, 165–179. [Google Scholar] [CrossRef]
- Spendley, W.; Hext, G.R.; Himsworth, F.R. Sequential Application of Simplex Designs in Optimisation and Evolutionary Operation. Technometrics 1962, 4, 441–461. [Google Scholar] [CrossRef]
- Nelder, J.; Mead, R. A Simplex Method for Function Minimization. Comput. J. 1965, 7, 308–313. [Google Scholar] [CrossRef]
- Dasgupta, S.; Imamura, M.; Gorstein, E.; Nakahara, T.; Tsuge, M.; Churkin, A.; Yardeni, D.; Etzion, O.; Uprichard, S.L.; Barash, D.; et al. Modeling-Based Response-Guided Glecaprevir-Pibrentasvir Therapy for Chronic Hepatitis C to Identify Patients for Ultrashort Treatment Duration. J. Infect. Dis. 2020, jiaa219. [Google Scholar] [CrossRef] [PubMed]
- Gorstein, E.; Martinello, M.; Churkin, A.; Dasgupta, S.; Walsh, K.; Applegate, T.; Yardeni, D.; Etzion, O.; Uprichard, S.L.; Barash, D.; et al. Modeling based response guided therapy in subjects with recent hepatitis C infection. Antivir. Res. 2020. [Google Scholar] [CrossRef]
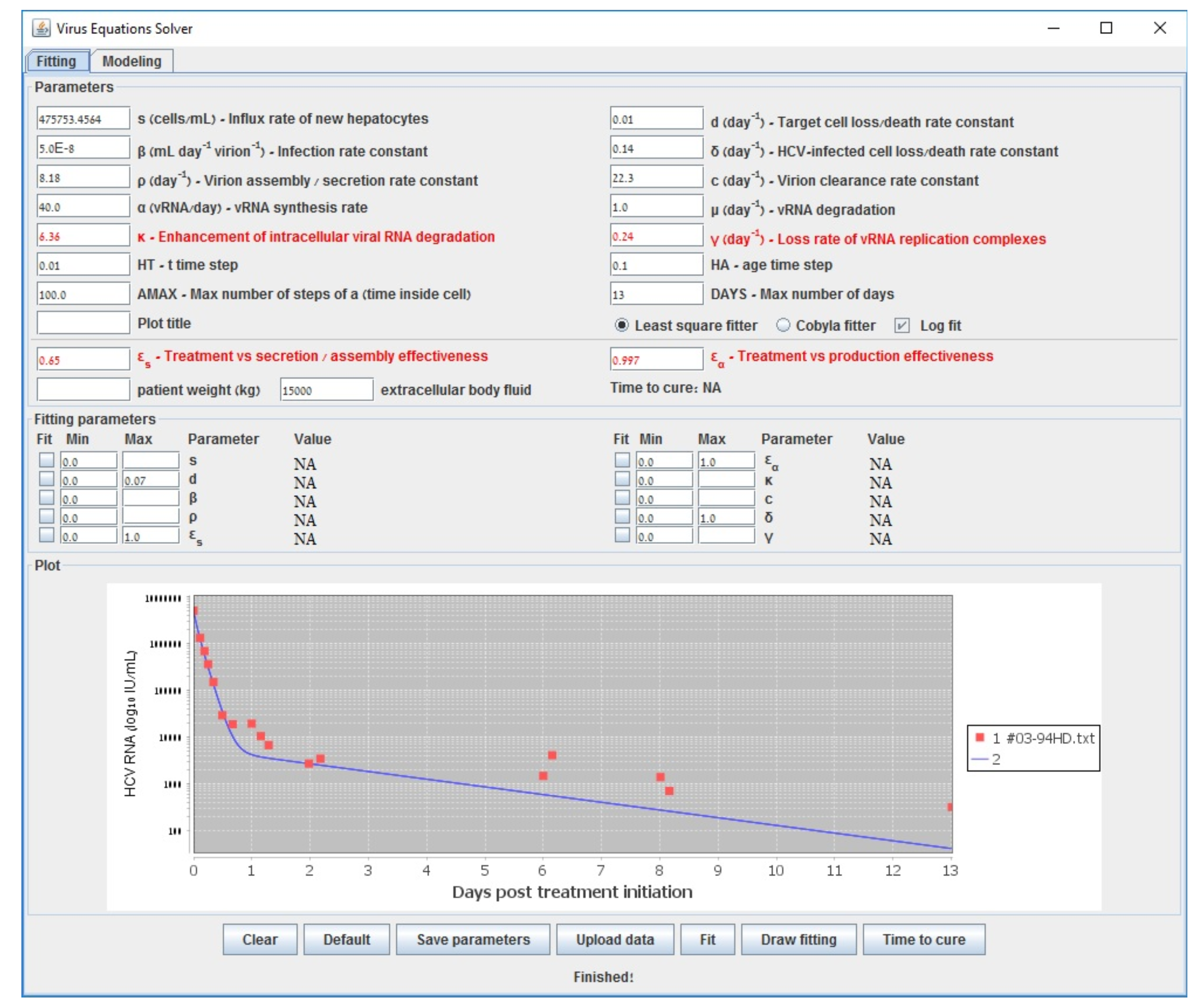
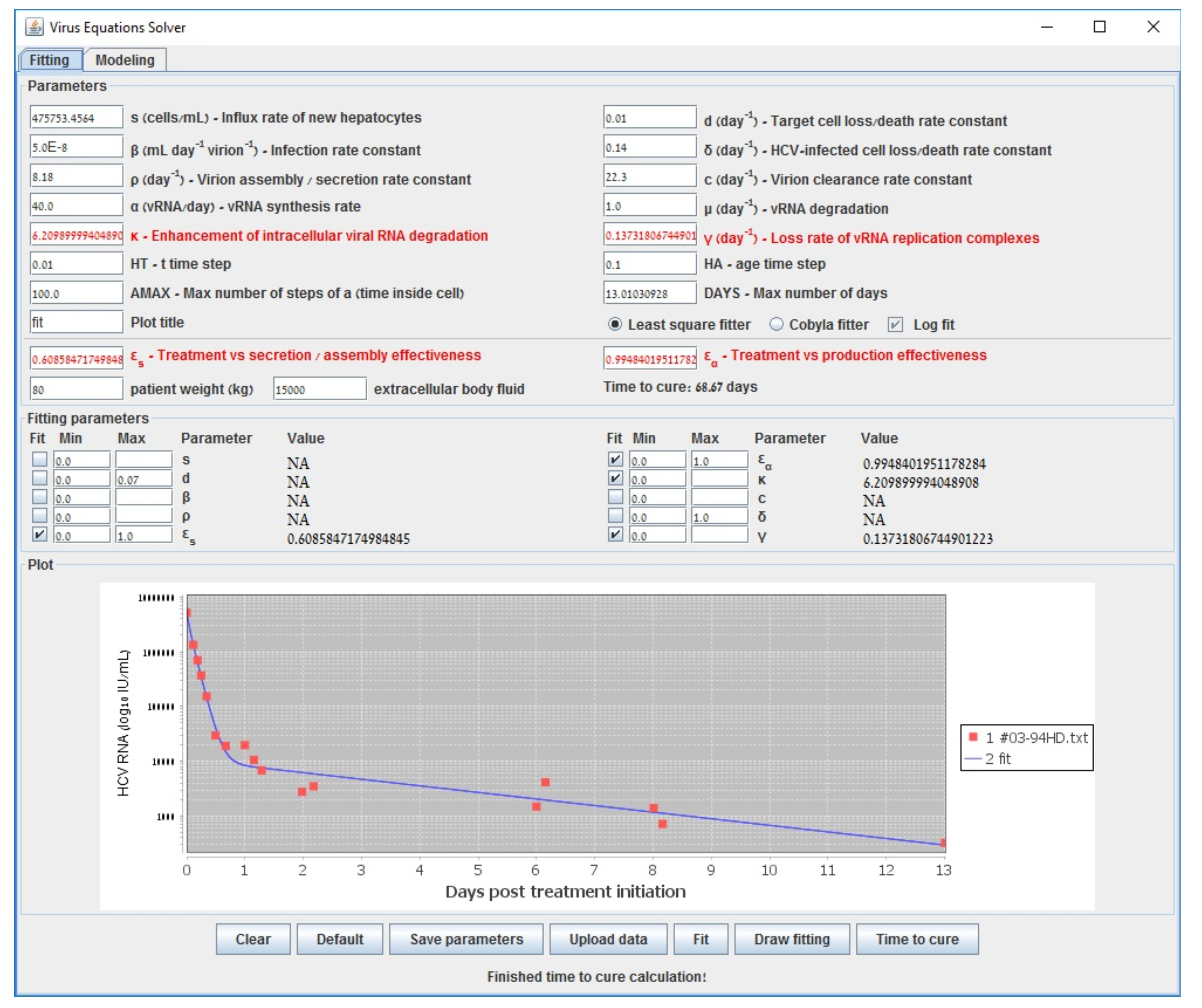
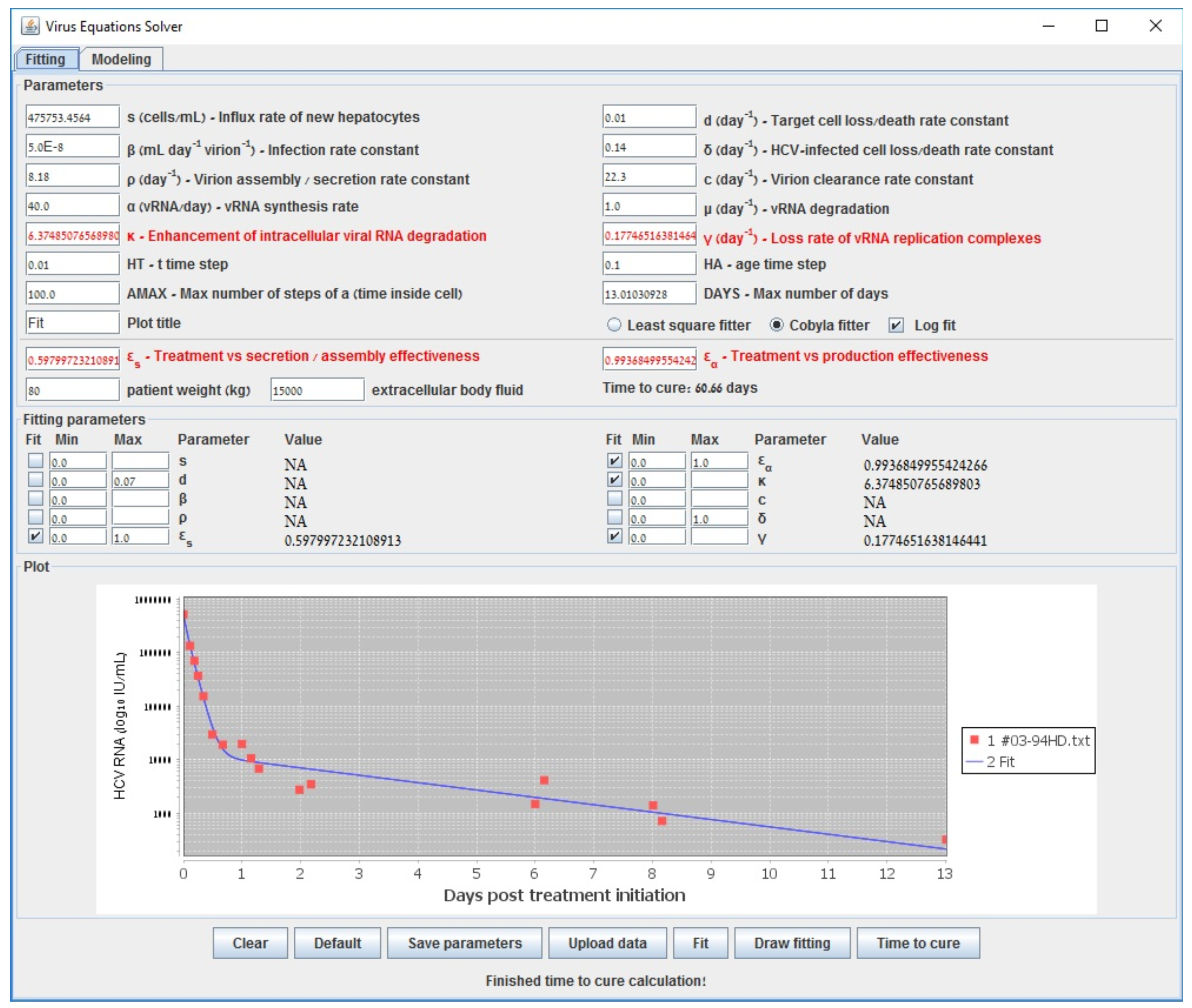
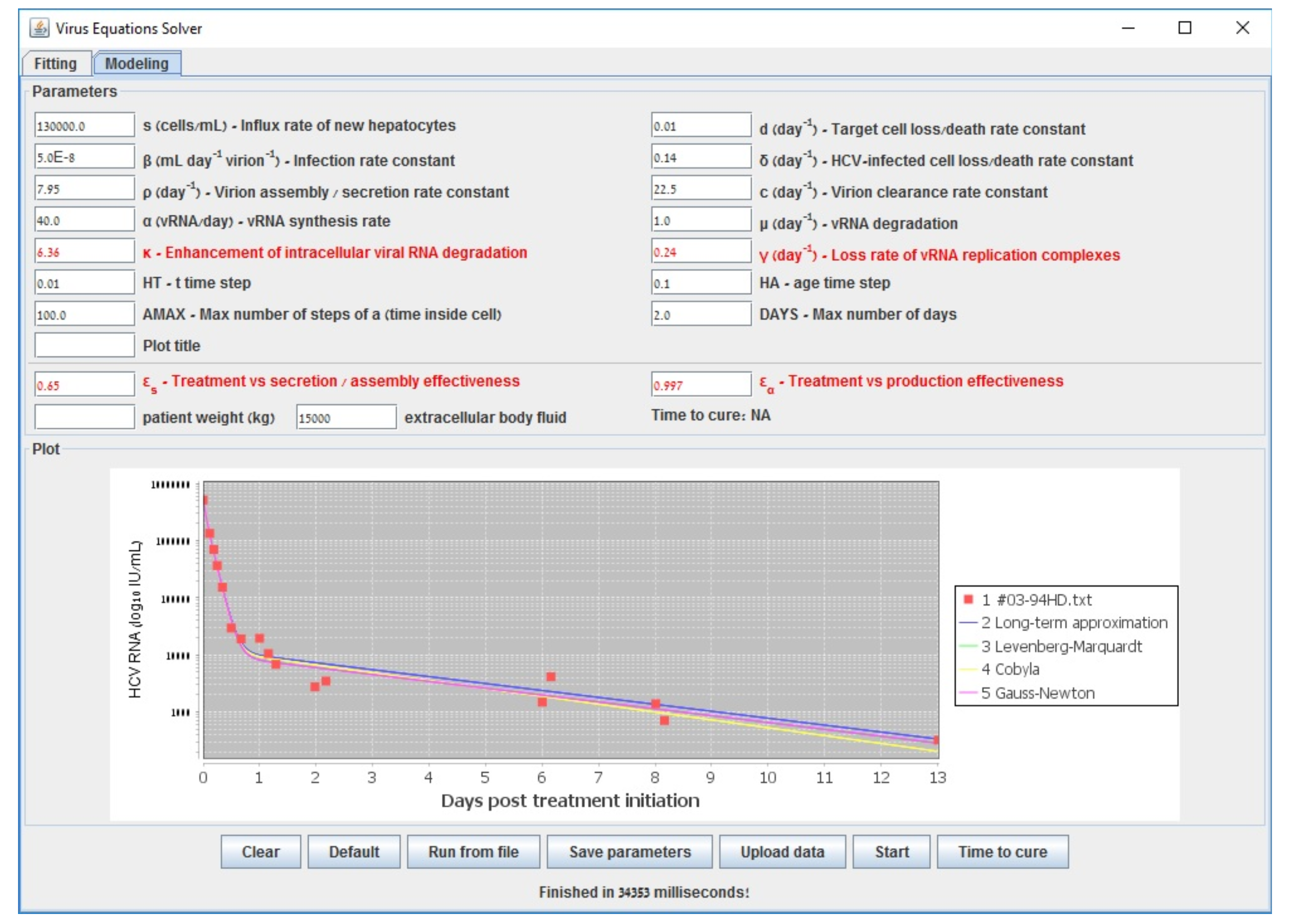
| s () | Influx rate of new hepatocytes |
| d () | Target cell loss/death rate constant |
| () | Infection rate constant |
| () | HCV-infected cell loss/death rate constant |
| () | Virion assembly/secretion rate constant |
| c () | Virion clearance rate constant |
| (vRNA) | vRNA synthesis rate |
| () | vRNA degradation |
| Enhancement of intracellular viral RNA degradation | |
| () | Loss rate of vRNA replication complexes |
| Treatment vs. secretion/assembly effectiveness | |
| Treatment vs. production effectiveness |
| 40 d−1 | d−1 | ||
| c | d−1 | d−1 | |
| 1 d−1 | d | d−1 | |
| d−1 | s | / |
| Gauss–Newton (LSF) | COBYLA | Levenberg–Marquardt | Long-Term | |
|---|---|---|---|---|
| 0.609 | 0.598 | 0.602 | 0.6000 | |
| 0.995 | 0.994 | 0.995 | 0.994 | |
| 6.210 | 6.375 | 6.219 | 6.160 | |
| (d−1) | 0.137 | 0.177 | 0.139 | 0.140 |
| accuracy (sum error) | 0.538 | 0.582 | 0.538 | 0.587 |
| run-time (s) | 194 | 3698 | 70118 | <1 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Churkin, A.; Lewkiewicz, S.; Reinharz, V.; Dahari, H.; Barash, D. Efficient Methods for Parameter Estimation of Ordinary and Partial Differential Equation Models of Viral Hepatitis Kinetics. Mathematics 2020, 8, 1483. https://doi.org/10.3390/math8091483
Churkin A, Lewkiewicz S, Reinharz V, Dahari H, Barash D. Efficient Methods for Parameter Estimation of Ordinary and Partial Differential Equation Models of Viral Hepatitis Kinetics. Mathematics. 2020; 8(9):1483. https://doi.org/10.3390/math8091483
Chicago/Turabian StyleChurkin, Alexander, Stephanie Lewkiewicz, Vladimir Reinharz, Harel Dahari, and Danny Barash. 2020. "Efficient Methods for Parameter Estimation of Ordinary and Partial Differential Equation Models of Viral Hepatitis Kinetics" Mathematics 8, no. 9: 1483. https://doi.org/10.3390/math8091483
APA StyleChurkin, A., Lewkiewicz, S., Reinharz, V., Dahari, H., & Barash, D. (2020). Efficient Methods for Parameter Estimation of Ordinary and Partial Differential Equation Models of Viral Hepatitis Kinetics. Mathematics, 8(9), 1483. https://doi.org/10.3390/math8091483






