Abstract
Colorectal cancer (CRC) is the second most common global malignancy with high mortality, and timely early polyp detection is critical to halt its progression. Yet, polyp image segmentation—an essential tool—faces challenges: blurred edges, small sizes, and artifacts from intestinal folds, bubbles, and mucus. To address these, we proposed a novel segmentation model with multi-scale feature extraction. Its encoder uses Multiscale Attention-based Pyramid Vision Transformer v2 (PVTv2) for hierarchical features (lower-stage modules expand receptive field), while the decoder adopts a Parallel Multi-level Aggregation structure, plus multi-branch and improved reverse attention modules. Ablation experiments validated key modules. Compared to nine state-of-the-art networks across five benchmarks, the model showed superiority: optimal mDice/mIoU on polyp datasets, 0.2% higher mDice than MEGANet on Kvasir-SEG, and outperformance over UHA-Net and CSCA-U-Net on CVC-ClinicDB.
Keywords:
polyp segmentation; pyramid vision transformer; multi-scale feature; colonoscopy; computer vision MSC:
92C55; 68U10; 68T45
1. Introduction
Colorectal cancer (CRC) is one of the leading causes of cancer-related death and is the second most common cancer worldwide, with a persistently high mortality rate [,]. Early detection and removal of polyps are critical measures to prevent the progression of colorectal cancer []. Colonoscopy is the gold standard for the detection of colorectal lesions [], and accurate localization of early-stage polyps is crucial for the clinical prevention of CRC.
However, this manual detection paradigm is prone to significant missed diagnoses—attributed to factors such as operator fatigue, subtle polyp morphology, and interference from intestinal physiological structures. To address this clinical bottleneck, integrating AI-driven deep learning and image segmentation techniques bridges subjective manual detection with objective computational analysis. This translates the clinical need for precise early polyp localization into a technical task of medical image segmentation, enabling quantitative polyp analysis to enhance diagnostic efficiency and accuracy [], while mitigating manual limitations through synergistic integration with traditional inspection. Image segmentation represents a fundamental yet challenging research area within the domain of computer science []. These challenges are visually exemplified in Figure 1:
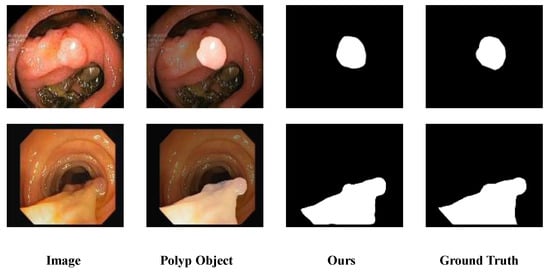
Figure 1.
Polyp segmentation challenges in colonoscopy. In images presented by colonoscopy, the differences between polyp and non-polyp regions are usually not obvious. Meanwhile, factors such as air bubbles, mucus and intestinal folds further increase the difficulty of image analysis. The highlighted area in the Polyp Object indicates the polyp region.
As demonstrated in the figure, existing polyp segmentation methods still face substantial hurdles due to the diverse shapes and sizes of colon polyps. For instance, some polyps have blurred boundaries that are difficult to distinguish from normal tissues [], while smaller polyp targets are easily overlooked or misidentified as intestinal folds—issues that also plague manual clinical inspection. Current methods are continuously addressing these challenges, exploring solutions, and achieving steady progress.
Here, we propose a novel approach, using a polyp segmentation model leveraging multi-scale feature extraction. It employs Multiscale Attention-based Pyramid Vision Transformer v2 (PVTv2) [] as the encoder and a Parallel Multi-level Aggregation Decoder composed of attention and multi-scale parallel convolutions, incorporating an improved Scale Aggregation Reverse Attention. This strategy enhances learning and generalization capabilities, overcoming the weaknesses of existing models in challenging environments and better addressing the issues faced by current polyp segmentation models.
- (1)
- We introduce a Multiscale Attention (MA) module at the encoder’s lower stages. This module utilizes multiscale attention mechanisms and dilated convolutions with multiple sizes to expand the receptive field, improving performance in low-layers feature map extraction.
- (2)
- The proposed method introduces an integrated decoder that incorporates an Attention Gate (AG) module and multi-branch convolution modules to enhance segmentation accuracy for polyps of varying sizes and shapes, leveraging hierarchical feature supplementation and refinement.
- (3)
- The proposed method integrates an Improved Reverse Attention module, which specializes in polyp boundary segmentation tasks via enhanced feature extraction and reverse attention mechanisms.
- (4)
- To verify the practical effectiveness of the proposed model, experiments were conducted on five challenging benchmark datasets. The results demonstrate that the model exhibits reliable performance and strong versatility as a segmentation tool.
To address the key challenges prevalent in existing polyp segmentation, such as unclear boundary segmentation and inadequate detailed feature extraction, this paper adopts a systematic research framework and arranges the specific structure with core contents as follows: Section 2 systematically reviews existing deep learning architectures and combs through previous research achievements in the field of polyp segmentation, clarifying the current technical pain points in boundary and detail extraction, and providing targeted directions for subsequent model design; Section 3 deeply elaborates on the design of the three core components of the proposed model, including the Multiscale Attention (MA) module deployed in the lower stages of the encoder to enhance feature extraction, the integrated decoder that improves the segmentation accuracy of diverse polyps, and the Improved Reverse Attention module focusing on boundary segmentation; Section 4 details the experimental procedures and results, comparing our model with the current state-of-the-art (SOTA) models to evaluate its performance. It also includes ablation experiments to assess the contribution of each core component. Finally, Section 5 discusses the effectiveness of the model and summarizes the contributions made in this paper.
2. Related Work
In recent years, deep neural networks have been widely and deeply used in the domain of image processing due to their excellent performance [,]. Especially in medical image segmentation, the convolutional neural network (CNN) framework forms the core cornerstone of most advanced models. Long et al. [] took the lead in making a breakthrough by replacing all linear layers with convolutional layers to design a fully convolutional network (FCN). Later, Ronneberger et al. [] constructed a symmetric structure and proposed a U-shaped architecture (U-Net) with shrinking paths and expanding paths. Because of its simple structure, few parameters, and rapid convergence speed, U-Net performs well as a benchmark model in this field. Zhou et al. [] put forward a modified variant of U-Net, known as U-Net++, which incorporates dense skip connections linking the encoder and decoder components. Oktay et al. [] added attention mechanisms to enhance spatial perception and developed Attention U-Net. In addition, Jin et al. [] cleverly combined residual learning and attention mechanisms to propose that residual attention U-Net (RA-U-Net) has achieved significant results in multiple organ classification tasks.
Transformer [], as a large semantic network model specifically designed to capture global semantic information in language, has garnered widespread academic attention due to its exceptional global semantic extraction capabilities and spatially adaptive information aggregation properties. Dosovitskiy et al. [] introduced Transformer to the field of computer vision by incorporating positional encoding and patch embedding techniques, pioneering the Vision Transformer (ViT). Chen et al. [] proposed a novel network structure combining Transformer with U-Net—the TransUNet model—which integrates Transformer into the encoder stage and upsamples encoder features in the decoder before combining them with CNN outputs. Building on this, Wang et al. [] further integrated the pyramid structure into the Transformer architecture, thereby proposing the Pyramid Visual Transformer (PVT), which offers new approaches for the multi-scale representation of image features. Base on PVT, optimized models like SS-Former [] and FCBFormer [] have shown remarkable performance in related tasks, further promoting the application of Transformer in the field of image segmentation.
However, Transformer-based models come with their own challenges. They possess a vast number of parameters and demand substantial computational resources, particularly exhibiting relatively insufficient capability in capturing local features when processing small datasets like medical images. The multi-scale pyramid structure design of CNNs (Convolutional Neural Networks) significantly enhances the model’s ability to handle multi-scale problems, improving performance across multiple dimensions such as multi-scale representation, local feature extraction, feature reuse, robustness, and interpretability. Wang et al. [] ingeniously integrated the pyramid structure from CNN architectures with the original advantages of ViT (Vision Transformer), aiming to optimize ViT ’s efficiency in processing high-resolution images, thereby giving rise to the Pyramid Vision Transformer (PVT) model. Unlike strategies such as VGGNet [] and ResNet [], which obtain multi-scale feature maps through varying convolutional strides, PVT adopts a progressive shrinking approach. It flexibly adjusts the scale of feature maps through patch embedding layers and encoders with built-in spatial reduction mechanisms, preserving the traditional CNN’s ability to generate multi-scale feature maps while effectively reducing computational overhead. Subsequently, in PVTv2 [], Wang et al. improved PVT’s performance by incorporating linear-complexity attention layers, overlapping patch embedding, and convolutional feedforward networks. UNeXt [] and Uformer [] drew inspiration from the UNet architecture from different perspectives to optimize Transformer models for computer vision tasks. CSWin Transformer [] employs a multi-layer Transformer structure and introduces residual-like connections between modules. HRFormer [] integrates Transformer modules based on the multi-resolution feature fusion architecture of HRNet []. Swin UNETR [] uses only Swin Transformer [] as the encoder and CNN as the decoder, where local and global features are alternately extracted, leading to potential confusion and incorrect feature acquisition.
Several methods in polyp segmentation showcase varied strengths and constraints, affecting their segmentation efficacy. PraNet [] Initially applies high-level features to generate a coarse prediction of polyp regions, then employs a Reverse attention module (RA) to progressively refine the edge details of polyps from the background. Inspired by PraNet, UACANet [] designs a novel parallel axial attention and introduces uncertain regions to the boundary information. CaraNet [] adopts an Axial reverse attention (A-RA) module to tackle segmentation issues involving diminutive polyp entities. This module effectively filters out disturbances and sharpens boundaries, although it lacks the capability to supplement and explore additional features. ADS_UNet [] integrates AdaBoost and a heuristic stepwise training strategy into the iterative construction of a collective predictive model, thereby achieving higher accuracy than other UNet-like architectures. M2SNet [] leverages pyramid aggregation based on its proposed intra-layer multi-scale subtraction unit, enabling cross-layer complementarity between low-layer and high-level features. UHA-Net [] introduces a hierarchical feature fusion module and an uncertainty-induced cross-level fusion module for feature aggregation, ultimately combining multi-scale features. EFA-Net [] improves the accuracy of polyp segmentation through the exploitation of inter-layer features for enhanced edge recognition. Texture traits also have a key role in telling polyp foregrounds apart from backgrounds. From the above polyp segmentation models, it is evident that feature extraction for polyp regions and boundary issues have consistently been focal points.
However, most of the SOTA polyp segmentation methods have difficulties in representing multi-scale features and are unable to draw the feature boundaries of polyps under problematic imaging conditions: U-Net variants (U-Net++, Attention U-Net) lack adaptive multi-scale fusion, leading to blurred boundary segmentation; Transformer-based models (TransUNet, PVT) suffer from high computational costs and weak local feature capture on medical small datasets; reverse attention methods (PraNet, CaraNet) fail to supplement discriminative features, resulting in incomplete small polyp segmentation.
The proposed model achieves superiority via targeted innovations: it adopts a PVTv2 encoder with Multiscale Attention to balance global semantics and local features, resolving Transformer’s inefficiency. Its Parallel Multi-level Aggregation decoder with Attention Gate and multi-branch convolutions enables dynamic hierarchical feature refinement for diverse polyp sizes and shapes, outperforming U-Net variants’ fixed skip connections. The Improved Reverse Attention module enhances edge extraction, mitigating artifacts (mucus, bubbles) that hinder PraNet and CaraNet.
3. Methodology
3.1. Overall Architecture
The inconsistency in defining segmentation targets in medical imaging is a well-documented issue, leading to suboptimal segmentation outcomes, poorly defined boundaries, and considerable challenges in achieving accurate diagnosis and the subsequent treatment planning []. To address these challenges, a novel multi-scale architectural framework is proposed, as shown in Figure 2.
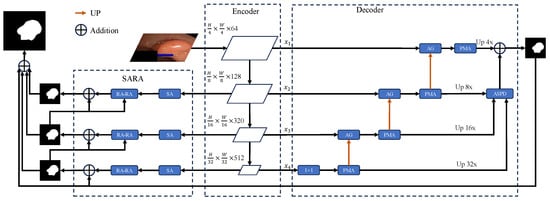
Figure 2.
Architecture of the proposed polyp segmentation model. Illustrates the architecture of the proposed model, which includes the backbone PVTv2, Parallel Multi-level Aggregation Decoder (PMAD), and Improved Reverse Attention Module.
The proposed model adopt the same structural design as U-net, featuring an encoder-decoder architecture, which is of the utmost importance for diagnostic and treatment planning tasks in medical imaging. This allows the network to the features that the image contains, as a process of image encoding is being undertaken. Subsequently, during the decoding phase, the segmentation results captured in the encoding phase must then be reinstated. Our model primarily consists of the following key components: the encoder, the decoder and the Improved Reverse Attention Module, as as illustrated.
In the encoder stage, the model employ PVTv2, which adapts to visual tasks by combining attention mechanisms adept at handling spatial and multi-scale features. This structured approach ensures that the model learns to identify polyp tissues with a high degree of precision. It achieves this by considering both the local context, such as the fine details of polyp edges and texture differences between lesions and surrounding mucosa, and the global context, including the overall distribution of polyps in the intestinal lumen and their positional relationships with large-scale anatomical structures within the image. Additionally, we introduce the Multiscale Attention module at the bottom stage of the encoder, which analyzes multi-level characteristics across various viewpoints, synthesizing both localized and extensive data for enhanced feature extraction compared to those that depend exclusively on single-scale cues. The decoder is specifically designed to leverage pyramid features—multi-scale feature maps extracted from the encoder backbone of the segmentation model. We combine attention with various parallel convolutional blocks to propose the Parallel Multi-level Aggregation Decoder, enabling the decoder to extract refined pyramid features as comprehensively as possible and further aggregate these features. Finally, through the Improved Reverse Attention module, we process the decoder output using the Scale Aggregation module, followed by reverse attention feature extraction, to further address boundary issues in polyp segmentation.
3.2. Multiscale Attention Module
Vision Transformer employs a global attention mechanism, facilitating the model’s comprehension of interconnections among diverse image spatches. However, computational costs grow rapidly as the number of image patches increases. For many typical computer vision tasks, modeling relationships between all image patches is not invariably essential. Some relationships may be redundant or have minimal impact on the final results.
By default, we use a convolution kernel size of 3 × 3 with a size and use hole convolutions with dilation ratios of 1, 2, and 3, which will bring the receptive field sizes of each head to 3 × 3, 5 × 5, and 7 × 7, respectively.The 3 × 3 convolution kernel is selected as the base due to its balance of effective local feature extraction, lower computational complexity compared to larger kernels such as 5 × 5 and 7 × 7, and flexibility in receptive field expansion via dilation. Dilation ratios 1, 2, and 3 are adopted to generate multi-scale receptive fields for each head, enabling adaptive capture of fine polyp boundaries, medium-sized regions, and large polyp structures while avoiding parameter redundancy.
Local attention [] reduces computational complexity but compromises long-range dependency modeling and global receptivity, presenting an inherent trade-off. To address this limitation, we propose a novel Multiscale Attention (MA) module, which is uniquely integrated into the multi-head attention of PVTv2’s bottom encoder stages, as shown in Figure 3.
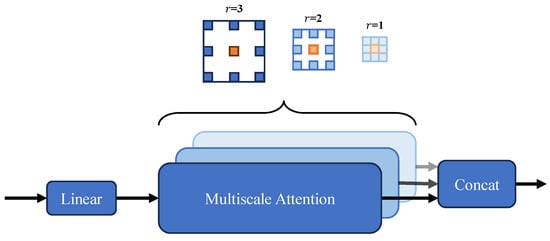
Figure 3.
Multiscale Attention (MA) module architecture schematic. The feature map’s channels are segmented into various components, and the self-attention process takes place among the blue blocks nestled in the vicinity of the red query block. This is all done while employing a range of dilation rates throughout the different segments. Additionally, concatenation of various section features is then presented to a linear processing layer. By default, we use a kernel size of 3 × 3, with dilation rates r = 1, 2, and 3, resulting in receptive field sizes of 3 × 3, 5 × 5, and 7 × 7 for the respective heads.
This module innovatively extends multi-head attention with flexible, cost-efficient sparse global attention: by incrementally expanding the receptive field without extra computation, it captures broader contextual dependencies while preserving fine-grained local details. Its tailored adaptation to PVTv2’s bottom layers specifically enhances low-level feature perception—critical for resolving subtle structures like small polyp edges and texture variations—thereby strengthening downstream segmentation precision. This design overcomes the limitations of conventional local attention in modeling long-range interactions, marking a distinct advance in optimizing encoder.
To formalize the computational workflow of the Multiscale Attention (MA) module, as shown in Algorithm 1, the following algorithm details its step-by-step process, including channel segmentation, multi-dilation convolution operations, self-attention computation, and feature fusion, with specific parameter settings as described.
| Algorithm 1 Multiscale Attention (MA) Module Algorithm |
| Input: Feature map (where H = height, W = width, C = channels) Output: Enhanced feature map
|
3.3. Parallel Multi-Level Aggregation Decoder
The proposed parallel multi-level aggregation decoder includes UP blocks, an Attention Gate (AG), a Parallel Multi-level Aggregation (PMA) module, and an Adjacent Supplement Partial Decoder (ASPD). The UP blocks enhance subdivision by preserving spatial details during upsampling, avoiding loss that blurs boundaries. The AG module boosts subdivision through cascaded fusion, focusing on critical cross-level features to sharpen fine-grained distinctions. The PMA module deepens subdivision via robust refinement and integration, extracting nuanced details beyond basic segmentation. The ASPD module enhances subdivision by aggregating final refined features, strengthening contextual coherence to clarify subtle subdivisions—each module’s originality lies in targeted, non-segmentation-based enhancement of feature subdivision. The parallel multi-level aggregation decoder streamlines segmentation by combining precise upsampling, focused attention, robust refinement, and contextual enhancement, ensuring sharp, fine-grained subdivisions and clear boundaries throughout the process.
3.3.1. Attention Gate (AG)
Low-layer features are considered to contain substantial redundant information, making them unsuitable for direct integration into subsequent feature extraction processes. Additionally, when low-layer features are directly upsampled and added to higher-layer features, it may lead to the loss of fine-grained details. Therefore, the model incorporated the Attention Gate module. Similar to Attention U-Net, the AG can assign varying degrees of weighting to different parts of the input data, enabling the network to focus on more critical regions.
Here, and correspond to the ReLU and Sigmoid activation functions, and denotes the process of computing the attention query vector based on the inputs, respectively. denotes batch normalization. Respectively g and x represent the upsampled and encoder output features. By incorporating the AG, nonlinearity can be significantly enhanced without altering the scale, refining the encoder’s output and enabling cross-channel information integration.
3.3.2. Parallel Multi-Level Aggregation (Pma)
Encoder convolution and downsampling increases feature dimensionality but reduces spatial scale, risking loss of small polyp details unrecoverable via decoder upsampling. Following the AG module, the novel Parallel Multi-level Aggregation (PMA) module includes Polarized Self-Attention (PSA) [] and a Multi-branch Feature Aggregation (MFA) module, as shown in Figure 4. PSA enhances subdivision by fusing channel and spatial attention to pinpoint and refine critical features and their positions, sharpening fine-grained distinctions without segmentation. MFA boosts subdivision via parallel multi-scale convolutional blocks, integrating diverse visual scales to capture nuanced details—both modules’ originality lies in non-segmentation-based, targeted enhancement of feature subdivision.
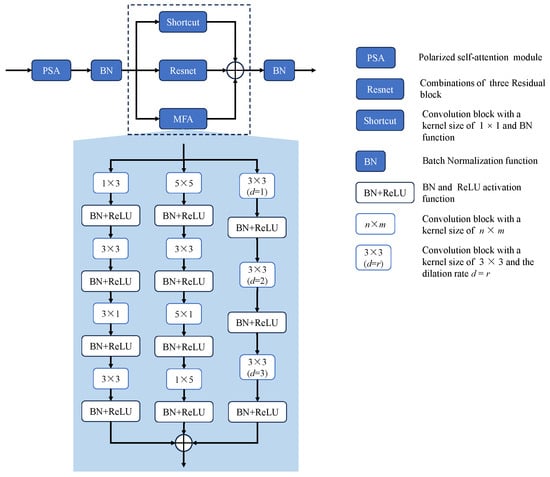
Figure 4.
The architecture of the Parallel Multi-level Aggregation (PMA) Module. The PMA module comprises a Polarized Self-Attention module and a feature refinement block, where the latter further consists of Multi-branch Feature Aggregation (MFA), a shortcut connection, and a residual connection.
To better refine feature maps, CBAM’s sequential spatial and channel attention structure [] is adopted: Channel Attention (CA) focuses on and optimizes key feature maps; Spatial Attention (SA) highlights critical locations. Unlike CBAM (relying on fully connected and convolutional layers, limiting information mining), we adopt PSA’s sequential design to enhance features and suppress noise. The self-attention module converts input x into query Q (fully compressed channels) and value V (retaining () channels), enabling efficient long-range modeling via Q-dimensionality reduction—ensuring computational efficiency, specific-direction feature compression, and perpendicular high resolution. Q is then enhanced via High Dynamic Range (HDR), with dimensionality increased through matrix multiplication and 1 × 1 convolution. By leveraging PSA’s efficient self-attention mechanism and the targeted enhancement of key features through Channel and Spatial Attention, the module refines feature maps and improves segmentation precision, particularly in capturing fine-grained spatial details and reducing noise interference. The entire process is as follows:
In Equation 3, and are the channel attention matrix and spatial attention matrix, respectively; and are the channel-wise multiplication operator and spatial multiplication operator.
The Multi-branch Feature Aggregation (MFA) module, a clever multi-convolutional setup, takes segmentation to the next level through three ingenious pathways. Borrowing a page from RFB Net [], it leverages various hole convolutions to beef up receptive fields, nailing down those all-important multi-scale features that make or break fine-grained segmentation edges. The first pathway stacks 3 × 3 dilated convolutions to weave together multi-scale details, giving object contours a much-needed sharpening. The second strikes a balance between scale variety and computational efficiency by employing 5 × 5 convolutions alongside diagonal ones, zeroing in on spatial subtleties that often define tricky or oddly shaped regions. The third pathway, armed with 3 × 3 diagonal convolutions and feature-organizing layers, keeps spatial information intact—a must-have for dead-on localization. This multi-pronged approach cranks up non-linearity, letting the network flex both global and local feature muscles; optimizing each branch independently also lightens the load, paving the way for deeper networks that pack richer segmentation insights. When you throw these aggregated features into the mix with smart shortcut and residual connections that keep the original data intact, the module really hits its stride, telling fine-grained segments apart rather than just carving up space willy-nilly. By strategically combining multi-scale features and enhancing spatial details through each pathway, the MFA module significantly improves segmentation accuracy, ensuring precise delineation of fine-grained boundaries and achieving sharper, more reliable object contours.
To clarify the operational mechanism of the MFA module, the following pseudocode formalizes its multi-branch convolutional design, receptive field expansion via dilated convolutions, and feature aggregation strategy, along with residual shortcut connections.
3.3.3. Adjacent Supplement Partial Decoder
The Adjacent Supplement Partial Decoder is added after the PMA module as a partial encoder. Due to the nature of pyramid features, multi-level pyramid features are generated at both the encoder and decoder stages. However, two key issues remain when aggregating multiple feature pyramids: how to maintain semantic consistency within layers and how to bridge context across layers. Here, we draw inspiration from the Cascaded Partial Decoder (CPD) [] by introducing a partial encoder to aggregate high-level output features. More specifically, we use the Adjacent Supplement Partial Decoder (ASPD) to combine these discriminative features through concatenation. Here, , , , and denote the multi-level pyramid features generated at different stages of the encoder and decoder, with representing the lowest-layer feature (rich in original details) and the highest-layer feature (more refined in semantics). Lower-layer features are upsampled to the same scale as the higher-layer features and then optimized via a 3 × 3 convolutional layer. These features are multiplied level-by-level with the higher-layer features, and the three resulting products are concatenated to obtain the final feature map. As shown in Figure 5, the computation involves processing , , and sequentially. Lower-layer features contain more original information, while higher-layer features are more refined. Multiplying the processed lower-layer features with higher-layer features and then combining multi-level features allows for better integration of contextual information, further highlighting the characteristics of each layer. Concatenating the results helps avoid losing optimized feature information.
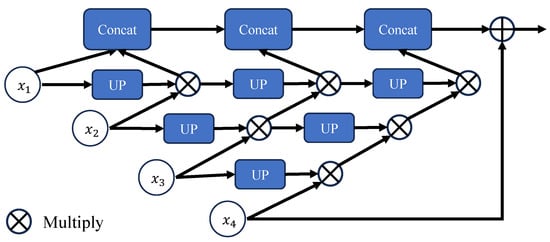
Figure 5.
The Adjacent Supplement Partial Decoder (ASPD) Module structure description. Illustrates the structure of the ASPD module, where UP represents a combination of a 3 × 3 convolutional layer and an upsampling operations. The input consists of lower-layer features processed through the UP block, which are then multiplied with higher-layer features. The outputs from each layer will be connected together to form the final output result.
The Algorithm 2 is shown below:
| Algorithm 2 Multi-branch Feature Aggregation (MFA) Module Algorithm |
| Input: Input feature map Output: Aggregated and refined feature map
|
3.4. Improved Reverse Attention Module
The decoder’s output is further processed by the Scale Aggregation (SA) module to attain a more accurate and comprehensive prediction map, as shown in Figure 6. The SA module consists of convolutions of different scales connected via concat. The reverse attention module is enhanced by a novel residual axial attention block for gradual correction of polyp boundaries. Therefore, we believe convolutional layers are crucial for the refinement module, as they emphasize high-frequency features and offer more detailed edge information for boundary refinement. The refinement module learns multi-scale features using convolutional kernels of varying sizes to handle scale variations effectively. To aggregate multi-scale information, features are concatenated along a specific dimension. The detailed flowchart of scale aggregation is illustrated in the figure. For convenience, we denote the input to SA as x. Specifically, spatial features are extracted from different scales by sequentially performing dilated convolutions, including 3 × 3, 5 × 5, and 7 × 7 operations. The features across various scales are subsequently concatenated. Additionally, a 1 × 1 convolutional layer is added for skip connections, ensuring the final aggregated features retain the original input information and effectively integrate the results. To reduce computational overhead, multiple 3 × 3 convolutions are used instead of 5 × 5 and 7 × 7 operations. Stacked convolutional layers provide more activation functions, enhancing the network’s nonlinearity and improving feature extraction and fusion.
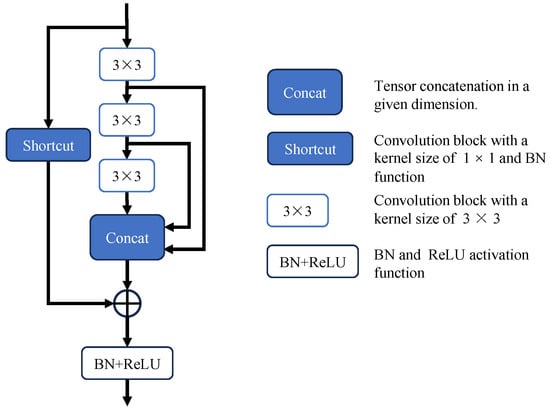
Figure 6.
Architecture of the Scale Aggregation (SA) module. Implements multi-scale convolutional operations using consecutive 3 × 3 convolutional layers and adds shortcuts to enhance residual connections.
The employment of the SA module’s multi-scale convolutional layers and residual axial attention block within the Improved Reverse Attention Module facilitates the refinement of polyp boundaries. This module enhances edge clarity, mitigates blurring via multi-scale feature aggregation, enables precise boundary correction in ambiguous regions, and achieves more accurate, detailed segmentation. The aforementioned process can be expressed as follows:
In Equation 4, denotes the concatenation operation. Subsequently, reverse attention is incorporated, drawing inspiration from PraNet’s parallel reverse attention network design. Starting from the saliency map generated at the deepest layer, existing estimated polyp regions are erased to compute the weights of the reverse attention module. Higher-layer output features are then subjected to element-wise dot products, using deeper features to sequentially mine complementary regions and details.
3.5. Loss Function
When analyzing polyp images, considering both the boundaries and polyp regions significantly contributes to segmentation performance. Specifically, the global loss optimizes overall polyp segmentation accuracy by focusing on the entire polyp region and background, while the local loss enhances boundary precision by assigning higher weights to edge pixels to address indistinct polyp edges in colonoscopy images. The proposed loss function employs a weighted combination [] of Intersection over Union (IoU) [] and Binary Cross-Entropy (BCE). The BCE effectively mitigates class imbalance through pixel-wise classification penalties, while IoU guarantees spatial consistency by quantifying regional overlap—both are critical for clinically relevant polyp segmentation—with their weighted formulation enabling adaptive alignment with task-specific challenges. Our loss calculation is as follows:
During loss computation, to better address boundary issues by increasing weights, encouraging the model to focus more intently on the edge regions, we introduce multiple prediction heads to separately calculate global and local losses. For model training, we use the decoder’s final aggregated output and the outputs of three reverse attention modules as prediction heads, and then use additive aggregation to calculate the final prediction map, as shown in Equation (6):
where , , and are the feature maps of the four prediction heads; a, b, c and d are the weights of each prediction head, all set to 1.0 here.
4. Experiments
4.1. Dataset
Experiments are conducted on five popular benchmark polyp segmentation datasets: Kvasir-SEG [], CVC-ClinicDB [], CVC-ColonDB [], Endoscene [] and ETIS-LaribPolypDB (also known as CVC-T) []. The details of this five datasets are described below:
As shown in Table 1, following the configuration of PraNet, the Kvasir-SEG and CVC-ClinicDB datasets are utilised for training purposes, with 80% of the data allocated for this purpose. Validation is performed using 10% of the dataset, and testing is conducted with a further 10% of the data. To assess generalisation-related performance, the model is tested on three unseen small datasets: EndoScene, CVC-ColonDB, and ETIS-LaribPolypDB. Due to varying image sizes in these three datasets, all exceeding 500 × 500, we resize all training images to 256 × 256 to conserve computational resources and accelerate convergence.

Table 1.
The number and size of five challenging benchmark polyp datasets is provided, in addition to the division of the train and test datasets followed the Pranet.
4.2. Implementation Details
Our experiments were conducted using PyTorch 1.11.0, torchvision 0.12.0, and Python 3.9.18 via the Anaconda command shell. The experimental platform for training and testing on a computer equipped with 13th Gen Intel(R) Core(TM) i7-13700KF @ 3.40 GHz and NVIDIA GeForce RTX 4080 with 32 GB of memory.
During training, the pre-trained weights derived from ImageNet dataset [] were utilised for the PVTv2-b2 backbone network for faster convergence, higher training efficiency, and enhanced performance in intestinal polyp segmentation. AdamW combined with weight decay effectively suppresses overfitting, while a 0.0001 learning rate balances convergence speed and stability; the 256 × 256 size achieves a trade-off between feature retention and computational efficiency. And multi-scale training was utilized with scales of 0.75, 1.0, and 1.25 to enhance the model’s adaptability to variations in input size is imperative, as is the improvement of its generalisation performance, serving as an alternative to data augmentation. Additionally, for better data augmentation, gradient clipping was set to 0.5, with a decay cycle of 200, and random flipping was applied to the images. All models in our tests made use of publicly available code, with the datasets being public and the pre-trained weights provided by the original authors.
Following the reverse attention mechanism module, our final prediction map , , and is generated by summation. The entire of the network is trained end-to-end, taking roughly 600 min to converge over 100 epochs with batches configured to a size of 16. To avoid underfitting or overfitting, we store the train-weights at the end of each epoch if performance on the validation set increases.
4.3. Evaluation Metrics
We primarily adopted the Mean Dice Similarity Coefficient (mDice) [] and Mean Intersection over Union (mIoU) as evaluation metrics to evaluate the model’s performance on the target dataset. The mDice is a set similarity measure function that is frequently employed to calculate the degree of similarity between two samples. It operates on a scale that varies from 0 to 1. The IoU metric is widely used in semantic segmentation, referring to the prediction accuracy for each category. This evaluation measure is a frequently used to gauge the models’ effectiveness in image segmentation, where increased values reflect a stronger correspondence between the segmented outputs and the actual ground truth.
Additionally, we employed other evaluation metrics such as the weighted F-measure () [], Structural measure () [], Enhanced-alignment measure () [] and Mean Absolute Error (). The calculates a weighted harmonic mean to balance precision and recall. The gauges structural resemblance between the prominence map and the reference standard. The employs an improved alignment matrix reflecting two binary map characteristics. The is the average absolute error between observed and true values, assessing pixel-level segmentation accuracy. Parameters (Params) refer to the total number of trainable variables in a model, directly related to its capacity and parameter efficiency. Floating-point operations (FLOPs) quantify the computational complexity of model training/inference, reflecting the required computing resources. Speed (fps, frames per second) denotes the number of input frames a model processes per second during inference, a key metric for evaluating real-time performance in latency-sensitive scenarios.
4.4. Results
Learning and generalization capability analysis: An evaluation was conducted for the purpose of determining the model’s performance when applied to five distinct datasets, with Kvasir-SEG and CVC-ClinicDB datasets used primarily to assess the capability of feature modeling, while the other unseen datasets were used to evaluate generalization ability. To assess model lightweighting, we compared FLOPs, Speed, and Params metrics, with experimental results.
The present study constitutes an experimental comparison of disparate network models on the Kvasir-SEG and CVC-ClinicDB datasets. The results on Kvasir-SEG are shown in Table 2. For ease of visualization, the two highest-scoring responses are distinguished by red and blue, respectively.

Table 2.
The results on the Kvasir-SEG for different models in a few noteworthy cases.
We compared the metric data with nine other state-of-the-art (SOTA) models. The comparative results are presented in Table 2 and Table 3, including reports on U-Net [], U-Net++ [], SFA [], PraNet [], MSNet [], CFA-Net [], MEGANet(Res2Net-50) [] and CSCA U-Net []. All comparative data were sourced directly from the original papers. The experimental results demonstrate that our model significantly outperforms both traditional and modern segmentation techniques. As shown in Table 2 and Table 3, our model exhibits improved feature modeling capabilities compared to other models and demonstrates better generalization across other datasets. Particularly in cross-dataset evaluations, our model outperformed the second-best CFA-Net model by 0.8% in the mDice metric on the Kvasir-SEG dataset and surpassed MEGANet and CSCA U-Net on the CVC-ClinicDB dataset. It also maintained strong performance in mIoU, leading significantly on the Kvasir-SEG dataset and trailing MEGANet by only 0.2% on the CVC-ClinicDB dataset. Additionally, our model performed well on other metrics, validating the algorithm’s robustness, stability, and generalization capabilities. Consistent progress over these multiple polyp datasets confirms the model’s robust generalisation under endoscopic imaging conditions.

Table 3.
The results on the CVC-ClinicDB for different models in a few noteworthy cases.
The experimental comparison of different network models on CVC-ColonDB, Endoscene, and ETIS-LaribPolypDB datasets. For ease of visualization, the two highest-scoring responses are distinguished by red and blue, respectively.
Since the datasets in Table 4, Table 5 and Table 6 were not included in the training set, all methods’ performance generally declined on these datasets, particularly on Endoscene and ETIS-LaribPolypDB. The model’s performance on the ETIS-LaribPolypDB dataset is demonstrated by an mDice score of 0.756 and an mIoU score of 0.673, with reaching 0.844 (the highest across all datasets) and at 0.709, both leading comparative models. Although on Endoscene, our metrics were slightly lower than MEGANet and marginally higher than CFA-Net, our proposed method still improved in boundary clarity and detail preservation. The visualization reveals that our model provides clearer boundary judgments, enabled by the Improved Reverse Attention module that optimizes edge entropy and reduces segmentation blur, allowing the segmentation of complex and subtle details in ambiguous cases and performing well even in challenging scenarios with uncertain regions—this is further validated by the optimal MAE (0.028) on CVC-ColonDB and 0.006 on CVC-ClinicDB, achieved through the synergy of MA module’s detail error reduction and reverse attention’s boundary optimization.

Table 4.
The results on the CVC-ColonDB for different models in a few noteworthy cases.

Table 5.
The results on the Endoscene for different models in a few noteworthy cases.

Table 6.
The results on the ETIS-LaribPolypDB for different models in a few noteworthy cases.
On the CVC-ColonDB dataset, our model outperformed all alternative methods across the majority of evaluation metrics, including mDice 0.820, mIoU 0.738, 0.805, and 0.869, demonstrating excellent segmentation performance. Despite the noteworthy segmentation capabilities demonstrated by MEGANet and CSCA U-Net, and the incorporation of cross-level feature fusion in CFA-Net, the efficacy of these methods in segmenting polyps with blurred edges was found to be unsatisfactory due to insufficient low-level detail extraction. n contrast, our model achieves precise polyp segmentation and superior results by integrating the MA module at the encoder’s lower stages—its multi-scale attention and dilated convolutions capture fine-grained features, avoid small target/weak boundary omission, and enhance effective feature weighting to suppress background interference. It is paired with hierarchical feature supplementation from the integrated decoder and edge-focused optimization from the Improved Reverse Attention module. These two components collectively boost overlap rate, positioning accuracy, and structural consistency across datasets, as evidenced by the standout metrics on Kvasir-SEG: mDice 0.923, mIoU 0.870, and 0.967.
The visual analysis results of the comparative experiments are shown in Figure 7, Figure 8 and Figure 9. The proposed model is compared with five SOTA polyp segmentation methods (PraNet, MSNet, CFA-Net, MEGANet, CSCA U-Net) across five datasets, showing superior performance in segmenting challenging targets. On Kvasir-SEG as shown in Figure 7, CSCA U-Net and CFA-Net lack fine texture restoration for small polyps and show slight thin edge discontinuities, while our model captures low-level micro-textures with continuous edges via the MA module’s multi-scale attention and dilated convolutions. For irregular, low-contrast polyps as shown in Figure 8, MEGANet and MSNet exhibit detail loss and blurred edges at contour turns, whereas our PMA and Improved Reverse Attention modules synergistically enhance boundary discrimination, achieving near-ground truth segmentation. In Figure 9, PraNet and CFA-Net present blurred edges and background missegmentation; our Improved Reverse Attention module (with a pre-added SA module for precise localization) effectively distinguishes polyp-background pixels, ensuring sharp, continuous edges with high ground truth alignment.
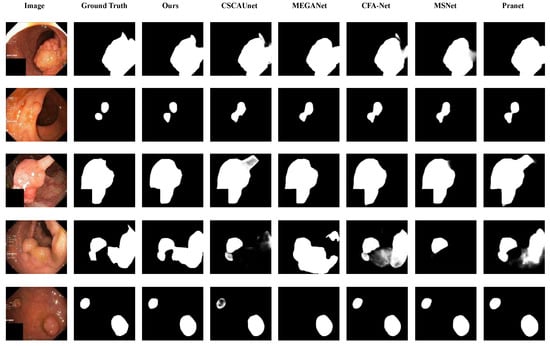

Figure 7.
Qualitative analysis on the Kvasir-SEG dataset for different models in several noteworthy cases.

Figure 8.
Qualitative analysis on the CVC-ClinicDB dataset for different models in several noteworthy cases.
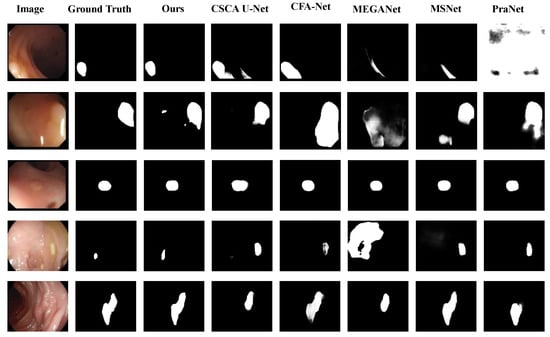
Figure 9.
Qualitative analysis on three Benchmark Datasets for different models in several noteworthy cases. The first and second rows belong to the CVC-ClinicDB dataset, the third row to the CVC-300 dataset, and the last two rows to the ETIS-LaribPolypDB dataset.
As shown in Figure 8, for the fourth input image, it can be observed that the segmentation results of CSCA U-Net, MSNet, and PraNet are incomplete, whereas our method achieves segmentation results closer to the ground truth.
Based on the validation of segmentation results, further analysis is conducted from the perspective of computational efficiency. The model’s lightweight nature and operational performance are quantified through metrics including FLOPs, Speed (FPS), and Params (M) presented in Table 7. The proposed model exhibits balanced performance in terms of parameter quantity, computational cost, and inference speed, which lays a solid foundation for its deployment in real-world hospital environments. The model comes in at 45.2 million trainable parameters—small enough to prevent hospital edge devices from being overwhelmed by memory demands, retaining sufficient capacity to learn fine-grained polyp features from medical images. When saved as a standard .pth file (only storing model weights), the file size is approximately 172 MB, which is easy to transmit and deploy across hospital internal systems without consuming excessive storage resources.

Table 7.
Comparison of FLOPs, Speed and Params for different models.
In terms of inference computational cost and speed (tested on NVIDIA RTX 4080, with 256 × 256 input size), the model achieves an average of 9.8 GFLOPs per test image, which effectively sidesteps the bogged-down inference times that typically plague systems with excessive computational demands. The proposed model achieves an average inference time of 5.88–5.90 ms per test image (derived from 170 FPS), which is far faster than the clinical real-time requirement of at least 30 FPS. Ensuring low latency in real-time endoscopic examinations minimizes imaging-segmentation display delays for doctors, a key guarantee for the continuity of clinical operations.
4.5. Ablation Study
In this section, we present a detailed account of the ablation experiments that were conducted on the model’s key modules. The purpose of these experiments was to demonstrate the roles of these modules in the context of the entire network. Table 8 summarizes the specific test results, where the Improved Reverse Attention module is denoted as SARA in tables and figures. Figure 10 displays the final segmentation results after removing or replacing modules. In the single-module ablation experiments, we directly removed the Improved Reverse Attention Module, replaced the MA module with the original attention module from PVTv2, and substituted the PAM module with a combination of 3 × 3 convolutional block, BN, and ReLU functions, while keeping other structures unchanged. We selected two metrics to evaluate our method: mDice and mIoU. The ablation study was conducted on two datasets used for training. Table 8 shows that in single-module ablation experiments, the PAM module contributed more to the model than other modules. This is likely due to its convolutions with varying receptive fields capturing multi-scale features and the improved attention mechanism enhancing feature selection and extraction. Notably, on the Kvasir-SEG dataset, mDice improved by 0.011. Similarly, the Improved Reverse Attention Module significantly aided the model in edge segmentation, as evidenced by Figure 10, where the model with this module exhibited clearer boundaries on the CVC-ClinicDB dataset compared to the model without it, which also underperformed in mDice and mIoU. The MA module contributed to the model but showed smaller improvements compared to the other two proposed modules. As seen in the first and third rows of Figure 10, it had certain limitations in polyp segmentation refinement, failing to effectively capture relationships among features across larger regions, resulting in segmentation that did not fully encompass the polyp area.

Table 8.
The results of the ablation study on Kvasir-SEG and CVC-ClinicDB datasets.
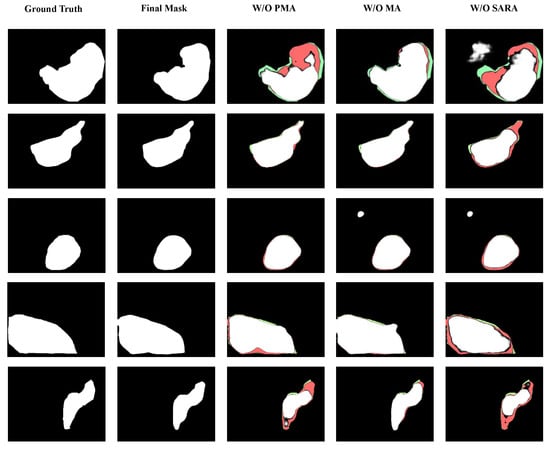
Figure 10.
Core module ablation impact on the proposed Polyp segmentation accuracy. Results of the ablation study, where the last three images in the horizontal row respectively show the outcomes after removing the corresponding modules. The first row displays results from the Kvasir-SEG dataset, the second row from the CVC-ClinicDB dataset, the third and fourth rows from the CVC-ColonDB dataset, and the final row from the ETIS-LaribPolypDB dataset. Green, red, and white regions represent the ground truth, final mask, and the false negative. It can be observed that removing any module affects segmentation accuracy, leading to missed or erroneous detection results. The proposed model can accurately locate and segment polyps regardless of size.
Overall, the PAM module significantly enhances the model’s ability to extract and aggregate polyp features from the decoder output, laying a foundation for subsequent work. Additionally, the Improved Reverse Attention module also contributes substantially to the overall network performance.
5. Conclusions
5.1. Discussion
In this paper, the proposed model dedicated to the task of polyp image segmentation. Currently, some classical polyp segmentation networks still struggle to address the challenges in this field, such as indistinct polyp boundaries, dispersed polyp regions, and image blurring or reflections. To tackle these issues, we incorporate more parallel multi-branch convolutional blocks and an enhanced reverse attention mechanism.
We utilize the traditional classic U-net encoder-decoder architecture, which is crucial for medical image diagnosis and treatment planning, as it captures image features during encoding and reconstructs segmentation results during decoding. The model consists of three core components:
- (1)
- The encoder employs PVTv2, leveraging spatial and multi-scale feature attention mechanisms for visual tasks. A Multiscale Attention module is introduced at the bottom stage of the encoder to optimize multi-scale feature extraction by fusing local and global information, improving polyp tissue recognition accuracy.
- (2)
- The parallel multi-level aggregation decoder is specifically designed for pyramid features, combining attention mechanisms with multi-convolutional blocks in parallel through residual convolutions to efficiently integrate and re-extract multi-scale feature maps from the encoder output.
- (3)
- The Scale Aggregation Reverse Attention module first processes decoder outputs via Scale Aggregation and then reinforces feature extraction using reverse attention, focusing on optimizing boundary recognition accuracy for polyp segmentation.
Evidently, the model demonstrates a high level of proficiency when applied to five polyp datasets, and we validated the effectiveness of the proposed three modules through ablation experiments. To evaluate our model’s performance, we compared it with nine SOTA segmentation networks. Table 2, Table 3, Table 4, Table 5, Table 6, Table 7 and Table 8 and Figure 7, Figure 8, Figure 9 and Figure 10 clearly demonstrate that our model achieves the best segmentation results on five small polyp target datasets, scoring high in both mDice and mIoU, indicating superior feature capture capability. Particularly in cross-dataset evaluations, our model outperforms the second-best MEGANet by 0.2% in mDice on the Kvasir-SEG dataset and surpasses UHA-Net and CSCA U-Net on the CVC-ClinicDB dataset. It also maintains strong performance in mIoU, trailing only slightly behind MEGANet. On the Endoscene dataset, compared to CFA-Net, the proposed model outperforms it in mIoU (0.829 vs. 0.827, +0.2%), while maintaining competitive mDice (0.895 vs. 0.893, +0.2%) and (0.878 vs. 0.875, +0.3%), and its MAE (0.007) is 0.1% lower than CFA-Net’s (0.008), highlighting its superior ability to capture polyp structural integrity and reduce prediction errors. On the CVC-ColonDB and ETIS-LaribPolypDB dataset, our proposed model achieves the best performance across key evaluation metrics—securing the highest mDice, mIoU, , and , while maintaining competitive scores in and , demonstrating its superior ability to accurately segment polyps even on this challenging dataset. These results validate the algorithm’s robustness, stability, and generalization capability. The model’s strong generalisation under diverse endoscopic imaging conditions has been confirmed by consistent progress across multiple independent datasets.
In summary, our proposed model has yielded satisfactory segmentation outcomes for small polyps, but it is imperative to acknowledge that the method still has shortcomings when dealing with complex and challenging samples. From the third row of Figure 8 and the fourth row of Figure 9, it can be seen that there is still significant room for improvement in segmentation accuracy between our method and SOTA segmentation methods. However, in terms of overall comparison, the segmentation results of the proposed model are the closest alignment with the ground truth.
5.2. Conclusions
In this paper, we propose a model that differs from other polyp segmentation methods. Utilizing PVTv2 as the backbone architecture, we integrate a Multiscale Attention module in initial encoder phases to enhance the receptive field for lower-layer feature discernment, and introduce an Improved Reverse Attention module to address the segmentation of polyp edges. We conducted ablation and comparative experiments on five small polyp target datasets, all achieving promising results, demonstrating the performance of our model in polyp segmentation tasks.
Author Contributions
Methodology, Project administration, Software, Validation, Visualization, Writing—original draft, R.Y.; Funding acquisition, Writing—review & editing, D.Z.; Formal analysis, Data curation, Resources, Supervision, Y.W. All authors have read and agreed to the published version of the manuscript.
Funding
This work was primarily supported by the National Natural Science Foundation of China under Grants 62066047 and 61966037.
Data Availability Statement
The original contributions presented in this study are included in the article. Further inquiries can be directed to the corresponding author.
Acknowledgments
The authors would like to express their gratitude to the anonymous reviewers whose efforts have resulted in the enhancement of the manuscript. The quality and presentation of their work are of paramount importance.
Conflicts of Interest
The authors declare no conflicts of interest.
References
- Bray, F.; Laversanne, M.; Sung, H.; Ferlay, J.; Siegel, R.L.; Soerjomataram, I.; Jemal, A. Global cancer statistics 2022: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2024, 74, 229–263. [Google Scholar] [CrossRef] [PubMed]
- Li, N.; Lu, B.; Luo, C.; Cai, J.; Lu, M.; Zhang, Y.; Chen, H.; Dai, M. Incidence, mortality, survival, risk factor and screening of colorectal cancer: A comparison among China, Europe, and northern America. Cancer Lett. 2021, 522, 255–268. [Google Scholar] [CrossRef]
- Bernal, J.; Tajkbaksh, N.; Sánchez, F.J.; Matuszewski, B.J.; Chen, H.; Yu, L.; Angermann, Q.; Romain, O.; Rustad, B.; Balasingham, I.; et al. Comparative Validation of Polyp Detection Methods in Video Colonoscopy: Results From the MICCAI 2015 Endoscopic Vision Challenge. IEEE Trans. Med Imaging 2017, 36, 1231–1249. [Google Scholar] [CrossRef]
- Jia, X.; Xing, X.; Yuan, Y.; Xing, L.; Meng, M.Q.H. Wireless Capsule Endoscopy: A New Tool for Cancer Screening in the Colon With Deep-Learning-Based Polyp Recognition. Proc. IEEE 2020, 108, 178–197. [Google Scholar] [CrossRef]
- Gupta, M.; Mishra, A. A systematic review of deep learning based image segmentation to detect polyp. Artif. Intell. Rev. 2024, 57. [Google Scholar] [CrossRef]
- Bang, C.S.; Lee, J.J.; Baik, G.H. Computer-Aided Diagnosis of Diminutive Colorectal Polyps in Endoscopic Images: Systematic Review and Meta-analysis of Diagnostic Test Accuracy. J. Med. Internet Res. 2021, 23, e29682. [Google Scholar] [CrossRef]
- Sánchez-González, A.; García-Zapirain, B.; Sierra-Sosa, D.; Elmaghraby, A. Automatized colon polyp segmentation via contour region analysis. Comput. Biol. Med. 2018, 100, 152–164. [Google Scholar] [CrossRef]
- Wang, W.; Xie, E.; Li, X.; Fan, D.P.; Song, K.; Liang, D.; Lu, T.; Luo, P.; Shao, L. PVT v2: Improved baselines with Pyramid Vision Transformer. Comput. Vis. Media 2022, 8, 415–424. [Google Scholar] [CrossRef]
- Jin, E.H.; Lee, D.; Bae, J.H.; Kang, H.Y.; Kwak, M.S.; Seo, J.Y.; Yang, J.I.; Yang, S.Y.; Lim, S.H.; Yim, J.Y.; et al. Improved Accuracy in Optical Diagnosis of Colorectal Polyps Using Convolutional Neural Networks with Visual Explanations. Gastroenterology 2020, 158, 2169–2179.e8. [Google Scholar] [CrossRef] [PubMed]
- Choi, S.J.; Kim, E.S.; Choi, K. Prediction of the histology of colorectal neoplasm in white light colonoscopic images using deep learning algorithms. Sci. Rep. 2021, 11, 5311. [Google Scholar] [CrossRef] [PubMed]
- Long, J.; Shelhamer, E.; Darrell, T. Fully convolutional networks for semantic segmentation. In Proceedings of the 2015 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Boston, MA, USA, 7–12 June 2015; pp. 3431–3440. [Google Scholar] [CrossRef]
- Ronneberger, O.; Fischer, P.; Brox, T. U-Net: Convolutional Networks for Biomedical Image Segmentation. In Proceedings of the Medical Image Computing and Computer-Assisted Intervention—MICCAI 2015, Munich, Germany, 5–9 October 2015; Volume 9351, pp. 234–241. [Google Scholar] [CrossRef]
- Zhou, Z.; Rahman Siddiquee, M.M.; Tajbakhsh, N.; Liang, J. Segmentation. In Deep Learning in Medical Image Analysis and Multimodal Learning for Clinical Decision Support; Springer International Publishing: Berlin/Heidelberg, Germany, 2018; pp. 3–11. [Google Scholar] [CrossRef]
- Oktay, O.; Schlemper, J.; Le Folgoc, L.; Lee, M.; Heinrich, M.; Misawa, K.; Mori, K.; McDonagh, S.; Y Hammerla, N.; Kainz, B.; et al. Attention U-Net: Learning Where to Look for the Pancreas. arXiv 2018, arXiv:1804.03999. [Google Scholar] [CrossRef]
- Jin, Q.; Meng, Z.; Sun, C.; Cui, H.; Su, R. RA-UNet: A Hybrid Deep Attention-Aware Network to Extract Liver and Tumor in CT Scans. Front. Bioeng. Biotechnol. 2020, 8, 605132. [Google Scholar] [CrossRef]
- Vaswani, A.; Shazeer, N.; Parmar, N.; Uszkoreit, J.; Jones, L.; Gomez, A.N.; Kaiser, L.; Polosukhin, I. Attention is all you need. In Neural Information Processing Systems; Curran Associates Inc.: Red Hook, NY, USA, 2017; pp. 6000–6010. [Google Scholar] [CrossRef]
- Dosovitskiy, A.; Beyer, L.; Kolesnikov, A.; Weissenborn, D.; Zhai, X.; Unterthiner, T.; Dehghani, M.; Minderer, M.; Heigold, G.; Gelly, S.; et al. An image is worth 16x16 words: Transformers for image recognition at scale. arXiv 2021, arXiv:2010.11929. [Google Scholar] [CrossRef]
- Chen, J.; Lu, Y.; Yu, Q.; Luo, X.; Adeli, E.; Wang, Y.; Lu, L.; Yuille, A.L.; Zhou, Y. Transunet: Transformers make strong encoders for medical image segmentation. arXiv 2021, arXiv:abs/2102.04306. [Google Scholar] [CrossRef]
- Wang, W.; Xie, E.; Li, X.; Fan, D.P.; Song, K.; Liang, D.; Lu, T.; Luo, P.; Shao, L. Pyramid Vision Transformer: A Versatile Backbone for Dense Prediction without Convolutions. In Proceedings of the 2021 IEEE/CVF International Conference on Computer Vision (ICCV), Montreal, QC, Canada, 11–17 October 2021; pp. 548–558. [Google Scholar] [CrossRef]
- Shi, W.; Xu, J.; Gao, P. SSformer: A Lightweight Transformer for Semantic Segmentation. In Proceedings of the 2022 IEEE 24th International Workshop on Multimedia Signal Processing (MMSP), Shanghai, China, 26–28 September 2022; pp. 1–5. [Google Scholar] [CrossRef]
- Sanderson, E.; Matuszewski, B.J. FCN-Transformer Feature Fusion for Polyp Segmentation. In Proceedings of the Medical Image Understanding and Analysis: 26th Annual Conference, MIUA 2022, Cambridge, UK, 27–29 July 2022; Proceedings. Volume 13413, pp. 892–907. [Google Scholar] [CrossRef]
- Simonyan, K.; Zisserman, A. Very deep convolutional networks for large-scale visual recognition. arXiv 2014, arXiv:1409.1556. [Google Scholar] [CrossRef]
- He, K.M.; Zhang, X.Y.; Ren, S.Q.; Sun, J. Deep Residual Learning for Image Recognition. In Proceedings of the 2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Las Vegas, NV, USA, 27–30 June 2016; pp. 770–778. [Google Scholar] [CrossRef]
- Valanarasu, J.M.J.; Patel, V.M. UNeXt: MLP-Based Rapid Medical Image Segmentation Network. In Proceedings of the Medical Image Computing and Computer Assisted Intervention—MICCAI 2022, Singapore, 18–22 September 2022; Springer Nature: Cham, Switzerland, 2022; pp. 23–33. [Google Scholar] [CrossRef]
- Wang, Z.; Cun, X.; Bao, J.; Zhou, W.; Liu, J.; Li, H. Uformer: A General U-Shaped Transformer for Image Restoration. In Proceedings of the 2022 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), New Orleans, LA, USA, 19–24 June 2022; pp. 17662–17672. [Google Scholar] [CrossRef]
- Dong, X.Y.; Bao, J.M.; Chen, D.D.; Zhang, W.M.; Yu, N.H.; Yuan, L.; Chen, D.; Guo, B.N.; Ieee Comp, S.O.C. CSWin Transformer: A General Vision Transformer Backbone with Cross-Shaped Windows. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), New Orleans, LA, USA, 18–24 June 2022; pp. 12114–12124. [Google Scholar] [CrossRef]
- Yuan, Y.; Fu, R.; Huang, L.; Lin, W.; Zhang, C.; Chen, X.; Wang, J. HRFormer: High-resolution transformer for dense prediction. In Proceedings of the 35th International Conference on Neural Information Processing Systems, Online, 6–14 December 2021; Curran Associates Inc.: Red Hook, NY, USA, 2021; p. 557. [Google Scholar] [CrossRef]
- Wang, J.; Sun, K.; Cheng, T.; Jiang, B.; Deng, C.; Zhao, Y.; Liu, D.; Mu, Y.; Tan, M.; Wang, X.; et al. Deep High-Resolution Representation Learning for Visual Recognition. IEEE Trans. Pattern Anal. Mach. Intell. 2021, 43, 3349–3364. [Google Scholar] [CrossRef]
- Hatamizadeh, A.; Nath, V.; Tang, Y.; Yang, D.; Roth, H.R.; Xu, D. Swin UNETR: Swin Transformers for Semantic Segmentation of Brain Tumors in MRI Images. In Brainlesion: Glioma, Multiple Sclerosis, Stroke and Traumatic Brain Injuries; Springer International Publishing: Berlin/Heidelberg, Germany, 2022; pp. 272–284. [Google Scholar] [CrossRef]
- Liu, Z.; Lin, Y.; Cao, Y.; Hu, H.; Wei, Y.; Zhang, Z.; Lin, S.; Guo, B. Swin Transformer: Hierarchical Vision Transformer using Shifted Windows. In Proceedings of the 2021 IEEE/CVF International Conference on Computer Vision (ICCV), Montreal, QC, Canada, 11–17 October 2021; pp. 9992–10002. [Google Scholar] [CrossRef]
- Fan, D.P.; Ji, G.P.; Zhou, T.; Chen, G.; Fu, H.; Shen, J.; Shao, L. PraNet: Parallel Reverse Attention Network for Polyp Segmentation. In Proceedings of the Medical Image Computing and Computer Assisted Intervention—MICCAI 2020, Lima, Peru, 4–8 October 2020; Springer International Publishing: Berlin/Heidelberg, Germany, 2020; pp. 263–273. [Google Scholar] [CrossRef]
- Kim, T.; Lee, H.; Kim, D. UACANet: Uncertainty Augmented Context Attention for Polyp Segmentation. In Proceedings of the 29th ACM International Conference on Multimedia, Virtual Event, 20–24 October 2021; Association for Computing Machinery: New York, NY, USA, 2021. MM ’21. pp. 2167–2175. [Google Scholar] [CrossRef]
- Lou, A.G.; Guan, S.Y.; Ko, H.; Loew, M. CaraNet: Context Axial Reverse Attention Network for Segmentation of Small Medical Objects. J. Med. Imaging 2022, 10, 014005. [Google Scholar] [CrossRef]
- Yang, Y.; Dasmahapatra, S.; Mahmoodi, S. ADS_UNet: A nested UNet for histopathology image segmentation. Expert Syst. Appl. 2023, 226, 120128. [Google Scholar] [CrossRef]
- Zhao, X.; Jia, H.; Pang, Y.; Lv, L.; Tian, F.; Zhang, L.; Sun, W.; Lu, H. M2SNet: Multi-scale in Multi-scale Subtraction Network for Medical Image Segmentation. arXiv 2023, arXiv:2303.10894. [Google Scholar] [CrossRef]
- Zhou, T.; Zhou, Y.; Li, G.; Chen, G.; Shen, J. Uncertainty-Aware Hierarchical Aggregation Network for Medical Image Segmentation. IEEE Trans. Circuits Syst. Video Technol. 2024, 34, 7440–7453. [Google Scholar] [CrossRef]
- Zhou, T.; Zhang, Y.; Chen, G.; Zhou, Y.; Wu, Y.; Fan, D.P. Edge-aware Feature Aggregation Network for Polyp Segmentation. Mach. Intell. Res. 2025, 22, 101–116. [Google Scholar] [CrossRef]
- Lee, H.J.; Kim, J.U.; Lee, S.; Kim, H.G.; Ro, Y.M. Structure Boundary Preserving Segmentation for Medical Image With Ambiguous Boundary. In Proceedings of the 2020 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), Seattle, WA, USA, 13–19 June 2020; pp. 4816–4825. [Google Scholar] [CrossRef]
- Luong, M.T.; Pham, H.; Manning, C.D. Effective Approaches to Attention-based Neural Machine Translation. In Proceedings of the 2015 Conference on Empirical Methods in Natural Language Processing, Lisbon, Portugal, 17–21 September 2015; pp. 1412–1421. [Google Scholar] [CrossRef]
- Liu, H.; Liu, F.; Fan, X.; Huang, D. Polarized self-attention: Towards high-quality pixel-wise mapping. Neurocomputing 2022, 506, 158–167. [Google Scholar] [CrossRef]
- Woo, S.; Park, J.; Lee, J.Y.; Kweon, I.S. CBAM: Convolutional Block Attention Module. In Proceedings of the Computer Vision—ECCV 2018, Munich, Germany, 8–14 September 2018; Springer: Berlin/Heidelberg, Germany, 2018; pp. 3–19. [Google Scholar] [CrossRef]
- Liu, S.T.; Huang, D.; Wang, Y.H. Receptive Field Block Net for Accurate and Fast Object Detection. In Proceedings of the European Conference on Computer Vision (ECCV), Munich, Germany, 8–14 September 2018; Volume 11215, pp. 404–419. [Google Scholar] [CrossRef]
- Wu, Z.; Su, L.; Huang, Q. Cascaded Partial Decoder for Fast and Accurate Salient Object Detection. In Proceedings of the 2019 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), Long Beach, CA, USA, 15–20 June 2019; pp. 3902–3911. [Google Scholar] [CrossRef]
- Wei, J.; Wang, S.; Huang, Q. F3Net: Fusion, feedback and focus for salient object detection. In Proceedings of the AAAI Conference on Artificial Intelligence, New York, NY, USA, 7–12 February 2020; Volume 34, pp. 12321–12328. [Google Scholar] [CrossRef]
- Yu, J.; Jiang, Y.; Wang, Z.; Cao, Z.; Huang, T. UnitBox: An Advanced Object Detection Network. In Proceedings of the 24th ACM International Conference on Multimedia, Amsterdam, The Netherlands, 15–19 October 2016; Association for Computing Machinery: New York, NY, USA, 2016; pp. 516–520. [Google Scholar] [CrossRef]
- Jha, D.; Smedsrud, P.H.; Riegler, M.A.; Halvorsen, P.; Lange, T.d.; Johansen, D.; Johansen, H.D. Kvasir-SEG: A Segmented Polyp Dataset. In Proceedings of the MultiMedia Modeling: 26th International Conference, MMM 2020, Daejeon, Republic of Korea, 5–8 January 2020; Proceedings, Part II. Springer: Berlin/Heidelberg, Germany, 2020; pp. 451–462. [Google Scholar] [CrossRef]
- Bernal, J.; Sánchez, F.J.; Fernández-Esparrach, G.; Gil, D.; Rodríguez, C.; Vilariño, F. WM-DOVA maps for accurate polyp highlighting in colonoscopy: Validation vs. saliency maps from physicians. Comput. Med Imaging Graph. 2015, 43, 99–111. [Google Scholar] [CrossRef]
- Tajbakhsh, N.; Gurudu, S.R.; Liang, J. Automated Polyp Detection in Colonoscopy Videos Using Shape and Context Information. IEEE Trans. Med Imaging 2016, 35, 630–644. [Google Scholar] [CrossRef]
- Vázquez, D.; Bernal, J.; Sánchez, F.J.; Fernández-Esparrach, G.; López, A.M.; Romero, A.; Drozdzal, M.; Courville, A. A Benchmark for Endoluminal Scene Segmentation of Colonoscopy Images. J. Healthc. Eng. 2017, 2017, 4037190. [Google Scholar] [CrossRef] [PubMed]
- Silva, J.; Histace, A.; Romain, O.; Dray, X.; Granado, B. Toward embedded detection of polyps in WCE images for early diagnosis of colorectal cancer. Int. J. Comput. Assist. Radiol. Surg. 2014, 9, 283–293. [Google Scholar] [CrossRef] [PubMed]
- Deng, J.; Dong, W.; Socher, R.; Li, L.J.; Kai, L.; Li, F.F. ImageNet: A large-scale hierarchical image database. In Proceedings of the 2009 IEEE Conference on Computer Vision and Pattern Recognition, Miami, FL, USA, 20–25 June 2009; pp. 248–255. [Google Scholar] [CrossRef]
- Milletari, F.; Navab, N.; Ahmadi, S. V-Net: Fully Convolutional Neural Networks for Volumetric Medical Image Segmentation. In Proceedings of the 2016 Fourth International Conference on 3D Vision (3DV), Stanford, CA, USA, 25–28 October 2016; pp. 565–571. [Google Scholar] [CrossRef]
- Margolin, R.; Zelnik-Manor, L.; Tal, A. How to Evaluate Foreground Maps. In Proceedings of the 2014 IEEE Conference on Computer Vision and Pattern Recognition, Columbus, OH, USA, 24–27 June 2014; pp. 248–255. [Google Scholar] [CrossRef]
- Fan, D.P.; Cheng, M.M.; Liu, Y.; Li, T.; Borji, A. Structure-Measure: A New Way to Evaluate Foreground Maps. In Proceedings of the 2017 IEEE International Conference on Computer Vision (ICCV), Venice, Italy, 22–29 October 2017; pp. 4558–4567. [Google Scholar] [CrossRef]
- Fan, D.P.; Gong, C.; Cao, Y.; Ren, B.; Cheng, M.M.; Borji, A. Enhanced-alignment measure for binary foreground map evaluation. In Proceedings of the 27th International Joint Conference on Artificial Intelligence, Stockholm, Sweden, 9–19 July 2018; AAAI Press: Washington, DC, USA, 2018; pp. 698–704. [Google Scholar] [CrossRef]
- Fang, Y.; Chen, C.; Yuan, Y.; Tong, K.y. Selective Feature Aggregation Network with Area-Boundary Constraints for Polyp Segmentation. In Proceedings of the Medical Image Computing and Computer Assisted Intervention—MICCAI 2019, Shenzhen, China, 13–17 October 2019; Springer International Publishing: Berlin/Heidelberg, Germany, 2019; pp. 302–310. [Google Scholar] [CrossRef]
- Zhao, X.; Zhang, L.; Lu, H. Automatic Polyp Segmentation via Multi-scale Subtraction Network. In Proceedings of the Medical Image Computing and Computer Assisted Intervention—MICCAI 2021, Strasbourg, France, 27 September–1 October 2021; Springer International Publishing: Berlin/Heidelberg, Germany, 2021; pp. 120–130. [Google Scholar] [CrossRef]
- Patel, K.; Bur, A.M.; Wang, G. Enhanced U-Net: A Feature Enhancement Network for Polyp Segmentation. Proc. Int. Robot Vis. Conf. 2021, 2021, 181–188. [Google Scholar] [CrossRef]
- Zhou, T.; Zhou, Y.; He, K.; Gong, C.; Yang, J.; Fu, H.; Shen, D. Cross-level Feature Aggregation Network for Polyp Segmentation. Pattern Recognit. 2023, 140, 109555. [Google Scholar] [CrossRef]
- Bui, N.T.; Dinh-Hieu, H.; Quang-Thuc, N.; Minh-Triet, T.; Le, N.; Soc, I.C. MEGANet: Multi-Scale Edge-Guided Attention Network for Weak Boundary Polyp Segmentation. In Proceedings of the 2024 IEEE/CVF Winter Conference on Applications of Computer Vision (WACV), Waikoloa, HI, USA, 3–8 January 2024; pp. 7970–7979. [Google Scholar] [CrossRef]
- Shu, X.; Wang, J.; Zhang, A.; Shi, J.; Wu, X.J. CSCA U-Net: A channel and space compound attention CNN for medical image segmentation. Artif. Intell. Med. 2024, 150, 102800. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).