A Novel Method to Use Coordinate Based Meta-Analysis to Determine a Prior Distribution for Voxelwise Bayesian Second-Level fMRI Analysis
Abstract
:1. Introduction
2. Experimental Procedure
2.1. Materials
2.1.1. Statistical Image Datasets for Analyses
2.1.2. Meta-Analysis Results for Prior Determination and Performance Evaluation
2.2. Basis of Voxelwise Second-Level fMRI Analysis
2.3. Voxelwise Bayesian Second-Level fMRI Analysis
2.4. Prior Determination Based on Results from Meta-Analyses
2.5. Performance Evaluation
2.5.1. Overlap Index for Evaluation
2.5.2. Statistical Analysis of Performance Outcomes
3. Results
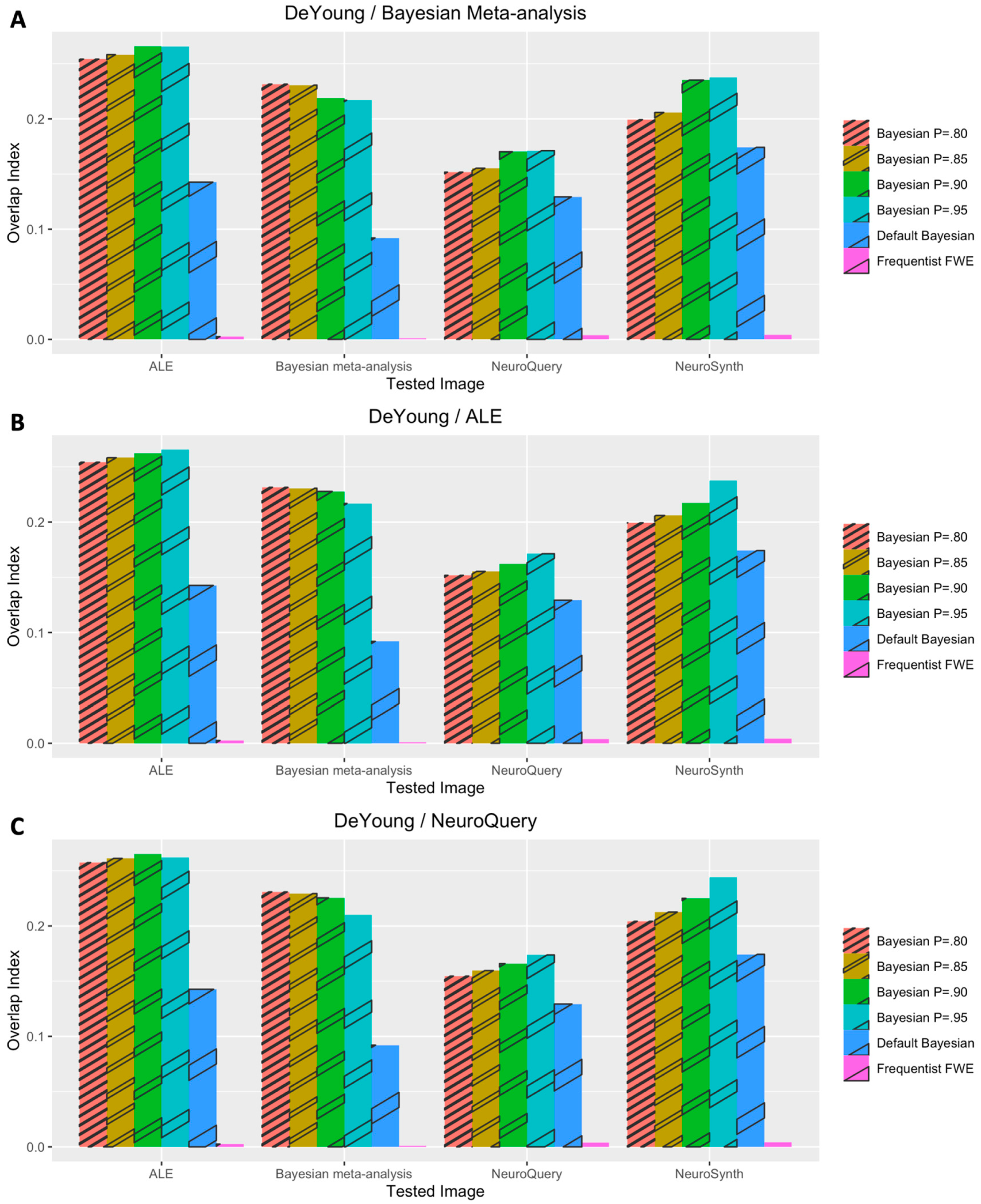
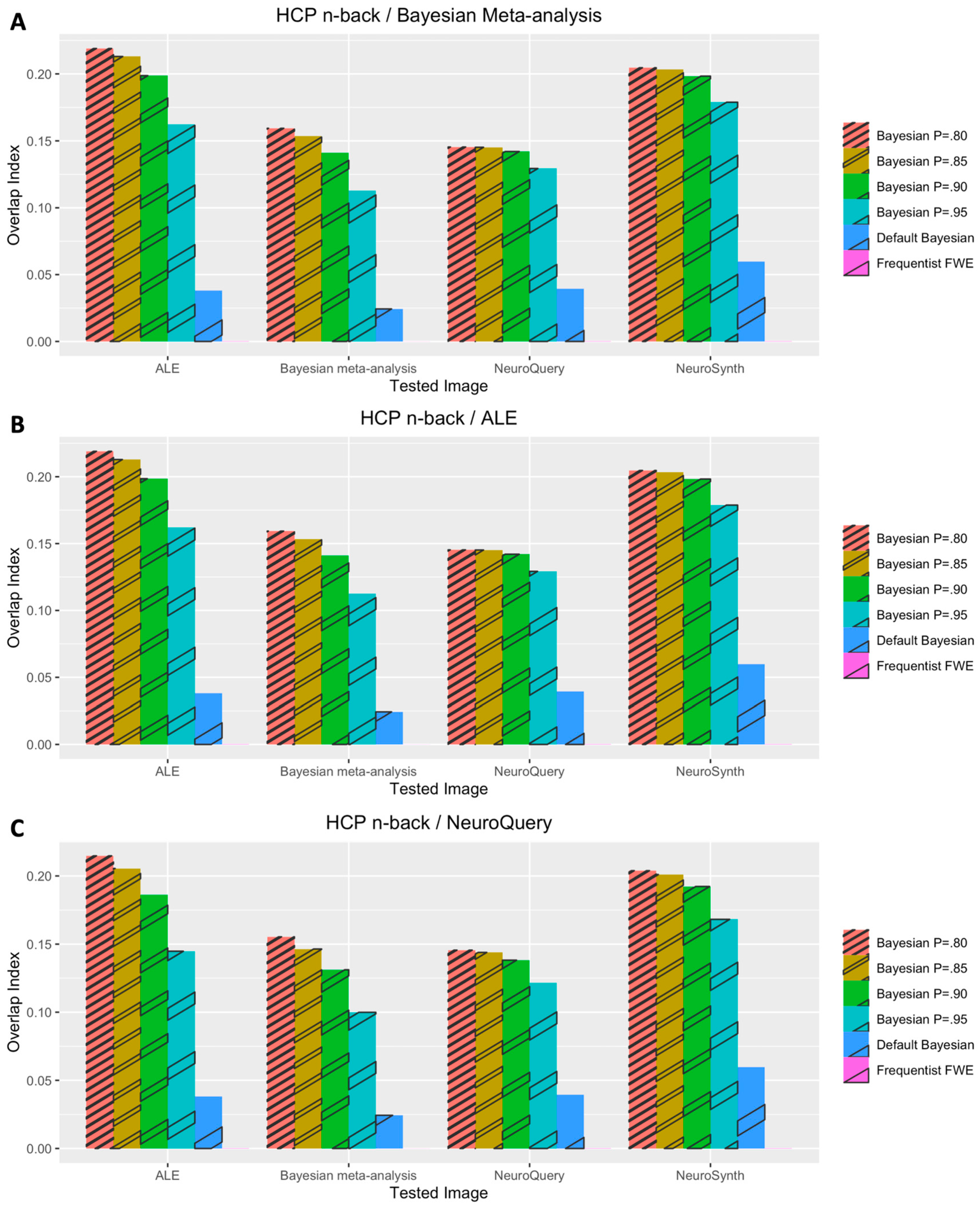
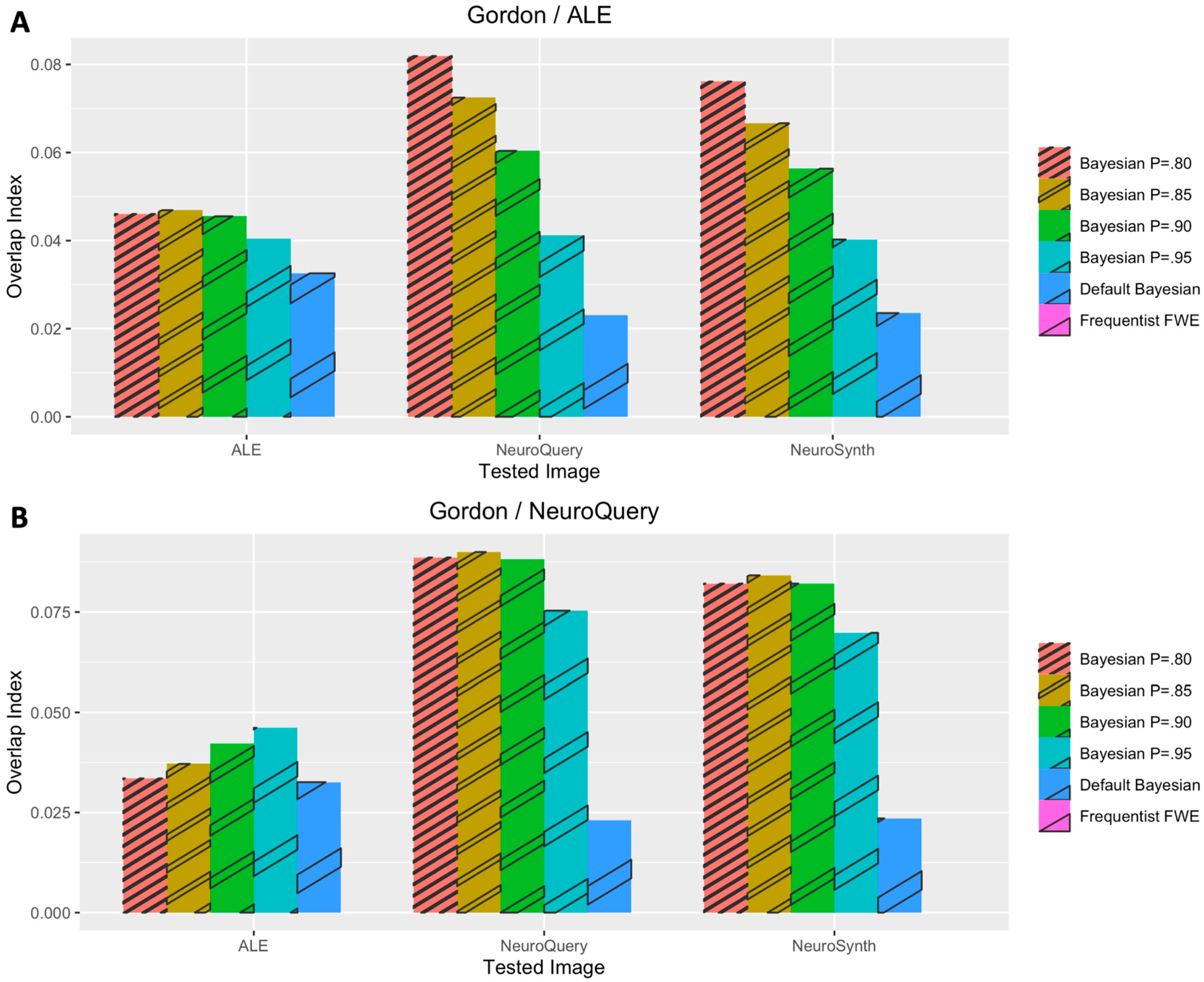
3.1. Voxelwise Second-Level fMRI Analyses
3.2. Statistical Analyses of Performance Outcomes
4. Discussion
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Bennett, C.M.; Miller, M.B.; Wolford, G.L. Neural Correlates of Interspecies Perspective Taking in the Post-Mortem Atlantic Salmon: An Argument for Multiple Comparisons Correction. NeuroImage 2009, 47, S125. [Google Scholar] [CrossRef] [Green Version]
- Eklund, A.; Nichols, T.E.; Knutsson, H. Cluster Failure: Why FMRI Inferences for Spatial Extent Have Inflated False-Positive Rates. Proc. Natl. Acad. Sci. USA 2016, 113, 7900–7905. [Google Scholar] [CrossRef] [Green Version]
- Mueller, K.; Lepsien, J.; Möller, H.E.; Lohmann, G. Commentary: Cluster Failure: Why FMRI Inferences for Spatial Extent Have Inflated False-Positive Rates. Front. Hum. Neurosci. 2017, 11, 345. [Google Scholar] [CrossRef]
- Nichols, T.E.; Eklund, A.; Knutsson, H. A Defense of Using Resting-State FMRI as Null Data for Estimating False Positive Rates. Cogn. Neurosci. 2017, 8, 144–149. [Google Scholar] [CrossRef] [Green Version]
- Cox, R.W.; Chen, G.; Glen, D.R.; Reynolds, R.C.; Taylor, P.A. FMRI Clustering in AFNI: False-Positive Rates Redux. Brain Connect. 2017, 7, 152–171. [Google Scholar] [CrossRef]
- Wagenmakers, E.-J.; Marsman, M.; Jamil, T.; Ly, A.; Verhagen, J.; Love, J.; Selker, R.; Gronau, Q.F.; Šmíra, M.; Epskamp, S.; et al. Bayesian Inference for Psychology. Part I: Theoretical Advantages and Practical Ramifications. Psychon. Bull. Rev. 2018, 25, 35–57. [Google Scholar] [CrossRef]
- Han, H.; Park, J.; Thoma, S.J. Why Do We Need to Employ Bayesian Statistics and How Can We Employ It in Studies of Moral Education?: With Practical Guidelines to Use JASP for Educators and Researchers. J. Moral Educ. 2018, 47, 519–537. [Google Scholar] [CrossRef] [Green Version]
- Wagenmakers, E.-J.; Love, J.; Marsman, M.; Jamil, T.; Ly, A.; Verhagen, J.; Selker, R.; Gronau, Q.F.; Dropmann, D.; Boutin, B.; et al. Bayesian Inference for Psychology. Part II: Example Applications with JASP. Psychon. Bull. Rev. 2018, 25, 58–76. [Google Scholar] [CrossRef] [Green Version]
- Rouder, J.N.; Speckman, P.L.; Sun, D.; Morey, R.D.; Iverson, G. Bayesian t Tests for Accepting and Rejecting the Null Hypothesis. Psychon. Bull. Rev. 2009, 16, 225–237. [Google Scholar] [CrossRef]
- Gelman, A.; Hill, J.; Yajima, M. Why We (Usually) Don’t Have to Worry about Multiple Comparisons. J. Res. Educ. Eff. 2012, 5, 189–211. [Google Scholar] [CrossRef] [Green Version]
- Woolrich, M.W. Bayesian Inference in FMRI. NeuroImage 2012, 62, 801–810. [Google Scholar] [CrossRef] [PubMed]
- Han, H.; Park, J. Using SPM 12’s Second-Level Bayesian Inference Procedure for FMRI Analysis: Practical Guidelines for End Users. Front. Neuroinform. 2018, 12, 1. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mejia, A.F.; Yue, Y.; Bolin, D.; Lindgren, F.; Lindquist, M.A. A Bayesian General Linear Modeling Approach to Cortical Surface FMRI Data Analysis. J. Am. Stat. Assoc. 2020, 115, 501–520. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Han, H. BayesFactorFMRI: Implementing Bayesian Second-Level FMRI Analysis with Multiple Comparison Correction and Bayesian Meta-Analysis of FMRI Images with Multiprocessing. J. Open Res. Softw. 2021, 9, 1. [Google Scholar] [CrossRef]
- Han, H. Implementation of Bayesian Multiple Comparison Correction in the Second-Level Analysis of FMRI Data: With Pilot Analyses of Simulation and Real FMRI Datasets Based on Voxelwise Inference. Cogn. Neurosci. 2020, 11, 157–169. [Google Scholar] [CrossRef]
- de Jong, T. A Bayesian Approach to the Correction for Multiplicity; The Society for the Improvement of Psychological Science: Charlottesville, VA, USA, 2019. [Google Scholar] [CrossRef]
- Westfall, P.H.; Johnson, W.O.; Utts, J.M. A Bayesian Perspective on the Bonferroni Adjustment. Biometrika 1997, 84, 419–427. [Google Scholar] [CrossRef] [Green Version]
- Liu, C.C.; Aitkin, M. Bayes Factors: Prior Sensitivity and Model Generalizability. J. Math. Psychol. 2008, 52, 362–375. [Google Scholar] [CrossRef]
- Sinharay, S.; Stern, H.S. On the Sensitivity of Bayes Factors to the Prior Distributions. Am. Stat. 2002, 56, 196–201. [Google Scholar] [CrossRef]
- Han, H. A Method to Adjust a Prior Distribution in Bayesian Second-Level FMRI Analysis. PeerJ 2021, 9, e10861. [Google Scholar] [CrossRef]
- Kruschke, J.K.; Liddell, T.M. Bayesian Data Analysis for Newcomers. Psychon. Bull. Rev. 2018, 25, 155–177. [Google Scholar] [CrossRef] [Green Version]
- van de Schoot, R.; Sijbrandij, M.; Depaoli, S.; Winter, S.D.; Olff, M.; van Loey, N.E. Bayesian PTSD-Trajectory Analysis with Informed Priors Based on a Systematic Literature Search and Expert Elicitation. Multivar. Behav. Res. 2018, 53, 267–291. [Google Scholar] [CrossRef] [Green Version]
- Avci, E. Using Informative Prior from Meta-Analysis in Bayesian Approach. J. Data Sci. 2017, 15, 575–588. [Google Scholar] [CrossRef]
- Zondervan-Zwijnenburg, M.; Peeters, M.; Depaoli, S.; van de Schoot, R. Where Do Priors Come From? Applying Guidelines to Construct Informative Priors in Small Sample Research. Res. Hum. Dev. 2017, 14, 305–320. [Google Scholar] [CrossRef] [Green Version]
- Han, H.; Park, J. Bayesian Meta-Analysis of FMRI Image Data. Cogn. Neurosci. 2019, 10, 66–76. [Google Scholar] [CrossRef] [PubMed]
- Salimi-Khorshidi, G.; Smith, S.M.; Keltner, J.R.; Wager, T.D.; Nichols, T.E. Meta-Analysis of Neuroimaging Data: A Comparison of Image-Based and Coordinate-Based Pooling of Studies. NeuroImage 2009, 45, 810–823. [Google Scholar] [CrossRef] [PubMed]
- Eickhoff, S.B.; Bzdok, D.; Laird, A.R.; Roski, C.; Caspers, S.; Zilles, K.; Fox, P.T. Co-Activation Patterns Distinguish Cortical Modules, Their Connectivity and Functional Differentiation. NeuroImage 2011, 57, 938–949. [Google Scholar] [CrossRef] [Green Version]
- Eickhoff, S.B.; Bzdok, D.; Laird, A.R.; Kurth, F.; Fox, P.T. Activation Likelihood Estimation Meta-Analysis Revisited. NeuroImage 2012, 59, 2349–2361. [Google Scholar] [CrossRef] [Green Version]
- Eickhoff, S.B.; Laird, A.R.; Grefkes, C.; Wang, L.E.; Zilles, K.; Fox, P.T. Coordinate-Based Activation Likelihood Estimation Meta-Analysis of Neuroimaging Data: A Random-Effects Approach Based on Empirical Estimates of Spatial Uncertainty. Hum. Brain Mapp. 2009, 30, 2907–2926. [Google Scholar] [CrossRef] [Green Version]
- Dockès, J.; Poldrack, R.A.; Primet, R.; Gözükan, H.; Yarkoni, T.; Suchanek, F.; Thirion, B.; Varoquaux, G. NeuroQuery, Comprehensive Meta-Analysis of Human Brain Mapping. eLife 2020, 9, e53385. [Google Scholar] [CrossRef]
- DeYoung, C.G.; Shamosh, N.A.; Green, A.E.; Braver, T.S.; Gray, J.R. Intellect as Distinct from Openness: Differences Revealed by FMRI of Working Memory. J. Personal. Soc. Psychol. 2009, 97, 883–892. [Google Scholar] [CrossRef] [Green Version]
- Henson, R.N.A.; Shallice, T.; Gorno-Tempini, M.L.; Dolan, R.J. Face Repetition Effects in Implicit and Explicit Memory Tests as Measured by FMRI. Cereb. Cortex 2002, 12, 178–186. [Google Scholar] [CrossRef]
- Kragel, P.A.; Kano, M.; van Oudenhove, L.; Ly, H.G.; Dupont, P.; Rubio, A.; Delon-Martin, C.; Bonaz, B.L.; Manuck, S.B.; Gianaros, P.J.; et al. Generalizable Representations of Pain, Cognitive Control, and Negative Emotion in Medial Frontal Cortex. Nat. Neurosci. 2018, 21, 283–289. [Google Scholar] [CrossRef]
- Pinho, A.L.; Amadon, A.; Gauthier, B.; Clairis, N.; Knops, A.; Genon, S.; Dohmatob, E.; Torre, J.J.; Ginisty, C.; Becuwe-Desmidt, S.; et al. Individual Brain Charting Dataset Extension, Second Release of High-Resolution FMRI Data for Cognitive Mapping. Sci. Data 2020, 7, 353. [Google Scholar] [CrossRef]
- Gordon, E.M.; Laumann, T.O.; Gilmore, A.W.; Newbold, D.J.; Greene, D.J.; Berg, J.J.; Ortega, M.; Hoyt-Drazen, C.; Gratton, C.; Sun, H.; et al. Precision Functional Mapping of Individual Human Brains. Neuron 2017, 95, 791–807.e7. [Google Scholar] [CrossRef] [Green Version]
- Laird, A.R.; Lancaster, J.L.; Fox, P.T. BrainMap: The Social Evolution of a Human Brain Mapping Database. Neuroinformatics 2005, 3, 65–78. [Google Scholar] [CrossRef]
- Turkeltaub, P.E.; Eickhoff, S.B.; Laird, A.R.; Fox, M.; Wiener, M.; Fox, P. Minimizing within-Experiment and within-Group Effects in Activation Likelihood Estimation Meta-Analyses. Hum. Brain Mapp. 2012, 33, 1–13. [Google Scholar] [CrossRef] [Green Version]
- Laird, A.R.; Fox, P.M.; Price, C.J.; Glahn, D.C.; Uecker, A.M.; Lancaster, J.L.; Turkeltaub, P.E.; Kochunov, P.; Fox, P.T. ALE Meta-Analysis: Controlling the False Discovery Rate and Performing Statistical Contrasts. Hum. Brain Mapp. 2005, 25, 155–164. [Google Scholar] [CrossRef]
- Yarkoni, T.; Poldrack, R.A.; Nichols, T.E.; van Essen, D.C.; Wager, T.D. Large-Scale Automated Synthesis of Human Functional Neuroimaging Data. Nat. Methods 2011, 8, 665–670. [Google Scholar] [CrossRef] [Green Version]
- Poldrack, R.A. Inferring Mental States from Neuroimaging Data: From Reverse Inference to Large-Scale Decoding. Neuron 2011, 72, 692–697. [Google Scholar] [CrossRef] [Green Version]
- Glymour, C.; Hanson, C. Reverse Inference in Neuropsychology. Br. J. Philos. Sci. 2016, 67, 1139–1153. [Google Scholar] [CrossRef]
- Dockès, J.; Poldrack, R.A.; Primet, R.; Gözükan, H.; Yarkoni, T.; Suchanek, F.; Thirion, B.; Varoquaux, G. About NeuroQuery. Available online: https://neuroquery.org/about (accessed on 13 January 2022).
- Kass, R.E.; Raftery, A.E. Bayes Factors. J. Am. Stat. Assoc. 1995, 90, 773–795. [Google Scholar] [CrossRef]
- Stefan, A.M.; Gronau, Q.F.; Schönbrodt, F.D.; Wagenmakers, E.-J. A Tutorial on Bayes Factor Design Analysis Using an Informed Prior. Behav. Res. Methods 2019, 51, 1042–1058. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Han, H. Neural Correlates of Moral Sensitivity and Moral Judgment Associated with Brain Circuitries of Selfhood: A Meta-Analysis. J. Moral Educ. 2017, 46, 97–113. [Google Scholar] [CrossRef]
- Cremers, H.R.; Wager, T.D.; Yarkoni, T. The Relation between Statistical Power and Inference in FMRI. PLoS ONE 2017, 12, e0184923. [Google Scholar] [CrossRef] [Green Version]
- Han, H.; Glenn, A.L. Evaluating Methods of Correcting for Multiple Comparisons Implemented in SPM12 in Social Neuroscience FMRI Studies: An Example from Moral Psychology. Soc. Neurosci. 2018, 13, 257–267. [Google Scholar] [CrossRef] [Green Version]
- Han, H.; Glenn, A.L.; Dawson, K.J. Evaluating Alternative Correction Methods for Multiple Comparison in Functional Neuroimaging Research. Brain Sci. 2019, 9, 198. [Google Scholar] [CrossRef] [Green Version]
- Ashburner, J.; Barnes, G.; Chen, C.-C.; Daunizeau, J.; Flandin, G.; Friston, K.; Kiebel, S.; Kilner, J.; Litvak, V.; Moran, R.; et al. SPM 12 Manual; Wellcome Trust Centre for Neuroimaging: London, UK, 2016. [Google Scholar]
- Han, H.; Dawson, K.J. Improved Model Exploration for the Relationship between Moral Foundations and Moral Judgment Development Using Bayesian Model Averaging. J. Moral Educ. 2021, 1–5. [Google Scholar] [CrossRef]
- Han, H. Exploring the Association between Compliance with Measures to Prevent the Spread of COVID-19 and Big Five Traits with Bayesian Generalized Linear Model. Personal. Individ. Differ. 2021, 176, 110787. [Google Scholar] [CrossRef]
- Gorgolewski, K.J.; Varoquaux, G.; Rivera, G.; Schwarz, Y.; Ghosh, S.S.; Maumet, C.; Sochat, V.V.; Nichols, T.E.; Poldrack, R.A.; Poline, J.-B.; et al. NeuroVault.Org: A Web-Based Repository for Collecting and Sharing Unthresholded Statistical Maps of the Human Brain. Front. Neuroinform. 2015, 9, 8. [Google Scholar] [CrossRef] [Green Version]
- Laird, A.R.; Eickhoff, S.B.; Fox, P.M.; Uecker, A.M.; Ray, K.L.; Saenz, J.J.; McKay, D.R.; Bzdok, D.; Laird, R.W.; Robinson, J.L.; et al. The BrainMap Strategy for Standardization, Sharing, and Meta-Analysis of Neuroimaging Data. BMC Res. Notes 2011, 4, 349. [Google Scholar] [CrossRef] [Green Version]
- Poldrack, R.A. Can Cognitive Processes Be Inferred from Neuroimaging Data? Trends Cogn. Sci. 2006, 10, 59–63. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ly, A.; Stefan, A.; van Doorn, J.; Dablander, F.; van den Bergh, D.; Sarafoglou, A.; Kucharský, S.; Derks, K.; Gronau, Q.F.; Raj, A.; et al. The Bayesian Methodology of Sir Harold Jeffreys as a Practical Alternative to the P Value Hypothesis Test. Comput. Brain Behav. 2020, 3, 153–161. [Google Scholar] [CrossRef] [Green Version]



| Category | Dataset Name | Sample Size | Compared Task Conditions | Link to Open Repository |
|---|---|---|---|---|
| Working memory | DeYoung et al. (2009) (in Kragel et al.’s (2018) repository) | 15 | 3-back vs. fixation | https://neurovault.org/collections/3324/ CBM (accessed on 13 January 2022) |
| Henson et al. (2002) | 12 | Famous vs. non-famous face memory | http://www.fil.ion.ucl.ac.uk/spm/download/data/face_rfx/face_rfx.zip (accessed on 13 January 2022) (under “cons_can” subfolder) | |
| Pinho et al. (2020) | 13 | 2-back vs. 0-back | https://neurovault.org/collections/6618/ (accessed on 13 January 2022) | |
| Speech | Pinho et al. (2020) | 18 | Speech vs. natural sound listening | https://neurovault.org/collections/2138/ (accessed on 13 January 2022) |
| Face | Gordon et al. (2017) | 10 | Face vs.word identification | https://neurovault.org/collections/2447/ (accessed on 13 January 2022) |
| Category | Type | Acquisition Method * |
|---|---|---|
| Working memory | Bayesian Meta-analysis | Han and Park’s (2018) meta-analysis Acquired from Han (2021) GitHub: https://github.com/hyemin-han/Prior-Adjustment-BayesFactorFMRI/tree/master/Working_memory_fMRI/Performance_evaluation (accessed on 13 January 2022) |
| BrainMap + Ginger ALE | Slueth: Normal Mapping & Activations Only & Paradigm Class = n-back Ginger ALE: cluster forming p < 0.001, cluster-level FWE p < 0.01 | |
| NeuroSynth | term = “working memory” https://neurosynth.org/analyses/terms/working%20memory/ (accessed on 13 January 2022) | |
| NeuroQuery | term = “working memory” https://neuroquery.org/query?text=working+memory+ (accessed on 13 January 2022) | |
| Speech | BrainMap + Ginger ALE | Slueth: Normal Mapping & Activations Only & Keywords = face | faces | face recognition | facial recognition Ginger ALE: cluster forming p < 0.001, cluster-level FWE p < 0.01 |
| NeuroSynth | term = “speech” https://neurosynth.org/analyses/terms/speech/ (accessed on 13 January 2022) | |
| NeuroQuery | term = “speech” https://neuroquery.org/query?text=speech+ (accessed on 13 January 2022) | |
| Face | BrainMap + Ginger ALE | Slueth: Normal Mapping & Activations Only & Keywords = speaking | Speech | speech | speech processing Ginger ALE: cluster forming p < 0.001, cluster-level FWE p < 0.01 |
| NeuroSynth | term = “face” https://neurosynth.org/analyses/terms/face/ (accessed on 13 January 2022) | |
| NeuroQuery | term = “face” https://neuroquery.org/query?text=face+ (accessed on 13 January 2022) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Han, H. A Novel Method to Use Coordinate Based Meta-Analysis to Determine a Prior Distribution for Voxelwise Bayesian Second-Level fMRI Analysis. Mathematics 2022, 10, 356. https://doi.org/10.3390/math10030356
Han H. A Novel Method to Use Coordinate Based Meta-Analysis to Determine a Prior Distribution for Voxelwise Bayesian Second-Level fMRI Analysis. Mathematics. 2022; 10(3):356. https://doi.org/10.3390/math10030356
Chicago/Turabian StyleHan, Hyemin. 2022. "A Novel Method to Use Coordinate Based Meta-Analysis to Determine a Prior Distribution for Voxelwise Bayesian Second-Level fMRI Analysis" Mathematics 10, no. 3: 356. https://doi.org/10.3390/math10030356
APA StyleHan, H. (2022). A Novel Method to Use Coordinate Based Meta-Analysis to Determine a Prior Distribution for Voxelwise Bayesian Second-Level fMRI Analysis. Mathematics, 10(3), 356. https://doi.org/10.3390/math10030356






