Abstract
Chest X-ray (CXR) imaging remains the most widely used radiological modality for assessing pulmonary and cardiothoracic disease, yet its interpretation is inherently constrained by tissue superposition, subtle radiographic findings and marked inter-observer variability. Recent advances in artificial intelligence (AI) have driven significant progress in automated CXR analysis, supported by large public datasets, evolving annotation strategies and increasingly expressive deep learning architectures. This review presents a comprehensive synthesis of approaches for pulmonary abnormality detection, encompassing convolutional neural networks, transformers, multimodal and vision–language models and self-supervised representation learning. We critically discuss their strengths, limitations and vulnerability to label noise, domain shift and shortcut learning. In parallel, we examine dataset properties, annotation practices, robustness challenges, explainability methods and the heterogeneity of evaluation protocols that hinder fair comparison and clinical translation. Building on these observations, the review identifies key future directions, including foundation models, multimodal integration, federated and domain-generalized training, longitudinal modeling, synthetic data generation and standardized clinical evaluation frameworks. By integrating methodological and clinical perspectives, this work offers an up-to-date reference for researchers and clinicians and outlines a roadmap toward reliable, interpretable and clinically deployable AI systems for chest radiography.
1. Introduction
Chest X-ray imaging is the most widely used radiological examination worldwide. It is the first-line diagnostic tool for evaluating pulmonary and cardiothoracic abnormalities due to its low cost, portability, rapid acquisition and relatively low radiation exposure [1,2]. It is indispensable in emergency departments, primary care, intensive care units and resource-limited settings. Chest radiography is routinely performed as the first-line imaging modality in patients presenting with acute dyspnea, persistent fever suspicious for pneumonia, oncologic staging to assess pulmonary metastases and as part of preoperative evaluation to ensure anesthetic safety. Its ubiquity across emergency, inpatient and outpatient settings further reinforces its central role in clinical decision-making.
However, CXR interpretation is challenging even for experienced radiologists. The projection of 3D chest anatomy onto a 2D X-ray image may result in tissue superposition, which can mask subtle findings, reduce sensitivity for early disease and contribute to marked inter-observer variability [3,4]. Diagnostic performance is further influenced by reader expertise, image quality, workload and availability of clinical context—factors that are often suboptimal in real-world practice [2,5].
In this context, artificial intelligence has emerged as a promising solution for automated CXR interpretation. Over the past decade, some deep learning models such as transformer-based architectures [6,7] and multimodal models have achieved high performance [8] across key tasks such as disease classification, finding localization, severity scoring and triage [9,10,11]. These advances have been catalyzed by the release of large public datasets such as ChestX-ray14 [12], CheXpert [13], MIMIC-CXR [14], VinDr-CXR [15] and PadChest [16], which provide training corpora of unprecedented size and heterogeneity.
Modern deep learning models can detect a broad spectrum of pulmonary abnormalities [4,9,17], including consolidation, interstitial opacities, atelectasis, pneumothorax, pleural effusion, edema, cardiomegaly and nodules. Vision–language models (VLMs) and self-supervised learning (SSL) strategies have further pushed the field forward by reducing dependence on manually curated labels and enabling broader generalization [11,18,19,20,21]. Despite this rapid algorithmic progress, the integration of AI systems into real-world CXR workflows remains limited. The role of AI in chest radiography should be considered less as a substitute for radiologist interpretation and more as an integrative support layer incorporated into clinical workflows, especially in areas with fewer resources where family doctors are responsible for evaluating radiographs.. Several challenges persist, including label noise inherent to report-derived annotations [13], domain shift across hospitals, scanners and patient populations [22,23] and confounding artifacts such as tubes, devices and text markers that introduce spurious correlations [3]. Many CXR abnormalities are subtle or ambiguous and heavily dependent on clinical context, which complicates fully automated interpretation. Moreover, the interpretability of deep learning predictions remains constrained, limiting trust and acceptance among clinicians [24].
Recent surveys have highlighted the fragmented nature of current research, where architectural advances, dataset characteristics, annotation quality and clinical deployment considerations are often addressed in isolation [25,26,27,28]. However, these works typically emphasize performance trends or application outcomes, without systematically analyzing how methodological choices and data-centric factors jointly shape robustness, generalization and clinical reliability.
Persistent challenges such as domain shift and cross-population generalization continue to limit real-world translation [29], while recent studies underline the critical role of annotation fidelity and pixel-level supervision for diagnostic robustness [30]. Nevertheless, a unified synthesis that connects explainability, model interpretability, dataset quality and clinical validation within a single methodological framework remains largely absent.
Although several reviews on AI for medical imaging and chest radiography have been published [31,32,33], most of them adopt a broad or application-driven perspective, focusing on general imaging pipelines, specific disease categories or high-level clinical use cases. In contrast, the work shows a methodologically oriented and problem-centric perspective, focusing specifically on pulmonary abnormality detection in chest X-ray imaging and on the interactions between dataset design, labeling strategies, learning paradigms and evaluation practices in order to integrate in the CXR workflows to improve the triage and decision-making. For that purpose, this review addresses this gap by explicitly integrating methodological, data-centric and clinical perspectives, moving beyond a catalog of architectures to analyze recurrent failure modes, evaluation inconsistencies and translational bottlenecks. It provides a structured roadmap for developing, evaluating and deploying more reliable and clinically aligned CXR-AI systems [34,35,36]. This review addresses three complementary and interrelated objectives. First, it provides a comprehensive and technically grounded overview of deep learning methods for pulmonary abnormality detection in chest radiographs, covering convolutional neural networks, transformer-based architectures, multimodal and vision–language models, as well as explainability techniques. Unlike prior surveys that primarily summarize architectural performance, this review emphasizes the methodological assumptions and limitations underlying these approaches. Second, it systematically analyzes major public CXR datasets and labeling methodologies, highlighting their strengths, weaknesses, sources of bias and downstream implications for model training, evaluation and generalization. Particular attention is paid to the impact of report-derived annotations, uncertainty handling and annotation granularity on algorithmic reliability. Third, the review discusses open challenges and emerging research directions that are critical for clinical translation, including domain generalization, interpretability, data quality, self-supervised learning, multimodal integration and the growing role of vision–language models in radiology.
By synthesizing methodological advances with clinical considerations, this work aims to serve as an up-to-date reference for researchers, clinicians and developers working on automated CXR interpretation, while also providing a methodological roadmap for the development and deployment of robust, trustworthy AI systems in real clinical environments. The review explicitly connects dataset design, labeling noise, learning paradigms and evaluation practices, highlighting how their interactions shape reported performance and clinical readiness. Rather than cataloging models in isolation, we focus on recurring methodological trade-offs and failure modes that continue to limit real-world translation.
2. Materials and Methods
2.1. Review Design and Methodological Positioning
This work was conducted as a structured narrative review [37] informed by a systematic literature search strategy. The objective was not to perform a quantitative meta-analysis or formal evidence grading, but to provide a critical methodological synthesis of artificial intelligence approaches for chest X-ray analysis. Elements of the Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA) 2020 guidelines [38] statement were adopted to enhance transparency in study identification, screening and reporting; however, the review does not constitute a full systematic review with prospective protocol registration or quantitative risk-of-bias scoring. The scope of the review encompasses methodological contributions, large-scale public datasets, clinical validation studies and recent advances in deep learning, self-supervised learning, multimodal models and VLMs for CXR analysis. The review protocol was not prospectively registered. A comprehensive literature search was independently performed by two reviewers using PubMed, Scopus and Web of Science (WoS) to identify studies evaluating artificial intelligence pipelines for chest X-ray interpretation.
The PubMed strategy combined both MeSH terms and free-text keywords related to chest radiographs and AI techniques: (“Chest X-Ray” [Mesh] OR “chest x-ray” OR “chest radiograph*” OR CXR) AND (“Artificial Intelligence” [Mesh] OR “artificial intelligence” OR “machine learning” OR “deep learning” OR “vision transformer*” OR “self-supervised learning” OR “computer-aided diagnosis” OR CAD). The Scopus query was constructed as follows: TITLE-ABS-KEY(“chest x-ray” OR “chest radiograph*” OR CXR) AND TITLE-ABS-KEY(“artificial intelligence” OR “deep learning” OR “machine learning” OR “vision transformer*” OR “self-supervised learning” OR “computer-aided diagnosis” OR CAD).
The Web of Science search used a Topic (TS) query: TS=(“chest x-ray” OR “chest radiograph*” OR CXR) AND TS=(“artificial intelligence” OR “deep learning” OR “machine learning” OR “vision transformer*” OR “self-supervised learning” OR “computer-aided diagnosis” OR CAD).
2.2. Data Extraction and Analysis
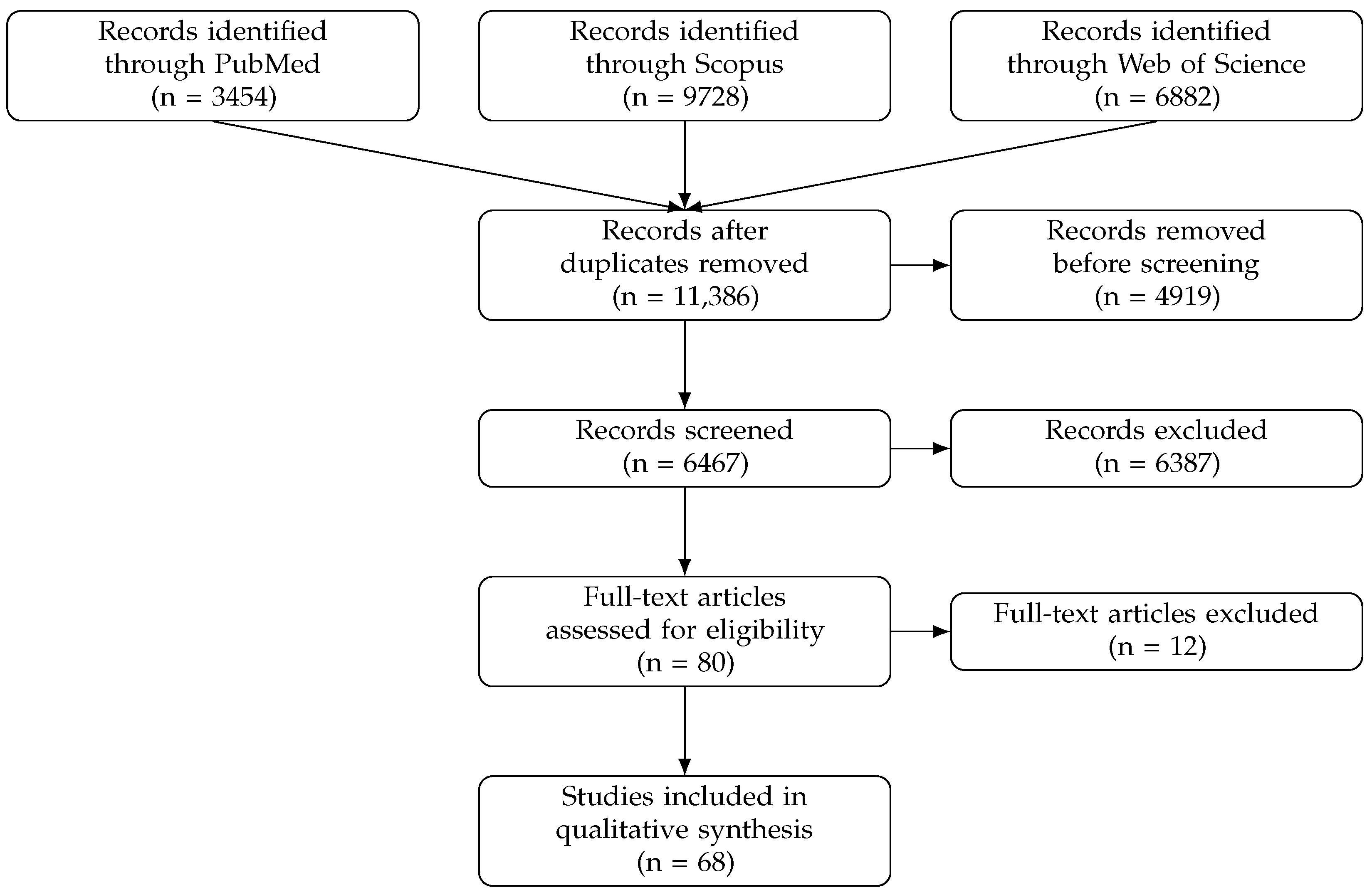
The searches retrieved 3454 records from PubMed, 9728 from Scopus and 6882 from Web of Science. No language restrictions were applied at the search stage and studies published between 2013 and 2025 were considered. To enhance comprehensiveness, the reference lists of all included articles were manually screened. Overall inter-reviewer agreement during the study selection process was 91% across title–abstract screening and full-text eligibility assessment. Disagreements between reviewers were primarily related to borderline cases, including studies combining multiple imaging modalities, papers with limited methodological description of the AI component or investigations focusing predominantly on prognostic rather than diagnostic tasks. All discrepancies were resolved through discussion and consensus and, when necessary, adjudicated by the senior author.
After duplicate records were removed, studies were excluded if they consisted of conference papers, books or book chapters, meta-analyses, editorials, letters, comments or meeting abstracts or lacked full-text availability in English. Additional exclusions applied to studies published before 2013, as well as to animal studies or experimental investigations based on physical phantoms. The restriction to peer-reviewed journal articles was adopted to ensure comprehensive methodological reporting and extended validation analyses, which are particularly relevant for assessing translational readiness in medical imaging. We acknowledge that influential methodological contributions in artificial intelligence are often published in conference proceedings and their exclusion may limit coverage of early-stage architectural innovations.
The remaining records were subsequently screened based on title and abstract. Studies were considered eligible if they evaluated the performance of AI-based models for detection, classification or segmentation tasks using primary chest radiographs. Inclusion was restricted to original research articles published in peer-reviewed journals with full-text availability in English. This language restriction was applied to ensure consistent methodological assessment and interpretability; however, it may introduce language bias and limit representation of research conducted in non-English-speaking contexts.
During full-text eligibility assessment, studies primarily focused on prognosis, biomarkers, non-imaging clinical data, alternative imaging modalities or lacking a clearly described AI methodology were excluded. The study selection process was summarized using a PRISMA flow diagram (Figure 1) [38].
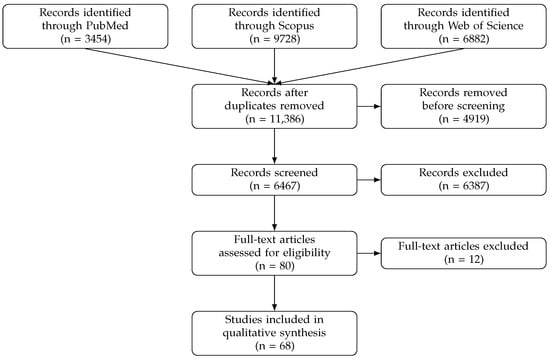
Figure 1.
PRISMA flow diagram of the study selection process.
2.3. Critical Appraisal and Content Synthesis
The methodological quality and potential sources of bias of the included studies were assessed using the Checklist for Artificial Intelligence in Medical Imaging (CLAIM) [39]. Each study was independently evaluated by two reviewers with respect to dataset characterization, annotation strategies, model architecture, training and validation procedures, performance reporting and reproducibility considerations.
CLAIM was employed as a qualitative appraisal framework, consistent with its original design and not used to derive a quantitative quality score or to exclude studies. Instead, its items served as an analytic lens to guide a structured and transparent assessment of reporting completeness and methodological rigor, enabling the identification of recurrent reporting gaps, common sources of bias and cross-study methodological patterns across the included literature [39,40]. The purpose of this appraisal was not to rank studies, but to characterize methodological strengths and limitations that directly informed the subsequent qualitative synthesis and interpretation of results.
Following study inclusion, a structured qualitative synthesis was conducted to integrate findings across the selected articles [37]. For each study, key methodological attributes were systematically extracted, including dataset composition and provenance, labeling and annotation strategies, learning paradigms, architectural design choices, evaluation protocols and reported clinical validation settings.
Content synthesis was performed using an inductive thematic analysis approach [41]. Studies were grouped according to recurring methodological themes, including dataset-centric challenges, supervision and labeling strategies, architectural trends, robustness to domain shift and evaluation practices. These themes formed the basis for the comparative analysis presented in Section 3, Section 4, Section 5 and Section 6, allowing methodological trade-offs and translational bottlenecks to be examined across heterogeneous study designs.
Application of the CLAIM framework revealed substantial variability in reporting practices and methodological rigor. While most articles provided clear descriptions of model architectures and reported conventional performance metrics, critical aspects related to data provenance, annotation quality and external validation were addressed inconsistently. In particular, detailed characterization of labeling strategies and annotation uncertainty was often limited in studies relying on report-derived labels, whereas works based on expert-annotated datasets tended to provide more transparent descriptions of annotation procedures. External validation across independent institutions was relatively uncommon, with the majority of studies relying on single-center or dataset-specific test splits. Similarly, calibration analysis, uncertainty estimation and assessment of potential confounders were rarely reported, despite their relevance for clinical deployment.
These observations, derived from CLAIM-guided qualitative appraisal rather than quantitative scoring, informed the thematic synthesis by highlighting recurrent methodological gaps that constrain reproducibility, generalization and clinical translation. As a result, the synthesis emphasizes not only reported performance, but also the methodological context in which such results should be interpreted, in line with best practices for qualitative synthesis in methodologically diverse and rapidly evolving research fields [37].
3. Clinical Background and Radiological Taxonomy
CXR imaging remains the most widely used radiological modality and continues to play a central role for assessing pulmonary and cardiothoracic conditions. Among its advantages are its low cost, wide availability, portability and low radiation exposure which make it indispensable in a broad range of clinical contexts. At the same time, the intrinsic limitation of projecting complex three-dimensional anatomy onto a two-dimensional image introduces tissue overlap and structural superposition. Subtle findings may therefore be partially obscured, leading to reduced sensitivity. In this setting, diagnostic performance can be further affected by inter-observer variability and by the cognitive burden imposed by increasing workloads, even among experienced radiologists. These challenges have driven growing interest in AI as a complementary tool for chest radiograph interpretation [1,3].
The availability of large public CXR datasets, such as ChestX-ray14, CheXpert, MIMIC-CXR, PadChest and VinDr-CXR, has fundamentally shaped the development of data-driven approaches in this field [12,13,14,15,16]. Although these resources differ substantially in scale, annotation strategy and label granularity, they converge on a core set of thoracic findings. Some of the most common findings are: consolidation, atelectasis, pleural effusion, pneumothorax, cardiomegaly, pulmonary edema and interstitial opacities. A clear understanding of these entities and their radiographic appearance is essential, not only for clinical interpretation, but also for the design, training and evaluation of AI models that aim to operate reliably in real-world settings.
Consolidation typically appears as a homogeneous increase in lung parenchymal opacity resulting from alveolar filling and is most often associated with pneumonia or acute lung injury. In practice, however, its radiographic appearance frequently overlaps with that of atelectasis, which also presents as an area of increased opacity but is accompanied by volume loss. This visual similarity remains a well-recognized source of diagnostic ambiguity, affecting both clinical interpretation and automated deep learning systems [4,9,13,42].
Pleural effusion, in contrast, is characterized by a more distinctive basal meniscus-shaped opacity. Owing to its relatively table and recognizable visual features, it consistently emerges as one of the best-performing categories in chest X-ray classification tasks, even in early deep learning studies [4,12]. Pneumothorax represents a different type of challenge. Defined by the presence of air in the pleural space, its radiographic signs can be subtle and highly dependent on patient positioning, particularly in supine acquisitions. These factors have made pneumothorax detection a prominent target for automatic triage and prioritization systems [2].
Cardiomegaly is commonly assessed using the cardiothoracic ratio, with values exceeding 0.5 typically considered abnormal. While structurally straightforward as a concept, its interpretation is highly sensitive to acquisition geometry. Differences between the projections, as well as patient positioning, can substantially alter apparent heart size and introduce systematic errors in both human and algorithmic assessments [43]. Pulmonary edema presents a different level of complexity. Its radiographic manifestations range from subtle vascular redistribution and Kerley lines to more conspicuous perihilar “bat-wing” patterns, often requiring contextual and temporal reasoning. Recent multimodal and vision–language approaches, which integrate image features with textual information, have begun to address this inherent ambiguity [11,18].
Diffuse interstitial abnormalities are particularly challenging to identify on chest radiographs. Their low contrast and frequent overlap with normal anatomical structures contribute to high inter-observer variability and inconsistent labeling [3,44]. Pulmonary nodules and masses pose similar difficulties, especially when small or poorly contrasted and remain considerably harder to detect on CXR than on computed tomography [10,45]. To mitigate these limitations, more recent algorithms have explored the incorporation of bone suppression techniques or prior anatomical segmentation as a means to improve finding-level sensitivity.
Finally, the presence of medical devices, such as endotracheal tubes, central venous catheters or pacemakers, introduces an additional source of visual confounding in chest radiographs. These elements are frequently correlated with disease severity or critical illness and deep learning models may inadvertently rely on them as shortcuts rather than on true pathological patterns. As a result, spurious associations can emerge, significantly impairing model generalization when applied across institutions, patient populations or acquisition settings [22,46,47].
Anatomical variability, projection-related artifacts and the influence of non-pathological visual cues illustrate the intrinsic complexity of CXR interpretation. This complexity challenges purely image-level classification approaches and reinforces the need for AI systems capable of integrating spatial relationships, contextual information and semantic reasoning. Approaches that explicitly model dependencies between findings or incorporate higher-level contextual understanding appear better suited to address these limitations [3,48].
These intrinsic differences in visual ambiguity, anatomical variability and contextual dependence are not merely descriptive radiological observations; they directly shape algorithmic behavior, influencing sensitivity to label noise, supervision requirements and the choice of architectural paradigms.
4. Datasets and Data Sources
The widespread adoption of deep learning for CXR interpretation has been closely tied to the emergence of large-scale public datasets. Some of the most commonly used are ChestX-ray14 [12], CheXpert [13], MIMIC-CXR [14], PadChest [16] and VinDr-CXR [15]. These datasets differ substantially in terms of patient population, acquisition protocols and labeling strategies. Such differences are often treated as secondary considerations, yet they have a direct and sometimes underappreciated impact on algorithm performance and generalizability [49,50]. An overview of the main characteristics of these datasets, with a particular focus on labeling strategies and data quality aspects, is provided in Table 1.

Table 1.
Overview of major public chest X-ray datasets, highlighting labeling strategies, annotation granularity, diagnostic confirmation level and data quality considerations relevant to deep learning–based CXR analysis.
Early efforts, most notably ChestX-ray14, relied on rule-based natural language processing (NLP) applied to radiology reports to automatically extract labels. This approach enabled an unprecedented scale but came at the cost of substantial label noise [51,52]. Subsequent datasets introduced refinements rather than abandoning automation altogether. As summarized in Table 1, CheXpert incorporated uncertainty-aware labeling to better reflect the ambiguity inherent in clinical reports, while MIMIC-CXR paired images with free-text reports, enabling multimodal learning and self-supervised representation learning paradigms. PadChest shifted the focus toward expert-verified hierarchical annotations, prioritizing label reliability and VinDr-CXR further extended supervision by providing finding-level bounding boxes that support detection and localization tasks.
Each of these design choices, whether related to label provenance, annotation granularity or supervision level, inevitably shapes how models are trained, evaluated and compared across studies. Across datasets, differences in label provenance, uncertainty modeling and annotation granularity directly influence robustness to shortcut learning and cross-institution generalization, reinforcing the need for careful dataset-aware interpretation of reported performance. Therefore, automated labeling pipelines should be interpreted with caution. Their contribution to initiating research is very valuable, but they are also prone to systematic errors, including false positives driven by historical mentions, missed subtle findings and the pervasive use of ambiguous categories such as “opacity” or “infiltrate”. As reflected by the heterogeneous uncertainty handling strategies reported in Table 1, the treatment of ambiguity varies widely across datasets, further complicating cross-study comparisons. In response, a growing body of work has explored weak supervision and attention-based approaches that attempt to infer spatial evidence from coarse image-level labels, sometimes leveraging text–image alignment to partially compensate for annotation noise.
Beyond labeling itself, dataset composition plays a decisive role in shaping model behavior. Systems trained predominantly on NLP-derived annotations may internalize institution-specific artifacts, resulting in overfitting to dataset biases rather than learning transferable pathological patterns. Common findings, such as pleural effusion or cardiomegaly, often achieve higher reported accuracy due to consistent labeling and relatively clear visual signatures. In contrast, rarer or more subjective conditions, including interstitial disease or fibrosis, remain considerably more challenging. The lack of standardized taxonomies and harmonized labeling schemes across major public datasets (Table 1) highlights the need for multi-source training strategies and unified annotation frameworks if reproducibility and cross-domain robustness are to be achieved [50].
To enable more structured comparison, major public datasets are contrasted in terms of label provenance (NLP-derived vs. expert), annotation granularity (image-level vs. spatial), uncertainty modeling, scale and expected implications for robustness, bias and external validity. It is important to note that most publicly available CXR datasets were constructed to benchmark disease classification performance rather than to model real-world radiology workflows. As a result, tasks such as longitudinal comparison, contextual reasoning across patient records, structured reporting or triage prioritization remain largely unsupported by current dataset design.
Table 1 summarizes the most widely adopted publicly available CXR datasets in deep learning research. Datasets were selected based on three criteria: (i) public accessibility, (ii) scale greater than approximately 10,000 radiographs and (iii) documented use in benchmarking or multi-study comparisons within the CXR-AI literature.
Terminology clarification: In this review, the following pragmatic definitions are adopted for consistency across studies: (i) Limited labeled data: fewer than approximately 5000 annotated radiographs; (ii) Moderate-to-large datasets: 5000–50,000 studies; (iii) Large-scale datasets: more than 50,000 studies; (iv) Weakly labeled data: image-level labels derived from radiology reports or indirect supervision without pixel-level confirmation or systematic cross-modality verification; (v) Local context: spatially contiguous image features occupying a restricted anatomical region (e.g., focal nodules or segmental opacities); (vi) Global context: diffuse, bilateral or multi-region patterns requiring modeling of long-range spatial dependencies across the thoracic cavity.
5. AI Methods for Chest X-Ray Analysis
To facilitate cross-study comparison, architectures and learning paradigms are analyzed across five recurring dimensions: (i) supervision level and annotation reliability, (ii) data requirements and scalability, (iii) robustness to domain shift and shortcut learning, (iv) interpretability and localization capacity and (v) clinical validation maturity. This structured lens allows methodological trade-offs to be examined beyond isolated performance metrics. Most published research focuses on detection-centered AI systems, which aim to replicate the identification of anomalies at the level of a radiologist. Studies on workflow improvement systems that aim to improve clinical efficiency are limited.
Table 2 operationalizes the analytical dimensions introduced above and highlights that no single paradigm is universally superior; instead, performance, robustness and translational potential are strongly contingent on supervision quality, dataset characteristics and intended clinical context.

Table 2.
Structured comparative framework of major AI paradigms for chest X-ray analysis across five analytical dimensions defined in this review.
Research on automated chest X-ray interpretation has expanded rapidly following the release of large-scale public datasets. These resources enabled the training of increasingly sophisticated deep learning systems capable of addressing a wide spectrum of thoracic abnormalities. Rather than treating architectural choices, learning paradigms and supervision strategies in isolation, this section provides an integrated overview of how these components have co-evolved in CXR AI systems.
Table 3 provides a comparative overview of the main AI paradigms adopted for chest X-ray analysis, summarizing their architectural choices, supervision strategies, strengths and limitations.

Table 3.
Comparative overview of major AI paradigms for chest X-ray analysis, highlighting architectural choices, supervision strategies, strengths and key limitations.
5.1. Deep Learning Architectures
Early work in CXR analysis relied almost exclusively on convolutional neural networks, typically pre-trained on ImageNet and fine-tuned for multi-label thoracic disease prediction. As summarized in the first row of Table 3, architectures such as ResNet, DenseNet, VGG and Inception quickly became standard baselines due to their strong performance on common findings and efficient optimization. The ChestX-ray14 benchmark established early performance references for weak supervision classification and localization using CNNs [12], while CheXNeXt demonstrated that a DenseNet-121 backbone could approach radiologist-level performance for several common findings [9]. Subsequent comparative studies explored the impact of backbone choice, transfer-learning strategies and data augmentation, highlighting the strong influence of pretraining schemes and class imbalance on reported performance [4]. In this context, class imbalance typically refers to prevalence ratios exceeding approximately 1:3 or 1:10 between positive and negative samples for a given pathology, with several thoracic findings occurring in fewer than 1–2% of cases in large public benchmarks.
Representative large-scale CNN baselines have reported strong discrimination performance on common thoracic findings, frequently achieving high AUROC values for conditions such as pleural effusion and cardiomegaly on established public benchmarks. However, these results vary substantially depending on label provenance, class prevalence and the rigor of external validation protocols.
As research progressed, architectural refinements began to address some of the limitations of purely convolutional designs. Attention mechanisms and multi-scale feature extraction improved sensitivity to subtle and spatially heterogeneous abnormalities, including pneumothorax and atelectasis [4,42]. EfficientNet family gained popularity due to favorable performance complexity trade-offs [44,53]. Nevertheless, CNN-based models remain fundamentally constrained by global pooling operations, which limit spatial precision and increase susceptibility to confounding visual cues [54].
Vision transformers (ViT) and hierarchical variants such as the Swin Transformer, were subsequently adapted to chest radiography. This paradigm (Table 3) explicitly captures long-range dependencies, an attractive property for abnormalities with diffuse or global radiographic manifestations. Compared to CNN-based baselines, transformer architectures tend to demonstrate improved sensitivity to diffuse or multi-region abnormalities [55,56], particularly interstitial patterns. However, these gains are typically associated with higher data requirements and increased sensitivity to annotation noise, which may limit robustness in small or heterogeneous datasets. In addition, the data-hungry nature of transformers has motivated hybrid architectures that combine convolutional layers for low-level feature extraction with transformer blocks for higher-level contextual reasoning. These hybrid designs often achieve a more favorable balance between data efficiency and representational capacity [7,28,57].
From a practical perspective, no single architectural paradigm consistently outperforms others across all chest X-ray analysis tasks and clinical settings. Convolutional neural networks remain a strong baseline choice when labeled data are limited, computational efficiency is a priority or rapid deployment is required. Their inductive bias toward local spatial patterns aligns well with many focal pulmonary abnormalities and their compatibility with established explainability techniques facilitates model inspection [4,44].
For example, abnormalities characterized by diffuse or bilateral patterns, such as interstitial disease or pulmonary edema, place greater emphasis on global contextual modeling, whereas focal findings such as nodules or masses require precise spatial localization and benefit from region-aware supervision.
Transformer-based architectures, by contrast, are better suited for scenarios involving large-scale datasets, multi-institutional training groups or abnormalities with diffuse and global radiographic manifestations [55]. Their ability to model long-range dependencies can improve sensitivity to complex patterns, but this advantage typically comes at the cost of increased data requirements, computational complexity and sensitivity to label noise.
Hybrid CNN–Transformer models offer a compromise between these paradigms by combining efficient local feature extraction with global contextual reasoning. In practice, such architectures are most beneficial when moderate-to-large datasets are available and when robustness to spatial heterogeneity is a key objective.
To bridge the gap between convolutional inductive bias and global attention, a subset of the included studies propose hybrid CNN–Transformer designs. This methods combine convolutional backbones for local feature extraction with transformer blocks for global contextual modeling [58].
Importantly, reported performance differences between architectures are often confounded by dataset characteristics, annotation quality and evaluation protocols. As a result, architectural choice should be guided less by marginal benchmark gains and more by practical considerations, including data availability, supervision quality, interpretability requirements and intended clinical use. This perspective is consistent with the comparative overview provided in Table 3 and underlines the need to interpret architectural advances within their broader methodological context.
While the following sections discuss these paradigms in detail, Table 4 provides a unified, practice-oriented summary spanning architectural designs, learning paradigms and multimodal frameworks.

Table 4.
Practical comparison of major deep learning architectures for chest X-ray analysis, highlighting key characteristics, strengths, limitations and recommended usage scenarios.
5.2. Learning Paradigms and Weak Supervision
Given the inherent noise and variability of large-scale CXR annotations, learning paradigms have progressively shifted toward self-supervised and semi-supervised strategies to reduce dependence on imperfect labels and improve representation robustness [13,14]. This limitation has driven a paradigm shift away from purely supervised learning toward self-supervised, semi-supervised and weak supervision approaches, which aim to decouple representation learning from imperfect labels. As summarized in Table 3, these paradigms play a central role in mitigating annotation noise and improving model robustness.
Self-supervised learning has emerged as a particularly influential strategy by enabling representation learning directly from large collections of unlabeled chest radiographs. Contrastive frameworks based on MoCo or SimCLR style objectives learn invariant visual representations that transfer effectively to downstream classification and localization tasks. Empirical evidence consistently shows that SSL pretraining improves downstream performance, calibration and robustness to distribution shifts when compared to models pretrained on natural images [3,59,60]. These gains are especially pronounced in low-label or noisy-label regimes, which are characteristic of most public CXR datasets.
Compared to strongly supervised learning on noisy report-derived labels, SSL and semi-supervised strategies are generally more label-efficient and often more robust under distribution shift; however, their downstream gains remain sensitive to pretraining objective choice, data diversity and fine-tuning protocol, limiting direct comparability across studies.
However, the benefits of self-supervised learning are not uniform across settings. Performance improvements depend strongly on the choice of pretraining objective, data diversity and fine-tuning strategy. In some cases, SSL representations may underperform task-specific supervised pretraining when high-quality expert annotations are available, highlighting the importance of aligning learning paradigms with data characteristics and clinical objectives.
Semi-supervised and weak supervision approaches further extend this paradigm by jointly leveraging limited expert annotations alongside large volumes of weak or unlabeled data. These methods have demonstrated improved generalization across institutions and reduced sensitivity to dataset-specific biases [53]. In practice, they offer a pragmatic compromise between purely supervised and fully self-supervised pipelines, particularly in settings where a small amount of reliable annotation can be obtained.
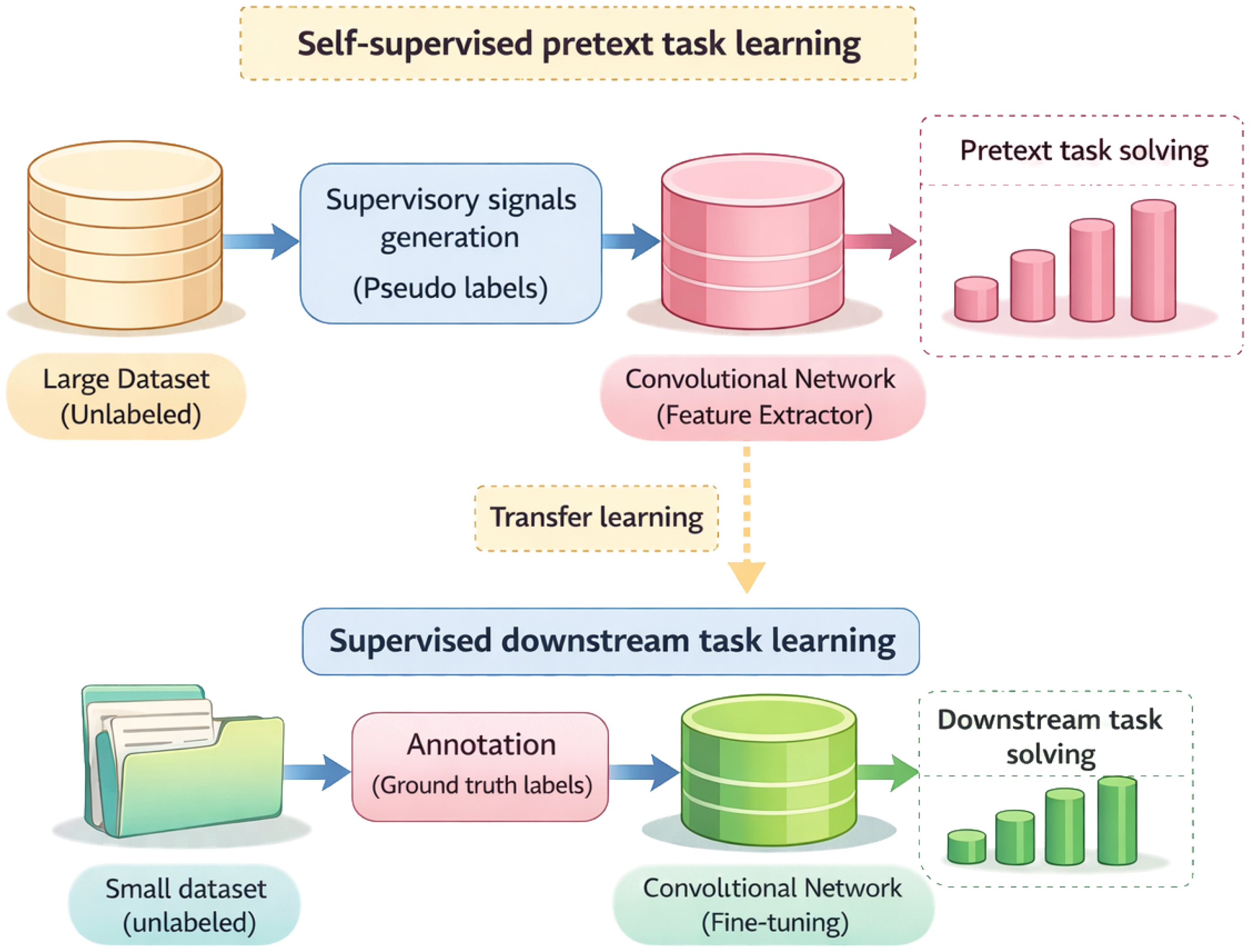
From a practical standpoint, self-supervised learning is most advantageous when expert annotations are scarce, inconsistent or costly to obtain, while semi-supervised strategies are preferable when limited high-quality labels are available to guide downstream optimization. In contrast, strongly supervised learning remains competitive when dense, expert-curated annotations exist and task-specific performance is prioritized. Consequently, learning paradigm selection should be driven by annotation reliability, dataset scale and intended clinical deployment, rather than by architectural considerations alone. Figure 2 illustrate self-supervised and semi-supervised learning paradigms.
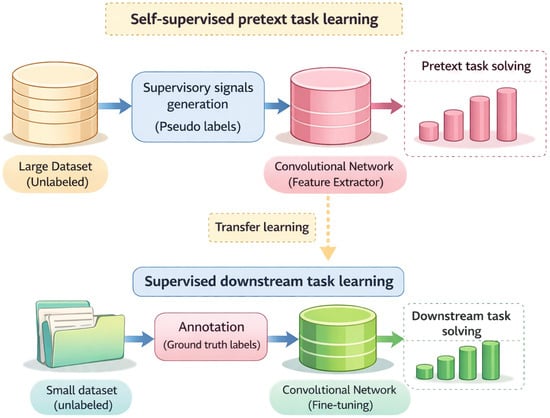
Figure 2.
Self-supervised learning paradigms for chest X-ray analysis. Based on [61]. In the context of publicly available CXR datasets, “ground truth” typically refers to radiologist-level or report-derived image annotations and does not systematically incorporate confirmation by CT, pathology or treatment response across the full dataset.
5.3. Multimodal Learning Frameworks
The availability of paired radiology reports in major CXR datasets has catalyzed the development of multimodal learning frameworks that jointly model images and text. As summarized in Table 3, vision–language models (VLMs) typically rely on weak or implicit supervision derived from image–report pairs, enabling flexible learning paradigms such as zero-shot and prompt-based inference. Early multimodal approaches combined CNN-based visual embeddings with textual representations, demonstrating improved classification performance and increased robustness to labeling noise when compared to purely image-based models [62].
More recent contrastive vision–language models, including BioViL [63,64] and CheXzero [11], learn aligned image–text representations directly from large-scale CXR report corpora without requiring explicit label supervision [18]. This shift toward contrastive multimodal pretraining has enabled scalable representation learning and improved transferability across tasks, particularly in settings where curated expert annotations are limited.
A key advantage of VLMs lies in their ability to support zero-shot generalization and flexible querying of radiographic findings using free-text prompts [18,21]. In contrast to fixed-label classification systems, vision–language representations can encode clinically relevant nuances such as laterality, chronicity and diagnostic uncertainty, which are often implicit in radiology reports but difficult to represent within predefined label sets.
Despite these advantages, current multimodal frameworks face important limitations that constrain clinical deployment. Model performance is strongly influenced by report quality, institutional reporting styles and language biases, which may limit cross-institution generalization. Furthermore, evaluation protocols for zero-shot and prompt-based inference remain insufficiently standardized and robust clinical validation of multimodal predictions is still limited. As a result, while VLMs represent a conceptual shift toward more expressive and clinically aligned CXR-AI systems, they are currently best viewed as enabling technologies for representation learning, weak supervision and exploratory analysis rather than as standalone diagnostic tools.
From a practical standpoint, multimodal learning frameworks are most beneficial when large-scale image–report datasets are available and when flexibility, label efficiency or semantic richness is prioritized over narrowly optimized task-specific performance. In contrast, unimodal image-based models may remain preferable for tightly defined clinical tasks with well-curated expert annotations and established evaluation protocols.
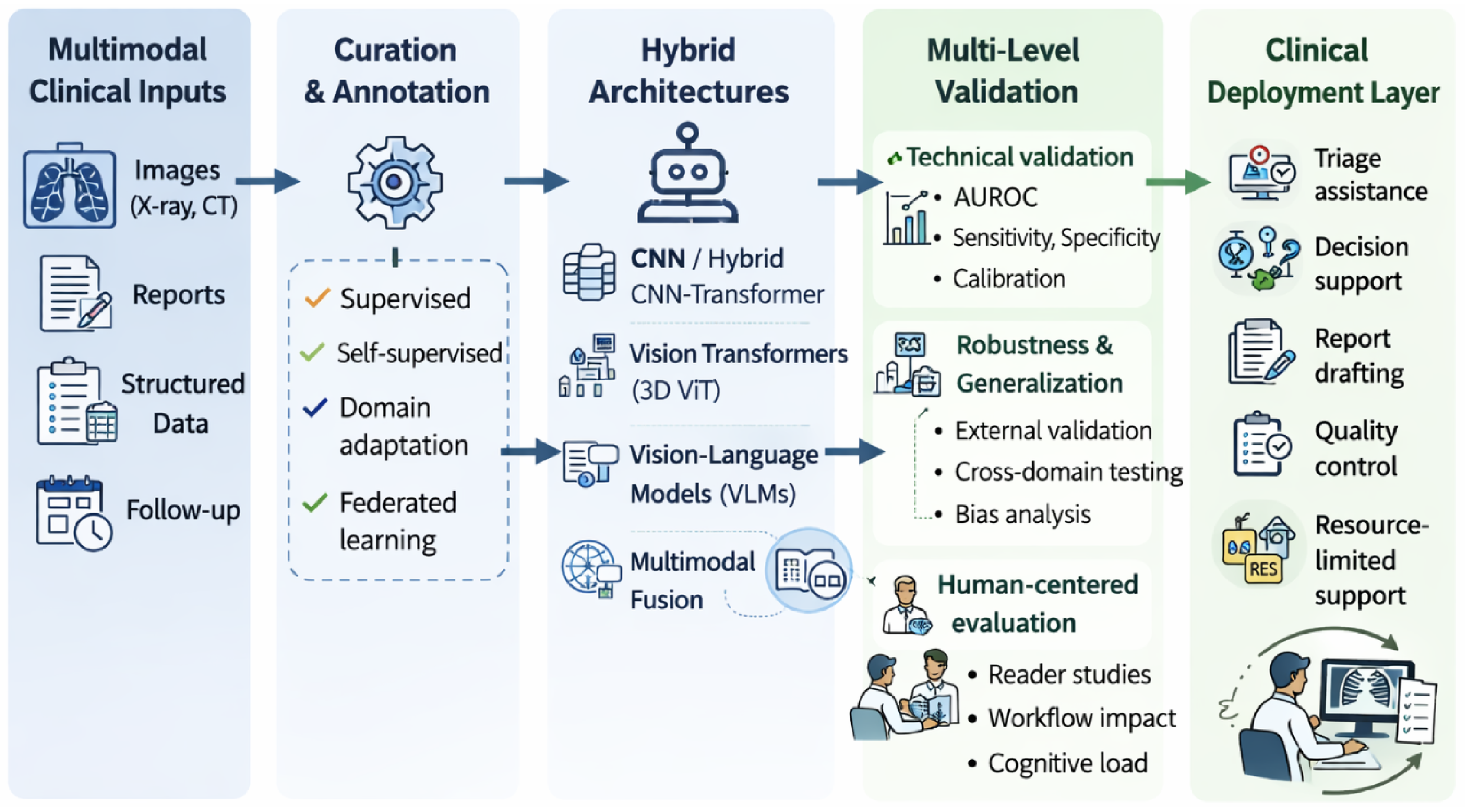
As illustrated in Figure 3, multimodal AI systems integrate visual and textual information within a unified representation framework, enabling cross-modal reasoning, robust generalization and clinically oriented inference strategies across different deployment settings [65].
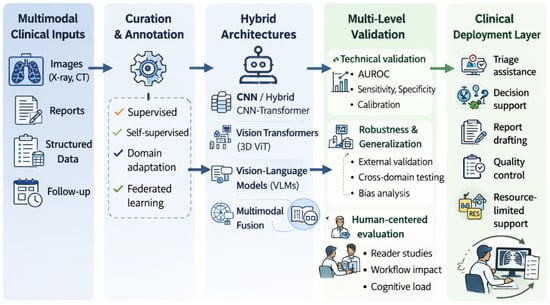
Figure 3.
Conceptual framework for multimodal AI systems in chest radiology. Visual and textual information are integrated within a unified representation space, enabling cross-modal reasoning, robust validation and clinically oriented deployment strategies across different workflow stages.
5.4. Anatomical Structure Localization
Efforts to move beyond purely global image-level predictions have motivated extensive research on regional localization and anatomical structure analysis in chest X-ray imaging. As summarized in Table 3, a common entry point for these approaches has been weak supervision localization, most often implemented through class activation maps or attention-based mechanisms. These techniques enable models to highlight image regions that contribute to specific predictions, thereby providing a degree of post-hoc interpretability. However, such localizations are typically coarse, spatially imprecise and poorly aligned with anatomical boundaries, which limits their suitability for fine-grained clinical interpretation and safety assessment [10].
To introduce stronger anatomical constraints, a defined subset of the 68 included studies incorporated explicit segmentation of lung fields, mediastinal structures and cardiac silhouettes as a preprocessing or auxiliary task. Fully convolutional networks and U-Net–based architectures remain the dominant choices for this purpose [66,67]. By isolating anatomically relevant regions, segmentation-based pipelines can reduce background noise, mitigate acquisition-related variability and provide more stable inputs for downstream classification or localization tasks. In practice, anatomical segmentation often improves robustness and interpretability, particularly in multi-center settings where imaging conditions vary substantially.
Beyond anatomical segmentation, finding-level detection of nodules, masses or focal opacities has been explored using adapted object-detection frameworks such as Faster R-CNN or YOLO. These approaches offer the potential for higher clinical granularity by explicitly localizing pathological findings. However, their progress is severely constrained by the limited availability of high-quality bounding-box or pixel-level annotations in public CXR datasets [68,69]. Annotation cost, inter-observer variability and inconsistent labeling protocols further complicate systematic evaluation and comparison across studies.
From a practical standpoint, weakly supervised localization methods are most appropriate when large-scale datasets with image-level labels are available and interpretability is desired at a coarse, qualitative level. Segmentation-based approaches are preferable when anatomical consistency, robustness and region-specific analysis are priorities, provided that suitable annotations or pretrained segmenters exist. In contrast, finding-level detection frameworks remain best suited for narrowly defined tasks with dedicated expert annotations and are currently less scalable for broad CXR screening applications. These constraints help explain why image-level classification remains the dominant paradigm in the literature, despite its limited clinical granularity.
5.5. Evaluation Strategies and Clinical Translation
Despite steady progress in reported model performance, robustness and generalizability remain persistent and unresolved challenges in chest X-ray AI. Performance degradation across hospitals, imaging systems and patient populations has been repeatedly documented, underscoring the sensitivity of data-driven models to domain shift [22,47]. Differences in acquisition protocols, scanner hardware, preprocessing pipelines and demographic composition can all lead to substantial drops in accuracy when models are evaluated outside their training environment. In response, a range of domain generalization and adaptation strategies has been explored, including style augmentation, adversarial feature alignment and contrastive learning of domain-invariant representations. While such approaches can mitigate performance loss in specific settings, no single strategy has demonstrated consistent superiority across datasets or institutions, highlighting the absence of universally robust solutions [23].
Across architectural categories, strong internal validation performance is commonly reported, particularly for high-prevalence findings on large public benchmarks. Reported discrimination metrics are generally high for conditions such as pleural effusion and cardiomegaly, although gains between architectural families are often incremental and performance varies substantially with dataset composition and validation design.
A closely related limitation is the tendency of deep learning systems to exploit shortcuts—spurious correlations arising from non-pathological cues rather than true disease patterns. In chest radiography, these confounders are particularly prevalent. Medical devices, projection markers, laterality labels or institution-specific acquisition artifacts may inadvertently become predictive signals. Well-documented examples include ventilator tubing acting as a proxy for disease severity or textual markers correlating with diagnostic labels [46]. Although such shortcuts can inflate benchmark performance, they fundamentally undermine model reliability, clinical trust and cross-domain generalization.
Explainability methods are frequently proposed as safeguards against these failure modes. Post-hoc interpretability techniques, most notably Grad-CAM and Grad-CAM++, are widely used to visualize image regions contributing to model predictions [70,71,72]. However, these methods exhibit important limitations, including noisy attribution maps, sensitivity to small perturbations and the absence of causal guarantees. Attribution maps may also highlight irrelevant or confounding regions, complicating their use for safety assessment. Consequently, recent work has shifted toward anatomically grounded explanations, counterfactual reasoning and uncertainty-aware interpretability frameworks [7]. Despite this progress, standardized evaluation protocols for explainability remain underdeveloped and consensus on best practices is still lacking.
Evaluation design further compounds these challenges. Most CXR-AI studies rely on single-center validation using internal test sets drawn from the same institution as the training data. While convenient, such evaluations fail to capture real-world variability and routinely overestimate generalization performance. Multiple studies have demonstrated marked performance degradation under cross-institution testing, reinforcing the need for routine multi-center external validation [22,47]. Nevertheless, external validation remains the exception rather than the norm.
Metric selection plays a critical role in shaping reported conclusions. Area under the receiver operating characteristic curve (AUROC) is by far the most commonly reported metric, yet it is insensitive to class imbalance and provides limited insight into clinically relevant operating points. As a result, high AUROC values may coexist with poor calibration or misleading decision thresholds. Alternative metrics—such as area under the precision–recall curve (AUPRC), sensitivity–specificity trade-offs, F1 score, likelihood ratios and decision-curve analysis—offer more clinically meaningful perspectives but are reported inconsistently across studies.
Importantly, the intrinsic clinical ambiguity of several thoracic findings implies that evaluation metrics must extend beyond aggregate discrimination performance and incorporate calibration, uncertainty quantification and external validation to reflect real-world diagnostic variability.
Calibration and uncertainty estimation remain particularly underexplored. Calibration, defined as the agreement between predicted probabilities and true outcome likelihoods, is essential for safe clinical deployment. Supervision noise and dataset-specific artifacts may inflate discrimination metrics under internal validation, while masking calibration errors and cross-institution generalization failures. Quantitative measures such as the Brier score or Expected Calibration Error (ECE) are still rarely included, limiting insight into predictive reliability.
Similar limitations affect the evaluation of localization and interpretability. Although saliency maps are frequently presented, few studies report quantitative localization metrics such as intersection-over-union (IoU), pointing game accuracy or region-level recall. In practice, attribution maps often emphasize non-pathological regions or confounding artifacts, further restricting their usefulness for model validation and safety assessment.
Finally, most evaluations treat AI systems as standalone classifiers, despite their intended role as decision-support tools within clinical workflows. Clinically meaningful endpoints, such as triage prioritization accuracy, reduction in reporting time, detection of overlooked findings or improvements in radiologist sensitivity, are seldom measured. Multi-reader studies, prospective evaluations and workflow-level assessments remain relatively rare, yet they are critical for regulatory approval and clinical integration.
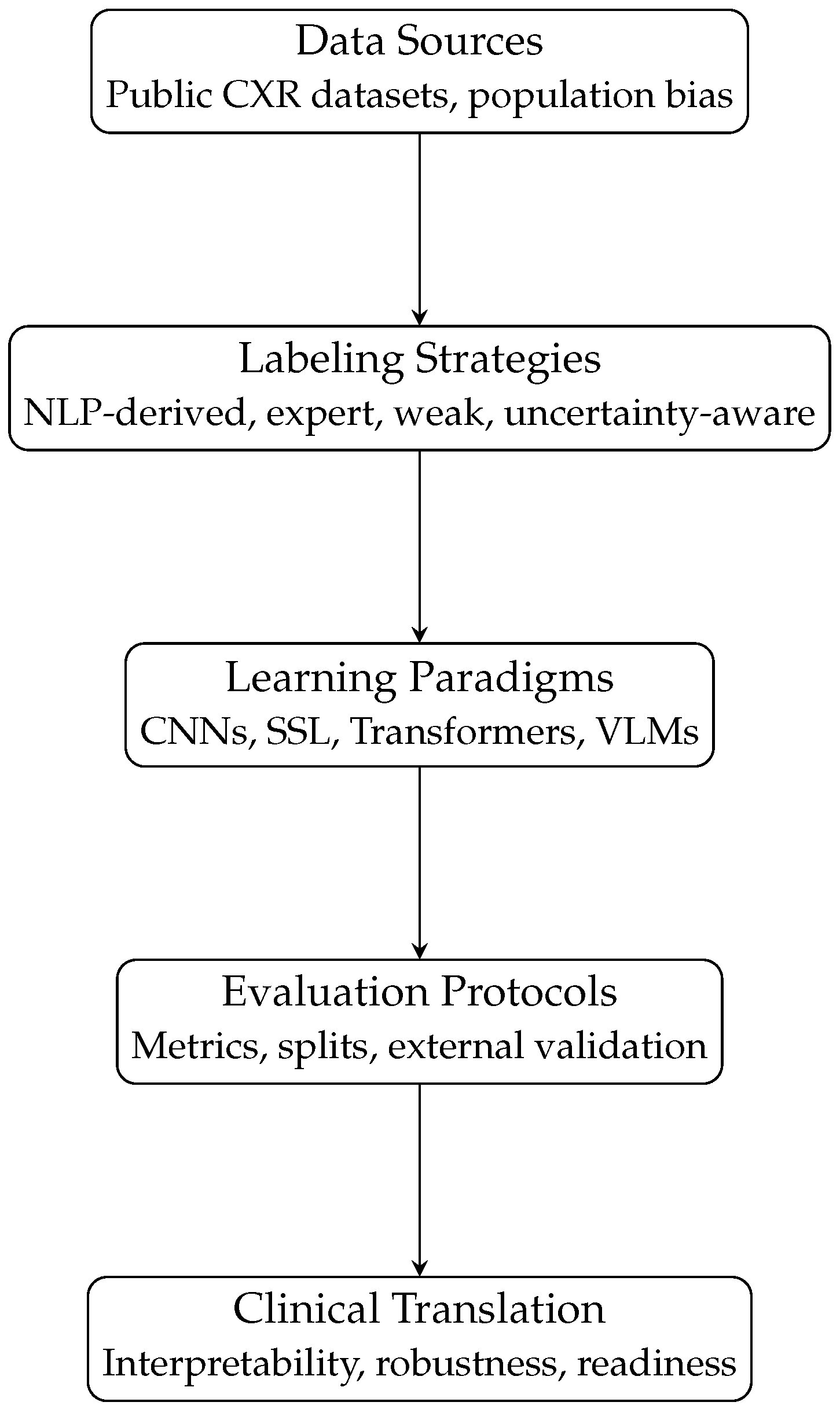
From a practical standpoint, robust clinical translation of CXR-AI systems requires evaluation frameworks that extend beyond algorithmic accuracy to explicitly address robustness, calibration, interpretability and real-world impact. Establishing standardized, multi-center and clinically grounded evaluation protocols is therefore essential to move CXR-AI systems beyond research prototypes and toward safe, deployable medical tools. The absence of standardized multi-dataset benchmarking protocols further complicates direct comparison across studies. Reported gains often reflect dataset-specific properties rather than universally transferable methodological improvements. Figure 4 summarizes the data-centric and model-centric factors discussed in this review and illustrates how their interactions jointly shape robustness, generalizability and clinical readiness in CXR-AI systems.
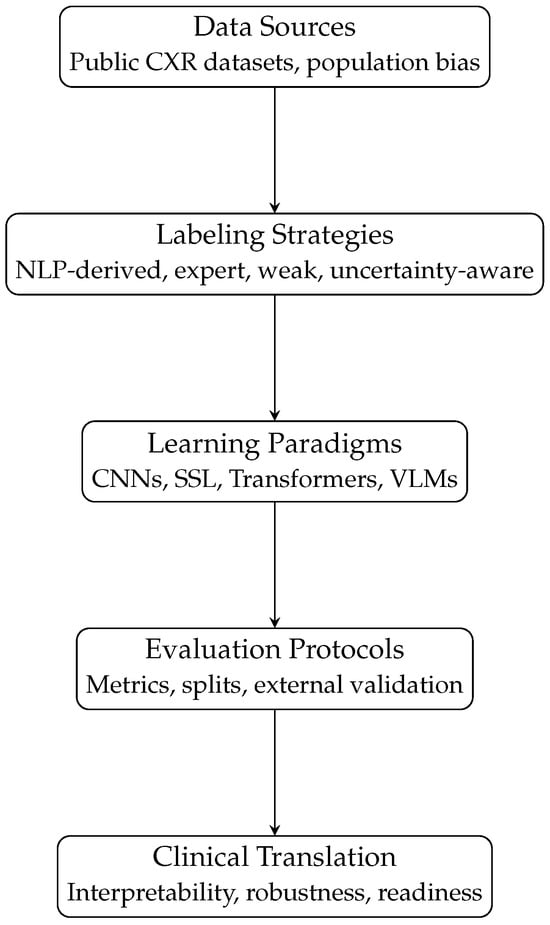
Figure 4.
Conceptual pipeline illustrating how data sources, labeling strategies, learning paradigms and evaluation protocols jointly shape the clinical readiness of AI systems for chest X-ray analysis.
Table 5 summarizes the most recurrent methodological limitations identified across the reviewed studies through CLAIM-guided qualitative appraisal, together with their practical implications for robustness, generalizability and clinical translation. These gaps were consistently observed across heterogeneous study designs, suggesting that they reflect systemic challenges in current CXR-AI research rather than isolated methodological shortcomings.

Table 5.
Recurrent methodological gaps identified through CLAIM-guided appraisal of CXR-AI studies and their practical implications for clinical translation.
6. Discussion
These inconsistencies underscore the need for structured comparative criteria that explicitly account for supervision quality, dataset heterogeneity and evaluation design when interpreting reported performance improvements. The methodological gaps identified through the CLAIM-guided appraisal provide a structural explanation for these persistent translational barriers in CXR-AI systems.
One of the most striking insights from the literature is that many of these limitations are not independent. Weak labels derived from radiology reports, for example, do more than introduce noise: they actively encourage shortcut learning. Models may latch onto artifacts such as medical devices, projection markers or acquisition-specific patterns instead of learning genuine pulmonary pathology. These shortcuts then amplify sensitivity to domain shift, making performance highly unstable when models are applied across institutions or imaging systems. Compounding the problem, such failure modes frequently remain hidden because evaluation protocols tend to prioritize aggregate metrics like AUROC, while overlooking calibration, external validation, spatial localization accuracy and impact on clinical workflows.
Recent architectures such as transformer-based models or hybrid convolutional-attention networks have undoubtedly expanded the representational capacity of CXR-AI. These approaches are better equipped to capture global context under sufficiently large and well-curated datasets, where their capacity to model long-range dependencies can translate into improved sensitivity to diffuse or multi-region abnormalities. However, in smaller or noisier datasets, their higher representational capacity may amplify annotation bias and reduce robustness. Yet improved accuracy alone does not resolve deeper questions of trust, interpretability or safety. Likewise, self-supervised learning has emerged as a powerful strategy to mitigate label noise and enhance generalization, but its clinical relevance ultimately depends on how rigorously these models are evaluated and how consistently training and testing conditions are reported.
Another persistent gap lies at the interface between algorithm development and clinical practice. Despite the intended role of AI as a decision-making support tool, relatively few studies examine its effects on radiologist performance, triage prioritization, diagnostic delays or error reduction. Prospective and multi-center evaluations remain rare and practical considerations, such as integration with PACS, latency constraints, responsibility attribution or ongoing monitoring for model drift, are often left unaddressed. Without attention to these factors, even technically sophisticated models risk remaining confined to research settings.
An additional conceptual consideration concerns whether current CXR-AI systems are primarily designed to replicate radiologist-level findings detection or to improve the radiological workflows. Most of studies emphasizes automated detection and classification of pulmonary findings, reporting strong discrimination metrics for common abnormalities. However, these approaches often target tasks that experienced radiologists already perform reliably, particularly in well-defined clinical scenarios.Alternatively, greater translational value can be achieved through workflow-oriented improvements, such as structured retrieval of relevant clinical context, triage organization, quantification of changes in medical history and support for structured report writing using multimodal or vision-language models. Thus, the future of CXR-AI may depend less on marginal gains in finding-level accuracy and more on its ability to integrate seamlessly into clinical ecosystems. This is of particular interest in areas with fewer resources or specialists.
From a methodological standpoint, this review should be interpreted within its intended scope. The absence of prospective protocol registration and formal quantitative risk-of-bias scoring constitutes a limitation, as the review prioritizes methodological synthesis and conceptual integration over evidence hierarchy assessment. While elements of PRISMA were adopted to enhance transparency in study identification and selection, the work was not designed as a formal systematic review with quantitative meta-analysis.
Overall, the current landscape reflects a field that has achieved substantial methodological sophistication but has yet to fully bridge the translational divide. Progress will likely require a shift away from narrowly benchmark-driven development toward more holistic approaches. Robust representation learning must be coupled with confounder-aware model design, evaluation frameworks that emphasize calibration and generalization and deployment strategies aligned with real clinical workflows. Only by addressing these interconnected challenges can CXR-AI systems move from promising prototypes to reliable tools in routine clinical care.
7. Conclusions and Future Work
This review provides a foundation for understanding the current state of CXR-AI and identifies critical avenues for future research. Continued interdisciplinary collaboration, between clinicians, computer scientists, engineers and regulatory experts, will be essential to translate these technologies into safe, effective and scalable tools for patient care.
Artificial intelligence has transformed the landscape of chest X-ray analysis, enabling automated detection of pulmonary abnormalities with unprecedented scale and accuracy. Technological advances, including convolutional networks, transformers, self-supervised learning and multimodal architectures, have substantially improved model performance and broadened the scope of CXR-AI applications.
However, challenges remain significant. Label noise, domain shift, shortcut learning, underrepresented pathologies and fragmented evaluation practices limit the reliability and clinical applicability of current systems. Future CXR-AI systems should evolve from image-level classifiers to integrated clinical systems capable of contextual reasoning, temporal assessments and workflow improvement, especially in high-volume, resource-constrained settings. Progress in algorithmic accuracy alone is insufficient. Robust clinical adoption will require comprehensive evaluation frameworks, high-quality datasets, interpretable and trustworthy models and seamless integration into radiologist workflows. Thus, the implementation or use in the real world of these tools must be done carefully and under strict controls.
Multimodal and vision–language integration represents a key frontier for the future. Future CXR-AI systems will increasingly incorporate radiology reports, structured clinical information, vital signs and electronic health records, allowing predictions that reflect richer clinical context rather than isolated image appearance. VLMs enable flexible prompt-based querying, support uncertainty-aware reasoning and capture semantic nuances, such as laterality or chronicity, that are often lost in discrete label taxonomies.
Temporal and longitudinal modeling remain underexplored despite their clinical relevance. Most existing systems treat each radiograph as an independent sample, ignoring the diagnostic value of prior imaging and temporal disease evolution. Integrating sequential radiographs and linking them to clinical events could substantially improve diagnostic accuracy, prognostic modeling and monitoring of chronic or progressive conditions.
Another key point for the next years is the real-world deployment, which will require substantial progress in domain generalization and adaptation. Equally critical is the establishment of standardized evaluation protocols and prospective clinical validation frameworks. Promising directions include domain-invariant feature learning, dynamic test-time adaptation mechanisms and style randomization techniques designed to minimize sensitivity to acquisition variability. Successful deployment will require close collaboration with PACS vendors, hospital IT departments, regulatory bodies and healthcare institutions to ensure that CXR-AI tools are not only accurate, but also usable, safe and sustainable in daily practice.
Author Contributions
Conceptualization, G.P.-C.; methodology, G.P.-C., J.J.J.-D. and F.D.P.-C.; software, G.P.-C. and F.D.P.-C.; validation, G.P.-C., F.D.P.-C. and J.J.J.-D.; formal analysis, G.P.-C.; investigation, G.P.-C., J.J.J.-D. and F.D.P.-C.; resources, G.P.-C. and F.D.P.-C.; data curation, G.P.-C. and F.D.P.-C.; writing—original draft preparation, G.P.-C.; writing—review and editing, G.P.-C., F.D.P.-C. and J.J.J.-D.; visualization, G.P.-C.; supervision, G.P.-C., F.D.P.-C. and J.J.J.-D.; project administration, J.J.J.-D.; funding acquisition, J.J.J.-D. All authors have read and agreed to the published version of the manuscript.
Funding
This research received no external funding.
Informed Consent Statement
Not applicable.
Conflicts of Interest
The authors declare no conflicts of interest.
References
- Hwang, E.J.; Park, C.M. Clinical Implementation of Deep Learning in Thoracic Radiology: Potential Applications and Challenges. Korean J. Radiol. 2020, 21, 511–525. [Google Scholar] [CrossRef]
- Annarumma, M.; Withey, S.J.; Bakewell, R.J.; Pesce, E.; Goh, V.; Montana, G. Automated Triaging of Adult Chest Radiographs with Deep Artificial Neural Networks. Radiology 2019, 291, 196–202. [Google Scholar] [CrossRef]
- Çallı, E.; Sogancioglu, E.; van Ginneken, B.; van Leeuwen, K.G.; Murphy, K. Deep learning for chest X-ray analysis: A survey. Med. Image Anal. 2021, 72, 102125. [Google Scholar] [CrossRef]
- Baltruschat, I.M.; Nickisch, H.; Grass, M.; Knopp, T.; Saalbach, A. Comparison of Deep Learning Approaches for Multi-Label Chest X-Ray Classification. Sci. Rep. 2019, 9, 6381. [Google Scholar] [CrossRef] [PubMed]
- Hwang, E.J.; Park, S.; Jin, K.N.; Kim, J.I.; Choi, S.Y.; Lee, J.H.; Goo, J.M.; Aum, J.; Yim, J.J.; Cohen, J.G.; et al. Development and Validation of a Deep Learning-Based Automated Detection Algorithm for Major Thoracic Diseases on Chest Radiographs. JAMA Netw. Open 2019, 2, e191095. [Google Scholar] [CrossRef] [PubMed]
- Ahmad, K.; Rehman, H.U.; Shah, B.; Ali, F.; Hussain, I. Dual-model approach for accurate chest disease detection using GViT and Swin Transformer V2. Sci. Rep. 2025, 15, 31717. [Google Scholar] [CrossRef] [PubMed]
- Yang, M.; Zhu, W. Embedding Radiomics into Vision Transformers for Multimodal Medical Image Classification. Front. Artif. Intell. 2025; in press. [Google Scholar] [CrossRef]
- Shamshad, F.; Khan, S.; Zamir, S.W.; Khan, M.H.; Hayat, M.; Khan, F.; Fu, H. Transformers in Medical Imaging: A Survey. arXiv 2022, arXiv:2201.09873. [Google Scholar] [CrossRef]
- Rajpurkar, P.; Irvin, J.; Ball, R.L.; Zhu, K.; Yang, B.; Mehta, H.; Duan, T.; Ding, D.; Bagul, A.; Langlotz, C.P.; et al. Deep Learning for Chest Radiograph Diagnosis: A Retrospective Comparison of the CheXNeXt Algorithm to Practicing Radiologists. PLoS Med. 2018, 15, e1002686. [Google Scholar] [CrossRef]
- Chowdary, G.J.; Kanhangad, V. A Dual-Branch Network for Diagnosis of Thorax Diseases From Chest X-rays. IEEE J. Biomed. Health Inform. 2022, 26, 6081–6092. [Google Scholar] [CrossRef]
- Tiu, E.; Talius, E.; Patel, P.; Langlotz, C.P.; Ng, A.Y.; Rajpurkar, P. Expert-level detection of pathologies from unannotated chest X-ray images via self-supervised learning. Nat. Biomed. Eng. 2022, 6, 1399–1406. [Google Scholar] [CrossRef]
- Wang, X.; Peng, Y.; Lu, L.; Lu, Z.; Bagheri, M.; Summers, R.M. ChestX-ray: Hospital-Scale Chest X-ray Database and Benchmarks on Weakly Supervised Classification and Localization of Common Thorax Diseases. In Deep Learning and Convolutional Neural Networks for Medical Imaging and Clinical Informatics; Springer International Publishing: Berlin/Heidelberg, Germany, 2019; pp. 369–392. [Google Scholar] [CrossRef]
- Irvin, J.; Rajpurkar, P.; Ko, M.; Yu, Y.; Ciurea-Ilcus, S.; Chute, C.; Marklund, H.; Haghgoo, B.; Ball, R.; Shpanskaya, K.; et al. CheXpert: A Large Chest Radiograph Dataset with Uncertainty Labels and Expert Comparison. In Proceedings of the AAAI Conference on Artificial Intelligence, Honolulu, HI, USA, 27 January–1 February 2019; Volume 33, pp. 590–597. [Google Scholar] [CrossRef]
- Johnson, A.E.W.; Pollard, T.J.; Berkowitz, S.J.; Greenbaum, N.R.; Lungren, M.P.; Deng, C.Y.; Mark, R.G.; Horng, S. MIMIC-CXR, a de-identified publicly available database of chest radiographs with free-text reports. Sci. Data 2019, 6, 317. [Google Scholar] [CrossRef]
- Nguyen, H.Q.; Lam, K.; Le, L.T.; Pham, H.H.; Tran, D.Q.; Nguyen, D.B.; Le, D.D.; Pham, C.M.; Tong, H.T.; Dinh, D.H.; et al. VinDr-CXR: An Open Dataset of Chest X-Rays with Radiologist’s Annotations. Sci. Data 2022, 9, 429. [Google Scholar] [CrossRef]
- Bustos, A.; Pertusa, A.; Salinas, J.M.; de la Iglesia-Vayá, M. PadChest: A large chest x-ray image dataset with multi-label annotated reports. Med. Image Anal. 2020, 66, 101797. [Google Scholar] [CrossRef]
- Anderson, P.G.; Tarder-Stoll, H.; Alpaslan, M.; Keathley, N.; Levin, D.L.; Venkatesh, S.; Bartel, E.; Sicular, S.; Howell, S.; Lindsey, R.V.; et al. Deep learning improves physician accuracy in the comprehensive detection of abnormalities on chest X-rays. Sci. Rep. 2024, 14, 25151. [Google Scholar] [CrossRef]
- Zhang, Z.; Jiang, A. Interactive Dual-Stream Contrastive Learning for Radiology Report Generation. J. Biomed. Inform. 2024, 157, 104718. [Google Scholar] [CrossRef]
- Jang, J.; Kyung, D.; Kim, S.; Lee, H.; Bae, K.; Choi, E. Significantly improving zero-shot X-ray pathology classification via fine-tuning pre-trained image–text encoders. Sci. Rep. 2024, 14, 23199. [Google Scholar] [CrossRef]
- Yuan, L.; Zhu, W. Rethinking Domain-Specific Pretraining by Supervised or Self-Supervised Learning for Chest Radiograph Classification. Health Care Sci. 2025, 4, e70009. [Google Scholar] [CrossRef] [PubMed]
- Di Folco, F.; Chan, S.; Xu, H.; Li, J.; Hu, M. Semantic Alignment of Unimodal Medical Text and Vision Representations. Med. Image Anal. 2025; in press. [Google Scholar] [CrossRef]
- Zech, J.R.; Badgeley, M.A.; Liu, M.; Costa, A.B.; Titano, J.J.; Oermann, E.K. Variable generalization performance of a deep learning model to detect pneumonia in chest radiographs: A cross-sectional study. PLoS Med. 2018, 15, e1002683. [Google Scholar] [CrossRef] [PubMed]
- Tayebi Arasteh, S.; Kühl, C.; Saehn, M.J.; Isfort, P.; Truhn, D.; Nebelung, S. Enhancing domain generalization in the AI-based analysis of chest radiographs with federated learning. Sci. Rep. 2023, 13, 22576. [Google Scholar] [CrossRef]
- Tjoa, E.; Guan, C. A Survey on Explainable Artificial Intelligence (XAI): Toward Medical XAI. IEEE Trans. Neural Netw. Learn. Syst. 2021, 32, 4793–4813. [Google Scholar] [CrossRef]
- Saha, P.; Nadeem, S.A.; Comellas, A. A survey on artificial intelligence in pulmonary imaging. Wiley Interdiscip. Rev. Data Min. Knowl. Discov. 2023, 13, e1510. [Google Scholar] [CrossRef]
- Sindhu, A.; Jadhav, U.S.; Ghewade, B.; Bhanushali, J.; Yadav, P. Revolutionizing Pulmonary Diagnostics: A Narrative Review of Artificial Intelligence Applications in Lung Imaging. Cureus 2024, 16, e57657. [Google Scholar] [CrossRef]
- Gonem, S.; Janssens, W.; Das, N.; Topalovic, M. Applications of artificial intelligence and machine learning in respiratory medicine. Thorax 2020, 75, 695–701. [Google Scholar] [CrossRef]
- Sturdza, O.; Filip, A.; Ionescu, R.T.; Coman, R. Deep Learning Network Selection and Optimized Information Fusion for Enhanced COVID-19 Detection. Diagnostics 2025, 15, 1830. [Google Scholar] [CrossRef]
- Musa, A.; Prasad, R. Addressing cross-population domain shift in chest X-ray classification through supervised adversarial domain adaptation. Sci. Rep. 2025, 15, 11383. [Google Scholar] [CrossRef]
- Kim, J.Y.; Ryu, W.S.; Kim, D.; Kim, E.Y. Better performance of deep learning pulmonary nodule detection using chest radiography with pixel level labels in reference to computed tomography: Data quality matters. Sci. Rep. 2024, 14, 15967. [Google Scholar] [CrossRef]
- Mongan, J.; Moy, L.; Kahn, C.E. Checklist for Artificial Intelligence in Medical Imaging (CLAIM): A Guide for Authors and Reviewers. Radiol. Artif. Intell. 2020, 2, e200029. [Google Scholar] [CrossRef]
- Lee, S.M.; Seo, J.; Yun, J.; Cho, Y.H.; Vogel-Claussen, J.; Schiebler, M.; Gefter, W.; van Beek, E.V.; Goo, J.; Lee, K.; et al. Deep Learning Applications in Chest Radiography and Computed Tomography: Current State of the Art. J. Thorac. Imaging 2019, 34, 75–85. [Google Scholar] [CrossRef]
- Ma, J.; Song, Y.; Tian, X.; Hua, Y.; Zhang, R.; Wu, J. Survey on deep learning for pulmonary medical imaging. Front. Med. 2019, 14, 450–469. [Google Scholar] [CrossRef]
- de Camargo, T.F.O.; Ribeiro, G.A.S.; da Silva, M.C.B.; da Silva, L.O.; Torres, P.P.T.E.S.; Rodrigues, D.D.S.D.S.; de Santos, M.O.N.; Filho, W.S.; Rosa, M.E.E.; Novaes, M.D.A.; et al. Clinical validation of an artificial intelligence algorithm for classifying tuberculosis and pulmonary findings in chest radiographs. Front. Artif. Intell. 2025, 8, 1512910. [Google Scholar] [CrossRef]
- de la Orden Kett Morais, S.R.; Felder, F.N.; Walsh, S.L.F. From pixels to prognosis: Unlocking the potential of deep learning in fibrotic lung disease imaging analysis. Br. J. Radiol. 2024, 97, 1517–1525. [Google Scholar] [CrossRef]
- Chassagnon, G.; de Margerie-Mellon, C.; Vakalopoulou, M.; Marini, R.; Hoang-Thi, T.N.; Revel, M.; Soyer, P. Artificial intelligence in lung cancer: Current applications and perspectives. Jpn. J. Radiol. 2022, 41, 235–244. [Google Scholar] [CrossRef]
- Ferrari, R. Writing narrative style literature reviews. Med. Writ. 2015, 24, 230–235. [Google Scholar] [CrossRef]
- Page, M.J.; McKenzie, J.E.; Bossuyt, P.M.; Boutron, I.; Hoffmann, T.C.; Mulrow, C.D.; Shamseer, L.; Tetzlaff, J.M.; Akl, E.A.; Brennan, S.E.; et al. The PRISMA 2020 statement: An updated guideline for reporting systematic reviews. BMJ 2021, 372, n71. [Google Scholar] [CrossRef]
- Tejani, A.S.; Klontzas, M.E.; Gatti, A.A.; Mongan, J.T.; Moy, L.; Park, S.H.; Kahn, C.E.; Abbara, S.; Afat, S.; Anazodo, U.C.; et al. Checklist for Artificial Intelligence in Medical Imaging (CLAIM): 2024 Update. Radiol. Artif. Intell. 2024, 6, e240300. [Google Scholar] [CrossRef]
- Warriner, D. How to Read a Paper: The Basics of Evidence-Based Medicine. BMJ 2008, 336, 1381. [Google Scholar] [CrossRef]
- Thomas, J.; Harden, A. Methods for the thematic synthesis of qualitative research in systematic reviews. BMC Med. Res. Methodol. 2008, 8, 45. [Google Scholar] [CrossRef]
- Nasser, A.A.; Akhloufi, M.A. Deep Learning Methods for Chest Disease Detection Using Radiography Images. SN Comput. Sci. 2023, 4, 388. [Google Scholar] [CrossRef]
- Mitsuyama, Y.; Takita, H.; Walston, S.L.; Watanabe, K.; Ishimaru, S.; Miki, Y.; Ueda, D. Deep learning models for radiography body-part classification and chest radiograph projection/orientation classification: A multi-institutional study. Eur. Radiol. 2025. [Google Scholar] [CrossRef]
- Ait Nasser, A.; Akhloufi, M.A. A Review of Recent Advances in Deep Learning Models for Chest Disease Detection Using Radiography. Diagnostics 2023, 13, 159. [Google Scholar] [CrossRef] [PubMed]
- Jaeger, S.; Candemir, S.; Antani, S.; Wáng, Y.X.; Lu, P.X.; Thoma, G. Two public chest X-ray datasets for computer-aided screening of pulmonary diseases. Quant. Imaging Med. Surg. 2014, 4, 475–477. [Google Scholar] [CrossRef]
- DeGrave, A.J.; Janizek, J.D.; Lee, S.I. AI for radiographic COVID-19 diagnosis exhibits shortcuts. Nat. Mach. Intell. 2021, 3, 610–619. [Google Scholar] [CrossRef]
- Kelly, C.J.; Karthikesalingam, A.; Suleyman, M.; Corrado, G.; King, D. Key challenges for delivering clinical impact with artificial intelligence. BMC Med. 2019, 17, 195. [Google Scholar] [CrossRef]
- Yao, L.; Poblenz, E.; Dagunts, D.; Covington, B.; Bernard, D.; Lyman, K. Learning to Diagnose from Scratch by Exploiting Dependencies Among Labels. arXiv 2017, arXiv:1710.10501. [Google Scholar] [CrossRef]
- Siddiqi, I.; Javaid, N. Deep Learning for Pneumonia Detection in Chest X-ray Images: A Comprehensive Survey. J. Imaging 2024, 10, 176. [Google Scholar] [CrossRef]
- Yanar, M.; Hardalaç, F. CELM: An Ensemble Deep Learning Model for Early Cardiomegaly Diagnosis. Diagnostics 2025, 15, 1602. [Google Scholar] [CrossRef]
- Abdullah, M.; Kim, J. Automated Radiology Report Labeling in Chest X-Ray Pathologies: Development and Evaluation of a Large Language Model Framework. JMIR Med. Inform. 2024, 13, e68618. [Google Scholar] [CrossRef]
- Uslu, E.E.; Sezer, E.; Guven, Z.A. NLP-Powered Healthcare Insights: A Comparative Analysis for Multi-Labeling Classification With MIMIC-CXR Dataset. IEEE Access 2024, 12, 67314–67324. [Google Scholar] [CrossRef]
- Al-qaness, M.A.A.; Zhu, J.; Al-Alimi, D.; Dahou, A.; Alsamhi, S.H.; Abd Elaziz, M.; Ewees, A.A. Chest X-ray Images for Lung Disease Detection Using Deep Learning Techniques: A Comprehensive Survey. Arch. Comput. Methods Eng. 2024, 31, 3267–3301. [Google Scholar] [CrossRef]
- Rguibi, Z.; Hajami, A.; Zitouni, D.; Elqaraoui, A.; Bedraoui, A. CXAI: Explaining Convolutional Neural Networks for Medical Imaging Diagnostic. Electronics 2022, 11, 1775. [Google Scholar] [CrossRef]
- Liu, Z.; Lin, Y.; Cao, Y.; Hu, H.; Wei, Y.; Zhang, Z.; Lin, S.; Guo, B. Swin Transformer: Hierarchical Vision Transformer using Shifted Windows. In Proceedings of the IEEE/CVF International Conference on Computer Vision (ICCV), Montreal, QC, Canada, 10–17 October 2021; pp. 9992–10002. [Google Scholar] [CrossRef]
- Hatamizadeh, A.; Tang, Y.; Nath, V.; Yang, D.; Myronenko, A.; Landman, B.A.; Roth, H.R.; Xu, D. UNETR: Transformers for 3D Medical Image Segmentation. In Proceedings of the IEEE/CVF Winter Conference on Applications of Computer Vision (WACV), Waikoloa, HI, USA, 3–8 January 2022; pp. 1748–1758. [Google Scholar] [CrossRef]
- Wu, Y.; Qi, S.; Sun, Y.; Xia, S. A vision transformer for emphysema classification using CT images. Phys. Med. Biol. 2021, 66, 245012. [Google Scholar] [CrossRef]
- Zeineldin, R.A.; Karar, M.E.; Elshaer, Z.; Coburger, J. Explainable hybrid vision transformers and convolutional network for multimodal glioma segmentation in brain MRI. Sci. Rep. 2024, 14, 3713. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Bentley, P.; Mori, K.; Misawa, K.; Fujiwara, M.; Rueckert, D. Self-supervised learning for medical image analysis using image context restoration. Med. Image Anal. 2019, 58, 101539. [Google Scholar] [CrossRef]
- Azizi, S.; Mustafa, B.; Ryan, F.; Beaver, Z.; Freyberg, J.; Deaton, J.; Loh, A.; Karthikesalingam, A.; Kornblith, S.; Chen, T.; et al. Big Self-Supervised Models Advance Medical Image Classification. In Proceedings of the IEEE/CVF International Conference on Computer Vision (ICCV), Montreal, QC, Canada, 10–17 October 2021. [Google Scholar] [CrossRef]
- Shurrab, S.; Duwairi, R. Self-supervised learning methods and applications in medical imaging analysis: A survey. PeerJ Comput. Sci. 2022, 8, e1045. [Google Scholar] [CrossRef] [PubMed]
- Huang, S.C.; Pareek, A.; Seyyedi, S.; Banerjee, I.; Lungren, M.P. Fusion of medical imaging and electronic health records using deep learning: A systematic review and implementation guidelines. npj Digit. Med. 2020, 3, 136. [Google Scholar] [CrossRef] [PubMed]
- Bannur, S.; Hyland, S.; Liu, Q.; Pérez-García, F.; Ilse, M.; Castro, D.C.; Boecking, B.; Sharma, H.; Bouzid, K.; Thieme, A.; et al. Learning to Exploit Temporal Structure for Biomedical Vision-Language Processing. In Proceedings of the 2023 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), Vancouver, BC, Canada, 17–24 June 2023; pp. 15016–15027. [Google Scholar] [CrossRef]
- Boecking, B.; Usuyama, N.; Bannur, S.; Castro, D.C.; Schwaighofer, A.; Hyland, S.; Wetscherek, M.; Naumann, T.; Nori, A.; Alvarez-Valle, J.; et al. Making the Most of Text Semantics to Improve Biomedical Vision–Language Processing. In Proceedings of the Computer Vision—ECCV 2022; Lecture Notes in Computer Science; Avidan, S., Brostow, G., Cissé, M., Farinella, G.M., Hassner, T., Eds.; Springer: Cham, Switzerland, 2022; Volume 13696, pp. 1–21. [Google Scholar] [CrossRef]
- Wu, J.; Wang, Y.; Zhong, Z.; Liao, W.; Trayanova, N.; Jiao, Z.; Bai, H.X. Vision-language foundation model for 3D medical imaging. npj Artif. Intell. 2025, 1, 17. [Google Scholar] [CrossRef]
- Ronneberger, O.; Fischer, P.; Brox, T. U-Net: Convolutional Networks for Biomedical Image Segmentation. In Proceedings of the Medical Image Computing and Computer-Assisted Intervention—MICCAI 2015; Lecture Notes in Computer Science; Navab, N., Hornegger, J., Wells, W.M., Frangi, A.F., Eds.; Springer: Cham, Switzerland, 2015; Volume 9351, pp. 234–241. [Google Scholar] [CrossRef]
- Novikov, A.A.; Lenis, D.; Major, D.; Hladůvka, J.; Wimmer, M.; Bühler, K. Fully Convolutional Architectures for Multiclass Segmentation in Chest Radiographs. IEEE Trans. Med. Imaging 2018, 37, 1865–1876. [Google Scholar] [CrossRef]
- Mansoor, A.; Bagci, U.; Xu, Z.; Foster, B.; Olivier, K.N.; Elinoff, J.M.; Suffredini, A.F.; Udupa, J.K.; Mollura, D.J. A Generic Approach to Pathological Lung Segmentation. IEEE Trans. Med. Imaging 2014, 33, 2293–2310. [Google Scholar] [CrossRef]
- Freedman, D.; Radke, R.J.; Zhang, T.; Jeong, Y.; Lovelock, D.M.; Chen, G.T. Model-based segmentation of medical imagery by matching distributions. IEEE Trans. Med. Imaging 2005, 24, 281–292. [Google Scholar] [CrossRef] [PubMed]
- Selvaraju, R.R.; Cogswell, M.; Das, A.; Vedantam, R.; Parikh, D.; Batra, D. Grad-CAM: Visual Explanations from Deep Networks via Gradient-Based Localization. Int. J. Comput. Vis. 2019, 128, 336–359. [Google Scholar] [CrossRef]
- Ghassemi, M.; Oakden-Rayner, L.; Beam, A.L. The false hope of current approaches to explainable artificial intelligence in health care. Lancet Digit. Health 2021, 3, e745–e750. [Google Scholar] [CrossRef]
- Groen, A.M.; Kraan, R.; Amirkhan, S.F.; Daams, J.G.; Maas, M. A systematic review on the use of explainability in deep learning systems for computer aided diagnosis in radiology: Limited use of explainable AI? Eur. J. Radiol. 2022, 157, 110592. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.