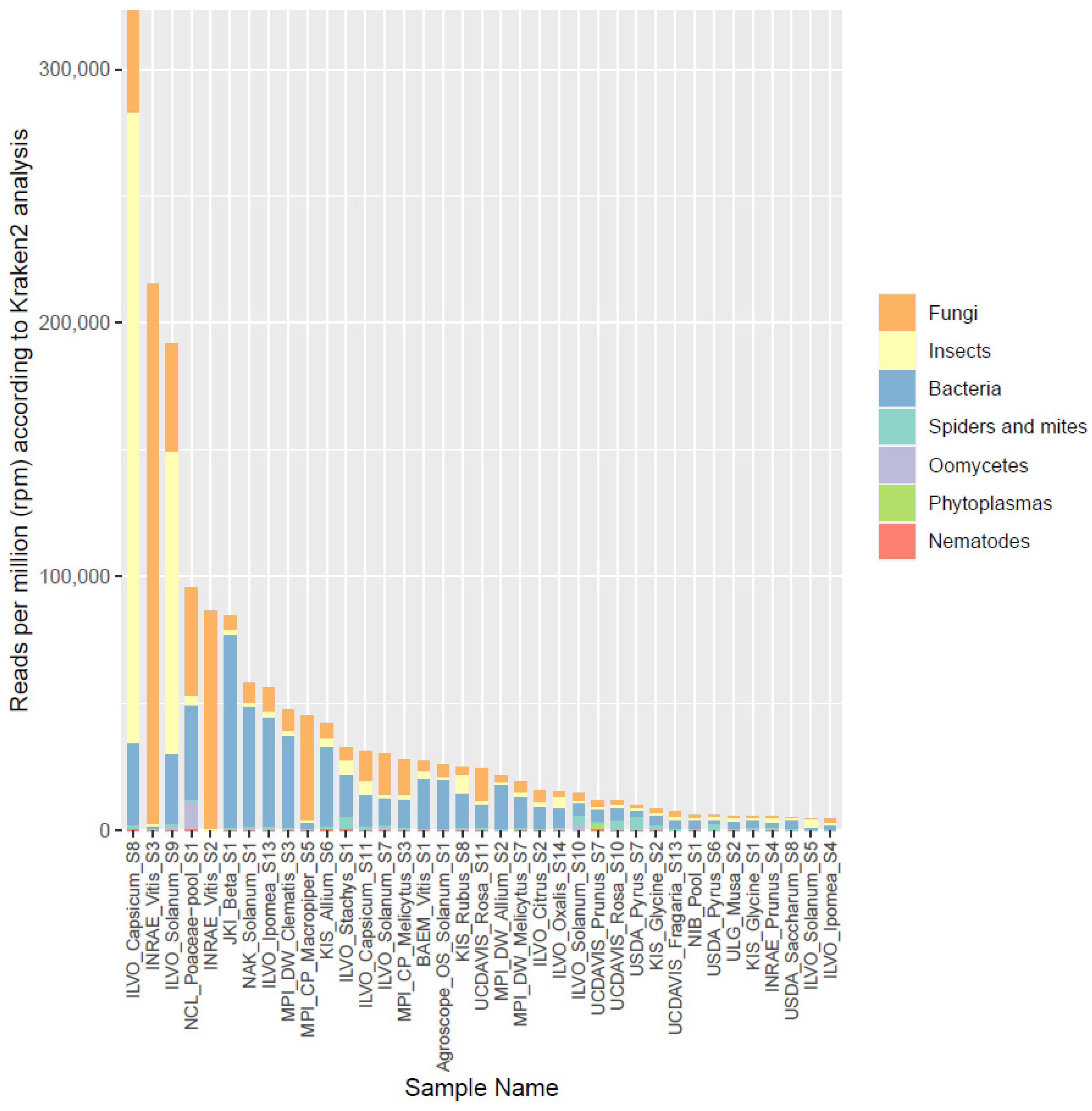
In the original publication [1], there were two mistakes in Figure 3 as published. The legend showed the wrong descriptions with the colors, and the Y-axis scale was not correctly labeled. The corrected Figure 3 with an adjusted description appears below.
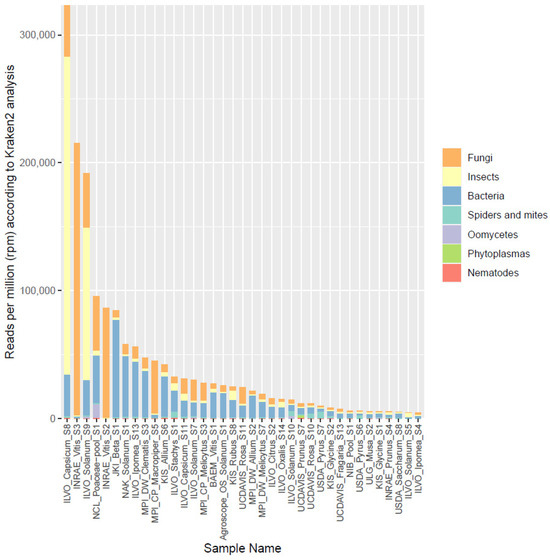
Figure 3.
Overview of the proportion of reads (in reads per million) assigned by Kraken2 to different broad organismal categories (fungi, insects, bacteria, spiders and mites, oomycetes, phytoplasmas, and nematodes) for the 37 selected datasets.
The authors state that the scientific conclusions are unaffected. This correction was approved by the Academic Editor. The original publication has also been updated.
Reference
- Haegeman, A.; Foucart, Y.; De Jonghe, K.; Goedefroit, T.; Al Rwahnih, M.; Boonham, N.; Candresse, T.; Gaafar, Y.Z.A.; Hurtado-Gonzales, O.P.; Kogej Zwitter, Z.; et al. Looking beyond Virus Detection in RNA Sequencing Data: Lessons Learned from a Community-Based Effort to Detect Cellular Plant Pathogens and Pests. Plants 2023, 12, 2139. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).