1. Introduction
The species commonly referred to as blueberry (
Vaccinium sect.
Cyanococcus) are perennial flowering plants native to North America. In the last thirty years, blueberry production has increased by over 600%, to 858,886 tonnes of blueberry harvested worldwide in 2020, with over 70% of that production occurring in the United States of America, Peru, and Canada [
1]. This increase in production has occurred in the wake of the publication of a number of studies, beginning in the mid-1990s, examining the potential health benefits associated with blueberry consumption, especially in regard to antioxidant activity, for which blueberry has among the highest levels compared to other fruits and vegetables [
2,
3,
4]. In addition to the use of the berries as fresh or processed foodstuff, the plants have also been used in traditional medicines [
5]. Even today in the realm of natural medicine, blueberries are recommended for use in treating Type II diabetes [
6].
Compounds with antioxidant activity, such as phenolic acids, anthocyanins, and anthocyanidins, have an important function in the plant. Plants regularly experience environmental stresses resulting in the production of reactive oxygen species (ROS), excessive levels of which disrupt normal metabolic function by damaging lipids, proteins, and nucleic acids, all of which negatively impact plant growth and development [
7,
8,
9]. Compounds with antioxidant activity scavenge ROS, protecting normal metabolic function [
10]. While the evidence for effect of antioxidant compounds obtained through food on human health is slim, the possibility of efficaciousness has encouraged substantial research on the topic.
One class of naturally occurring compounds known to have significant antioxidant capacity are phenolic acids, compounds containing a phenol moiety highly suited for trapping free radicals [
10]. Chlorogenic acids are a family of polyphenol esters formed between
trans-cinnamic acids and quinic acids [
11]. Chlorogenic acids are one of the most well-studied families of polyphenols, due to their abundance in plant-based food and drinks [
12]. They are widespread in plants and can be found in nearly all plant species [
13,
14,
15,
16].
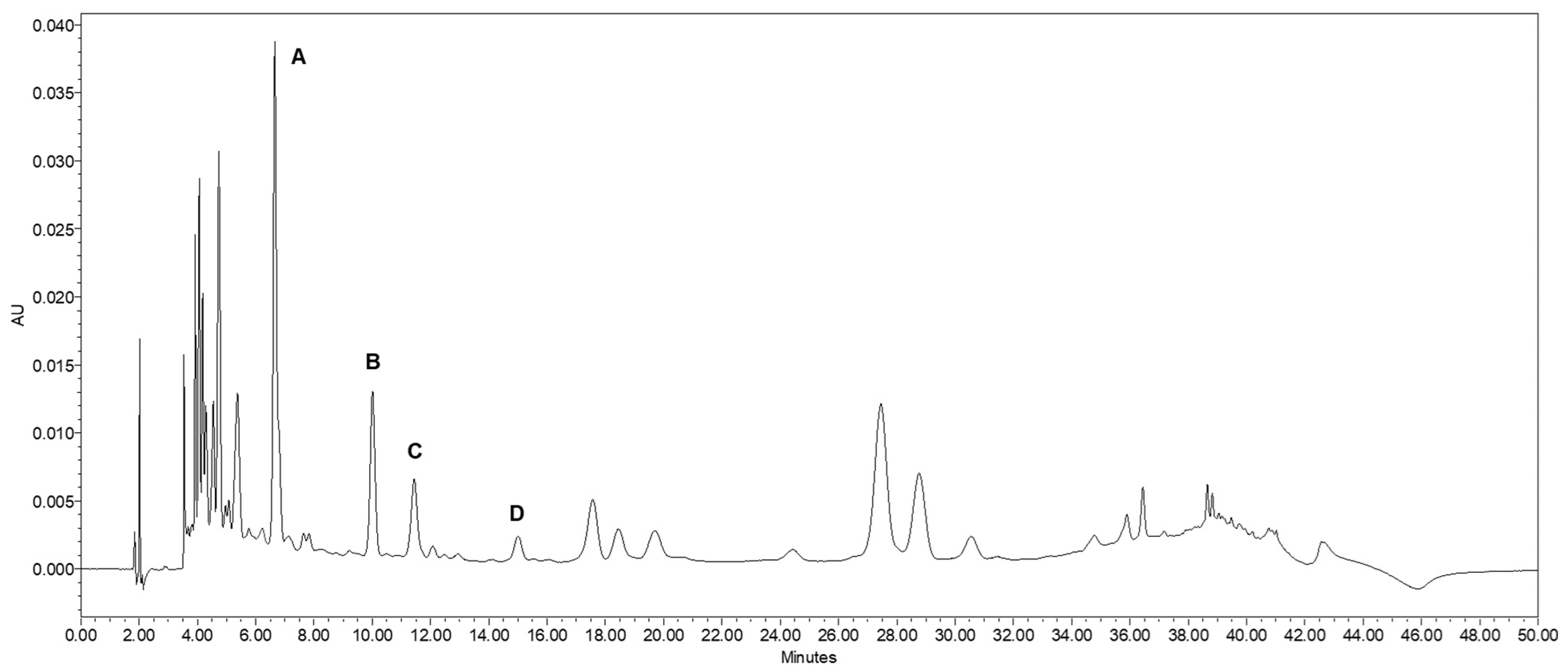
One important subgroup of chlorogenic acids is the caffeoylquinic acids, which consist of esterifications of caffeic acid (
Figure 1A), of which there are several isomeric forms. The most abundant of these isomers present in plants is 5-
O-caffeoylquinic acid (5-CQA) [
12], and it is the most widely studied due to its commercial availability [
14]. Generally, the term “chlorogenic acid” (CGA) refers to 5-CQA [
12]. It should be noted, however, that the nomenclature of the isomers 5-CQA (
Figure 1B) and 3-CQA (3-
O-cafffeoylquinic acid) (
Figure 1C) can create some confusion. In 1976, the International Union of Pure and Applied Chemistry (IUPAC) reversed the order of the numbering of atoms on the quinic acid ring [
17]. Consequently, the previously identified 3-CQA was renamed as 5-CQA [
17], and the current 3-CQA refers to neochlorogenic acid in accordance with the new numbering system [
12,
16]. However, due to the name change, there is still considerable confusion in the literature, and many papers do not specify, when they use the term “chlorogenic acid”, whether they mean to refer to 5-CQA or 3-CQA [
16]. This paper follows Clifford et al. [
16] and uses the current IUPAC numbering, with CGA as 5-CQA.
Chlorogenic acids are the major hydroxycinnamic acids present in blueberries [
18], likely accounting for a large proportion of their antioxidant activity [
19,
20], with CGA being the major component [
21]. CGA constitutes 10–16% of total acids in the blueberry fruit [
22,
23,
24]. It is present in concentrations of 98–208 mg per 100 g FW (fresh weight) in
V. corymbosum cultivars [
21,
24,
25]. Other phenolic acids such as caffeic acid, p-coumaric acid, and ferulic acid are present in concentrations under 1% [
21,
22].
CGA is one of the most abundant beneficial polyphenols in the human diet and is well known as a nutritional antioxidant in plant-based foods [
26,
27]. Dietary consumption of CGA is associated with the prevention of certain oxidative and degenerative, age-related diseases [
28,
29,
30]. Compelling evidence indicates that dietary CGA can promote a wide range of pharmacological effects and biological activities in various tissues and organs [
12]. Numerous studies have demonstrated the antioxidant activities of CGA, which include inhibiting the formation or scavenging of ROS [
31]. CGA is also negatively correlated with the risk of various harmful conditions, such as oxidative and inflammatory stresses [
32], type 2 diabetes mellitus [
33,
34], cardiovascular disease [
35], neurodegenerative disease [
36], and cancer [
37].
Compounds closely related to CGA are the acetylated caffeoylquinic acids (ACQA, C
18H
20O
10), which have previously been identified in blueberry [
38]. These compounds have not been well characterized and the configurations of the compounds are not known, but the acetylation is likely to be on the quinic acid moiety [
38].
Figure 1D shows a potential chemical structure for ACQA proposed by Jaiswal et al. [
38]. Note that the regiochemistry of the acetyl group in
Figure 1D is an arbitrarily selected example; the compound identified in the present analysis may not be the 4-acetyl caffeoylquinic acid isomer.
While CGA has been studied for its potential for improving human health, similar efforts have yet to be made to understand the potential health impacts of consumption of ACQA1 and ACQA2. Further research is required to understand the potential benefits of the compounds, as well as their bioavailability when consumed in food or drink.
Another related compound to CGA is caffeoylarbutin (CA). As with CGA, CA is an ester of caffeic acid, though with an arbutin group instead of quinic acid (
Figure 1E). CA has been previously identified in the leaves of blueberry [
39], as well as in the leaves of other
Vaccinium species, such as lingonberry (
V. vitis-idaea) [
40,
41],
V. dunalianum [
42,
43], and bilberry (
V. myrtillus) [
39,
41].
Other minor and specialty crops have seen great advances in the availability of genetic resources over the past decade. However, blueberry has lagged behind, with only a few published genome sequences [
44,
45,
46] and a limited number of genotyped mapping populations [
46,
47,
48,
49], all of which were constructed through crosses between tetraploid cultivars of
V. corymbosum. The development of mapping populations in blueberry is complicated by long generation time and partial self-sterility but is of exceeding importance to crop improvement efforts, as the identification of genetic markers and trait loci can increase the speed at which new varieties can be developed, and aid in understanding the mechanisms of compound biosynthesis.
The blueberry (
Cyanoccocus) section of
Vaccinium is highly diverse, including species of differing ploidy levels and adapted to different environments. The highly interfertile nature of the section offers great opportunities for geneticists and plant breeders alike to identify potentially valuable variations which could be used to develop new blueberry cultivars with improved nutritional value and increased resilience in the face of climate change. The most commercially important blueberries are tetraploid (4n = 48) highbush plants (
V. corymbosum). Accordingly, most of the available evidence on the beneficial health effects of blueberry deals with compounds identified in tetraploids. However, many wild species exist throughout North America [
50], naturally occurring as diploids, tetraploids, and hexaploids [
20]. Among these are numerous diploid blueberry species which offer diverse germplasm [
51], though most have limited analyses of their genetics. Two species likely to be of use in this effort are
V. darrowii and
V. corymbosum.
V. darrowii is a subtropical lowbush diploid blueberry native to the American southeast which has been used as a source of variation in breeding programs [
52,
53], while
V. corymbosum is a temperate highbush blueberry native to the mid-Atlantic region of the United States. The varying climates to which these two species are adapted, as well as genetic drift over evolutionary time, have resulted in wide differentiation on many traits including its fruit chemistry which is quite distinct from that of the highbush species [
32,
36]. This differentiation is exploited in this study by using hybrid populations of these two species for trait mapping. By identifying quantitative trait loci (QTL) in a population containing
V. darrowii ancestry, greater insight can be obtained into the genetic architecture of blueberry.
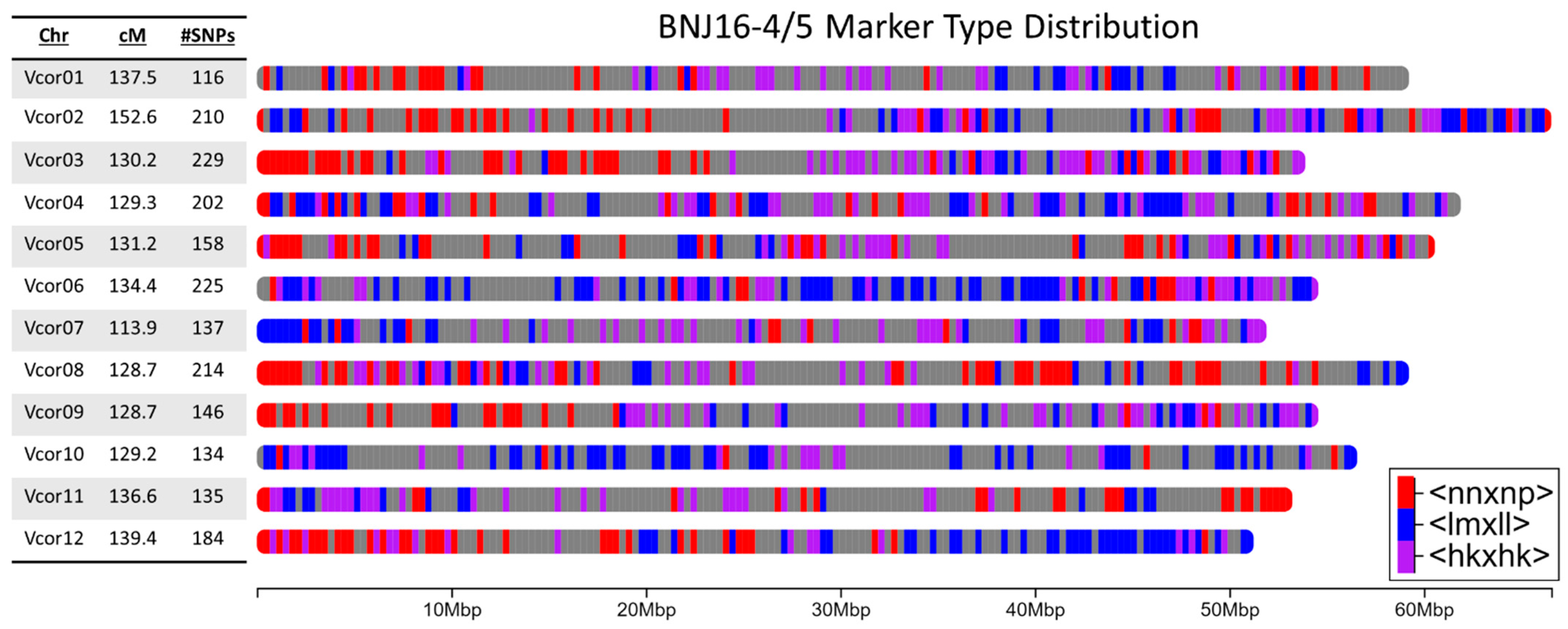
A cost-effective method to produce genetic information for newly developed populations is genotyping-by-sequencing (GBS) as it simultaneously performs single nucleotide polymorphism (SNP) discovery and genotyping, eliminating a step required by other methods [
54]. Using the newly available genome sequences, we can create highly accurate maps based on the physical ordering of the observed SNP markers. To the authors’ knowledge, phenolic acid content in blueberry has only been genetically mapped once previously, by Mengist et al. [
48], who identified a QTL on Vc02 for CGA in an F
1 tetraploid
V. corymbosum population. While CA has been identified in a number of different species, as yet no QTL for CA has been identified. The ACQA compounds have not been fully characterized, and as yet have not been identified outside of blueberry, let alone been mapped.
In this paper, we present the first genotyped diploid interspecific mapping population derived from crosses between V. darrowii and V. corymbosum var. caesariense (a diploid variety of V. corymbosum), two divergent species. This large population, recently developed at the Marucci Blueberry and Cranberry Research Center in Chatsworth, NJ, segregates for many traits of interest to breeders, including fruit chemistry. We present here mapping of the genetic control of phenolic acid content in blueberry fruit.
3. Discussion
Phenolic acids are known to be potent antioxidants [
10] with implications for human health. Breeding programs can make use of the marker-trait association presented in this study to develop marker-assisted breeding programs, potentially increasing the speed of the selection process and leading to the development of improved varieties with increased antioxidant potential.
The phenylpropanoid biosynthesis pathways have been extensively studied, with the pathways well understood [
16,
56,
57,
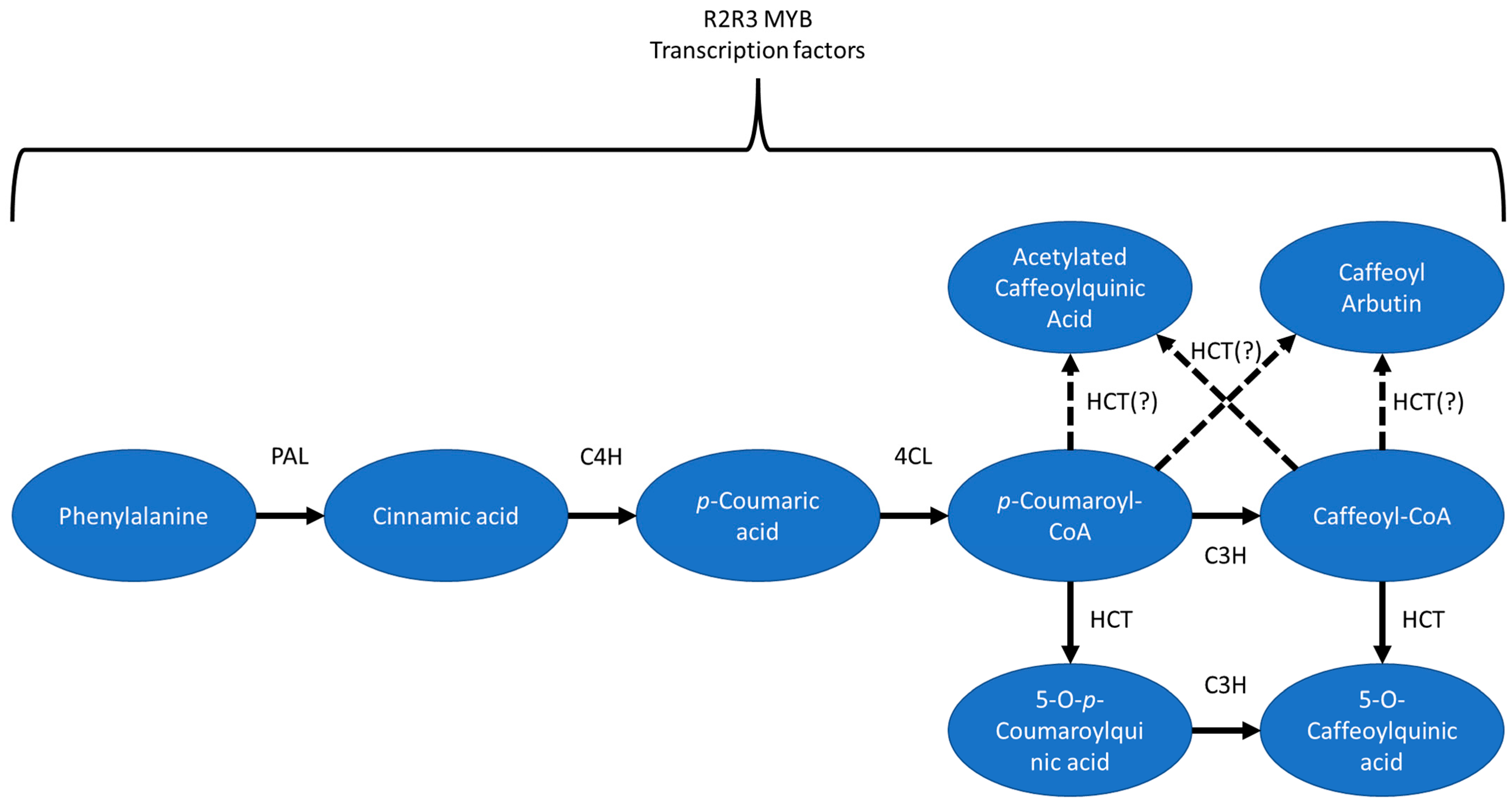
58], including both the enzymes involved, such as various transferases and hydroxylases and potential transcriptional regulators, such as those in the R2R3 MYB transcription family. Some potential pathways for the biosynthesis of ACQA1, ACQA2, and CA can be found in
Figure 6.
Using the BLUPs, CGA, ACQA1, and ACQA2 had QTL located on the proximal arm of Vc02. Mapping for CA using BLUPs did not identify a peak on Vc02, though mapping with 2020 data did identify a QTL close to the others, and the 2019 data showed elevated correlation between CA and the ACQA compounds but one which did not rise to the level of significance. The overlapping QTL on Vc02 suggest that a single gene or several genes located in that region are involved in the biosynthesis of all four compounds. Notably, seven genes in the significant region on Vc02 had BLAST hits on HCT from tea (
Camellia sinensis) and coffee (
Coffea arabica). Previous studies have shown that HCT performs the final step in the main CGA biosynthesis pathway, producing caffeoyl-CoA, the penultimate compound in the path [
16,
56]. While this is the main pathway, others have been proposed [
56]. Of relevance is the alternate pathway wherein cinnamic acid is bonded with glucose by UGCT. Two gene models in the significant region have BLAST hits on UGCT identified from poplar (
Populus spp.). Unfortunately, while Mengist et al. [
46] mapped CGA content and produced transcriptomes in tetraploid blueberry, they did not make a comparison between genotypes with varying levels of CGA and as yet, there is no searchable database with expression data in different tissues (J. L. Humann, personal communication).
R2R3 MYBs have been associated with the phenylpropanoid biosynthesis pathway [
58,
59]. Within the significant region identified on Vc02 are fifteen gene models annotated as MYB-encoding (
Table S4), offering additional avenues of potential exploration. Future studies should focus on the potential activity of such transcription factors in the biosynthesis of CGA, ACQA1, ACQA2, and CA in
Vaccinium, perhaps identifying markers which can be used for breeding purposes.
Considerably less is known regarding the biosynthesis of other caffeoylquinic acid isomers. Our current understanding is that other caffeoylquinic acid isomers are derived from CGA, but there is little data on the enzymes that would be involved in such conversions [
16]. In addition to the QTL identified on Vc02, CA also mapped to regions on Vc07 and Vc12, whereas the other compounds did not. This suggests that while CA biosynthesis is related to biosynthesis of the remaining compounds, it is more complicated. Further research is required to elucidate the biosynthesis pathways of the studied phenolic acids.
Comparing the results of trait mapping presented here with the previous findings of Mengist et al. [
46] showcases the mapping power of the interspecific population. While Mengist et al. [
46] used a population of 196 individuals and found QTL for CGA with LOD scores of 5.9–6.9, the present study used a comparable number of individuals and identified QTL for CGA with LOD scores of ~16.0. This difference is likely partially explained by the larger difference between the parents in the present study. While the parents in Mengist et al. [
46] population, the cultivars Jewel and Draper-44392, differed by ~25mg/100g FW, BNJ05-237-8 and BNJ05-218-9 differed by ~110mg/100g FW. It is unlikely, however, that this would explain the 10-magnitude difference in probabilities. It is more likely that the diploid nature of the BNJ16-4 population allows for greater clarity. Amadeu et al. [
60] demonstrates the increased accuracy of trait mapping in diploid as compared to tetraploid populations, suggesting that trait mapping efforts should be focused on diploid populations when possible. The BNJ16-4 population shows segregation for a wide range of traits, beyond those presented here. Future studies should make use of the population to map those traits with a similar degree of confidence.
In addition to its interspecific nature, the population used in this study offers advantages over the tetraploid populations used in other studies. This primarily stems from the diploid nature of the population, which greatly simplifies the genetic analysis. With just two copies of the genome, constructing maps is far simpler, as well as determining inheritance, dosage effects, and performance of QTL mapping [
60]. Additionally, due to the relatively common failure of diploid male
Vaccinium gametes to complete meiosis, ~13.5% of pollen grains have unreduced gametes [
61], meaning that pollen from a diploid plant can be directly crossed with a tetraploid plant to bring the desired trait into the breeding population. Further, diploid blueberry can be induced to undergo polyploidization using oryzalin or colchicine [
62]. Consequently, the bulk of breeding could potentially occur at the diploid level and the findings of mapping studies done in diploid plants, such as the present one, could be directly applied to the breeding program.
The nature of the map conformed to expectations from the literature. The centromeric regions, as defined by a region with large physical distances and minimal genetic distances, and hence a slope near zero, was generally found at or near the center of the physical map (
Figure S2). This accords with the karyotype data from Hall and Galleta [
63], who reported that centromeres in
Vaccinium were, when observed, median to submedian. The one exception was Vc06, where the centromere was shifted toward the proximal arm of the chromosome.
The present study identifies a strong QTL for the tested phenolic acids, and its overlap with the QTL region identified by Mengist et al. [
46] lends credence to our finding. However, the lack of independent populations in which markers could be tested means that any genetic selection, such as in a breeding program, would require using an individual or parent from the BNJ16-4 population as the donor. Further analysis on phenolic acid content in independent populations, such as biparental, multiparental, or germplasm collections, is required to develop widely applicable markers which could be used in marker-assisted selection.
The observed near 1:1 correlation between ACQA1 and ACQA2, in conjunction with both traits showing highly significant associations with the same region on Vc02, suggests the possibility that biosynthesis of both compounds is carried out by a single enzyme, and which of the two compounds is produced is random. The minor positive correlation between CGA and CA observed in the BNJ16-4 population (0.26 in 2019 and 0.37 in 2020,
Table 4) and the minor negative correlation between CGA and the ACQA compounds (
Table 4) suggest a relationship in the biosynthetic pathways of these compounds.
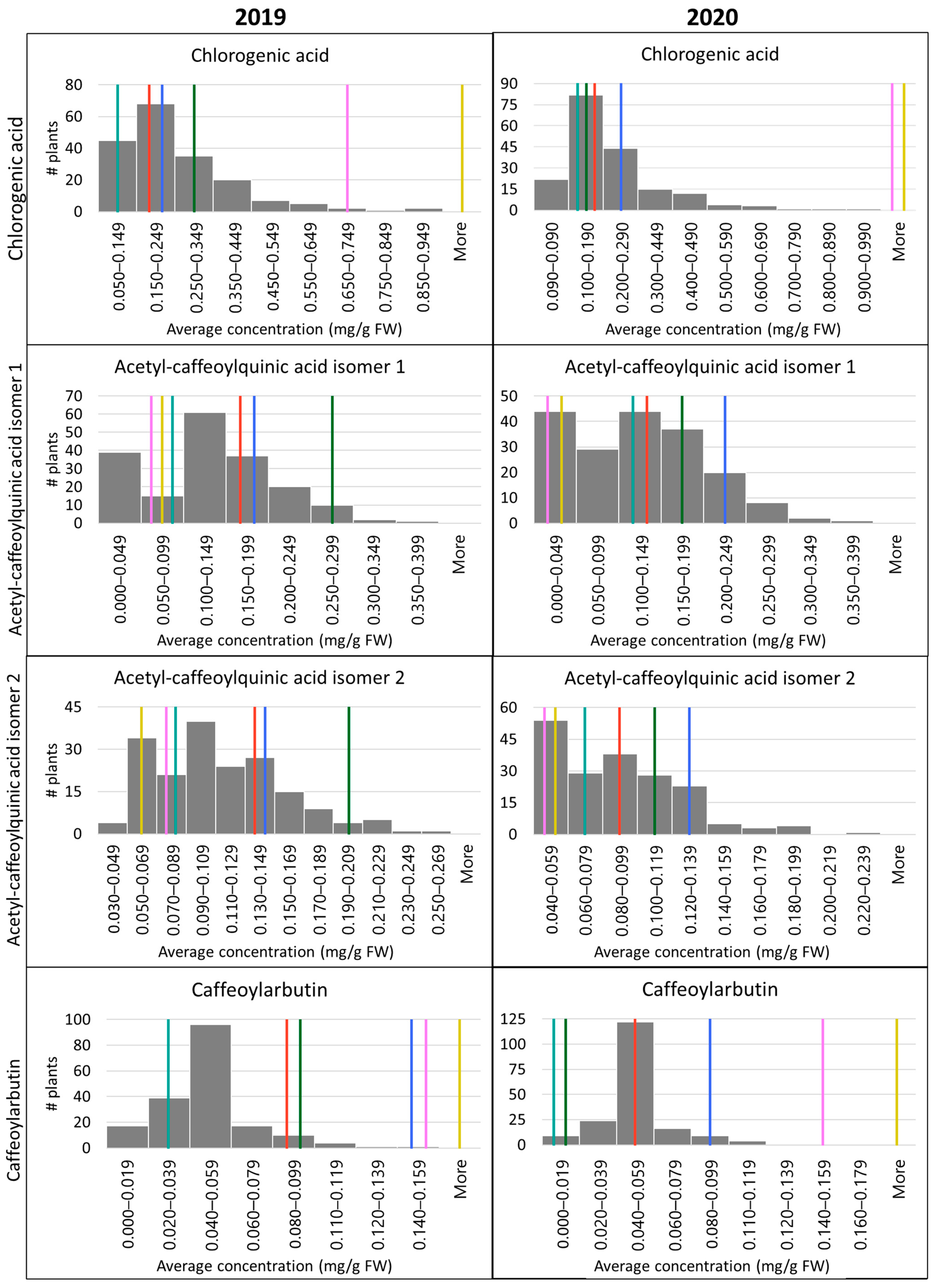
Phenolic content in OPB-15 varied dramatically between 2019 and 2020, with nearly double the concentration or more of each compound observed in 2019 as in 2020 (
Table 2). This pattern was not observed in any of the other parents or grandparents. Previous studies have found that blueberries collected later in the season show increased levels of phenols, anthocyanins, and antioxidants [
3,
64,
65,
66]. Indeed, harvesting began and ended later in 2019 than in 2020. Another possible cause is elevated UV radiation in 2019 compared to 2020, which may have affected the phenolic content, though UV radiation was not measured in the greenhouse in which the plants were grown. Previous studies in carrot [
67] and blueberry [
68] have indicated that CGA content increases in correlation with increased UV radiation. It should also be noted that OPB-15 had low fruit yields in 2020, perhaps contributing or relating to the observed difference in CGA content. Furthermore, it is worth noting that the BNJ16-4 population also showed a general drop in phenolic compound concentrations in 2020 compared to 2019, with the exception of ACQA1, though not to the same extent as observed in OPB-15 (
Table 2). This general decrease was significant, as evident in the results from the Kruskal–Wallis test (
Table 3). Environmental variation could be involved in the inconsistent identification of a peak on Vc02 for CA. Further investigation on the possible environmental factors affecting phenolic content in blueberry is required, such as additional years of data or trials in different locations.
Similarly, CA is thought to have potentially beneficial effects for human health. CA levels are high in the leaves of bearberry and
V. dunalium, and a tisane is made from the leaves of both in traditional medicine [
39,
69]. Arbutin and its derivatives, such as CA, are used as skin-whitening agents, due to their activity in inhibiting melanogenesis by inhibiting tyrosinase [
39,
70]; indeed, it has been shown to inhibit melanogenesis in zebrafish [
71]. Arbutin has also been shown to have anticancer properties, likely due to its high level of antioxidant activity [
72,
73]. CA appears to be bioavailable, being present in urine following consumption [
39]. This paper demonstrates the presence of CA in blueberry and suggests possible breeding avenues to increase CA content, with potential health benefits for consumers.
Each of the tested phenolic compounds contains a caffeoyl moiety (
Figure 1A) as a component, suggesting that the significantly associated region encodes a gene responsible for the esterification of caffeic acid with various R groups. Further investigation could be directed at identifying other esters of caffeic acid in blueberry and mapping them to see if they co-locate.
Cultivated blueberry consists mostly of tetraploid
V. corymbosum, while the population tested here was developed from crosses between diploids,
V. corymbosum var.
caesariense and
V. darrowii plants. The observed difference in the grandparents, where OPB-8 and OPB-15 (
V. corymbosum var.
caesariense) showed elevated ACQA1 and ACQA2 concentrations and NJ88-14-03 and NJ88-12-41 (
V. darrowii) showed elevated CGA and CA concentrations, could represent different strategies by the species to combat oxidative stress. In addition, CGA is known to be bioavailable in humans [
14,
74,
75] while to the authors’ knowledge the bioavailability of ACQA1 and ACQA2 has not been tested. While further studies are required to determine the bioavailability of the ACQA compounds, should they be less bioavailable in humans or show lower antioxidant activity, breeding to increase CGA levels in blueberry cultivars could improve nutritional value of the berries. Wang et al. [
51] compared various wild diploid species along with tetraploid cultivars of
V. corymbosum, showing that the chemical composition of the tetraploids was distinct from that of the tested diploid species for anthocyanin and flavanol glycoside content. This could indicate that introgression from a variety collection, such as the grandparents of the BNJ16-4 population, would be of use in introducing variation to breeding programs.