Exploration of the Misfolding Mechanism of Transthyretin Monomer: Insights from Hybrid-Resolution Simulations and Markov State Model Analysis
Abstract
1. Introduction
2. Materials and Methods
2.1. Preparation of Initial Structures
2.2. Molecular Dynamics Simulations
2.3. Markov State Model and Construction and Validation
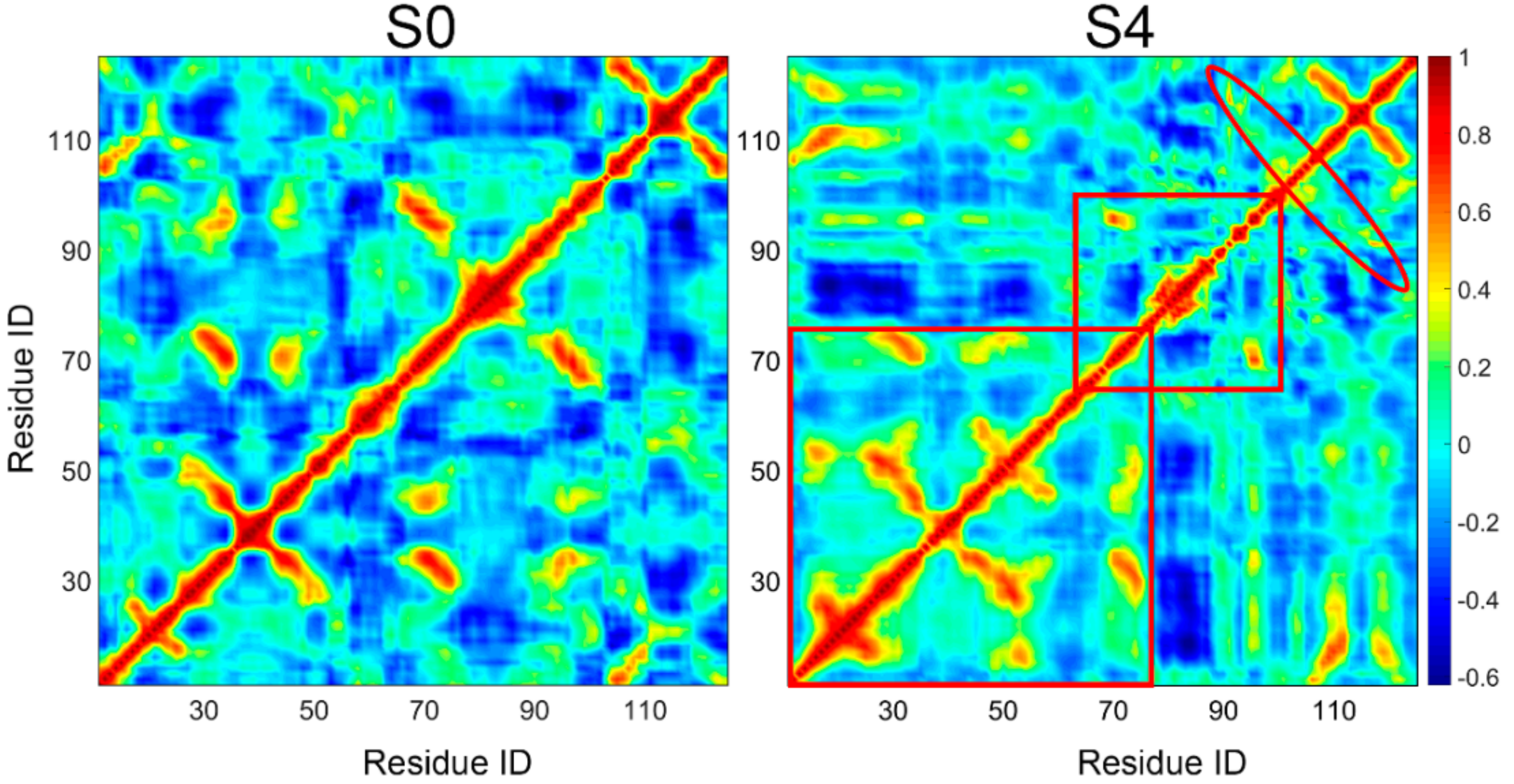
2.4. Dynamical Cross-Correlation Map Analysis
3. Results and Discussion
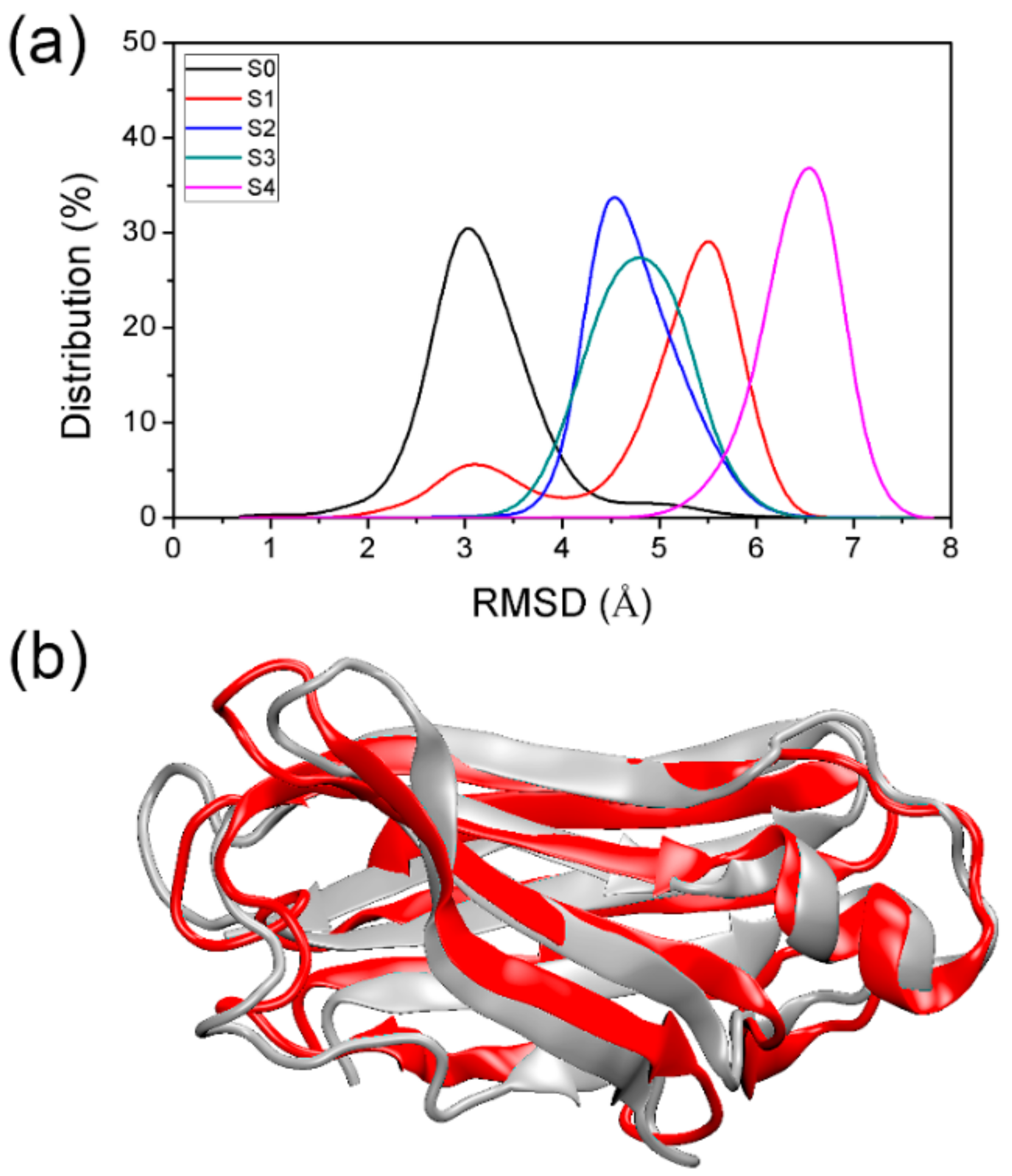
3.1. Comparison of PACE Simulation and All-Atom Simulation
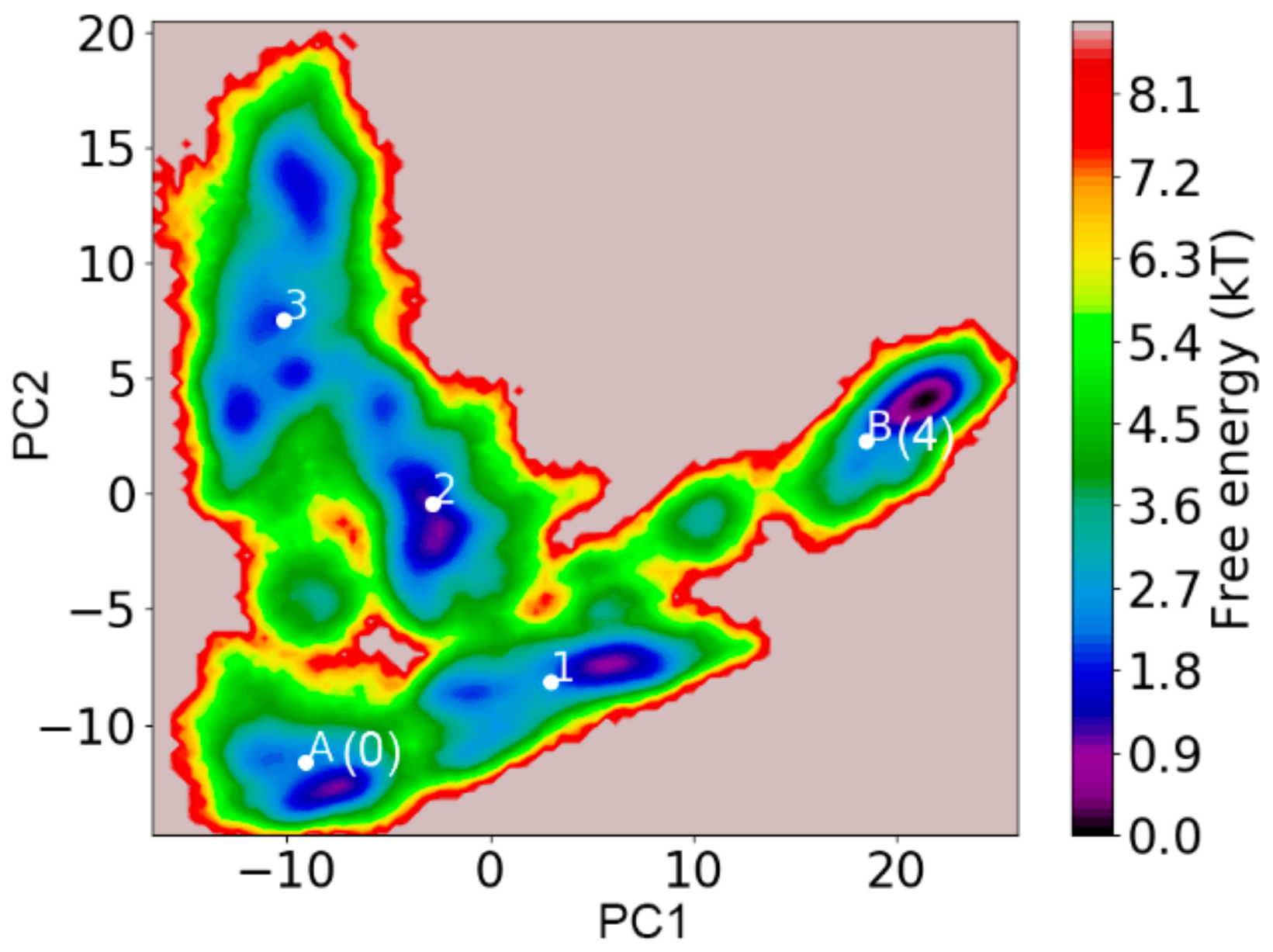
3.2. Validation of Constructed Markov State Model
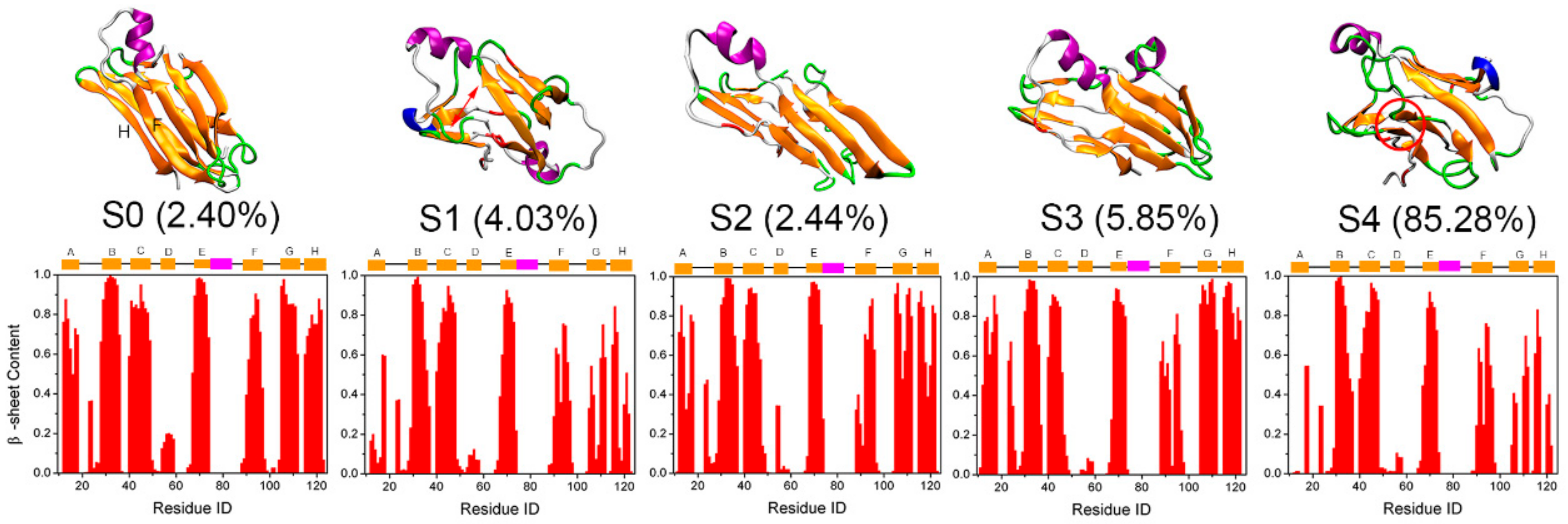
3.3. Structural Ensemble of Key States of Transthyretin Monomer Misfolding
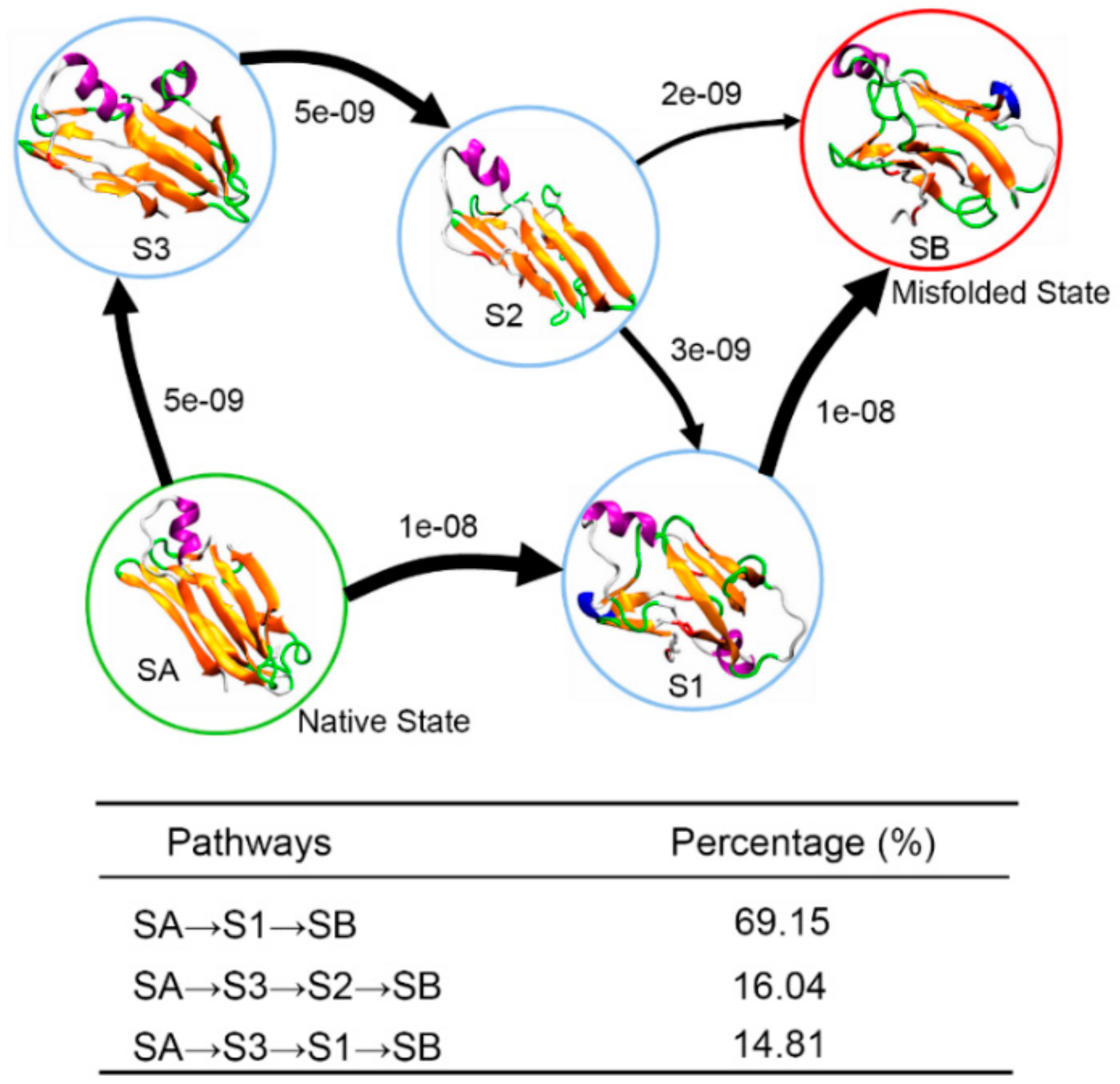
3.4. Insights into the Misfolding Mechanisms of TTR Monomer
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Koike, H.; Katsuno, M. Ultrastructure in transthyretin amyloidosis: From pathophysiology to therapeutic insights. Biomedicines 2019, 7, 11. [Google Scholar] [CrossRef]
- Westermark, P.; Sletten, K.; Johansson, B.; Cornwell, G.G. Fibril in senile systemic amyloidosis is derived from normal transthyretin. Proc. Natl. Acad. Sci. USA 1990, 87, 2843–2845. [Google Scholar] [CrossRef]
- Ando, Y.; Nakamura, M.; Araki, S. Transthyretin-related familial amyloidotic polyneuropathy. Arch. Neurol. 2005, 62, 1057–1062. [Google Scholar] [CrossRef]
- Planté-Bordeneuve, V.; Said, G. Familial amyloid polyneuropathy. Lancet Neurol. 2011, 10, 1086–1097. [Google Scholar] [CrossRef]
- Benson, M.D. Pathogenesis of transthyretin amyloidosis. Amyloid 2012, 19, 14–15. [Google Scholar] [CrossRef]
- Benson, M.D.; Uemichi, T. Transthyretin amyloidosis. Amyloid 1996, 3, 44–56. [Google Scholar] [CrossRef]
- Schneider, F.; Hammarström, P.; Kelly, J.W. Transthyretin slowly exchanges subunits under physiological conditions: A convenient chromatographic method to study subunit exchange in oligomeric proteins. Protein Sci. 2001, 10, 1606–1613. [Google Scholar] [CrossRef] [PubMed]
- Johnson, S.M.; Wiseman, R.L.; Sekijima, Y.; Green, N.S.; Adamski-Werner, S.L.; Kelly, J.W. Native state kinetic stabilization as a strategy to ameliorate protein misfolding diseases: A focus on the transthyretin amyloidoses. Acc. Chem. Res. 2005, 38, 911–921. [Google Scholar] [CrossRef] [PubMed]
- Lai, Z.; Colón, W.; Kelly, J.W. The acid-mediated denaturation pathway of transthyretin yields a conformational intermediate that can self-assemble into amyloid. Biochemistry 1996, 35, 6470–6482. [Google Scholar] [CrossRef]
- Quintas, A.; Vaz, D.C.; Cardoso, I.; Saraiva, M.J.M.; Brito, R.M. Tetramer dissociation and monomer partial unfolding precedes protofibril formation in amyloidogenic transthyretin variants. J. Biol. Chem. 2001, 276, 27207–27213. [Google Scholar] [CrossRef]
- Lim, K.H.; Dasari, A.K.; Hung, I.; Gan, Z.; Kelly, J.W.; Wemmer, D.E. Structural changes associated with transthyretin misfolding and amyloid formation revealed by solution and solid-state NMR. Biochemistry 2016, 55, 1941–1944. [Google Scholar] [CrossRef] [PubMed]
- Lim, K.H.; Dyson, H.J.; Kelly, J.W.; Wright, P.E. Localized structural fluctuations promote amyloidogenic conformations in transthyretin. J. Mol. Biol. 2013, 425, 977–988. [Google Scholar] [CrossRef] [PubMed]
- Lim, K.H.; Dasari, A.K.; Ma, R.; Hung, I.; Gan, Z.; Kelly, J.W.; Fitzgerald, M.C. Pathogenic Mutations Induce Partial Structural Changes in the Native β-Sheet Structure of Transthyretin and Accelerate Aggregation. Biochemistry 2017, 56, 4808–4818. [Google Scholar] [CrossRef] [PubMed]
- Dasari, A.K.; Hung, I.; Gan, Z.; Lim, K.H. Two distinct aggregation pathways in transthyretin misfolding and amyloid formation. BBA-Proteins Proteom. 2019, 1867, 344–349. [Google Scholar] [CrossRef]
- Koike, H.; Ikeda, S.; Takahashi, M.; Kawagashira, Y.; Iijima, M.; Misumi, Y.; Ando, Y.; Ikeda, S.-i.; Katsuno, M.; Sobue, G. Schwann cell and endothelial cell damage in transthyretin familial amyloid polyneuropathy. Neurology 2016, 87, 2220–2229. [Google Scholar] [CrossRef] [PubMed]
- Koike, H.; Nishi, R.; Ikeda, S.; Kawagashira, Y.; Iijima, M.; Sakurai, T.; Shimohata, T.; Katsuno, M.; Sobue, G. The morphology of amyloid fibrils and their impact on tissue damage in hereditary transthyretin amyloidosis: An ultrastructural study. J. Neurol. Sci. 2018, 394, 99–106. [Google Scholar] [CrossRef]
- Ortore, G.; Martinelli, A. Computational studies on transthyretin. Curr. Med. Chem. 2012, 19, 2380–2387. [Google Scholar] [CrossRef]
- Das, J.K.; Mall, S.S.; Bej, A.; Mukherjee, S. Conformational Flexibility Tunes the Propensity of Transthyretin to Form Fibrils Through Non-Native Intermediate States. Angew. Chem. Int. Ed. 2014, 53, 12781–12784. [Google Scholar] [CrossRef]
- Yee, A.W.; Aldeghi, M.; Blakeley, M.P.; Ostermann, A.; Mas, P.J.; Moulin, M.; De Sanctis, D.; Bowler, M.W.; Mueller-Dieckmann, C.; Mitchell, E.P. A molecular mechanism for transthyretin amyloidogenesis. Nat. Commun. 2019, 10, 925. [Google Scholar] [CrossRef]
- Rodrigues, J.R.; Simões, C.J.; Silva, C.G.; Brito, R.M. Potentially amyloidogenic conformational intermediates populate the unfolding landscape of transthyretin: Insights from molecular dynamics simulations. Protein Sci. 2010, 19, 202–219. [Google Scholar] [CrossRef]
- Armen, R.S.; Alonso, D.O.; Daggett, V. Anatomy of an amyloidogenic intermediate: Conversion of β-sheet to α-sheet structure in transthyretin at acidic pH. Structure 2004, 12, 1847–1863. [Google Scholar] [CrossRef] [PubMed]
- Ben-Nissan, G.; Sharon, M. Capturing protein structural kinetics by mass spectrometry. Chem. Soc. Rev. 2011, 40, 3627–3637. [Google Scholar] [CrossRef] [PubMed]
- Kumar, A.; Balbach, J. Real-time protein NMR spectroscopy and investigation of assisted protein folding. BBA Gen. Subj. 2015, 1850, 1965–1972. [Google Scholar] [CrossRef] [PubMed]
- Han, W.; Schulten, K. Further optimization of a hybrid united-atom and coarse-grained force field for folding simulations: Improved backbone hydration and interactions between charged side chains. J. Chem. Theory Comput. 2012, 8, 4413–4424. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Han, W.; Schulten, K. Characterization of folding mechanisms of Trp-cage and WW-domain by network analysis of simulations with a hybrid-resolution model. J. Phys. Chem. B 2013, 117, 13367–13377. [Google Scholar] [CrossRef]
- Han, W.; Wan, C.-K.; Wu, Y.-D. PACE force field for protein simulations. 2. Folding simulations of peptides. J. Chem. Theory Comput. 2010, 6, 3390–3402. [Google Scholar] [CrossRef]
- Qi, Y.; Cheng, X.; Han, W.; Jo, S.; Schulten, K.; Im, W. CHARMM-GUI PACE CG Builder for solution, micelle, and bilayer coarse-grained simulations. J. Chem. Inf. Model. 2014, 54, 1003–1009. [Google Scholar] [CrossRef]
- Yu, H.; Han, W.; Ma, W.; Schulten, K. Transient β-hairpin formation in α-synuclein monomer revealed by coarse-grained molecular dynamics simulation. J. Chem. Phys. 2015, 143, 12B623_621. [Google Scholar] [CrossRef]
- Han, W.; Schulten, K. Fibril elongation by Aβ17–42: Kinetic network analysis of hybrid-resolution molecular dynamics simulations. J. Am. Chem. Soc. 2014, 136, 12450–12460. [Google Scholar] [CrossRef]
- Prinz, J.-H.; Wu, H.; Sarich, M.; Keller, B.; Senne, M.; Held, M.; Chodera, J.D.; Schütte, C.; Noé, F. Markov models of molecular kinetics: Generation and validation. J. Chem. Phys. 2011, 134, 174105. [Google Scholar] [CrossRef]
- Shukla, D.; Hernández, C.X.; Weber, J.K.; Pande, V.S. Markov state models provide insights into dynamic modulation of protein function. Acc. Chem. Res. 2015, 48, 414–422. [Google Scholar] [CrossRef] [PubMed]
- Bowman, G.R.; Pande, V.S.; Noé, F. An Introduction to Markov State Models and Their Application to Long Timescale Molecular Simulation; Springer Science & Business Media: Berlin/Heidelberg, Germany, 2013; Volume 797. [Google Scholar]
- Haupt, M.; Blakeley, M.P.; Fisher, S.J.; Mason, S.A.; Cooper, J.B.; Mitchell, E.P.; Forsyth, V.T. Binding site asymmetry in human transthyretin: Insights from a joint neutron and X-ray crystallographic analysis using perdeuterated protein. IUCrJ 2014, 1, 429–438. [Google Scholar] [CrossRef] [PubMed]
- Jo, S.; Kim, T.; Iyer, V.G.; Im, W. CHARMM-GUI: A web-based graphical user interface for CHARMM. J. Comput. Chem. 2008, 29, 1859–1865. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.; MacKerell, A.D., Jr. CHARMM36 all-atom additive protein force field: Validation based on comparison to NMR data. J. Comput. Chem. 2013, 34, 2135–2145. [Google Scholar] [CrossRef]
- Humphrey, W.; Dalke, A.; Schulten, K. VMD: Visual molecular dynamics. J. Mol. Graph. 1996, 14, 33–38. [Google Scholar] [CrossRef]
- Phillips, J.C.; Braun, R.; Wang, W.; Gumbart, J.; Tajkhorshid, E.; Villa, E.; Chipot, C.; Skeel, R.D.; Kale, L.; Schulten, K. Scalable molecular dynamics with NAMD. J. Comput. Chem. 2005, 26, 1781–1802. [Google Scholar] [CrossRef]
- Jorgensen, W.L.; Chandrasekhar, J.; Madura, J.D.; Impey, R.W.; Klein, M.L. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 1983, 79, 926–935. [Google Scholar] [CrossRef]
- Essmann, U.; Perera, L.; Berkowitz, M.L.; Darden, T.; Lee, H.; Pedersen, L.G. A smooth particle mesh Ewald method. J. Chem. Phys. 1995, 103, 8577–8593. [Google Scholar] [CrossRef]
- Beauchamp, K.A.; McGibbon, R.; Lin, Y.-S.; Pande, V.S. Simple few-state models reveal hidden complexity in protein folding. Proc. Natl. Acad. Sci. USA 2012, 109, 17807–17813. [Google Scholar] [CrossRef]
- Schor, M.; Mey, A.S.; Noé, F.; MacPhee, C.E. Shedding light on the dock–lock mechanism in amyloid fibril growth using Markov state models. J. Phys. Chem. Lett. 2015, 6, 1076–1081. [Google Scholar] [CrossRef]
- Swope, W.C.; Pitera, J.W.; Suits, F. Describing protein folding kinetics by molecular dynamics simulations. 1. Theory. J. Phys. Chem. B 2004, 108, 6571–6581. [Google Scholar] [CrossRef]
- Bowman, G.R.; Huang, X.; Pande, V.S. Using generalized ensemble simulations and Markov state models to identify conformational states. Methods 2009, 49, 197–201. [Google Scholar] [CrossRef] [PubMed]
- Zheng, W.; Gallicchio, E.; Deng, N.; Andrec, M.; Levy, R.M. Kinetic network study of the diversity and temperature dependence of trp-cage folding pathways: Combining transition path theory with stochastic simulations. J. Phys. Chem. B 2011, 115, 1512–1523. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Röblitz, S.; Weber, M. Fuzzy spectral clustering by PCCA+: Application to Markov state models and data classification. Adv. Data Anal. Classi. 2013, 7, 147–179. [Google Scholar] [CrossRef]
- Scherer, M.K.; Trendelkamp-Schroer, B.; Paul, F.; Pérez-Hernández, G.; Hoffmann, M.; Plattner, N.; Wehmeyer, C.; Prinz, J.-H.; Noé, F. PyEMMA 2: A software package for estimation, validation, and analysis of Markov models. J. Chem. Theory Comput. 2015, 11, 5525–5542. [Google Scholar] [CrossRef]
- Kasahara, K.; Fukuda, I.; Nakamura, H. A novel approach of dynamic cross correlation analysis on molecular dynamics simulations and its application to Ets1 dimer–DNA complex. PLoS ONE 2014, 9, e112419. [Google Scholar] [CrossRef]
- Jin, X.; Shao, Y.; Bai, Q.; Xue, W.; Liu, H.; Yao, X. Insights into conformational regulation of PfMATE transporter from Pyrococcus furiosus induced by alternating protonation state of Asp41 residue: A molecular dynamics simulation study. BBA Gen. Subj. 2016, 1860, 1173–1180. [Google Scholar] [CrossRef]
- Mhlongo, N.N.; Ebrahim, M.; Skelton, A.A.; Kruger, H.G.; Williams, I.H.; Soliman, M.E. Dynamics of the thumb-finger regions in a GH11 xylanase Bacillus circulans: Comparison between the Michaelis and covalent intermediate. RSC Adv. 2015, 5, 82381–82394. [Google Scholar] [CrossRef]
- Ichiye, T.; Karplus, M. Collective motions in proteins: A covariance analysis of atomic fluctuations in molecular dynamics and normal mode simulations. Proteins 1991, 11, 205–217. [Google Scholar] [CrossRef]
- Jazaj, D.; Ghadami, S.A.; Bemporad, F.; Chiti, F. Probing conformational changes of monomeric transthyretin with second derivative fluorescence. Sci. Rep. 2019, 9, 10988. [Google Scholar] [CrossRef]
- Freddolino, P.L.; Park, S.; Roux, B.; Schulten, K. Force field bias in protein folding simulations. Biophys. J. 2009, 96, 3772–3780. [Google Scholar] [CrossRef]
- Higo, J.; Ito, N.; Kuroda, M.; Ono, S.; Nakajima, N.; Nakamura, H. Energy landscape of a peptide consisting of α-helix, 310-helix, β-turn, β-hairpin, and other disordered conformations. Protein Sci. 2001, 10, 1160–1171. [Google Scholar] [CrossRef]
- Malmstrom, R.D.; Lee, C.T.; Van Wart, A.T.; Amaro, R.E. Application of molecular-dynamics based markov state models to functional proteins. J. Chem. Theory Comput. 2014, 10, 2648–2657. [Google Scholar] [CrossRef]
- Liu, K.; Cho, H.S.; Hoyt, D.W.; Nguyen, T.N.; Olds, P.; Kelly, J.W.; Wemmer, D.E. Deuterium-proton exchange on the native wild-type transthyretin tetramer identifies the stable core of the individual subunits and indicates mobility at the subunit interface. J. Mol. Biol. 2000, 303, 555–565. [Google Scholar] [CrossRef] [PubMed]
- Saelices, L.; Johnson, L.M.; Liang, W.Y.; Sawaya, M.R.; Cascio, D.; Ruchala, P.; Whitelegge, J.; Jiang, L.; Riek, R.; Eisenberg, D.S. Uncovering the mechanism of aggregation of human transthyretin. J. Biol. Chem. 2015, 290, 28932–28943. [Google Scholar] [CrossRef] [PubMed]
- Conti, S.; Li, X.; Gianni, S.; Ghadami, S.A.; Buxbaum, J.; Cecchi, C.; Chiti, F.; Bemporad, F. A complex equilibrium among partially unfolded conformations in monomeric transthyretin. Biochemistry 2014, 53, 4381–4392. [Google Scholar] [CrossRef]
- Berezhkovskii, A.; Hummer, G.; Szabo, A. Reactive flux and folding pathways in network models of coarse-grained protein dynamics. J. Chem. Phys. 2009, 130, 05B614. [Google Scholar] [CrossRef] [PubMed]
- Metzner, P.; Schütte, C.; Vanden-Eijnden, E. Transition path theory for Markov jump processes. Multiscale Model. Simul. 2009, 7, 1192–1219. [Google Scholar] [CrossRef]
- Huang, X.; Yao, Y.; Bowman, G.R.; Sun, J.; Guibas, L.J.; Carlsson, G.; Pande, V.S. Constructing multi-resolution Markov state models (MSMs) to elucidate RNA hairpin folding mechanisms. In Biocomputing 2010; World Scientific: Singapore, 2010; pp. 228–239. [Google Scholar]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhou, S.; Cheng, J.; Yang, T.; Ma, M.; Zhang, W.; Yuan, S.; Lo, G.V.; Dou, Y. Exploration of the Misfolding Mechanism of Transthyretin Monomer: Insights from Hybrid-Resolution Simulations and Markov State Model Analysis. Biomolecules 2019, 9, 889. https://doi.org/10.3390/biom9120889
Zhou S, Cheng J, Yang T, Ma M, Zhang W, Yuan S, Lo GV, Dou Y. Exploration of the Misfolding Mechanism of Transthyretin Monomer: Insights from Hybrid-Resolution Simulations and Markov State Model Analysis. Biomolecules. 2019; 9(12):889. https://doi.org/10.3390/biom9120889
Chicago/Turabian StyleZhou, Shuangyan, Jie Cheng, Ting Yang, Mingyue Ma, Wenying Zhang, Shuai Yuan, Glenn V. Lo, and Yusheng Dou. 2019. "Exploration of the Misfolding Mechanism of Transthyretin Monomer: Insights from Hybrid-Resolution Simulations and Markov State Model Analysis" Biomolecules 9, no. 12: 889. https://doi.org/10.3390/biom9120889
APA StyleZhou, S., Cheng, J., Yang, T., Ma, M., Zhang, W., Yuan, S., Lo, G. V., & Dou, Y. (2019). Exploration of the Misfolding Mechanism of Transthyretin Monomer: Insights from Hybrid-Resolution Simulations and Markov State Model Analysis. Biomolecules, 9(12), 889. https://doi.org/10.3390/biom9120889






