Abstract
Effective gene feature selection is critical for enhancing the interpretability and accuracy of genetic data analysis, particularly in the realm of disease prediction and precision medicine. Most evolutionary feature selection algorithms tend to become stuck in local optima and incur high computational costs, particularly when dealing with the complex and high-dimensional nature of genetic data. To address these issues, this study proposes a multitasking feature selection method based on clone selection for high-dimensional microarray data, which identifies optimal features by transferring useful knowledge across two related tasks derived from the same microarray dataset. First, a dual-task generation strategy is designed, where one task selects features based on the Relief-F method, and the other task is generated from the original features. Second, a new mutation operator is introduced to share useful information between the multiple tasks. Finally, an improved clonal selection algorithm is proposed to strengthen the global and local search abilities. The experimental results on six high-dimensional microarray datasets demonstrate that our method significantly outperforms four state-of-the-art feature selection methods, highlighting its effectiveness and efficiency in tackling complex feature selection problems.
1. Introduction
In the fields of medicine and biology, gene feature selection (FS) plays a crucial role in disease diagnosis [1], enhances the accuracy of genetic association studies [2], and advances personalized medicine [3]. This process involves identifying the most relevant genetic features from the original microarray datasets, which helps mitigate the challenges associated with the ‘curse of dimensionality’, thereby typically enhancing classification accuracy and decreasing computational expenses [4].
Feature selection methods are commonly classified into four categories: filter, wrapper, embedded, and hybrid [5]. Filter methods [6] evaluate features based on their inherent characteristics, employing statistical metrics to determine their relevance to the target variable. Wrapper methods [7] select feature subsets by directly evaluating the performance of a particular learning algorithm. Embedded methods [8] integrate feature selection into the model training process itself. They automatically select relevant features while building the model. In general, filter methods are independent of any specific algorithm and offer high computational efficiency, while wrapper methods achieve better accuracy by considering feature interactions; however, they are computationally expensive. In contrast, embedded methods balance efficiency and accuracy by considering feature interactions during training, although they can be complex to implement and are dependent on the chosen model. Hybrid methods [9] combine the strengths of different feature selection techniques, enhancing both accuracy and efficiency.
Clonal selection algorithms (CSAs) [10], based on the natural immune response of the human body to foreign antigens, have been widely used in the field of feature selection [11]. These algorithms have several advantages for feature selection, such as the ability to find optima due to the diversity introduced by the cloning and mutation processes. Moreover, they are easy to implement for feature selection. For example, a two-stage hybrid FS method was proposed in [12], in which Fisher scoring is used to reduce the dimensionality of the search space in the first stage, and then an improved CSA method is employed to select a feature subset. Chai et al. [13] proposed a decomposition multi-objective CSA method for FS, which initializes the population based on symmetric uncertainty and uses a population update strategy to improve the evolutionary process. However, most CSAs are limited by premature convergence and significant computational costs, particularly in the context of high-dimensional data.
Evolutionary multitasking (EMT) [14,15] is an innovative approach for tackling optimization problems, allowing for the simultaneous resolution of multiple related tasks by sharing information across different tasks. Consequently, these approaches provide benefits such as robust search capabilities and rapid convergence rates. Chen et al. [16] proposed a novel high-dimensional feature selection method based on particle swarm optimization, in which evolutionary multitasking is used to enhance feature selection performance and decrease computational costs. It generates two tasks from the same dataset: one task involves selecting features from the whole dataset, while the other focuses on choosing promising features based on Relief-F. Li et al. [17] introduced three filtering methods to improve a multitask generation strategy and employed a competitive swarm optimizer to tackle the four associated tasks by facilitating the transfer of useful knowledge. Lin et al. [18] integrated an innovative searching technique and a transformation method into the evolutionary multitasking framework, significantly boosting the efficacy of the multi-objective feature selection algorithm.
Based on the aforementioned analysis, we propose a multitasking feature selection method based on clone selection for high-dimensional microarray data, called CSA-EMT, which aims to improve the quality of feature subsets and reduce computational costs by introducing evolutionary multitasking. To the best of our knowledge, little attention has been given to combining a clonal selection algorithm with multitasking to improve the performance of high-dimensional feature selection algorithms. The key contributions of this study are summarized as follows:
- (1)
- An effective multitasking feature selection method is proposed, which improves the performance of FS by transferring useful knowledge across two related tasks;
- (2)
- A dual-task generation strategy is adopted, in which one task is constructed using the Relief-F method, while the other is derived from the original features;
- (3)
- A clonal selection algorithm is improved to enhance search capabilities and accelerate the rate of evolution by introducing a new mutation operator that shares useful information between multiple tasks.
The structure of the rest of this paper is as follows. Section 2 reviews the related works on evolutionary multitasking, clonal selection algorithms, and feature selection. Section 3 introduces the proposed CSA-EMT method in detail. Section 4 presents the experimental setup and results. Finally, Section 5 concludes this paper with a discussion of the findings and future directions.
2. Related Works
2.1. Evolutionary Multitasking
Evolutionary multitasking is an emerging research field in optimization that aims to solve multiple optimization tasks simultaneously. Instead of focusing on a single problem, EMT leverages the synergies between different tasks, enabling knowledge transfer across tasks during the evolutionary process. EMT methods are usually classified into two categories: multifactorial-based methods and multi-population methods [19]. Multifactorial-based methods are a type of evolutionary multitasking that use a single population to solve multiple optimization tasks simultaneously. In these methods, each individual in the population is evaluated on multiple tasks, and the knowledge transfer between tasks happens implicitly as the population evolves. Gupta et al. [20] proposed a famous multifactorial evolutionary algorithm (MFEA) that optimizes multiple tasks simultaneously by evolving a single population within a shared search space. The method begins by initializing a population where each individual is associated with a task based on its skill factor. Crossover operations are then performed between individuals with high skill factors from different tasks, enabling cross-domain knowledge transfer. Since then, a large number of variant algorithms have been proposed. For example, Bali et al. [21] proposed an enhanced MFEA, in which an online transfer parameter estimation mechanism is used to allow effective multitasking across multiple tasks, even with varying intertask relationships. Xing et al. [22] developed an adaptive multifactorial evolutionary algorithm for constrained optimization problems, which incorporates an archiving strategy, an adaptive mutation probability, and a mutation strategy. Additionally, empirical research has explored multifactorial differential evolution [23] and multifactorial particle swarm optimization [24].
Multi-population EMT methods are another class of multitasking algorithms, where each task is assigned to a separate population, which evolves independently. The populations are designed to solve specific tasks, but controlled interactions between populations allow for knowledge transfer (such as migration or crossover), enabling the sharing of useful information between tasks. For example, Feng et al. [25] employed explicit genetic transfer using autoencoders to transfer useful information between different tasks, helping improve overall performance across tasks. Lin et al. [26] used an effective positive transfer strategy to improve the convergence of the target task, wherein solutions are transferred from the neighbors of those that achieved successful knowledge transfer. Zhang et al. [27] introduced adaptive dual knowledge transfer into differential evolution, which integrates unified search space-based transfer with domain adaptation-based transfer to enhance the performance of EMT. Li et al. [28] employed dual information transfer alongside a mating strategy to minimize the negative effects of migration. Compared with multifactorial-based methods, multi-population methods allow for task-specific evolution, enabling each population to focus on solving its own task without interference from others. They also carefully control the interactions between populations, allowing for more flexible and targeted knowledge sharing between tasks. Therefore, our algorithm adopts a multi-population EMT pattern.
2.2. Clonal Selection Algorithm
The clonal selection algorithm is an important technique derived from the immune system’s adaptive mechanisms, particularly focusing on the clonal selection theory [29]. In this theory, immune cells recognize antigens and undergo proliferation and mutation to better adapt to detected foreign entities. This biological process is emulated in computational applications by cloning candidate solutions, introducing controlled variations, and then selecting the best-adapted clones. Through iterative selection, clonal expansion, and mutation, the algorithm can search the solution space effectively, avoiding local optima while enhancing global search capabilities.
Initially introduced by de Castro [30], the CSA has been widely used in various optimization tasks due to its ability to maintain diversity while ensuring convergence. In recent years, a large number of variant algorithms have been proposed. Yan et al. [31] used a non-uniform mutation along with an arithmetic crossover to improve the performance of the CSA. In [32], a CSA was combined with a negative selection algorithm to accelerate searches, and network suppression was employed to reduce premature convergence. Wang et al. [33] employed multiple mutation strategies based on differential evolution and an adaptive parameter mechanism derived from [34] to mitigate the semi-blindness caused by hypermutations. To enhance population diversity, Li et al. [35] adopted vertical distances instead of crowding distances to determine the number of clones for each antibody. The effectiveness of CSAs in the field of optimization has been proven, especially when applied to high-dimensional datasets. Therefore, this study utilizes a CSA for feature selection, aiming to identify the most relevant features while maintaining diversity within the population and improving overall search efficiency.
2.3. Feature Selection for Microarray Data
Feature selection is a critical step in the analysis of microarray data, which typically involves a high-dimensional dataset with a large number of gene expressions and a relatively small sample size. The primary goal of feature selection in microarray data is to identify the most relevant genes that contribute significantly to classification or prediction tasks, thereby improving model performance and interpretability while reducing computational costs.
Various methods have been proposed to address the challenge of feature selection in microarray data. These methods can be broadly classified into four categories: filter, wrapper, embedded, and hybrid methods. Filter methods are typically based on statistical techniques that evaluate the relevance of features independently of the learning algorithm. Ouaderhman et al. [36] proposed the Psitop filter FS method for mixed-type DNA microarray data, which ranks features based on Psitop scores derived from two nonparametric association coefficients: Psicor and partial Psicor. Lee et al. [37] proposed a Markov blanket-based multivariate feature ranking method, which considers both feature relevance and redundancy to improve gene selection and microarray data classification accuracy. A least-loss feature selection method was proposed in [38], which reduces data dimensionality by removing weakly correlated variables while maintaining classifier performance through probability-based similarity scoring. Filter methods are computationally efficient and easy to implement, but they evaluate features independently, often ignoring feature interactions, which can result in suboptimal subsets for complex tasks.
Wrapper methods incorporate a learning algorithm to evaluate subsets of features by directly optimizing the performance of a classifier. Techniques such as particle swarm optimization [39], CSAs [40], and gray wolf optimization [41] have been extensively used for wrapper-based feature selection in microarray data. They provide high accuracy by evaluating feature subsets using a learning algorithm but are computationally expensive, especially with high-dimensional data.
Embedded methods perform feature selection during the learning process of a model, and examples include decision trees and regularization techniques. For example, in [42], four embedded FS methods were compared, and the most effective one was selected for the diagnosis of Alzheimer’s disease. Fu et al. [8] proposed an embedded FS method for GMM, which accounts for feature interdependencies and offers feature relevance ranking as a byproduct. These FS methods strike a balance between computational efficiency and effectiveness, as they take into account interactions between features while being integrated with the learning algorithm.
Hybrid methods integrate multiple types of feature selection algorithms, combining the strengths of both filter and wrapper approaches to enhance overall performance. These methods usually first use filter techniques to quickly reduce the number of irrelevant features or improve solution quality. Then, a wrapper method is applied to the reduced feature subset to further refine and select the optimal set by evaluating the performance of a learning algorithm on the selected features. For example, Vommi et al. [43] proposed a filter-wrapper FS method for COVID-19 case classification, which uses Relief-F and fuzzy entropy to remove unimportant features, followed by an improved equilibrium optimizer to further refine the feature subset. Ke et al. [44] proposed a two-stage FS method, which improves the quality of the initial population in the genetic algorithm or ant colony optimization by utilizing the feature ranking information from the first stage.
In summary, compared to related FS approaches, the proposed method offers several key contributions. While most multitasking feature selection methods rely on particle swarm optimization, genetic algorithms, or competitive swarm optimizers, integrating a clonal selection algorithm with multitasking remains relatively unexplored. This integration presents a promising alternative for optimizing high-dimensional feature selection, thanks to its robust search capabilities and ease of implementation. Additionally, we introduce an enhanced clonal selection algorithm that enables efficient knowledge transfer across tasks, population initialization based on feature weights, and an adaptive parameter control mechanism, all of which support robust global and local search performance.
3. The Proposed CSA-EMT Method
This section introduces the CSA-EMT method, covering its general framework, dual-task generation strategy, an improved clonal selection algorithm for enhanced performance, and a runtime complexity analysis, offering a comprehensive feature selection approach for high-dimensional data.
3.1. The General Framework of CSA-EMT
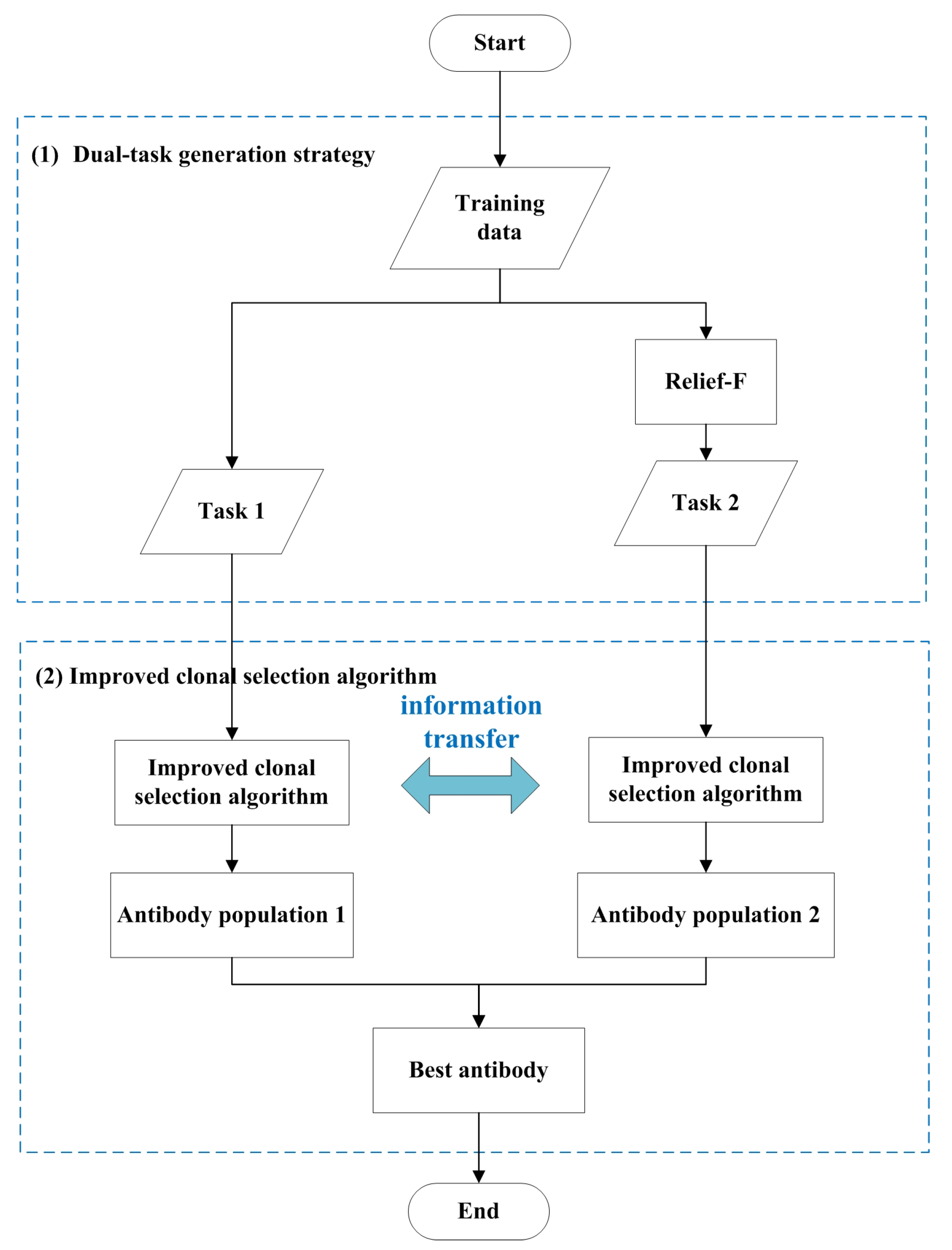
The proposed method integrates an immune algorithm with evolutionary multitasking to optimize gene feature selection. The flowchart of CSA-EMT is shown in Figure 1. It primarily encompasses two critical steps: a dual-task generation strategy and an improved clonal selection algorithm tailored for gene selection. The dual-task generation strategy is initially employed to create two related tasks, as explained in detail in Section 3.2. This approach allows the algorithm to effectively explore diverse search spaces. Next, an improved clonal selection algorithm is applied to optimize feature selection by facilitating knowledge transfer between the two tasks, thereby enhancing the search process and preventing the algorithm from becoming trapped in local optima, as discussed in Section 3.3. In the final step, the best-performing antibodies from both tasks are selected and output as the final solution, ensuring robust and efficient feature selection. The pseudocode for the CSA-EMT framework is shown in Algorithm 1.
| Algorithm 1 Framework of CSA-EMT |
|
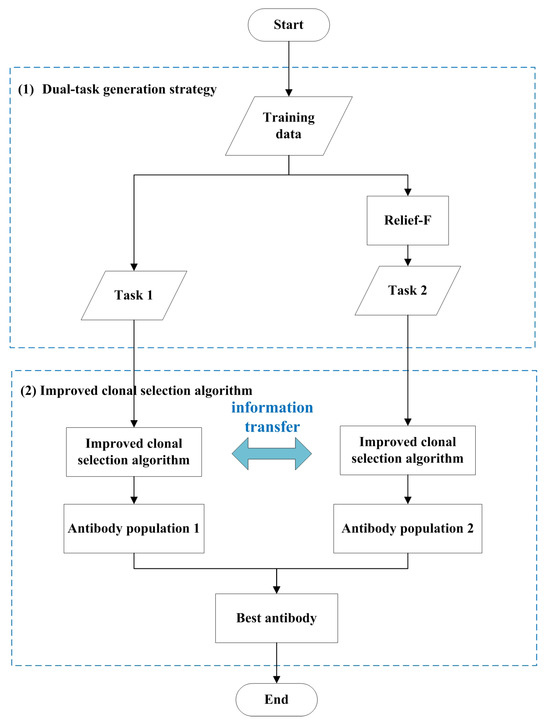
Figure 1.
The general framework of CSA-EMT.
3.2. Dual-Task Generation Strategy
In this study, Task 1 retains all features from the original dataset. Task 2 contains important features identified by the Relief-F and knee point methods. In Task 2, features are ranked according to their weight values, which are computed using the Relief-F algorithm. Then, the knee point method, based on these weights, is used to automatically determine the important features. Features are included in Task 2 if their weight values exceed the threshold defined by the knee point method.
3.3. An Improved Clonal Selection Algorithm
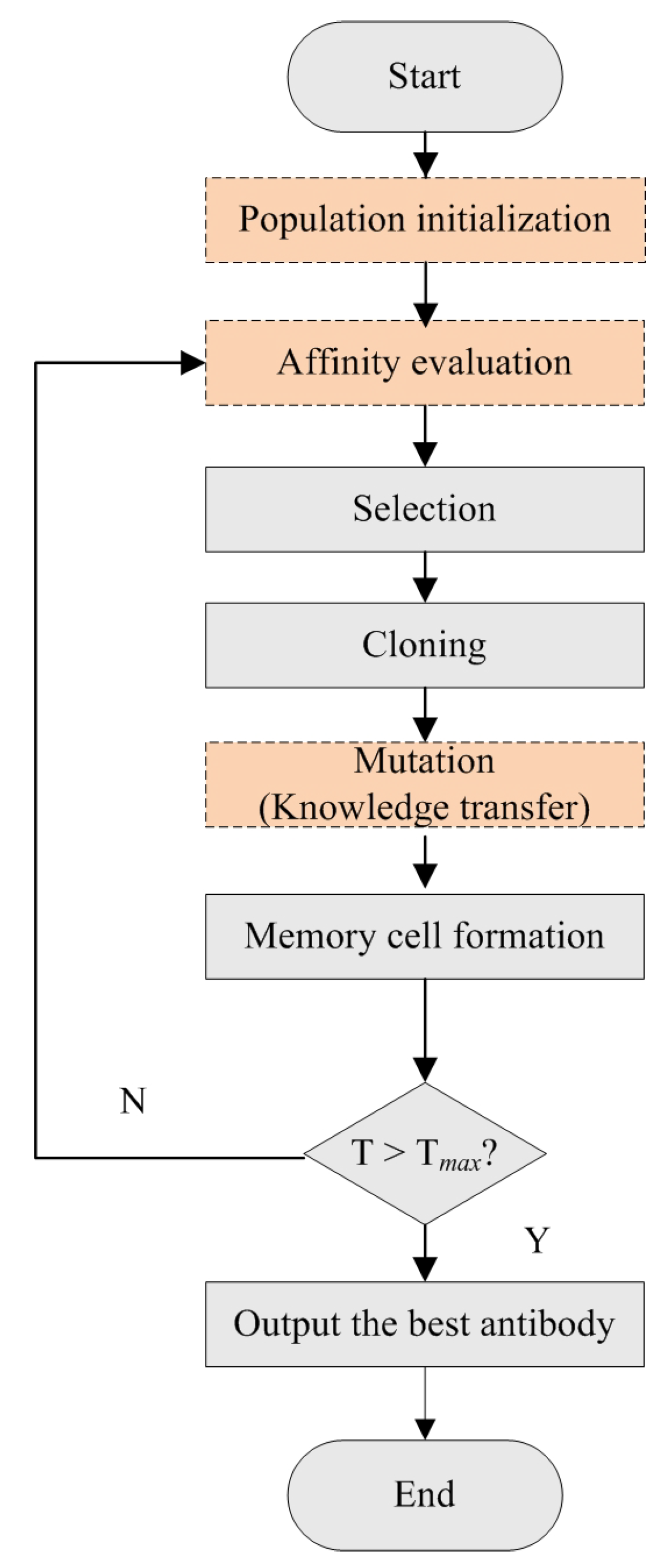
The CSA-EMT method uses a modified clonal selection algorithm to simultaneously address two relevant tasks. The flowchart is shown in Figure 2. It primarily includes population initialization, affinity evaluation, selection, cloning, mutation, and memory cell formation. Specifically, the process begins by generating an initial population of antibodies for Task 1, selected randomly, and for Task 2, based on feature weights. This initialization provides a diverse starting set for optimization. In each iteration, antibodies are evaluated for their affinities, and those with higher affinity values are selected for cloning. Following cloning, each clone undergoes a mutation process. This step introduces diversity by making small alterations and facilitates knowledge transfer between the two tasks, enabling the algorithm to explore a broader range of potential solutions and avoid local optima. Finally, the best clones are selected to form the updated population, replacing the weakest antibodies from the previous generation. This iterative process ensures that the population’s overall quality improves with each generation and continues until a termination condition is met. Here, we focus on introducing the improvement steps (population initialization, an affinity function, and a mutation strategy), which are described as follows:
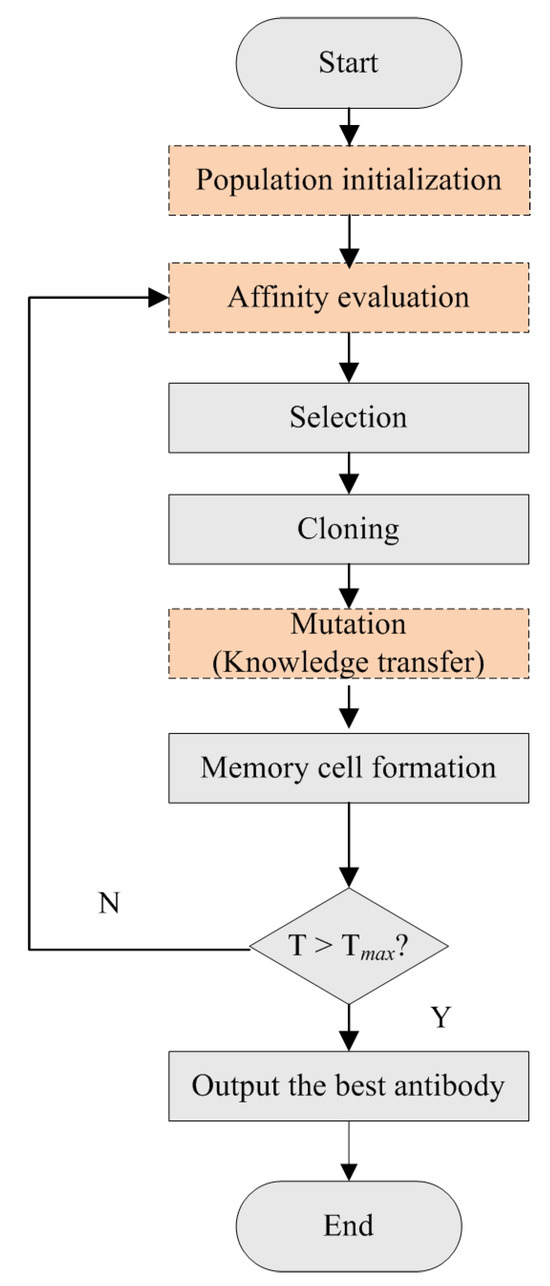
Figure 2.
The flowchart of the improved clonal selection algorithm.
(1) Population initialization
Real numbers are used to characterize the feature masks of antibodies, which allow for smoother mutation operations in continuous space. When the value is greater than 0.5, the corresponding feature is selected; otherwise, it is not selected. For Task 1, each component of the antibody is randomly generated from a uniform distribution in the [0, 1] range. For Task 2, each component corresponds to a normalized weight value determined by the Relief-F method.
(2) Affinity function
CSA-EMT uses Equation (1) as the affinity function, which aims to minimize the error rate and the number of selected features:
where represents the balanced error rate, which offers particular benefits in handling imbalanced datasets. It calculates the average error rate for each class and ensures that all classes contribute equally to the final error rate, regardless of their frequency in the dataset. C is the number of classes, and is the accuracy for class i. The right-hand side of the equation represents the ratio of the number of selected features () to the total number of features (). The value of is set to 0.999999.
(3) Mutation strategy
Knowledge is transferred between Task 1 and Task 2 through the mutation strategy applied to each antibody, which can improve the quality of antibodies and lead to faster convergence. During the mutation process, a transfer probability, , is predefined to determine whether to perform information exchange between the two tasks. Then, a random number is generated between 0 and 1. If its value is greater than , the knowledge transfer will occur using Equation (3); otherwise, antibodies will undergo mutation without knowledge transfer, using Equation (4).
These mutation operators are DE-based mutation strategies, which use superior and inferior antibodies to guide the search directions. Two integers, and , are randomly selected from the range of 1 to the population size. They are distinct from each other and neither equals i. The antibodies and are randomly selected from the best-performing antibodies of the current task and from all tasks, respectively. The antibody is randomly selected from a set that combines an external archive of inferior solutions with the current population. is a scaling factor for the ith antibody within the range [0, 1]. After mutation, the mutant antibodies will undergo crossover:
where j denotes the jth feature of the ith antibody. The parameter refers to the crossover probability of the ith antibody, with a value that ranges between 0 and 1. The term is randomly selected between 1 and the number of features.
The parameter update process for and in SHADE 1.1 [45] is retained, with the modification that the last elements in the memory pools start with a value of 0.9. In particular, it initializes with two memory pools that store the historically successful values of and . Each element, except for the last one, is initialized with a value of 0.5, while the last element is set to 0.9. In each generation, the control parameters and are computed based on the following equations:
is sampled from a Cauchy distribution with a location parameter and a scale parameter of 0.1. If the sampled value exceeds 1, it is clipped to 1. If the sampled value is less than 0, it is resampled from the Cauchy distribution until it falls within the valid range (0, 1]. The index k is randomly selected from 1 to the memory pool size. is sampled from a normal distribution, with a mean of and a standard deviation of 0.1, and the clipped value stays within [0, 1].
After each generation, our method collects information about F and from the successful individuals and updates the memory pools using Algorithm 2.
| Algorithm 2 Memory update process |
|
The function computes the weighted Lehmer mean of the successful F and values, as follows:
|.| is the improvement in fitness between the parent and the offspring.
3.4. Runtime Complexity Analysis
The analysis of the time complexity of CSA-EMT is as follows:
Step 1: Generate dual tasks by separately applying Relief-F and using the original training data with m samples and d features. The time complexity follows , where s represents the number of times the algorithm randomly samples instances from the dataset.
Step 2.1: Initialize the population with N antibodies and evaluate the affinity values. The time complexity follows , where E represents the time taken for affinity evaluation.
Step 2.2: Select the best antibodies for cloning, which has a time complexity of .
Step 2.3: Each selected antibody is cloned times, which has a time complexity of .
Step 2.4: Each clone undergoes mutation and has its affinity value evaluated. The time complexity follows .
Step 2.5: Update the population, which has a time complexity of .
Step 2.6: The evolutionary process runs for G generations.
The total time complexity is as follows:
=
From this, it is clear that the population size N, the number of selected antibodies , the number of clones , and the number of evolutionary iterations G are all key factors that impact the algorithm’s time complexity. As our algorithm can achieve faster convergence with a smaller population size N, it can reduce the number of evolutionary iterations and effectively decrease the training time.
4. Experimental Results and Discussion
This section presents three experiments to evaluate the performance of CSA-EMT. First, we compare CSA-EMT with the 1NN classifier using the full feature set. Second, we benchmark CSA-EMT against other well-established feature selection methods. Finally, we examine the impact of different generation strategies on the performance of CSA-EMT.
4.1. Experimental Setup
For this study, we utilized six microarray datasets to evaluate the performance of the proposed CSA-EMT method. The details are provided in Table 1 and are available at https://github.com/lyceia/FS-DB (accessed on 20 October 2024). The training and test sets were generated using 10-fold cross-validation, which is widely used in feature selection due to its ability to effectively balance bias and variance [17,46]. In this method, nine folds are designated for training, and the remaining fold is used for testing. Furthermore, we employed stratified sampling to ensure that the class distribution remained consistent during the splitting of the datasets, which helps mitigate the impact of class imbalance during cross-validation.

Table 1.
The details of the benchmark datasets.
The experiments were conducted on a system with an Intel Core i9 processor and 32 GB of RAM in a Windows 11 environment. The CSA-EMT algorithm was implemented using the JAVA WekaClassalgos development package, which can be accessed at http://wekaclassalgos.sourceforge.net (accessed on 20 October 2024). The main parameters were as follows: the population size for each task was set to 5; the top 20% of antibodies were selected for cloning, with each selected antibody producing one clone; the historical memory pool size for each task was set to 5; the maximum number of evolutions was set to 50; and the 1NN classifier was used to evaluate the performance of the methods. Each method was run 20 times on each dataset to reduce bias.
4.2. Performance Evaluation
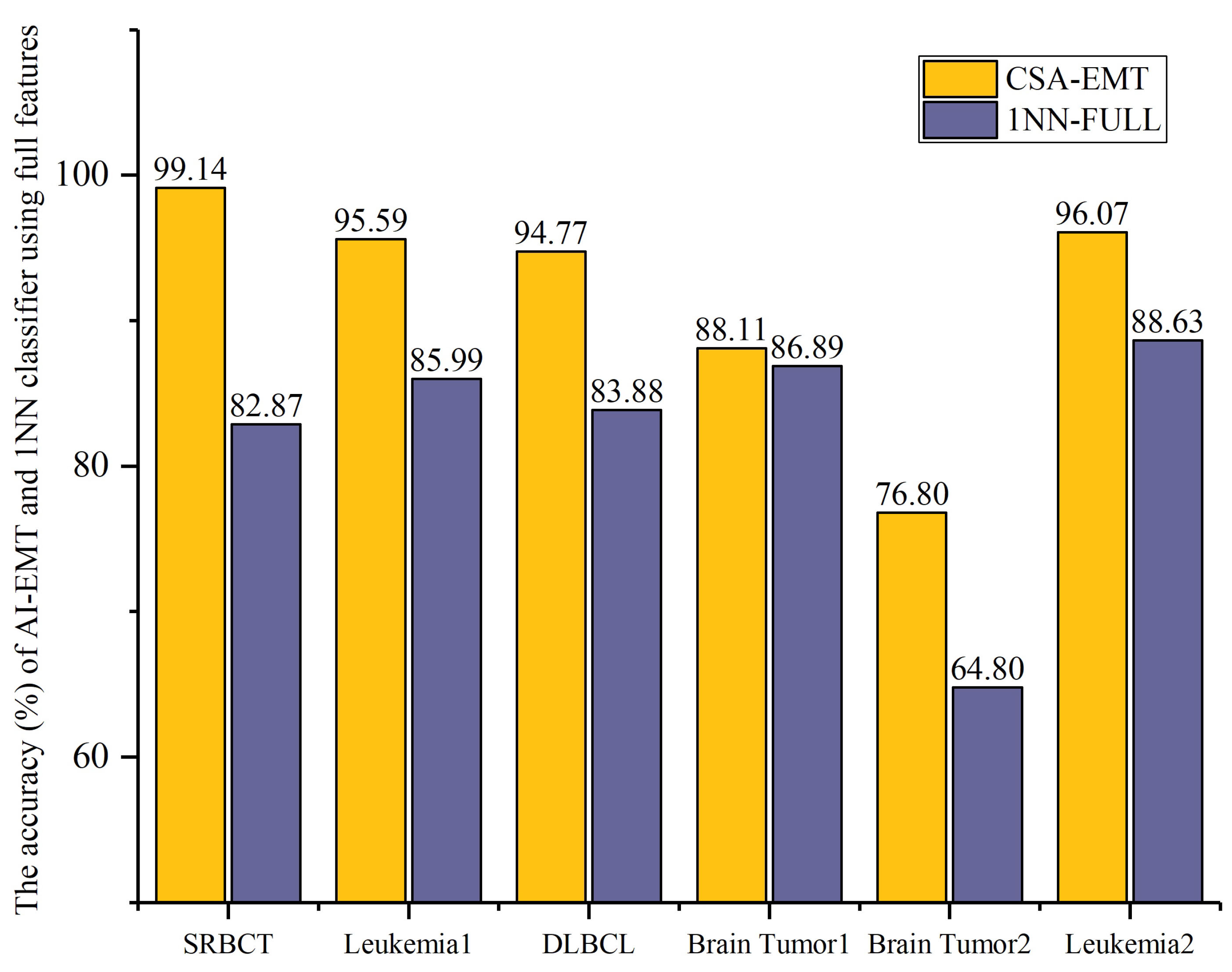
We evaluated the performance of CSA-EMT based on accuracy and the number of selected features. Figure 3 and Figure 4 illustrate the comparative results between CSA-EMT and the 1NN classifier using all features.
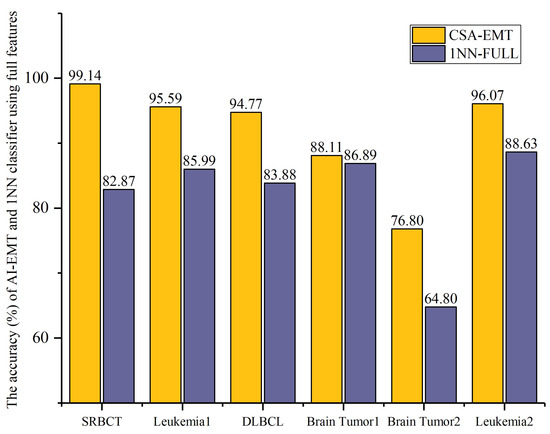
Figure 3.
The accuracy (%) of CSA-EMT vs. that of the 1NN classifier using all features.
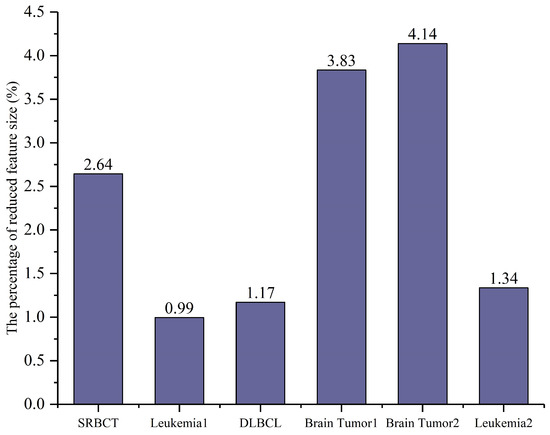
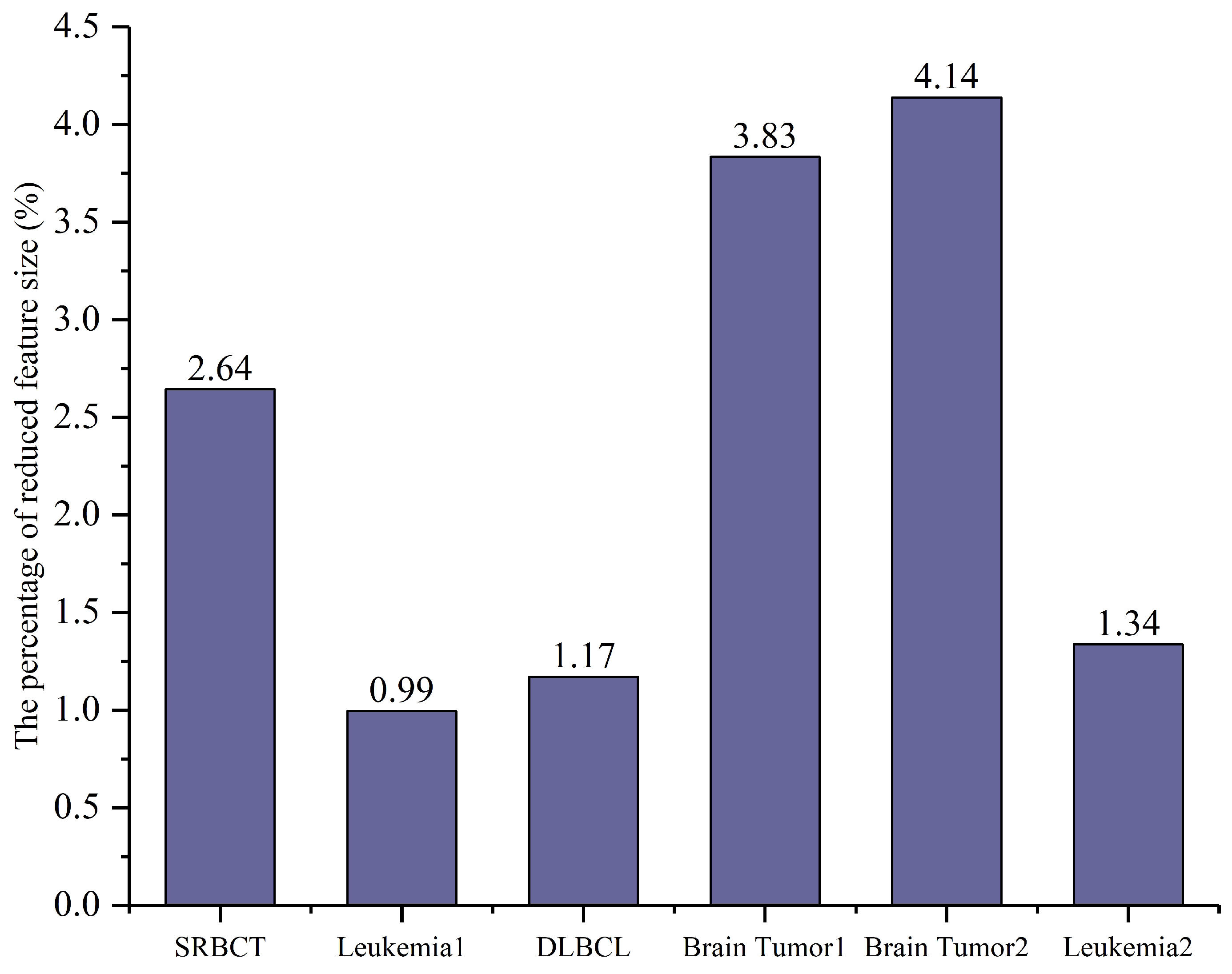
Figure 4.
The percentage of selected features relative to the total number of features (%).
In Figure 3, the accuracy (%) of the proposed CSA-EMT method is compared with that of the 1NN classifier using all feature sets across six datasets. The x-axis represents the datasets, while the y-axis indicates the classification accuracy as a percentage. As shown, CSA-EMT consistently outperformed the 1NN classifier across all datasets, demonstrating its effectiveness in selecting high-quality feature subsets, which improved classification accuracy. Compared to using all features, the classification accuracy increased by at least 7.44% on five datasets, with SRBCT showing the largest improvement, achieving a 16.27% increase.
In Figure 4, the percentage of the average number of selected features relative to the total number of features is presented, with the x-axis representing each dataset and the y-axis indicating the proportion of selected features. The results show that CSA-EMT significantly reduced the feature subset by at least 95.86%, selecting a smaller subset while retaining relevant information. Leukemia1 exhibited the most significant reduction, decreasing by 99.01%.
In summary, these results highlight the robust feature selection capability of CSA-EMT, which significantly reduces dimensionality while enhancing the predictive power of the classifier.
4.3. Comparison with Feature Selection Methods
To demonstrate the effectiveness and efficiency of CSA-EMT, we compared it with four state-of-the-art, well-known feature selection algorithms, including MF-CSO [17], VLPSO-LS [47], the GA (genetic algorithm), and the CSA (clonal selection algorithm). MF-CSO is an evolutionary multitasking-based feature selection method that employs a competitive swarm optimizer to tackle the four tasks generated using three filtering methods. VLPSO-LS is a one-stage feature selection method that employs a variable-length representation to accelerate the search speed. The GA and CSA are well-known evolutionary algorithms with powerful global search abilities.
Table 2 presents the comparison results with other state-of-the-art feature selection methods, where represents the number of selected features, represents the average accuracy, and represents the running time. The symbols “−”, “+”, and “≈” denote whether the compared methods outperformed, underperformed, or performed similarly to CSA-EMT.

Table 2.
Comparison of CSA-EMT with four feature selection methods.
The results indicate that CSA-EMT outperformed the comparative methods in both accuracy and running time on five out of six datasets. Compared with the traditional GA and CSA-based feature selection methods, CSA-EMT not only achieved the highest accuracy but also required fewer features across all datasets. The average accuracy improved by at least 8.07%, except for Brain Tumor1. Compared with VLPSO-LS, although CSA-EMT selected more features on four datasets, it enhanced the performance across all datasets except for SRBCT, surpassing VLPSO-LS in overall effectiveness. This is because CSA-EMT adopts the evolutionary multitasking technique to share information across different tasks. Compared with MF-CSO using EMT, CSA-EMT showed obvious advantages in average accuracy across all datasets. The most notable enhancement was observed in Brain Tumor1, where the accuracy increased by at least 13.18%. Regarding the average running time, CSA-EMT was the top performer, followed by GA, VLPSO-LS, CSA, and MF-CSO.
4.4. Impact of Different Generation Strategies
In this section, we analyze the effect of three generation strategies on the performance of CSA-EMT. We implemented three generation strategies: information gain, mRMR, and Relief-F. Each strategy was tested using the same initial parameters to ensure a fair comparison. Table 3 shows the comparison results of CSA-EMT using different generation strategies. CSA-EMT-IG employs information gain and the knee point method to generate features in Task 2. CSA-EMT-mRMR utilizes the top 10% of the important features generated by mRMR to produce Task 2.

Table 3.
Comparison of CSA-EMT performance using different generation strategies.
The results indicate that CSA-EMT and CSA-EMT-mRMR achieved similar accuracies; however, CSA-EMT selected fewer features and had a shorter running time. CSA-EMT-IG, on the other hand, performed worse overall. These findings suggest that while the Relief-F and knee point methods are effective, other methods, such as mRMR, are also viable for feature selection. The choice of method can impact both the efficiency and accuracy of the results. Therefore, it is essential to explore the use of various filter-based feature selection methods to generate multiple tasks to improve overall performance. Here, we have used only the Relief-F and knee point methods, but we are not limited to these.
5. Conclusions
In this study, we proposed an evolutionary multitasking feature selection approach based on an immune algorithm to address the challenges of high-dimensional microarray data analysis. Multitask learning is introduced to leverage the clonal selection algorithm to solve multiple feature selection tasks simultaneously. By sharing information across tasks, the method enhances convergence and reduces computational costs, resulting in more robust feature selection. The dual-task generation strategy and the improved clonal selection algorithm are the key components of our proposed algorithm. The dual-task generation strategy uses the Relief-F method and the original features to generate two tasks. Meanwhile, the improved clonal selection algorithm enhances the exploration and exploitation capabilities of the CSA. Through experiments on six high-dimensional microarray datasets, the CSA-EMT method demonstrated a better trade-off between the number of selected features, classification accuracy, and computational time when compared to other feature selection methods across most datasets. However, it presented some limitations. Notably, it tended to select slightly more features, as observed in the Brain Tumor1 and Brain Tumor2 datasets. For future research, we plan to enhance the search capabilities of the clone selection process to address this limitation, potentially by incorporating local search techniques or developing a more effective knowledge transfer strategy.
Author Contributions
Conceptualization, Y.W.; Methodology, Y.W.; Validation, Y.W., D.L., and J.Y.; Resources, J.Y.; Writing—original draft, Y.W. and D.L.; Supervision, D.L.; Funding acquisition, Y.W. All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by the National Natural Science Foundation of China (Grant No. 62102146).
Data Availability Statement
The original contributions presented in the study are included in the article. Further inquiries can be directed to the corresponding author.
Conflicts of Interest
The authors declare no conflicts of interest.
References
- Umirzakova, S.; Ahmad, S.; Mardieva, S.; Muksimova, S.; Whangbo, T.K. Deep learning-driven diagnosis: A multi-task approach for segmenting stroke and Bell’s palsy. Pattern Recognit. 2023, 144, 109866. [Google Scholar] [CrossRef]
- Sun, S.; Dong, B.; Zou, Q. Revisiting genome-wide association studies from statistical modelling to machine learning. Briefings Bioinform. 2021, 22, bbaa263. [Google Scholar] [CrossRef] [PubMed]
- Masciocchi, C.; Gottardelli, B.; Savino, M.; Boldrini, L.; Martino, A.; Mazzarella, C.; Massaccesi, M.; Valentini, V.; Damiani, A. Federated cox proportional hazards model with multicentric privacy-preserving LASSO feature selection for survival analysis from the perspective of personalized medicine. In Proceedings of the 2022 IEEE 35th International Symposium on Computer-Based Medical Systems (CBMS), Shenzhen, China, 21–23 July 2022; IEEE: Piscataway, NJ, USA, 2022; pp. 25–31. [Google Scholar]
- Osama, S.; Shaban, H.; Ali, A.A. Gene reduction and machine learning algorithms for cancer classification based on microarray gene expression data: A comprehensive review. Expert Syst. Appl. 2023, 213, 118946. [Google Scholar] [CrossRef]
- Firdaus, F.F.; Nugroho, H.A.; Soesanti, I. A review of feature selection and classification approaches for heart disease prediction. IJITEE (Int. J. Inf. Technol. Electr. Eng.) 2021, 4, 75–82. [Google Scholar] [CrossRef]
- Bommert, A.; Welchowski, T.; Schmid, M.; Rahnenführer, J. Benchmark of filter methods for feature selection in high-dimensional gene expression survival data. Briefings Bioinform. 2022, 23, bbab354. [Google Scholar] [CrossRef] [PubMed]
- Maldonado, J.; Riff, M.C.; Neveu, B. A review of recent approaches on wrapper feature selection for intrusion detection. Expert Syst. Appl. 2022, 198, 116822. [Google Scholar] [CrossRef]
- Fu, Y.; Liu, X.; Sarkar, S.; Wu, T. Gaussian mixture model with feature selection: An embedded approach. Comput. Ind. Eng. 2021, 152, 107000. [Google Scholar] [CrossRef]
- Pramanik, R.; Pramanik, P.; Sarkar, R. Breast cancer detection in thermograms using a hybrid of GA and GWO based deep feature selection method. Expert Syst. Appl. 2023, 219, 119643. [Google Scholar] [CrossRef]
- Haktanirlar Ulutas, B.; Kulturel-Konak, S. A review of clonal selection algorithm and its applications. Artif. Intell. Rev. 2011, 36, 117–138. [Google Scholar] [CrossRef]
- Wang, Y.; Tian, H.; Li, T.; Liu, X. A two-stage clonal selection algorithm for local feature selection on high-dimensional data. Inf. Sci. 2024, 677, 120867. [Google Scholar] [CrossRef]
- Zhu, Y.; Li, W.; Li, T. A hybrid artificial immune optimization for high-dimensional feature selection. Knowl.-Based Syst. 2023, 260, 110111. [Google Scholar] [CrossRef]
- Chai, Z.; Li, W.; Li, Y. Symmetric uncertainty based decomposition multi-objective immune algorithm for feature selection. Swarm Evol. Comput. 2023, 78, 101286. [Google Scholar] [CrossRef]
- Wei, T.; Wang, S.; Zhong, J.; Liu, D.; Zhang, J. A review on evolutionary multitask optimization: Trends and challenges. IEEE Trans. Evol. Comput. 2021, 26, 941–960. [Google Scholar] [CrossRef]
- Wu, X.; Wang, W.; Zhang, T.; Han, H.; Qiao, J. Improved evolutionary multitasking optimization algorithm with similarity evaluation of search behavior. IEEE Trans. Evol. Comput. 2024. [Google Scholar] [CrossRef]
- Chen, K.; Xue, B.; Zhang, M.; Zhou, F. An evolutionary multitasking-based feature selection method for high-dimensional classification. IEEE Trans. Cybern. 2022, 52, 7172–7186. [Google Scholar] [CrossRef]
- Li, L.; Xuan, M.; Lin, Q.; Jiang, M.; Ming, Z.; Tan, K.C. An evolutionary multitasking algorithm with multiple filtering for high-dimensional feature selection. IEEE Trans. Evol. Comput. 2023, 27, 802–816. [Google Scholar] [CrossRef]
- Lin, J.; Chen, Q.; Xue, B.; Zhang, M. Evolutionary multitasking for multi-objective feature selection in classification. IEEE Trans. Evol. Comput. 2023, 1. [Google Scholar] [CrossRef]
- Li, J.Y.; Zhan, Z.H.; Tan, K.C.; Zhang, J. A meta-knowledge transfer-based differential evolution for multitask optimization. IEEE Trans. Evol. Comput. 2021, 26, 719–734. [Google Scholar] [CrossRef]
- Gupta, A.; Ong, Y.S.; Feng, L. Multifactorial evolution: Toward evolutionary multitasking. IEEE Trans. Evol. Comput. 2015, 20, 343–357. [Google Scholar] [CrossRef]
- Bali, K.K.; Ong, Y.S.; Gupta, A.; Tan, P.S. Multifactorial evolutionary algorithm with online transfer parameter estimation: MFEA-II. IEEE Trans. Evol. Comput. 2019, 24, 69–83. [Google Scholar] [CrossRef]
- Xing, C.; Gong, W.; Li, S. Adaptive archive-based multifactorial evolutionary algorithm for constrained multitasking optimization. Appl. Soft Comput. 2023, 143, 110385. [Google Scholar] [CrossRef]
- Yu, Y.; Zhu, A.; Zhu, Z.; Lin, Q.; Yin, J.; Ma, X. Multifactorial differential evolution with opposition-based learning for multi-tasking optimization. In Proceedings of the 2019 IEEE Congress on Evolutionary Computation (CEC), Wellington, New Zealand, 10–13 June 2019; IEEE: Piscataway, NJ, USA, 2019; pp. 1898–1905. [Google Scholar]
- Tang, Z.; Gong, M.; Xie, Y.; Li, H.; Qin, A.K. Multi-task particle swarm optimization with dynamic neighbor and level-based inter-task learning. IEEE Trans. Emerg. Top. Comput. Intell. 2021, 6, 300–314. [Google Scholar] [CrossRef]
- Feng, L.; Zhou, L.; Zhong, J.; Gupta, A.; Ong, Y.S.; Tan, K.C.; Qin, A.K. Evolutionary multitasking via explicit autoencoding. IEEE Trans. Cybern. 2018, 49, 3457–3470. [Google Scholar] [CrossRef] [PubMed]
- Lin, J.; Liu, H.L.; Tan, K.C.; Gu, F. An effective knowledge transfer approach for multiobjective multitasking optimization. IEEE Trans. Cybern. 2020, 51, 3238–3248. [Google Scholar] [CrossRef] [PubMed]
- Zhang, T.; Gong, W.; Li, Y. Multitask differential evolution with adaptive dual knowledge transfer. Appl. Soft Comput. 2024, 165, 112040. [Google Scholar] [CrossRef]
- Li, X.; Wang, L.; Jiang, Q. Multipopulation-based multi-tasking evolutionary algorithm. Appl. Intell. 2023, 53, 4624–4647. [Google Scholar] [CrossRef]
- Tauber, A.I.; Podolsky, S.H. The Generation of Diversity: Clonal Selection Theory and the Rise of Molecular Immunology; Harvard University Press: Cambridge, MA, USA, 2000. [Google Scholar]
- De Castro, L.N.; Von Zuben, F.J. Learning and optimization using the clonal selection principle. IEEE Trans. Evol. Comput. 2002, 6, 239–251. [Google Scholar] [CrossRef]
- Yan, X.; Li, P.; Tang, K.; Gao, L.; Wang, L. Clonal selection based intelligent parameter inversion algorithm for prestack seismic data. Inf. Sci. 2020, 517, 86–99. [Google Scholar] [CrossRef]
- Etaati, B.; Ghorrati, Z.; Ebadzadeh, M.M. A full-featured cooperative coevolutionary memory-based artificial immune system for dynamic optimization. Appl. Soft Comput. 2022, 117, 108389. [Google Scholar] [CrossRef]
- Wang, Y.; Li, T.; Liu, X.; Yao, J. An adaptive clonal selection algorithm with multiple differential evolution strategies. Inf. Sci. 2022, 604, 142–169. [Google Scholar] [CrossRef]
- Awad, N.H.; Ali, M.Z.; Suganthan, P.N.; Reynolds, R.G. An ensemble sinusoidal parameter adaptation incorporated with L-SHADE for solving CEC2014 benchmark problems. In Proceedings of the 2016 IEEE Congress on Evolutionary Computation (CEC), Vancouver, BC, Canada, 24–29 July 2016; IEEE: Piscataway, NJ, USA, 2016; pp. 2958–2965. [Google Scholar]
- Li, L.; Lin, Q.; Li, K.; Ming, Z. Vertical distance-based clonal selection mechanism for the multiobjective immune algorithm. Swarm Evol. Comput. 2021, 63, 100886. [Google Scholar] [CrossRef]
- Ouaderhman, T.; Chamlal, H.; Janane, F.Z. A new filter-based gene selection approach in the DNA microarray domain. Expert Syst. Appl. 2024, 240, 122504. [Google Scholar] [CrossRef]
- Lee, J.; Choi, I.Y.; Jun, C.H. An efficient multivariate feature ranking method for gene selection in high-dimensional microarray data. Expert Syst. Appl. 2021, 166, 113971. [Google Scholar] [CrossRef]
- Thabtah, F.; Kamalov, F.; Hammoud, S.; Shahamiri, S.R. Least Loss: A simplified filter method for feature selection. Inf. Sci. 2020, 534, 1–15. [Google Scholar] [CrossRef]
- Alrefai, N.; Ibrahim, O. Optimized feature selection method using particle swarm intelligence with ensemble learning for cancer classification based on microarray datasets. Neural Comput. Appl. 2022, 34, 13513–13528. [Google Scholar] [CrossRef]
- Zhu, Y.; Li, T.; Lan, X. Feature selection optimized by the artificial immune algorithm based on genome shuffling and conditional lethal mutation. Appl. Intell. 2023, 53, 13972–13992. [Google Scholar] [CrossRef]
- Mafarja, M.; Thaher, T.; Too, J.; Chantar, H.; Turabieh, H.; Houssein, E.H.; Emam, M.M. An efficient high-dimensional feature selection approach driven by enhanced multi-strategy grey wolf optimizer for biological data classification. Neural Comput. Appl. 2023, 35, 1749–1775. [Google Scholar] [CrossRef]
- Mahendran, N.; PM, D.R.V. A deep learning framework with an embedded-based feature selection approach for the early detection of the Alzheimer’s disease. Comput. Biol. Med. 2022, 141, 105056. [Google Scholar] [CrossRef]
- Vommi, A.M.; Battula, T.K. A hybrid filter-wrapper feature selection using Fuzzy KNN based on Bonferroni mean for medical datasets classification: A COVID-19 case study. Expert Syst. Appl. 2023, 218, 119612. [Google Scholar] [CrossRef]
- Ke, L.; Li, M.; Wang, L.; Deng, S.; Ye, J.; Yu, X. Improved swarm-optimization-based filter-wrapper gene selection from microarray data for gene expression tumor classification. Pattern Anal. Appl. 2023, 26, 455–472. [Google Scholar] [CrossRef]
- Tanabe, R.; Fukunaga, A. Success-history based parameter adaptation for differential evolution. In Proceedings of the 2013 IEEE Congress on Evolutionary Computation, Cancun, Mexico, 20–23 June 2013; IEEE: Piscataway, NJ, USA, 2013; pp. 71–78. [Google Scholar]
- Braik, M.; Hammouri, A.; Alzoubi, H.; Sheta, A. Feature selection based nature inspired capuchin search algorithm for solving classification problems. Expert Syst. Appl. 2024, 235, 121128. [Google Scholar] [CrossRef]
- Tran, B.; Xue, B.; Zhang, M. Variable-length particle swarm optimization for feature selection on high-dimensional classification. IEEE Trans. Evol. Comput. 2019, 23, 473–487. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).