Differential Expression of the Tetraspanin CD9 in Normal and Leukemic Stem Cells
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
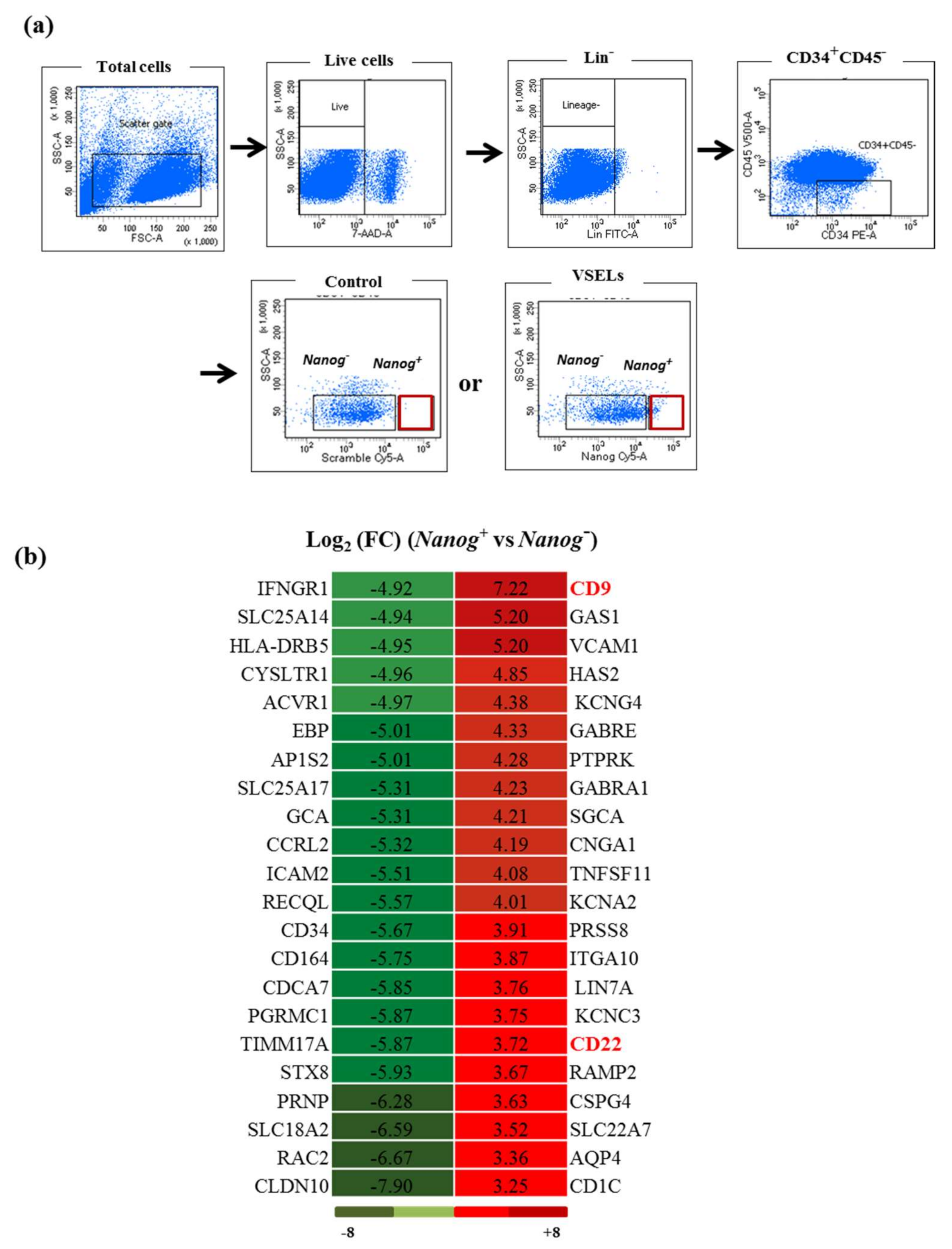
2.1. Isolation of VSELs and Flow Cytometry
2.2. RNA-Sequencing and Data Processing
2.3. Real-Time RT–PCR Analysis
2.4. In Vitro Migration Assay
2.5. Peripheral Blood Samples from Patients and Controls
2.6. Statistical Analysis
3. Results
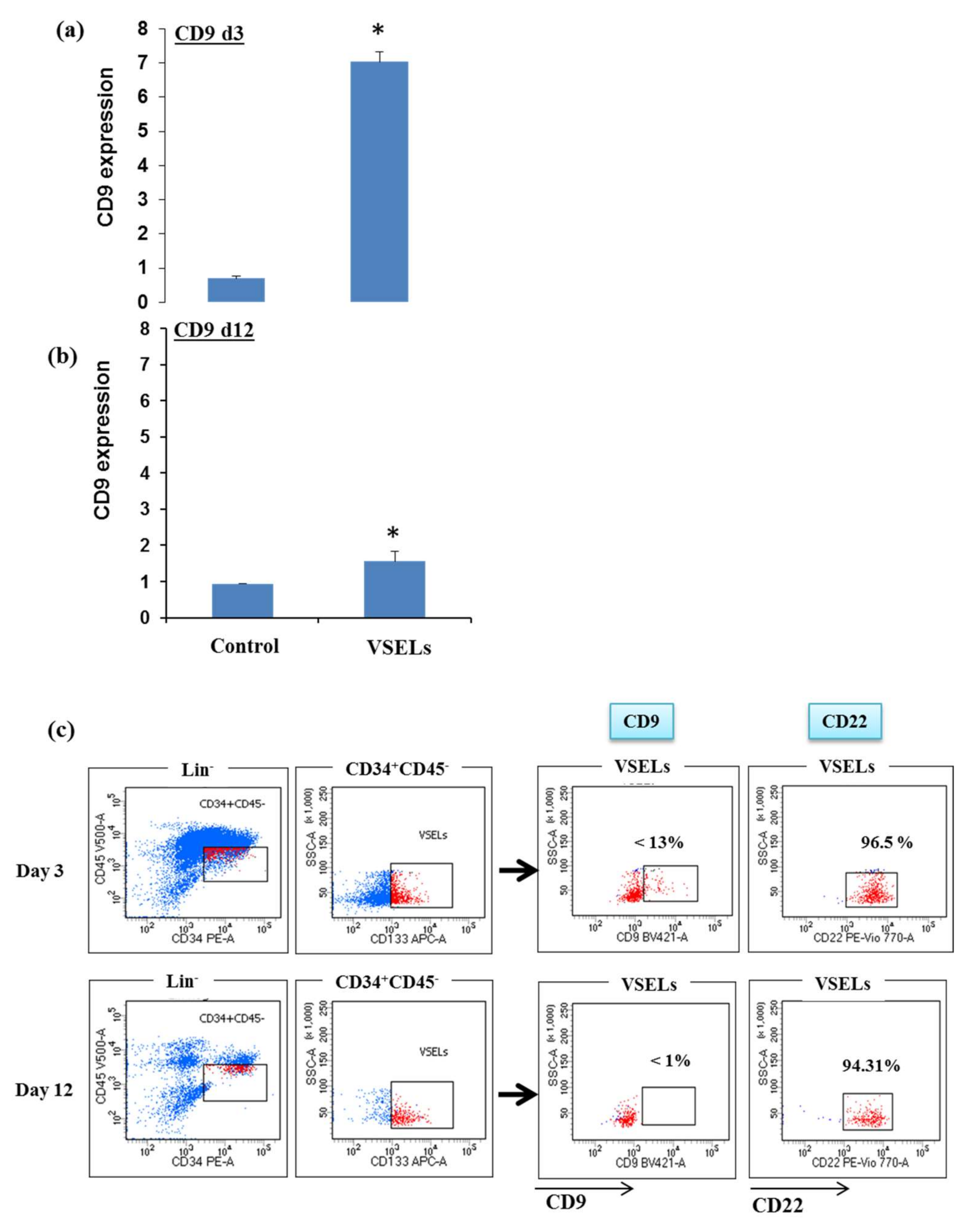
3.1. The CD9 Receptor Expression in the VSEL Subpopulation
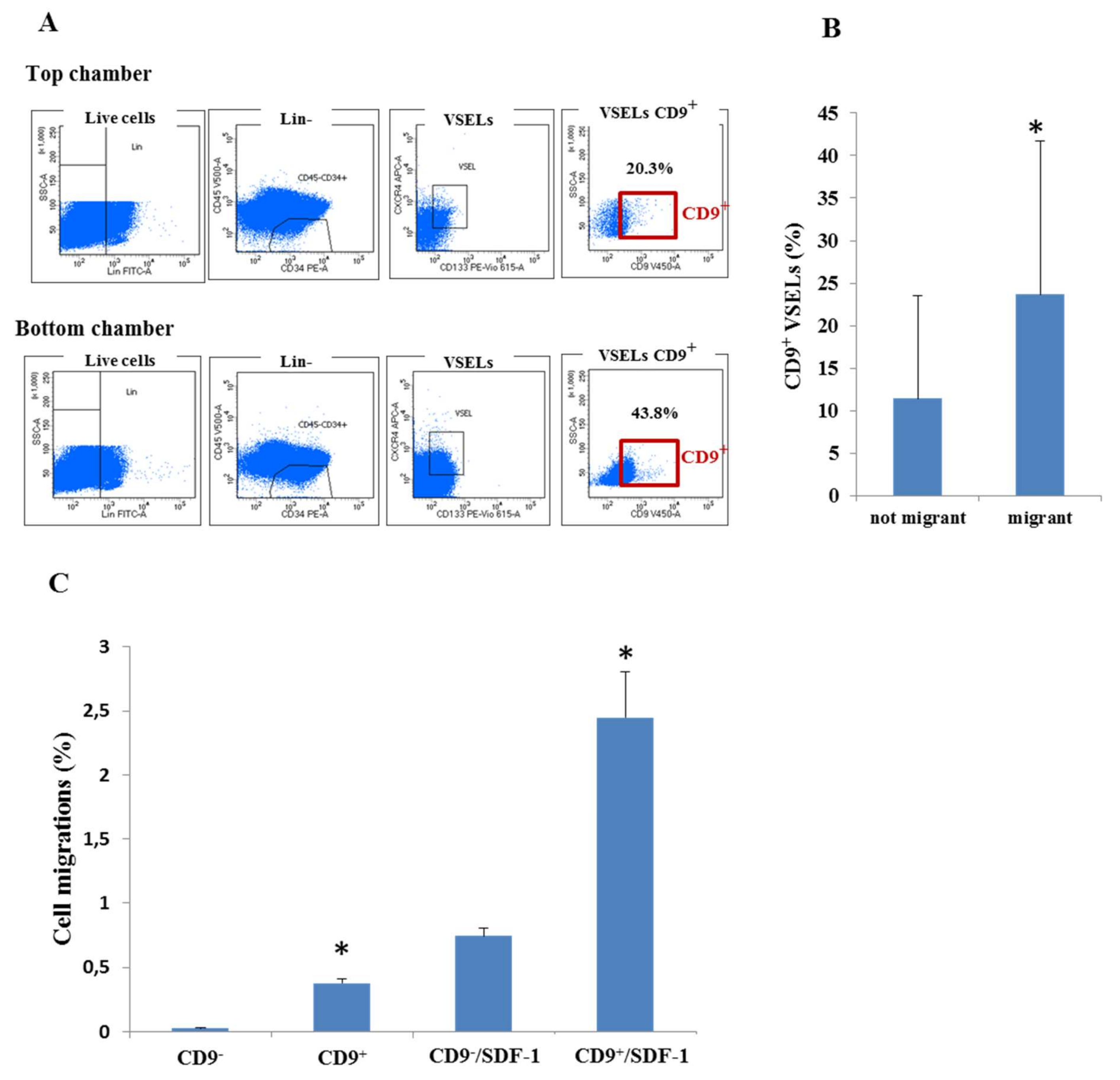
3.2. The SDF-1 Effect on CD9 Receptor Expression in VSEL Subpopulations
3.3. CD9 Receptor Expression and VSELs Migration
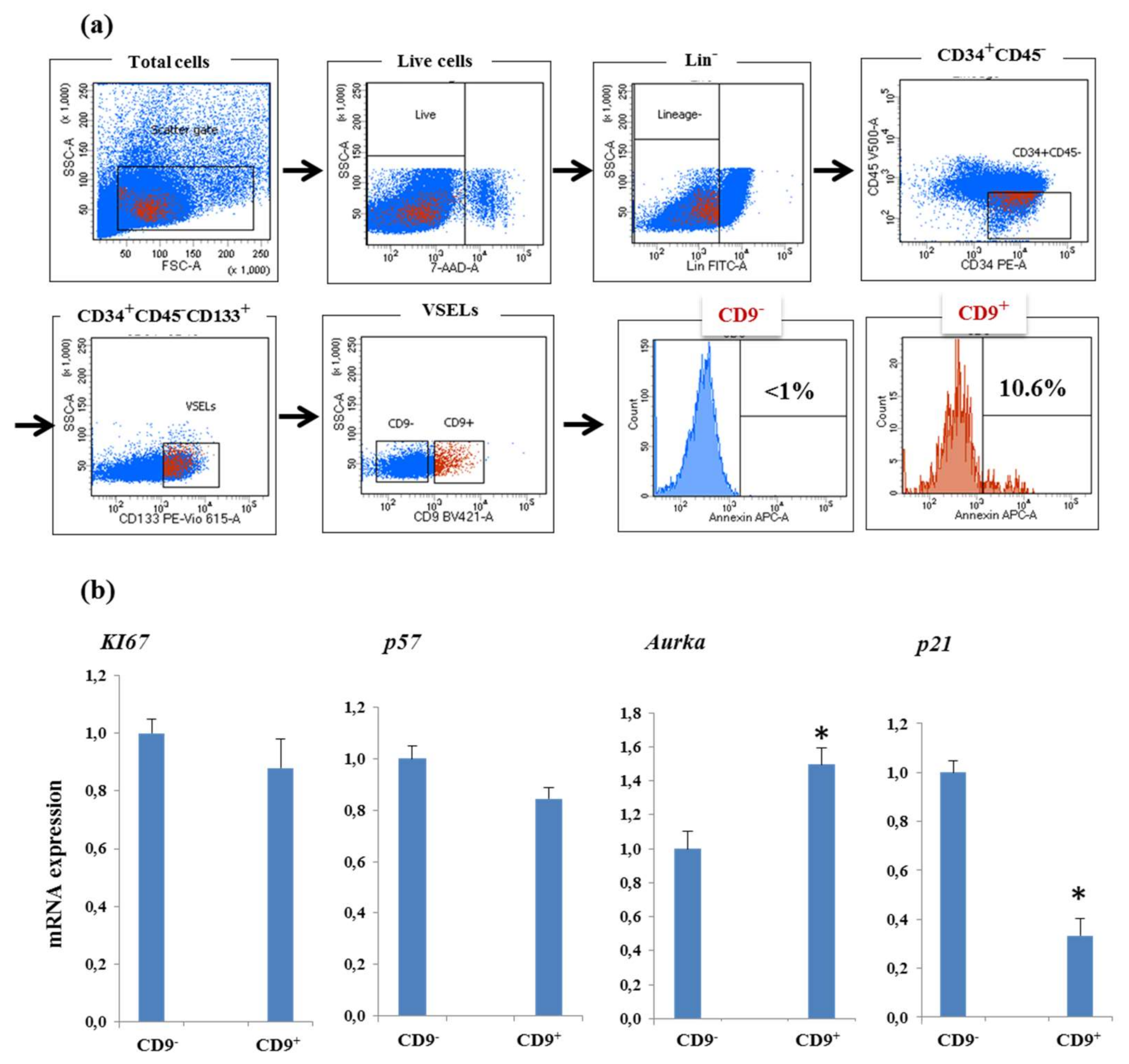
3.4. CD9 Receptor Expression and VSELs Survival and Proliferation
3.5. CD9 Receptor Expression in VSELs and Leukemia
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- De Luca, M.; Aiuti, A.; Cossu, G.; Parmar, M.; Pellegrini, G.; Robey, P.G. Advances in stem cell research and therapeutic development. Nat. Cell Biol. 2019, 21, 801–811. [Google Scholar] [CrossRef] [PubMed]
- Ratajczak, M.; Zuba-Surma, E.; Kucia, M.; Poniewierska, A.; Suszynska, M.; Ratajczak, J. Pluripotent and multipotent stem cells in adult tissues. Adv. Med. Sci. 2012, 57, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Larijani, B.; Esfahani, E.N.; Amini, P.; Nikbin, B.; Alimoghaddam, K.; Amiri, S.; Malekzadeh, R.; Yazdi, N.M.; Ghodsi, M.; Dowlati, Y.; et al. Stem Cell Therapy in Treatment of Different Diseases. Acta Med. Iran. 2012, 50, 79–96. [Google Scholar]
- Yoo, J.; Kim, H.S.; Hwang, D.Y. Stem cells as promising therapeutic options for neurological disorders. J. Cell. Biochem. 2013, 114, 743–753. [Google Scholar] [CrossRef] [PubMed]
- Kucia, M.; Reca, R.; Campbell, F.R.; Zuba-Surma, E.; Majka, M.; Ratajczak, J.; Ratajczak, M.Z. A population of very small embryonic-like (VSEL) CXCR4+SSEA-1+Oct-4+ stem cells identified in adult bone marrow. Leukemia 2006, 20, 857–869. [Google Scholar] [CrossRef] [PubMed]
- Sovalat, H.; Scrofani, M.; Eidenschenk, A.; Pasquet, S.; Rimelen, V.; Hénon, P. Identification and isolation from either adult human bone marrow or G-CSF−mobilized peripheral blood of CD34+/CD133+/CXCR4+/ Lin−CD45− cells, featuring morphological, molecular, and phenotypic characteristics of very small embryonic-like (VSEL) stem cells. Exp. Hematol. 2011, 39, 495–505. [Google Scholar] [CrossRef] [PubMed]
- Dawn, B.; Tiwari, S.; Kucia, M.J.; Zuba-Surma, E.K.; Guo, Y.; SanganalMath, S.K.; Abdel-Latif, A.; Hunt, G.; Vincent, R.J.; Taher, H.; et al. Transplantation of Bone Marrow-Derived Very Small Embryonic-Like Stem Cells Attenuates Left Ventricular Dysfunction and Remodeling After Myocardial Infarction. Stem Cells 2008, 26, 1646–1655. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Zuba-Surma, E.K.; Guo, Y.; Taher, H.; Sanganalmath, S.K.; Hunt, G.; Vincent, R.J.; Kucia, M.; Abdel-Latif, A.; Tang, X.L.; Ratajczak, M.Z.; et al. Transplantation of expanded bone marrow-derived very small embryonic-like stem cells (VSEL-SCs) improves left ventricular function and remodelling after myocardial infarction. J. Cell. Mol. Med. 2011, 15, 1319–1328. [Google Scholar] [CrossRef]
- Lee, S.J.; Park, S.H.; Kim, Y.I.; Hwang, S.; Kwon, P.M.; Han, I.S.; Kwon, B.S. Adult Stem Cells from the Hyaluronic Acid-Rich Node and Duct System Differentiate into Neuronal Cells and Repair Brain Injury. Stem Cells Dev. 2014, 23, 2831–2840. [Google Scholar] [CrossRef] [PubMed]
- Guerin, C.L.; Rossi, E.; Saubamea, B.; Cras, A.; Mignon, V.; Silvestre, J.; Smadja, D.M. Human very Small Embryonic-like Cells Support Vascular Maturation and Therapeutic Revascularization Induced by Endothelial Progenitor Cells. Stem Cell Rev. Rep. 2017, 13, 552–560. [Google Scholar] [CrossRef] [PubMed]
- Sovalat, H.; Scrofani, M.; Eidenschenk, A.; Hénon, P. Human Very Small Embryonic-Like Stem Cells Are Present in Normal Peripheral Blood of Young, Middle-Aged, and Aged Subjects. Stem Cells Int. 2016, 2016, 7651645. [Google Scholar] [CrossRef] [PubMed]
- Guerin, C.L.; Blandinières, A.; Planquette, B.; Silvestre, J.S.; Israel-Biet, D.; Sanchez, O.; Smadja, D.M. Very Small Embryonic-like Stem Cells Are Mobilized in Human Peripheral Blood during Hypoxemic COPD Exacerbations and Pulmonary Hypertension. Stem Cell Rev. Rep. 2017, 13, 561–566. [Google Scholar] [CrossRef] [PubMed]
- Eljaszewicz, A.; Bolkun, L.; Grubczak, K.; Rusak, M.; Wasiluk, T.; Dabrowska, M.; Radziwon, P.; Marlicz, W.; Kamiński, K.; Kloczko, J.; et al. Very Small Embryonic-Like Stem Cells, Endothelial Progenitor Cells, and Different Monocyte Subsets Are Effectively Mobilized in Acute Lymphoblastic Leukemia Patients after G-CSF Treatment. Stem Cells Int. 2018, 2018, 1943980. [Google Scholar] [CrossRef]
- Hénon, P.; Lahlil, R.; Aries, A. CD34+ Very Small Embryonic-like Stem Cells or Induced Pluripotent Stem Cells for Cardiac Repair? J. Cardiol. Vasc. Med. 2020, 6, 1–12. [Google Scholar]
- Lahlil, R.; Scrofani, M.; Barbet, R.; Tancredi, C.; Aries, A.; Hénon, P. VSELs Maintain their Pluripotency and Competence to Differentiate after Enhanced Ex Vivo Expansion. Stem Cell Rev. Rep. 2018, 14, 510–524. [Google Scholar] [CrossRef]
- Ratajczak, M.Z.; Ratajczak, J.; Suszynska, M.; Miller, D.M.; Kucia, M.; Shin, D.M. A Novel View of the Adult Stem Cell Compartment from the Perspective of a Quiescent Population of Very Small Embryonic-Like Stem Cells. Circ. Res. 2017, 120, 166–178. [Google Scholar] [CrossRef]
- Yunta, M.; Lazo, P.A. Tetraspanin proteins as organisers of membrane microdomains and signalling complexes. Cell. Signal. 2003, 15, 559–564. [Google Scholar] [CrossRef]
- De Bruyne, E.; Andersen, T.L.; De Raeve, H.; Van Valckenborgh, E.; Caers, J.; Van Camp, B.; Delaissé, J.M.; Van Riet, I.; Vanderkerken, K. Endothelial cell-driven regulation of CD9 or motility-related protein-1 expression in multiple myeloma cells within the murine 5T33MM model and myeloma patients. Leukemia 2006, 20, 1870–1879. [Google Scholar] [CrossRef]
- Leung, K.T.; Chan, K.Y.; Ng, P.C.; Lau, T.K.; Chiu, W.M.; Tsang, K.S.; Li, C.K.; Kong, C.K.; Li, K. The tetraspanin CD9 regulates migration, adhesion, and homing of human cord blood CD34+ hematopoietic stem and progenitor cells. Blood 2011, 117, 1840–1850. [Google Scholar] [CrossRef]
- Karlsson, G.; Rörby, E.; Pina, C.; Soneji, S.; Reckzeh, K.; Miharada, K.; Karlsson, C.; Guo, Y.; Fugazza, C.; Gupta, R.; et al. The Tetraspanin CD9 Affords High-Purity Capture of All Murine Hematopoietic Stem Cells. Cell Rep. 2013, 4, 642–648. [Google Scholar] [CrossRef] [PubMed]
- Guglielmelli, P.; Zini, R.; Bogani, C.; Salati, S.; Pancrazzi, A.; Bianchi, E.; Mannelli, F.; Ferrari, S.; Le Bousse-Kerdilès, M.C.; Bosi, A.; et al. Molecular Profiling of CD34+ Cells in Idiopathic Myelofibrosis Identifies a Set of Disease-Associated Genes and Reveals the Clinical Significance of Wilms’ Tumor Gene 1 (WT1). Stem Cells 2007, 25, 165–173. [Google Scholar] [CrossRef]
- Desterke, C.; Martinaud, C.; Guerton, B.; Pieri, L.; Bogani, C.; Clay, D.; Torossian, F.; Lataillade, J.J.; Hasselbach, H.C.; Gisslinger, H.; et al. Tetraspanin CD9 participates in dysmegakaryopoiesis and stromal interactions in primary myelofibrosis. Haematologica 2015, 100, 757–767. [Google Scholar] [CrossRef]
- Gandemer, V.; Aubry, M.; Roussel, M.; Rio, A.G.; de Tayrac, M.; Vallee, A.; Mosser, J.; Ly-Sunnaram, B.; Galibert, M.D. CD9 expression can be used to predict childhood TEL/AML1-positive acute lymphoblastic leukemia: Proposal for an accelerated diagnostic flowchart. Leuk. Res. 2010, 34, 430–437. [Google Scholar] [CrossRef]
- Arnaud, M.P.; Vallee, A.; Robert, G.; Bonneau, J.; Leroy, C.; Varin-Blank, N.; Rio, A.G.; Troadec, M.B.; Galibert, M.D.; Gandemer, V. CD9, a key actor in the dissemination of lymphoblastic leukemia, modulating CXCR4-mediated migration via RAC1 signaling. Blood 2015, 126, 1802–1812. [Google Scholar] [CrossRef] [PubMed]
- Arber, D.A.; Orazi, A.; Hasserjian, R.; Thiele, J.; Borowitz, M.J.; Le Beau, M.M.; Bloomfield, C.D.; Cazzola, M.; Vardiman, J.W. The 2016 revision to the World Health Organization classification of myeloid neoplasms and acute leukemia. Blood 2016, 127, 2391–2405. [Google Scholar] [CrossRef]
- Brzoska, E.; Kowalski, K.; Markowska-Zagrajek, A.; Kowalewska, M.; Archacki, R.; Plaskota, I.; Stremińska, W.; Jańczyk-Ilach, K.; Ciemerych, M.A. Sdf-1 (CXCL12) induces CD9 expression in stem cells engaged in muscle regeneration. Stem Cell Res. Ther. 2015, 6, 46. [Google Scholar] [CrossRef] [PubMed]
- Liang, P.; Miao, M.; Liu, Z.; Wang, H.; Jiang, W.; Ma, S.; Li, C.; Hu, R. CD9 expression indicates a poor outcome in acute lymphoblastic leukemia. Cancer Biomark. 2018, 21, 781–786. [Google Scholar] [CrossRef] [PubMed]
- Peled, A.; Petit, I.; Kollet, O.; Magid, M.; Ponomaryov, T.; Byk, T.; Nagler, A.; Ben-Hur, H.; Many, A.; Shultz, L.; et al. Dependence of human stem cell engraftment and repopulation of NOD/SCID mice on CXCR4. Science 1999, 283, 845–848. [Google Scholar] [CrossRef] [PubMed]
- Shivtiel, S.; Lapid, K.; Kalchenko, V.; Avigdor, A.; Goichberg, P.; Kalinkovich, A.; Nagler, A.; Kollet, O.; Lapidot, T. CD45 regulates homing and engraftment of immature normal and leukemic human cells in transplanted immunodeficient mice. Exp. Hematol. 2011, 39, 1161–1170.e1. [Google Scholar] [CrossRef]
- Psatha, N.; Georgolopoulos, G.; Phelps, S.; Papayannopoulou, T. Brief Report: A Differential Transcriptomic Profile of Ex Vivo Expanded Adult Human Hematopoietic Stem Cells Empowers Them for Engraftment Better than Their Surface Phenotype: Ex Vivo Expansion of Adult HSCs. Stem Cells Transl. Med. 2017, 6, 1852–1858. [Google Scholar] [CrossRef]
- Kollet, O.; Spiegel, A.; Peled, A.; Petit, I.; Byk, T.; Hershkoviz, R.; Guetta, E.; Barkai, G.; Nagler, A.; Lapidot, T. Rapid and efficient homing of human CD34+CD38−/lowCXCR4+ stem and progenitor cells to the bone marrow and spleen of NOD/SCID and NOD/SCID/B2mnull mice. Blood 2001, 97, 3283–3291. [Google Scholar] [CrossRef]
- Kawabata, K.; Ujikawa, M.; Egawa, T.; Kawamoto, H.; Tachibana, K.; Iizasa, H.; Katsura, Y.; Kishimoto, T.; Nagasawa, T. A cell-autonomous requirement for CXCR4 in long-term lymphoid and myeloid reconstitution. Proc. Natl. Acad. Sci. USA 1999, 96, 5663–5667. [Google Scholar] [CrossRef] [PubMed]
- García-Bernal, D.; Redondo-Muñoz, J.; Dios-Esponera, A.; Chèvre, R.; Bailón, E.; Garayoa, M.; Arellano-Sánchez, N.; Gutierrez, N.C.; Hidalgo, A.; García-Pardo, A.; et al. Sphingosine-1-phosphate activates chemokine-promoted myeloma cell adhesion and migration involving α4β1 integrin function. J. Pathol. 2013, 229, 36–48. [Google Scholar] [CrossRef] [PubMed]
- Yamazaki, H.; Wilson Xu, C.; Naito, M.; Nishida, H.; Okamoto, T.; Ghani, F.I.; Iwata, S.; Inukai, T.; Sugita, K.; Morimoto, C. Regulation of cancer stem cell properties by CD9 in human B-acute lymphoblastic leukemia. Biochem. Biophys. Res. Commun. 2011, 409, 14–21. [Google Scholar] [CrossRef]
- Touzet, L.; Dumezy, F.; Roumier, C.; Berthon, C.; Bories, C.; Quesnel, B.; Preudhomme, C.; Boyer, T. CD9 in acute myeloid leukemia: Prognostic role and usefulness to target leukemic stem cells. Cancer Med. 2019, 8, 1279–1288. [Google Scholar] [CrossRef]
- Yamashita, M.; Dellorusso, P.V.; Olson, O.C.; Passegué, E. Dysregulated haematopoietic stem cell behaviour in myeloid leukaemogenesis. Nat. Rev. Cancer 2020, 20, 365–382. [Google Scholar] [CrossRef] [PubMed]
- Zou, Y.-R.; Kottmann, A.H.; Kuroda, M.; Taniuchi, I.; Littman, D.R. Function of the chemokine receptor CXCR4 in haematopoiesis and in cerebellar development. Nature 1998, 393, 595–599. [Google Scholar] [CrossRef]

| CD9 Fw | AAGTTAGCCCTCACCATGCC |
| CD9 Rv | TCCAATGGCAAGGACAGCA |
| KI67 Fw | ATTGAACCTGCGGAAGAGCTGA |
| KI67 Rv | GGAGCGCAGGGATATTCCCTTA |
| P57 Kip2 Fw | GCGGCGATCAAGAAGCTGT |
| P57 Kip2 Rv | TGGCGAAGAAATCGGAGATCA |
| AURKA-Fw | GCATTTCAGGACCTGTTAAGGCTA |
| AURKA-Rv | TGCTGAGTCACGAGAACACGTTT |
| GAPDH Fw | CATCGCTCAGACACCATGG |
| GAPDH Rv | ATGTAGTTGAGGTCAATGAAGGG |
| β2-microglobulin Fw | TGACTTTGTCACAGCCCAAGATA |
| β2-microglobulin Rv | AATGCGGCATCTTCAAACCT |
| Controls (n = 13) | MPNs Diagnosis (n = 7) | |
|---|---|---|
| Age (year) | 37.4 ± 7 | 64.6 ± 17.6 |
| WBC × 109/L | 3.9–10.9 # | 8.7 ± 6.6 |
| RBC × 1012/L | 4.4–5.6 # | 5.1 ± 112 |
| Hemoglobin g/dL | 13.5–16.9 # | 14.9 ± 2.9 |
| hematocrit | 40–49.4 # | 44.6 ± 8.6 |
| Platelet × 109/L | 166–308 # | 529.7 ± 424.8 |
| CD9+ VSELs (%)/Lin– | 0.16 ± 0.18 | 1.11 ± 1.02 * |
| CD9+ HSCs (%)/Lin– | 0.14 ± 0.1 | 0.17 ± 0.15 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lahlil, R.; Scrofani, M.; Aries, A.; Hénon, P.; Drénou, B. Differential Expression of the Tetraspanin CD9 in Normal and Leukemic Stem Cells. Biology 2021, 10, 312. https://doi.org/10.3390/biology10040312
Lahlil R, Scrofani M, Aries A, Hénon P, Drénou B. Differential Expression of the Tetraspanin CD9 in Normal and Leukemic Stem Cells. Biology. 2021; 10(4):312. https://doi.org/10.3390/biology10040312
Chicago/Turabian StyleLahlil, Rachid, Maurice Scrofani, Anne Aries, Philippe Hénon, and Bernard Drénou. 2021. "Differential Expression of the Tetraspanin CD9 in Normal and Leukemic Stem Cells" Biology 10, no. 4: 312. https://doi.org/10.3390/biology10040312
APA StyleLahlil, R., Scrofani, M., Aries, A., Hénon, P., & Drénou, B. (2021). Differential Expression of the Tetraspanin CD9 in Normal and Leukemic Stem Cells. Biology, 10(4), 312. https://doi.org/10.3390/biology10040312







