Isolation, Characterization, and Evaluation of a Lytic Jumbo Phage Z90 Against Aeromonas hydrophila in American Eels (Anguilla rostrata)
Abstract
1. Introduction
2. Results
2.1. Isolation and Identification of A. hydrophila 2408
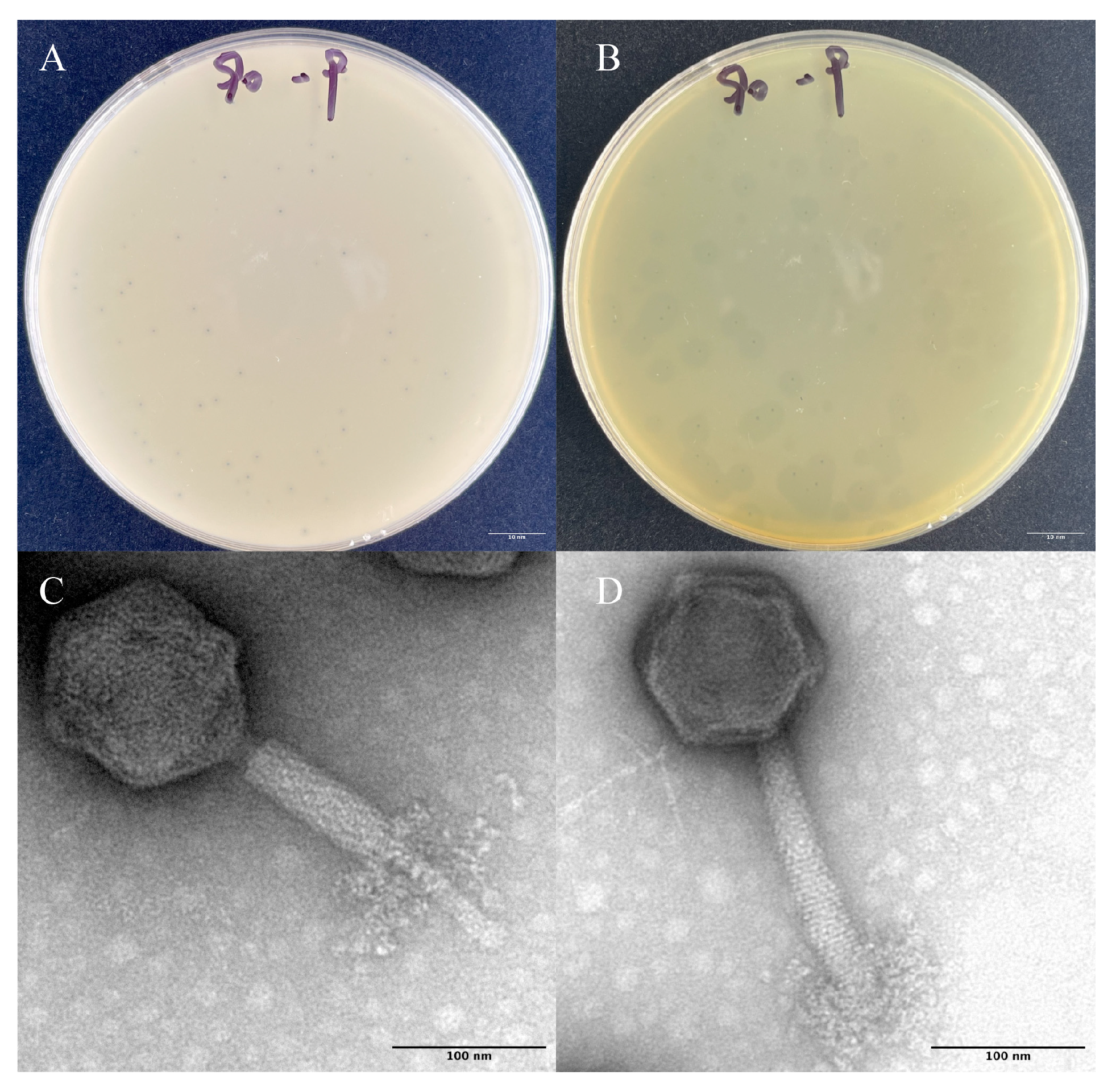
2.2. Morphological Characteristics of the Phage Z90
2.3. Host Range of the Phage Z90
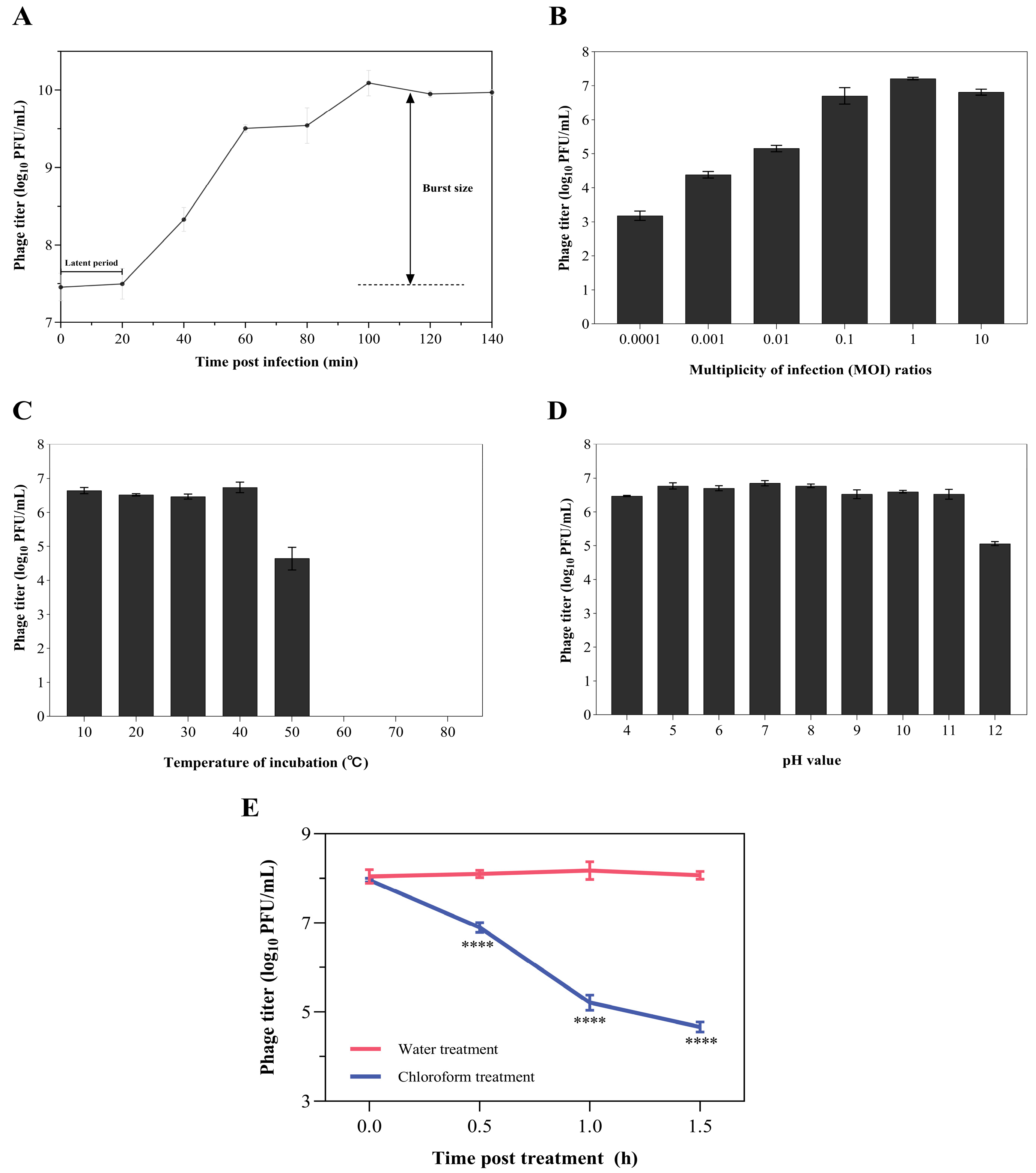
2.4. One-Step Growth Curve Analysis of the Phage Z90
2.5. Optimal MOI and Temperature, pH, and Chloroform Stability of the Phage Z90
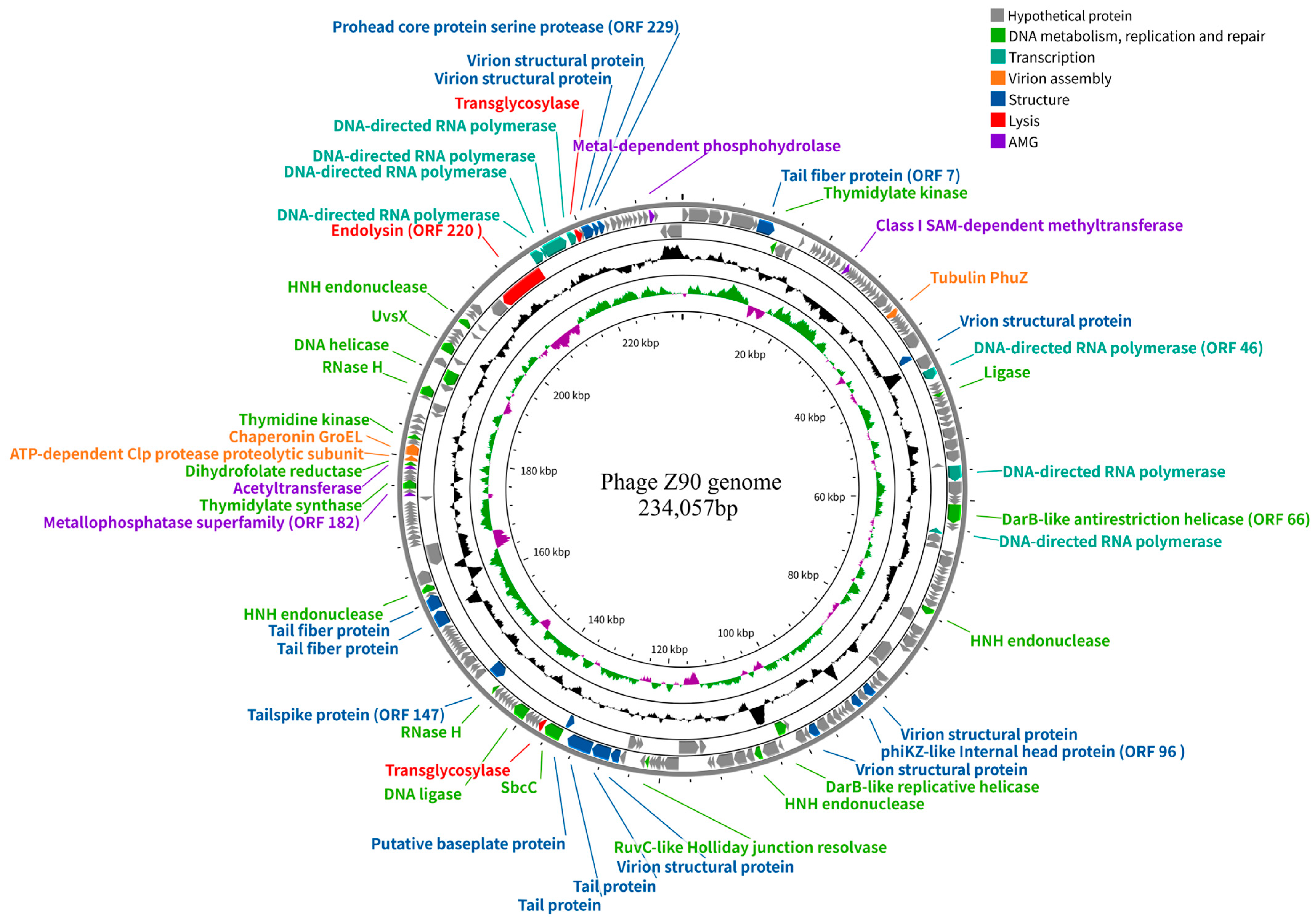
2.6. Genomic Characteristics of the Phage Z90
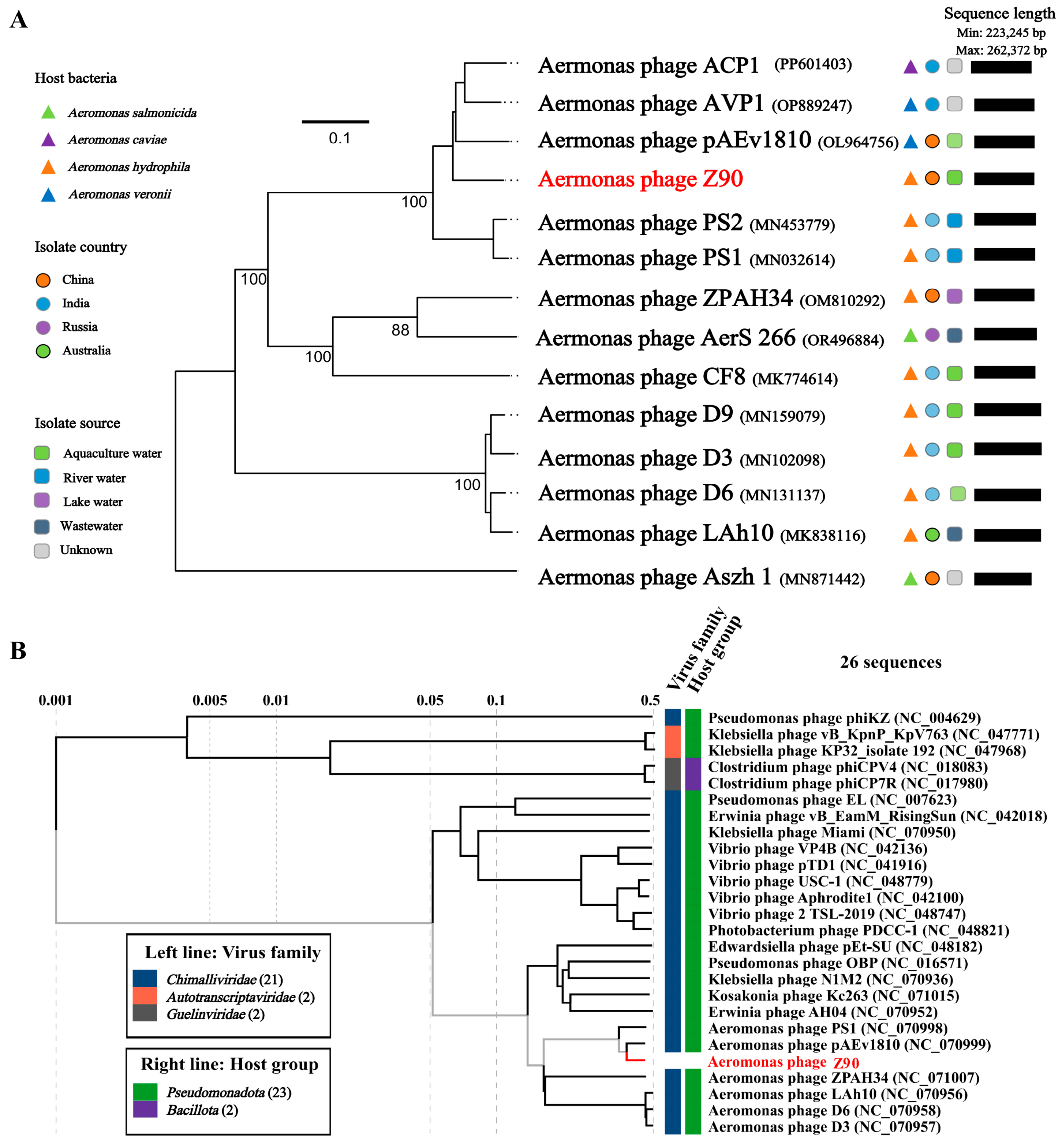
2.7. Phylogenetic and Taxonomic Classification of the Phage Z90
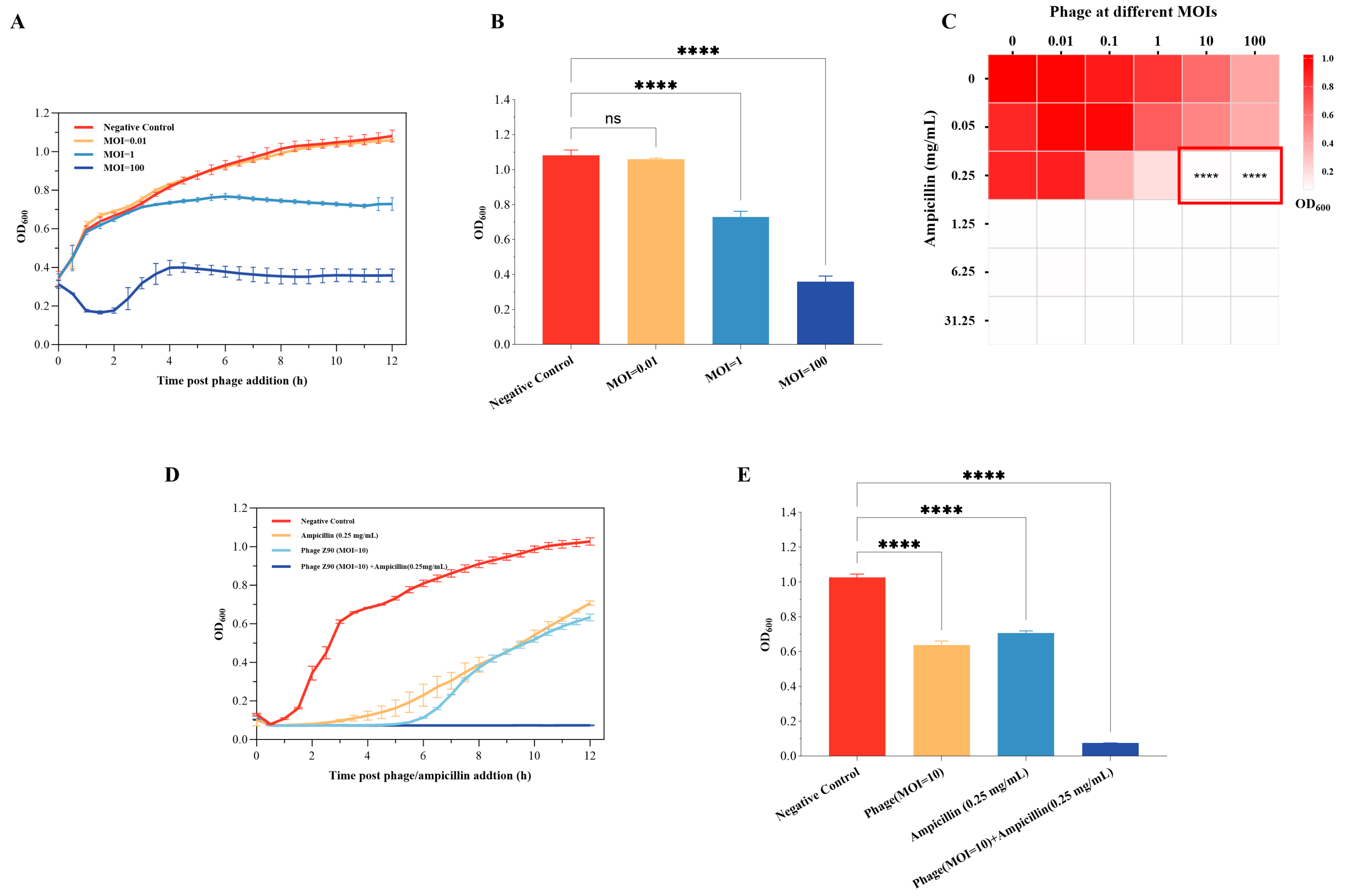
2.8. In Vitro Bactericidal Effects of the Phage Z90
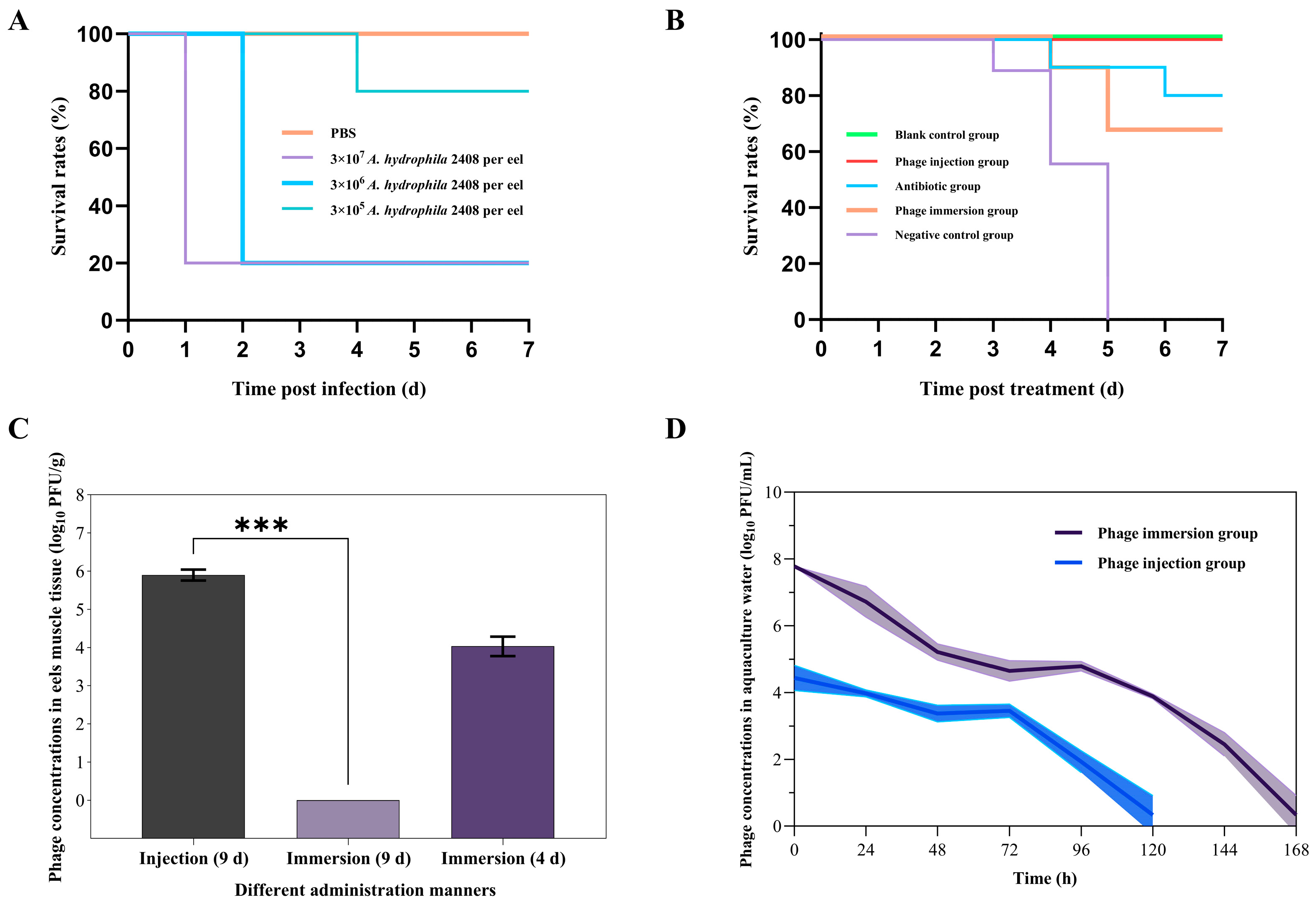
2.9. In Vivo Therapeutic Effects of the Phage Z90
3. Discussion
4. Materials and Methods
4.1. Isolation and Identification of Bacteria Pathogens
4.2. Phage Isolation, Purification, and Amplification
4.3. Transmission Electron Microscopy (TEM)
4.4. One-Step Growth Curve Assay
4.5. Determination of Optimal MOI
4.6. Determination of Temperature, pH, and Chloroform Stability
4.7. Determination of Host Range
4.8. Genomic Analysis of the Phage Z90
4.9. In Vitro Evaluation of Phage Bactericidal Effects
4.10. In Vivo Evaluation of Phage Therapeutic Effects
4.11. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Huang, Y.; Ge, R.; Lou, G.; Jiang, N.; Zhu, X.; Guo, Y.; Liu, H.; Liu, X.; Chen, X. The influence of dietary Coenzyme Q10 on growth performance, antioxidant capacity and resistance against Aeromonas hydrophila of juvenile European eel (Anguilla anguilla). Fish Shellfish. Immunol. 2023, 138, 108834. [Google Scholar] [CrossRef] [PubMed]
- Łuczyńska, J.; Nowosad, J.; Łuczyński, M.J.; Kucharczyk, D. Evaluation of Chemical Elements, Lipid Profiles, Nutritional Indices and Health Risk Assessment of European Eel (Anguilla anguilla L.). Int. J. Environ. Res. Public Health 2023, 20, 2257. [Google Scholar] [CrossRef]
- Qi, T.; Wei, S.; Li, Z.; Ribas, L.; Cao, Q. Current research on bacterial diseases in eel: An immunological perspective. Aquaculture 2025, 595, 741599. [Google Scholar] [CrossRef]
- He, L.; Wu, L.; Lin, P.; Zhai, S.; Guo, S.; Xiao, Y.; Wan, Q. First expression and immunogenicity study of a novel trivalent outer membrane protein (OmpII-U-A) from Aeromonas hydrophila, Vibrio vulnificus and Edwardsiella anguillarum. Aquaculture 2020, 519, 734932. [Google Scholar] [CrossRef]
- He, L.; Wu, L.; Tang, Y.; Lin, P.; Zhai, S.; Xiao, Y.; Guo, S. Immunization of a novel outer membrane protein from Aeromonas hydrophila simultaneously resisting A. hydrophila and Edwardsiella anguillarum infection in European eels (Angullia angullia). Fish Shellfish. Immunol. 2020, 97, 300–312. [Google Scholar] [CrossRef]
- Liang, S.; Liang, R.; Raza, S.H.A.; Huang, Q.; Li, T.; Bai, H.; Feng, C.; Xin, L.; Guo, H.; Wang, S.; et al. Biological analysis of phage vB_AhaP_PT2 and treatment rescued crucian carp infected with Aeromonas hydrophila. Aquaculture 2025, 595, 741654. [Google Scholar] [CrossRef]
- Antonova, D.; Nichiporenko, A.; Sobinina, M.; Wang, Y.; Vishnyakov, I.E.; Moiseenko, A.; Kurdyumova, I.; Chesnokov, Y.M.; Stepanchikova, E.; Bourkaltseva, M.; et al. Genomic transfer via membrane vesicle: A strategy of giant phage phiKZ for early infection. J. Virol. 2024, 98, e0020524. [Google Scholar] [CrossRef] [PubMed]
- Bakiyev, S.; Smekenov, I.; Zharkova, I.; Kobegenova, S.; Sergaliyev, N.; Absatirov, G.; Bissenbaev, A. Isolation, identification, and characterization of pathogenic Aeromonas hydrophila from critically endangered Acipenser baerii. Aquac. Rep. 2022, 26, 101293. [Google Scholar] [CrossRef]
- Hossain, M.J.; Sun, D.; McGarey, D.J.; Wrenn, S.; Alexander, L.M.; Martino, M.E.; Xing, Y.; Terhune, J.S.; Liles, M.R. An Asian origin of virulent Aeromonas hydrophila responsible for disease epidemics in United States-farmed catfish. mBio 2014, 5, e00848-14. [Google Scholar] [CrossRef] [PubMed]
- Peterman, M.A.; Posadas, B.C. Direct Economic Impact of Fish Diseases on the East Mississippi Catfish Industry. N. Am. J. Aquac. 2019, 81, 222–229. [Google Scholar] [CrossRef]
- Guo, S.; Wan, Q.; Xu, M.; Chen, M.; Chen, Z. Transcriptome analysis of host anti-Aeromonas hydrophila infection revealed the pathogenicity of A. hydrophila to American eels (Anguilla rostrata). Fish Shellfish. Immunol. 2024, 148, 109504. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; Yang, Q.E.; Zhou, X.; Wang, F.-H.; Muurinen, J.; Virta, M.P.; Brandt, K.K.; Zhu, Y.-G. Antibiotic resistome in the livestock and aquaculture industries: Status and solutions. Crit. Rev. Environ. Sci. Technol. 2021, 51, 2159–2196. [Google Scholar] [CrossRef]
- Rai, S.; Kaur, B.; Singh, P.; Singh, A.; Benjakul, S.; Reddy, S.V.K.; Nagar, V.; Tyagi, A. Perspectives on phage therapy for health management in aquaculture. Aquac. Int. 2024, 32, 1349–1393. [Google Scholar] [CrossRef]
- Sokolova, M.L.; Misovetc, I.; Severinov, K.V. Multisubunit RNA Polymerases of Jumbo Bacteriophages. Viruses 2020, 12, 1064. [Google Scholar] [CrossRef]
- Korn Abby, M.; Hillhouse Andrew, E.; Sun, L.; Gill Jason, J. Comparative Genomics of Three Novel Jumbo Bacteriophages Infecting Staphylococcus aureus. J. Virol. 2021, 95, JVI0239120. [Google Scholar] [CrossRef]
- Nazir, A.; Ali, A.; Qing, H.; Tong, Y. Emerging Aspects of Jumbo Bacteriophages. Infect. Drug Resist. 2021, 14, 5041–5055. [Google Scholar] [CrossRef]
- Fischetti, V.A. Development of Phage Lysins as Novel Therapeutics: A Historical Perspective. Viruses 2018, 10, 310. [Google Scholar] [CrossRef]
- Maciejewska, B.; Olszak, T.; Drulis-Kawa, Z. Applications of bacteriophages versus phage enzymes to combat and cure bacterial infections: An ambitious and also a realistic application? Appl. Microbiol. Biotechnol. 2018, 102, 2563–2581. [Google Scholar] [CrossRef]
- da Silva, J.D.; Melo, L.D.R.; Santos, S.B.; Kropinski, A.M.; Xisto, M.F.; Dias, R.S.; da Silva Paes, I.; Vieira, M.S.; Soares, J.J.F.; Porcellato, D.; et al. Genomic and proteomic characterization of vB_SauM-UFV_DC4, a novel Staphylococcus jumbo phage. Appl. Microbiol. Biotechnol. 2023, 107, 7231–7250. [Google Scholar] [CrossRef]
- Zhang, K.; Wei, X.; Liu, H.; He, B.; Zhao, R.; Zhou, J.; Deng, J.; Xie, F.; Xiong, X.; Li, G.; et al. Characterization of a novel Jumbo phage JP4 with potential to control pathogenic Escherichia coli. Virol. J. 2025, 22, 386. [Google Scholar] [CrossRef]
- Guan, J.; Bondy-Denomy, J. Intracellular Organization by Jumbo Bacteriophages. J. Bacteriol. 2020, 203, e00362-20. [Google Scholar] [CrossRef] [PubMed]
- Jo, D.; Kim, H.; Lee, Y.; Kim, J.; Ryu, S. Characterization and genomic study of EJP2, a novel jumbo phage targeting antimicrobial resistant Escherichia coli. Front. Microbiol. 2023, 14, 1194435. [Google Scholar] [CrossRef]
- Imam, M.; Alrashid, B.; Patel, F.; Dowah, A.S.A.; Brown, N.; Millard, A.; Clokie, M.R.J.; Galyov, E.E. vB_PaeM_MIJ3, a Novel Jumbo Phage Infecting Pseudomonas aeruginosa, Possesses Unusual Genomic Features. Front. Microbiol. 2019, 10, 2772. [Google Scholar] [CrossRef]
- Leskinen, K.; Blasdel, B.G.; Lavigne, R.; Skurnik, M. RNA-Sequencing Reveals the Progression of Phage-Host Interactions between φR1-37 and Yersinia enterocolitica. Viruses 2016, 8, 111. [Google Scholar] [CrossRef]
- Orekhova, M.; Koreshova, A.; Artamonova, T.; Khodorkovskii, M.; Yakunina, M. The study of the phiKZ phage non-canonical non-virion RNA polymerase. Biochem. Biophys. Res. Commun. 2019, 511, 759–764. [Google Scholar] [CrossRef]
- Iyer, L.M.; Anantharaman, V.; Krishnan, A.; Burroughs, A.M.; Aravind, L. Jumbo Phages: A Comparative Genomic Overview of Core Functions and Adaptions for Biological Conflicts. Viruses 2021, 13, 63. [Google Scholar] [CrossRef]
- Ceyssens, P.J.; Minakhin, L.; Van den Bossche, A.; Yakunina, M.; Klimuk, E.; Blasdel, B.; De Smet, J.; Noben, J.P.; Bläsi, U.; Severinov, K.; et al. Development of giant bacteriophage ϕKZ is independent of the host transcription apparatus. J. Virol. 2014, 88, 10501–10510. [Google Scholar] [CrossRef] [PubMed]
- Yoshikawa, G.; Askora, A.; Blanc-Mathieu, R.; Kawasaki, T.; Li, Y.; Nakano, M.; Ogata, H.; Yamada, T. Xanthomonas citri jumbo phage XacN1 exhibits a wide host range and high complement of tRNA genes. Sci. Rep. 2018, 8, 4486. [Google Scholar] [CrossRef]
- Le, T.S.; Nguyen, T.H.; Vo, H.P.; Doan, V.C.; Nguyen, H.L.; Tran, M.T.; Tran, T.T.; Southgate, P.C.; Kurtböke, D. Protective Effects of Bacteriophages against Aeromonas hydrophila Species Causing Motile Aeromonas Septicemia (MAS) in Striped Catfish. Antibiotics 2018, 7, 16. [Google Scholar] [CrossRef] [PubMed]
- Liang, R.; Cui, Z.; Abbas Raza, S.H.; Li, T.; Zhang, Z.; Huang, Q.; Bai, H.; Cheng, Y.; Du, B.; Li, J.; et al. Protective immune-response of Aeromonas hydrophila phage lysate in crucian carp against direct virulent challenge with A. hydrophila-TPS. Fish Shellfish. Immunol. 2024, 155, 110011. [Google Scholar] [CrossRef]
- Feng, C.; Jia, K.; Chi, T.; Chen, S.; Yu, H.; Zhang, L.; Haidar Abbas Raza, S.; Alshammari, A.M.; Liang, S.; Zhu, Z.; et al. Lytic Bacteriophage PZL-Ah152 as Biocontrol Measures Against Lethal Aeromonas hydrophila Without Distorting Gut Microbiota. Front. Microbiol. 2022, 13, 898961. [Google Scholar] [CrossRef]
- Janelidze, N.; Jaiani, E.; Didebulidze, E.; Kusradze, I.; Kotorashvili, A.; Chalidze, K.; Porchkhidze, K.; Khukhunashvili, T.; Tsertsvadze, G.; Jgenti, D.; et al. Phenotypic and Genetic Characterization of Aeromonas hydrophila Phage AhMtk13a and Evaluation of Its Therapeutic Potential on Simulated Aeromonas Infection in Danio rerio. Viruses 2022, 14, 412. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.; Yang, Q.; Hu, R.-G.; Cong, W.; Li, S.; Kang, Y.-H. The evaluation of bacteriophage therapy in aquaculture: A systematic review and meta-analysis. Aquaculture 2024, 588, 740925. [Google Scholar] [CrossRef]
- Rai, S.; Tyagi, A.; Kumar, B.T.N. Oral feed-based administration of phage cocktail protects rohu fish (Labeo rohita) against Aeromonas hydrophila infection. Arch. Microbiol. 2024, 206, 219. [Google Scholar] [CrossRef]
- Islam, M.S.; Yang, X.; Euler, C.W.; Han, X.; Liu, J.; Hossen, M.I.; Zhou, Y.; Li, J. Application of a novel phage ZPAH7 for controlling multidrug-resistant Aeromonas hydrophila on lettuce and reducing biofilms. Food Control. 2021, 122, 107785. [Google Scholar] [CrossRef]
- Prichard, A.; Lee, J.; Laughlin, T.G.; Lee, A.; Thomas, K.P.; Sy, A.E.; Spencer, T.; Asavavimol, A.; Cafferata, A.; Cameron, M.; et al. Identifying the core genome of the nucleus-forming bacteriophage family and characterization of Erwinia phage RAY. Cell Rep. 2023, 42, 112432. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Zhang, F.; Liang, Y.; Zheng, K.; Gu, C.; Zhang, W.; Liu, Y.; Zhang, X.; Shao, H.; Jiang, Y.; et al. Genome and Ecology of a Novel Alteromonas Podovirus, ZP6, Representing a New Viral Genus, Mareflavirus. Microbiol. Spectr. 2021, 9, e0046321. [Google Scholar] [CrossRef]
- Zhang, M.; Kang, J.; Wu, B.; Qin, Y.; Huang, L.; Zhao, L.; Mao, L.; Wang, S.; Yan, Q. Comparative transcriptome and phenotype analysis revealed the role and mechanism of ompR in the virulence of fish pathogenic Aeromonas hydrophila. Microbiologyopen 2020, 9, e1041. [Google Scholar] [CrossRef]
- Feng, J.; Lin, P.; Guo, S.; Jia, Y.; Wang, Y.; Zadlock, F.; Zhang, Z. Identification and characterization of a novel conserved 46 kD maltoporin of Aeromonas hydrophila as a versatile vaccine candidate in European eel (Anguilla anguilla). Fish Shellfish. Immunol. 2017, 64, 93–103. [Google Scholar] [CrossRef]
- Ding, H.; Shi, K.; Hsiao, M.; Li, W.; Liu, X.; Xu, J.; Yang, Y.; Zhang, R. Two virulent Vibrio campbellii phages with potential for phage therapy in aquaculture. BMC Microbiol. 2025, 25, 99. [Google Scholar] [CrossRef]
- Lindberg, H.M.; McKean, K.A.; Wang, I.N. Phage fitness may help predict phage therapy efficacy. Bacteriophage 2014, 4, e964081. [Google Scholar] [CrossRef]
- Shi, K.; Ding, H.; Zhong, W.; Li, W.; Hsiao, M.; Xu, B.; Guo, H.; Zhao, J.; Xu, J.; Zhang, D.; et al. A broad-spectrum vibriophage with potential for phage therapy against Vibrio alginolyticus in shrimp aquaculture. Aquaculture 2026, 612, 743215. [Google Scholar] [CrossRef]
- Hou, Y.; Wu, Z.; Ren, L.; Chen, Y.; Zhang, Y.A.; Zhou, Y. Characterization and application of a lytic jumbo phage ZPAH34 against multidrug-resistant Aeromonas hydrophila. Front. Microbiol. 2023, 14, 1178876. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Zhong, W.; Ding, H.; Shi, K.; Guo, H.; Zhang, D.; Cai, L.; Hsiao, M.; Xu, J.; Zhang, R. A lytic vibriophage as a potential biocontrol agent against vibriosis in aquaculture. Fish. Shellfish. Immunol. 2025, 165, 110506. [Google Scholar] [CrossRef] [PubMed]
- Silva, Y.J.; Costa, L.; Pereira, C.; Cunha, Â.; Calado, R.; Gomes, N.C.; Almeida, A. Influence of environmental variables in the efficiency of phage therapy in aquaculture. Microb. Biotechnol. 2014, 7, 401–413. [Google Scholar] [CrossRef]
- Thanki, A.M.; Brown, N.; Millard, A.D.; Clokie, M.R.J. Genomic Characterization of Jumbo Salmonella Phages That Effectively Target United Kingdom Pig-Associated Salmonella Serotypes. Front. Microbiol. 2019, 10, 1491. [Google Scholar] [CrossRef]
- Armbruster, E.G.; Rani, P.; Lee, J.; Klusch, N.; Hutchings, J.; Hoffman, L.Y.; Buschkaemper, H.; Enustun, E.; Adler, B.A.; Inlow, K.; et al. Sequential membrane- and protein-bound organelles compartmentalize genomes during phage infection. Cell Host Microbe 2025, 33, 484–497.e486. [Google Scholar] [CrossRef]
- Harding, K.R.; Malone, L.M.; Kyte, N.A.P.; Jackson, S.A.; Smith, L.M.; Fineran, P.C. Genome-wide identification of bacterial genes contributing to nucleus-forming jumbo phage infection. Nucleic Acids Res. 2025, 53, gkae1194. [Google Scholar] [CrossRef]
- Bondy-Denomy, J.; Pawluk, A.; Maxwell, K.L.; Davidson, A.R. Bacteriophage genes that inactivate the CRISPR/Cas bacterial immune system. Nature 2013, 493, 429–432. [Google Scholar] [CrossRef]
- Malone, L.M.; Warring, S.L.; Jackson, S.A.; Warnecke, C.; Gardner, P.P.; Gumy, L.F.; Fineran, P.C. A jumbo phage that forms a nucleus-like structure evades CRISPR-Cas DNA targeting but is vulnerable to type III RNA-based immunity. Nat. Microbiol. 2020, 5, 48–55. [Google Scholar] [CrossRef]
- Mendoza, S.D.; Nieweglowska, E.S.; Govindarajan, S.; Leon, L.M.; Berry, J.D.; Tiwari, A.; Chaikeeratisak, V.; Pogliano, J.; Agard, D.A.; Bondy-Denomy, J. A bacteriophage nucleus-like compartment shields DNA from CRISPR nucleases. Nature 2020, 577, 244–248. [Google Scholar] [CrossRef]
- Rodriguez, Z.K.; Laughlin, T.G.; VanderWal, A.; Aindow, A.; Razavilar, A.; Prichard, A.M.; Lee, J.; Armbruster, E.; Pogliano, J.; Villa, E. Investigating the early stages of infection of nucleus-forming jumbo phage. Biophys. J. 2023, 122, 414a. [Google Scholar] [CrossRef]
- Mozumdar, D.; Fossati, A.; Stevenson, E.; Guan, J.; Nieweglowska, E.; Rao, S.; Agard, D.; Swaney, D.L.; Bondy-Denomy, J. Characterization of a lipid-based jumbo phage compartment as a hub for early phage infection. Cell Host Microbe 2024, 32, 1050–1058.e1057. [Google Scholar] [CrossRef]
- Yang, L.; Zhong, W.; Tang, T.; He, M.; Zhang, T.; Zhou, B.; Yin, Y.; Guo, J.; Gao, Z. Phage-Based Biocontrol Strategies and Application in Aquatic Animal Disease Prevention and Control. Rev. Aquac. 2025, 17, e70024. [Google Scholar] [CrossRef]
- Hegarty, B. Making waves: Intelligent phage cocktail design, a pathway to precise microbial control in water systems. Water Res. 2025, 268, 122594. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Zhan, C.; Sun, Y.; Pan, T.; Zhou, X. Exploring the efficacy of bacteriophage cocktails for mitigating Vibrio contamination within the seafood production chain: A feasible approach to microbial risk management. Food Control. 2025, 177, 111424. [Google Scholar] [CrossRef]
- Kim, M.K.; Chen, Q.; Echterhof, A.; Pennetzdorfer, N.; McBride, R.C.; Banaei, N.; Burgener, E.B.; Milla, C.E.; Bollyky, P.L. A blueprint for broadly effective bacteriophage-antibiotic cocktails against bacterial infections. Nat. Commun. 2024, 15, 9987. [Google Scholar] [CrossRef] [PubMed]
- Lu, H.; Li, Z.; Elbaz, A.; Ni, S.Q. Synergistic action of phages and lytic proteins with antibiotics: A combination strategy to target bacteria and biofilms. BMC Microbiol. 2023, 23, 149. [Google Scholar] [CrossRef]
- Cao, Y.; Li, S.; Han, S.; Wang, D.; Zhao, J.; Xu, L.; Liu, H.; Lu, T. Characterization and application of a novel Aeromonas bacteriophage as treatment for pathogenic Aeromonas hydrophila infection in rainbow trout. Aquaculture 2020, 523, 735193. [Google Scholar] [CrossRef]
- Kumari, R.; Yadav, R.; Kumar, D.; Chaube, R.; Nath, G. Evaluation of bacteriophage therapy of Aeromonas hydrophila infection in a freshwater fish, Pangasius buchanani. Front. Aquac. 2023, 2, 1201466. [Google Scholar] [CrossRef]
- Ryan, E.M.; Gorman, S.P.; Donnelly, R.F.; Gilmore, B.F. Recent advances in bacteriophage therapy: How delivery routes, formulation, concentration and timing influence the success of phage therapy. J. Pharm. Pharmacol. 2011, 63, 1253–1264. [Google Scholar] [CrossRef]
- Donati, V.L.; Dalsgaard, I.; Sundell, K.; Castillo, D.; Er-Rafik, M.; Clark, J.; Wiklund, T.; Middelboe, M.; Madsen, L. Phage-Mediated Control of Flavobacterium psychrophilum in Aquaculture: In vivo Experiments to Compare Delivery Methods. Front. Microbiol. 2021, 12, 628309. [Google Scholar] [CrossRef]
- Fang, D.; Jin, L.; Song, Q.; Zou, X.; Feng, L.; Li, L.; Zhang, H.; Hu, K.; Lin, M.; Fang, W. Pharmacokinetic study of Vibrio parahaemolyticus phage Φvp140 in Litopenaeus vannamei following oral administration. Aquaculture 2025, 606, 742619. [Google Scholar] [CrossRef]
- Cui, H.; Cong, C.; Wang, L.; Li, X.; Li, J.; Yang, H.; Li, S.; Xu, Y. Protective effectiveness of feeding phage cocktails in controlling Vibrio harveyi infection of turbot Scophthalmus maximus. Aquaculture 2021, 535, 736390. [Google Scholar] [CrossRef]
- Zhong, H.; Huang, W.; Lin, K.T.; Zhang, Q.; Deng, Y.; Zhang, R.; Ma, R. A virulent phage vB_VpaP_R28Z infecting Vibrio parahaemolyticus with potential for therapeutic application. BMC Microbiol. 2025, 25, 433. [Google Scholar] [CrossRef]
- Jin, M.; Yu, M.; Feng, X.; Li, Y.; Zhang, M. Characterization and comparative genomic analysis of a marine Bacillus phage reveal a novel viral genus. Microbiol. Spectr. 2024, 12, e00037-24. [Google Scholar] [CrossRef]
- Feng, X.; Yan, W.; Wang, A.; Ma, R.; Chen, X.; Lin, T.H.; Chen, Y.L.; Wei, S.; Jin, T.; Jiao, N.; et al. A Novel Broad Host Range Phage Infecting Alteromonas. Viruses 2021, 13, 987. [Google Scholar] [CrossRef] [PubMed]
- Delcher, A.L.; Bratke, K.A.; Powers, E.C.; Salzberg, S.L. Identifying bacterial genes and endosymbiont DNA with Glimmer. Bioinformatics 2007, 23, 673–679. [Google Scholar] [CrossRef] [PubMed]
- Besemer, J.; Lomsadze, A.; Borodovsky, M. GeneMarkS: A self-training method for prediction of gene starts in microbial genomes. Implications for finding sequence motifs in regulatory regions. Nucleic Acids Res. 2001, 29, 2607–2618. [Google Scholar] [CrossRef] [PubMed]
- Meier-Kolthoff, J.P.; Göker, M. VICTOR: Genome-based phylogeny and classification of prokaryotic viruses. Bioinformatics 2017, 33, 3396–3404. [Google Scholar] [CrossRef]
- Nishimura, Y.; Yoshida, T.; Kuronishi, M.; Uehara, H.; Ogata, H.; Goto, S. ViPTree: The viral proteomic tree server. Bioinformatics 2017, 33, 2379–2380. [Google Scholar] [CrossRef] [PubMed]
- Moraru, C.; Varsani, A.; Kropinski, A.M. VIRIDIC-A Novel Tool to Calculate the Intergenomic Similarities of Prokaryote-Infecting Viruses. Viruses 2020, 12, 1268. [Google Scholar] [CrossRef] [PubMed]
- Shein, A.M.S.; Wannigama, D.L.; Hurst, C.; Monk, P.N.; Amarasiri, M.; Wongsurawat, T.; Jenjaroenpun, P.; Phattharapornjaroen, P.; Ditcham, W.G.F.; Ounjai, P.; et al. Phage cocktail amikacin combination as a potential therapy for bacteremia associated with carbapenemase producing colistin resistant Klebsiella pneumoniae. Sci. Rep. 2024, 14, 28992. [Google Scholar] [CrossRef] [PubMed]

| VF Category | Related Genes |
|---|---|
| Nutritional/Metabolic factor | allB, allS, argP, asbC, basB, basC, basD, basF, basG, bauA, bauB, bauC, bauD, bauE, bauE, bioA, bioB, bioD, bplA, carA, carB, ccmA, ccmB, ccmC, ccmE, ccmF, chuA, chuT, chuU, chuV, chuW, dhbE, dhbF, entA, entB, entC, entE, fbpC, feoB, fepA, fepC, fepD, fpvA, fpvI, hgpB, hitC, hpt, iroC, lbtC, lplA1, mgtB, mgtC, PA2383, panC, phzG1, ptxR, purCD, purM, pvcA, pvcB, pvdD, pvdE, pvdH, pvdI, pvdL, pyrB, shuA, shuU, shuV, shuY, ybtP, ybtQ |
| Motility | AHML_RS07540, cheA, cheA-2, cheB, cheB-2, cheD, cheR, cheR-3, cheV, cheV3, cheW, cheY, cheZ, eptC, flaA, flaG, flaH, flaJ, fleN, fleQ, fleR, fleR/flrC, fleS, fleS/flrB, flgA, flgB, flgC, flgD, flgE, flgF, flgG, flgH, flgI, flgJ, flgK, flgL, flgM, flgN, flgO, flgP, flhA, flhB, flhF, flh, fliA, fliC, fliE, fliF, fliG, fliH, fliI, fliJ, fliK, fliL, fliM, fliN, fliO, fliP, fliQ, fliR, flmD, flmH, flrA, flrB, flrC, lafK, motA, motC, motX, motY, nueA, nueB, PA1459, PA1464, PA3349, pdxJ, pomA2, pomB2, pseB |
| Adherence | cadF, cgsD, fimA, fimB, fimC, fimD, fimE, fimF, fimZ, frpC, gbpA, htpB, IlpA, lap, mam7, mshA, mshB, mshC, mshD, mshE, mshF, mshG, mshH, mshI, mshJ, mshK, mshL, mshM, mshN, mshO, mshP, mshQ, pebA, pilG, pilJ, pilR, psaC, rpoN, tapB, tapC, tapD/pilD, tapF, tapM, tapN, tapO, tapP, tapQ, tapT, tapU, tapV, tapW, tcpI, tppF, tsaP, tufA, vfr, yagX/ecpC |
| Immune modulation | acpXL, bexA, bplF, cap8J, cap8P, cpsA/uppS, cpsB/cdsA, cysC, ddrA, fabZ, flmK, FTT_RS04140, galE, galU, gmhA, gmhA/lpcA, gtrB, hisH2, htrB, kdkA, kdsA, kdtB, KP1_RS17330, kpsF, lgtF, lpxA, lpxA/glmU, lpxB, lpxC, lpxD, lpxH, lpxK, lpxL, lsgD, msbA, ompA, ompP2, opsX/rfaC, orfM, pbpG, pgi, pks1, pks15, ppsB, ppsC, rfaD, rfaE, rfbB, rfbC, rfbD, rffG, rpe, Rv2962c, ugd, waaA, waaF, waaQ, wbpK, wbpL, wbpM, wbtL, wbuZ, wcbN, wcbP, wcbT, wecA |
| Effector delivery system | AHA_RS09305, ascN, atsA, atsB, atsD, atsG, atsH, atsI, atsJ, atsK(Aerolysin A), atsL, atsP, atsQ, atsS, bprB, BPS_RS26875, BPS_RS26880, bsaN, CBU_0270, CBU_1079, CBU_1434, CBU_1566, CBU_1594, clpB, dotA, dotU, exeA, exeB, exeC, exeD, exeE, exeF, exeG, exeH, exeI, exeJ, exeK, exeL, exeM, exeN, exsA, glgX, hcp1, ipaH4.5, lasA, lidL, lirB, LPG_RS00105, LPG_RS00200, LPG_RS14840, lpnE, PA1663, ricA, ssrA, ssrB, tagT, vasH, vasK/icmF, vgrG1, vgrG2, vgrG3, vipA, vipB |
| Regulation | bvgA, bvgS, bvrR, cdpA, csrA, devR/dosR, fur, letA, letS, mprA, phoP, phoQ, phoR, rcsB, relA, rpoS, sigA/rpoV |
| Biofilm | adeF, adeG, algB, algC, algQ, algR, algU, algW, algZ, bopD, uxS, mucD, mucP, pmlR/bspR1, vpsG |
| Exotoxin | acpC, aerA/act, clbF, clbG, clbK, clbS, cyaB, cylA, cylB, hlyA, hlyB, hlyD, nheB, plcD, rtxA, rtxB, rtxC, rtxD, tlh, toxA, zot |
| Stress survival | ahpC, clpC, clpE, clpP, katA, LPG_RS11000, recN, sodB |
| Antimicrobial activity | acrA, acrB, farA, farB, mtrC, mtrD |
| Exoenzyme | BAS_RS06430, hap/vvp, nagI, stcE, tlyC |
| Invasion | ibeB, ompA |
| Post-translational modification | Mip, prsA2 |
| Others | aatC, acfB, icl |
| Bacterial Species | Strain | Source | Lytic Ability |
|---|---|---|---|
| Aeromonas hydrophila | 2408 | diseased American eels (Anguilla rostrata) b | + |
| Aermonas hydrophila | 1A00007 | Mullet (Mugil cephalus) a | − |
| Aermonas hydrophila | 1A00032 | Starry flounder (Platichthys stellatus) a | − |
| Aermonas hydrophila | 1A00179 | Yellowfin Seabream (Acanthopagrus latus) a | − |
| Aermonas veronii | 1A00013 | Mud a | − |
| Aermonas veronii | 1A02245 | Yellowfin Seabream (Acanthopagrus latus) a | − |
| Aermonas salmonicida | 1K03261 | Yellowfin Seabream (Acanthopagrus latus) a | − |
| Aeromonas caviae | FJML01 | diseased American eels (Anguilla rostrata) b | − |
| Aeromonas encheleia | FJML02 | diseased American eels (Anguilla rostrata) b | − |
| Aeromonas allosaccharophila | FJML03 | diseased American eels (Anguilla rostrata) b | − |
| Acinetobacter piscicola | FJML04 | diseased American eels (Anguilla rostrata) b | − |
| Citrobacter arsenatis | FJML05 | diseased American eels (Anguilla rostrata) b | − |
| Exiguobacterium enclense | FJML06 | diseased American eels (Anguilla rostrata) b | − |
| Raoultella ornithinolytica | FJML07 | diseased American eels (Anguilla rostrata) b | − |
| Pseudomonas pharyngis | FJML08 | diseased American eels (Anguilla rostrata) b | − |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Zhang, M.; Feng, X.; Wang, J.; Qu, W.; Jin, M. Isolation, Characterization, and Evaluation of a Lytic Jumbo Phage Z90 Against Aeromonas hydrophila in American Eels (Anguilla rostrata). Antibiotics 2026, 15, 27. https://doi.org/10.3390/antibiotics15010027
Zhang M, Feng X, Wang J, Qu W, Jin M. Isolation, Characterization, and Evaluation of a Lytic Jumbo Phage Z90 Against Aeromonas hydrophila in American Eels (Anguilla rostrata). Antibiotics. 2026; 15(1):27. https://doi.org/10.3390/antibiotics15010027
Chicago/Turabian StyleZhang, Miaosen, Xuejin Feng, Jianxin Wang, Wu Qu, and Min Jin. 2026. "Isolation, Characterization, and Evaluation of a Lytic Jumbo Phage Z90 Against Aeromonas hydrophila in American Eels (Anguilla rostrata)" Antibiotics 15, no. 1: 27. https://doi.org/10.3390/antibiotics15010027
APA StyleZhang, M., Feng, X., Wang, J., Qu, W., & Jin, M. (2026). Isolation, Characterization, and Evaluation of a Lytic Jumbo Phage Z90 Against Aeromonas hydrophila in American Eels (Anguilla rostrata). Antibiotics, 15(1), 27. https://doi.org/10.3390/antibiotics15010027






