Abstract
Xylella fastidiosa subsp. pauca (Xfp) infects olive trees and other hosts in Southern Apulia (Italy), devastating agriculture and landscape. A containment strategy of the disease requires quick and sensitive detection tools. Therefore, a colorimetric LAMP protocol was developed using as a template a crude alkaline sap obtained from incubation of 50–60 mg of thin slices of olive twigs in a NaOH-containing buffer. This rapid molecular assay can be performed directly in the field, as it needs only a portable isothermal block. Tissues of the same olive trees analysed by this technique were also compared to qPCR (using purified total plant DNA as template) as well as digital droplet PCR (on the same crude alkaline extracts used in cLAMP). A titration of the cLAMP reaction with healthy olive sap, spiked with dilutions of in vitro cultivated Xfp cells and plasmid DNA containing the target sequence, gave positive detection results as low as 102 CFU/mL and up to 169.2 target copies/µL, equivalent to about 0.9 pg of the genomic DNA. A portable, sensitive and target-specific Xfp field test was developed, which has a 40 min sample-to-answer time and does not require any DNA isolation procedure or laboratory equipment. The application of this detection assay could help the monitoring and containment of the disease spread.
1. Introduction
Xylella fastidiosa (Xf) is a non-flagellate, Gram-negative phytopathogenic bacterium belonging to the Xanthomonadaceae family [1], evaluated as a ‘priority pest’ in Europe (Regulation EU 2019/1702), since it causes severe diseases worldwide, such as Pierce’s disease in grapevines, citrus variegated chlorosis (CVC), and olive quick decline syndrome (OQDS) [2]. The bacterium multiplies and colonizes the vascular system of the host plants [3,4], where it produces cell aggregates which cause the occlusion of xylem vessels and affect the water and minerals carriage. The main symptoms visible are leaf scorching, twig desiccation, and gradual plant dieback. In natural conditions, it spreads locally by xylem-feeding insects of the Cicadellidae and Aphrophoridae families [5]. Conversely, the long-distance bacterium dissemination and establishment occurs through international movements of infected plant materials. The bacterium host range consists of over 600 plant species [6], mainly perennial, with most of the infected species that do not show remarkable alterations.
For this reason, it is important to understand the disease distribution and have affordable, reliable, and prompt diagnostic methods to avoid further introductions in the Xf-free areas. There are several PCR-based detection techniques available to characterize and differentiate between Xf subspecies and strains, widely used in the laboratory, but not easily transferable to the field.
Although qPCR assay represents the golden standard for bacterium detection and is recognized as an official method in the EU (Reg (EU) 2020/1201—Annex IV), due to its high sensitivity and specificity, it requires a molecular biology laboratory, highly qualified personnel, and bulky and expensive instruments. Moreover, the need for high-purity nucleic acid extraction (EPPO PM 7/24 (4) [7]) limits the testing capacity of this method, as it requires laborious and time-consuming DNA purification steps from infected plant samples, followed by one hour qPCR run time. Hence, the qPCR technique is not able to meet the current demand for a rapid and simple field-based or point-of-care testing (POCT) assay, which could be used at the entry points of plant materials or in the containment area, to facilitate faster detection of Xf and to reduce or avoid further spread.
Loop-mediated isothermal amplification (LAMP) is a low-cost and portable point-of-care diagnostic technology that provides nucleic acid amplification in a short time. It uses 4 to 6 specially designed primers complementary to genomic DNA and takes advantage of a particular DNA polymerase, obtained from Bacillus stearothermophilus (Bst), which has a strand displacement activity and is more tolerant to inhibitors present in the samples [8]. Therefore, the DNA denaturation by heating and purification phases can be overcome, using—as a template—crude extracts in a simple extraction buffer, facilitating the chemical disruption of plant tissues [9].
The amplification reaction is carried out for 15–60 min under constant temperature (ranging from 60° to 70 °C) and produces a stem-loop DNA with several inverted repeats and cauliflower-like structures that make amplicons more stable [10]. The isothermal feature of the reaction needs only a heat block or water bath, equipped with a rechargeable battery, set at the target temperature to successfully amplify DNA. Therefore, it can be adequately implemented in situ with minimal requirements [11].
For this reason, LAMP is continuously being implemented in the medical, agriculture, and food sectors. Several LAMP methods have been developed for an easy molecular detection based on fluorescence signal [12], turbidity [13], colorimetric [14] and even by visualization through the naked eye [15], bypassing the agarose gel electrophoresis examination and preventing the carry-over contamination.
LAMP has been already applied to identify phytopathogens in host plants and in insect vectors, such as Spiroplasma citri [16] and Guignardia citricarpa [9] in citrus tree, Xanthomonas gardneri in tomato and pepper [17], Olea europea geminivirus (OEGV) and Pleurostoma richardsiae in Olea europea [18,19], Phytophthora infestans in tomato and potato [20], and Flavescence Dorée phytoplasma (FD) both in grapevine [21,22] and in its Scaphoideus titanus vector [23].
Even though the detection of Xf is difficult, due to the uneven distribution in host plants and the bacterial concentration varying at different disease stages, some protocols of LAMP molecular tool for early in-field detection have been successfully applied, using as template either purified DNA or even crude extracts, with a fluorescent measure of the amplification [24,25,26,27,28,29]. Some of these tools rely on a fluorometric analysis, while others rely on a colorimetric evaluation of the results. Recently, other advanced techniques for the comparison of serological lab-on-a-chip detection of Xylella fastidiosa subsp. pauca (Xfp) from plant tissues were applied in the studies of [30,31]. Conversely, a comparison of the detection between digital droplet PCR (ddPCR) and qPCR on extracted Xf DNA from bacterial culture was performed by Dupas et al., (2019) [32].
The goal of this study—in the frame of the EU FREE@POC project (https://www.freepoc.eu; accessed on 10 December 2022)—is the fine-tuning and validation of a novel colorimetric LAMP (cLAMP) assay based on primer sequences already published [24]. In the current work, the rapid detection of Xfp was pursued, starting from alkaline crude olive extracts obtained at room temperature without any DNA purification step, and employing a pH-sensitive dye for an unaided visual examination of the results. Overall, this colorimetric assay could be easily extended to other plant pathogens and be implemented for surveillance of both emerging and widespread agricultural threats for on-site diagnostic testing.
2. Materials and Methods
2.1. Bacterial Strains and Growth Conditions
For this study, the Xfp strain ‘De Donno’, isolated from infected olive trees in Salento (Apulia, Italy) [33,34] and grown on PD3 agar plates at 28 °C for seven days, was used. Bacterial suspensions of scraped cultured colonies were prepared in sterile phosphate-buffered saline (PBS 1X, pH 7.4) and their titer was estimated by measuring the optical density value at 600 nm (OD600) and adjusted to 0.5, equivalent to 4 × 108 CFU/mL [35], before a further confirmation by plate counting. To assess the specificity of the cLAMP assay, several non-target bacterial species available in our laboratory were tested. Xanthomonas campestris pv campestris, a Gram-negative bacterium that causes black rot of Brassicaceae, Paraburkholderia phytofirmans strain PsJN, a rhizosphere-colonizing bacterium known to be effective in reducing Pierce’s disease symptom severity in grapevines [36], and Pseudomonas marginalis, a Gram-negative bacterium responsible for field and storage soft rots in different plant species, were cultivated on Tryptone Soy Agar (TSA) plate medium at 28 °C for one, two, or three days, respectively. The Gram-negative epiphytic bacterium Rahnella aquatilis [37] was cultivated on Luria Bertani (LB) solid medium for one day at 28 °C and Escherichia coli strain BL21(DE3) was grown at 37 °C overnight. All these non-target bacterial species were used in LB suspensions at concentrations of no less than 0.5 OD600 (Table S1).
2.2. Preparation of Cloned Plasmid Standard
A fragment (1414 bp-long) encoding part of the ribosome maturation factor rimM (a single copy gene of the Xfp genome) [24] was amplified from genomic DNA of Xfp ‘De Donno’ strain. The purified PCR amplicon was cloned into the pSC-A vector (Agilent Technologies, Santa Clara, CA, USA). Insert-containing colonies were identified, and their plasmid DNA was sequenced for confirmation. The concentration of plasmid DNA was measured using a NanoPhotometer™ N60 UV/Vis spectrophotometer (Implen, München Germany) and recalculated to obtain a final value expressed as plasmid copies/μL. This cloned plasmid DNA, containing the target sequence designed in LAMP and qPCR tests as in reference [24], was used to assess the analytical sensitivity of the cLAMP and ddPCR techniques.
2.3. Plant Materials
For preliminary tests and method development, olive samples (young, lignified twigs 1 or 2 years old) were collected in groves placed in the demarcated area in Apulia, where OQDS symptoms are prevalent. The Xfp-infection status of the selected plant material was assessed by qPCR on purified DNA extracts [24,38]. Negative control samples were collected from Xf-free olive trees. According to quarantine recommendations, samples were stored at 4 °C in closed plastic bags to prevent cross-contamination until laboratory processing.
2.4. Crude Plant Extracts Preparation
For the preparation of crude plant extract, an extraction buffer (EB) [NaOH 0.4M; Sodium diethyldithiocarbamate (Na-DIECA) 2%; Polyvinylpyrrolidone (PVP 25–30k) 1%; 0.1% Triton-X100] was set up to break the bacterial cells and release target DNA molecules without interfering with the reaction mixture pH, thus potentially causing false positive outcomes. After the incubation with olive tissues, the neutralization of the highly alkaline condition in the crude sap was achieved by adding an equal volume of neutralization buffer (NB) made of HCl 0.4 M and NaCl 50 mM. Approximately 50–60 mg of thin slices (0.3–0.5 mm length) of stem or leaf midribs/petioles were trimmed from olive twigs with sodium hypochlorite-disinfected scissors and put in a 1.5 mL microcentrifuge tube. The slices were soon incubated with 500 µL EB for about 15 min at room temperature and manually shaken; 50 µL of the incubated crude sap were neutralized by shortly mixing with the same quantity of NB. Finally, 10 µL of the neutralized mixture were diluted 1:10 in sterile water to further reduce plant debris and potential inhibitor effect. One and two µL of this mixture were used as templates in the cLAMP and ddPCR assays, respectively.
2.5. cLAMP Assay and Device
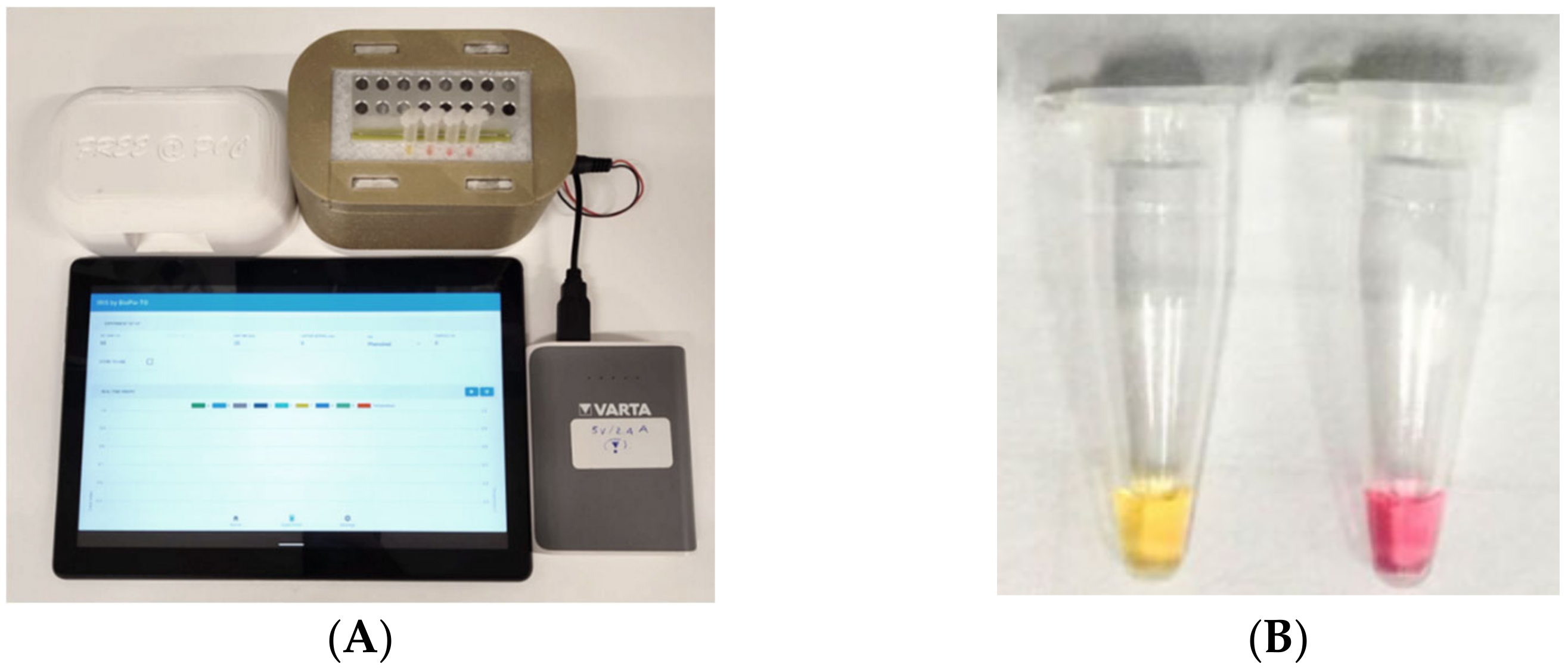
The cLAMP assay was performed in standard 0.2 mL microtubes in a final volume of 25 μL, which comprises 12.5 μL Bst 2.0 WarmStart® DNA Polymerase 2x Master Mix (New England Biolabs, Ipswich, MA) containing the Phenol red indicator, 2.5 μL LAMP primer [24] mix (0.2 μM of primer F3/B3, 1.6 μM of primers FIP/BIP, 0.4 μM of primers LF/LB), 1 μL of neutralized and diluted crude DNA extract, double-distilled water to a final volume; 25 μL of mineral oil were added to each tube to cover the LAMP mix. Additionally, a healthy control (a crude alkaline extract from a Xf-free olive tree) and a no-template control (NTC) were also included. The reaction mixture was incubated for 30 min at 65 °C in a portable incubator device (Figure 1A) endowed with eight PCR tubes, recharged by a power bank, connected via Bluetooth to a smartphone or tablet, and activated by an Android application. The device was manufactured by using 3D-printing technology. The system was designed using a CAD software (Fusion 365, Autodesk, accessed on 15 March 2022). Two different materials were implemented in the device. An HT from Colorfabb which has a high temperature resistance is located in the centre of the system (clear color). This part has a slot in order to receive the heating element, while on top of the heating element a metal part was fixed with a thermally conductive glue. The metal block has cylindrical slots for housing PCR tubes. The main case of the system consists of PLA from Prusa (yellow-gold color). The system is connected through a Bluetooth protocol to a smart device, through which time and temperature of the system can be regulated. An external power bank provides power to the system (5V, 2A). The phenol red indicator in the master mix allows the visual detection of the result immediately after the removal of the tubes from the device. It relies on the production of protons and a pH decrease, which takes place when the DNA polymerase activity specifically operates on the target, revealing a colour shift from pink to yellow (Figure 1B). End-point picture records of the reaction tubes were taken with a smartphone camera. Occasionally, reaction products were analysed in agarose gel electrophoresis (1.5% agarose in Tris-acetate-EDTA buffer).

Figure 1.
(A) Overview of the 3D-printed portable thermal block device. The plastic box contains a heating metal block, set to operate at 65 °C for 30 min. The device is powered by a portable power bank and connected via Bluetooth to a tablet. (B) The positive reaction (amplified DNA) is visualized by the color change from pink (negative signal) to yellow (positive signal).
2.6. Analytical Sensitivity of cLAMP Assay
To evaluate the limit of detection (LoD) of the cLAMP assay for the amplification of the Xfp target DNA on crude alkaline extracts, both the purified plasmid DNA and the cultured Xfp cell suspension were employed. Plasmid was 10-fold serially diluted (from 90 ng/µL to 0.9 fg/µL) in distilled water, and 1 µL of each dilution was added into 100 µL of healthy alkaline olive extract. After neutralization and dilution in water of the extracts, the final plasmid amount in the reaction tubes was further reduced of 2 × 103 times from the starting concentration.
The bacterial suspension (10 µL) was similarly diluted in 90 µL of healthy alkaline olive sap to prepare 10-fold serial dilutions at concentration ranging from 106 to 10 CFU/mL. One µL of each dilution (after 1:1 neutralization and 1:10 dilution in water) was used as a template in the cLAMP reaction mixture. Negative internal control and no-template control (NTC) were also included in each experiment.
2.7. Evaluation of cLAMP Performance on Field-Grown Olive Samples
Validation of the cLAMP assay was conducted on 282 field-grown olive trees, randomly selected and collected in the Xfp-infected demarcated area as well as in the border buffer and containment zones. The geo-positioning of the orchards, or at least their location as municipality, was always recorded. From all the selected trees, regardless of the symptom expression and the irregular pathogen distribution, twigs were collected from the four cardinal points of the canopy. Samples collected in plastic bags, were immediately processed when in-field tests were performed, or alternatively stored at 5 °C for 1–2 days before being processed in the laboratory. Crude plant extracts were obtained with the alkaline buffer as described above, and scissors were rinsed with sodium hypochlorite before processing each sample. To establish accuracy and reliability of cLAMP assay in comparison with the golden standard qPCR [24], purified DNA was extracted from the same field samples tissues (EPPO PM 7/24 (4) [7]) and subsequently tested for Xfp detection by qPCR. The performance criteria diagnostic sensitivity, diagnostic specificity, and accuracy were calculated according to EPPO PM 7/76 (5) [39].
2.8. DNA Extraction and Real Time PCR (qPCR) Assay
About 0.5 g of small pieces of olive twigs were crushed in 5 mL of CTAB extraction buffer with a semi-automated Homex homogenizer (Bioreba, AG, Switzerland). After a 15 min incubation at room temperature, 1 mL of homogenate was recovered from plastic bag, heated at 65 °C for 20 min, and centrifuged for 10 min at 16,000× g. DNA was purified with the automatic Maxwell® RSC PureFood GMO and Authentication Kit (Promega Corporation, Madison, WI, USA) following manufacturer’s instructions, eluted in 100 µL of Elution buffer and stored at −20 °C until use. The qPCR tests were performed following the standard EPPO (PM7/24 (4)—Annex 5), using the primer set already described [24] with a TaqManTM Fast Universal Master Mix (Applied Biosystems, Waltham, MA, USA), in a CFX96 thermal cycler (BioRad Laboratories, Hercules, CA, USA). All the samples were run in duplicate wells to obtain the final average Cq value. Negative internal (NIC) and no-template controls (NTC) were included in the run. Cq values were analysed using the CFX Maestro 1.1 software v. 4.1 (BioRad Laboratories) with automatic calculation of the threshold baseline. An internal laboratory cutoff was established and samples with a Cq ≤ 32 were considered positive, while Cq > 32 were considered as undetermined [38]; when they produced not absorption signals (NA), they were assessed as negative.
2.9. Droplet Digital PCR Assay
The same crude extracts, obtained from a subset of olive tree samples (as described in Section 2.4) and tested in cLAMP, were submitted to a parallel absolute quantification of target copy number per microliter through amplification by digital droplet PCR; ddPCR assay was performed with the QX200TM ddPCR system (BioRad Laboratories), according to the manufacturer’s instructions, by adapting the qPCR protocol [24] to detect Xfp. At first, 10-fold serial dilutions of plasmid DNA spiked in healthy olive alkaline sap were submitted to ddPCR quantification in order to assess the detection limit of the assay. Then, a subset of 115 out of the full set of 282 olive samples was selected to compare results generated by qPCR, cLAMP, and ddPCR. The ddPCR-optimized reaction mixture contained the same primers and probes (at a modified concentration) as the above qPCR mixture [24]. In detail: 2X ddPCR™ Supermix for Probes (No dUTP) (Bio-Rad), 1.2 μL (600 nM) Xf forward and reverse primers (XF-F and XF-R), 0.6 μL (300 nM) 6’FAM/BHQ-1 labelled probe (XF-P), 2 µL of crude neutralized and diluted plant extract and double-distilled water to a final volume of 20 μL. A negative internal control (NIC) and a no-template control (NTC) were included in each run. Each sample was tested in duplicate reaction. QX-Manager-V.1.2-STD software (Bio-Rad Laboratories) was used for data acquisition and analysis; ddPCR reactions with less than 10,000 generated droplets were excluded from the analysis. The diagnostic sensitivity, specificity, and accuracy parameters of ddPCR test, in comparison with qPCR results of the same samples, were also calculated.
2.10. Statistical Analysis
The agreement between the different detection methods was evaluated by Cohen’s kappa statistical analysis [40]. A linear regression analysis was performed to allow an estimate of the values of positive droplets obtained in ddPCR based on the Cq values resulting from the qPCR assay. Their correlation was also evaluated based on Pearson’s correlation coefficient [41]. All statistical analyses were performed with the SPSS 23.0 software program (IBM, Armonk, NY, USA). The diagnostic performances were statistically evaluated based on the area under the curve (AUC) values, calculated on the Receiver operating characteristic (ROC) curves constructed for each of the cLAMP, ddPCR, and qPCR diagnostic assays.
3. Results
3.1. Specificity of Xfp Colorimetric LAMP Assay
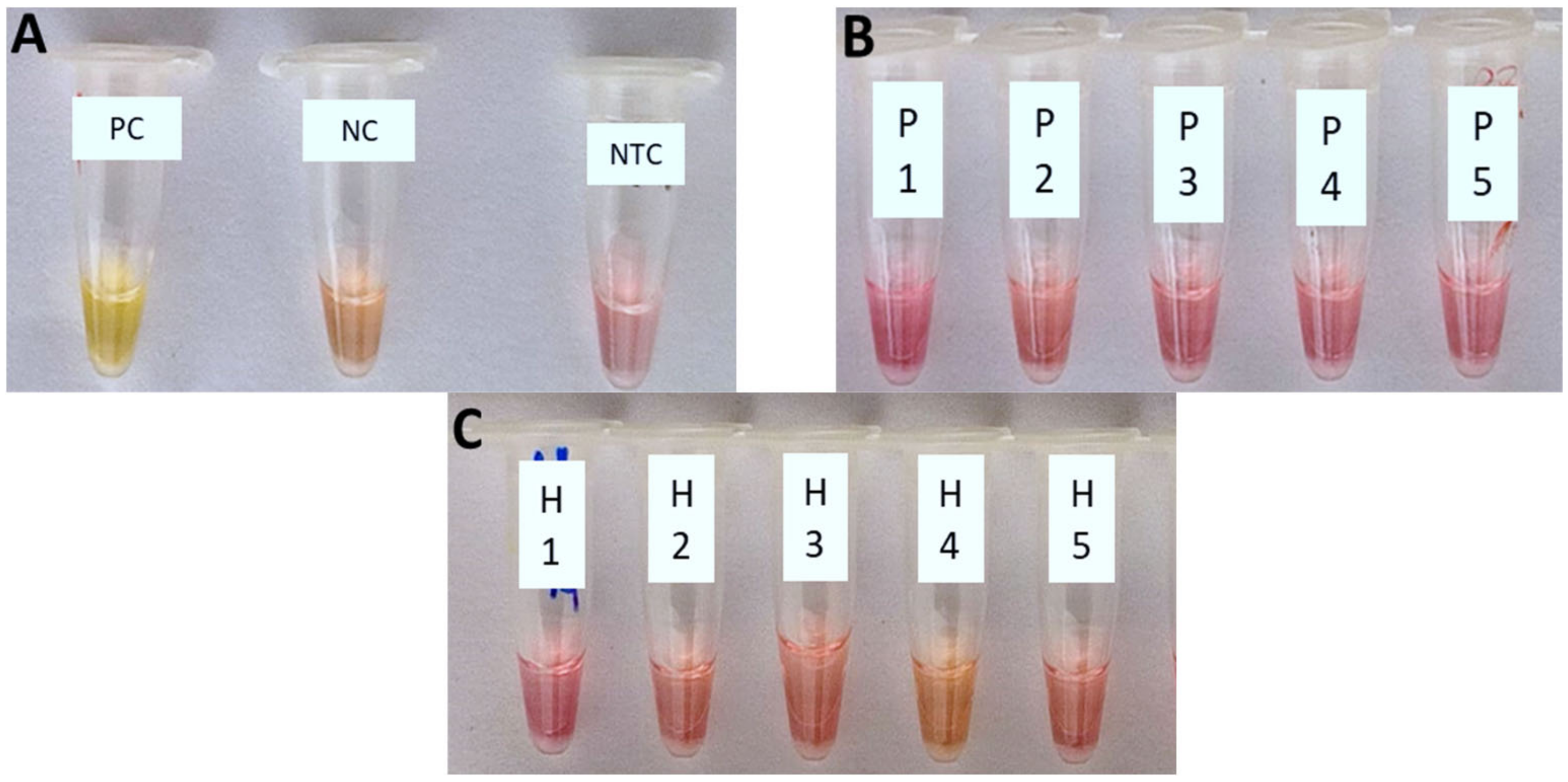
The cLAMP assay was demonstrated to have a high specificity detecting only the target DNA of Xfp (Figure 2A), with no cross-reaction with the other non-target bacterial species tested, spiked in PBS (Figure 2B) or healthy olive crude sap (Figure 2C).
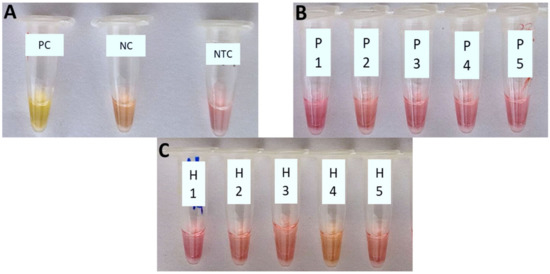
Figure 2.
The cLAMP reactions. (A) PC (positive Xfp control): bacterial suspension of Xylella fastidiosa subsp. pauca strain ‘De Donno’ (approx. 105 CFU/mL); NC (negative control): healthy olive sap; NTC: No-Template Control; non-target bacteria spiked in PBS 1X (B) and healthy olive crude sap (C). 1: Rahnella aquatilis; 2: Escherichia coli BL21(DE3); 3: Paraburkholderia phytofirmans strain PsJN; 4: Pseudomonas marginalis; 5: Xanthomonas campestris pv. campestris.
3.2. Sensitivity of Xfp Colorimetric LAMP Assay
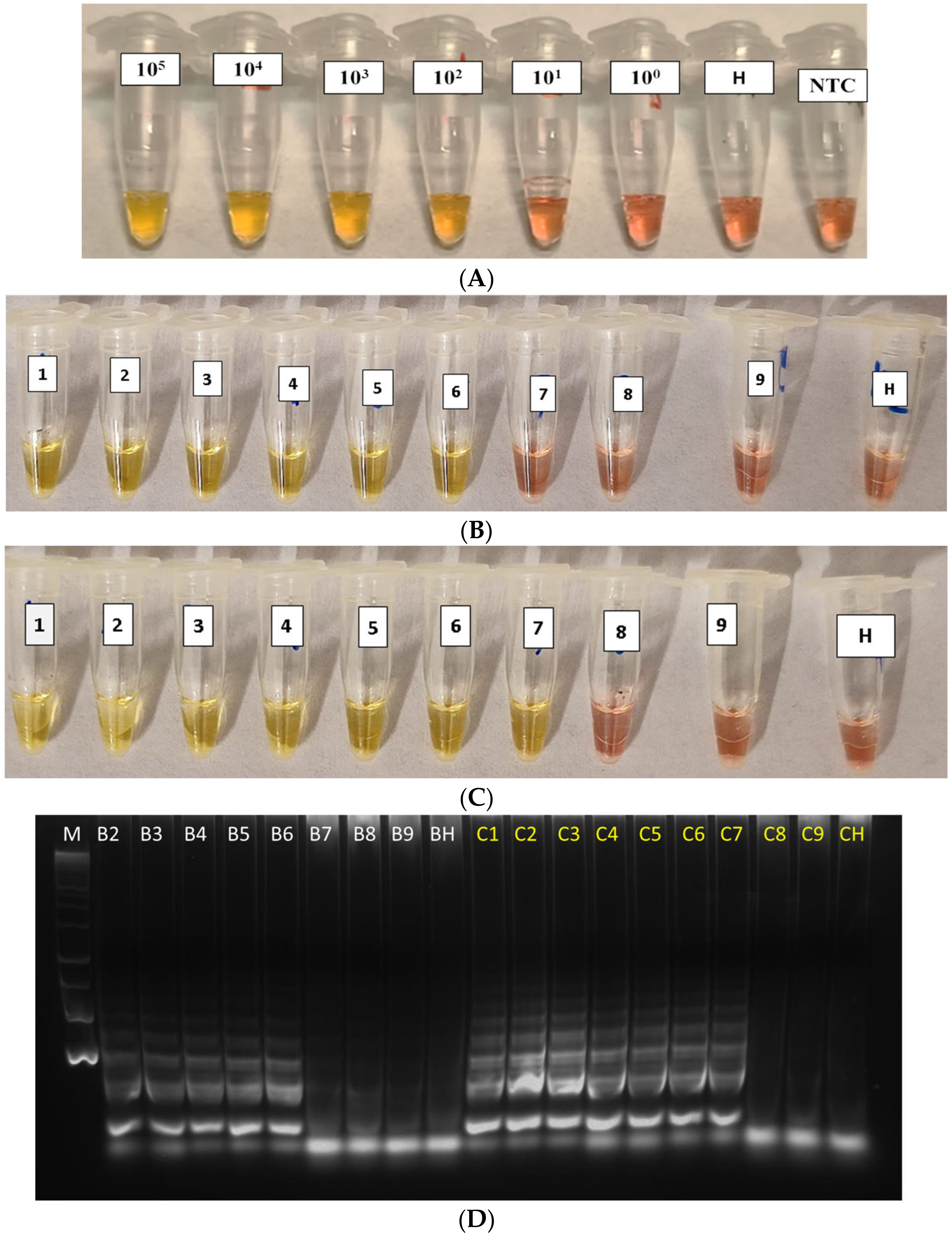
The analytical sensitivity of the cLAMP assay, using the 10-fold dilutions of the Xfp bacterial suspension and of the rimM- recombinant plasmid DNA, both spiked in healthy alkaline olive sap, showed that the detection limit (LoD) was 102 CFU/mL for the bacterial culture (Figure 3A) and 0.9 pg/µL for rimM-plasmid DNA (equivalent to 169.2 target copies/µL; Figure 3B, tube 6). A difference in sensitivity of one lower 10-fold dilution (at 0.09 pg/µL, Figure 3C, tube 7) was sometimes obtained when experiments were run in duplicate. To verify this result, amplification products were run in agarose gel electrophoresis (Figure 3D, lanes C7 vs. B6). A reaction time of 30 min was enough to generate clear positive reactions.
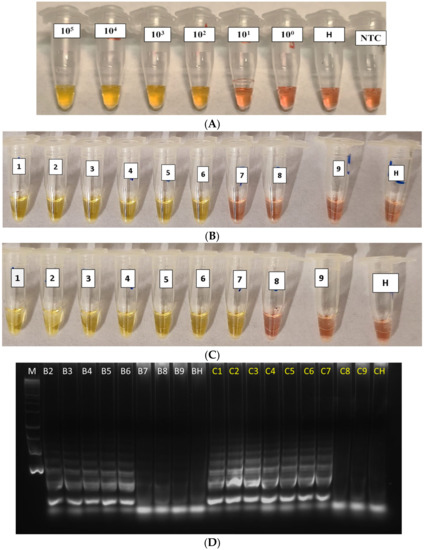
Figure 3.
Healthy crude sap spiked with 10-fold serial dilution of (A) bacterial pure culture from 105 to 10 CFU/mL (H: Healthy control; NTC: NoTemplate Control) and (B–C) two replicates of purified plasmid DNA dilutions in healthy alkaline olive sap (from 90 ng/µL [tube 1] to 0.9 fg/µL [tube 9]). (D) Amplified products of the panel B (in white) and C (in yellow) were analysed by gel electrophoresis. M: Molecular DNA ladder; B2–B9: 9 ng/µL to 0.9 fg/µL; C1–C9: 90 ng/µL to 0.9 fg/µL; BH and CH: Healthy controls.
3.3. Validation of cLAMP Assay on Field-Grown Olive Samples by Comparison with qPCR
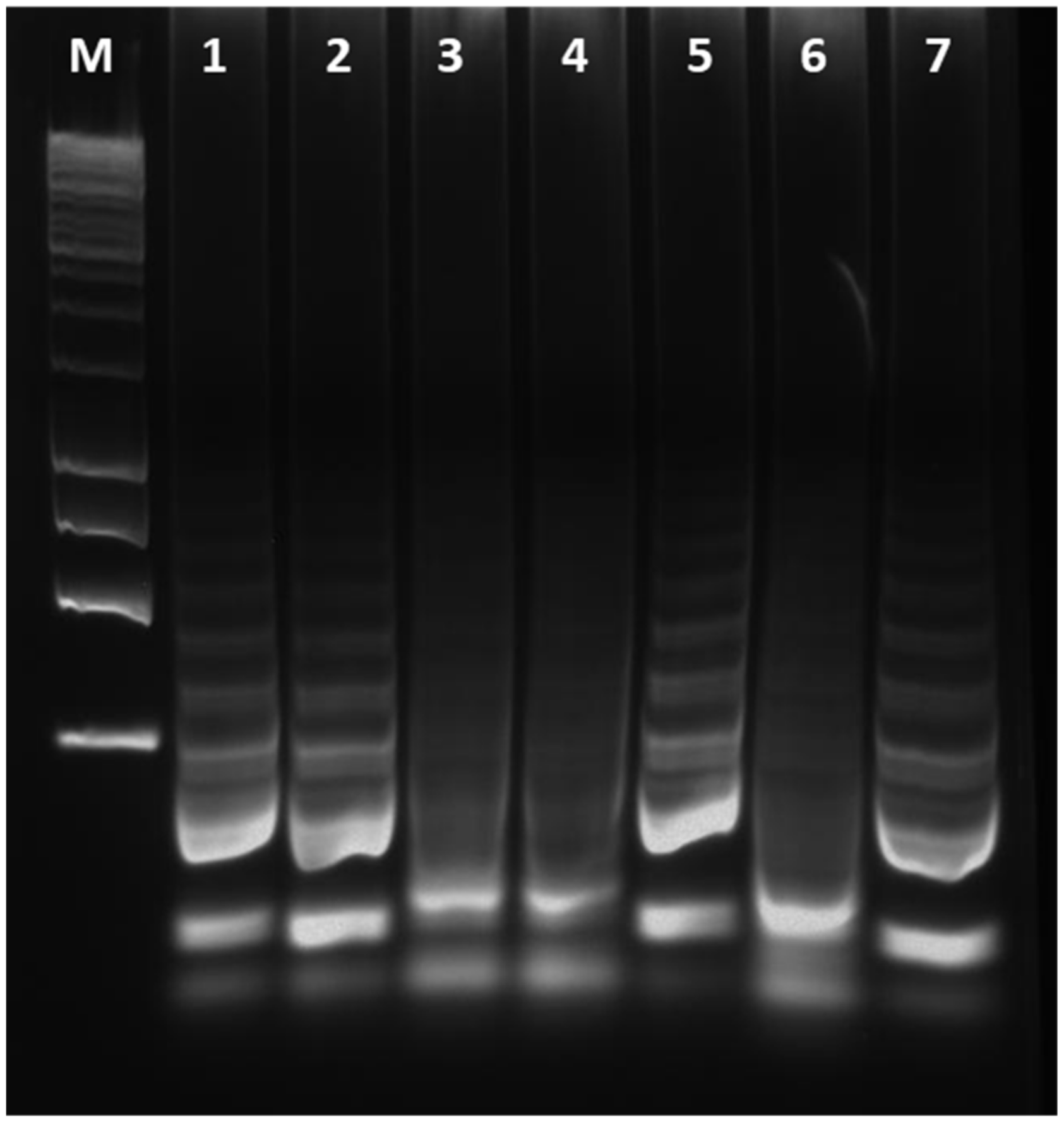
To validate the cLAMP assay, field olive samples were tested both by cLAMP and qPCR assay, considered as a benchmark for Xf detection. The latter test used the extracted DNA as template. Sampled trees were classified according to their Cq values generated in qPCR assay. Xfp infection was confirmed by qPCR in 244 out of 282 samples (Table 1; Supplementary Figure S2). The other 38 samples resulted as undetermined with Cq values > 32 or as negative. The presence of the bacterium was determined by the cLAMP assay in 148 out of 282 samples tested (52.48% along with all the Cq-classified samples). For the 29 out of 38 samples with Cq > 32 and NA in qPCR, the results obtained by cLAMP were properly confirmed as negative (76.31 %). The residual nine samples (23.68%), producing a positive result by cLAMP assay, were considered as false positives according to the statement about qPCR standard. However, a visualization in agarose gels of cLAMP-positive reaction tubes, from samples testing negative in qPCR, shows that a positive amplification is indeed obtained in most of these few ‘yellow-colour’ reactions (Figure 4, lane 5 and 7 vs. lane 6).

Table 1.
Comparative performance of cLAMP vs. qPCR assay for the detection of Xfp in olive alkaline crude extracts. Positive samples obtained by cLAMP are indicated for each group of Cq values obtained from qPCR. Percentage of diagnostic sensitivity and specificity generated by cLAMP assay are reported.
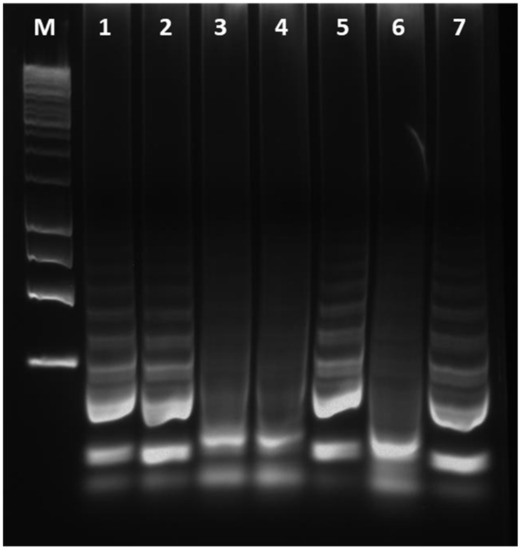
Figure 4.
Electrophoresis of the cLAMP products from field-olive samples. 1–2: samples positive both in qPCR and in cLAMP; 3–4: samples resulted negative in both assays; 5 and 7: positive DNA amplification in samples which produce no amplification in qPCR but a colour change in cLAMP; 6: sample testing negative in qPCR with a yellow-colour change in cLAMP; M: DNA size marker.
As illustrated in Table 1, the cLAMP diagnostic sensitivity was higher if the group of samples producing Cq values < or equal to 23 was considered (ranging from 78.57% for Cq values 18–21 to 82.76% for Cq value 23), while it decreased for samples with Cq values above 23. In addition, for the samples showing Cq values beyond 33, considered undetermined by qPCR, the diagnostic specificity (recognition of truly negative samples) yet remains around 70%. Thus, the comprehensive accuracy calculated for the whole set of samples is 72.21%.
3.4. Evaluation of ddPCR Performance and Comparison with cLAMP
The detection limit of the ddPCR assay was determined to be 169.2 copies/µL using plasmid DNA dilutions, corresponding to 3384 target copies per reaction in a 20 μL total volume (Figure S1). At the first dilution (9 ng/µL; Figure S1, lane A) tested in ddPCR, the Poisson’s algorithm failed the quantification because the copious number of positive droplets saturated the fluorescence signal. As expected, no positive droplets were counted in NIC and NTC. A randomly selected subset of 115 DNA samples from olive trees (already analysed with cLAMP and qPCR) were tested in duplicate with ddPCR, using their crude alkaline extracts as template. A total of 80 out of 105 samples, having a qPCR Cq value below 32, were positive in ddPCR (76.2%), while the other 10 samples gave negative results by ddPCR assay on crude extract. Among these 105 samples, only 55 samples tested positive in cLAMP (52.38%) (Table 2; Figure S3). For ddPCR, the percentage of positive samples was variable when correlated to Cq values, ranging from 90.00% (in the Cq range 19–21) to 77.78% (for Cq 22). However, samples generating Cq values up to 27 still produced 76.92% of positive results in ddPCR. A sudden drop in the number of positive ddPCR results (an average of 40.91% among the different Cq ranges) was then ascertained for those samples with higher Cq values (range 28–32) (Table 2). In only 1 out of the 10 negative qPCR samples (NA) the ddPCR gave a false positive result with more than two droplets counted. In the latter subset of analysis (samples with qPCR with NA result) diagnostic specificity was equal to 90.00%. Among the 81 olive samples which tested positive by ddPCR, 55 of them were also positive by cLAMP (equal to 67.90%) (Table 3 and Figure S4). As expected, the ddPCR assay is able to detect a higher number of positive samples compared to the cLAMP assay. This points out that ddPCR is a more robust method for the detection of Xfp in crude samples since a diagnostic specificity of 90.00% occurred, and the diagnostic sensitivity ranged from 40.00% for samples with Cq values of approx. 28–32 to 100.00% for samples with Cq value of approx. 23.

Table 2.
Comparison of ddPCR and qPCR assays for the Xfp detection in a subset of 115 olive samples. Number of plants testing positive and negative in ddPCR and diagnostic parameters of the assay are reported. The samples are grouped per each Cq value.

Table 3.
Average values (with standard deviation, SD) of positive droplets counted by ddPCR, for olive samples grouped according to similar qPCR Cq values. The number of olive plants positive in the cLAMP assay (and their percentage) are reported, similarly grouped per each Cq value. PA: positive agreement.
3.5. Statistical Data Analysis
Cohen’s kappa statistical analysis was run to determine if there was an agreement between positive detection in cLAMP and qPCR assays. The kappa coefficient was 0.16, which represents a slight strength of agreement, according to the definition and the reference parameters proposed by [42]. Statistical significance was proved at the 0.001 alpha level (p-value < 0.001). When the agreement between qPCR and ddPCR was tested, it was found that the k coefficient rose to 0.29 (p-value < 0.001). Although this value still represents a ‘fair agreement’, according to the standard mentioned earlier, it also proves that there was a higher percentage of statistically supported agreement, between the detection performance in qPCR and ddPCR. The mean value of positive droplets analysed in ddPCR, generated by samples belonging to the same Cq range, is reported (Table 3, Figure S5), as well as the associated standard deviation. This value (number of positive amplification events) is essentially halved from a Cq range group of samples to the next lower one. Pearson’s analysis showed a strong, negative correlation (r = 0.954), significant at the 0.001 alpha level (p-value < 0.001) between average values of positive droplets obtained in ddPCR and Cq values resulting from qPCR assay. Our regression analysis produced the linear equation y = 0.1216x + 4.5665, allowing an estimate of the outcome variable, y (ddPCR positive droplets), based on the predictor variable, x (qPCR Cq values). ROC curves were constructed with results of cLAMP, ddPCR, and qPCR (considered as the reference assay) including both symptomatic and asymptomatic samples. The area under curves values (AUC) for each assay indicated its performance in differentiating Xfp-infected trees from the negative ones in term of sensitivity and specificity. As shown in Figure S6, the AUCs were 0.7316, 0.8196, and 0.9708 for cLAMP, ddPCR, and qPCR, respectively.
4. Discussion
Rapid and reliable detection of Xf is required for timely control measures and to prevent the further spread of the disease in the current pandemic behaviour in olive orchards in Apulia [43]. In our study, the performance of the colorimetric LAMP assay has been set up using, as molecular template for the amplification, a diluted crude sap recovered from olive tissues and a well-validated primer set [24]. This sap, obtained by simple incubation of a tiny amount of olive twigs in an alkaline buffer at room temperature, can easily extract the genomic DNA of the pathogen from the infected tissues. The performance of cLAMP was evaluated on field-grown olive samples in comparison with a qPCR assay [24] included as official test in the EU Reg. 2020/1201. The detection limit of the cLAMP assay was determined to be at femtogram amount of a cloned plasmid containing a Xf-specific gene sequence and at 102 CFU/mL of Xfp bacterial suspension, added to a crude alkaline sap from Xfp-free olive trees. The accuracy of the cLAMP assay, compared to qPCR, was approx. 71%, while the diagnostic sensitivity, calculated considering as positive all the samples producing Cq values up to 32 in the qPCR assay, was 52.54%. The diagnostic specificity of the cLAMP on the truly negative samples is around 78%. Similarly, ddPCR tests were performed using as a template the same crude alkaline sap. We demonstrate that ddPCR, which produced an accuracy of 81.56% vs. qPCR, can also work with a rough template to detect Xfp. ROC curves analysis also confirms this matter. Both qPCR and ddPCR assays can discriminate between positive and non-infected olive samples, as proved by their high AUC values (Figure S6).
Several fluorescent LAMP detection methods were recently published for plant pathogenic bacteria using a crude sap preparation. Yaseen et al. [25] developed a real-time LAMP assay able to detect Xfp in olive plants and insects, but it needed a preliminary heating denaturation step. Kogovsek et al., (2015) and (2017) described a LAMP assay with a low detection limit in the range of 9–27 copies of genomic template for Flavescence Dorèe (FD) and Bois noir (BN) phytoplasma agents in grapevine when using a crude extract. The latter method [21,22], however, needs a thorough homogenization of the plant tissues by a small shaking apparatus. These authors underline that the extraction of the low concentrated DNA of a pathogen is the most critical issue for the application of this approach in field conditions.
The aim of the present work was the development of a colorimetric visualization of the amplification results from rough extracts, so as to allow a user-friendly detection method of the LAMP amplified product, without any apparatus or specialized personnel for the analysis of the results. Some software applications have been described for the quantification of the cLAMP results through colour image decomposition [44,45]. An improvement of the method developed in this study, with the application of a quantitative evaluation of the end-point results, could certainly help in a better definition and discrimination of borderline colour evidence that is not appreciable by the naked-eye inspection. However, our preliminary results in the direction of an unaided colorimetric visualization, while certainly suffering from a merely qualitative and ‘yes-or-no’ diagnostic answer, are substantially able to reveal the pathogen presence roughly in one infected olive tree tested out of two. This is relevant since severe drawbacks are encountered in such kinds of diagnostic effort: (i) the low pathogen titer and erratic distribution in the centennial olive trees induce a long latency period (around 14 months after vector infection; [43]) and delay the symptom expression; (ii) an exhaustive sampling of the olive orchards requires continuous monitoring and a high number of sensitive molecular tests, avoiding costs, machinery-application and time-consuming procedures. The possibility to apply an instrument-free method at the field with a really portable system and an overall cost of about 3 euros per reaction could allow a timely detection of new infection foci and tests of surrounding plants with a minimum effort and a reasonable reliability. We observed a detection rate generated by cLAMP in agreement with the qPCR assay, for samples producing Cq values up to 23. For samples producing Cq values in the range 24–27, it decreased at 50%, still assessing at 33% for Cq 28. Samples having the latter Cq values are generally considered in the case of field-grown olive trees at an early step of Xfp infection and at the limit of any visual inspection for symptom scoring, and therefore likely to be classified as symptomless.
It is conceivable that the variability observed in the number of positive samples detected by cLAMP, including samples with Cq values in the range 18–24, could be mainly due to the kind of crude extraction or/and to the erratic distribution of the bacterium in the small size of sampled tissue. Additionally, at lower target concentrations, the detection of a target sequence could be a variable providing sometimes positive and negative results for the same sample in different analyses, due to a stochastic target copy distribution in the replicates [22]. In this regard, the random amplification results sometimes obtained in cLAMP reactions could be considered not unusual, that is, when target concentration close to detection limit are concerned (as denoted in Figure 3B–D). Similarly, the few discrepancy cases between cLAMP versus qPCR results (Figure 4; lanes 5–7) could be ascribed to the sampling in erratically infected tissues, mostly when dealing with undetermined and negative qPCR results.
As expected, no cross-reactivity was observed with artificial contamination of olive healthy sap with other bacterial species, while the load of epiphytic/endophytic microflora colonizing healthy field-grown olive trees does not seem to affect the expected negative reaction. This indicates that the cLAMP assay applied is highly specific, in agreement with the previous studies [24,25]. The cLAMP allows the achievement of a full result in about 40 min (from the sample preparation to visual observation of the outcome). Conversely, qPCR assay requires a longer time to be carried out (usually 3–4 h in well-equipped laboratory facilities). Notably, the sensitivity and LOD of direct cLAMP assay were not strongly affected using a crude alkaline extraction, which significantly reduces the time and cost of the assay, making cLAMP a valuable alternative for the on-site detection of Xfp. An additional advantage of this method is avoiding the need to move plant material from the contaminated sites to centralized laboratories, thus limiting the risk of pathogen dissemination in Xfp-free areas. Further evaluation of the cLAMP assay performance for large-scale surveys and in insect vectors should be foreseen and addressed in future studies.
5. Conclusions
A protocol for the fast and specific detection of Xfp in olive trees was herein developed. To the best of our knowledge, this is the first study evaluating the cLAMP using crude alkaline extracts for the detection of Xfp from infected olive samples. The advantage cLAMP provides is the possibility of working in field with limited testing facilities and employing rough samples without any DNA extraction and purification procedures. For this reason, this assay has a great potential to evaluate the disease spreading. Therefore, the alkaline extraction and cLAMP protocol developed represent a reliable and alternative approach to real-time qPCR for a timely control strategy of the pathogen.
Supplementary Materials
The following supporting information can be downloaded at: https://www.mdpi.com/article/10.3390/agriculture13020448/s1, Figure S1: Limit of detection of ddPCR by 10-fold dilution of plasmid DNA in the olive alkaline sap. Figure S2: Validation of cLAMP assay on field-grown olive samples by comparison with qPCR; Figure S3: Evaluation of ddPCR performance compared with qPCR; Figure S4: Comparison of the results obtained in cLAMP and ddPCR assays on infected olive alkaline crude extracts according to the qPCR Cq values; Figure S5: Statistical analysis of data correlation (qPCR vs.ddPCR); Figure S6: Receiver Operating Characteristic (ROC) curves for qPCR, ddPCR and cLAMP. Table S1: Optical density of the non-target bacterial species used to assess the cLAMP assay specificity.
Author Contributions
Planning of experimental design, A.M., S.S.A. and G.L.; plant maintenance, S.S.A. and A.L.; preparation of alkaline-crude extracts A.M.; DNA extraction and qPCR assay, A.L.; setting up of cLAMP and ddPCR reactions A.M. and S.S.A.; system development A.K.P. and G.P.; writing—draft preparation, S.S.A. and A.M.; writing—review and editing, A.M., G.L. and E.G.; funding management and project administration, G.L. and E.G. All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by the EU’s Horizon 2020 Project No. 862840 ‘FREE@POC Towards an instrument-FREE future of molecular diagnostics at the Point-Of-Care’.
Institutional Review Board Statement
Not applicable.
Data Availability Statement
Not applicable.
Acknowledgments
The authors express their gratitude to Enza Dongiovanni (CRSFA—Centro di Ricerca, Formazione e Sperimentazione in Agricoltura “Basile Caramia”, Locorotondo, Italy), Vincenzo Cavalieri, Raied Abou Koubaa and Antony Surano (IPSP-CNR Sede secondaria di Bari) for providing olive samples collected in the field monitoring surveys. The critical reading and the suggestions about statistical analysis of the data is kindly acknowledged to Massimiliano Morelli and Carmine del Grosso (IPSP-CNR Sede secondaria di Bari).
Conflicts of Interest
The authors declare no conflict of interest.
References
- Wells, J.M.; Raju, B.C.; Hung, H.-Y.; Weisburg, W.G.; Mandelco-Paul, L.; Brenner, D.J. Xylella Fastidiosa Gen. Nov., Sp. Nov: Gram-Negative, Xylem-Limited, Fastidious Plant Bacteria Related to Xanthomonas Spp. Int. J. Syst. Bacteriol. 1987, 37, 136–143. [Google Scholar] [CrossRef]
- Landa, B.B.; Saponari, M.; Feitosa, O.R., Jr.; Giampetruzzi, A.; Vieira, F.J.D.; Mor, E.; Robatzek, S. Xylella Fastidiosa’s Relationships: The Bacterium, the Host Plants, and the Plant Microbiome. New Phytol. 2022, 234, 1598–1605. [Google Scholar] [CrossRef] [PubMed]
- Hopkins, D.L. Xylella Fastidiosa: Xylem-Limited Bacterial Pathogen of Plants. Annu. Rev. Phytopathol. 1989, 27, 271–290. [Google Scholar] [CrossRef]
- Purcell, A.H.; Hopkins, D.L. Fastidious Xylem-Limited Bacterial Plant Pathogens. Annu. Rev. Phytopathol. 1996, 34, 131–151. [Google Scholar] [CrossRef]
- Hill, B.L.; Purcell, A.M. Populations of Xylella Fastidiosa in Plants Required for Transmission by an Efficient Vector. Phytopathology 1997, 87, 1197–1201. [Google Scholar] [CrossRef]
- Delbianco, A.; Gibin, D.; Pasinato, L.; Morelli, M. Update of the Xylella Spp. Host Plant Database—Systematic Literature Search up to 30 June 2021. EFSA J. 2022, 20, e07039. [Google Scholar] [CrossRef]
- PM 7/24 (4) Xylella Fastidiosa. EPPO Bull. 2019, 49, 175–227. [CrossRef]
- Notomi, T.; Okayama, H.; Masubuchi, H.; Yonekawa, T.; Watanabe, K.; Amino, N.; Hase, T. Loop-mediated isothermal amplification of DNA. Nucleic Acids Res. 2000, 28, e63. [Google Scholar] [CrossRef]
- Tomlinson, J.A.; Ostoja-Starzewska, S.; Webb, K.; Cole, J.; Barnes, A.; Dickinson, M.; Boonham, N. A Loop-Mediated Isothermal Amplification-Based Method for Confirmation of Guignardia Citricarpa in Citrus Black Spot Lesions. Eur. J. Plant Pathol. 2013, 136, 217–224. [Google Scholar] [CrossRef]
- Becherer, L.; Borst, N.; Bakheit, M.; Frischmann, S.; Zengerle, R.; Von Stetten, F. Loop-Mediated Isothermal Amplification (LAMP)-Review and Classification of Methods for Sequence-Specific Detection. Anal. Methods 2020, 12, 717–746. [Google Scholar] [CrossRef]
- Baldi, P.; La Porta, N. Molecular Approaches for Low-Cost Point-of-Care Pathogen Detection in Agriculture and Forestry. Front. Plant Sci. 2020, 11, 570862. [Google Scholar] [CrossRef]
- Gadkar, V.J.; Goldfarb, D.M.; Gantt, S.; Tilley, P.A.G. Real-Time Detection and Monitoring of Loop Mediated Amplification (LAMP) Reaction Using Self-Quenching and De-Quenching Fluorogenic Probes. Sci. Rep. 2018, 8, 5548. [Google Scholar] [CrossRef]
- Wang, D.; Yu, J.; Wang, Y.; Zhang, M.; Li, P.; Liu, M.; Liu, Y. Development of a Real-Time Loop-Mediated Isothermal Amplification (LAMP) Assay and Visual LAMP Assay for Detection of African Swine Fever Virus (ASFV). J. Virol. Methods 2020, 276, 113775. [Google Scholar] [CrossRef]
- Papadakis, G.; Pantazis, A.K.; Fikas, N.; Chatziioannidou, S.; Tsiakalou, V.; Michaelidou, K.; Pogka, V.; Megariti, M.; Vardaki, M.; Giarentis, K.; et al. Portable Real-Time Colorimetric LAMP-Device for Rapid Quantitative Detection of Nucleic Acids in Crude Samples. Sci. Rep. 2022, 12, 3775. [Google Scholar] [CrossRef]
- Cibecchini, G.; Cecere, P.; Tumino, G.; Morcia, C.; Ghizzoni, R.; Carnevali, P.; Terzi, V.; Pompa, P.P. A Fast, Naked-Eye Assay for Varietal Traceability in the Durum Wheat Production Chain. Foods 2020, 9, 1691. [Google Scholar] [CrossRef]
- Drais, M.I.; Maheshwari, Y.; Selvaraj, V.; Varvaro, L.; Yokomi, R.; Djelouah, K. Development and Validation of a Loop-Mediated Isothermal Amplification Technique (LAMP) for the Detection of Spiroplasma Citri, the Causal Agent of Citrus Stubborn Disease. Eur. J. Plant Pathol. 2019, 155, 125–134. [Google Scholar] [CrossRef]
- Stehlíková, D.; Beran, P.; Cohen, S.P.; Čurn, V. Development of Real-Time and Colorimetric Loop Mediated Isothermal Amplification Assay for Detection of Xanthomonas Gardneri. Microorganisms 2020, 8, 1301. [Google Scholar] [CrossRef]
- Bertacca, S.; Caruso, A.G.; Trippa, D.; Marchese, A.; Giovino, A.; Matic, S.; Noris, E.; Ambrosio, M.I.F.S.; Alfaro, A.; Panno, S.; et al. Development of a Real-Time Loop-Mediated Isothermal Amplification Assay for the Rapid Detection of Olea Europaea Geminivirus. Plants 2022, 11, 660. [Google Scholar] [CrossRef]
- Sadallah, A.; Minutillo, S.A.; Valentini, F.; Raimondo, M.L.; Lops, F.; Carlucci, A.; Ippolito, A.; D’Onghia, A.M. A Real Time Loop-Mediated Isothermal Amplification (RealAmp) Assay for Rapid Detection of Pleurostoma Richardsiae in Declining Olive Plants. Phytopathol. Mediterr. 2022, 61, 259–267. [Google Scholar] [CrossRef]
- Hansen, Z.R.; Knaus, B.J.; Tabima, J.F.; Press, C.M.; Judelson, H.S.; Grünwald, N.J.; Smart, C.D. Loop-Mediated Isothermal Amplification for Detection of the Tomato and Potato Late Blight Pathogen, Phytophthora Infestans. J. Appl. Microbiol. 2016, 120, 1010–1020. [Google Scholar] [CrossRef]
- Kogovšek, P.; Hodgetts, J.; Hall, J.; Prezelj, N.; Nikolić, P.; Mehle, N.; Lenarčič, R.; Rotter, A.; Dickinson, M.; Boonham, N.; et al. LAMP Assay and Rapid Sample Preparation Method for On-Site Detection of Flavescence Dorée Phytoplasma in Grapevine. Plant Pathol. 2015, 64, 286–296. [Google Scholar] [CrossRef] [PubMed]
- Kogovšek, P.; Mehle, N.; Pugelj, A.; Jakomin, T.; Schroers, H.J.; Ravnikar, M.; Dermastia, M. Rapid Loop-Mediated Isothermal Amplification Assays for Grapevine Yellows Phytoplasmas on Crude Leaf-Vein Homogenate Has the Same Performance as QPCR. Eur. J. Plant Pathol. 2017, 148, 75–84. [Google Scholar] [CrossRef]
- Matić, S.; Candian, V.; D’Errico, C.; Pierro, R.; Panno, S.; Davino, S.; Noris, E.; Tedeschi, R. In-Field LAMP Detection of Flavescence Dorée Phytoplasma in Crude Extracts of the Scaphoideus Titanus Vector. Agronomy 2022, 12, 1645. [Google Scholar] [CrossRef]
- Harper, S.J.; Ward, L.I.; Clover, G.R.G. Development of LAMP and Real-Time PCR Methods for the Rapid Detection of Xylella Fastidiosa for Quarantine and Field Applications. Phytopathology 2010, 100, 1282–1288. [Google Scholar] [CrossRef] [PubMed]
- Aglietti, C.; Luchi, N.; Pepori, A.L.; Bartolini, P.; Pecori, F.; Raio, A.; Capretti, P.; Santini, A. Real-Time Loop-Mediated Isothermal Amplification: An Early-Warning Tool for Quarantine Plant Pathogen Detection. AMB Express 2019, 9, 50. [Google Scholar] [CrossRef]
- Yaseen, T.; Drago, S.; Valentini, F.; Elbeaino, T.; Stampone, G.; D’Onghia, A.M. On-Site Detection of Xylella Fastidiosa in Host Plants and in “Spy Insects” Using the Real-Time Loop-Mediated Isothermal Amplification Method. Phytopathol. Mediterr. 2015, 54, 488–496. [Google Scholar]
- Elbeaino, T.; Incerti, O.; Dakroub, H.; Valentini, F.; Huang, Q. Development of an FTP-LAMP Assay Based on TaqMan Real-Time PCR and LAMP for the Specific Detection of Xylella Fastidiosa De Donno and Mulberry Strains in Both Plants and Insect Vectors. J. Microbiol. Methods 2020, 175, 105992. [Google Scholar] [CrossRef]
- Kim, S.; Park, Y.; Kim, G. Development of Diagnostic Technology of Xylella Fastidiosa Using Loop-Mediated Isothermal Amplification and PCR Methods. Res. Plant Dis. 2021, 27, 38–44. [Google Scholar] [CrossRef]
- Waliullah, S.; Di Genova, D.; Oliver, J.E.; Ali, M.E. Development of a CAPS Marker and a LAMP Assay for Rapid Detection of Xylella Fastidiosa Subsp. Multiplex and Differentiation from X. Fastidiosa Subsp. Fastidiosa on BlueBerry. Int. J. Mol. Sci. 2022, 23, 1937. [Google Scholar] [CrossRef]
- Chiriacò, M.S.; Luvisi, A.; Primiceri, E.; Sabella, E.; De Bellis, L.; Maruccio, G. Development of a Lab-on-a-Chip Method for Rapid Assay of Xylella Fastidiosa Subsp. Pauca Strain CoDiRO. Sci. Rep. 2018, 8, 7376. [Google Scholar] [CrossRef]
- Sarcina, L.; Macchia, E.; Loconsole, G.; D’Attoma, G.; Bollella, P.; Catacchio, M.; Leonetti, F.; Di Franco, C.; Elicio, V.; Scamarcio, G.; et al. Fast and Reliable Electronic Assay of a Xylella Fastidiosa Single Bacterium in Infected Plants Sap. Adv. Sci. 2022, 9, 2203900. [Google Scholar] [CrossRef]
- Dupas, E.; Legendre, B.; Olivier, V.; Poliakoff, F.; Manceau, C.; Cunty, A. Comparison of Real-Time PCR and Droplet Digital PCR for the Detection of Xylella Fastidiosa in Plants. J. Microbiol. Methods 2019, 162, 86–95. [Google Scholar] [CrossRef]
- Saponari, M.; Boscia, D.; Nigro, F.; Martelli, G.P. Identification Of Dna Sequences Related To Xylella Fastidiosa In Oleander, Almond And Olive Trees Exhibiting Leaf Scorch Symptoms In Apulia (Southern Italy). J. Plant Pathol. 2013, 95, 668. [Google Scholar] [CrossRef]
- Giampetruzzi, A.; Saponari, M.; Almeida, R.P.P.; Essakhi, S.; Boscia, D.; Loconsole, G.; Saldarelli, P. Complete Genome Sequence of the Olive-Infecting Strain Xylella Fastidiosa Subsp. Pauca De Donno. Genome Announc. 2017, 5, e00569-17. [Google Scholar] [CrossRef]
- D’attoma, G.; Morelli, M.; Saldarelli, P.; Saponari, M.; Giampetruzzi, A.; Boscia, D.; Savino, V.N.; Fuente, L.D.L.; Cobine, P.A. Ionomic Differences between Susceptible and Resistant Olive Cultivars Infected by Xylella Fastidiosa in the Outbreak Area of Salento, Italy. Pathogens 2019, 8, 272. [Google Scholar] [CrossRef]
- Baccari, C.; Antonova, E.; Lindow, S. Biological Control of Pierce’s Disease of Grape by an Endophytic Bacterium. Phytopathology 2019, 109, 248–256. [Google Scholar] [CrossRef]
- Sun, X.; Ma, W.; Xu, Y.; Jin, X.; Ni, H. Complete Genome Sequence of Rahnella Aquatilis MEM40, a Plant Growth-Promoting Rhizobacterium Isolated from Rice Rhizosphere Soil, with Antagonism against Magnaporthe Oryzae and Fusarium Graminearum. Microbiol. Resour. Announc. 2020, 9, e00651-20. [Google Scholar] [CrossRef]
- Loconsole, G.; Potere, O.; Boscia, D.; Altamura, G.; Djelouah, K.; Elbeaino, T.; Frasheri, D.; Lorusso, D.; Palmisano, F.; Pollastro, P.; et al. Detection of Xylella Fastidiosa in Olive Trees By Molecular And Serological Methods. J. Plant Pathol. 2014, 96, 7–14. [Google Scholar] [CrossRef]
- PM 7/76 (5) Use of EPPO Diagnostic Standards. EPPO Bull. 2018, 48, 373–377. [CrossRef]
- Landis, J.R.; Koch, G.G. The Measurement of Observer Agreement for Categorical Data. Biometrics 1977, 33, 159–174. [Google Scholar] [CrossRef]
- Cleophas, T.J.; Zwinderman, A.H. Modern Bayesian Statistics in Clinical Research; Springer International Publishing: Cham, Switzerland, 2018; ISBN 978-3-319-92746-6. [Google Scholar]
- Altman, D.G. Practical Statistics for Medical Research; Chapman & Hall/CRC Press: New York, NY, USA, 1999. [Google Scholar]
- Saponari, M.; Giampetruzzi, A.; Loconsole, G.; Boscia, D.; Saldarelli, P. Xylella Fastidiosa in Olive in Apulia: Where We Stand. Phytopathology 2019, 109, 175–186. [Google Scholar] [CrossRef] [PubMed]
- González-González, E.; Lara-Mayorga, I.M.; Rodríguez-Sánchez, I.P.; Zhang, Y.S.; Martínez-Chapa, S.O.; Santiago, G.T.; Alvarez, M.M. Colorimetric Loop-Mediated Isothermal Amplification (LAMP) for Cost-Effective and Quantitative Detection of SARS-CoV-2: The Change in Color in LAMP-Based Assays Quantitatively Correlates with Viral Copy Number. Anal. Methods Adv. Methods Appl. 2021, 13, 169–178. [Google Scholar] [CrossRef] [PubMed]
- García-Bernalt Diego, J.; Fernández-Soto, P.; Márquez-Sánchez, S.; Santos Santos, D.; Febrer-Sendra, B.; Crego-Vicente, B.; Muñoz-Bellido, J.L.; Belhassen-García, M.; Corchado Rodríguez, J.M.; Muro, A. SMART-LAMP: A Smartphone-Operated Handheld Device for Real-Time Colorimetric Point-of-Care Diagnosis of Infectious Diseases via Loop-Mediated Isothermal Amplification. Biosensors 2022, 12, 424. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).