A Convenient Co-Dominant Marker for Height-Reducing Ddw1 Allele Useful for Marker-Assisted Selection
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Material
2.2. DNA Isolation and PCR Amplification of REMS1218 Microsatellite Fragment
2.3. Cloning and Sequencing REMS1218 PCR Products
2.4. CAPS Analysis of REMS1218
2.5. Rht-B1b Genotyping
2.6. Statistical Analysis
3. Results
3.1. PCR Amplification of Microsatellite REMS1218 Marker Results in Several Product Patterns
3.2. Cloning and Sequencing of REMS1218 Amplicons Revealed a Common 257 bp Sequence with Six Polymorphic Nucleotides
3.3. Selection and Verification of a Set of Restrictases for CAPS Analysis of Ddw1 Alleles
3.4. Comparison of CAPS Marker with Capillary Electrophoresis Analysis of REMS1218 PCR Products on 91 F2 Crosses of Khongor and Dublet
3.5. Association of Plant Height and Ddw1 Genotype Determined by Ddw1 CAPS Marker in Two Segregating Populations Obtained from Four Parent Cultivars
4. Discussion
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Ayalew, H.; Kumssa, T.T.; Butler, T.J.; Ma, X.-F. Triticale Improvement for Forage and Cover Crop Uses in the Southern Great Plains of the United States. Front. Plant Sci. 2018, 9, 1130. [Google Scholar] [CrossRef] [PubMed]
- Ammar, K.; Mergoum, M.; Rajaram, S. The history and evolution of triticale. In Triticale Improvement and Production. FAO Plant Production and Protection Paper No. 179; Mergoum, M., Gómez-Macpherson, H., Eds.; Food and Agriculture Organization of United Nations: Rome, Italy, 2004; pp. 1–9. [Google Scholar]
- Mergoum, M.; Pfeiffer, W.H.; Peña, R.J.; Ammar, K.; Rajaram, S. Triticale crop improvement: the CIMMYT programme. In Triticale Improvement and Production. FAO Plant Production and Protection Paper No. 179; Mergoum, M., Gómez-Macpherson, H., Eds.; Food and Agriculture Organization of United Nations: Rome, Italy, 2004; pp. 11–26. [Google Scholar]
- Van De Velde, K.; Ruelens, P.; Geuten, K.; Rohde, A.; Van Der Straeten, D. Exploiting DELLA Signaling in Cereals. Trends Plant Sci. 2017, 22, 880–893. [Google Scholar] [CrossRef] [PubMed]
- Korshunova, A.D.; Divashuk, M.G.; Soloviev, A.; Karlov, G.I. Analysis of wheat and rye semidwarfing gene distribution in spring hexaploid triticale (Triticosecale Wittm.) cultivars and breeding lines. Russ. J. Genet. 2015, 51, 272–277. [Google Scholar] [CrossRef]
- Kobyliansky, V.D. On genetics of the dominant factor of short-strawed rye. Genetika 1972, 8, 12–17. [Google Scholar]
- Braun, E.-M.; Tsvetkova, N.; Rotter, B.; Siekmann, D.; Schwefel, K.; Krezdorn, N.; Plieske, J.; Winter, P.; Melz, G.; Voylokov, A.V.; et al. Gene Expression Profiling and Fine Mapping Identifies a Gibberellin 2-Oxidase Gene Co-segregating With the Dominant Dwarfing Gene Ddw1 in Rye (Secale Cereale L.). Front. Plant Sci. 2019, 10, 857. [Google Scholar] [CrossRef] [PubMed]
- Kroupin, P.Y.; Chernook, A.; Karlov, G.I.; Soloviev, A.; Divashuk, M.G. Effect of Dwarfing Gene Ddw1 on Height and Agronomic Traits in Spring Triticale in Greenhouse and Field Experiments in a Non-Black Earth Region of Russia. Plants 2019, 8, 131. [Google Scholar] [CrossRef] [PubMed]
- Kroupin, P.; Chernook, A.; Karlov, G.I.; Soloviev, A.; Korshunova, A.D.; Divashuk, M.G. Effects of Dwarfing Wheat (Triticum Aestivum L.) and rye (Secale Cereale L.) genes in spring triticale segregating population as studied in pot trials. Sel’skokhozyaistvennaya Boil. 2019, 54, 920–933. [Google Scholar] [CrossRef]
- Chernook, A.; Kroupin, P.Y.; Karlov, G.I.; Soloviev, A.; Korshunova, A.D.; Rubets, V.; Igonin, V.; Divashuk, M.G. Effects of Rht-B1b and Ddw1 Dwarfing Genes in Two Connecting Populations of Spring Triticale under Greenhouse Experiment Conditions. Agriculture 2019, 9, 119. [Google Scholar] [CrossRef]
- Khlestkina, E.; Than, M.H.M.; Pestsova, E.; Malyshev, S.V.; Korzun, V. Mapping of 99 new microsatellite-derived loci in rye (Secale Cereale L.) including 39 expressed sequence tags. Theor. Appl. Genet. 2004, 109, 725–732. [Google Scholar] [CrossRef] [PubMed]
- Korzun, V.; Börner, A.; Melz, G. RFLP mapping of the dwarfing (Ddw1) and hairy peduncle (Hp) genes on chromosome 5 of rye (Secale Cereale L.). Theor. Appl. Genet. 1996, 92, 1073–1077. [Google Scholar] [CrossRef] [PubMed]
- Tenhola-Roininen, T.; Tanhuanpää, P. Tagging the dwarfing gene Ddw1 in a rye population derived from doubled haploid parents. Euphytica 2009, 172, 303–312. [Google Scholar] [CrossRef]
- Murray, M.; Thompson, W.F. Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res. 1980, 8, 4321–4326. [Google Scholar] [CrossRef] [PubMed]
- Nicholas, K.B.; Nicholas, H.B., Jr.; Deerfield, D.W., II. GeneDoc: Analysis and visualization of genetic variation. EMB Net. News 1997, 4, 1–4. [Google Scholar]
- Ellis, M.H.; Spielmeyer, W.; Gale, K.; Rebetzke, G.J.; Richards, R.A. “Perfect” markers for the Rht-B1b and Rht-D1b dwarfing genes in wheat. Theor. Appl. Genet. 2002, 105, 1038–1042. [Google Scholar] [CrossRef] [PubMed]


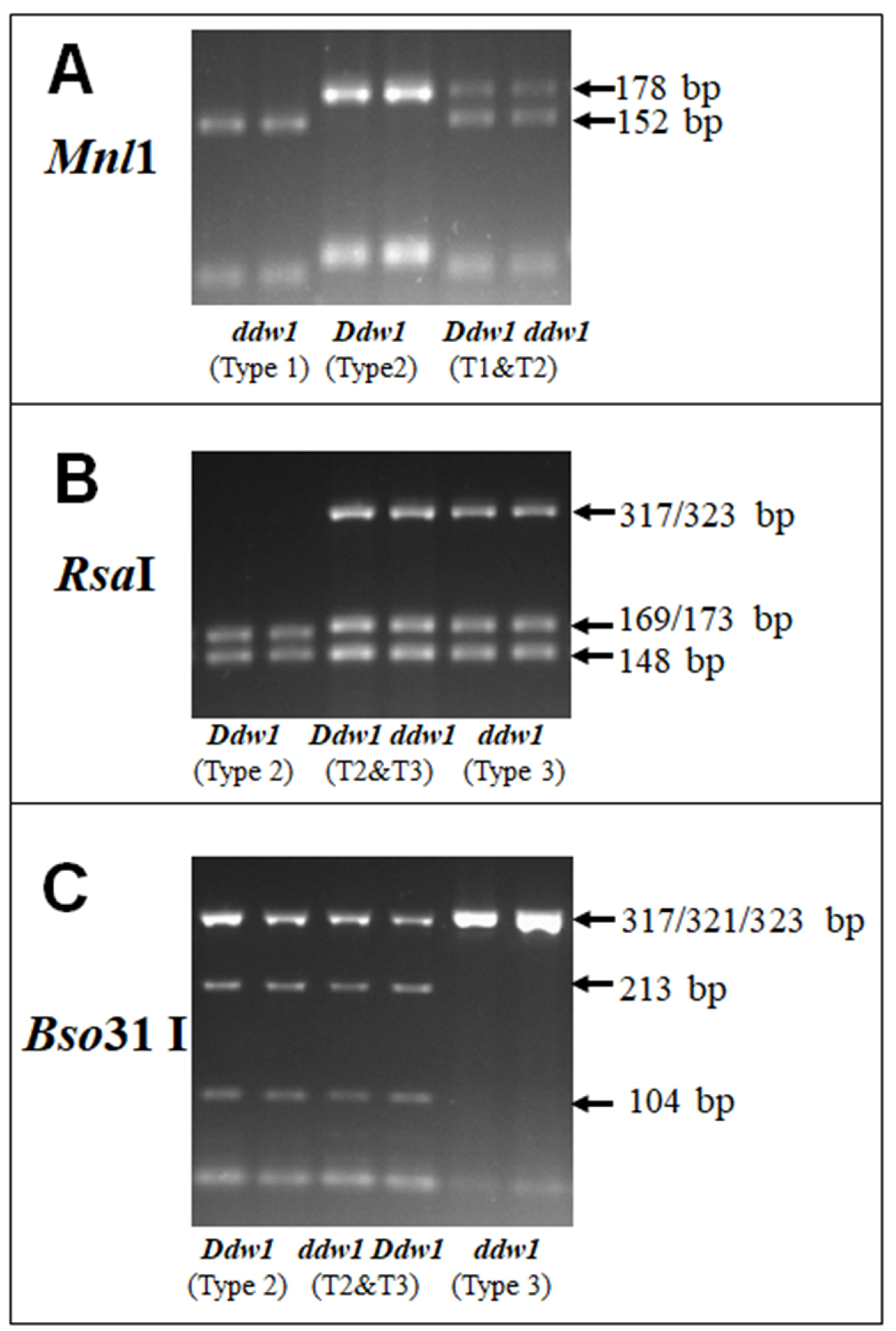
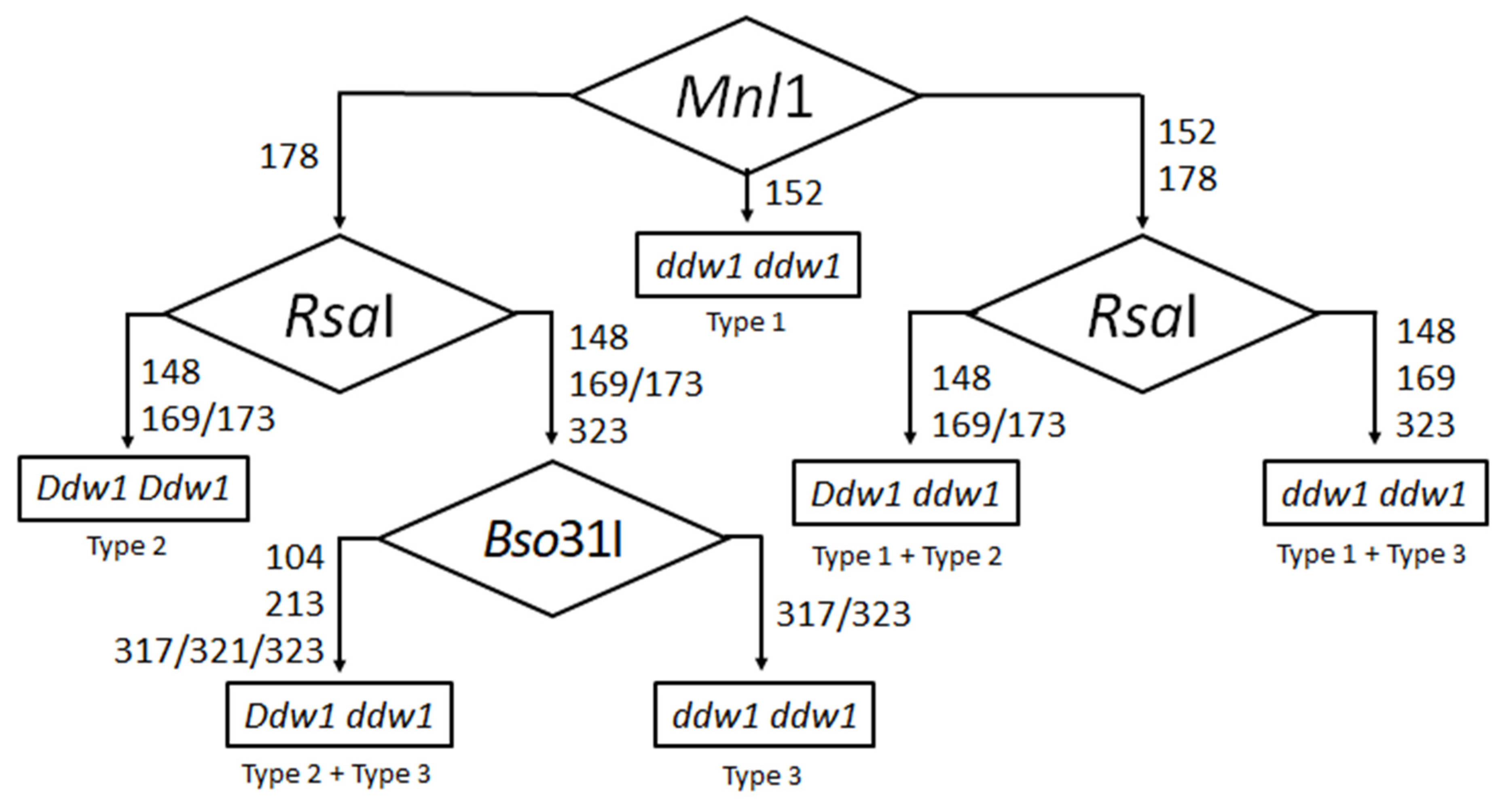
| Type | Product Size, bp | Fragment Sizes (bp) after Digestions by: | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| MnlI | RsaI | Bso31I | |||||||||
| T1a (ddw1) | 317 | 26 | 152 | 15 | 44 | 14 | 66 | 148 | 169 | 213 | 104 |
| T1b (ddw1) | 317 | 26 | 152 | 15 | 44 | 14 | 66 | 148 | 169 | 213 | 104 |
| T2a (Ddw1) | 321 | 178 | 15 | 44 | 14 | 70 | 148 | 173 | 321 | ||
| T2b (Ddw1) | 317 | 178 | 15 | 44 | 14 | 66 | 148 | 169 | 213 | 104 | |
| T3a (ddw1) | 321 | 178 | 15 | 44 | 14 | 72 | 323 | 323 | |||
| T3b (ddw1) | 317 | 178 | 15 | 44 | 14 | 66 | 148 | 169 | 317 | ||
| Ddw1 Genotype | Height, cm |
|---|---|
| A/SKh plants | |
| ddw1 ddw1 (26 plants) | 101.7 ± 4.6 |
| Ddw1 ddw1 (29 plants) | 89.1 ± 4.1 |
| Ddw1 Ddw1 (22 plants) | 82.8 ± 3.4 * |
| Kh/D plants | |
| ddw1 ddw1 (13 plants) | 101.9 ± 4.6 |
| Ddw1 ddw1 (57 plants) | 83.4 ± 2.2 ** |
| Ddw1 Ddw1 (21 plants) | 74.9 ± 3.6 *** |
| Parent Line, Crossed with Ddw1 (Type 2) | Restrictase(s) for the CAPS Analysis | Restriction Product(s) Size, bp | Genotype |
|---|---|---|---|
| ddw1 Type 1 317 bp | Mnl1 | 152 | Homozygous Type 1 (ddw1 ddw1) |
| 152 + 178 | Heterozygous Types 1 & 2 (Ddw1 ddw1) | ||
| 178 | Homozygous Type 2 (Ddw1 Ddw1) | ||
| ddw1 Type 3 317 + 323 bp | RsaI, Bso31I | RsaI: 148 + 169/173 Bso31I: 104 + 213 + 321 | Homozygous Type 2 (Ddw1 Ddw1) |
| RsaI: 148 + 169/173 + 323 Bso31I: 104 +213 + 317/321/323 | Heterozygous Types 2 & 3 (Ddw1 ddw1) | ||
| RsaI: 148 + 169 + 323 Bso31I: 317/323 | Homozygous Type 3 (ddw1 ddw1) |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Litvinov, D.Y.; Chernook, A.G.; Kroupin, P.Y.; Bazhenov, M.S.; Karlov, G.I.; Avdeev, S.M.; Divashuk, M.G. A Convenient Co-Dominant Marker for Height-Reducing Ddw1 Allele Useful for Marker-Assisted Selection. Agriculture 2020, 10, 110. https://doi.org/10.3390/agriculture10040110
Litvinov DY, Chernook AG, Kroupin PY, Bazhenov MS, Karlov GI, Avdeev SM, Divashuk MG. A Convenient Co-Dominant Marker for Height-Reducing Ddw1 Allele Useful for Marker-Assisted Selection. Agriculture. 2020; 10(4):110. https://doi.org/10.3390/agriculture10040110
Chicago/Turabian StyleLitvinov, Dmitry Y., Anastasiya G. Chernook, Pavel Yu. Kroupin, Mikhail S. Bazhenov, Gennady I. Karlov, Sergey M. Avdeev, and Mikhail G. Divashuk. 2020. "A Convenient Co-Dominant Marker for Height-Reducing Ddw1 Allele Useful for Marker-Assisted Selection" Agriculture 10, no. 4: 110. https://doi.org/10.3390/agriculture10040110
APA StyleLitvinov, D. Y., Chernook, A. G., Kroupin, P. Y., Bazhenov, M. S., Karlov, G. I., Avdeev, S. M., & Divashuk, M. G. (2020). A Convenient Co-Dominant Marker for Height-Reducing Ddw1 Allele Useful for Marker-Assisted Selection. Agriculture, 10(4), 110. https://doi.org/10.3390/agriculture10040110





