Confocal Laser Microscopy in Neurosurgery: State of the Art of Actual Clinical Applications
Abstract
1. Introduction
2. Materials and Methods
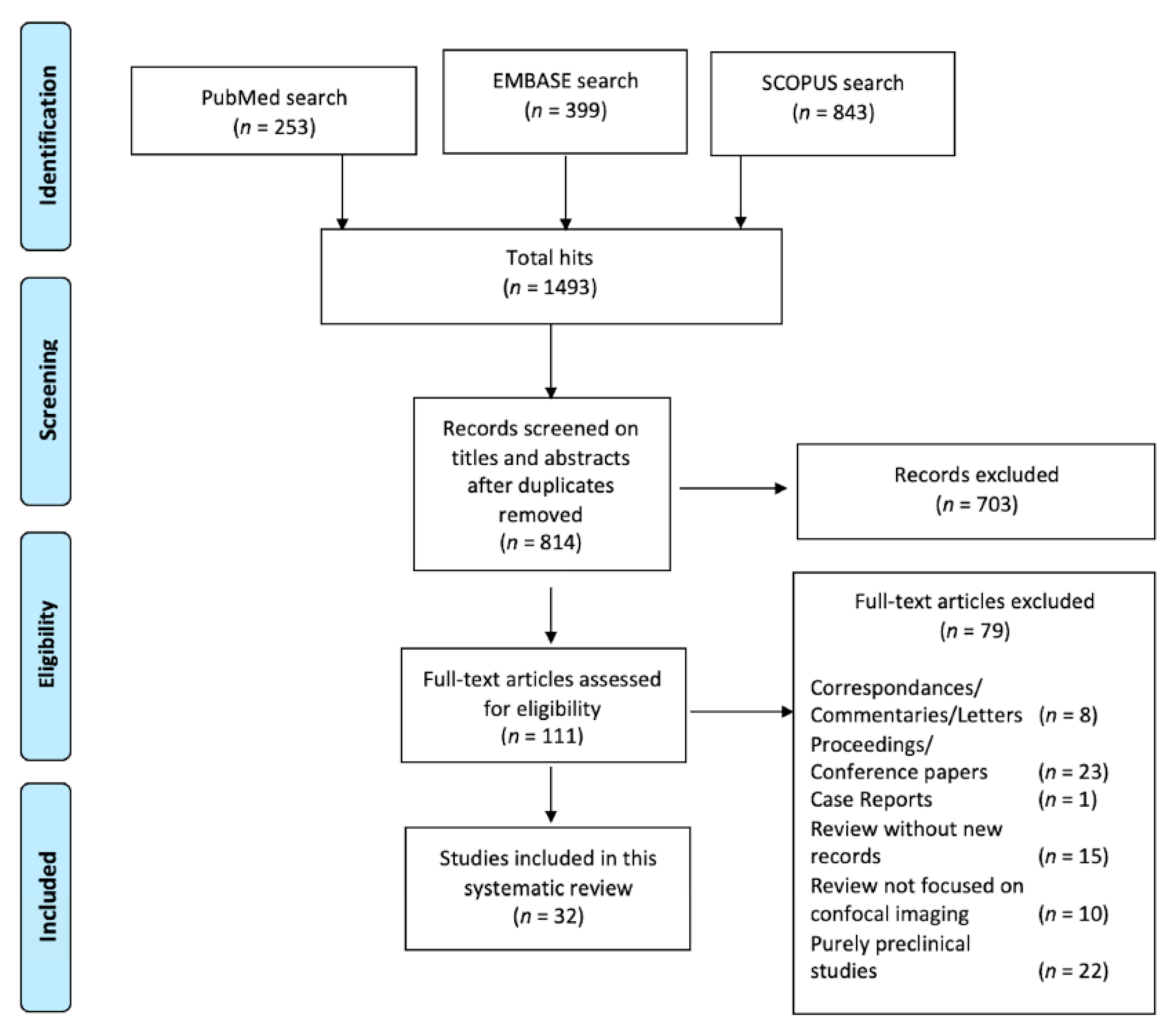
2.1. Literature Search and Screening Process
2.2. Eligibility Criteria
- -
- Clinical works focused on confocal imaging technology application in neurosurgery.
- -
- Preclinical works with a subsection on clinical application of confocal imaging technology in neurosurgery.
- -
- Correspondences.
- -
- Comments.
- -
- Letters to the editor.
- -
- Proceedings/conference papers.
- -
- Case reports.
- -
- Reviews.
- -
- Purely preclinical studies.
2.3. Data Extraction
2.4. Statistical Analysis
3. Results
3.1. In Vivo Experiences
3.2. Ex Vivo Experiences
3.3. Contrast Enhancers Used during In Vivo and Ex Vivo CLE Imaging
3.4. Summary on Sensibility, Specificity and Diagnostic Accuracy of Confocal Imaging Technology in Brain Tumors Surgery
3.5. Descriptive Confocal Imaging Patterns of Normal Brain and Different Brain Tumors
3.5.1. Normal Brain
3.5.2. HGGs
3.5.3. LGGs
3.5.4. Ependimomas and Subependimomas
3.5.5. Meningiomas
3.5.6. Schwannomas
3.5.7. Craniopharyngiomas
3.5.8. Pituitary Adenomas
3.5.9. Central Neurocytoma
3.5.10. Paraganglioma
3.5.11. Hemangioblastoma
3.5.12. Metastases
3.5.13. Epidermoid Tumors
3.5.14. Choroid Plexus Papilloma
3.5.15. Plasmacytoma
3.5.16. Confocal Imaging in Non-CNS Tumors
3.6. The Clinical Role of Machine Learning in Confocal Imaging: A New Frontier for More Interpretable Images
4. Summary
- Confocal imaging may be used similarly to a frozen section analysis: to obtain an intraoperative differential diagnosis. Looking at this specific purpose, scarce but pertinent results are available in neurosurgical literature, although data should be considered and interpreted in light of two aspects: the “blindness grade” of each study and the place where the hypothetical diagnosis was made (i.e., in OR or in the lab or in office), to check for its possible role to be implemented in routine clinical practice.For instance, in the work of Snuderl the bioptic samples were transferred to another institute to be studied on a benchtop microscope and, although the design of the study was built to simulate an intraoperative frozen section evaluation process, testing sensitivity and specificity, the real design of the study was far from being possibly integrated in a neurosurgical OR [28]. Foersch in 2012 used Optiscan (now dismissed) to make diagnosis using a benchtop confocal microscope [26]. Additionally, in this case, although raters were blinded to histology and macroscopic appearance of the tumor, diagnosis was not given in the OR, during neurosurgical procedure. Both Daali and Breuskin reported interesting results regarding sensibility/specificity of this technology in obtaining intraoperative diagnosis, with an acceptable grade of blindness of pathologists. No sensibility/specificity data were given in both studies regarding morphological characterization of different tumors [19,33]. Similarly, no morphological quantitative characterization was carried out by Eschbacher in 2017, where pathologists were aware of eventual contrast enhancement and location of tumors analyzed (lower grade of blindness) [32]. In this context, the work of Belykh in 2020 should be considered as a miliary stone, due to the extremely rigorous blinded study in analyzing CLE images from both neuropathologist and experienced/unexperienced neurosurgeons’ points of view and the huge amount of tumors investigated [41]. Looking at our work, in 2020 we reported the results of a rigorous blinded and prospective study where an optimal diagnostic concordance was obtained at the central core of 15 HGGs analyzed [42]. Even though one of the major limits of this study was the fact that just one hystotipe of tumor was analyzed, hence rendering it unsuitable to study the differential diagnosis ability of Convivo (programmed for the next in vivo study), we reported an optimal concordance among CLE imaging and histopathological/frozen section analysis at the central core of the different cases, with a rigorous blinded design. Moreover, we performed for the first time a different analysis for specimens taken at central core or at tumor margins (see the second point of this discussion). In conclusion, still more data on the real possibility of obtaining prospectively and blindly an intraoperative differential diagnosis using confocal imaging are necessary and awaited.
- Confocal imaging may be used to check for the presence of tumor at the tumor margins, possibly increasing EOR. As stated before, from a methodological point of view, a real sensibility and specificity calculation would be possible only for pure in vivo experiences, given the need of optical biopsies on healthy brain parenchyma (negatives, see above). To date, the only solid statistic study that prospectively analyzed in vivo sensibility and specificity of CLE imaging obtained at both tumor central core and margins was the one of Martirosyan in 2016 [11]. Nevertheless, in this study no specific statistic differentiation was made among specimens taken at central core and at transition zones. Hence, a real ability in identifying tumor tissue at borders was not investigated. In addition, considering ex vivo studies, a very paucity of data regarding biopsies taken at tumor margins is present. As anticipated, our work in 2020 was the first prospective and blinded ex vivo study to specifically check for ability of Convivo system to identify tumor tissue at borders in HGGs [42]. Comparing Convivo and frozen section analyses, we obtained a high rate of concordance at tumor borders in both obtaining a diagnosis and categorizing morphological patterns. Unfortunately, we were not able to extend this study also to meningiomas, metastasis and other tumor types, reason for which a prospective, blinded in vivo study using Convivo analyzing different CNS tumors at both central core and borders is ongoing in our Institute. Considering the paucity of data regarding this specific issue, further studies in this field are awaited to discover how sensitive and specific such systems may be when used to check for tumor tissue at borders.
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Wen, P.Y.; Kesari, S. Malignant gliomas in adults. N. Engl. J. Med. 2008, 359, 492–507. [Google Scholar] [CrossRef]
- Bondy, M.L.; Scheurer, M.E.; Malmer, B.; Barnholtz-Sloan, J.S.; Davis, F.G.; Il’yasova, D.; Kruchko, C.; McCarthy, B.J.; Rajaraman, P.; Schwartzbaum, J.A.; et al. Brain tumor epidemiology: Consensus from the Brain Tumor Epidemiology Consortium. Cancer 2008, 113, 1953–1968. [Google Scholar] [CrossRef] [PubMed]
- Stupp, R.; Mason, W.P.; van den Bent, M.J.; Weller, M.; Fisher, B.; Taphoorn, M.J.; Belanger, K.; Brandes, A.A.; Marosi, C.; Bogdahn, U. Radiotherapy plus concomitant and adjuvant temozolomide for glioblastoma. N. Engl. J. Med. 2005, 352, 987–996. [Google Scholar] [CrossRef]
- Stupp, R.; Hegi, M.E.; Mason, W.P.; van den Bent, M.J.; Taphoorn, M.J.; Janzer, R.C.; Ludwin, S.K.; Allgeier, A.; Fisher, B.; Belanger, K.; et al. Effects of radiotherapy with concomitant and adjuvant temozolomide versus radiotherapy alone on survival in glioblastoma in a randomised phase III study: 5-year analysis of the EORTC-NCIC trial. Lancet Oncol. 2009, 10, 459–466. [Google Scholar] [CrossRef]
- Kuhnt, D.; Becker, A.; Ganslandt, O.; Bauer, M.; Buchfelder, M.; Nimsky, C. Correlation of the extent of tumor volume resection and patient survival in surgery of glioblastoma multiforme with high-field intraoperative MRI guidance. Neuro-Oncology 2011, 13, 1339–1348. [Google Scholar] [CrossRef] [PubMed]
- Acerbi, F.; Broggi, M.; Schebesch, K.-M.; Höhne, J.; Cavallo, C.; De Laurentis, C.; Eoli, M.; Anghileri, E.; Servida, M.; Boffano, C.; et al. Fluorescein-guided surgery for resection of high-grade gliomas: A multicentric prospective phase II study (FLUOGLIO). Clin. Cancer Res. 2018, 24, 52–61. [Google Scholar] [CrossRef]
- Janson, M.; Michael, K.; Berg, J.; Anderson, J.C. The role of intraoperative sonography in neurosurgery. J. Diagn. Med. Sonogr. 2005, 21, 148–151. [Google Scholar] [CrossRef]
- Lara-Velazquez, M.; Al-Kharboosh, R.; Jeanneret, S.; Vazquez-Ramos, C.; Mahato, D.; Tavanaiepour, D.; Rahmathulla, G.; Quinone-Hinojosa, A. Advances in brain tumor surgery for glioblastoma in adults. Brain Sci. 2017, 7, 166. [Google Scholar] [CrossRef]
- Chatterjee, S. Artefacts in histopathology. J. Oral Maxillofac. Pathol. 2014, 18, S111–S116. [Google Scholar] [CrossRef]
- Plesec, T.P.; Prayson, R.A. Frozen section discrepancy in the evaluation of central nervous system tumors. Arch. Pathol. Lab. Med. 2007, 131, 1532–1540. [Google Scholar] [CrossRef]
- Martirosyan, N.L.; Eschbacher, J.M.; Yashar, M.; Turner, J.D.; Belykh, E.; Spetzler, R.F.; Nakaji, P.; Preul, M.C.; Kalani, M.Y.; Turner, J.D.; et al. Prospective evaluation of the utility of intraoperative confocal laser endomicroscopy in patients with brain neoplasms using fluorescein sodium: Experience with 74 cases. Neurosurg. Focus 2016, 40, E11. [Google Scholar] [CrossRef]
- Dunbar, K.B.; Okolo, P.; Montgomery, E.; Canto, M.I. Confocal laser endomicroscopy in Barrett’s esophagus and endoscopically inapparent Barrett’s neoplasia: A prospective, randomized, double-blind, controlled, crossover trial. Gastrointest. Endosc. 2009, 70, 645–654. [Google Scholar] [CrossRef]
- Tan, J.; Quinn, M.A.; Pyman, J.M.; Delaney, P.M.; McLaren, W.J. Detection of cervical intraepithelial neoplasia in vivo using confocal endomicroscopy. BJOG Int. J. Obstet. Gynaecol. 2009, 116, 1663–1670. [Google Scholar] [CrossRef] [PubMed]
- Martirosyan, N.L.; Georges, J.; Eschbacher, J.M.; Cavalcanti, D.D.; Elhadi, A.M.; Abdelwahab, M.G.; Scheck, A.C.; Nakaji, P.; Spetzler, R.F.; Preul, M.C. Potential application of a handheld confocal endomicroscope imaging system using a variety of fluorophores in experimental gliomas and normal brain. Neurosurg. Focus 2014, 36, E16. [Google Scholar] [CrossRef]
- Martirosyan, N.L.; Georges, J.; Kalani, M.Y.S.; Nakaji, P.; Spetzler, R.F.; Feuerstein, B.G.; Preul, M.C. Handheld confocal laser endomicroscopic imaging utilizing tumor-specific fluorescent labeling to identify experimental glioma cells in vivo. Surg. Neurol. Int. 2016, 7, S995–S1003. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Sankar, T.; Delaney, P.M.; Ryan, R.W.; Eschbacher, J.; Abdelwahab, M.; Nakaji, P.; Coons, S.W.; Scheck, A.C.; Smith, K.A.; Spetzler, R.F.; et al. Miniaturized handheld confocal microscopy for neurosurgery: Results in an experimental glioblastoma model. Neurosurgery 2010, 66, 410–417. [Google Scholar] [CrossRef]
- Pavlov, V.; Meyronet, D.; Meyer-Bisch, V.; Armoiry, X.; Pikul, B.; Dumot, C.; Beuriat, P.A.P.-A.; Signorelli, F.; Guyotat, J. Intraoperative Probe-Based Confocal Laser Endomicroscopy in Surgery and Stereotactic Biopsy of Low-Grade and High-Grade Gliomas: A Feasibility Study in Humans. Neurosurgery 2016, 79, 604–611. [Google Scholar] [CrossRef] [PubMed]
- Charalampaki, P.; Javed, M.; Daali, S.; Heiroth, H.-J.; Igressa, A.; Weber, F. Confocal laser endomicroscopy for real-time histomorphological diagnosis: Our clinical experience with 150 brain and spinal tumor cases. Neurosurgery 2015, 62, 171–176. [Google Scholar] [CrossRef]
- Breuskin, D.; Szczygielski, J.; Urbschat, S.; Kim, Y.; Oertel, J. Confocal Laser Endomicroscopy in Neurosurgery-An Alternative to Instantaneous Sections? World Neurosurg. 2017, 100, 180–185. [Google Scholar] [CrossRef]
- Sanai, N.; Eschbacher, J.; Hattendorf, G.; Coons, S.W.; Preul, M.C.; Smith, K.A.; Nakaji, P.; Spetzler, R.F. Intraoperative confocal microscopy for brain tumors: A feasibility analysis in humans. Neurosurgery 2011, 68, 282–289. [Google Scholar] [CrossRef] [PubMed]
- Sanai, N.; Snyder, L.A.; Honea, N.J.; Coons, S.W.; Eschbacher, J.M.; Smith, K.A.; Spetzler, R.F. Intraoperative confocal microscopy in the visualization of 5-aminolevulinic acid fluorescence in low-grade gliomas: Clinical article. J. Neurosurg. 2011, 115, 740–748. [Google Scholar] [CrossRef]
- Eschbacher, J.; Martirosyan, N.L.; Nakaji, P.; Sanai, N.; Preul, M.C.; Smith, K.A.; Coons, S.W.; Spetzler, R.F. In vivo intraoperative confocal microscopy for real-time histopathological imaging of brain tumors: Clinical article. J. Neurosurg. 2012, 116, 854–860. [Google Scholar] [CrossRef] [PubMed]
- Charalampaki, P.; Nakamura, M.; Athanasopoulos, D.; Heimann, A. Confocal-Assisted Multispectral Fluorescent Microscopy for Brain Tumor Surgery. Front. Oncol. 2019, 9, 583. [Google Scholar] [CrossRef] [PubMed]
- Höhne, J.; Schebesch, K.; Zoubaa, S.; Proescholdt, M.; Riemenschneider, M.J.; Schmidt, N.O. Intraoperative imaging of brain tumors with fluorescein: Confocal laser endomicroscopy in neurosurgery. Clinical and user experience. Neurosurg. Focus 2021, 50, E19–E24. [Google Scholar] [CrossRef] [PubMed]
- Schlosser, H.G.; Suess, O.; Vajkoczy, P.; Landeghem, F.K.H.V.; Zeitz, M.; Bojarski, C. Confocal neurolasermicroscopy in human brain perspectives for neurosurgery on a cellular level (including additional Comments to this article). Cent. Eur. Neurosurg.-Zent. Für Neurochir. 2010, 71, 13–16. [Google Scholar] [CrossRef] [PubMed]
- Foersch, S.; Heimann, A.; Ayyad, A.; Spoden, G.A.; Florin, L.; Mpoukouvalas, K.; Kiesslich, R.; Kempski, O.; Goetz, M.; Charalampaki, P. Confocal Laser Endomicroscopy for Diagnosis and Histomorphologic Imaging of Brain Tumors In Vivo. PLoS ONE 2012, 7, e41760. [Google Scholar] [CrossRef]
- Wirth, D.; Snuder, M.; Sheth, S.; Kwon, C.-S.; Frosch, M.P.; Curry, W.; Yaroslavsky, A.N. Identifying brain neoplasms using dye-enhanced multimodal confocal imaging. J. Biomed. Opt. 2012, 17, 026012. [Google Scholar] [CrossRef]
- Snuderl, M.; Wirth, D.; Sheth, S.A.; Bourne, S.K.; Kwon, C.S.; Ancukiewicz, M.; Curry, W.T.; Frosch, M.P.; Yaroslavsky, A.N. Dye-enhanced multimodal confocal imaging as a novel approach to intraoperative diagnosis of brain tumors. Brain Pathol. 2013, 23, 73–81. [Google Scholar] [CrossRef]
- Wirth, D.; Smith, T.W.; Moser, R.; Yaroslavsky, A.N. Demeclocycline as a contrast agent for detecting brain neoplasms using confocal microscopy. Phys. Med. Biol. 2015, 60, 3003–3011. [Google Scholar] [CrossRef]
- Breuskin, D.; Divincenzo, J.; Kim, Y.Y.-J.; Urbschat, S.; Oertel, J. Confocal Laser Endomicroscopy in Neurosurgery: A New Technique with Much Potential. Minim. Invasive Surg. 2013, 2013, 851819. [Google Scholar] [CrossRef]
- Georges, J.; Zehri, A.; Carlson, E.; Nichols, J.; Mooney, M.A.; Martirosyan, N.L.; Ghaffari, L.; Kalani, M.Y.S.; Eschbacher, J.; Feuerstein, B.; et al. Label-free microscopic assessment of glioblastoma bioposy specimens prior to biobanking. Neurosurg. Focus 2014, 36, E8. [Google Scholar] [CrossRef]
- Eschbacher, J.M.; Georges, J.F.; Belykh, E.; Yazdanabadi, M.I.; Martirosyan, N.L.; Szeto, E.; Seiler, C.Y.; Mooney, M.A.; Daniels, J.K.; Goehring, K.Y.; et al. Immediate label-free ex vivo evaluation of human brain tumor biopsies with confocal reflectance microscopy. J. Neuropathol. Exp. Neurol. 2017, 76, 1008–1022. [Google Scholar] [CrossRef] [PubMed]
- Daali, S.; Javed, M.; Altekoester, A.-K.; Linxweiler, M.; Bostelmann, R.; Schlegel, J.; Igressa, A.; Charalampaki, P. Analysis of 258 Different Lesions of the Central Nervous System for Real Time Histopathological Diagnosis Using Confocal Laser Endomicroscopy. J. Mult. Scler. 2016, 3, 2. [Google Scholar] [CrossRef]
- Forest, F.; Cinotti, E.; Yvorel, V.; Habougit, C.; Vassal, F.; Nuti, C.; Perrot, J.-L.; Labeille, B.; Péoc’h, M. Ex vivo confocal microscopy imaging to identify tumor tissue on freshly removed brain sample. J. Neurooncol. 2015, 124, 157–164. [Google Scholar] [CrossRef] [PubMed]
- Wei, L.; Chen, Y.; Yin, C.; Borwege, S.; Sanai, N.; Liu, J.T.C. Optical-sectioning microscopy of protoporphyrin IX fluorescence in human gliomas: Standardization and quantitative comparison with histology. J. Biomed. Opt. 2017, 22, 046005. [Google Scholar] [CrossRef] [PubMed]
- Yoneyama, T.; Watanabe, T.; Kagawa, H.; Hayashi, Y.; Nakada, M. Fluorescence intensity and bright spot analyses using a confocal microscope for photodynamic diagnosis of brain tumors. Photodiagnosis Photodyn. Ther. 2017, 17, 13–21. [Google Scholar] [CrossRef]
- Yoneyama, T.; Watanabe, T.; Tamai, S.; Miyashita, K.; Nakada, M. Bright spot analysis for photodynamic diagnosis of brain tumors using confocal microscopy. Photodiagnosis Photodyn. Ther. 2019, 25, 463–471. [Google Scholar] [CrossRef] [PubMed]
- Martirosyan, N.L.; Georges, J.; Eschbacher, J.M.; Belykh, E.; Carotenuto, A.; Spetzler, R.F.; Nakaji, P.; Preul, M.C. Confocal scanning microscopy provides rapid, detailed intraoperative histological assessment of brain neoplasms: Experience with 106 cases. Clin. Neurol. Neurosurg. 2018, 169, 21–28. [Google Scholar] [CrossRef]
- Mooney, M.A.; Georges, J.; Yazdanabadi, M.I.; Goehring, K.Y.; White, W.L.; Little, A.S.; Preul, M.C.; Coons, S.W.; Nakaji, P.; Eschbacher, J.M. Immediate ex-vivo diagnosis of pituitary adenomas using confocal reflectance microscopy: A proof-of-principle study. J. Neurosurg. 2018, 128, 1072–1075. [Google Scholar] [CrossRef]
- Belykh, E.; Patel, A.A.; Miller, E.J.; Bozkurt, B.; Yağmurlu, K.; Woolf, E.C.; Scheck, A.C.; Eschbacher, J.M.; Nakaji, P.; Preul, M.C. Probe-based three-dimensional confocal laser endomicroscopy of brain tumors: Technical note. Cancer Manag. Res. 2018, 10, 3109–3123. [Google Scholar] [CrossRef]
- Belykh, E.; Zhao, X.; Ngo, B.; Farhadi, D.S.; Byvaltsev, V.A.; Eschbacher, J.M.; Nakaji, P.; Preul, M.C. Intraoperative Confocal Laser Endomicroscopy Ex Vivo Examination of Tissue Microstructure During Fluorescence-Guided Brain Tumor Surgery. Front. Oncol. 2020, 10, 2584. [Google Scholar] [CrossRef]
- Acerbi, F.; Pollo, B.; De Laurentis, C.; Restelli, F.; Falco, J.; Vetrano, I.G.; Broggi, M.; Schiariti, M.; Tramacere, I.; Ferroli, P.; et al. Ex Vivo Fluorescein-Assisted Confocal Laser Endomicroscopy (CONVIVO® System) in Patients With Glioblastoma: Results From a Prospective Study. Front. Oncol. 2020, 10, 606574. [Google Scholar] [CrossRef] [PubMed]
- Beleña, J.M.; Núñez, M.; Rodríguez, M. Adverse Reactions Due to Fluorescein during Retinal Angiography. JSM Ophtalmol. 2013, 1, 4–7. [Google Scholar]
- Dilek, O.; Ihsan, A.; Tulay, H. Anaphylactic reaction after fluorescein sodium administration during intracranial surgery. J. Clin. Neurosci. 2011, 18, 430–431. [Google Scholar] [CrossRef]
- Belykh, E.; Ngo, B.; Farhadi, D.S.; Zhao, X.; Mooney, M.A.; White, W.L.; Daniels, J.K.; Little, A.S.; Eschbacher, J.M.; Preul, M.C. Confocal Laser Endomicroscopy Assessment of Pituitary Tumor Microstructure: A Feasibility Study. J. Clin. Med. 2020, 9, 3146. [Google Scholar] [CrossRef] [PubMed]
- Shay, J.E.S.; Imtiyaz, H.Z.; Sivanand, S.; Durham, A.C.; Skuli, N.; Hsu, S.; Mucaj, V.; Eisinger-Mathason, T.S.K.; Krock, B.L.; Giannoukos, D.N.; et al. Inhibition of Hypoxia-Inducible Factors Limits Tumor Progression in a Mouse Model of Colorectal Cancer; Oxford University Press: New York, NY, USA, 2014; Volume 35, pp. 1067–1077. [Google Scholar]
- Lee, C.-J.; Yue, C.-H.; Lin, Y.-Y.; Wu, J.-C.; Liu, J.-Y. Antitumor activity of acriflavine in human hepatocellular carcinoma cells. Anticancer Res. 2014, 34, 3549–3556. [Google Scholar]
- Yaroslavsky, A.N.; Neel, V.; Anderson, R.R. Demarcation of nonmelanoma skin cancer margins in thick excisions using multispectral polarized light imaging. J. Investig. Dermatol. 2003, 121, 259–266. [Google Scholar] [CrossRef]
- Wirth, D.; Snuderl, M.; Curry, W.; Yaroslavsky, A. Comparative evaluation of methylene blue and demeclocycline for enhancing optical contrast of gliomas in optical images. J. Biomed. Opt. 2014, 19, 090504. [Google Scholar] [CrossRef] [PubMed]
- Peyre, M.; Clermont-Taranchon, E.; Stemmer-Rachamimov, A.; Kalamarides, M. Miniaturized handheld confocal microscopy identifies focal brain invasion in a mouse model of aggressive meningioma. Brain Pathol. 2013, 23, 371–377. [Google Scholar] [CrossRef] [PubMed]
- Martirosyan, N.L.; Cavalcanti, D.D.; Eschbacher, J.M.; Delaney, P.M.; Scheck, A.E.C.; Abdelwahab, M.G.; Nakaji, P.; Spetzler, R.F.; Preul, M.C. Use of in vivo near-infrared laser confocal endomicroscopy with indocyanine green to detect the boundary of infiltrative tumor: Laboratory investigation. J. Neurosurg. 2011, 115, 1131–1138. [Google Scholar] [CrossRef]
- Martirosyan, N.L.; Eschbacher, J.M.; Nakaji, P.; Sanai, N.; Smith, K.A.; Preul, M.C.; Spetzler, R.F. Histopathologic features of brain tumors revealed with in-vivo intraoperative fiber-optic confocal laser endomicroscopy using fluorescein sodium in humans. J. Neurosurg. 2011, 115, A424. [Google Scholar]
- Fotteler, M.; Heninger, L.; Holl, F.; Schlegel, J.; Swoboda, W. Confocal Laser Endomicroscopy for Intraoperative Tumor Assessment: Development of a Conceptual Model for an Evaluation Study. Stud. Health Technol. Inform. 2019, 262, 376–379. [Google Scholar] [CrossRef] [PubMed]
- Izadyyazdanabadi, M.; Belykh, E.; Mooney, M.; Martirosyan, N.; Eschbacher, J.; Nakaji, P.; Preul, M.C.; Yang, Y. Convolutional neural networks: Ensemble modeling, fine-tuning and unsupervised semantic localization for neurosurgical CLE images. J. Vis. Commun. Image Represent. 2018, 54, 10–20. [Google Scholar] [CrossRef]
- Kamen, A.; Sun, S.; Wan, S.; Kluckner, S.; Chen, T.; Gigler, A.M.; Simon, E.; Fleischer, M.; Javed, M.; Daali, S.; et al. Automatic Tissue Differentiation Based on Confocal Endomicroscopic Images for Intraoperative Guidance in Neurosurgery. BioMed Res. Int. 2016, 2016, 6183218. [Google Scholar] [CrossRef] [PubMed]
- Izadyyazdanabadi, M.; Belykh, E.; Zhao, X.; Moreira, L.B.; Gandhi, S.; Cavallo, C.; Eschbacher, J.; Nakaji, P.; Preul, M.C.; Yang, Y. Fluorescence Image Histology Pattern Transformation Using Image Style Transfer. Front. Oncol. 2019, 9, 519. [Google Scholar] [CrossRef]
- Kaur, P.; Sharma, M.; Mittal, M. Big Data and Machine Learning Based Secure Healthcare Framework. Procedia Comput. Sci. 2018, 132, 1049–1059. [Google Scholar] [CrossRef]
- Kaissis, G.A.; Makowski, M.R.; Ruckert, D.; Braren, R.F. Secure, privacy-preserving and federated machine learning in medical imaging. Nat. Mach. Intell. 2020, 2, 305–311. [Google Scholar] [CrossRef]
- Belykh, E.; Cavallo, C.; Gandhi, S.; Zhao, X.; Veljanoski, D.; Yazdanabadi, M.I.; Martirosyan, N.L.; Byvaltsev, V.A.; Eschbacher, J.; Preul, M.C.; et al. Utilization of intraoperative confocal laser endomicroscopy in brain tumor surgery. J. Neurosurg. Sci. 2018, 62, 704–717. [Google Scholar] [CrossRef] [PubMed]
- Mooney, M.A.; Zehri, A.H.; Georges, J.F.; Nakaji, P. Laser scanning confocal endomicroscopy in the neurosurgical operating room: A review and discussion of future applications. Neurosurg. Focus 2014, 36, E9. [Google Scholar] [CrossRef]
| Study, Year | Confocal System Used | Fluorophore Used (Dosages, Protocol) | N. of pts | Pathologies Treated | Main Findings | Sensibility/Specificity; Diagnostic Accuracy | Blinding Level of the Study |
|---|---|---|---|---|---|---|---|
| Sanai et al. (2011) [20] | Optiscan 5.1 | SF (5 mL, 10% in saline solution, injection immediately before imaging) | 35 | 13 LGGs 8 HGGs 8 Meningiomas 3 radiation necrosis | First in vivo experience in humans with confocal laser technology, using SF as contrast enhancer, demonstrating feasibility of in vivo confocal imaging in neurosurgery | / | / |
| Sanai et al. (2011) [21] | Optiscan 5.1 | 5-ALA (20 mg/kg, 3 h before surgery) | 10 | 10 LGGs | First in vivo experience with 5-ALA as contrast enhancer: CLE may be used in conjunction with 5-ALA to detect LGGs at borders | / | / |
| Eschbacher et al. (2012) [22] | Optiscan 5.1 | SF (25 mg iv, 2–5 min before CLE imaging) | 50 | 24 Meningiomas 12 HGGs 8 LGGs 4 Schwannomas 2 Other tumors | First report of CLE accuracy in obtaining in vivo diagnosis | ACCURACY (ex vivo analysis): 92.9% in obtaining correct diagnosis (26/28) | Blindness of pathologist regarding type of tumor operated, available information on location and radiological enhancement; possibility to choose diagnosis among a list of possible tumors |
| Martirosyan et al. (2016) [11] | Optiscan 5.1 | SF (5 mL, 10% in saline solution, iv injection 5 min before imaging) | 74 | 30 Meningiomas 14 Other tumors 13 HGGs 7 No tumors 4 Schwannomas 1 Metastasis | First report of operative data about CLE imaging in vivo; first report of sensibility and specificity for in vivo imaging of HGGs and meningiomas | SENSIBILITY/SPECIFICITY for:→HGGs: 91%/94% →Meningiomas: 97%/93% | Blindness of the study not specifically defined (“Images were reviewed by a neuropathologist and 2 neurosurgeons who were not involved in the surgeries”) |
| Pavlov et al. (2016) [17] | Cellvizio | 5-ALA (4 h before surgery; SF 500 mg/5 mL) | 18 | 6 HGGs 2 LGGs 1 Lymphoma | Feasibility of CLE with 5-ALA and SF; first application of CLE to in vivo brain stereotactic biopsy | / | / |
| Charalampaki et al. (2019) [23] | Multispectral fluorecence microscope + Cellvizio | ICG (50 mg iv 1 h before tumor removal) | 13 | 5 Gliomas 3 Meningiomas 3 Metastases 2 Schwannomas | Feasibility of in vivo concomitant use of multispectral surgical microscope with Glow800 software and CLE imaging in brain tumor surgery; feasibility of inserting CLE probe in endoscopic channel for looking “behind the corner” in brain tumor surgery | / | / |
| Hohne et al. (2021) [24] | Convivo | SF (5 mg/kg, 10% in saline solution, injected intraoperatively (various times) | 12 | 5 Metastasis 4 HGGs 2 LGGs 1 Gliosis/recurrent hemangiopericytoma | Feasibility of in vivo Convivo system in humans in different kind of brain tumors | / | / |
| Study, Year | Confocal System Used | Fluorophore Used (Dosages, Protocol) | N. of pts | Pathologies Treated | Main Findings | Sensibility/Specificity; Diagnostic Accuracy | Blinding Level of the Study |
|---|---|---|---|---|---|---|---|
| Schlosser et al. (2009) [25] | Optiscan | AF (0.05%, topical administration) | 12 | 9 HGGs 3 Meningiomas | Pilot study demonstrating the feasibility of CLE imaging ex vivo in brain tumor surgery | / | / |
| Foersch et al. (2012) [26] | Optiscan | AF (50 microliters topical or SF 50 microliters topical) | 15 | 6 Meningiomas 3 Other tumors 2 Schwannomas 2 Healthy brain specimens 1 HGG 1 Metastasis | First attempt to calculate and report a diagnostic accuracy for ex vivo CLE imaging with AF | ACCURACY in diagnosis on 35 preselected images from 7 different tissues→87% for non-clinicians →93% for clinical experts | 5 raters blinded to histology and macroscopic appearance of tumors |
| Wirth et al. (2012) [27] | Multimodal confocal microscope | MB (1% topical around 2–5 min before imaging) | 119 | 41 HGGs 25 Metastases 14 Meningiomas 11 LGGs | Feasibility of multimodal confocal reflectance and fluorescence imaging for histologic assessment of brain tumors ex vivo | / | / |
| Snuderl et al. (2012) [28] | Multimodal confocal microscope | MB (1% topical around 2–5 min before imaging) | 37 | 10 Metastases 9 HGGs 8 Non tumors 3 LGGs | Sensibility/specificity available for ex vivo imaging with a multimodal confocal reflectance/fluorescence imaging | SENSIBILITY/SPECIFICITY: →in identifying normal vs abnormal tissue of 95%/100% →in identifying glial vs non-glial tumor of 83%/90% →in making final correct diagnosis (neuropathologists) of 88%/100% | 13 pathologists (9 generalists and 4 neuropathologists), all without previous exposure to confocal imaging technique, assessed each case answering to such questions: →is the tissue normal brain or abnormal; →if abnormal, is it a glial or nonglial neoplasm; →if glial, is it a LGG or a HGG; →if nonglial, is it meningioma or metastatic carcinoma. |
| Breuskin et al. (2013) [30] | EndoMag1 | MB (topical around 20 min before imaging) | >50 | Not further described | First feasibility study on red light CLE in brain tumors surgery | / | / |
| Georges et al. (2014) [31] | Benchtop confocal microscope | Only reflectance microscopy | 2 | 1 HGG 1 Radiation necrosis | CLE proposed as a feasible method to distinguish among tumor and radiation necrosis prior to specimens biobanking | / | / |
| Wirth et al. (2015) [29] | Multimodal confocal microscope | DMN (0.75 mg/mL, staining for 20 min 1–2 h after surgery) | 14 | 7 HGGs 4 Metastases 2 Meningiomas 1 Pituitary adenoma | Feasibility study of DMN used as optical contrast enhancer for HGG tumor cells; morphological characteristics of different CNS tumors are presented | / | / |
| Forest et al. (2015) [34] | VivaScope 2500 | AO (undefined dosage) | 19 | 9 Meningiomas 7 HGGs 3 Metastases | Feasibility study of the implementation of a dermatological confocal microscope on a routine use for the most frequent brain tumors. First ex vivo human study to implement AO as contrast enhancer. | / | / |
| Charalampaki et al. (2015) [18] | Cellvizio | AF (0.05% topical) | 150 | 47 HGGs 32 Metastases 30 Meningiomas 16 Other tumors 13 Schwannomas 12 LGGs | Descriptive study showing CLE features of different CNS tumors | / | To establish accuracy and interobserver agreement a set of confocal images (n = 100) of 20 different tissues were selected and presented to 2 groups of raters: nonclinical and clinical experts, blinded to the macroscopic appearance and the histopathological diagnosis of routine pathology. |
| Daali et al. (2016) [33] | Cellvizio | AF (0.01 mg/mL topical) | 258 | 74 Meningiomas 69 Metastases 50 Other tumors 47 HGGs 7 LGGs | Prospective descriptive study reporting the overall accuracy in making diagnosis for ex vivo AF CLE imaging | ACCURACY in obtaining diagnosis of 89%, calculated on preselected images from 258 cases | Images were evaluated by 4 different evaluators (surgeons and neuropathologists). The traditional histopathological findings were blinded to both groups. |
| Breuskin et al. (2017) [19] | EndoMag1 | Only reflectance microscopy | 100 | 34 Meningiomas 32 HGGs 16 Metastases 10 LGGs 8 Schwannomas | First study assessing sensibility and specificity for identifying brain tumors ex vivo with red light confocal imaging without prior contrast administration | SENSIBILITY/SPECIFICITY for diagnosis of: →HGGs 85%/81% →Meningiomas 82%/95% →LGGs 90/93% →Schwannomas 87%/100% →Metastases 7%/94% | The CLE investigator was blinded for patient data and for results of instantaneous sections. |
| Eschbacher et al. (2017) [32] | Benchtop confocal microscope | Only reflectance microscopy | 76 | 25 Meningiomas 24 Other tumors 10 Pituitary adenomas 8 HGGs 7 LGGs 5 Normal pituitary glands 4 Schwannomas 4 Metastases 3 Treatment effect | Excellent image quality study, also reporting a blinded interpretation of acquired images by neuropathologists and general pathologists | ACCURACY: in→asserting tumors vs non tumor: 91.5 % for general pathologists 97.9 % for neuropathologists →labeling lesions with corrected diagnosis: 85.1% for general pathologist 95.8% for neuropathologist | Preselected 47 images, analyzed blindly by neuro and general pathologists without prior experience in confocal laser imaging. Pathologists were aware of eventual contrast enhancement and location of tumors. |
| Wei et al. (2017) [35] | Benchtop confocal laser microscope | 5-ALA (20 mg/kg before surgery per os) | 14 | 14 HGGs and LGGs | Feasibility of ex vivo confocal microscopy analysis after PpIX administration | / | / |
| Yoneyama et al. (2017) [36] | Benchtop confocal laser microscope | 5-ALA (20 mg/kg before surgery per os) | More than 20 | More than 20 HGGs | Fluorescence intensity and bright-spot analysis using 5-ALA as contrast enhancer may help in distinguishing a tumor region, differentiating between infiltrating tumor and normal regions. | / | / |
| Martirosyan et al. (2018) [38] | Benchtop confocal laser microscope | AF (0.05%) AO (0.01%) CV (0.02%) MB (82%) ICG (0.6%) | 106 | 32 Other tumors 30 Meningiomas 19 Gliomas 13 Pituitary adenomas 9 Metastases 3 Non tumor | First study to implement different fluorescent dyes for ex vivo imaging, setting procedural “standards” and reporting high image quality, especially for AF and AO staining. AF and AO staining resulted to be the best option for the great majority of tumors investigated | ACCURACY: Correct identification of: ➔86.7% meningiomas (also correct subtype) ➔89% gliomas (21% correct subtyping) ➔100% pituitary adenomas ➔55.6% metastases | / |
| Mooney et al. (2018) [39] | Benchtop confocal laser microscope | Only reflectance microscopy | 11 | 11 Pituitary tumors | Feasibility study for confocal reflectance microscopy without contrast enhancers for pituitary adenomas, reporting also accuracy in their identification | ACCURACY in making proper diagnosis on 16 preselected images: 94% | 16 representative confocal images from the 11 cases were selected by the neuropathologist (7 images of normal adenohypophysis and 9 images of pituitary adenoma), then presented in a blinded fashion to a second dedicated neuropathologist who had no prior knowledge of the cases. |
| Belykh et al. (2018) [40] | Convivo | SF (2–5 mg/kg iv 5–60 min before imaging) AO (0.1%) AF (0.1%) | 31 | 11 Other tumors 9 HGGs 4 Metastases 3 Schwannomas 3 Meningiomas 1 LGG | Feasibility study for Convivo ex vivo analysis of brain tumors. | / | / |
| Yoneyama et al. (2019) [37] | Benchtop confocal laser microscope | 5-ALA (20 mg/kg before surgery per os) | > 9 | 6 GBMs characterized by 5-ALA induced fluorescence 3GBMs characterized by no 5-ALA induced fluorescence 5 additional specimens (2 HGGs, 1 LGG, 1 recurrent GBM, 1 nerve sarcoma, 1 normal tissue) | Bright-spot analysis may be of help in distinguishing tumorous vs non tumorous tissue also in GBM without 5-ALA induced fluorescence and in other tumor subtypes | / | / |
| Belykh et al. (2020) [45] | Convivo | SF (2 mg/Kg iv before biopsy collection; optimal timing reported to be from 1 min to 10 min before biopsy) | 9 | 9 Pituitary adenomas (13 biopsies) | Feasibility of portable Convivo probe implementation in cadaver heads through a trans-sphenoidal corridor for pituitary adenomas; ex-vivo study of accuracy in their identification with SF as contrast enhancer. | ACCURACY in diagnosing pituitary adenomas with frozen sections as standard: →“definitively” for 13/16 specimens. →“favoring” for 3/16 specimens. | A neuropathologist with experience interpreting CLE images, but who was not involved in the surgical procedures, reviewed the CLE digital images as well as frozen and permanent section slides. |
| Belykh et al. (2020) [41] | Convivo | SF (2 to 5 mg/kg iv upon induction of anesthesia, 5 mg/kg during surgery for CLE contrast improvement) | 47 | 29 HGGs 7 Meningiomas 4 Metastasis 3 LGGs 1 Choroid plexus carcinoma 1 Cranyopharingioma 1 Schwannoma 1 Arterovenous malformation | Very detailed quantitative and descriptive analysis of different brain tumors, along with autofluorescent cells characteristics classification; first time where a second SF injections was used to improve diagnostic power of CLE; blinded study in analyzing CLE images from both neuropathologist and experienced/unexperienced neurosurgeons (regarding CLE imaging interpretation) | DIAGNOSTIC ACCURACY: positive predictive value of: 98% for gliomas 91% for meningiomas 83% for metastasis with an overall diagnostic accuracy of 75% for blinded neuropathologist. DIAGNOSTIC ACCURACY for all biopsies blinded for analysis by a neurosurgeon with experience interpreting CLE images was 78% vs 71% for the neurosurgeon without CLE image-reading experience. | →The neuropathologist had no clinical information except that the biopsy had been performed during an intracranial procedure →Two neurosurgeons (experienced and unexperienced) reviewed a set of CLE images, after being instructed on the key histologic features on CLE images. No information was provided to the neurosurgeons or general pathologist regarding case history, imaging or diagnosis |
| Acerbi et al. (2020) [42] | Convivo | SF (5 mg/kg at anesthesia induction) | 15 | 15 HGGs | First available study where the ability of Convivo in obtaining intraoperative diagnosis and categorizing morphological patterns at both central core and tumor margins was assessed prospectively and based on a near real-time, blinded interpretation of the pathologist during surgery in OR. | ACCURACY in: →Obtaining a diagnosis (compared to frozen sections): 80% at central core, 80% at tumor border →Categorizing patterns (compared to frozen sections): 93.3% at central core and 80% at tumor margins →Obtaining a diagnosis (compared to permanent sections): 80% at central core, 67% at tumor border →Categorizing patterns (compared to permanent sections): 86% at central core and 67% at tumor margins. | A dedicated pathologist was asked to judge in near real-time intraoperatively if the tissue represented tumor tissue, to provide a possible intraoperative tumor diagnosis, and to categorize eventual morphological patterns (blinded to frozen and permanent section results) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Restelli, F.; Pollo, B.; Vetrano, I.G.; Cabras, S.; Broggi, M.; Schiariti, M.; Falco, J.; de Laurentis, C.; Raccuia, G.; Ferroli, P.; et al. Confocal Laser Microscopy in Neurosurgery: State of the Art of Actual Clinical Applications. J. Clin. Med. 2021, 10, 2035. https://doi.org/10.3390/jcm10092035
Restelli F, Pollo B, Vetrano IG, Cabras S, Broggi M, Schiariti M, Falco J, de Laurentis C, Raccuia G, Ferroli P, et al. Confocal Laser Microscopy in Neurosurgery: State of the Art of Actual Clinical Applications. Journal of Clinical Medicine. 2021; 10(9):2035. https://doi.org/10.3390/jcm10092035
Chicago/Turabian StyleRestelli, Francesco, Bianca Pollo, Ignazio Gaspare Vetrano, Samuele Cabras, Morgan Broggi, Marco Schiariti, Jacopo Falco, Camilla de Laurentis, Gabriella Raccuia, Paolo Ferroli, and et al. 2021. "Confocal Laser Microscopy in Neurosurgery: State of the Art of Actual Clinical Applications" Journal of Clinical Medicine 10, no. 9: 2035. https://doi.org/10.3390/jcm10092035
APA StyleRestelli, F., Pollo, B., Vetrano, I. G., Cabras, S., Broggi, M., Schiariti, M., Falco, J., de Laurentis, C., Raccuia, G., Ferroli, P., & Acerbi, F. (2021). Confocal Laser Microscopy in Neurosurgery: State of the Art of Actual Clinical Applications. Journal of Clinical Medicine, 10(9), 2035. https://doi.org/10.3390/jcm10092035








