Characteristics of the Urinary Proteome in Women with Overactive Bladder Syndrome: A Case-Control Study
Abstract
1. Introduction
2. Materials and Methods
3. Results
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Haylen, B.T.; de Ridder, D.; Freeman, R.M.; Swift, S.E.; Berghmans, B.; Lee, J.; Monga, A.; Petri, E.; Rizk, D.E.; Sand, P.K.; et al. An International Urogynecological Association (IUGA)/International Continence Society (ICS) joint report on the terminology for female pelvic floor dysfunction. Int. Urogynecol. J. 2010, 21, 5–26. [Google Scholar] [CrossRef] [PubMed]
- Cheung, W.W.; Khan, N.H.; Choi, K.K.; Bluth, M.H.; Vincent, M.T. Prevalence, evaluation and management of overactive bladder in primary care. BMC Fam. Pract. 2009, 10, 8. [Google Scholar] [CrossRef]
- Melville, J.L.; Katon, W.; Delaney, K.; Newton, K. Urinary incontinence in US women: A population-based study. Arch. Intern. Med. 2005, 165, 537–542. [Google Scholar] [CrossRef]
- Irwin, D.E.; Mungapen, L.; Milsom, I.; Kopp, Z.; Reeves, P.; Kelleher, C. The economic impact of overactive bladder syndrome in six Western countries. BJU Int. 2009, 103, 202–209. [Google Scholar] [CrossRef]
- Corcos, J.; Przydacz, M.; Campeau, L.; Gray, G.; Hickling, D.; Honeine, C.; Radomski, S.B.; Stothers, L.; Wagg, A.; Lond, F. CUA guideline on adult overactive bladder. Can. Urol. Assoc. J. J. L’association Urol. Can. 2017, 11, E142–E173. [Google Scholar] [CrossRef]
- Cardozo, L.; Chapple, C.R.; Dmochowski, R.; Fitzgerald, M.P.; Hanno, P.; Michel, M.C.; Staskin, D.; Van Kerrebroeck, P.; Wyndaele, J.J.; Yamaguchi, O.; et al. Urinary urgency—Translating the evidence base into daily clinical practice. Int. J. Clin. Pract. 2009, 63, 1675–1682. [Google Scholar] [CrossRef]
- Bhide, A.A.; Cartwright, R.; Khullar, V.; Digesu, G.A. Biomarkers in overactive bladder. Int. Urogynecol. J. 2013, 24, 1065–1072. [Google Scholar] [CrossRef]
- Tyagi, P.; Tyagi, V.; Qu, X.; Lin, H.T.; Kuo, H.C.; Chuang, Y.C.; Chancellor, M. Association of inflammaging (inflammation + aging) with higher prevalence of OAB in elderly population. Int. Urol. Nephrol. 2013. [Google Scholar] [CrossRef]
- Liu, H.T.; Jiang, Y.H.; Kuo, H.C. Increased serum adipokines implicate chronic inflammation in the pathogenesis of overactive bladder syndrome refractory to antimuscarinic therapy. PLoS ONE 2013, 8, e76706. [Google Scholar] [CrossRef]
- Rachaneni, S.; Arya, P.; Latthe, P. Urinary nerve growth factor: A biomarker of detrusor overactivity? A systematic review. Int. Urogynecol. J. 2013, 24, 1603–1609. [Google Scholar] [CrossRef] [PubMed]
- Kashyap, M.; Kawamorita, N.; Tyagi, V.; Sugino, Y.; Chancellor, M.; Yoshimura, N.; Tyagi, P. Down-regulation of nerve growth factor expression in the bladder by antisense oligonucleotides as new treatment for overactive bladder. J. Urol. 2013, 190, 757–764. [Google Scholar] [CrossRef]
- Seth, J.H.; Sahai, A.; Khan, M.S.; van der Aa, F.; de Ridder, D.; Panicker, J.N.; Dasgupta, P.; Fowler, C.J. Nerve growth factor (NGF): A potential urinary biomarker for overactive bladder syndrome (OAB)? BJU Int. 2013, 111, 372–380. [Google Scholar] [CrossRef]
- Antunes-Lopes, T.; Pinto, R.; Barros, S.C.; Botelho, F.; Silva, C.M.; Cruz, C.D.; Cruz, F. Urinary neurotrophic factors in healthy individuals and patients with overactive bladder. J. Urol. 2013, 189, 359–365. [Google Scholar] [CrossRef]
- Cho, K.J.; Kim, H.S.; Koh, J.S.; Kim, J.C. Changes in urinary nerve growth factor and prostaglandin E2 in women with overactive bladder after anticholinergics. Int. Urogynecol. J. 2013, 24, 325–330. [Google Scholar] [CrossRef]
- Wang, L.W.; Han, X.M.; Chen, C.H.; Ma, Y.; Hai, B. Urinary brain-derived neurotrophic factor: A potential biomarker for objective diagnosis of overactive bladder. Int. Urol. Nephrol. 2013. [Google Scholar] [CrossRef]
- Koch, M.; Mitulovic, G.; Hanzal, E.; Umek, W.; Seyfert, S.; Mohr, T.; Koelbl, H.; Laterza, R.M. Urinary proteomic pattern in female stress urinary incontinence: A pilot study. Int. Urogynecol. J. 2016, 27, 1729–1734. [Google Scholar] [CrossRef][Green Version]
- Koch, M.; Umek, W.; Hanzal, E.; Mohr, T.; Seyfert, S.; Koelbl, H.; Mitulović, G. Serum proteomic pattern in female stress urinary incontinence. Electrophoresis 2018, 39, 1071–1078. [Google Scholar] [CrossRef]
- Futyma, K.; Nowakowski, Ł.; Ziętek-Strobl, A.; Kamińska, A.; Taoussi, N.; Rechberger, T. Urine Proteomic Study in OAB Patients-Preliminary Report. J. Clin. Med. 2020, 9, 1389. [Google Scholar] [CrossRef]
- Wessel, D.; Flügge, U.I. A method for the quantitative recovery of protein in dilute solution in the presence of detergents and lipids. Anal. Biochem. 1984, 138, 141–143. [Google Scholar] [CrossRef]
- Mitulović, G.; Stingl, C.; Steinmacher, I.; Hudecz, O.; Hutchins, J.R.; Peters, J.M.; Mechtler, K. Preventing carryover of peptides and proteins in nano LC-MS separations. Anal. Chem. 2009, 81, 5955–5960. [Google Scholar] [CrossRef]
- Jung, S.H. Sample size for FDR-control in microarray data analysis. Bioinformatics 2005, 21, 3097–3104. [Google Scholar] [CrossRef] [PubMed]
- Jassal, B.; Matthews, L.; Viteri, G.; Gong, C.; Lorente, P.; Fabregat, A.; Sidiropoulos, K.; Cook, J.; Gillespie, M.; Haw, R.; et al. The reactome pathway knowledgebase. Nucleic Acids Res. 2020, 48, D498–D503. [Google Scholar] [CrossRef]
- Fabregat, A.; Sidiropoulos, K.; Viteri, G.; Forner, O.; Marin-Garcia, P.; Arnau, V.; D’Eustachio, P.; Stein, L.; Hermjakob, H. Reactome pathway analysis: A high-performance in-memory approach. BMC Bioinform. 2017, 18, 142. [Google Scholar] [CrossRef]
- Kültz, D. Molecular and evolutionary basis of the cellular stress response. Annu. Rev. Physiol. 2005, 67, 225–257. [Google Scholar] [CrossRef]
- Yang, Y.; Yu, X. Regulation of apoptosis: The ubiquitous way. FASEB J. Off. Publ. Fed. Am. Soc. Exp. Biol. 2003, 17, 790–799. [Google Scholar] [CrossRef]
- Stanley, P.; Schachter, H.; Taniguchi, N. N-Glycans. In Essentials of Glycobiology; Varki, A., Cummings, R.D., Esko, J.D., Freeze, H.H., Stanley, P., Bertozzi, C.R., Hart, G.W., Etzler, M.E., Eds.; Cold Spring Harbor Laboratory Press: Cold Spring Harbor, NY, USA, 2009. [Google Scholar]
- Sparks, S.E.; Krasnewich, D.M. Congenital Disorders of N-Linked Glycosylation and Multiple Pathway Overview. In GeneReviews(®); Adam, M.P., Ardinger, H.H., Pagon, R.A., Wallace, S.E., Bean, L.J.H., Mirzaa, G., Amemiya, A., Eds.; University of Washington: Seattle, WA, USA, 1993. [Google Scholar]
- Depner, C.M.; Melanson, E.L.; McHill, A.W.; Wright, K.P., Jr. Mistimed food intake and sleep alters 24-hour time-of-day patterns of the human plasma proteome. Proc. Natl. Acad. Sci. USA 2018, 115, E5390–E5399. [Google Scholar] [CrossRef]
| OAB (N = 20) | Control (N = 20) | p-Value | |
|---|---|---|---|
| (Mean ± SD) | |||
| Age (years) | 57.5 (±16) | 55.3 (±17) | 0.904 |
| BMI (kg/m2) | 27.7 (±5) | 25.5 (±5) | 0.740 |
| Vaginal deliveries (n) | 1.15 | 0.99 | 0.484 |
| Chronic diseases * (n) | 12/20 (60%) | 8/20 (40%) | 0.206 |
| Menopausal status (n) | 0.744 | ||
| premenopausal | 7/20 (35%) | 8/20 (40%) | |
| postmenopausal | 13/20 (65%) | 12/20 (60%) | |
| Smoking (yes) | 2/20 (10%) | 3/20 (15%) | 0.633 |
| Caffeine consumption | 17/20 (85%) | 16/20 (80%) | 0.677 |
| ICIQ sum score | 31 | 0 | - |
| Protein | Accession | Gene Symbol |
|---|---|---|
| 26S proteasome non-ATPase regulatory subunit 11 | O00231 | PSD11 |
| 26S proteasome regulatory subunit 10B | A0A087 × 2I1 | A0A087 × 2I1 |
| 3-hydroxyacyl-CoA dehydrogenase type-2 | Q99714 | HCD2 |
| 60S ribosomal protein L15 | P61313 | RL15 |
| Alpha-centractin | R4GMT0 | R4GMT0 |
| Alpha-mannosidase 2 | Q16706 | MA2A1 |
| Asparagine--tRNA ligase, cytoplasmic | O43776 | SYNC |
| Beta/gamma crystallin domain-containing protein 1 | A0A0J9YWL0 | A0A0J9YWL0 |
| Bleomycin hydrolase | Q13867 | BLMH |
| Coatomer subunit beta | P53618 | COPB |
| Cytoplasmic dynein 1 heavy chain 1 | Q14204 | DYHC1 |
| Cytosolic phospholipase A2 delta | Q86XP0 | PA24D |
| Electron transfer flavoprotein subunit alpha, mitochondrial | P13804 | ETFA |
| Eukaryotic translation initiation factor 2 subunit 1 | P05198 | IF2A |
| Eukaryotic translation initiation factor 3 subunit A | Q14152 | EIF3A |
| Heat shock 70 kDa protein 4L | O95757 | HS74L |
| Isoform 2 of 26S proteasome regulatory subunit 6B | P43686-2 | PRS6B |
| Isoform 2 of Alpha-S1-casein | P47710-2 | CASA1 |
| Isoform 2 of Glutamine--tRNA ligase | P47897-2 | SYQ |
| Isoform 2 of Nck-associated protein 1 | Q9Y2A7-2 | NCKP1 |
| Isoform 2 of Serine/threonine-protein phosphatase 2A 55 kDa regulatory subunit B alpha isoform | P63151-2 | 2ABA |
| Isoform 2 of Threonine--tRNA ligase, cytoplasmic | P26639-2 | SYTC |
| Kappa-casein | P07498 | CASK |
| Mitogen-activated protein kinase 1 | P28482 | MK01 |
| Mucin-4 | Q99102 | MUC4 |
| Peroxisomal acyl-coenzyme A oxidase 1 | Q15067 | ACOX1 |
| Polypyrimidine tract-binding protein 1 | A0A0U1RRM4 | A0A0U1RRM4 |
| Prenylcysteine oxidase 1 | Q9UHG3 | PCYOX |
| Proteasome subunit alpha type-2 | P25787 | PSA2 |
| Protein flightless-1 homolog | Q13045 | FLII |
| Spectrin beta chain, non-erythrocytic 2 | O15020 | SPTN2 |
| Spliceosome RNA helicase DDX39B | Q13838 | DX39B |
| Tripartite motif-containing protein 16 | O95361 | TRI16 |
| Tropomodulin-3 | Q9NYL9 | TMOD3 |
| UDP-glucose 4-epimerase | Q14376 | GALE |
| Vesicle-fusing ATPase | P46459 | NSF |
| X-ray repair cross-complementing protein 5 | P13010 | XRCC5 |
| Protein | Accession | Gene Symbol |
|---|---|---|
| Trefoil factor 3 | X6R3S7 | X6R3S7 |
| Latent-transforming growth factor beta-binding protein 2 | G3V3 × 5 | G3V3 × 5 |
| Meprin A subunit | B7ZL91 | B7ZL91 |
| Testican-1 | A0A0A0MQX7 | A0A0A0MQX7 |
| IgGFc-binding protein | Q9Y6R7 | FCGBP |
| Frizzled-4 | Q9ULV1 | FZD4 |
| Mucin-5B | Q9HC84 | MUC5B |
| Growth/differentiation factor 15 | Q99988 | GDF15 |
| Isoform 2 of Phosphoinositide-3-kinase-interacting protein 1 | Q96FE7-2 | P3IP1 |
| BPI fold-containing family B member 1 | Q8TDL5 | BPIB1 |
| CD99 antigen-like protein 2 | Q8TCZ2 | C99L2 |
| Adhesion G protein-coupled receptor F5 | Q8IZF2 | AGRF5 |
| Protein S100-A7A | Q86SG5 | S1A7A |
| Elongation factor Tu | Q83ES6 | EFTU |
| Protocadherin Fat 4 | Q6V0I7 | FAT4 |
| Na(+)/H(+) exchange regulatory cofactor NHE-RF3 | Q5T2W1 | NHRF3 |
| FRAS1-related extracellular matrix protein 2 | Q5SZK8 | FREM2 |
| Trefoil factor 2 | Q03403 | TFF2 |
| Fibulin-2 | P98095 | FBLN2 |
| Hepcidin | P81172 | HEPC |
| Brain acid soluble protein 1 | P80723 | BASP1 |
| Hemoglobin subunit gamma-2 | P69892 | HBG2 |
| NPC intracellular cholesterol transporter 2 | P61916 | NPC2 |
| Beta-defensin 1 | P60022 | DEFB1 |
| Secreted Ly-6/uPAR-related protein 1 | P55000 | SLUR1 |
| Isoform 2 of Cysteine-rich secretory protein 3 | P54108-2 | CRIS3 |
| Afamin | P43652 | AFAM |
| Fibrillin-1 | P35555 | FBN1 |
| Isoform 2 of Syndecan-4 | P31431-2 | SDC4 |
| Granulins | P28799 | GRN |
| Serum paraoxonase/arylesterase 1 | P27169 | PON1 |
| CD27 antigen | P26842 | CD27 |
| Fibulin-1 | P23142 | FBLN1 |
| Small proline-rich protein 2D | P22532 | SPR2D |
| Cornifin-B | P22528 | SPR1B |
| Azurocidin | P20160 | CAP7 |
| Elafin | P19957 | ELAF |
| Inter-alpha-trypsin inhibitor heavy chain H2 | P19823 | ITIH2 |
| Complement component C7 | P10643 | CO7 |
| Matrilysin | P09237 | MMP7 |
| Tumor necrosis factor receptor superfamily member 16 | P08138 | TNR16 |
| Calbindin | P05937 | CALB1 |
| Heparin cofactor 2 | P05546 | HEP2 |
| Major prion protein | P04156 | PRIO |
| Trefoil factor 1 | P04155 | TFF1 |
| Antileukoproteinase | P03973 | SLPI |
| Cystatin-A | P01040 | CYTA |
| Complement C5 | P01031 | CO5 |
| Plasminogen | P00747 | PLMN |
| Superoxide dismutase [Cu-Zn] | P00441 | SODC |
| CD5 antigen-like | O43866 | CD5L |
| Ribonuclease T2 | O00584 | RNT2 |
| Isoform 2 of Deoxyribonuclease-2-alpha | O00115-2 | DNS2A |
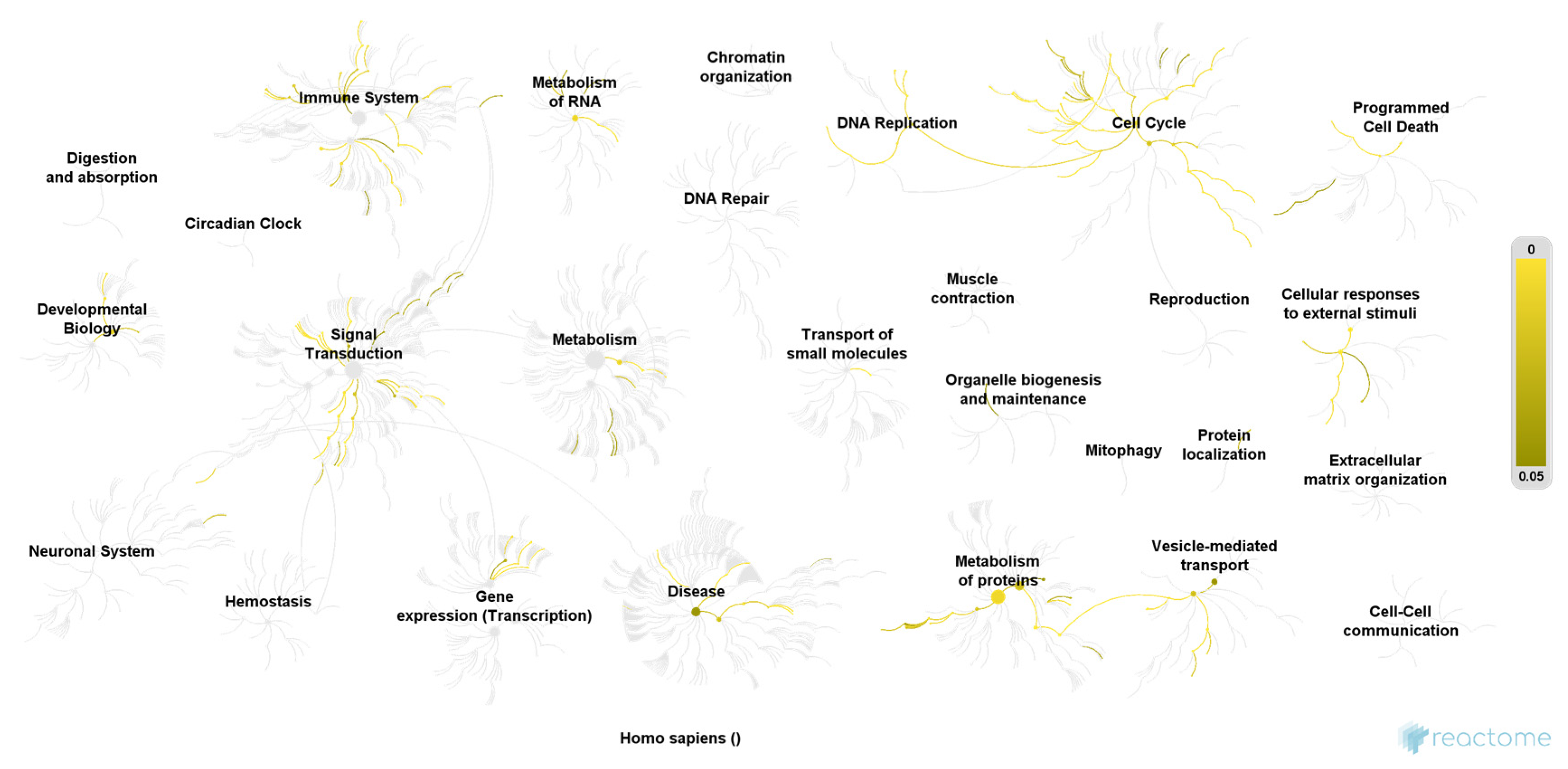
| Pathway | Ratio of Identified Proteins | p-Value | False Discovery Rate |
|---|---|---|---|
| COPI-mediated anterograde transport | 5/102 | 0.00003 | 0.006 |
| Transport to the Golgi and subsequent modification | 6/186 | 0.00004 | 0.007 |
| ER to Golgi Anterograde Transport | 5/155 | 0.0002 | 0.012 |
| ABC-family proteins mediated transport | 4/103 | 0.0005 | 0.012 |
| Cellular responses to stress | 7/408 | 0.0005 | 0.012 |
| G2/M Transition | 5/198 | 0.0006 | 0.012 |
| Asparagine N-linked glycosylation | 6/305 | 0.0006 | 0.012 |
| Mitotic G2-G2/M phases | 5/200 | 0.0007 | 0.012 |
| Regulation of activated PAK-2p34 by proteasome mediated degradation | 3/50 | 0.0007 | 0.012 |
| Cross-presentation of soluble exogenous antigens (endosomes) | 3/50 | 0.0007 | 0.012 |
| Intra-Golgi and retrograde Golgi-to-ER traffic | 5/206 | 0.0007 | 0.012 |
| Regulation of ornithine decarboxylase (ODC) | 3/51 | 0.0008 | 0.012 |
| Autodegradation of the E3 ubiquitin ligase COP1 | 3/52 | 0.0008 | 0.012 |
| Ubiquitin-dependent degradation of Cyclin D1 | 3/52 | 0.0008 | 0.012 |
| Ubiquitin-dependent degradation of Cyclin D | 3/52 | 0.0008 | 0.012 |
| Transcriptional regulation by RUNX2 | 4/121 | 0.0009 | 0.012 |
| Vpu mediated degradation of CD4 | 3/53 | 0.0009 | 0.012 |
| Regulation of Apoptosis | 3/53 | 0.0009 | 0.012 |
| p53-Independent DNA Damage Response | 3/53 | 0.0009 | 0.012 |
| p53-Independent G1/S DNA damage checkpoint | 3/53 | 0.0009 | 0.012 |
| Ubiquitin Mediated Degradation of Phosphorylated Cdc25A | 3/53 | 0.0009 | 0.012 |
| FBXL7 down-regulates AURKA during mitotic entry and in early mitosis | 3/55 | 0.0010 | 0.012 |
| SCF-beta-TrCP mediated degradation of Emi1 | 3/55 | 0.0010 | 0.012 |
| Degradation of AXIN | 3/55 | 0.0010 | 0.012 |
| Negative regulation of NOTCH4 signaling | 3/55 | 0.0010 | 0.012 |
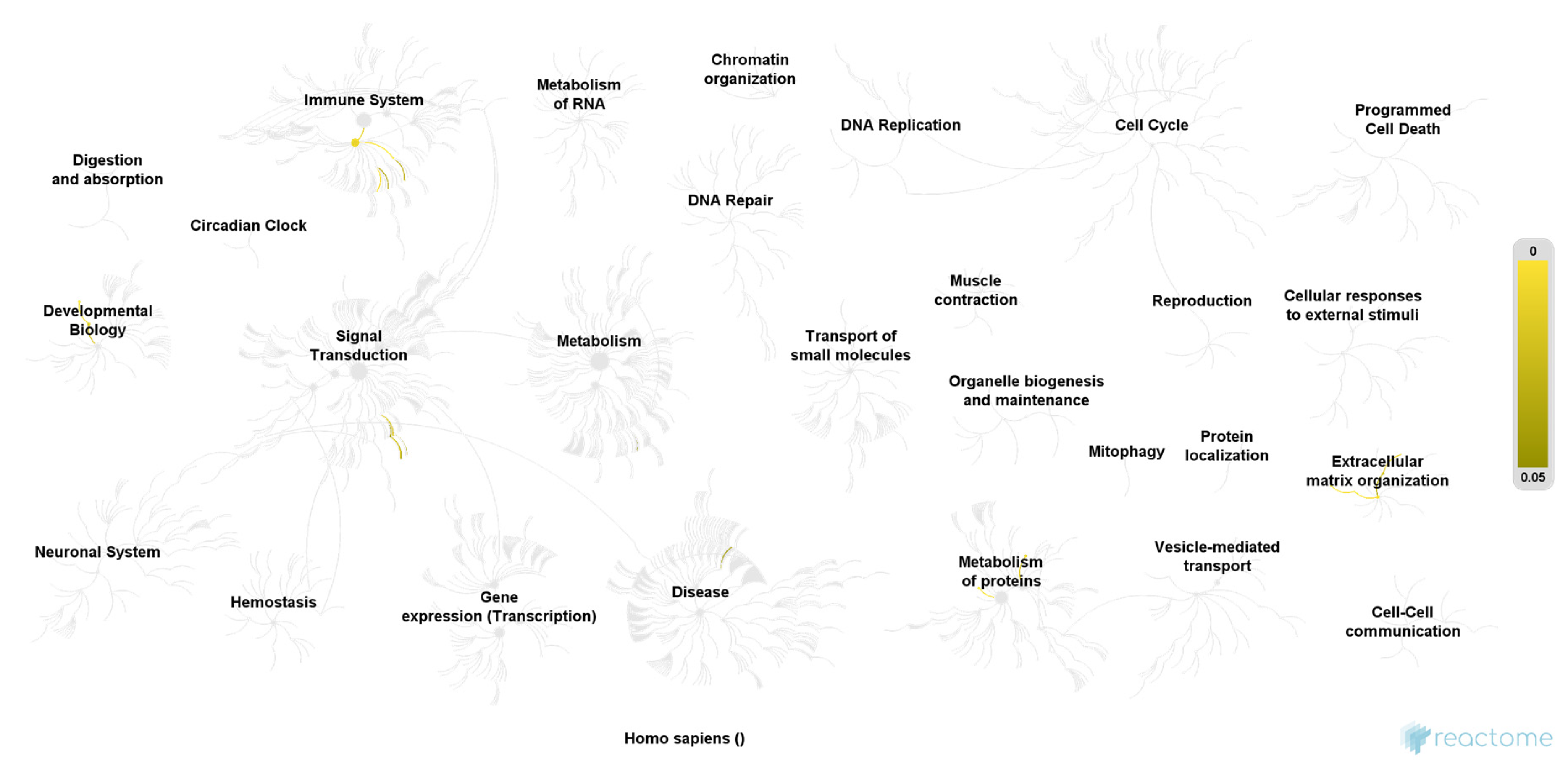
| Pathway | Ratio of Identified Proteins | p-Value | False Discovery Rate |
|---|---|---|---|
| Molecules associated with elastic fibers | 4/38 | 0.00004 | 0.004 |
| Elastic fiber formation | 4/45 | 0.00007 | 0.004 |
| Terminal pathway of complement | 2/8 | 0.0007 | 0.026 |
| Antimicrobial peptides | 4/95 | 0.001 | 0.033 |
| Regulation of Insulin-like Growth Factor (IGF) transport and uptake by Insulin-like Growth Factor Binding Proteins (IGFBPs) | 4/124 | 0.003 | 0.054 |
| Extracellular matrix organization | 6/301 | 0.003 | 0.054 |
| Formation of the cornified envelope | 4/129 | 0.003 | 0.054 |
| Innate Immune System | 12/1183 | 0.009 | 0.122 |
| Activation of Matrix Metalloproteinases | 2/33 | 0.011 | 0.132 |
| NFG and proNGF binds to p75NTR | 1/3 | 0.014 | 0.132 |
| Ceramide signaling | 1/3 | 0.014 | 0.132 |
| Post-translational protein phosphorylation | 3/107 | 0.015 | 0.132 |
| Axonal growth stimulation | 1/4 | 0.019 | 0.151 |
| Keratinization | 4/217 | 0.020 | 0.160 |
| NADE modulates death signaling | 1/6 | 0.028 | 0.169 |
| p75NTR negatively regulates cell cycle via SC1 | 1/6 | 0.028 | 0.169 |
| Metal sequestration by antimicrobial proteins | 1/6 | 0.028 | 0.169 |
| Degradation of the extracellular matrix | 3/140 | 0.029 | 0.176 |
| Activation of C3 and C5 | 1/7 | 0.033 | 0.187 |
| RNF mutants show enhanced WNT signaling and proliferation | 1/8 | 0.037 | 0.187 |
| Axonal growth inhibition (RHOA activation) | 1/9 | 0.042 | 0.205 |
| Synthesis of 5-eicosatetraenoic acids | 1/9 | 0.042 | 0.205 |
| p75NTR regulates axonogenesis | 1/10 | 0.047 | 0.205 |
| Regulated proteolysis of p75NTR | 1/11 | 0.051 | 0.205 |
| Dissolution of Fibrin Clot | 1/13 | 0.06 | 0.224 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Koch, M.; Lyatoshinsky, P.; Mitulovic, G.; Bodner-Adler, B.; Lange, S.; Hanzal, E.; Umek, W. Characteristics of the Urinary Proteome in Women with Overactive Bladder Syndrome: A Case-Control Study. J. Clin. Med. 2021, 10, 2446. https://doi.org/10.3390/jcm10112446
Koch M, Lyatoshinsky P, Mitulovic G, Bodner-Adler B, Lange S, Hanzal E, Umek W. Characteristics of the Urinary Proteome in Women with Overactive Bladder Syndrome: A Case-Control Study. Journal of Clinical Medicine. 2021; 10(11):2446. https://doi.org/10.3390/jcm10112446
Chicago/Turabian StyleKoch, Marianne, Pavel Lyatoshinsky, Goran Mitulovic, Barbara Bodner-Adler, Sören Lange, Engelbert Hanzal, and Wolfgang Umek. 2021. "Characteristics of the Urinary Proteome in Women with Overactive Bladder Syndrome: A Case-Control Study" Journal of Clinical Medicine 10, no. 11: 2446. https://doi.org/10.3390/jcm10112446
APA StyleKoch, M., Lyatoshinsky, P., Mitulovic, G., Bodner-Adler, B., Lange, S., Hanzal, E., & Umek, W. (2021). Characteristics of the Urinary Proteome in Women with Overactive Bladder Syndrome: A Case-Control Study. Journal of Clinical Medicine, 10(11), 2446. https://doi.org/10.3390/jcm10112446







