COVID-19 Vaccination: From Interesting Agent to the Patient
Abstract
1. Introduction
1.1. Covid-19 Breakdown and Background
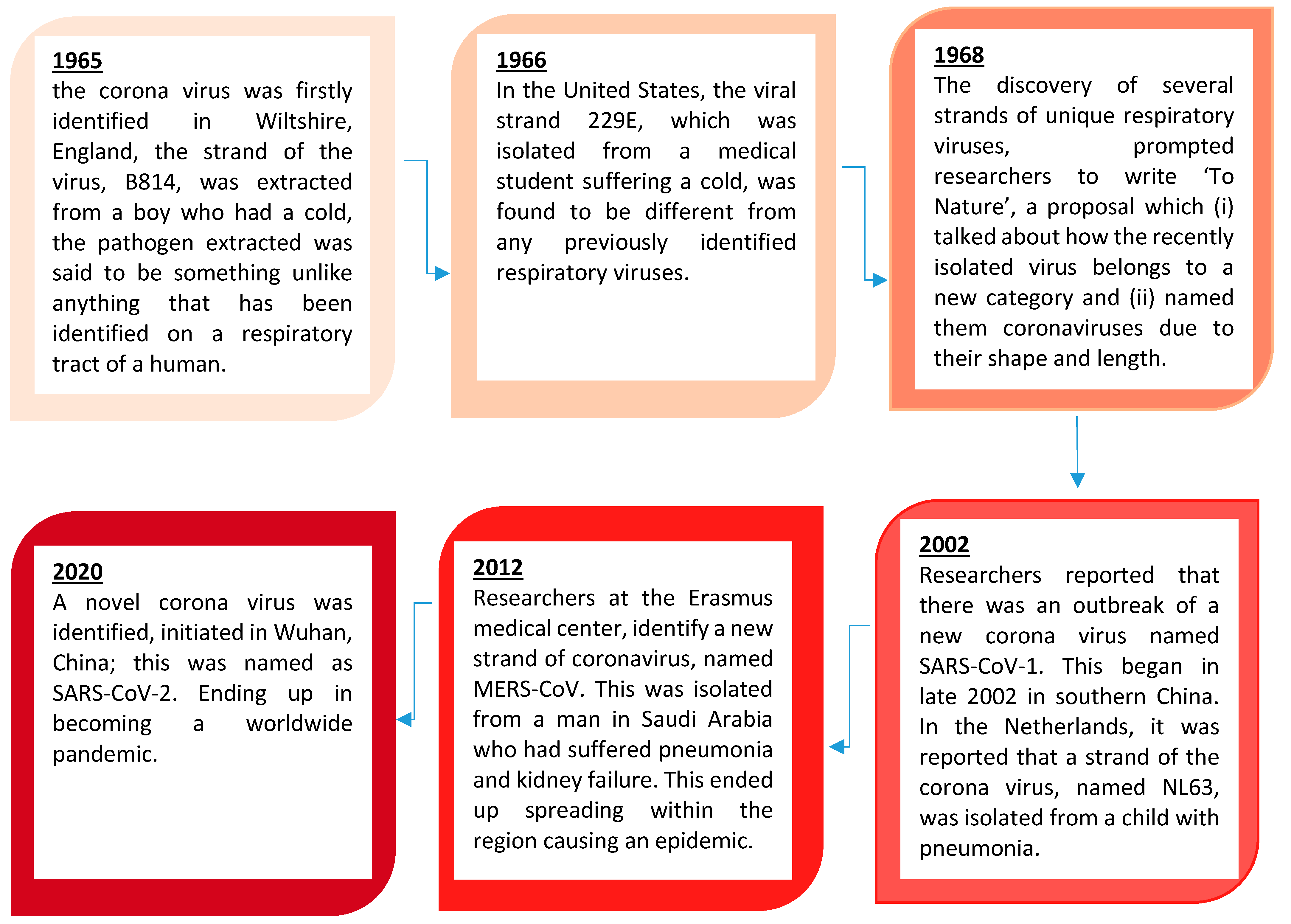
1.2. The Research and History Ortho-Coronavirinae
1.3. Vaccinations Proposed for SARS-CoV-1
- Inactivated SARS-CoV based vaccine: this vaccine expressed several structural proteins such as nucleo-capsid, membranes and spike proteins [27]. These are thought to induce an immune reaction that is capable of stimulating an immune response. The inactivated virus was intended for use as a first-generation vaccine, this is due to the ease of generation of these inactivated viral particles. The next step was the replacement of the inactivated viral vaccine by a second vaccine based around fragments containing neutralizing epitopes that are safer and more efficacious to use. Several reports have shown that SARS-CoV-1 was inactivated with formaldehyde, UV light, and β-propiolactone which can induce virus-neutralizing antibodies in immunized animals [28,29,30].
- S-protein based vaccines: several recombinant based vaccines that have expressed the spike protein in SARS-CoV-1 were assessed in pre-clinical studies [31]. Reports have shown that candidate DNA vaccines encoding the spike protein stimulated an immune response. This led to the study showing that injected mice are protected for SARS-CoV-1. Wang et al. have produced higher titres of neutralizing antibodies and demonstrated that major and minor neutralizing epitopes are located in the S1 and S2 subunits, respectively [32]. Other groups also found neutralizing epitopes in the S2 subunit [33,34]. Bisht et al. [35] have shown that intranasal or intramuscular inoculations of mice with highly attenuated Modified Vacciniavirus Ankara (MVA) vaccines encoding full-length SARS-CoV-1 S protein. This also produced neutralizing antibodies. Bukreyev et al. [36] reported that mucosal immunization of African green monkeys with an attenuated parainfluenza virus expressing S protein resulted in the production of neutralizing antibodies and protected animals from infection by challenge with SARS-CoV-1. These data suggest that the S protein can induce neutralizing antibodies and protective responses in immunized animals [37].
- Vaccines based on fragments containing neutralizing epitopes: fragments that were responsible for the virus binding to receptors within a host cell were targeted. Patients and animals that became infected with SAR-CoV-1 reacted strongly to this type of vaccine. They were immunized and inactivated with a receptor-binding domain (RBD) [29,38]. Absorption of antibodies by RBD showed the capability for removal of most of the neutralizing antibodies, RBD-specific antibodies isolated from these antisera have potent neutralizing activity [38,39]. The immunized mice were protected from SARS-CoV-1. The antibodies purified from the antisera against SARS-CoV-1 significantly inhibited RBD binding to ACE2 [29,38,40,41]. This suggested that RBD contains the major neutralizing epitopes in the S protein and is an ideal SARS vaccine candidate because RBD contains the receptor-binding site, which is critical for virus attachment to the target cell for infection [42,43,44]. Antibodies specific for RBD are expected to block the binding of the virus to the target cell. Therefore, RBD induces higher titers of neutralizing antibodies than those vaccines expressing the full-length S protein [31,32,35,37,43].
1.4. Vaccinations Proposed for MERS-CoV
- Recombinant MERS-CoV: unlike the SARS-CoV-1 vaccine, the MERS-CoV vaccine was constructed based on the recombinant viruses using reverse genetics. This resulted in expressed marker mutations, which allowed for replication-competent, propagation-defective MERS-CoV vaccines [45].
- Viral-Vector-based MERS vaccine: this is similar to the vector-based SARS vaccine; MERS vaccines can also be constructed using viral vectors that express major MERS-CoV proteins, normally the S protein. Several such MERS vaccine candidates have been developed and/or tested for efficacy in mouse models or camels [46,47,48,49]. Viral vectors expressing full-length S protein of MERS-CoV induced S-specific antibody responses and/or T-cell responses in a mouse model via the intramuscular route, showed effective in vitro neutralization for MERS-CoV infection [47,50]. Additionally, vaccination of mice with an MVA-based full-length S vaccine-elicited MERS-CoV-specific CD8+ T cell response and neutralizing antibodies, protecting mice against MERS-CoV [48,49]. Intra-nasally or intra-muscularly administered MVA-S vaccine-induced mucosal immunity resulted in a significant reduction of excreted infectious viruses and viral RNA transcripts [45,46].
- Nanoparticle-based MERS vaccine: in recent years nanoparticles have been at the forefront of many research projects, this has allowed them to have the potential to develop a MERS vaccine. Nanoparticles containing MERS-CoV full-length S proteins can be prepared and purified from pellets of infected baculovirus insect cells. In the absence of adjuvants, these nanoparticles induced a lower level of MERS-CoV producing antibodies in mice. While in the presence of adjuvants, such as aluminium hydroxide (Alum) or Matrix M1, such antibodies were significantly increased and maintained. Thus, adjuvants are required for MERS nanoparticle vaccines and different adjuvants function differently in promoting the immunogenicity of these vaccines [45].
- DNA-based MERS vaccine: Like the full-length S gene of SARS-CoV-1, DNA encoding full-length S protein of MERS-CoV is utilized to develop MERS vaccines [31,51]. Indeed, intramuscular injections of mice with a synthetic DNA encoding full-length S proteins of MERS-CoV elicited potent virus-neutralizing antibodies and cellular immune responses, as represented by the secretion of INF-γ, TNF-α, and/or IL-2 cytokines in CD4+ and/or CD8+ T cells, as well as the production of antibodies in immunized camels.
- Subunit MERS vaccines: Protein-based subunit vaccines against MERS-CoV have been developed [52,53,54]. While some are designed on the basis of the full-length S1 proteins [55], the majority of them are based on viral RBD [53,54,56,57]. These RBD-based vaccines are evaluated for immunogenicity and protective immunity in a number of MERS-CoV mice models. The antigenicity and functionality of these RBD proteins have also been extensively investigated. In general, subunit vaccines might not induce immune responses as strong as those induced by other vaccine types mentioned above. However, the immunogenicity of subunit vaccines could be significantly promoted in the presence of an ideal adjuvant via an appropriate route [52]. In addition, it is also essential to maintain a suitable conformation of the protein antigen in the vaccine, such as the MERS-CoV RBD proteins [53,56]. In terms of safety consideration, subunit vaccines should be accounted as the safest vaccine type. They do not contain viral genetic materials, but only include essential antigens for eliciting protective immune responses, thus excluding the possibility of recovering virulence or inducing adverse reactions [58,59,60].
1.5. Vaccinations Proposed for SARS-CoV-2
2. Drug Research and Development: Promising Analogue to Patient
2.1. Drug Development and Discovery
2.2. Target Identification
- SAR defined;
- Drug ability (preliminary toxicity);
- Synthetic feasibility;
- In-Vitro assessment of drug resistance and efflux potential;
- Evidence of in vivo efficacy of chemical class;
- Pharmacokinetics of chemical entity.
2.3. Lead Optimization
2.4. Formulation and Development Process
2.5. Pre-Clinical Research
2.6. Clinical Research
2.6.1. Phase 0
2.6.2. Phase 1 (Safety and Dosage)
2.6.3. Phase 2 (Efficacy and Side Effects)
2.6.4. Phase 3 (Efficacy and Monitoring of Adverse Reactions)
2.6.5. Phase 4
3. FDA Review
- Proposed labeling;
- Safety updates;
- Drug abuse information;
- Patent information;
- Any data from studies that may have been conducted outside the United States;
- Institutional review board compliance information;
- Directions for use.
- Each member of the board conducts a full review of their section of the application.
- FDA instructors travel to clinical study sites to conduct an inspection of the facilities. The FDA looks for evidence of fabrication, manipulation or withholding of data.
- The project manager assigned will oversee all the individual reviews into a combined action package. The review team recommends a decision and a senior figure will make the final decision.
4. FDA Approval
5. FDA Post Market Drug Safety Monitoring
6. Conclusions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Jiang, F.; Deng, L.; Zhang, L.; Cai, Y.; Cheung, C.W.; Xia, Z. Review of the clinical characteristics of coronavirus disease 2019 (COVID-19). J. Gen. Intern. Med. 2020, 4, 1–5. [Google Scholar] [CrossRef] [PubMed]
- Pan, F.; Ye, T.; Sun, P.; Gui, S.; Liang, B.; Li, L.; Zheng, D.; Wang, J.; Hesketh, R.L.; Yang, L.; et al. Time Course of Lung Changes at Chest CT during Recovery from Coronavirus Disease 2019 (COVID-19). Radiology 2020, 295, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Zhu, N.; Zhang, D.; Wang, W.; Li, X.; Yang, B.; Song, J.; Zhao, X.; Huang, B.; Shi, W.; Lu, R.; et al. A Novel Coronavirus from Patients with Pneumonia in China, 2019. N. Eng. J. Med. 2020, 382, 727–733. [Google Scholar] [CrossRef] [PubMed]
- Davies, N.G.; Barnard, R.C.; Jarvis, C.I.; Kucharski, A.J.; Munday, J.; Pearson, C.A.B.; Russell, T.W.; Tully, D.C.; Abbott, S.; Gimma, A.; et al. Estimated Transmissibility and Severity of Novel SARS-CoV-2 Variant of Concern 202012/01 in England. 2020. Available online: https://cmmid.github.io/topics/covid19/reports/uk-novel-variant/2020_12_23_Transmissibility_and_severity_of_VOC_202012_01_in_England.pdf (accessed on 29 December 2020).
- Huang, C.; Wang, Y.; Li, X.; Ren, L.; Zhao, J.; Hu, Y.; Zhang, L.; Fan, G.; Xu, J.; Gu, X.; et al. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet 2020, 395, 497–506. [Google Scholar] [CrossRef]
- Oran, D.P.; Topol, E.J. Prevalence of Asymptomatic SARS-CoV-2 Infection. Ann. Intern. Med. 2020, 173, 362–367. [Google Scholar] [CrossRef]
- World Health Organization (WHO). Coronavirus Disease (COVID-19). 2020. Available online: https://www.who.int/emergencies/diseases/novel-coronavirus-2019/question-and-answers-hub/q-a-detail/coronavirus-disease-covid-19#:~:text=symptoms (accessed on 29 December 2020).
- Adhikari, S.P.; Meng, S.; Wu, Y.-J.; Mao, Y.-P.; Ye, R.-X.; Wang, Q.-Z.; Sun, C.; Sylvia, S.; Rozelle, S.; Raat, H.; et al. Epidemiology, causes, clinical manifestation and diagnosis, prevention and control of coronavirus disease (COVID-19) during the early outbreak period: A scoping review. Infect. Dis. Poverty 2020, 9, 1–12. [Google Scholar] [CrossRef]
- Xiao, F.; Tang, M.; Zheng, X.; Liu, Y.; Li, X.; Shan, H. Evidence for Gastrointestinal Infection of SARS-CoV-2. Gastroenterology 2020, 158, 1831–1833. [Google Scholar] [CrossRef]
- World Health Organization. Consensus Document on the Epidemiology of Severe Acute Respiratory Syndrome (SARS). 2003. Available online: https://www.who.int/csr/sars/WHOconsensus.pdf?ua=1 (accessed on 29 November 2020).
- Anthony, R. Fehr and Stanley Perlman; Coronaviruses: An Overview of Their Replication and Pathogenesis. Coronaviruses 2015, 1282, 1–23. [Google Scholar]
- Singhal, T. A Review of Coronavirus Disease-2019 (COVID-19). Indian J. Pediatr. 2020, 87, 281–286. [Google Scholar] [CrossRef]
- Richman, D.; Whitley, R.J.; Hayden, F.G. Infectious Disease in the Ageing: A Clinical Handbook, 3rd ed.; Springer: Berlin/Heidelberg, Germany, 2009; Volume 12, p. 1692. [Google Scholar]
- Liu, Y.; Gayle, A.A.; Wilder-Smith, A.; Rocklöv, J. The reproductive number of COVID-19 is higher compared to SARS coronavirus. J. Travel Med. 2020, 27, 1–4. [Google Scholar] [CrossRef]
- Ullah, H.; Ullah, A.; Gull, A.; Mousavi, T.; Khan, M.W. Novel Coronavirus 2019 (COVID-19) Pandemic Outbreak: A Comprehensive Review of the Current Literature. Vacunas 2020, 20, 1–24. [Google Scholar] [CrossRef] [PubMed]
- Corley, J.U.S. Government Response To COVID-19 Was Slow. But How Does It Compare To Other Countries? 2020. Available online: https://www.forbes.com/sites/jacquelyncorley/2020/04/10/us-government-response-to-covid-19-was-slow-but-how-does-it-compare-to-other-countries/?sh=515dc7a46dc2 (accessed on 29 November 2020).
- Coronavirus Disease (COVID-19) Advice for the Public. 2020. Available online: https://www.who.int/emergencies/diseases/novel-coronavirus-2019/advice-for-public (accessed on 29 November 2020).
- Agarwal, K.M.; Mohapatrab, S.; Sharmaa, P.; Sharmac, S.; Bhatia, D.; Mishrae, A. Study and overview of the novel corona virus disease (COVID-19). Sens. Int. 2020, 1, 100037. [Google Scholar] [CrossRef]
- Li, F. Structure, Function, and Evolution of Coronavirus Spike Proteins. Annu. Rev. Virol. 2016, 3, 237–261. [Google Scholar] [CrossRef] [PubMed]
- Mousavizadeh, L.; Ghasemi, S. Genotype and phenotype of COVID-19: Their roles in pathogenesis. J. Microbiol. Immunol. Infect. 2020, 52, 1–6. [Google Scholar] [CrossRef]
- Snijder, E.J.; Decroly, E.; Ziebuhr, J. The Nonstructural Proteins Directing Coronavirus RNA Synthesis and Processing. Adv. Clin. Chem. 2016, 96, 59–126. [Google Scholar] [CrossRef]
- Shereen, M.A.; Khan, S.; Kazmi, A.; Bashir, N.; Siddique, R. COVID-19 infection: Origin, transmission, and characteristics of human coronaviruses. J. Adv. Res. 2020, 24, 91–98. [Google Scholar] [CrossRef]
- Lai, C.C.; Shih, T.P.; Ko, W.C.; Tang, H.J.; Hsueh, P.R. Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) and coronavirus disease-2019 (COVID-19): The epidemic and the challenges. Int. J. Antimicrob. Agents 2020, 55, 105924. [Google Scholar] [CrossRef]
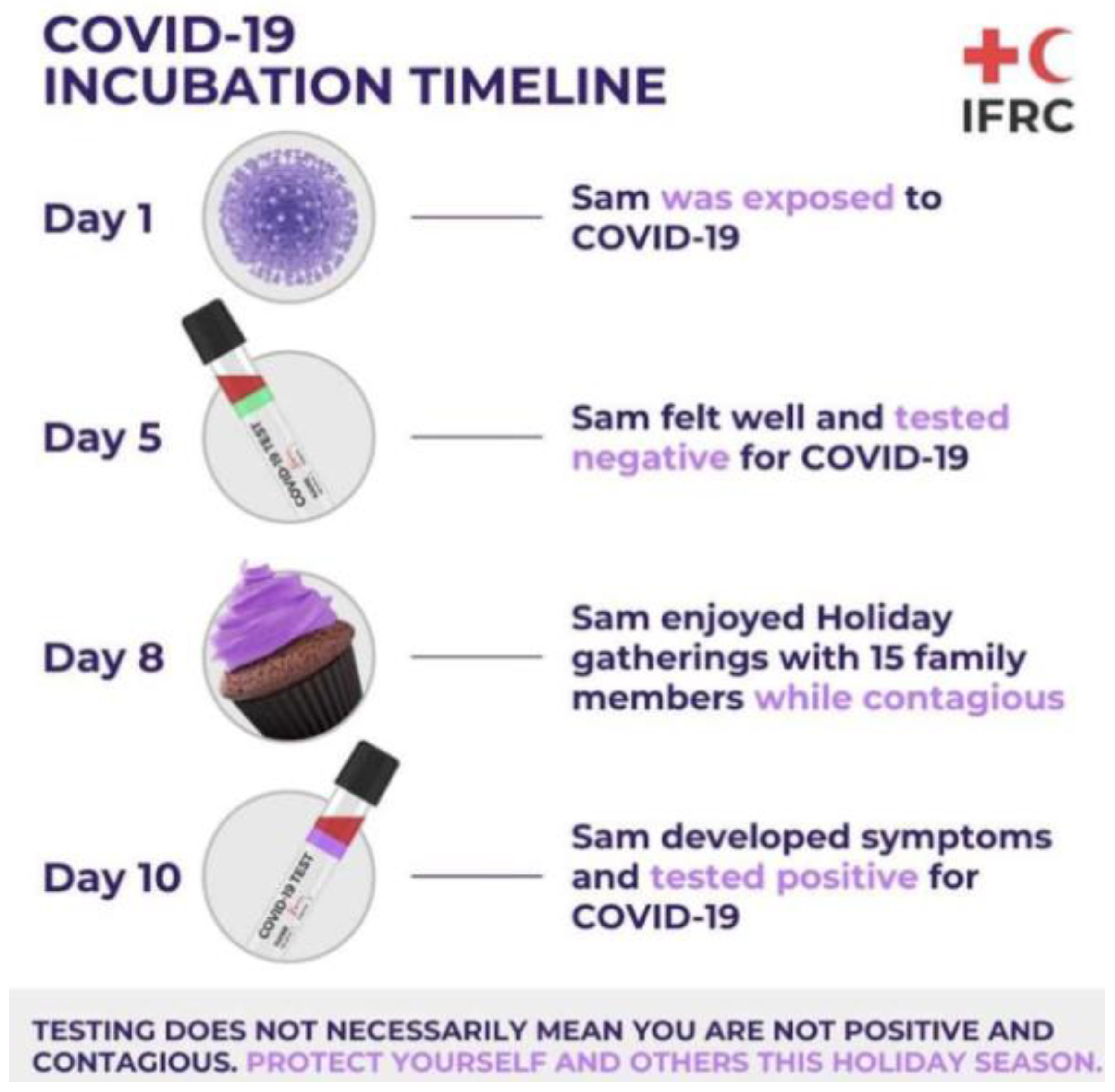
- Binnicker, M.J. Challenges and Controversies to Testing for COVID-19. J. Clin. Microbiol. 2020, 58, e01695-20. [Google Scholar] [CrossRef]
- IFRC. COVID-19 Symptoms. 2020. Available online: https://media.ifrc.org/ifrc/ (accessed on 1 January 2021).
- Williams, S. A Brief History of Human Coronaviruses. 2020. Available online: https://www.the-scientist.com/news-opinion/a-brief-history-of-human-coronaviruses-67600 (accessed on 29 November 2020).
- Peiris, J.S.M.; Guan, Y.; Yuen, K.Y. Severe acute respiratory syndrome. Nat. Med. 2004, 10, S88–S97. [Google Scholar] [CrossRef]
- Xiong, S.; Wang, Y.-F.; Zhang, M.-Y.; Liu, X.-J.; Zhang, C.-H.; Liu, S.-S.; Qian, C.-W.; Li, J.-X.; Lu, J.-H.; Wan, Z.-Y.; et al. Immunogenicity of SARS inactivated vaccine in BALB/c mice. Immunol. Lett. 2004, 95, 139–143. [Google Scholar] [CrossRef]
- He, Y.; Zhou, Y.; Siddiqui, P.; Jiang, S. Inactivated SARS-CoV vaccine elicits high titers of spike protein-specific antibodies that block receptor binding and virus entry. Biochem. Biophys. Res. Commun. 2004, 325, 445–452. [Google Scholar] [CrossRef]
- Qu, D.; Zheng, B.; Yao, X.; Guan, Y.; Yuan, Z.-H.; Zhong, N.-S.; Lu, L.-W.; Xie, J.-P.; Wen, Y.-M. Intranasal immunization with inactivated SARS-CoV (SARS-associated coronavirus) induced local and serum antibodies in mice. Vaccine 2005, 23, 924–931. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z.-Y.; Kong, W.-P.; Huang, Y.; Roberts, A.; Murphy, B.R.; Subbarao, K.; Nabel, G.J. A DNA vaccine induces SARS coronavirus neutralization and protective immunity in mice. Nature 2004, 428, 561–564. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Chou, T.-H.W.; Sakhatskyy, P.V.; Huang, S.; Lawrence, J.M.; Cao, H.; Huang, X.; Lu, S. Identification of Two Neutralizing Regions on the Severe Acute Respiratory Syndrome Coronavirus Spike Glycoprotein Produced from the Mammalian Expression System. J. Virol. 2005, 79, 1906–1910. [Google Scholar] [CrossRef] [PubMed]
- Keng, C.-T.; Zhang, A.; Shen, S.; Lip, K.-M.; Fielding, B.C.; Tan, T.H.P.; Chou, C.-F.; Loh, C.B.; Wang, S.; Fu, J.; et al. Amino Acids 1055 to 1192 in the S2 Region of Severe Acute Respiratory Syndrome Coronavirus S Protein Induce Neutralizing Antibodies: Implications for the Development of Vaccines and Antiviral Agents. J. Virol. 2005, 79, 3289–3296. [Google Scholar] [CrossRef]
- Zhong, X.; Yang, H.; Guo, Z.-F.; Sin, W.-Y.F.; Chen, W.; Xu, J.; Fu, L.; Wu, J.; Mak, C.-K.G.; Cheng, C.-S.S.; et al. B-Cell Responses in Patients Who Have Recovered from Severe Acute Respiratory Syndrome Target a Dominant Site in the S2 Domain of the Surface Spike Glycoprotein. J. Virol. 2005, 79, 3401–3408. [Google Scholar] [CrossRef]
- Bisht, H.; Roberts, A.; Vogel, L.; Bukreyev, A.; Collins, P.L.; Murphy, B.R.; Subbarao, K.; Moss, B. Severe acute respiratory syndrome coronavirus spike protein expressed by attenuated vaccinia virus protectively immunizes mice. Proc. Natl. Acad. Sci. USA 2004, 101, 6641–6646. [Google Scholar] [CrossRef]
- Bukreyev, A.; Lamirande, E.W.; Buchholz, U.J.; Vogel, L.N.; Elkins, W.R.; Claire, M.S.; Murphy, B.R.; Subbarao, K.; Collins, P.L. Mucosal immunisation of African green monkeys (Cercopithecus aethiops) with an attenuated parainfluenza virus expressing the SARS coronavirus spike protein for the prevention of SARS. Lancet 2004, 363, 2122–2127. [Google Scholar] [CrossRef]
- Jiang, S.; He, Y.; Liu, S. SARS Vaccine Development. Emerg. Infect. Dis. 2005, 11, 1016–1020. [Google Scholar] [CrossRef]
- He, Y.; Zhu, Q.; Liu, S.; Zhou, Y.; Yang, B.; Li, J.; Jiang, S. Identification of a critical neutralization determinant of severe acute respiratory syndrome (SARS)-associated coronavirus: Importance for designing SARS vaccines. Virology 2005, 334, 74–82. [Google Scholar] [CrossRef]
- Chen, Z.; Zhang, L.; Qin, C.; Ba, L.; Yi, C.E.; Zhang, F.; Wei, Q.; He, T.; Yu, W.; Yu, J.; et al. Recombinant Modified Vaccinia Virus Ankara Expressing the Spike Glycoprotein of Severe Acute Respiratory Syndrome Coronavirus Induces Protective Neutralizing Antibodies Primarily Targeting the Receptor Binding Region. J. Virol. 2005, 79, 2678–2688. [Google Scholar] [CrossRef] [PubMed]
- Du, L.; Zhao, G.; He, Y.; Guo, Y.; Zheng, B.-J.; Jiang, S.; Zhou, Y. Receptor-binding domain of SARS-CoV spike protein induces long-term protective immunity in an animal model. Vaccine 2007, 25, 2832–2838. [Google Scholar] [CrossRef] [PubMed]
- He, Y.; Lu, H.; Siddiqui, P.; Zhou, Y.; Jiang, S. Receptor-Binding Domain of Severe Acute Respiratory Syndrome Coronavirus Spike Protein Contains Multiple Conformation-Dependent Epitopes that Induce Highly Potent Neutralizing Antibodies. J. Immunol. 2005, 174, 4908–4915. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Moore, M.J.; Vasilieva, N.; Sui, J.; Wong, S.K.; Berne, M.A.; Somasundaran, M.; Sullivan, J.L.; Luzuriaga, K.; Greenough, T.C.; et al. Angiotensin-converting enzyme 2 is a functional receptor for the SARS coronavirus. Nature 2003, 426, 450–454. [Google Scholar] [CrossRef] [PubMed]
- Wong, S.K.; Li, W.; Moore, M.J.; Choe, H.; Farzan, M. A 193-Amino Acid Fragment of the SARS Coronavirus S Protein Efficiently Binds Angiotensin-converting Enzyme 2. J. Biol. Chem. 2004, 279, 3197–3201. [Google Scholar] [CrossRef] [PubMed]
- Xiao, X.; Chakraborti, S.; Dimitrov, A.S.; Gramatikoff, K.; Dimitrov, D.S. The SARS-CoV S glycoprotein: Expression and functional characterization. Biochem. Biophys. Res. Commun. 2003, 312, 1159–1164. [Google Scholar] [CrossRef] [PubMed]
- Du, L.; Tai, W.; Zhou, Y.; Jiang, S. Vaccines for the prevention against the threat of MERS-CoV. Expert Rev. Vaccines 2017, 15, 1123–1134. [Google Scholar] [CrossRef] [PubMed]
- Haagmans, B.L.; Brand, J.M.A.V.D.; Raj, V.S.; Volz, A.; Wohlsein, P.; Smits, S.L.; Schipper, D.; Bestebroer, T.M.; Okba, N.; Fux, R.; et al. An orthopoxvirus-based vaccine reduces virus excretion after MERS-CoV infection in dromedary camels. Science 2016, 351, 77–81. [Google Scholar] [CrossRef]
- Kim, E.; Okada, K.; Kenniston, T.; Raj, V.S.; Alhajri, M.M.; Farag, E.A.; Alhajri, F.; Osterhaus, A.D.; Haagmans, B.L.; Gambotto, A. Immunogenicity of an adenoviral-based Middle East Respiratory Syndrome coronavirus vaccine in BALB/c mice. Vaccine 2014, 32, 5975–5982. [Google Scholar] [CrossRef]
- Song, F.; Fux, R.; Provacia, L.; Volz, A.; Eickmann, M.; Becker, S.; Osterhaus, A.; Haagmans, B.; Suttera, G. Middle East Respiratory Syndrome Coronavirus Spike Protein Delivered by Modified Vaccinia Virus Ankara Efficiently Induces Virus-Neutralizing Antibodies. J. Virol. 2013, 87, 11950–11954. [Google Scholar] [CrossRef]
- Volz, A.; Kupke, A.; Song, F.; Jany, S.; Fux, R.; Shams-Eldin, H.; Schmidt, J.C.; Becker, C.; Eickmann, M.; Becker, S.; et al. Protective Efficacy of Recombinant Modified Vaccinia Virus Ankara Delivering Middle East Respiratory Syndrome Coronavirus Spike Glycoprotein. J. Virol. 2015, 89, 8651–8656. [Google Scholar] [CrossRef] [PubMed]
- Guo, X.; Deng, Y.; Chen, H.; Lan, J.; Wang, W.; Zou, X.; Hung, T.; Lu, Z.; Tan, W. Systemic and mucosal immunity in mice elicited by a single immunization with human adenovirus type 5 or 41 vector-based vaccines carrying the spike protein of Middle East respiratory syndrome coronavirus. Immunology 2015, 145, 476–484. [Google Scholar] [CrossRef] [PubMed]
- Muthumani, K.; Falzarano, D.; Reuschel, E.L.; Tingey, C.; Flingai, S.; Villarreal, D.O.; Wise, M.C.; Patel, A.; Izmirly, A.; Aljuaid, A.; et al. A synthetic consensus anti–spike protein DNA vaccine induces protective immunity against Middle East respiratory syndrome coronavirus in nonhuman primates. Sci. Transl. Med. 2015, 7, 301ra132. [Google Scholar] [CrossRef] [PubMed]
- Ma, C.; Li, Y.; Wang, L.; Zhao, G.; Tao, X.; Tseng, C.-T.K.; Zhou, Y.; Du, L.; Jiang, S. Intranasal vaccination with recombinant receptor-binding domain of MERS-CoV spike protein induces much stronger local mucosal immune responses than subcutaneous immunization: Implication for designing novel mucosal MERS vaccines. Vaccine 2014, 32, 2100–2108. [Google Scholar] [CrossRef]
- Ma, C.; Wang, L.; Tao, X.; Zhang, N.; Yang, Y.; Tseng, C.-T.K.; Li, F.; Zhou, Y.; Jiang, S.; Du, L. Searching for an ideal vaccine candidate among different MERS coronavirus receptor-binding fragments—The importance of immunofocusing in subunit vaccine design. Vaccine 2014, 32, 6170–6176. [Google Scholar] [CrossRef]
- Lan, J.; Deng, Y.; Chen, H.; Lu, G.; Wang, W.; Guo, X.; Lu, Z.; Gao, G.F.; Tan, W. Tailoring Subunit Vaccine Immunity with Adjuvant Combinations and Delivery Routes Using the Middle East Respiratory Coronavirus (MERS-CoV) Receptor-Binding Domain as an Antigen. PLoS ONE 2014, 9, e112602. [Google Scholar] [CrossRef]
- Wang, L.; Shi, W.; Joyce, M.G.; Modjarrad, K.; Zhang, Y.; Leung, K.; Lees, C.R.; Zhou, T.; Yassine, H.M.; Kanekiyo, M.; et al. Evaluation of candidate vaccine approaches for MERS-CoV. Nat. Commun. 2015, 6, 7712. [Google Scholar] [CrossRef]
- Du, L.; Kou, Z.; Ma, C.; Tao, X.; Wang, L.; Zhao, G.; Chen, Y.; Yu, F.; Tseng, C.-T.K.; Zhou, Y.; et al. A Truncated Receptor-Binding Domain of MERS-CoV Spike Protein Potently Inhibits MERS-CoV Infection and Induces Strong Neutralizing Antibody Responses: Implication for Developing Therapeutics and Vaccines. PLoS ONE 2013, 8, e81587. [Google Scholar] [CrossRef]
- Mou, H.; Raj, V.; Van Kuppeveld, F.; Rottier, P.; Haagmans, B.; Bosch, B.J. The Receptor Binding Domain of the New Middle East Respiratory Syndrome Coronavirus Maps to a 231-Residue Region in the Spike Protein That Efficiently Elicits Neutralizing Antibodies. J. Virol. 2013, 87, 9379–9383. [Google Scholar] [CrossRef]
- Chlibek, R.; Pauksens, K.; Rombo, L.; Van Rijckevorsel, G.; Richardus, J.H.; Plassmann, G.; Schwarz, T.F.; Catteau, G.; Lal, H.; Heineman, T.C. Long-term immunogenicity and safety of an investigational herpes zoster subunit vaccine in older adults. Vaccine 2016, 34, 863–868. [Google Scholar] [CrossRef]
- Berkowitz, E.M.; Moyle, G.; Stellbrink, H.-J.; Schürmann, D.; Kegg, S.; Stoll, M.; El Idrissi, M.; Oostvogels, L.; Heineman, T.C.; Brockmeyer, N.; et al. Safety and Immunogenicity of an Adjuvanted Herpes Zoster Subunit Candidate Vaccine in HIV-Infected Adults: A Phase 1/2a Randomized, Placebo-Controlled Study. J. Infect. Dis. 2015, 211, 1279–1287. [Google Scholar] [CrossRef] [PubMed]
- Sindoni, D.; La Fauci, V.; Squeri, R.; Cannavò, G.; Bacilieri, S.; Panatto, D.; Gasparini, R.; Amicizia, D. Comparison between a conventional subunit vaccine and the MF59-adjuvanted subunit influenza vaccine in the elderly: An evaluation of the safety, tolerability and immunogenicity. J. Prev. Med. Hyg. 2009, 50, 121–126. [Google Scholar] [PubMed]
- About the Oxford COVID-19 Vaccine. 2020. Available online: https://www.research.ox.ac.uk/Article/2020-07-19-the-oxford-covid-19-vaccine (accessed on 4 January 2021).
- Convidicea (Ad5-nCoV) Vaccine Description. 2020. Available online: https://www.precisionvaccinations.com/vaccines/convidicea-ad5-ncov-vaccine (accessed on 4 January 2021).
- Pfizer and BioNTech Choose Lead mRNA Vaccine Candidate against COVID-19 and Commence Pivotal Phase 2/3 Global Study. 2020. Available online: https://www.pfizer.com/news/press-release/press-release-detail/pfizer-and-biontech-choose-lead-mrna-vaccine-candidate-0 (accessed on 4 January 2021).
- Sinovac’s Coronavac™, SARS-CoV-2 Vaccine (Vero Cell), Inactivated, Announces Approval for Phase I/II Clinical Trial in Adolescents and Children. 2020. Available online: https://www.biospace.com/article/releases/sinovac-s-coronavac-sars-cov-2-vaccine-vero-cell-inactivated-announces-approval-for-phase-i-ii-clinical-trial-in-adolescents-and-children/ (accessed on 4 January 2021).
- Jackson, I.A.; Anderson, E.J.; Rouphael, N.G.; Roberts, P.C.; Makhene, M.; Coler, R.N.; McCullough, M.P.; Chappell, J.D.; Denison, M.R.; Stevens, L.J.; et al. An mRNA Vaccine against SARS-CoV-2—Preliminary Report. N. Eng. J. Med. 2020, 1, 1920–1931. [Google Scholar] [CrossRef] [PubMed]
- Ad26.COV2-S (Janssen COVID-19) Vaccine. 2020. Available online: https://www.precisionvaccinations.com/vaccines/ad26cov2-s-janssen-covid-19-vaccine (accessed on 4 January 2021).
- Moffat, J.G.; Vincent, F.; Lee, J.A.; Eder, J.; Prunotto, M. Opportunities and challenges in phenotypic drug discovery: An industry perspective. Nat. Rev. Drug Discov. 2017, 16, 531–543. [Google Scholar] [CrossRef]
- DiMasi, J.A.; Hansen, R.W.; Grabowski, H.G. The price of innovation: New estimates of drug development costs. J. Health Econ. 2003, 22, 151–185. [Google Scholar] [CrossRef]
- Lindsay, M.A. Target discovery. Nat. Rev. Drug Discov. 2003, 2, 831–838. [Google Scholar] [CrossRef]
- Imming, P.; Sinning, C.; Meyer, A. Drugs, their targets and the nature and number of drug targets. Nat. Rev. Drug Discov. 2006, 5, 821–834. [Google Scholar] [CrossRef]
- Patidar, A.K.; Selvam, G.; Jeyakanda, M.; Mobiya, A.K.; Bagherwal, A.; Sanadya, G.; Mehta, R. Lead Discovery and Lead Optimization: A Useful Strategy in Molecular Modification of Lead Compound in Analog Design. Int. J. Drug Design Discov. 2011, 2, 458–463. [Google Scholar]
- Huber, W. A new strategy for improved secondary screening and lead optimization using high-resolution SPR characterization of compound-target interactions. J. Mol. Recognit. 2005, 18, 273–281. [Google Scholar] [CrossRef]
- Faqi, A.S. A Comprehensive Guide to Toxicology in Preclinical Drug Development, 2nd ed.; Elsevier: Detroit, MI, USA, 2012. [Google Scholar]
- Honek, J. Preclinical Research in Drug Development; Honek Communications Consulting & Medical Writing: Stockholm, Sweden, 2017; Volume 26, pp. 6–8. [Google Scholar]
- FDA. Step 3: Clinical Research. 2018. Available online: https://www.fda.gov/patients/drug-development-process/step-3-clinical-research (accessed on 30 November 2020).

| Vaccine Candidate, Developers | Technology Used | Current Phase | Completed Phase (Findings) | Clinical Trial Sites |
|---|---|---|---|---|
| AZD1222 University of Oxford, AstraZeneca [61] | Modified chimp adenovirus vector (ChAdOx1) | Phase III (30,000) Interventional; randomized, placebo-controlled study for efficacy, safety, and immunogenicity. Brazil (5000) International enrolment of the Phase III trial was paused on 8 September 2020, due to an adverse neurological event in one participant, but resumed on 12 September in the UK. On 23 October, AstraZeneca said it will resume the trial in the US | Phase I-II (543) Spike-specific antibodies at day 28; neutralizing antibodies after a booster dose at day 56. Adverse effects: pain at the injection site, headache, fever, chills, muscle ache, malaise in more than 60% of participants; paracetamol allowed for some participants to increase tolerability | 20 in the UK, São Paulo |
| Ad5-nCoV CanSinoBIO, Beijing Institute of Biotechnology of the Academy of Military Medical Sciences [62] | Recombinant adenovirus type 5 vector | Phase III (40,000) global multi-center, randomized, double-blind, placebo-controlled to evaluate efficacy, safety and immunogenicity in Mexico, Pakistan, Russia, Saudi Arabia | Phase II (508) Neutralizing antibody and T cell responses. Adverse effects: moderate over 7 days: 74% had fever, pain, fatigue | China and Pakistan |
| BNT162b2 BioNTech, Fosun Pharma, Pfizer [63] | mRNA | Phase III (30,000) Randomized, placebo-controlled | Phase I-II (45) Strong RBD-binding IgG and neutralizing antibody response peaked 7 days after a booster dose, robust CD4+ and CD8+ T cell responses, undetermined durability. Adverse effects: dose-dependent and moderate including pain at the injection site, fatigue, headache, chills, muscle and joint pain, fever | 62 in the USA and Germany |
| CoronaVac Sinovac [64] | Inactivated SARS-CoV-2 | Phase III (33,620) Double-blind, randomized, placebo-controlled to evaluate efficacy and safety in Brazil (15,000); Chile (3000); Indonesia (1620); Turkey (13,000) Brazil paused Phase III trials on November 10 after the suicide of a volunteer in the trials before resuming them on November 11. | Phase II (600) Preprint. Immunogenicity eliciting 92% seroconversion at lower dose; Adverse effects: mild in severity, pain at injection site | 2 in China; 22 in Brazil; Bandung, Indonesia |
| mRNA-1273 Moderna, NIAID, BARDA [65] | Lipid nanoparticle dispersion containing mRNA | Phase III (30,000) Interventional; randomized, placebo-controlled study for efficacy, safety, and immunogenicity | Phase I (45) Dose-dependent neutralizing antibody response on two-dose schedule; undetermined durability. Adverse effects: fever, fatigue, headache, muscle ache, and pain at the injection site | 89 sites in the USA |
| Ad26.COV2.S Janssen Pharmaceuticals (Johnson and Johnson), BIDMC [66] | Non-replicating viral vector | Phase III (60,000) Randomized, double-blinded, placebo-controlled Temporarily paused on 13 October 2020, due to an unexplained illness in a participant. Johnson and Johnson announced, on 23 October, that they are preparing to resume the trial in the US. | Phase I-II (1045) Preprint. Seroconversion for S antibodies over 95%. Adverse effects: injection site pain, fatigue, headache and myalgia | 291 in US, Argentina, Brazil, Chile, Colombia, Mexico, Peru, Philippines, South Africa and Ukraine |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Daou, A. COVID-19 Vaccination: From Interesting Agent to the Patient. Vaccines 2021, 9, 120. https://doi.org/10.3390/vaccines9020120
Daou A. COVID-19 Vaccination: From Interesting Agent to the Patient. Vaccines. 2021; 9(2):120. https://doi.org/10.3390/vaccines9020120
Chicago/Turabian StyleDaou, Anis. 2021. "COVID-19 Vaccination: From Interesting Agent to the Patient" Vaccines 9, no. 2: 120. https://doi.org/10.3390/vaccines9020120
APA StyleDaou, A. (2021). COVID-19 Vaccination: From Interesting Agent to the Patient. Vaccines, 9(2), 120. https://doi.org/10.3390/vaccines9020120






