A Targeted Radiotheranostic Agent for Glioblastoma: [64Cu]Cu-NOTA-TP-c(RGDfK)
Abstract
1. Introduction
2. Materials and Methods
2.1. Chemistry
2.1.1. General
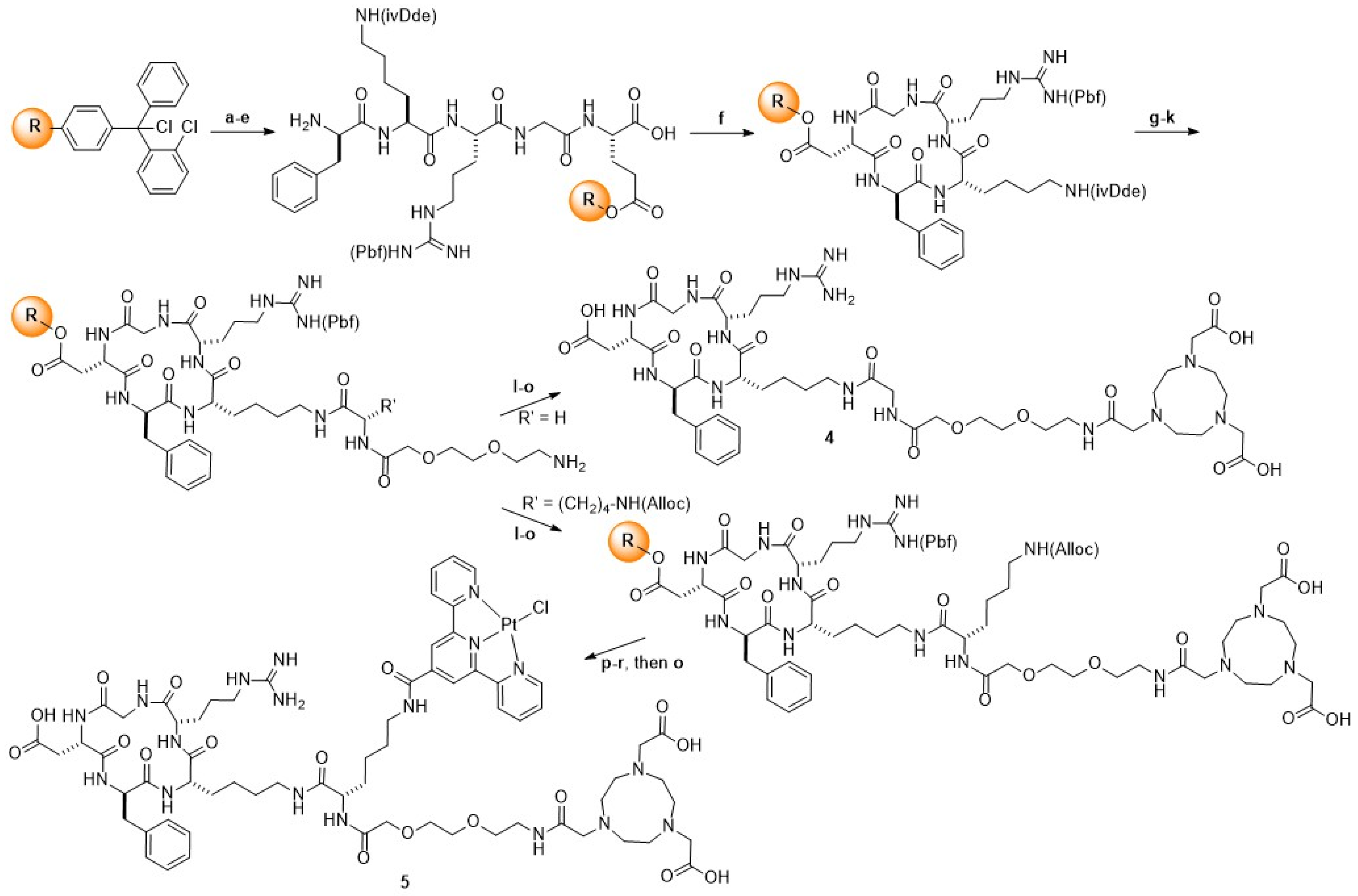
2.1.2. Synthesis of NOTA-c(RGDfK) 4 and NOTA-TP-c(RGDfK) 5
2.1.3. Synthesis of NOTA-TP
2.1.4. Synthesis of natCu-NOTA-c(RGDfK) and natCu-NOTA-TP-c(RGDfK)
2.1.5. Radiolabeling of NOTA-Conjugates
2.2. Stability Studies
Plasma Stability and Protein Binding
2.3. Cellular Assays
2.3.1. Cell Culture
2.3.2. Competition Assay
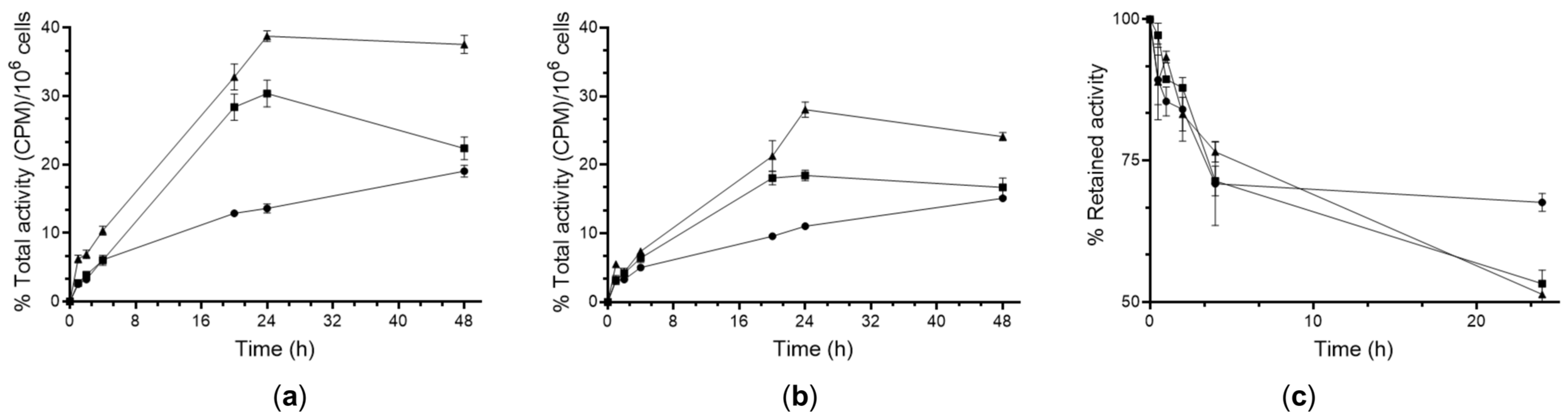
2.3.3. Cellular Uptake, Internalization, and Efflux Kinetics of the 64Cu-NOTA Conjugates
2.3.4. Comparative Subcellular Localization of [64Cu]Cu-NOTA-TP-c(RGDfK) in U87 MG Versus SVG p12 Cells
2.3.5. Cytotoxicity Assay
2.4. Data Analysis
3. Results
3.1. Synthesis and Radiolabeling
3.2. Stability Studies
3.3. Cellular Assays
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| CH3CN | Acetonitrile |
| CPM | Count per minute |
| c(RGDfK | Cyclic arginine-glycine-aspartate-D-phenylalanine-lysine |
| CRT | Chemoradiotheranostic |
| 64Cu | Copper-64 |
| 64Cu(OAc)2 | 64Cu-Acetate |
| 64CuCl2 | 64Cu-Chloride |
| DCM | Dichlorometane |
| DIPEA | N,N-Diisopropylethylamine |
| DMEM | Dulbecco’s modified Eagle medium |
| DMF | N,N-Dimethylformamide |
| DNA | Deoxyribonucleic acid |
| EC50 | Effective concentration |
| EMEM | Eagle’s minimal essential medium |
| Equiv | Equivalent |
| ESI | Electrospray ionization |
| Fmoc | Fluorenylmethyloxycarbonyl |
| GBM | Glioblastoma multiforme |
| H2O | Water |
| HATU | Dimethylamino-N,N-dimethyl(3H-[1,2,3]triazolo[4,5-b]pyridin-3-yloxy)methaniminium hexafluorophosphate |
| HOBt | Hydroxybenzotriazole |
| HPLC | High performance liquid chromatography |
| HRMS | High-resolution mass spectrometry |
| IC50 | Inhibitory concentration |
| iTLC | Instant thin layer chromatography |
| ivDde | 1-(4,4-Dimethyl-2,6-dioxocyclohex-1-ylidene)-3-methylbutyl |
| MeOH | Methanol |
| MG | Malignant glioblastoma |
| MS | Mass spectrometry |
| 64Ni | Nickel-64 |
| NOTA | 2,2′,2″-(1,4,7-Triazacyclononane-1,4,7-triyl)triacetic acid |
| PET | Positron Emission Tomography |
| Pd(PPh3)4 | Tetrakis(triphenylphosphine)palladium(0) |
| PhSiH3, | Phenylsilane |
| PyBOP | Benzotriazol-1-yloxytripyrrolidinophosphonium hexafluorophosphate |
| RGD | Arginine-glycine-aspartate |
| Rt | Retention time |
| TACN | Triazacyclononane |
| TFA | Trifluoroacetic acid |
| TIPS | Triisopropylsilane |
| TMZ | Temozolomide |
| TP | Terpyridine platinum |
| μM | Micromolar |
References
- Delgado-Martín, B.; Medina, M.A. Advances in the Knowledge of the Molecular Biology. Adv. Sci. 2020, 7, 1902971. [Google Scholar] [CrossRef]
- Déry, L.; Charest, G.; Guérin, B.; Akbari, M.; Fortin, D. Chemoattraction of Neoplastic Glial Cells with CXCL10, CCL2 and CCL11 as a Paradigm for a Promising Therapeutic Approach for Primary Brain Tumors. Int. J. Mol. Sci. 2021, 22, 12150. [Google Scholar] [CrossRef] [PubMed]
- Ostrom, Q.T.; Gittleman, H.; Liao, P.; Rouse, C.; Chen, Y.; Dowling, J.; Wolinsky, Y.; Kruchko, C.; Barnholtz-Sloan, J.S.; Xu, J.; et al. CBTRUS Statistical Report: Primary Brain and Central Nervous System Tumors Diagnosed in the United States in 2007–2011. Neuro-Oncology 2014, 16 (Suppl. S4), iv1–iv63. [Google Scholar] [CrossRef]
- Mathew, E.N.; Berry, B.C.; Yang, H.W.; Carroll, R.S.; Johnson, M.D. Delivering Therapeutics to Glioblastoma: Overcoming Biological Constraints. Int. J. Mol. Sci. 2022, 23, 1711. [Google Scholar] [CrossRef]
- Liu, Y.; Zhan, Z.; Kang, Z.; Li, M.; Lv, Y.; Li, S.; Tong, L.; Feng, F.; Li, Y.; Zhang, M.; et al. Preclinical and early clinical studies of a novel compound SYHA1813 that efficiently crosses the blood–brain barrier and exhibits potent activity against glioblastoma. Acta Pharm. Sin. B 2023, 13, 4748–4764. [Google Scholar] [CrossRef]
- Stupp, R.; Mason, W.P.; Van den Bent, M.J.; Weller, M.; Fisher, B.; Taphoorn, M.J.B.; Belanger, K.; Brandes, A.A.; Marosi, C.; Bogdahn, U.; et al. Radiotherapy plus Concomitant and Adjuvant Temozolomide for Glioblastoma. N. Engl. J. Med. 2005, 352, 987–996. [Google Scholar] [CrossRef]
- Wu, W.; Klockow, J.L.; Zhang, M.; Lafortune, F.; Chang, E.; Jin, L.; Wu, Y.; Daldrup-Link, H.E. Glioblastoma multiforme (GBM): An overview of current therapies and mechanisms of resistance. Pharmacol. Res. 2021, 171, 105780. [Google Scholar] [CrossRef]
- Roth, P. The role of integrins in glioma biology and anti-glioma therapies. SpringerPlus 2015, 4 (Suppl. S1), L12. [Google Scholar] [CrossRef][Green Version]
- Takada, Y.; Ye, X.; Simon, S. The integrins. Genome Biol. 2007, 8, 215. [Google Scholar] [CrossRef] [PubMed]
- Echavidre, W.; Picco, V.; Faraggi, M.; Montemagno, C. Integrin-αvβ3 as a Therapeutic Target in Glioblastoma: Back to the Future? Pharmaceutics 2022, 14, 1053. [Google Scholar] [CrossRef] [PubMed]
- Shi, J.; Wang, F.; Liu, S. Radiolabeled cyclic RGD peptides as radiotracers for tumor imaging. Biophys. Rep. 2016, 2, 1–20. [Google Scholar] [CrossRef] [PubMed]
- Fu, S.; Xu, X.; Ma, Y.; Zhang, S.; Zhang, S. RGD peptide-based non-viral gene delivery vectors targeting integrin αvβ3 for cancer therapy. J. Drug Target. 2019, 27, 1–11. [Google Scholar] [CrossRef]
- Haubner, R.; Weber, W.A.; Beer, A.J.; Vabuliene, E.; Reim, D.; Sarbia, M.; Becker, K.-F.; Goebel, M.; Hein, R.; Wester, H.-J.; et al. Noninvasive visualization of the activated alphavbeta3 integrin in cancer patients by positron emission tomography and [18F]Galacto-RGD. PLoS Med. 2005, 2, e70. [Google Scholar] [CrossRef] [PubMed]
- Haubner, R.; Kuhnast, B.; Mang, C.; Weber, W.A.; Kessler, H.; Wester, H.-J.; Schwaiger, M. [18F]Galacto-RGD: Synthesis, Radiolabeling, Metabolic Stability, and Radiation Dose Estimates. Bioconjug. Chem. 2004, 15, 61–69. [Google Scholar] [CrossRef]
- Novy, Z.; Stepankova, J.; Hola, M.; Flasarova, D.; Popper, M.; Petrik, M. Preclinical Evaluation of Radiolabeled Peptides for PET Imaging of Glioblastoma Multiforme. Molecules 2019, 24, 2496. [Google Scholar] [CrossRef]
- Li, D.; Zhao, X.; Zhang, L.; Li, F.; Ji, N.; Gao, Z.; Wang, J.; Kang, P.; Liu, Z.; Shi, J.; et al. 68Ga-PRGD2 PET/CT in the Evaluation of Glioma: A Prospective Study. Mol. Pharm. 2014, 11, 3923–3929. [Google Scholar] [CrossRef]
- Zhang, S.; Wang, X.; Gao, X.; Chen, X.; Li, L.; Li, G.; Liu, C.; Miao, Y.; Wang, R.; Hu, K. Radiopharmaceuticals and their applications in medicine. Signal Transduct. Target. Ther. 2025, 10, 1. [Google Scholar] [CrossRef]
- Meng, L.; Fang, J.; Lin, X.; Zhuang, R.; Huang, L.; Li, Y.; Zhang, X.; Guo, Z. Development of radioligands with an albumin-binding moiety of 4-(P-Iodophenyl) butyric acid for theranostic applications. J. Control. Release 2025, 382, 113757. [Google Scholar] [CrossRef]
- Tippayamontri, T.; Kotb, R.; Sanche, L.; Paquette, B. New Therapeutic Possibilities of Combined Treatment of Radiotherapy with Oxaliplatin and its Liposomal Formulation, Lipoxal™, in Rectal Cancer Using Xenograft in Nude Mice. Anticancer Res. 2014, 34, 5303–5312. [Google Scholar] [PubMed]
- Tippayamontri, T.; Kotb, R.; Paquette, B.; Sanche, L. Synergism in concomitant chemoradiotherapy of cisplatin and oxaliplatin and their liposomal formulation in the human colorectal cancer HCT116 model. Anticancer Res. 2012, 32, 4395–4404. [Google Scholar] [PubMed]
- Tippayamontri, T.; Kotb, R.; Paquette, B.; Sanche, L. Efficacy of cisplatin and Lipoplatin™ in combined treatment with radiation of a colorectal tumor in nude mouse. Anticancer Res. 2013, 33, 3005–3014. [Google Scholar] [PubMed]
- Seiwert, T.Y.; Salama, J.K.; Vokes, E. The concurrent chemoradiation paradigm--general principles. Nat. Clin. Pract. Oncol. 2007, 4, 86–100. [Google Scholar] [CrossRef] [PubMed]
- Tippayamontri, T.; Kotb, R.; Paquette, B.; Sanche, L. Cellular Uptake and Cytoplasm/DNA Distribution of Cisplatin and Oxaliplatin and Their Liposomal Formulation in Human Colorectal Cancer Cell HCT116. Investig. New Drugs 2011, 29, 1321–1327. [Google Scholar] [CrossRef] [PubMed]
- Le, X.; Hanna, E.Y. Optimal regimen of cisplatin in squamous cell carcinoma of head and neck yet to be determined. Ann. Transl. Med. 2018, 6, 229. [Google Scholar] [CrossRef] [PubMed]
- Oun, R.; Moussa, Y.E.; Wheate, N.J. The side effects of platinum-based chemotherapy drugs: A review for chemists. Dalton trans. 2018, 47, 7848. [Google Scholar] [CrossRef]
- Shen, D.-W.; Pouliot, L.M.; Hall, M.D.; Gottesman, M.M. Cisplatin resistance: A cellular self-defense mechanism resulting from multiple epigenetic and genetic changes. Pharmacol. Rev. 2012, 64, 706–721. [Google Scholar] [CrossRef]
- Harper, B.W.J.; Aldrich-Wright, J.R. The synthesis, characterisation and cytotoxicity of bisintercalating (2,2′:6′,2′-terpyridine) platinum(ii) complexes. Dalton Trans. 2015, 44, 87–96. [Google Scholar] [CrossRef]
- Ndagi, U.; Mhlongo, N.; Soliman, M.E. Metal complexes in cancer therapy—An update from drug design perspective. Drug Des. Devel. Ther. 2017, 3, 599–616. [Google Scholar] [CrossRef]
- Stafford, V.S.; Suntharalingam, K.; Shivalingam, A.; White, A.J.P.; Mann, D.J.; Vilar, R. Syntheses of polypyridyl metal complexes and studies of their interaction with quadruplex DNA. Dalton Trans. 2014, 44, 3686–3700. [Google Scholar] [CrossRef]
- Savić, A.; Marzo, T.; Scaletti, F.; Massai, L.; Bartoli, G.; Hoogenboom, R.; Messori, L.; Van Deun, R.; Van Hecke, K. New platinum(II) and palladium(II) complexes with substituted terpyridine ligands: Synthesis and characterization, cytotoxicity and reactivity towards biomolecules. Biometals 2019, 32, 33–47. [Google Scholar] [CrossRef]
- Georgiades, S.N.; Abd Karim, N.H.; Suntharalingam, K.; Vilar, R. Interaction of metal complexes with G-quadruplex DNA. Angew. Chem. Int. Ed. 2010, 49, 4020–4034. [Google Scholar] [CrossRef]
- Di Antonio, M.; Ponjavic, A.; Radzevičius, A.; Ranasinghe, R.T.; Catalano, M.; Zhang, X.; Shen, J.; Needham, L.-M.; Lee, S.F.; Klenerman, D.; et al. Single-molecule visualization of DNA G-quadruplex formation in live cells. Nat. Chem. 2020, 12, 832–837. [Google Scholar] [CrossRef]
- Bertrand, H.; Bombard, S.; Monchaud, D.; Talbot, E.; Guédin, A.; Mergny, J.-L.; Grünert, R.; Bednarskid, P.J.; Teulade-Fichou, M.-P. Exclusive platination of loop adenines in the human telomeric G-quadruplex. Org. Biomol. Chem. 2009, 7, 2864–2871. [Google Scholar] [CrossRef]
- Khosravifarsani, M.; Ait-Mohand, S.; Paquette, B.; Sanche, L.; Guérin, B. High cytotoxic effect by combining copper-64 with a NOTA-terpyridine platinum conjugate. J. Med. Chem. 2021, 64, 6765–6776. [Google Scholar] [CrossRef]
- Khosravifarsani, M.; Ait-Mohand, S.; Paquette, B.; Sanche, L.; Guérin, B. Design, Synthesis, and Cytotoxicity Assessment of [64Cu]Cu-NOTA-Terpyridine Platinum Conjugate: A Novel Chemoradiotherapeutic Agent with Flexible Linker. Nanomaterials 2021, 11, 2154. [Google Scholar] [CrossRef] [PubMed]
- Khosravifarsani, M.; Ait-Mohand, S.; Paquette, B.; Sanche, L.; Guérin, B. In vivo behavior of [64Cu]NOTA-terpyridine platinum, a novel chemo-radio-theranostic agent for imaging, and therapy of colorectal cancer. Front. Med. 2022, 9, 975213. [Google Scholar] [CrossRef] [PubMed]
- de Paola, I. Investigation of the Best Conditions to Obtain c(RGDfK) Peptide on Solid Phase. Int. J. Pept. Res. Ther. 2011, 17, 39–45. [Google Scholar] [CrossRef]
- Guérin, B.; Ait-Mohand, S.; Tremblay, M.C.; Dumulon-Perreault, V.; Fournier, P.; Bénard, F. Total solid-phase synthesis of NOTA-functionalized peptides for PET imaging. Org Lett. 2010, 12, 280–283. [Google Scholar] [CrossRef]
- Hahn, E.M.; Estrada-Ortiz, N.; Han, J.; Ferreira, V.F.C.; Kapp, T.G.; Correia, J.D.G.; Casini, A.; Kühn, F.E. Functionalization of Ruthenium(II) Terpyridine Complexes with Cyclic RGD Peptides to Target Integrin Receptors in Cancer Cells. Eur. J. Inorg. Chem. 2016, 2017, 1667–1672. [Google Scholar] [CrossRef]
- Xu, M.; McCanna, D.J.; Sivak, J.G. Use of the viability reagent PrestoBlue in comparison with alamarBlue and MTT to assess the viability of human corneal epithelial cells. J. Pharmacol. Toxicol. Methods 2015, 71, 1–7. [Google Scholar] [CrossRef]
- Lo, W.-L.; Lo, S.-W.; Chen, S.-J.; Chen, M.-W.; Huang, Y.-R.; Chen, L.-C.; Chang, C.-H.; Li, M.-H. Molecular Imaging and Preclinical Studies of Radiolabeled Long-Term RGD Peptides in U-87 MG Tumor-Bearing Mice. Int. J. Mol. Sci. 2021, 22, 5459. [Google Scholar] [CrossRef] [PubMed]
- Chen, B.; He, X.-Y.; Yi, X.-Q.; Zhuo, R.-X.; Cheng, S.-X. Dual-Peptide-Functionalized Albumin-Based Nanoparticles with pH-Dependent Self-Assembly Behavior for Drug Delivery. ACS Appl. Mater. Interfaces 2015, 7, 15148–15153. [Google Scholar] [CrossRef]
- Spada, A.; Emami, J.; Tuszynski, J.A.; Lavasanifar, A. The Uniqueness of Albumin as a Carrier in Nanodrug Delivery. Mol. Pharm. 2021, 18, 1862–1894. [Google Scholar] [CrossRef]
- Aguzzi, M.S.; Fortugno, P.; Giampietri, C.; Ragone, G.; Capogrossi, M.C.; Facchiano, A. Intracellular targets of RGDS peptide in melanoma cells. Mol. Cancer 2010, 22, 84. [Google Scholar] [CrossRef]
- Watt, H.L.; Rachid, Z.; Jean-Claude, B.J. Receptor activation and inhibition in cellular response to chemotherapeutic combinational mimicries: The concept of divergent targeting. J. Neurooncol. 2010, 100, 345–361. [Google Scholar] [CrossRef]
- Wu, Y.; Zhang, X.; Xiong, Z.; Cheng, Z.; Fisher, D.R.; Liu, S.; Gambhir, S.S.; Chen, X. microPET imaging of glioma integrin αvβ3 expression using 64Cu-labeled tetrameric RGD peptide. J. Nucl. Med. 2005, 46, 1707–1718. [Google Scholar] [PubMed]
- Chen, X.; Hou, Y.; Tohme, M.; Park, R.; Khankaldyyan, V.; Gonzales-Gomez, I.; Bading, J.R.; Laug, W.E.; Conti, P.S. Pegylated Arg-Gly-Asp peptide: 64Cu labeling and PET imaging of brain tumor alphavbeta3-integrin expression. J. Nucl. Med. 2004, 45, 1776–1783. [Google Scholar] [PubMed]

| Entry | c(RGDfK) Ligand a | IC50 on U87 MG (nM) |
|---|---|---|
| 1 | natCu-NOTA-c(RGDfK) | 10.6 ± 3.2 |
| 2 | natCu-NOTA-TP-c(RGDfK) | 16.2 ± 6.1 |
| Entry | Compound | 24 h | 48 h | 72 h | |||
|---|---|---|---|---|---|---|---|
| U87 MG | SVG p12 | U87 MG | SVG p12 | U87 MG | SVG p12 | ||
| 1 | natCu-NOTA-TP-c(RGDfK) | 42 ± 4 | 75 ± 7 | 27 ± 4 | 40 ± 7 | 13 ± 7 | 35 ± 3 |
| 2 | [64Cu]Cu-NOTA-TP-c(RGDfK) | 0.017 ± 0.002 | >0.300 | 0.010 ± 0.002 | 0.089 ± 0.008 | 0.007 ± 0.002 | 0.040 ± 0.005 |
| 3 | [64Cu]Cu-NOTA-TP | 0.026 ± 0.002 | >0.300 | 0.021 ± 0.001 | 0.081 ± 0.003 | 0.020 ± 0.003 | 0.041 ± 0.004 |
| 4 | [64Cu]Cu-NOTA-c(RGDfK) | 0.025 ± 0.007 | >0.300 | 0.032 ± 0.009 | >0.200 | 0.045 ± 0.002 | 0.118 ± 0.008 |
| 5 | Cisplatin | 90 ± 8 | 35 ± 4 | 25 ± 6 | 15 ± 7 | 20 ± 5 | 15 ± 5 |
| 6 | Temozolomide | 800 ± 11 | 80 ± 10 | 580 ± 8 | 76 ± 7 | 575 ± 6 | 72 ± 5 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mirzaei, A.; Ait-Mohand, S.; Ignatius Arokia Doss, P.M.; Rousseau, É.; Guérin, B. A Targeted Radiotheranostic Agent for Glioblastoma: [64Cu]Cu-NOTA-TP-c(RGDfK). Brain Sci. 2025, 15, 844. https://doi.org/10.3390/brainsci15080844
Mirzaei A, Ait-Mohand S, Ignatius Arokia Doss PM, Rousseau É, Guérin B. A Targeted Radiotheranostic Agent for Glioblastoma: [64Cu]Cu-NOTA-TP-c(RGDfK). Brain Sciences. 2025; 15(8):844. https://doi.org/10.3390/brainsci15080844
Chicago/Turabian StyleMirzaei, Alireza, Samia Ait-Mohand, Prenitha Mercy Ignatius Arokia Doss, Étienne Rousseau, and Brigitte Guérin. 2025. "A Targeted Radiotheranostic Agent for Glioblastoma: [64Cu]Cu-NOTA-TP-c(RGDfK)" Brain Sciences 15, no. 8: 844. https://doi.org/10.3390/brainsci15080844
APA StyleMirzaei, A., Ait-Mohand, S., Ignatius Arokia Doss, P. M., Rousseau, É., & Guérin, B. (2025). A Targeted Radiotheranostic Agent for Glioblastoma: [64Cu]Cu-NOTA-TP-c(RGDfK). Brain Sciences, 15(8), 844. https://doi.org/10.3390/brainsci15080844








